1. Introduction
Cotton, a member of the Malvaceae family, stands as one of the most economically significant crops globally, providing the primary source of natural fiber for the textile industry and contributing substantially to the production of edible oil and animal feed [
1,
2,
3]. Originating from tropical and subtropical regions, cultivated cotton species, particularly upland cotton (
Gossypium hirsutum L.), are inherently sensitive to a range of abiotic stresses, which pose a significant threat to their growth, development, and productivity [
4,
5]. Among these environmental challenges, low-temperature or cold stress is a major limiting factor, especially during the critical stages of seed germination and seedling establishment [
6]. Exposure to suboptimal temperatures can trigger a cascade of detrimental physiological and biochemical responses, leading to stunted growth, reduced biomass accumulation, impaired reproductive development, and ultimately, significant losses in both fiber yield and quality [
7,
8].
In many cotton-producing regions, such as Xinjiang in China, the planting season in early spring is often marked by unpredictable cold spells, colloquially known as “late spring frosts”. Exposure to chilling temperatures (below 15 °C) during seed germination and seedling emergence can lead to a cascade of detrimental physiological and developmental effects. One of the primary targets is the photosynthetic apparatus. Low temperatures can lead to a decrease in the efficiency of photosystem II (PSII), disrupt electron transport, and inhibit the activity of key carbon fixation enzymes like RuBisCO, ultimately reducing the net photosynthetic rate [
9]. This inhibition is often exacerbated by the overproduction of reactive oxygen species (ROS), such as superoxide radicals and hydrogen peroxide, which cause oxidative damage to cellular components including lipids, proteins, and nucleic acids [
10]. Furthermore, cold stress alters the physical properties of cellular membranes by decreasing their fluidity, which impairs the function of membrane-bound proteins and transport systems, disrupting cellular homeostasis and signaling [
11]. These cellular-level disturbances manifest as macroscopic anatomical changes, including alterations in the thickness of leaf tissues like the palisade and spongy mesophyll, and damage to vascular structures, which further compromise plant function and survival.
To counteract the deleterious effects of cold stress, plants have evolved a sophisticated and integrated network of defense mechanisms. These adaptive responses span from anatomical modifications to complex biochemical and molecular reprogramming [
12]. At the molecular level, the perception of a cold signal initiates a signaling cascade that leads to a massive reconfiguration of the transcriptome. This transcriptional reprogramming is central to cold acclimation and involves the differential expression of thousands of genes [
13,
14]. These genes encode a wide array of functional proteins, including transcription factors that act as master regulators of downstream stress-responsive genes, enzymes involved in the biosynthesis of protective compounds, and proteins that directly protect cellular structures. Recent studies have highlighted the complexity of this regulation, showing that it extends beyond simple transcription to include post-transcriptional modifications like alternative splicing [
15] and epigenetic mechanisms such as histone modifications, which add further layers of control to the plant stress response [
16].
Different cotton varieties exhibit considerable differences in their ability to withstand low temperatures, suggesting a genetic basis for this differential tolerance [
17]. Identifying the genes and pathways that contribute to the superior performance of cold-tolerant cultivars is a critical step toward developing more resilient cotton varieties through molecular breeding. Comparative transcriptomics, which contrasts the gene expression profiles of tolerant and sensitive genotypes under stress, is a powerful strategy for pinpointing these key genetic determinants. Previous studies in cotton have identified candidate genes involved in cold response, such as those related to lipid metabolism [
18], transcription factors [
19], and hormone signaling [
20]. However, a comprehensive understanding requires integrating this molecular data with physiological and anatomical evidence to build a holistic picture of the adaptive strategies employed by tolerant varieties. For instance, the link between the transcriptional activation of genes involved in secondary metabolism or structural development and the resulting phenotypic traits, such as altered cell wall composition or the density of protective structures like glandular trichomes, needs to be more firmly established.
In this study, we aimed to dissect the molecular and anatomical basis of differential cold tolerance in cotton. We selected two upland cotton varieties, the cold-tolerant ‘Xinluzhong 61’ (C61) and the cold-sensitive ‘Tahe 2’ (C2), which are widely cultivated in the Xinjiang region of China, an area prone to early-season cold spells. By performing a time-course comparative transcriptomic analysis under severe cold stress (0 °C), we sought to identify the key genes, biological processes, and metabolic pathways that are differentially regulated between these two varieties. We hypothesized that the superior cold tolerance of C61 is underpinned by a more rapid and robust activation of specific defense pathways, including antioxidant systems, secondary metabolite biosynthesis, and pathways related to the reinforcement of physical barriers. Furthermore, we investigated the anatomical changes in the cotyledons, true leaves, and stems of both varieties in response to cold stress to correlate the observed transcriptional changes with tangible structural adaptations. By integrating these multi-level analyses, this research aims to provide a comprehensive understanding of the adaptive mechanisms contributing to cold tolerance in cotton, offering valuable insights and potential genetic targets for the future improvement of this vital crop.
2. Materials and methods
2.1. Plant Materials and Growth Conditions
Two upland cotton (Gossypium hirsutum L.) cultivars, Tahe 2(cold-sensitive) and Xinluzhong 61(cold-tolerant), were used in this study. Seeds were surface-sterilized with 1% sodium hypochlorite for 10 minutes, rinsed thoroughly with sterile distilled water, and germinated on moist filter paper in the dark at 28 °C for 48 hours. Uniformly germinated seeds were then transferred to plastic pots (10 cm diameter) filled with a sterile mixture of vermiculite and nutrient soil (1:1, v/v). The seedlings were grown in a controlled environment growth chamber under a 16-hour light (28 °C)/8-hour dark (22 °C) cycle, with a light intensity of 300 µmol m-2s-1 and 60% relative humidity.
2.2. Cold Stress Treatment and Sample Collection
When the seedlings reached the three-leaf stage (approximately 21 days after sowing), they were divided into two groups. The control group was maintained under the normal growth conditions described above (28 °C). The cold stress group was transferred to another growth chamber set to a constant temperature of 0 °C for 12 and 24 hours, with the same light and humidity conditions. For each cultivar and treatment, three biological replicates were established, with each replicate consisting of a pool of five individual seedlings. After the 12 and 24-hour treatment period, the second true leaves and stem segments (1 cm below the cotyledons) were collected from both control and cold-stressed seedlings. A portion of the collected tissue was immediately frozen in liquid nitrogen and stored at -80 °C for RNA extraction. The remaining portion was fixed for anatomical analysis.
2.3. RNA Extraction, Library Construction, and Sequencing
Total RNA was extracted from the frozen leaf samples using the TRIzol reagent (Invitrogen, USA) according to the manufacturer’s protocol. RNA quality and quantity were assessed using a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific, USA) and an Agilent 2100 Bioanalyzer (Agilent Technologies, USA). Only RNA samples with an RNA Integrity Number (RIN) > 8.0 were used for library construction. For each sample, 1 µg of total RNA was used to construct a sequencing library using the NEBNext®® Ultra™ RNA Library Prep Kit for Illumina®® (NEB, USA) following the manufacturer’s instructions. Briefly, mRNA was enriched using oligo(dT)-attached magnetic beads. The enriched mRNA was then fragmented and used as a template for first-strand cDNA synthesis using random hexamer primers. Second-strand cDNA was synthesized using DNA Polymerase I and RNase H. The resulting double-stranded cDNA was end-repaired, A-tailed, and ligated to sequencing adapters. The ligated products were purified and amplified by PCR to create the final cDNA libraries. The quality of the libraries was assessed on the Agilent Bioanalyzer 2100 system. The qualified libraries were sequenced on an Illumina NovaSeq 6000 platform, generating 150 bp paired-end reads.
2.4. Functional Annotation and Enrichment Analysis
Raw sequencing reads were first processed to remove adapter sequences, low-quality reads, and reads containing poly-N using Trimmomatic. The quality of the clean reads was verified using FastQC. The clean reads were then aligned to the Gossypium hirsutum L. TM-1 reference genome (v2.1) using HISAT2 with default parameters. The number of reads mapped to each gene was counted using featureCounts. Differential gene expression analysis was performed using the DESeq2 R package (Rapaport, 2013). Genes with an adjusted p-value (padj) < 0.05 and an absolute log2(Fold Change) > 1 were identified as DEGs. This analysis was performed for each cultivar by comparing the cold-stressed samples to their respective controls.
To understand the biological functions of the identified DEGs, Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were performed using the clusterProfiler R package. GO terms and KEGG pathways with a padj < 0.05 were considered significantly enriched.
2.5. Quantitative Real-Time PCR (qRT-PCR) Validation
To validate the RNA-seq results, the expression levels of ten selected DEGs were quantified by qRT-PCR. First-strand cDNA was synthesized from 1µg of the same total RNA samples used for sequencing, using the PrimeScript™ RT Reagent Kit with gDNA Eraser (Takara, Japan). Gene-specific primers were designed using Primer3Plus. The cotton GhUBQ7 gene was used as the internal reference for normalization. The qRT-PCR reactions were performed on a CFX96 Real-Time PCR Detection System (Bio-Rad, USA) using TB Green®® Premix Ex Taq™ II (Tli RNaseH Plus) (Takara, Japan). The reaction volume was 20µL, containing 10µL of TB Green Premix, 0.8µL of each primer (10µM), 2µL of diluted cDNA, and 6.4µL of nuclease-free water. The thermal cycling conditions were: 95 °C for 30s, followed by 40 cycles of 95 °C for 5s and 60 °C for 30s. The relative expression levels were calculated using the 2-ΔΔCt method. Three technical replicates were performed for each of the three biological replicates.
2.6. Paraffin Section Preparation
Stem segments and leaf samples were fixed in FAA solution (formalin:acetic acid:70% ethanol, 5:5:90, v/v/v) for at least 24 hours at 4 °C. The fixed samples were then dehydrated through a graded ethanol series (70%, 85%, 95%, 100%), cleared with xylene, and embedded in paraffin. Transverse sections of 8 µm thickness were prepared using a rotary microtome (Leica RM2235, Germany). The sections were mounted on glass slides, deparaffinized with xylene, rehydrated through a reverse ethanol series, and stained with Safranin-Fast Green. The stained sections were observed and photographed using a light microscope (Olympus BX53, Japan) equipped with a digital camera. For quantitative analysis, ImageJ software was used to measure various anatomical parameters from at least 10 different sections per biological replicate. The measured parameters included leaf thickness, palisade tissue thickness, spongy tissue thickness, stem diameter, cortex thickness, xylem thickness, and pith diameter.
2.7. Statistical Analysis
All statistical analyses for anatomical and qRT-PCR data were performed using SPSS software (version 22.0). Data are presented as the mean±standard deviation (SD). Significant differences between control and cold-stressed groups were determined using an independent samples t-test, with p < 0.05 considered statistically significant.
3. Results
3.1. Effects of Low-Temperature Stress on Cotton Seedlings of Xinluzhong 61 and Tahe 2
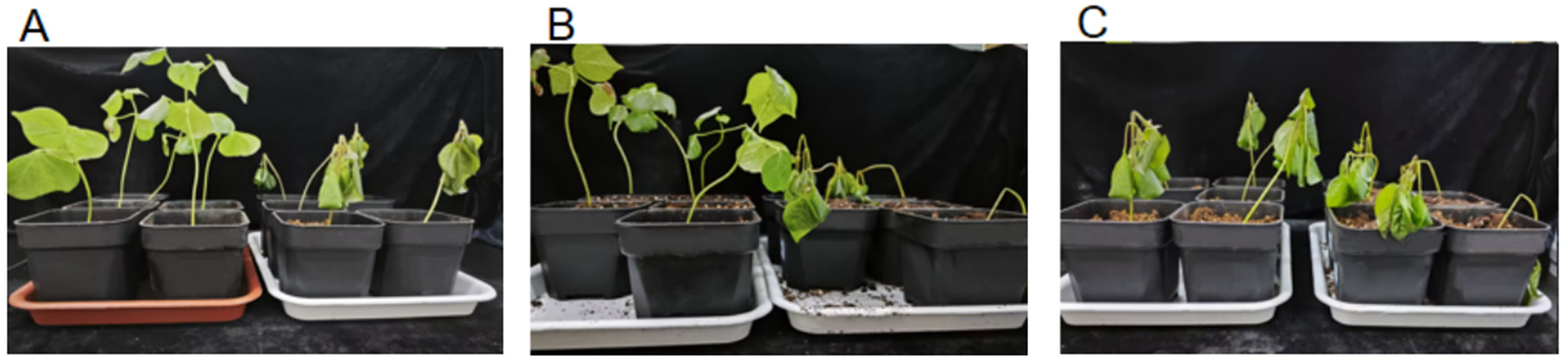
To clarify the response differences of different cotton varieties to low-temperature stress, this study selected the main cultivated varieties in southern Xinjiang, Xinluzhong 61 and Tahe 2 as materials. During the seedling stage, the plants were subjected to 0 °C low-temperature treatment to observe their phenotypic changes. As shown in
Figure 1, after 0 °C low-temperature treatment, both varieties displayed significant responses to low-temperature stress, but the extent varied noticeably:
For Xinluzhong 61 under low-temperature treatment, the plants remained relatively upright overall, with only some leaves showing slight wilting. The supportive structures of the main stem and lateral branches were relatively intact, and the stress symptoms were relatively mild. In contrast, Tahe 2 under low-temperature treatment exhibited severe lodging, large areas of leaf wilting and drooping, some leaves showing water-soaked lesions, and a significant decrease in stem toughness, indicating more severe stress symptoms (
Figure 1B). In a direct comparison under the same low-temperature treatment condition (
Figure 1C), the plant morphology and leaf integrity of Xinluzhong 61 were significantly better than those of Tahe 2.
Overall phenotypic observation results indicate that low-temperature stress significantly inhibits normal growth of cotton seedlings and that different varieties show significant differences in sensitivity to low temperature. Among them, while Xinluzhong 61 exhibits stronger adaptability and tolerance under low-temperature stress, while Tahe 2 is more sensitive to low temperature.
3.2. GO Enrichment Analysis of DEGs Induced by Low-Temperature Stress in Two Cotton Varieties
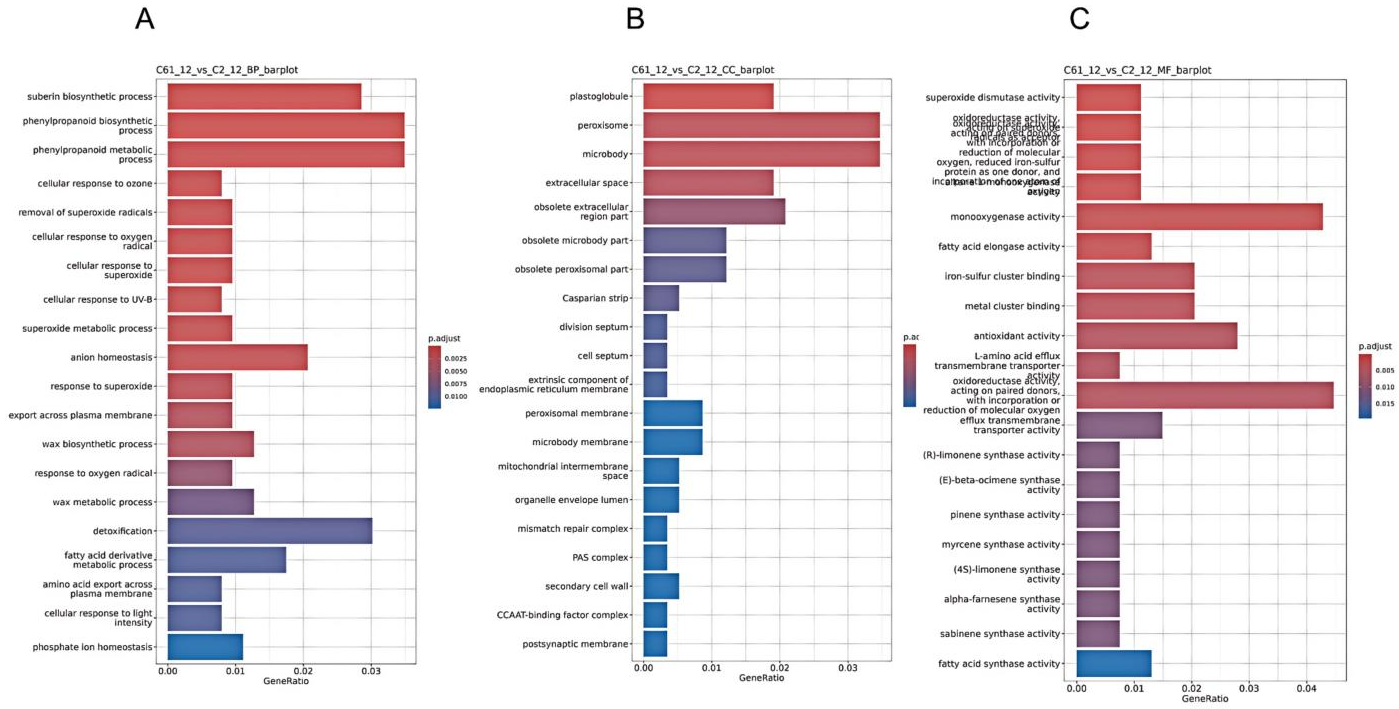
To dissect the molecular mechanisms underlying the differential low-temperature responses of two cotton varieties (Xinluzhong 61, C61, and Tahe 2, C2), we performed Gene Ontology (GO) enrichment analysis on differentially expressed genes (DEGs) under 0 °C treatment for 12 h and 24 h.The DEGs were categorized into three primary functional categories: biological process (BP), cellular component (CC), and molecular function (MF).
At 12 h, terms related to phenylpropanoid metabolism, ROS response, and cell wall biosynthesis were significantly enriched in biological process (BP), indicating enhanced stress tolerance and metabolic reprogramming in C61. Detoxification and ion homeostasis were also enriched, reflecting differences in cellular homeostasis and secondary metabolite accumulation(
Figure 2A). In cellular component (CC), DEGs were predominantly enriched in peroxisome, microbody, and plastoglobule, with the highest significance, along with extracellular space and membrane components, suggesting differences in subcellular compartmentalization(
Figure 2B). In molecular function (MF), oxidoreductase, monooxygenase, superoxide dismutase, and antioxidant activities were most prominently enriched, supporting enhanced ROS scavenging in C61, while terpene and fatty acid synthase activities indicated divergent secondary metabolism regulation(
Figure 2C).
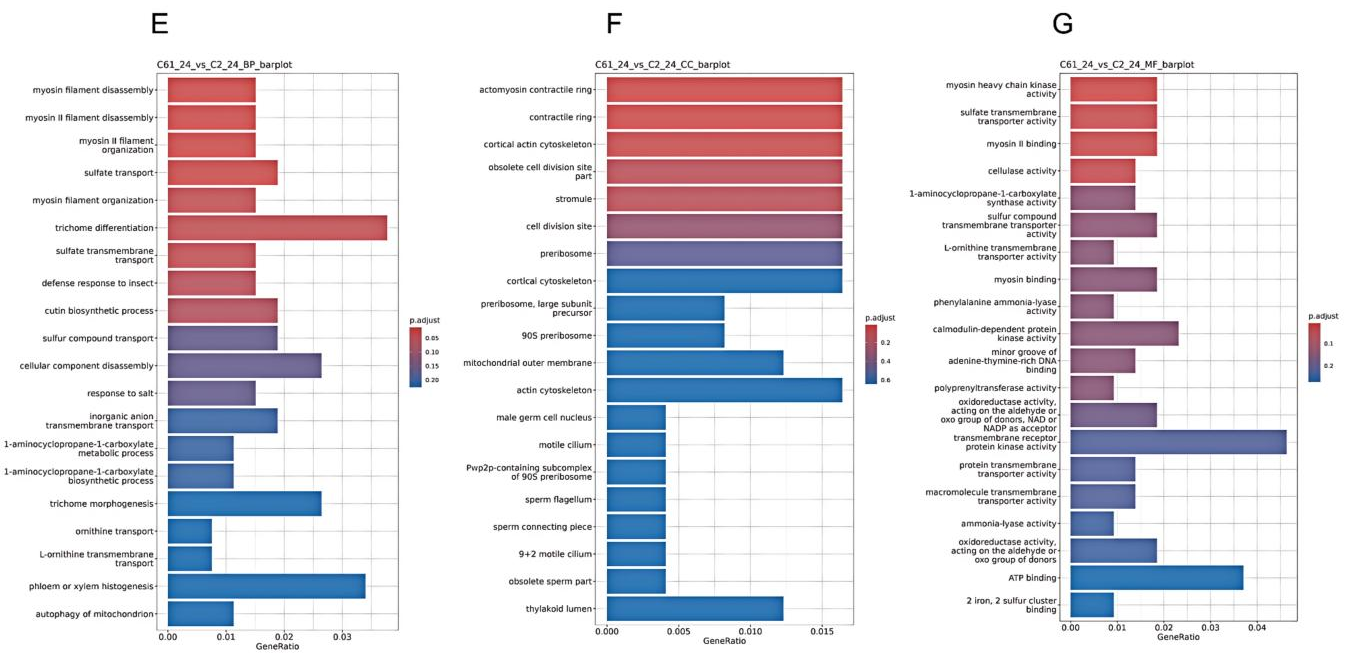
At 24 h, BP terms were mainly associated with trichome differentiation (with the highest gene ratio), phloem/xylem histogenesis, myosin filament dynamics, sulfur transport, and defense response to insects. CC terms focused on the actin cytoskeleton and contractile apparatus, as well as ribosome-related components, implying impacts on protein synthesis. MF terms involved motor activity, transmembrane transport, and catalytic activity, while receptor kinase activity and ATP binding were negatively enriched, suggesting downregulation under prolonged cold stress(
Figure 3).
3.3. KEGG Enrichment Analysis of DEGs Induced by Low-Temperature Stress in Two Cotton Varieties
To elucidate the key metabolic pathways underlying the differential cold tolerance of two cotton varieties (Xinluzhong 61 and Tahe 2), KEGG orthology mapping was conducted on the differentially expressed genes (DEGs) under 0 °C treatment (FDR-adjusted P < 0.05).
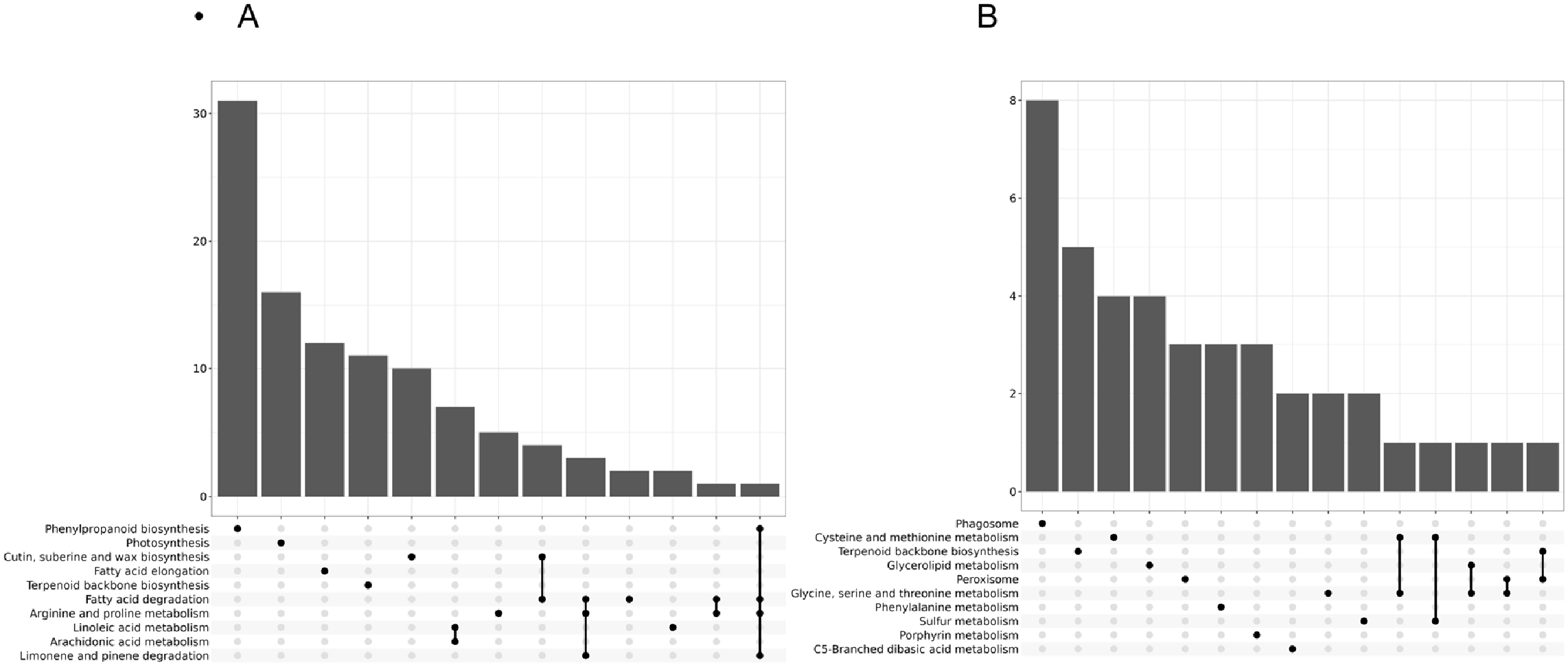
KEGG pathway analysis of DEGs between the two cotton varieties under 12 h of low—temperature stress indicated that the most significantly enriched pathways were phenylpropanoid biosynthesis, photosynthesis, and cutin, suberine, and wax biosynthesis. Moreover, pathways associated with fatty acid metabolism, terpenoid backbone biosynthesis, and linoleic acid metabolism were also enriched, suggesting that C61 enhances cold tolerance by strengthening cell wall barriers, optimizing energy metabolism, and activating secondary metabolic pathways.
In the comparison of the two varieties under 24 h of low—temperature stress, DEGs were significantly enriched, with the most prominent enrichment observed in the Phagosome pathway, followed by Cysteine and methionine metabolism, Terpenoid backbone biosynthesis, and Glycerolipid metabolism (
Figure 4). In contrast to the 12 h treatment, which emphasized phenylpropanoid biosynthesis and photosynthesis, the 24 h treatment shifted towards immune activation and antioxidant metabolism, reflecting a time—dependent adaptive strategy for maintaining cold tolerance.
3.4. Effects of Low Temperature Treatment on the Glandular Trichomes of Two Cotton Varieties
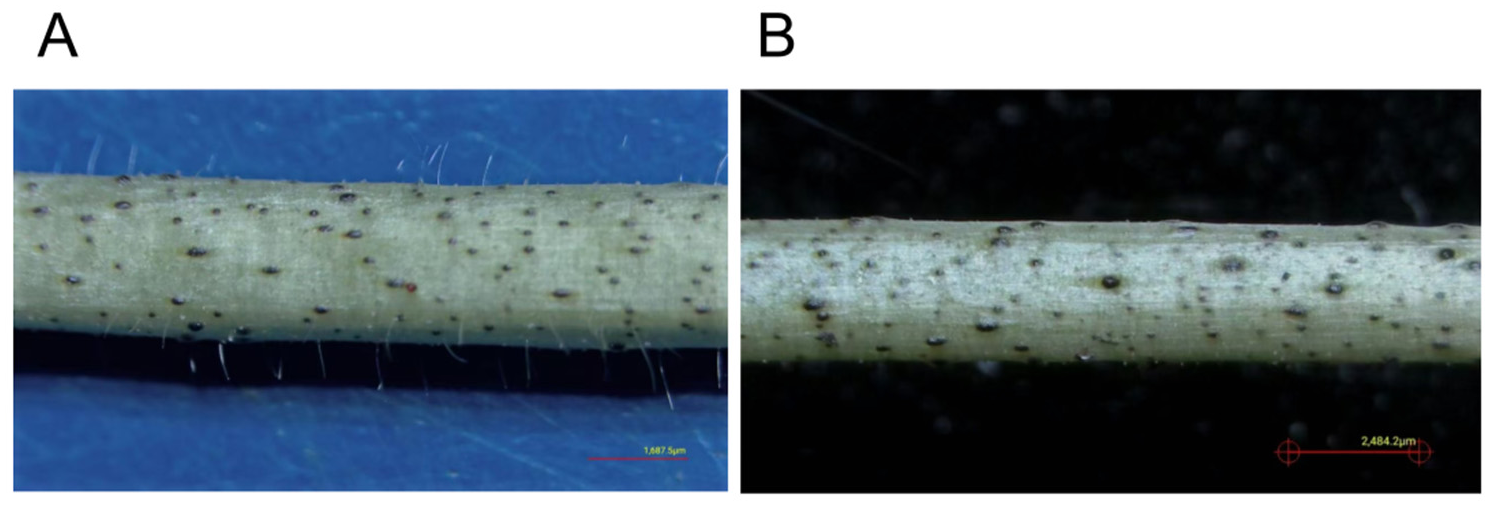
In the biological process category, we noticed that trichome differentiation emerged as the most highly enriched term with the largest gene ratio, indicating that this developmental process was transcriptionally activated in C61 in response to prolonged cold stress. Phenotypic observation of stem tissues revealed striking differences in glandular trichome development between the two cotton varieties under low-temperature stress. As shown in
Figure 5, Xinluzhong 61 (C61) exhibited dense and prominent glandular trichomes distributed across the stem surface, whereas Tahe 2 (C2) displayed a relatively smooth stem with significantly fewer trichomes (Fig. B). This morphological divergence was strongly supported by our transcriptomic GO enrichment analysis, particularly the results from the 24 h low-temperature treatment.
Collectively, the abundant glandular trichomes in C61 represent a key adaptive phenotype shaped by transcriptional reprogramming. This structural modification, coupled with enhanced secondary metabolism, forms a comprehensive physical and chemical defense system, which likely contributes significantly to the superior low-temperature tolerance of C61 compared to C2.
3.5. Cotyledon Anatomical Structure Response Under Cold Stress
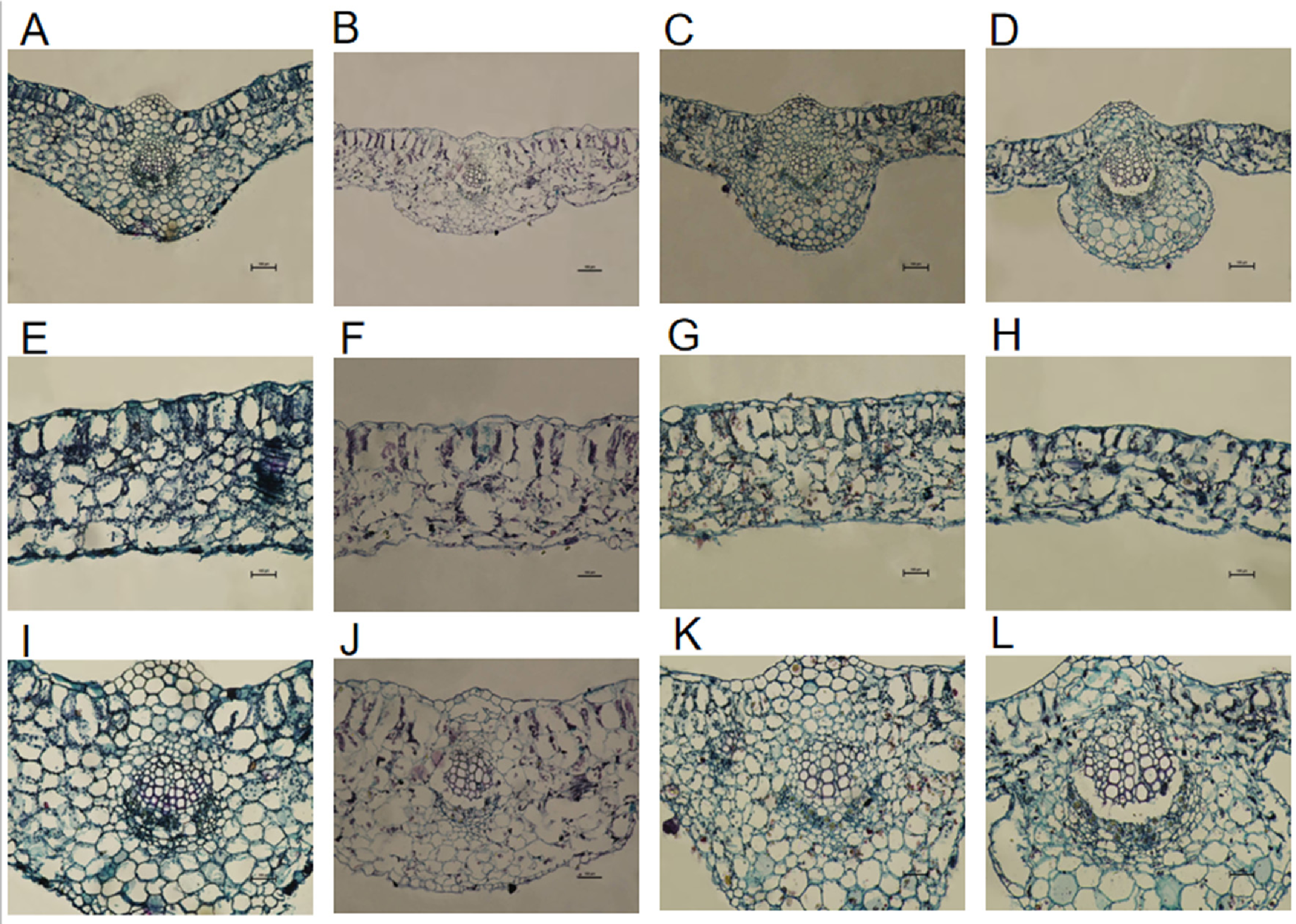
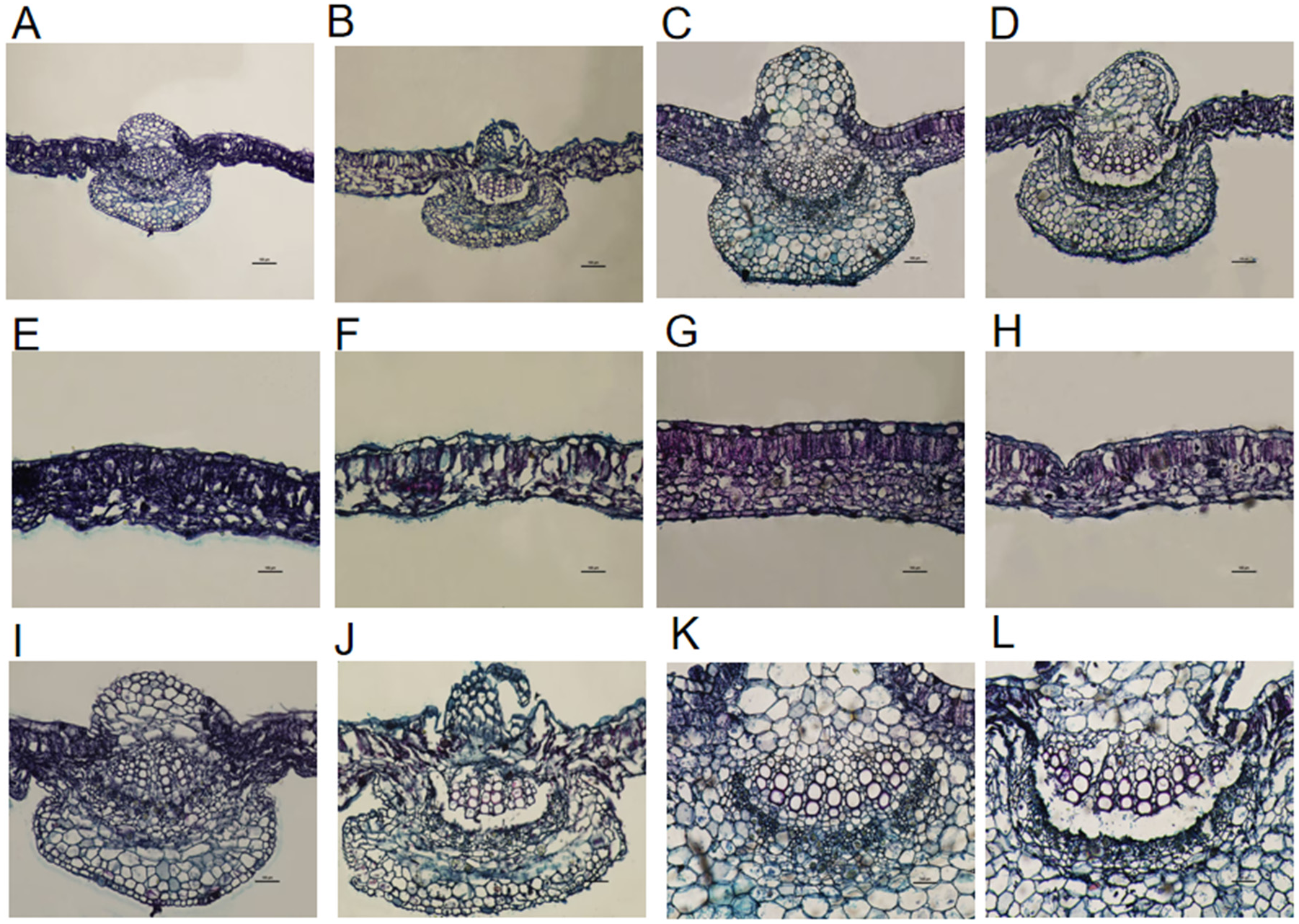
Under the effect of low-temperature stress, the thickness of the palisade and spongy tissues in leaves undergoes significant changes, which are key anatomical indicators reflecting the impact of low temperature. As shown in
Table 1 and
Figure 6, low-temperature stress significantly reduces the leaf thickness, palisade tissue thickness, and spongy tissue thickness of cotton cotyledons, and causes rupture in the midrib vascular bundles. Under low-temperature conditions of 0 °C for 12 hours, in Xinluzhong 61 leaves, the thickness of palisade tissue, spongy tissue, leaf thickness, palisade-to-spongy ratio, and leaf structural density decreased by 32.14%, 16.15%, 15.55%, 19.15%, and 19.35% respectively compared with the control (CK). For Tahe 2 leaves, the thickness of palisade tissue, spongy tissue, and leaf thickness decreased by 22.36%, 35.21%, and 28.78% respectively, while the palisade-to-spongy ratio and leaf structural density increased by 18.92% and 12%, with differences being statistically significant (P < 0.05).
Comparing these two cotton varieties, the cotyledon leaf palisade tissue thickness, spongy tissue thickness, leaf thickness, the palisade-spongy ratio, and leaf structural compactness of Xinluzhong 61 are all higher than those of Tahe 2. This indicates that Xinluzhong 61 has stronger photosynthetic capacity and a slower rate of water loss. In the case of low-temperature treatment, the cotyledon leaf palisade tissue thickness, spongy tissue thickness, and leaf thickness of Xinluzhong 61 are also higher than those of Tahe 2, which may be related to the stronger cold tolerance of this variety.
3.6. Response of True Leaf Anatomical Structure Under Cold Stress
Similar to the cotyledon situation, the effect of low-temperature stress on true leaves is also have relationship with the thickness of the palisade and spongy tissues. As shown in
Table 2 and
Figure 7, under a low-temperature treatment, the palisade tissue thickness, spongy tissue thickness, and leaf thickness of the Xinluzhong 61 leaf decreased by 20.11%, 42.82%, and 34.47% respectively compared to the control (CK), while the palisade-spongy ratio increased by 40%. For Taha 2 leaves, the palisade tissue thickness, spongy tissue thickness, and leaf thickness also slight decreased by 14.73%, 20.58%, and 16.86% respectively, but the palisade-spongy ratio increased by only 7.02%.
The palisade-spongy ratio of plant leaves usually positively correlated with plant stress resistance. As seen from
Table 2, the palisade tissue thickness, spongy tissue thickness, and leaf thickness of Xinluzhong 61 true leaves are all higher than those of Taha 2 under normal condition, while there is no significant difference in the palisade-spongy ratio (P > 0.05) and leaf structural density between them. After low-temperature treatment, the leaf thickness and palisade-spongy ratio of Xinluzhong 61 leaves are significantly higher than those of Taha 2, indicating that Xinluzhong 61 has stronger cold resistance than Taha 2.
Overall, low-temperature stress caused more severe damage to the anatomical structure of Taha 2true leaves. This is consistent with the more sensitive low-temperature response observed in phenotypic observations, further revealing the differences in cold tolerance between the two varieties at the anatomical level.
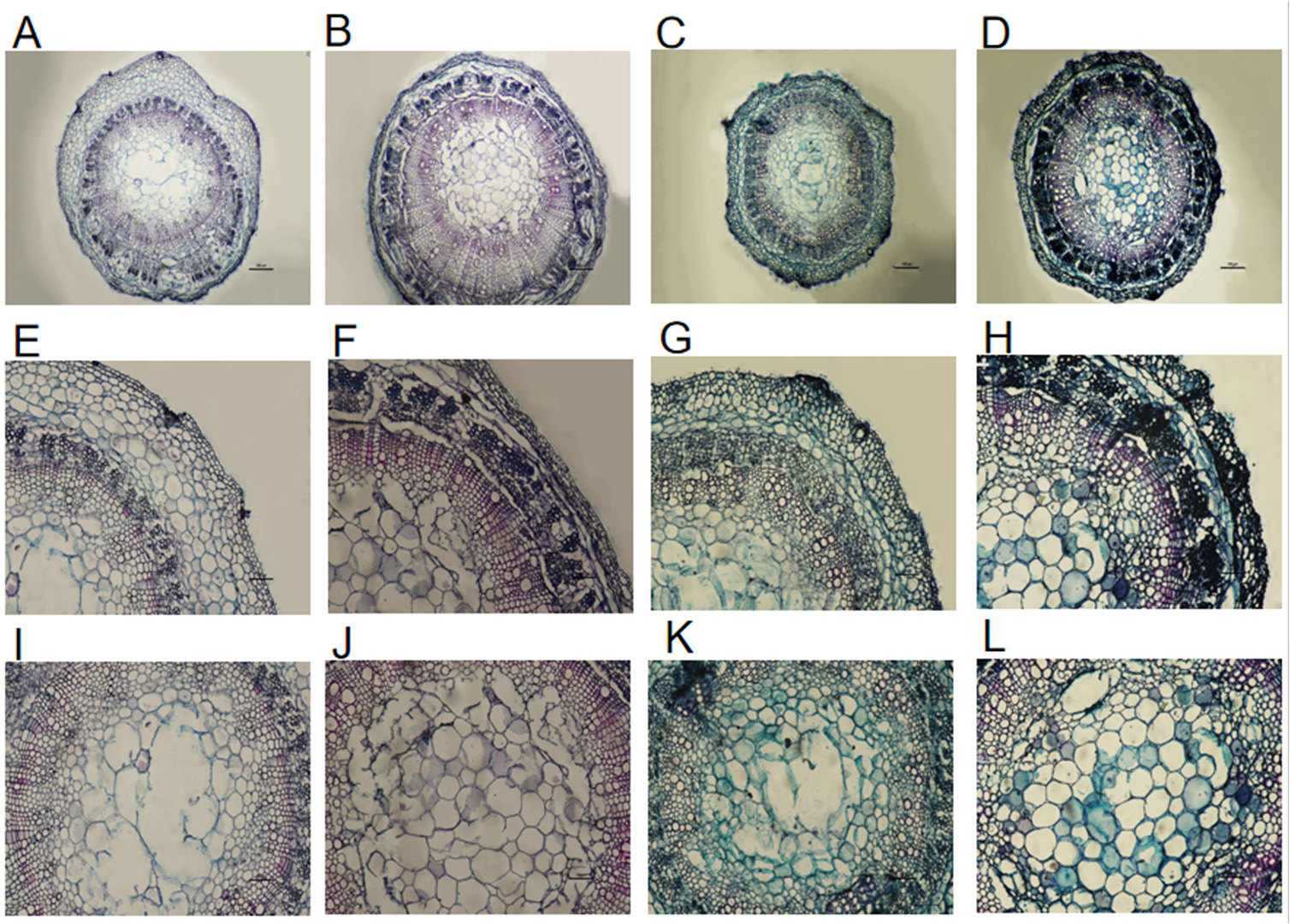
3.7. Response of Stem Anatomical Structure
Through the analysis of the data in
Table 3 and the microstructure in
Figure 8, it is evident that after 24 hours of 0 °C low-temperature stress treatment, the anatomical structure of cotton stems underwent significant changes.The cortex thickness decreased, while the thicknesses of the epidermis, phloem, xylem, and the diameter of the pith increased, and fractures appeared between the xylem, phloem, and cortex. Specifically, under low-temperature treatment, the cortex thickness of Xinluzhong 61 decreased by only 4.23% compared with the control (CK), and the thicknesses of the epidermis, phloem, xylem, and pith diameter increased by 14.97%, 21.14%, 35.98%, and 8.96%, respectively. For Tahe 2, the cortex thickness decreased by 47.17%, the phloem thickness remained basically stable, and the xylem thickness and pith diameter increased by 19.77% and 12.56%, respectively.
Xylem and phloem of plants are responsible for transporting water and inorganic salts, respectively. Therefore, plants usually resist adversity by expanding their xylem and phloem. A comparative analysis of the two varieties shows that the epidermis thickness, cortex thickness, and xylem thickness of Xinluzhong 61 are smaller than those of Tahe 2, while the phloem thickness and pith diameter are greater. After low-temperature treatment, the xylem thickness of Xinluzhong 61 increased greater than that of Tahe 2, indicating more tolerance to cold temperature.
4. Discussion
Low-temperature stress is a major abiotic factor that severely limits the productivity, quality, and geographical distribution of cotton, a crop of tropical and subtropical origin [
21]. Understanding the intricate molecular and physiological mechanisms that underpin cold tolerance is crucial for developing resilient cotton varieties. This study provides a comprehensive, multi-level analysis by integrating transcriptomic, anatomical, and morphological data to dissect the differential cold stress responses of two cotton varieties, the tolerant Xinluzhong 61 (C61) and the sensitive Tahe 2 (C2). Our findings reveal that the superior cold tolerance of C61 is not attributable to a single mechanism but rather a coordinated, time-dependent strategy involving rapid metabolic reprogramming for defense and subsequent structural adaptations for long-term protection.
The initial response to cold stress is critical for plant survival, as it sets the stage for subsequent acclimation [
22]. Our transcriptomic analysis at 12 hours post-stress revealed a robust and swift activation of defense-related pathways in the tolerant C61 variety. The significant enrichment of Gene Ontology (GO) terms and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways associated with phenylpropanoid biosynthesis, reactive oxygen species (ROS) response, and cell wall modifications underscores a proactive defense strategy [
23,
24].The phenylpropanoid pathway is a cornerstone of plant defense, producing a vast array of secondary metabolites including lignin, flavonoids, and suberin [
25]. The upregulation of this pathway in C61 likely contributes to the reinforcement of cell walls, creating a physical barrier that can limit cellular dehydration and the propagation of ice crystals. Concurrently, the enrichment of pathways for cutin, suberin, and wax biosynthesis further strengthens this physical barrier, reducing non-stomatal water loss and enhancing cellular integrity. This metabolic fortification is complemented by a highly active antioxidant system. The enrichment of GO terms such as oxidoreductase, monooxygenase, and superoxide dismutase activity in C61 points to an enhanced capacity to scavenge ROS, which are inevitably produced under cold stress and can cause significant oxidative damage [
26,
27,
28]. Furthermore, the observed enrichment in fatty acid and linoleic acid metabolism pathways is consistent with previous findings that lipid remodeling is a key component of the cold response in cotton, affecting membrane fluidity and generating signaling molecules like oxylipins [
29]. This rapid and multi-pronged metabolic response in C61 likely mitigates the initial shock of cold stress, preserving cellular homeostasis and providing a foundation for long-term survival.
As the duration of cold stress extended to 24 hours, we observed a significant shift in the transcriptomic landscape of C61, moving from immediate metabolic defense to developmental and structural adaptations. The most striking finding from our GO analysis was the dramatic enrichment of the “trichome differentiation” biological process, which was accompanied by a visible increase in the density of glandular trichomes on the stems of C61. Plant trichomes serve multiple functions in stress tolerance; they can create a microenvironment that insulates the plant surface, reduce water loss by trapping a layer of still air, and, in the case of glandular trichomes, secrete secondary metabolites that can act as chemical protectants [
30,
31,
32]. The development of a dense trichome layer in C61 represents a significant morphological adaptation, forming a physical and chemical barrier against the cold. This finding highlights a less-explored mechanism of cold tolerance in cotton and suggests that developmental plasticity plays a crucial role in long-term acclimation.
The transcriptomic and morphological adaptations observed in C61 were strongly correlated with the maintenance of anatomical integrity in its leaves and stems. Under cold stress, C61 retained significantly greater leaf thickness, as well as thicker palisade and spongy tissues, compared to the sensitive C2 variety [
33]. The palisade tissue is the primary site of photosynthesis, and its structural integrity is vital for maintaining energy production under stress [
34]. The thicker tissues and higher palisade-to-spongy ratio in C61 suggest a superior ability to maintain photosynthetic capacity and regulate water loss, which are critical for survival when cold temperatures impair water uptake from the soil. These anatomical features are likely the result of the observed transcriptional programs that reinforce cell walls and maintain cellular homeostasis.
Similarly, the stem anatomy of C61 showed adaptive changes, including a greater xylem thickness after cold treatment compared to the sensitive variety. A robust xylem is essential for maintaining water transport, which can be compromised by low temperatures due to increased water viscosity and the risk of embolism [
35]. The ability of C61 to preserve and even reinforce its vascular structures underpins its physiological resilience. In contrast, the more severe degradation of leaf and stem tissues in C2 reflects its inability to mount an effective and sustained defense, leading to cellular collapse and functional impairment. This integration of molecular data with anatomical evidence provides a holistic view of cold tolerance, where gene expression networks translate into tangible structural advantages that confer resilience [
36].
Significance, Limitations, and Future Directions
This study significantly advances our understanding of cold tolerance mechanisms in cotton by demonstrating a coordinated, multi-level response in the tolerant variety C61. We have identified not only key metabolic pathways but also a critical role for developmental plasticity, particularly trichome formation, as a key adaptive trait. The differentially expressed genes and pathways uncovered here, such as those controlling phenylpropanoid biosynthesis and trichome differentiation, represent valuable targets for molecular breeding programs aimed at enhancing cold tolerance in elite cotton cultivars [
37,
38].
However, this study has certain limitations. The findings are based on a comparison of two specific cultivars at discrete time points and a single stress temperature. Future research should expand this analysis to a broader range of cotton germplasm and environmental conditions to validate these findings and uncover additional tolerance mechanisms. While our transcriptomic analysis provides a powerful snapshot of gene expression, it is correlational. Functional validation of key candidate genes using techniques such as virus-induced gene silencing (VIGS) or CRISPR/Cas9 is necessary to establish their causal roles in cold tolerance [
39]. Furthermore, integrating proteomics and metabolomics would provide a more complete picture of the molecular landscape, revealing post-transcriptional regulation and the specific biochemical compounds that contribute to protection [
40,
41,
42,
43]. Investigating the specific chemical composition of the glandular trichome exudates in C61 could also unveil novel protective compounds.
In conclusion, our integrated analysis reveals that the superior cold tolerance of the cotton variety Xinluzhong 61 is orchestrated by a sophisticated and dynamic defense system. This system combines a rapid metabolic response, characterized by the activation of antioxidant and cell wall fortification pathways, with a subsequent phase of morphological adaptation, most notably the development of a dense protective layer of glandular trichomes. These molecular and developmental changes are reflected in the preservation of anatomical integrity, which underpins the plant’s physiological resilience. These findings provide valuable insights into the complex nature of plant stress adaptation and offer promising avenues for the genetic improvement of cold tolerance in cotton.
5. Conclusions
The ability of plants to survive low temperatures depends on a complex interplay of physiological, anatomical, and molecular adaptations. In this study, we combined anatomical and transcriptomic approaches to dissect the cold stress response in seedlings of two cotton cultivars. Our findings reveal that cotton employs a multifaceted strategy, involving significant structural remodeling of tissues and a comprehensive reprogramming of gene expression, to cope with cold.
Author Contributions
H.M.W. and Y.H.H. contributed equally to this work. They performed the experiments, analyzed the data and drafted the manuscript. F.J.W. and M.M.J conceived and designed the experiments., Z.Y.Y., Z.J.W. and F.L.W. revised the manuscript. All authors read and approved the final manuscript.
Funding
This work was supported by ‘Tianchi Talents’ of XPCC(230000339), Guided Science and Technology Program Project of XPCC (2023ZD108), the President’s Fund of Tarim University (TDZKBS202302) and National College Student Innovation and Entrepreneurship Training Program(202610757018).
References
- Jan M, Liu Z, Guo C, Sun X. Molecular Regulation of Cotton Fiber Development: A Review. Int J Mol Sci. 2022;23(9):5004. [CrossRef]
- Lin Z, Wang Z, Zhang Y, Tan S, Masangano M, Kang M, Cao X, Huang P, Gao Y, Pei X, Ren X, He K, Liang Y, Ji G, Tian Z, Wang X, Ma X. Gene expression modules during the emergence stage of upland cotton under low-temperature stress and identification of the GhSPX9 cold-tolerance gene. Plant Physiol Biochem. 2025;218:109320. [CrossRef]
- Qamer Z, Chaudhary M, Du X, Hinze L, Azhar M. Review of oxidative stress and antioxidative defense mechanisms in Gossypium hirsutum L. in response to extreme abiotic conditions. J Cotton Res. 2021;4(1). [CrossRef]
- Li Y, Zhu J, Xu J, Zhang X, Xie Z, Li Z. Effect of cold stress on photosynthetic physiological characteristics and molecular mechanism analysis in cold-resistant cotton (ZM36) seedlings. Front Plant Sci. 2024;15:1396666. [CrossRef]
- Tang S, Xian Y, Wang F, Luo C, Song W, Xie S, Chen X, Cao A, Li H, Liu H. Comparative transcriptome analysis of leaves during early stages of chilling stress in two different chilling-tolerant brown-fiber cotton cultivars. PLoS One. 2021;16(2):e0246801. [CrossRef]
- Yang P, Lv S, Liang F, Tian J, Zhang Y, Jiang C, Zhang W. Photoprotective Responses of Cotton Non-Leaf Green Tissues to Short-Term Low-Temperature Stress. Physiol Plantarum. 2025;177(3):e70246. [CrossRef]
- Salvucci M, Crafts-Brandner S. Inhibition of photosynthesis by heat stress: the activation state of Rubisco as a limiting factor in photosynthesis. Physiol Plantarum. 2004;120(2):179-186. [CrossRef]
- Wang Q, Du X, Zhou Y, Xie L, Bie S, Tu L, Zhang N, Yang X, Xiao S, Zhang X. The β-ketoacyl-CoA synthase KCS13 regulates the cold response in cotton by modulating lipid and oxylipin biosynthesis. J Exp Bot. 2020;71(18):5615-5630. [CrossRef]
- Dev W, Sultana F, He S, Waqas M, Hu D, Aminu IM, Geng X, Du X. An insight into heat stress response and adaptive mechanism in cotton. J Plant Physiol. 2024;302:154324. [CrossRef]
- Cai X, Magwanga RO, Xu Y, Zhou Z, Wang X, Hou Y, Wang Y, Zhang Y, Liu F, Wang K. Comparative transcriptome, physiological and biochemical analyses reveal response mechanism mediated by CBF4 and ICE2 in enhancing cold stress tolerance in Gossypium thurberi. AoB Plants. 2019;11(6):plz045. [CrossRef]
- Geng Z, Liu J, Zhao G, Zhou Y, Feng T, Geng X, Liu X, Liu X, Zhang H, Wang Y. Global profiling of alternative splicing landscapes in response to cold stress during the cotyledon stage in cotton. BMC Genomics. 2026. [CrossRef]
- Ma X, Chen Z, Xiao G, Huang J, Lin L, Xu Q. An evolutionarily conserved histone modification H3K37ac activates gene transcription in response to salt stress in rice. New Phytol. 2025. [CrossRef]
- Zhu F, Xu Q, Fan J, Meng L, Wang R, Niu J, Wang J, Zhang G, Shi S, Wang F, Li H. Genome-Wide Identification of BPC Gene Family in Ten Cotton Species and Function Analysis of GhBPC4 Involved in Cold Stress Response. Int J Mol Sci. 2025;26(16):7978. [CrossRef]
- Cheng G, Zhang L, Wei H, Wang H, Lu J, Yu S. Transcriptome Analysis Reveals a Gene Expression Pattern Associated with Fuzz Fiber Initiation Induced by High Temperature in Gossypium barbadense. Genes. 2020;11(9):1066. [CrossRef]
- Yang N, Xu C, Liang Y, Zheng J, Geng S, Sun F, Li S, Lai C, Yusuyin M, Gong Z, Wang J. Comparative Transcriptome and Volatile Metabolome Analysis of Gossypium hirsutum Resistance to Verticillium Wilt. Genes (Basel). 2025;16(8):877. [CrossRef]
- Yang N, Gong Z, Liang Y, Geng S, Sun F, Li X, Qian S, Lai C, Yusuyin M, Wang J, Zheng J. RNA-Seq Analysis and Candidate Gene Mining of Gossypium hirsutum Stressed by Verticillium dahliae Cultured at Different Temperatures. Plants (Basel). 2024;13(19):2688. [CrossRef]
- Wu J, Zhang F, Liu G, Abudureheman R, Bai S, Wu X, Zhang C, Ma Y, Wang X, Zha Q. Transcriptome and coexpression network analysis reveals properties and candidate genes associated with grape (Vitis vinifera L.) heat tolerance. Front Plant Sci. 2023;14:1270933. [CrossRef]
- Shen Q, Zhang S, Liu S, Chen J, Ma H, Cui Z, Zhang X, Ge C, Liu R, Li Y, Zhao X, Yang G, Song M, Pang C. Comparative Transcriptome Analysis Provides Insights into the Seed Germination in Cotton in Response to Chilling Stress. Int J Mol Sci. 2020;21(6):2067. [CrossRef]
- Zhang T, Xie Z, Zhou J, Feng H, Zhang T. Temperature impacts on cotton yield superposed by effects on plant growth and verticillium wilt infection in China. Int J Biometeorol. 2024;68:199-209. [CrossRef]
- Zhu H, Song J, Dhar N, Shan Y, Ma XY, Wang XL, Chen JY, Dai XF, Li R, Wang ZS. Transcriptome Analysis of a Cotton Cultivar Provides Insights into the Differentially Expressed Genes Underlying Heightened Resistance to the Devastating Verticillium Wilt. Cells. 2021;10(11):2961. [CrossRef]
- Mi H, Muruganujan A, Ebert D, Huang X, Thomas PD. PANTHER version 14: more genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res. 2019;47(D1):D419-D426. [CrossRef]
- Xia Z, Yin Q, Qiu C, Yang Y, Guo D, Huang H, Tang Q, Jiang S, Zhao X, Chen Z. Transcriptome Profiling of the Leaf Spot Pathogen, Pestalotiopsis trachicarpicola, and Its Host, Tea (Camellia sinensis), During Infection. Plant Dis. 2022;106(8):2247-2252. [CrossRef]
- Dai Y, Zhou J, Zhang B, Zheng D, Wang K, Han J. Time-course transcriptome analysis reveals gene co-expression networks and transposable element responses to cold stress in cotton. BMC Genomics. 2025;26(1):235. [CrossRef]
- Yuan J, Liao YS, Zhang TC, Tang YQ, Yu P, Liu YN, Cai DJ, Yu SG, Zhao L. Integrating Bulk RNA and Single-Cell Sequencing Data Unveils Efferocytosis Patterns and ceRNA Network in Ischemic Stroke. Transl Stroke Res. 2025;16(3):733-746. [CrossRef]
- Zhong X, Han A, Liusui Y, Zhang X, Fu W, Wang Z, Li Y, Cao J, Guo Y, Zhang J. Identification of key pathways and genes underlying melatonin-enhanced drought tolerance in cotton. PeerJ. 2025;13:e20005. [CrossRef]
- Adams WW III, Muller O, Cohu CM, Demmig-Adams B. May photoinhibition be a consequence, rather than a cause, of limited plant productivity? Photosynth Res. 2013;117(1):31-44.
- Adams WW III, Stewart JJ, Cohu CM, Muller O, Demmig-Adams B. Habitat temperature and precipitation of Arabidopsis thaliana ecotypes determine the response of foliar vasculature, photosynthesis, and transpiration to growth temperature. Front Plant Sci. 2016;7:1026.
- Ashraf MA, Rasheed R, Hussain I, Iqbal M, Farooq MU, Saleem MH, Ali S. Taurine modulates dynamics of oxidative defense, secondary metabolism, and nutrient relation to mitigate boron and chromium toxicity in Triticum aestivum L. plants. Environ Sci Pollut Res Int. 2022;29:45527-45548. [CrossRef]
- Anwar A, Liu Y, Dong R, Bai L, Yu X, Li Y. The physiological and molecular mechanism of brassinosteroid in response to stress: a review. Biol Res. 2018;51(1):1-14.
- Barrero-Gil J, Salinas J. Post-translational regulation of cold acclimation response. Plant Sci. 2013;205:48-54.
- Chaumont F, Barrieu F, Wojcik E, Chrispeels MJ, Jung R. Aquaporins constitute a large and highly divergent protein family in maize. Plant Physiol. 2001;125(3):1206-1212.
- Sakurai J, Ishikawa F, Yamaguchi T, Uemura M, Maeshima M. Identification of 33 rice aquaporin genes and analysis of their expression and function. Plant Cell Physiol. 2005;46(10):1568-1577.
- Fu QS, Yang RC, Wang HS, Zhao B, Zhou CL, Ren SX, Guo YD. Leaf morphological and ultrastructural performance of eggplant (Solanum melongena L.) in response to water stress. Photosynthetica. 2013;51:109-114. [CrossRef]
- Binks O, Meir P, Rowland L, da Costa AC, Vasconcelos SS, de Oliveira AA, Ferreira L, Mencuccini M. Limited acclimation in leaf anatomy to experimental drought in Tropical Rainforest Trees. Tree Physiol. 2016;36:1550-1561. [CrossRef]
- Hacke UG, Spicer R, Schreiber SG, Plavcová L. An ecophysiological and developmental perspective on variation in vessel diameter. Plant Cell Environ. 2016;40:831-845. [CrossRef]
- Mo Y, Yang R, Liu L, Gu X, Yang X, Wang Y, Zhang X, Li H. Growth, photosynthesis and adaptive responses of wild and domesticated watermelon genotypes to drought stress and subsequent re-watering. Plant Growth Regul. 2015;79:229-241. [CrossRef]
- Luo J, Zhou J, Li H, Shi W, Polle A, Lu M, Sun X, Luo ZB. Global poplar root and leaf transcriptomes reveal links between growth and stress responses under nitrogen starvation and excess. Tree Physiol. 2015;35(12):1283-1302. [CrossRef]
- Cavender-Bares J. Impacts of freezing on long-distance transport in woody plants. Coll Biol Sci, Univ Minnesota. 2005.
- Jensen KH, Savage JA, Holbrook NM. Optimal concentration for sugar transport in plants. J R Soc Interface. 2013;10(84):20130055.
- Jyske T, Hölttä T. Comparison of phloem and xylem hydraulic architecture in Picea abies stems. New Phytol. 2015;205(1):102-115.
- Mencuccini M, Hölttä T, Sevanto S, Nikinmaa E. Concurrent measurements of change in the bark and xylem diameters of trees reveal a phloem-generated turgor signal. New Phytol. 2013;198(4):1143-1154.
- Molz FJ, Peterson CM. Location of the low temperature water flow barrier in stems. Plant Physiol. 1968;43(1):652-656.
- Adams WW III, Stewart JJ, Polutchko SK, Demmig-Adams B. Leaf vasculature and the upper limit of photosynthesis. In: Adams WW III, Terashima I, eds. Photosynthesis: Plastid Biology, Energy Conversion and Carbon Assimilation. Springer; 2018:3-24.
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).