Submitted:
03 March 2026
Posted:
04 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Structure of VPS34 and the Two Complexes
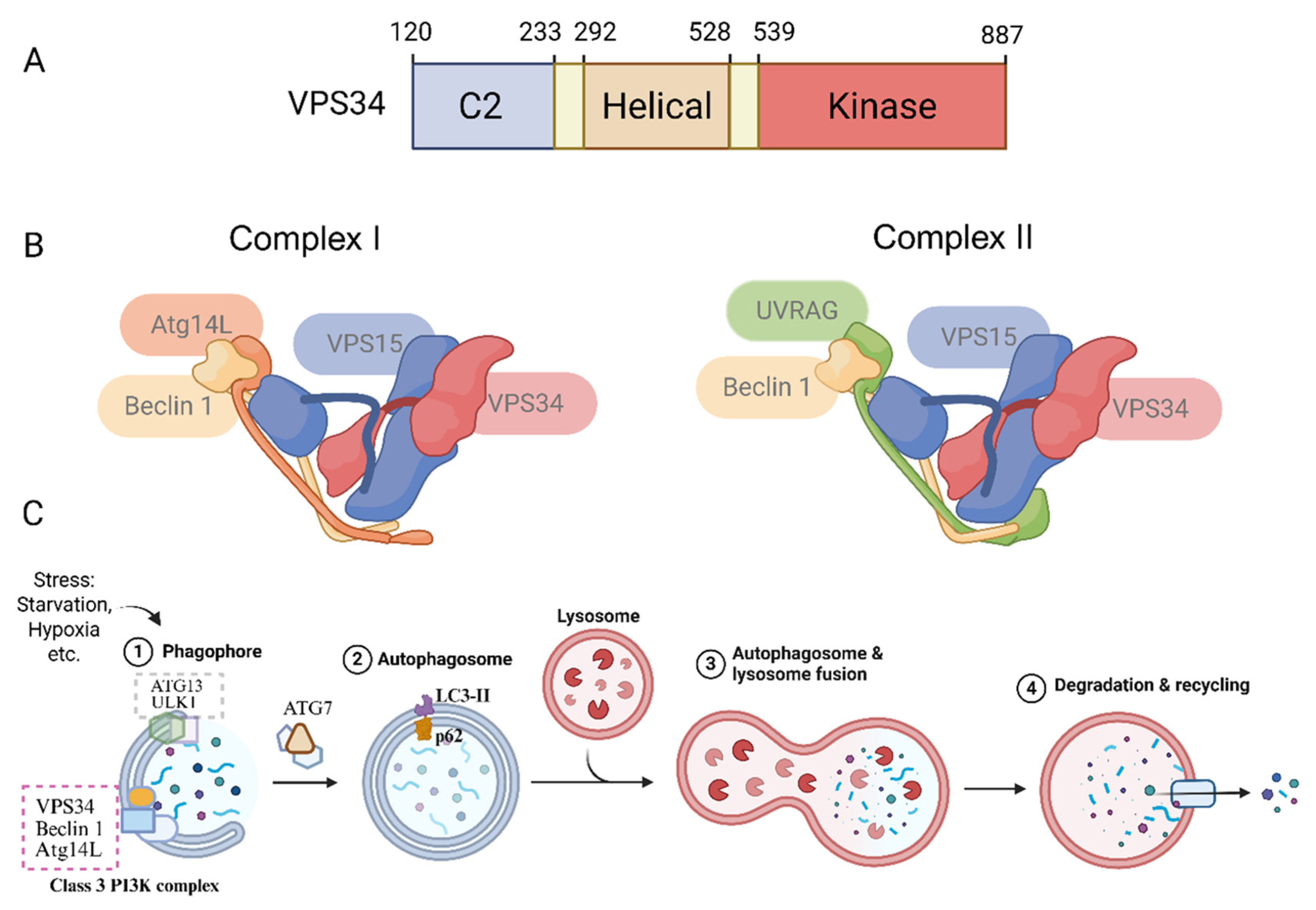
2.1. Molecular and Structural Features of VPS34
2.2. VPS34 Complexes: Complex I and Complex II
3. Canonical Functions of VPS34 in Autophagy
3.1. Role of VPS34 in Macroautophagy
3.2. Role of VPS34 in Selective Autophagy
3.3. Regulation of VPS34 Complexes and VPS34 Activity
3.3.1. Regulation of VPS34 Complex I and Complex II
3.3.2. Post-Translational Modifications of VPS34
3.3.3. Regulation of VPS34 by Hypoxia in a HIF-1α-Dependent Manner
4. Non-Autophagic Functions of VPS34
4.1. Role of VPS34 in Endosomal Trafficking and Vesicular Transport
4.2. Role of VPS34 in Phagocytosis, Lysosomal Biology and Nutrient Sensing
4.3. Role of VPS34 in Exocytosis and Secretion
5. Role of VPS34 in Cancer
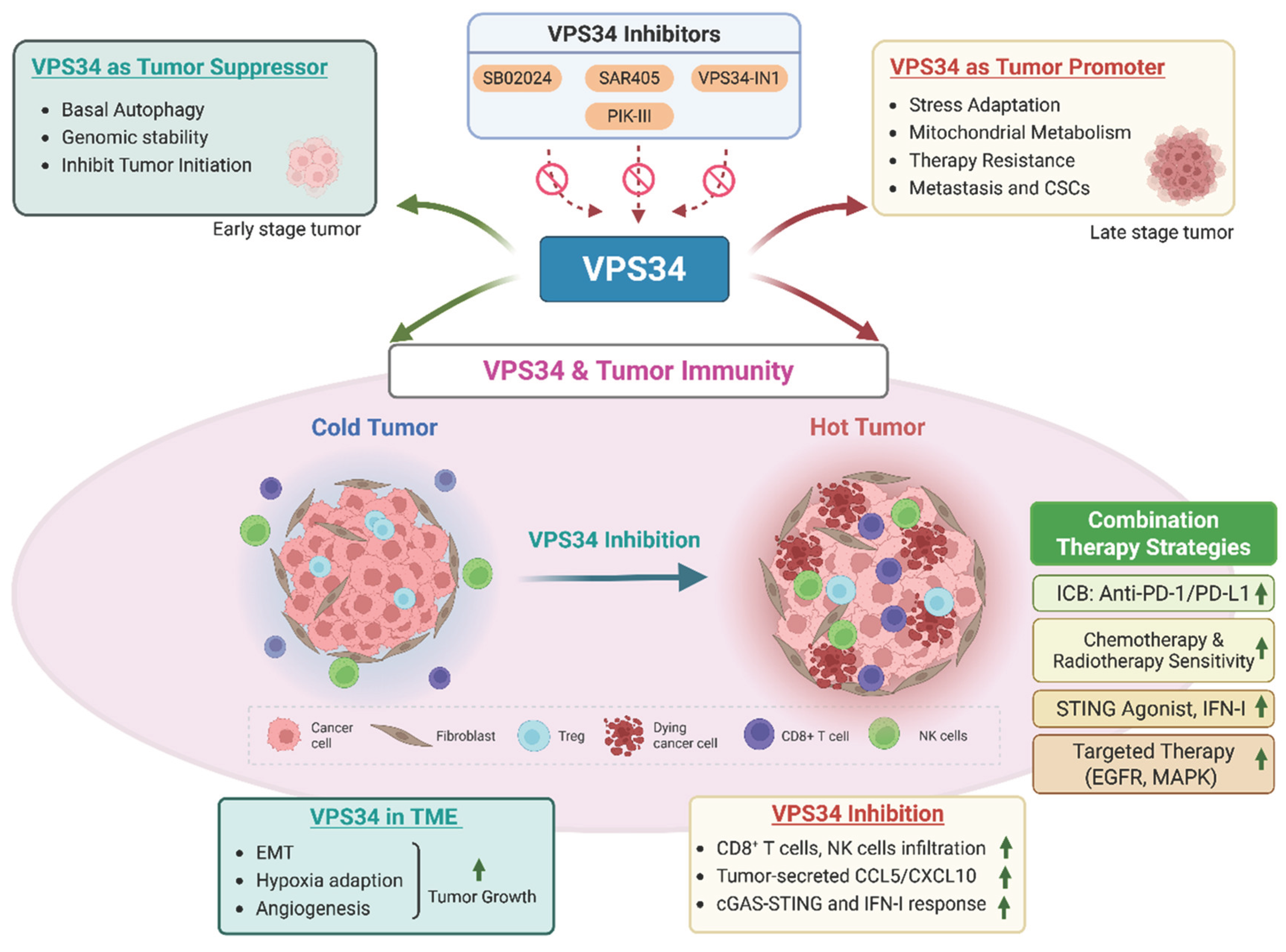
5.1. VPS34 as a Tumor Suppressor
5.2. VPS34 as a Tumor Promoter
5.3. VPS34’s Role on Cancer Cell Metabolism, Survival and Metastasis
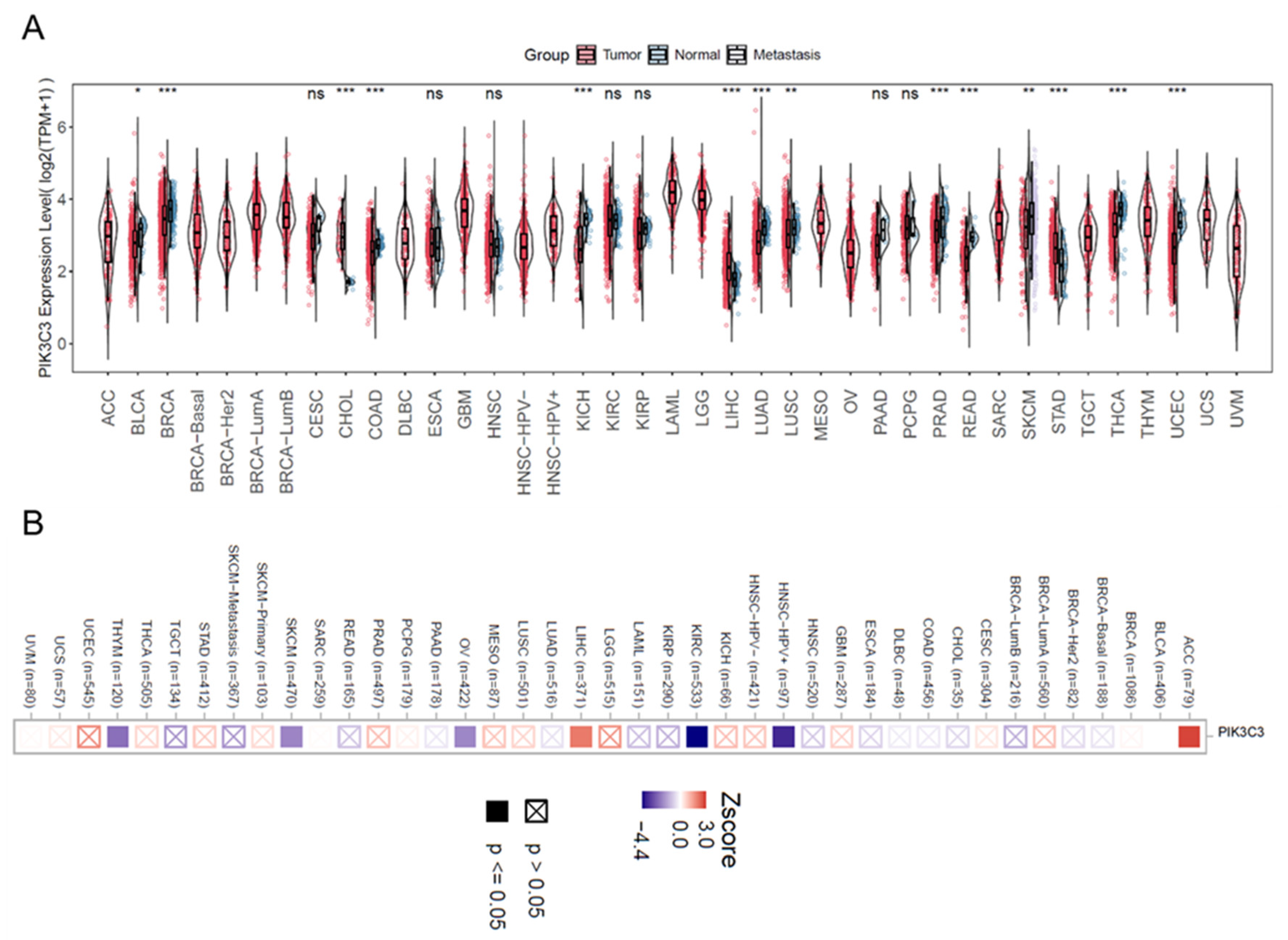
5.4. Genetic Depletion of VPS34 in Mouse Models and VPS34’s Clinical and Prognostic Relevance in Human Cancers
6. VPS34/Autophagy in Immunity and Cancer Immunotherapy
6.1. Role of VPS34/Autophagy in the Regulation of Innate Immunity
6.1.1. VPS34/Autophagy and cGAS-STING Pathway
6.1.2. Role of VPS34 in Dendritic Cells (DCs) and NK Cells
6.2. Role of VPS34/Autophagy in the Regulation of Adaptive Immunity
6.3. VPS34’s Role in Shaping the Tumor Microenvironment
7. Pharmacological Targeting of VPS34
7.1. Overview of VPS34 Inhibitors
7.2. Other VPS34 Inhibitors
8. Combination Strategies: Synergizing VPS34 Inhibition with Other Therapies in Cancers
8.1. Synergizing VPS34 Inhibition with ICB
8.2. Synergizing VPS34 Inhibition with Sting Agonists
8.3. Synergizing VPS34 Inhibition with Other Therapies
9. Conclusion and Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Cianciulli, A., Porro, C., Calvello, R., Trotta, T., Lofrumento, D. D., & Panaro, M. A. Microglia Mediated Neuroinflammation: Focus on PI3K Modulation. Biomolecules 2020, 10(11). [CrossRef]
- Len Stephens, P.H. Signalling via class IA PI3Ks. 2011; pp. Volume 51, Issue 51, Pages 27-36, ISSN 0065-2571. [CrossRef]
- He, Y., Sun, M.M., Zhang, G.G. et al. Targeting PI3K/Akt signal transduction for cancer therapy. Sig Transduct Target Ther 2021, 6, 425. [CrossRef]
- Thibault, B.; Ramos-Delgado, F.; Guillermet-Guibert, J. Targeting Class I-II-III PI3Ks in Cancer Therapy: Recent Advances in Tumor Biology and Preclinical Research. Cancers 2023, 15, 784. [CrossRef]
- Paul Workman, R.L.M.v.M. Unveiling the Secrets of the Ancestral PI3 Kinase Vps34. Cancer Cell, 2010, Volume 17, Issue 15, Pages 421-423, ISSN 1535-6108. [CrossRef]
- N. Jaber, Z.D., J. Chen, J. Catanzaro, Y. Jiang, L.M. Ballou, E. Selinger, X. Ouyang, R.Z. Lin, J. Zhang, & W. Zong. Class III PI3K Vps34 plays an essential role in autophagy and in heart and liver function. Proc. Natl. Acad. Sci. U.S.A. 2012, 109 (106) 2003-2008. [CrossRef]
- Françoise Hullin-Matsuda, T.T., Peter Greimel, Toshihide Kobayashi,. Lipid compartmentalization in the endosome system. In Seminars in Cell & Developmental Biology, Volume 31, 2014; pp. Pages 48-56, ISSN 1084-9521,.
- Furuya, T., Kim, M., Lipinski, M., Li, J., Kim, D., Lu, T., Shen, Y., Rameh, L., Yankner, B., Tsai, L. H., & Yuan, J. Negative regulation of Vps34 by Cdk mediated phosphorylation. Molecular cell 2010, 38(34), 500–511. [CrossRef]
- Gillooly, D.J., Simonsen, A., & Stenmark, H. Cellular functions of phosphatidylinositol 3-phosphate and FYVE domain proteins. The Biochemical journal 2001, 355(Pt 352), 249–258. [CrossRef]
- Bechtel, W., Helmstädter, M., Balica, J., Hartleben, B., Kiefer, B., Hrnjic, F., Schell, C., Kretz, O., Liu, S., Geist, F., Kerjaschki, D., Walz, G., & Huber, T. B. Vps34 deficiency reveals the importance of endocytosis for podocyte homeostasis. Journal of the American Society of Nephrology: JANS 2013, 24(25), 727-743. [CrossRef]
- Yu, Y., Bogdan, M., Noman, M. Z., Parpal, S., Bartolini, E., Van Moer, K., Kleinendorst, S. C., Bilgrav Saether, K., Trésaugues, L., Silvander, C., Lindström, J., Simeon, J., Timson, M. J..Janji, B. Combining VPS34 inhibitors with STING agonists enhances type I interferon signaling and anti-tumor efficacy. Molecular oncology 2024. [CrossRef]
- Bartolini, E., Van Moer, K., & Janji, B. Improving STING agonist-based cancer therapy by inhibiting the autophagy-related protein VPS34. Oncoimmunology 2024. [CrossRef]
- Bartolini, E., Van Moer, K., & Janji, B. Unleashing anti-tumor immunity: Targeting the autophagy-related protein VPS34 to enhance STING agonist-based therapy. Autophagy reports 2024.
- Noman, M.Z., Parpal, S., Van Moer, K., Xiao, M., Yu, Y., Viklund, J., De Milito, A., Hasmim, M., Andersson, M., Amaravadi, R. K., Martinsson, J., Berchem, G., & Janji, B. Inhibition of Vps34 reprograms cold into hot inflamed tumors and improves anti-PD-1/PD-L1 immunotherapy. Science advances 2020. [CrossRef]
- Sulochanadevi Baskaran, L.-A.C., Goran Stjepanovic, Lindsey N Young, Do Jin Kim, Patricia Grob, Robin E Stanley, Eva Nogales, James H Hurley. Architecture and dynamics of the autophagic phosphatidylinositol 3-kinase complex,. eLife 2014 3:e05. [CrossRef]
- Lo, W., Zhang, Y., Vadas, O. et al. Structural basis of phosphatidylinositol 3-kinase C2α function. Nat Struct Mol Biol 2022, 29, 218–228. [CrossRef]
- Morris, D.H., Yip, C. K., Shi, Y., Chait, B. T., & Wang, Q. J. BECLIN 1-VPS34 COMPLEX ARCHITECTURE: UNDERSTANDING THE NUTS AND BOLTS OF THERAPEUTIC TARGETS. Frontiers in biology 2015, 10(15), 398–426. [CrossRef]
- Ohashi, Y. Activation Mechanisms of the VPS34 Complex-es. Cells 2021, 10(11), 3124. [CrossRef]
- Yuan Liu, Q.Y., Siwei Chen, Zixiang Li, Leilei Fu. Targeting VPS34 in autophagy: An update on pharmacological small-molecule compounds. European Journal of Medicinal Chemistry, Volume 256, 2023, 115467, ISSN 110223-115234,. [CrossRef]
- Lee, Y., Tuan, N. M., Lee, G. J., Kim, B., Park, J. H., & Lee, C. H. Regulatory Mechanisms Governing the Autophagy-Initiating VPS34 Complex and Its inhibitors. Biomolecules & therapeu-tics 2024, 32(36), 723–735.
- Ohashi, Y., Tremel, S., & Williams, R. L. VPS34 complexes from a structural perspective. Journal of lipid research 2019, 60(62), 229–241. [CrossRef]
- Cao Zhumin, T.K., Ran Yincheng, Zhou Haonan, Zhou Lei, Ding Yana, Tang Xiaowei. Beclin-1: a therapeutic target at the intersection of autophagy, immunotherapy, and cancer treatment. Frontiers in Immunology.: 2024; p. Volume 15.
- Jing Ye, J.Z., Yanghui Zhu, Lian Wang, Xian Jiang, Bo Liu, Gu He. Targeting autophagy and be-yond: Deconvoluting the complexity of Beclin-1 from biological function to cancer therapy. Acta Phar-maceutica Sinica B: 2023; pp. Volume 13, Issue 12, Pages 468.
- Sargeet Kaur, H.C. The beclin 1 interactome: Modification and roles in the pathology of autophagy-related disorders,. In Biochimie,; ELSEVIER: 2020; pp. Pages 34-49,.
- Yohei Ohashi, S.T., Glenn Robert Masson, Lauren McGinney, Jerome Boulanger, Ksenia Rostislavleva, Christopher M Johnson, Izabella Niewczas, Jonathan Clark, Roger L Williams. Membrane characteristics tune activities of endosomal and autophagic human VPS34 complexes. eLife 2020, 9:e58281. [CrossRef]
- David M. Hollenstein, C.K. Autophagosomes are formed at a distinct cellular structure,. In Membrane Trafficking, Frances M Brodsky, J.L.S., Ed.; Current Opinion in Cell Biology: 2020; pp. Volume 65, 2020, Pages 2050-2057, ISSN 0955-0674,.
- Parzych, K.R., & Klionsky, D. J. An overview of autophagy: morphology, mechanism, and regulation. Antioxidants & redox signaling, 2014, 20(23), 460–473.
- Itakura, E., & Mizushima, N. Characterization of autophagosome formation site by a hierarchical analysis of mammalian Atg proteins. Autophagy, 2010, 6(6), 764–776. [CrossRef]
- Dooley, H.C., Razi, M., Polson, H. E., Girardin, S. E., Wilson, M. I., & Tooze, S. A. WIPI2 links LC3 conjugation with PI3P, autophagosome formation, and pathogen clearance by recruiting Atg12-5-16L1. Molecular cell 2014, 55(52), 238–252. [CrossRef]
- Hu, M., Ladowski, J. M., & Xu, H. The Role of Autophagy in Vascular Endothelial Cell Health and Physiology. Cells 2024, 13(10), 825. [CrossRef]
- Ornatowski, W., Lu, Q., Yegambaram, M., Garcia, A. E., Zemskov, E. A., Maltepe, E., Fineman, J. R., Wang, T., & Black, S. M. Complex interplay between autophagy and oxidative stress in the development of pulmonary disease. Redox biology, 2020, 36, 101679. [CrossRef]
- Camuzard, O., Santucci-Darmanin, S., Carle, G. F., & Pierrefite-Carle, V. Autophagy in the crosstalk between tumor and microenvironment. Cancer letters 2020, 490, 143–153. [CrossRef]
- Alessio Reggio, V.B., Paolo Grumati. Eating the unknown: Xenophagy and ER-phagy are cytoprotective defenses against pathogens. experimental cell research: 2020; pp. Volume 396, Issue 391, 112276, ISSN 110014-114827. [CrossRef]
- Gubas, A.a.D., I. A guide to the regulation of selective autophagy receptors. FEBS J 2022, 289: 275-289. [CrossRef]
- Claudia Dall’Armi, K.A.D., Gilbert Di Paolo. The Role of Lipids in the Control of Autophagy. Current Biology,: 2013; pp. Volume 23, Issue 21,Pages R33-R45, ISSN 0960-9822.
- Lorentzen, K.C., Prescott, A. R., & Ganley, I. G. Artificial tar-geting of autophagy components to mitochondria reveals both conventional and unconventional mitophagy pathways Autophagy 2025, 21(22), 315–337. [CrossRef]
- Mao, K., & Klionsky, D. J. Xenophagy: A battlefield between host and microbe, and a possible avenue for cancer treatment Autophagy 2017, 13(12), 223–224. [CrossRef]
- Pablo Sanz-Martinez, A.S. Mechanisms and physiological functions of ER-phagy. In Current Opinion in Physiology; 2022; pp. Volume 30,100613,ISSN 102468-108673,. [CrossRef]
- Hayat, M.A. Chapter 1 - Introduction to Autophagy: Cancer, Other Pathologies, Inflammation, Immunity, Infection, and Aging,. In Autophagy: Cancer, Other Pathologies, Inflamma-tion, Immunity, Infection and Aging; Academic Press: 2014; pp. Volumes 1–4, 1-35, ISB.
- John RP Knight, T.S., Mark Stoneley, Anne E Willis. Ribosomes and Stress - Linked from Birth to Death. In Encyclopedia of Cell Biology (Second Edition); Academic Press: 2023; pp. Pages 44-56, ISBN 9780128216248.
- Wold, M.S., Lim, J., Lachance, V., Deng, Z., & Yue, Z. ULK1-mediated phosphorylation of ATG14 promotes autophagy and is impaired in Huntington’s disease models. Molecular neurodegeneration 2016, 11(11), 76. [CrossRef]
- Ki Eun Pyo, C.R.K., Minkyoung Lee, Jong-Seo Kim, Keun Il Kim, Sung Hee Baek,. ULK1 O-GlcNAcylation Is Crucial for Activating VPS34 via ATG14L during Autophagy Initiation,. In Cell Reports,; 2018; Volume 2825, pp. Pages 2878-2890.e2874,. [CrossRef]
- Zhang, S., Lin, X., Hou, Q., Hu, Z., Wang, Y., & Wang, Z. Regulation of mTORC1 by amino acids in mammalian cells: A general picture of recent advances. Animal nutrition 2021. [CrossRef]
- Garcia, D., & Shaw, R. J. AMPK: Mechanisms of Cellular Energy Sensing and Restoration of Metabolic Balance. Molecular cell 2017. [CrossRef]
- Lin, M.G., & Hurley, J. H. Structure and function of the ULK1 com-plex in autophagy. Current opinion in cell biology, 2016, 39, 61–68. [CrossRef]
- Yun, Z.; Wang, Q.J.; Yue, Z. Atg14L and Rubicon: yin and yang of Beclin 1-mediated autophagy control. Autophagy 2009, 5, 890-891. [CrossRef]
- Magné, J., & Green, D. R. LC3-associated endocytosis and the functions of Rubicon and ATG16L1. Science advances 2022, 8. eabo5600. [CrossRef]
- Bhargava, H.K., Tabata, K., Byck, J. M., Hamasaki, M., Farrell, D. P., Anishchenko, I., DiMaio, F., Im, Y. J., Yoshimori, T., & Hurley, J. H. Structural basis for autophagy inhibition by the human Rubicon-Rab7 complex. Proceedings of the National Academy of Sciences of the United States of America 2020. [CrossRef]
- Wong, S.-W., Sil, P. and Martinez, J. Rubicon: LC3-associated phagocytosis and beyond. FEBS J, 2018, 285: 1379-1388.
- Nah, J., Zablocki, D. & Sadoshima, J. The roles of the inhibitory autophagy regulator Rubicon in the heart: A new therapeutic target to prevent cardiac cell death. Exp Mol Med 2021, 53, 528–536. [CrossRef]
- Kim, J., Kim, Y. C., Fang, C., Russell, R. C., Kim, J. H., Fan, W., Liu, R., Zhong, Q., & Guan, K. L. Differential regulation of distinct Vps34 complexes by AMPK in nutrient stress and autophagy. Cell 2013. [CrossRef]
- Yohei Ohashi, S.T., Roger L. Williams,. VPS34 complexes from a structural perspective,. Journal of Lipid Research, 2019. [CrossRef]
- Hill, S.M., Wrobel, L., & Rubinsztein, D. C. Post-translational modifications of Beclin 1 provide multiple strategies for autophagy regulation. Cell death and differentiation 2019.
- Jia M., Y., X., Sun, W., Zhou, Q., Chang, C., Gong, W., Feng, J., Li, X., Zhan, R., Mo, K., Zhang, L., Qian, Y., Sun, Y., Wang, A., Zou, Y., Chen, W., Li, Y., Huang, L., Yang, Y., Zhao, Y., … Cheng, X.,. ULK1-mediated metabolic reprogramming regulates Vps34 lipid kinase activity by its lactylation. Science advances 2023. [CrossRef]
- Ye Y., H., Q., Chen, H., Liang, K., Yuan, Y., Xiang, Y., Ruan, H., Zhang, Z., Song, A., Zhang, H., Liu, L., Diao, L., Lou, Y., Zhou, B., Wang, L., Zhou, S., Gao, J., Jonasch, E., Lin, S. H., Xia, Y., … Han, L. Characterization of Hypoxia-associated Molecular Features to Aid Hypoxia-Targeted Therapy. Nature metabolism 2019. [CrossRef]
- Chen Z., H., F., Du, Y. et al. Hypoxic microenvironment in cancer: molecular mechanisms and therapeutic interventions. Sig Transduct Target Ther 2023.
- Ziello J. E., J., I. S., & Huang, Y. Hypoxia-Inducible Factor (HIF)-1 regulatory pathway and its potential for therapeutic intervention in malignancy and ischemia. The Yale journal of biology and medicine 2007.
- Zaarour R. F., A., B., Hajam, E. Y., Nawafleh, H., Zeinelabdin, N. A., Engelsen, A. S. T., Thiery, J., Jamora, C., & Chouaib, S. Role of Hypoxia-Mediated Autophagy in Tumor Cell Death and Survival. Cancers 2021. [CrossRef]
- Zhao L., Q., H., Lv, H., Liu, W., Zhang, R., Yang, A. Lactylation in health and disease: physiological or pathological?. Theranostics 2025. [CrossRef]
- Kierans S.J. and Taylor, C.T. Regulation of glycolysis by the hypoxia-inducible factor (HIF): implications for cellular physiology. J Physiol 2021. [CrossRef]
- Dong, J., Tong, W., Liu, M. et al. Endosomal traffic disorders: a driving force behind neuro-degenerative diseases. Transl Neurodegener 13, 66 2024. [CrossRef]
- Tremel, S., Ohashi, Y., Morado, D.R. et al. Structural basis for VPS34 kinase activation by Rab1 and Rab5 on membranes. Nat Commun 12, 1564 2021. [CrossRef]
- Law, F., & Rocheleau, C. E. Vps34 and the Armus/TBC-2 Rab GAPs: Putting the brakes on the endosomal Rab5 and Rab7 GTPases. Cellular Logistics 7(4). 2017. [CrossRef]
- W. Liu, K.W., Y. Lin, L. Wang, X. Jin, Y. Qiu, W. Sun, L. Zhang, Y. Sun, X. Dou, S. Luo, Y. Su, Q. Sun, W. Xiang, F. Diao, J. Li. VPS34 Governs Oocyte Developmental Competence by Regulating Mito/Autophagy: A Novel Insight into the Significance of RAB7 Activity and its subcellular location. Adv. Sci. 2024. [CrossRef]
- He, F., Nichols, R. M., Agosto, M. A., & Wensel, T. G. Roles of class III phosphatidylinositol 3-kinase, Vps34, in phagocytosis, autophagy, and endocytosis in retinal pigmented epithelium. iScience, 28(5), 112371. 2025. [CrossRef]
- Mélanie Mansat, R.J.B. Lysosome identity crisis: Phosphoinositides and mTORC1 negotiate lysosomal behavior. Molecular Cell, Volume 84, Issue 1 2024, Pages 17-19. [CrossRef]
- Wu, S., Chen, JW., Liu, HY. et al. Secretory autophagy promotes Rab37-mediated exocyto-sis of tissue inhibitor of metalloproteinase 1. J Biomed Sci 29, 103 2022.
- Buratta, S., Tancini, B., Sagini, K., Delo, F., Chiaradia, E., Urbanelli, L., & Emiliani, C. Lysosomal Exocytosis, Exosome Release and Secretory Autophagy: The Autophagic- and Endo-Lysosomal Systems Go Extracellular. International journal of molecular Sciences, 21(7), 2576 2020. [CrossRef]
- Paulina Kaminska, A.T., Ela Scholz, Anna R. Malik. Cytokines on the way to secretion,Cytokine & Growth Factor Reviews. 2024; pp. Volume 79, Pages 52-65,ISSN 1359-6101.
- Weigert A, H.L. Immune modulation through secretory autophagy. J Cell Biochem. 2024. [CrossRef]
- Li, X., Zhao, H. Targeting secretory autophagy in solid cancers: mechanisms, immune regulation and clinical insights. Exp Hematol Oncol 2025. [CrossRef]
- Jaber, N., Dou, Z., Chen, J. S., Catanzaro, J., Jiang, Y. P., Ballou, L. M., Selinger, E., Ouyang, X., Lin, R. Z., Zhang, J., & Zong, W. X. Class III PI3K Vps34 plays an essential role in autophagy and in heart and liver function. Proceedings of the National Academy of Sciences of the United States of America 2012.
- Qi, C., Zou, L., Wang, S., Mao, X., Hu, Y., Shi, J., Zhang, Z., & Wu, H. Vps34 Inhibits Hepatocellular Carcinoma Invasion by Regulating Endosome-Lysosome Trafficking via Rab7-RILP and Rab11. Cancer research and treatment 2022. [CrossRef]
- Lv, Q., Wang, W., Xue, J., Hua, F., Mu, R., Lin, H., Yan, J., Lv, X., Chen, X., & Hu, Z. W. DEDD interacts with PI3KC3 to activate autophagy and attenuate epithelial-mesenchymal transition in human breast cancer. Cancer research 2012. [CrossRef]
- Elkholi, I.E., Robert, A., Malouf, C., Wu, J. L., Kuasne, H., Drapela, S., Macleod, G., Hébert, S., Pacis, A., Calderon, V., Kleinman, C. L., Gomes, A. P., Alvarez, J. V., Aguirre-Ghiso, J. A., Park, M., Angers, S., & Côté, J. F. Targeting the Dependence on PIK3C3-mTORC1 Signaling in Dormancy-Prone Breast Cancer Cells Blunts Metastasis Initiation. Cancer research 2025. [CrossRef]
- F. Ramos-Delgado, H.S., C. Guyon, C. Handschin, P. Cerapio-Arroyo, R. D’Angelo, N. Therville, A. Villard, C. Cayron, C. Valle, E. Sarot, N Dussere, M. Di-Luoffo, V. Rebours, A. Couvelard, C. Joffre,H. de Oliveira,M. Dufresne,B. Thibault,J. Guilbert. Autophagy driven by VPS34 enables differentiated cell plasticity and cancer initiation. bioRxiv 2025. [CrossRef]
- Liu, F., Wu, X., Qian, Y., Jiang, X., Wang, Y., & Gao, J. PIK3C3 regulates the expansion of liver CSCs and PIK3C3 inhibition counteracts liver cancer stem cell activity induced by PI3K inhibitor. Cell death & disease 2020.
- Jiang, X., Bao, Y., Liu, H., Kou, X., Zhang, Z., Sun, F., Qian, Z., Lin, Z., Li, X., Liu, X., Jiang, L., & Yang, Y. Oncogene. VPS34 stimulation of p62 phosphorylation for cancer progression 2017.
- Zhou, X., Takatoh, J., & Wang, F. The mammalian class 3 PI3K (PIK3C3) is required for early embryogenesis and cell proliferation. PloS one 2011.
- Zhou, X., Wang, L., Hasegawa, H., Amin, P., Han, B. X., Kaneko, S., He, Y., & Wang, F. Deletion of PIK3C3/Vps34 in sensory neurons causes rapid neurodegeneration by disrupting the endosomal but not the autophagic pathway. Proceedings of the National Academy of Sciences of the United States of America 2010.
- Grieco, G., Janssens, V., Gaide Chevronnay, H. P., N’Kuli, F., Van Der Smissen, P., Wang, T., Shan, J., Vainio, S., Bilanges, B., Jouret, F., Vanhaesebroeck, B., Pierreux, C. E., & Courtoy, P. J. Vps34/PI3KC3 deletion in kidney proximal tubules impairs apical trafficking and blocks autophagic flux, causing a Fanconi-like syndrome and renal insufficiency. Scientific reports 2018. [CrossRef]
- Yue, Z., Jin, S., Yang, C., Levine, A. J., & Heintz, N. Beclin 1, an autophagy gene essential for early embryonic development, is a haploinsufficient tumor suppressor. Proceedings of the National Academy of Sciences of the United States of America 2003.
- Takamura, A., Komatsu, M., Hara, T., Sakamoto, A., Kishi, C., Waguri, S., Eishi, Y., Hino, O., Tanaka, K., & Mizushima, N. Autophagy-deficient mice develop multiple liver tumors. Genes & development 2011.
- Cho, K.J., Shin, S. Y., Moon, H., Kim, B. K., & Ro, S. W. Knockdown of Atg7 suppresses Tumorigenesis in a murine model of liver cancer. Translational oncology 2021. [CrossRef]
- Zhu, C., Liu, W., & Da, M. Expression, Characteristics, and Clinical Target Prediction of PIK3C3/ vps34 in Gastric Cancer. Current cancer drug targets 2024. [CrossRef]
- Di Donato, M., Giovannelli, P., Migliaccio, A., & Bilancio, A. Inhibition of Vps34 and p110δ PI3K Impairs Migration, Invasion and Three-Dimensional Spheroid Growth in Breast Cancer Cells. International journal of molecular sciences 2022. [CrossRef]
- Deng, C., Chen, D., Yang, L., Zhang, Y., Jin, C., Li, Y., Lin, Q., Luo, M., Zheng, R., Huang, B., & Liu, S. The role of cGAS-STING pathway ubiquitination in innate immunity and multiple diseases. Frontiers in immunology 2025. [CrossRef]
- Liu, Y., Chen, X., Zhao, Y., Wang, X. Y., Luo, Y. W., Chen, L., Wang, W., Zhong, S., Hu, M., Dai, Z., Jiang, J., Wang, X., Ji, H., Cheng, X. X., Zheng, A., Zuo, J., Liu, H., Ma, D., Luo, Z., Cao, F., … Tang, K. F. Small cytosolic double-stranded DNA represses cyclic GMP-AMP synthase activation and induces autophagy. Cell reports 2023. [CrossRef]
- Chen, M., Meng, Q., Qin, Y., Liang, P., Tan, P., He, L., Zhou, Y., Chen, Y., Huang, J., Wang, R. F., & Cui, J. TRIM14 Inhibits cGAS Degradation Mediated by Selective Autophagy Receptor p62 to Promote Innate Immune Responses. Molecular cell 2016. [CrossRef]
- Huang, T., Sun, C., Du, F., & Chen, Z. J. STING-induced noncanonical autophagy regulates endolysosomal homeostasis. Proceedings of the National Academy of Sciences of the United States of America 2025.
- Kuchitsu, Y., Mukai, K., Uematsu, R., Takaada, Y., Shinojima, A., Shindo, R., Shoji, T., Hamano, S., Ogawa, E., Sato, R., Miyake, K., Kato, A., Kawaguchi, Y., Nishitani-Isa, M., Izawa, K., Nishikomori, R., Yasumi, T., Suzuki, T., Dohmae, N., Uemura, T., …. STING signalling is terminated through ESCRT-dependent microautophagy of vesicles originating from recycling endosomes. Nature cell biology 2023. [CrossRef]
- Tokatly Latzer, I., Roullet, J. B., Cesaro, S., DiBacco, M. L., Arning, E., Rotenberg, A., Lee, H. H. C., Opladen, T., Jeltsch, K., García-Cazorla, À., Juliá-Palacios, N., Gibson, K. M., Bertoldi, M., & Pearl, P. L. Phenotypic correlates of structural and functional protein impairments resultant from ALDH5A1 variants. Human genetics 2023. [CrossRef]
- Gonugunta VK, S.T., Pokatayev V, et al. Trafficking-Mediated STING Degradation Requires Sorting to Acidified Endolysosomes and Can Be Targeted to Enhance Anti-tumor Response. Cell Reports 2017. [CrossRef]
- Paulo Antas, M.D.M., Fátima Leite-Pinheiro, Daniela Barros, Carlota Ramalhinho, Andreia Mendes, Beatriz H. Ferreira, Daniela Carvoeiro, Marisa Reverendo, Iola F. Duarte, Miwako Narita, Bing Su, Rafael J. Argüello, Beatrice Nal,..Evelina Gatti. VPS34-IN1 inhibits cap-mediated translation and synergizes with STING to drive type-I IFN expression in human plasmacytoid DCs. bioRxiv 2024.
- Chen, S., Li, Z., Feng, J., Quan, Y., He, J., Hao, J., & Dong, Z. Dual Activity of Type III PI3K Kinase Vps34 is Critical for NK Cell Development and Senescence. Advanced science 2024. [CrossRef]
- Crotzer, V.L., & Blum, J. S. Autophagy and adaptive immunity. Immunology, 2010.
- Metur, S.P., & Klionsky, D. J. Adaptive immunity at the crossroads of autophagy and metabolism. Cellular & molecular immunology 2021.
- Willinger, T., & Flavell, R. A. Canonical autophagy dependent on the class III phosphoinositide-3 kinase Vps34 is required for naive T-cell homeostasis. Proceedings of the National Academy of Sciences of the United States of America 2012.
- Courreges, C.J.F., Davenport, E. C. M., Bilanges, B., Rebollo-Gomez, E., Hukelmann, J., Schoenfelder, P., Edgar, J. R., Sansom, D., Scudamore, C. L., Roychoudhuri, R., Garden, O. A., Vanhaesebroeck, B., & Okkenhaug, K. Lack of phosphatidylinositol 3-kinase VPS34 in regulatory T cells leads to a fatal lymphoproliferative disorder without affecting their development. Frontiers in immunology, 2024. [CrossRef]
- Postoak, J.L., Song, W., Yang, G., Guo, X., Xiao, S., Saffold, C. E., Zhang, J., Joyce, S., Manley, N. R., Wu, L., & Van Kaer, L. Thymic epithelial cells require lipid kinase Vps34 for CD4 but not CD8 T cell selection. The Journal of experimental medicine 2022. [CrossRef]
- Postoak, J.L., Song, W., Wu, L., & Van Kaer, L. PIK3C3/VPS34 helps school T cells in the thymus. Autophagy 2023. [CrossRef]
- Yang, G., Song, W., Postoak, J. L., Chen, J., Martinez, J., Zhang, J., Wu, L., & Van Kaer, L. Autophagy-related protein PIK3C3/VPS34 controls T cell metabolism and function. Autophagy 2021. [CrossRef]
- Yang G, V.K.L. PIK3C3/VPS34 links T-cell autophagy to autoimmunity. Cell Death & Disease. 2020.
- Ghislat, G., & Lawrence, T. Autophagy in dendritic cells. Cellular & molecular immunology, 2018.
- Parekh, V.V., Pabbisetty, S. K., Wu, L., Sebzda, E., Martinez, J., Zhang, J., & Van Kaer, L. Autophagy-related protein Vps34 controls the homeostasis and function of antigen cross-presenting CD8α+ dendritic cells. Proceedings of the National Academy of Sciences of the United States of America 2017.
- Monaci S, C.F., Rossi D, Giuntini G, Filippi I, Marotta G, Sozzani S, Carraro F, Naldini A. Hypoxia Induces Autophagy in Human Dendritic Cells: Involvement of Class III PI3K/Vps34. Cells 2022. [CrossRef]
- Yiwen Wang, L.P., Sung Hoon Cho, Wenqiang Song, Lan Wu, Mark Robin Boothby, Luc Van Kaer. Autophagy-related lipid kinase Vps34 is required for B cell homeostasis and humoral immunity 2555. In The Journal of Immunology, Volume 214; 2025. [CrossRef]
- Hu, Y.L., DeLay, M., Jahangiri, A., Molinaro, A. M., Rose, S. D., Carbonell, W. S., & Aghi, M. K. Hypoxia-induced autophagy promotes tumor cell survival and adaptation to antiangiogenic treatment in glioblastoma. Cancer research 2012. [CrossRef]
- Schaaf, M.B., Houbaert, D., Meçe, O., & Agostinis, P. Autophagy in endothelial cells and tumor angiogenesis. Cell death and differentiation 2019.
- Ronan, B., Flamand, O., Vescovi, L., Dureuil, C., Durand, L., Fassy, F., Bachelot, M. F., Lamberton, A., Mathieu, M., Bertrand, T., Marquette, J. P., El-Ahmad, Y., Filoche-Romme, B., Schio, L., Garcia-Echeverria, C., Goulaouic, H., & Pasquier, B. A highly potent and selective Vps34 inhibitor alters vesicle trafficking and autophagy. Nature chemical biology, 2014. [CrossRef]
- Bago, R., Malik, N., Munson, M. J., Prescott, A. R., Davies, P., Sommer, E., Shpiro, N., Ward, R., Cross, D., Ganley, I. G., & Alessi, D. R. Characterization of VPS34-IN1, a selective inhibitor of Vps34, reveals that the phosphatidylinositol 3-phosphate-binding SGK3 protein kinase is a downstream target of class III phosphoinositide 3-kinase. The Biochemical journal, 2014. [CrossRef]
- Dyczynski, M., Yu, Y., Otrocka, M., Parpal, S., Braga, T., Henley, A. B., Zazzi, H., Lerner, M., Wennerberg, K., Viklund, J., Martinsson, J., Grandér, D., De Milito, A., & Pokrovskaja Tamm, K. Targeting autophagy by small molecule inhibitors of vacuolar protein sorting 34 (Vps34) improves the sensitivity of breast cancer cells to Sunitinib. Cancer letters 2018. [CrossRef]
- Dowdle, W.E., Nyfeler, B., Nagel, J., Elling, R. A., Liu, S., Triantafellow, E., Menon, S., Wang, Z., Honda, A., Pardee, G., Cantwell, J., Luu, C., Cornella-Taracido, I., Harrington, E., Fekkes, P., Lei, H., Fang, Q., Digan, M. E.,.. Murphy, L. O. Selective VPS34 inhibitor blocks autophagy and uncovers a role for NCOA4 in ferritin degradation and iron homeostasis in vivo. Nature cell biology 2014. [CrossRef]
- New, J., Arnold, L., Ananth, M., Alvi, S., Thornton, M., Werner, L., Tawfik, O., Dai, H., Shnayder, Y., Kakarala, K., Tsue, T. T., Girod, D. A., Ding, W. X., Anant, S., & Thomas, S. M. Secretory Autophagy in Cancer-Associated Fibroblasts Promotes Head and Neck Cancer Progression and Offers a Novel Therapeutic Target. Cancer research 2017. [CrossRef]
- David Schlütermann, M.A.S., Niklas Berleth, Philip Böhler, Jana Deitersen, Fabian Stuhldreier, Nora Wallot-Hieke, Wenxian Wu, Christoph Peter, Michèle J Hoffmann, Günter Niegisch, Björn Stork. Targeting urothelial carcinoma cells by combining cisplatin with a specific inhibitor of the autophagy-inducing class III PtdIns3K complex. Urologic oncology 2017. [CrossRef]
- Kuwabara, Y.; Sakai, K.; Ishi, S.; Yokosuka, S.; Abe, M.; Takahashi, T.; Kawano, Y.; Nishimura, H.; Toda-Sasaki, M.; Kobayashi-Ogawa, Y.; et al. Evaluation of effect of autophagy inhibition by SAR405, a selective Vps34 inhibitor, on proliferation of pleural mesothelioma cells. Cancer Res 15 March 2024; 84 (6_Supplement): 4711. 2024. [CrossRef]
- Liu, X., Wang, A., Liang, X., Liu, J., Zou, F., Chen, C., Zhao, Z., Deng, Y., Wu, H., Qi, Z., Wang, B., Wang, L., Liu, F., Xu, Y., Wang, W., Fernandes, S. M., Stone, R. M., Galinsky, I. A., Brown, J. R., Loh, T., … Liu, Q. Simultaneous inhibition of Vps34 kinase would enhance PI3Kδ inhibitor cytotoxicity in the B-cell malignancies. Oncotarget 2016. [CrossRef]
- Bago, R., Sommer, E., Castel, P., Crafter, C., Bailey, F. P., Shpiro, N., Baselga, J., Cross, D., Eyers, P. A., & Alessi, D. R. The hVps34-SGK3 pathway alleviates sustained PI3K/Akt inhibition by stimulating mTORC1 and tumour growth. The EMBO journal 2016. [CrossRef]
- Meunier, G., Birsen, R., Cazelles, C. et al. Antileukemic activity of the VPS34-IN1 inhibitor in acute myeloid leukemia. Oncogenesis 2020. [CrossRef]
- Wu, Q., Zhou, D., Shen, Z., Chen, B., Wang, G., Wu, L., Zhang, L., Li, X., Yuan, L., Wu, Y., Qu, N., & Zhou, W. VPS34-IN1 induces apoptosis of ER+ breast cancer cells via activating PERK/ATF4/CHOP pathway. Biochemical pharmacology 2023. [CrossRef]
- Kobylarz, M.J., Goodwin, J. M., Kang, Z. B., Annand, J. W., Hevi, S., O’Mahony, E., McAllister, G., Reece-Hoyes, J., Wang, Q., Alford, J., Russ, C., Lindeman, A., Beibel, M., Roma, G., Carbone, W., Knehr, J., Loureiro, … Nyfeler, B. An iron-dependent metabolic vulnerability underlies VPS34-dependence in RKO cancer cells. PloS one 2020.
- Chen, C.H., Liu, Y. M., Pan, S. L., Liu, Y. R., Liou, J. P., & Yen, Y. Trichlorobenzene-substituted azaaryl compounds as novel FGFR inhibitors exhibiting potent antitumor activity in bladder cancer cells in vitro and in vivo. Oncotarget 2016. [CrossRef]
- Chen, C.H., Changou, C. A., Hsieh, T. H., Lee, Y. C., Chu, C. Y., Hsu, K. C., Wang, H. C., Lin, Y. C., Lo, Y. N., Liu, Y. R., Liou, J. P., & Yen, Y. Dual Inhibition of PIK3C3 and FGFR as a New Therapeutic Approach to Treat Bladder Cancer. Clinical cancer research: an official journal of the American Association for Cancer Research 2018.
- Chen, C.H., Hsieh, T. H., Lin, Y. C., Liu, Y. R., Liou, J. P., & Yen, Y. Targeting Autophagy by MPT0L145, a Highly Potent PIK3C3 Inhibitor, Provides Synergistic Interaction to Targeted or Chemotherapeutic Agents in Cancer Cells. Cancers 2019. [CrossRef]
- Janji, B., Hasmim, M., Parpal, S., De Milito, A., Berchem, G., & Noman, M. Z. Lighting up the fire in cold tumors to improve cancer immunotherapy by blocking the activity of the autophagy-related protein PIK3C3/VPS34. Autophagy 2020. [CrossRef]
- Zhang, J.; Chen, L.; Takahashi, Y.; Wang, H.-G. Abstract LB033: Targeting autophagy in neuroblastoma: Inhibiting VPS34 to enhance anti-GD2 immunotherapy. Cancer Res 15 April 2025; 85 (8_Supplement_2): LB033. 2025. [CrossRef]
- Mele, L., Del Vecchio, V., Liccardo, D., Prisco, C., Schwerdtfeger, M., Robinson, N., Desiderio, V., Tirino, V., Papaccio, G., & La Noce, M. The role of autophagy in resistance to targeted therapies. Cancer treatment reviews 2020. [CrossRef]
- Xiao, M., Benoit, A., Hasmim, M., Duhem, C., Vogin, G., Berchem, G., Noman, M. Z., & Janji, B. Targeting Cytoprotective Autophagy to Enhance Anticancer Therapies. Frontiers in oncology 2021. [CrossRef]
- Hassan, A.M.I.A., Zhao, Y., Chen, X., & He, C. Blockage of Autophagy for Cancer Therapy: A Comprehensive Review. International journal of molecular sciences, 2024. [CrossRef]
- Schlütermann, D., Skowron, M. A., Berleth, N., Böhler, P., Deitersen, J., Stuhldreier, F., Wallot-Hieke, N., Wu, W., Peter, C., Hoffmann, M. J., Niegisch, G., & Stork, B. Targeting urothelial carcinoma cells by combining cisplatin with a specific inhibitor of the autophagy-inducing class III PtdIns3K complex. Urologic oncology 2018. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



