Submitted:
03 March 2026
Posted:
04 March 2026
You are already at the latest version
Abstract
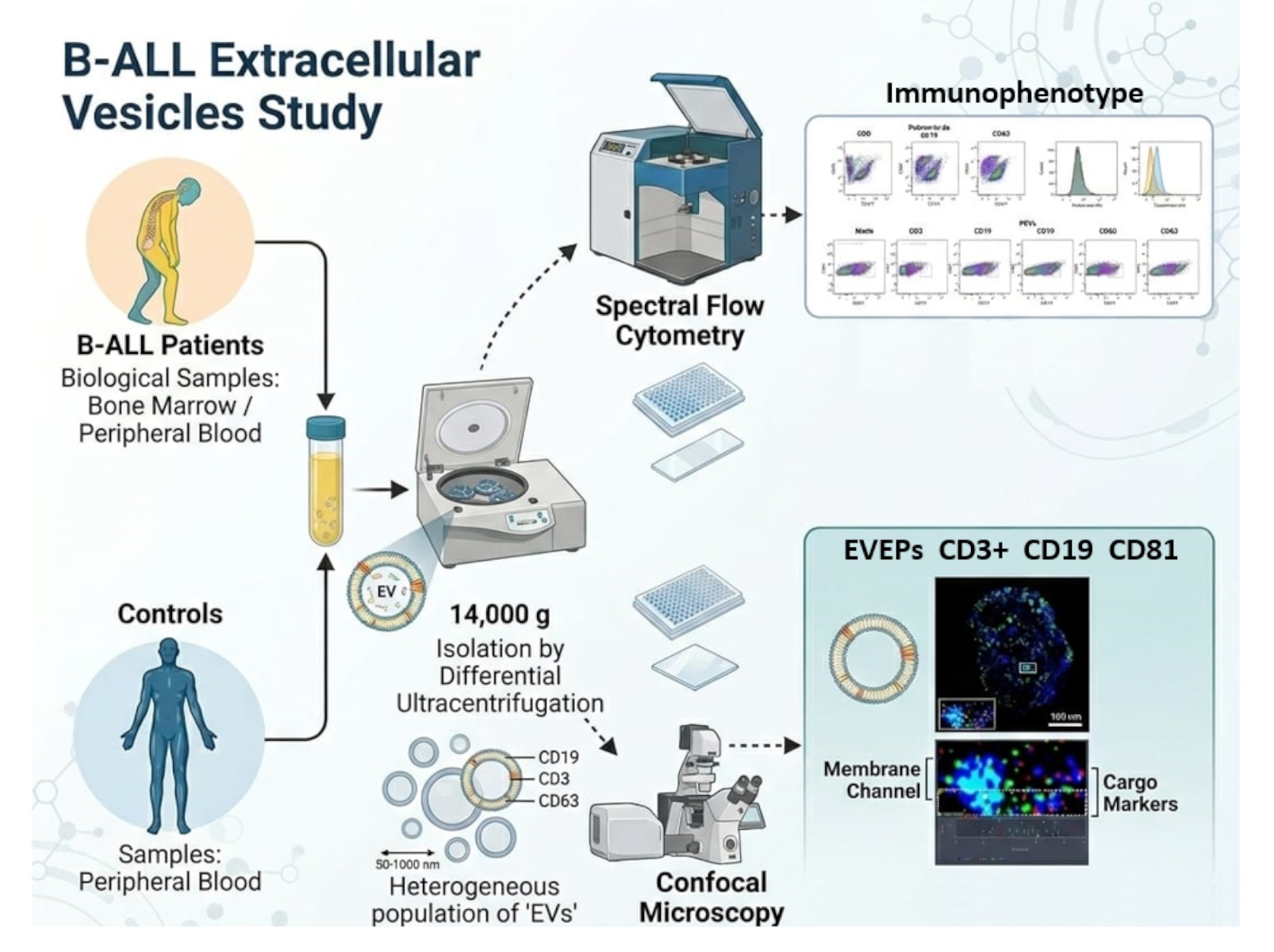
Keywords:
1. Introduction
2. Materials and Methods
2.1. Patient and Control Samples
2.2. Isolation of EVEPs
2.3. Immunophenotype of EVEPs
2.4. Confocal Microscopy of EVEPs
2.5. Statistical Analysis
3. Results
3.1. Clinical Information
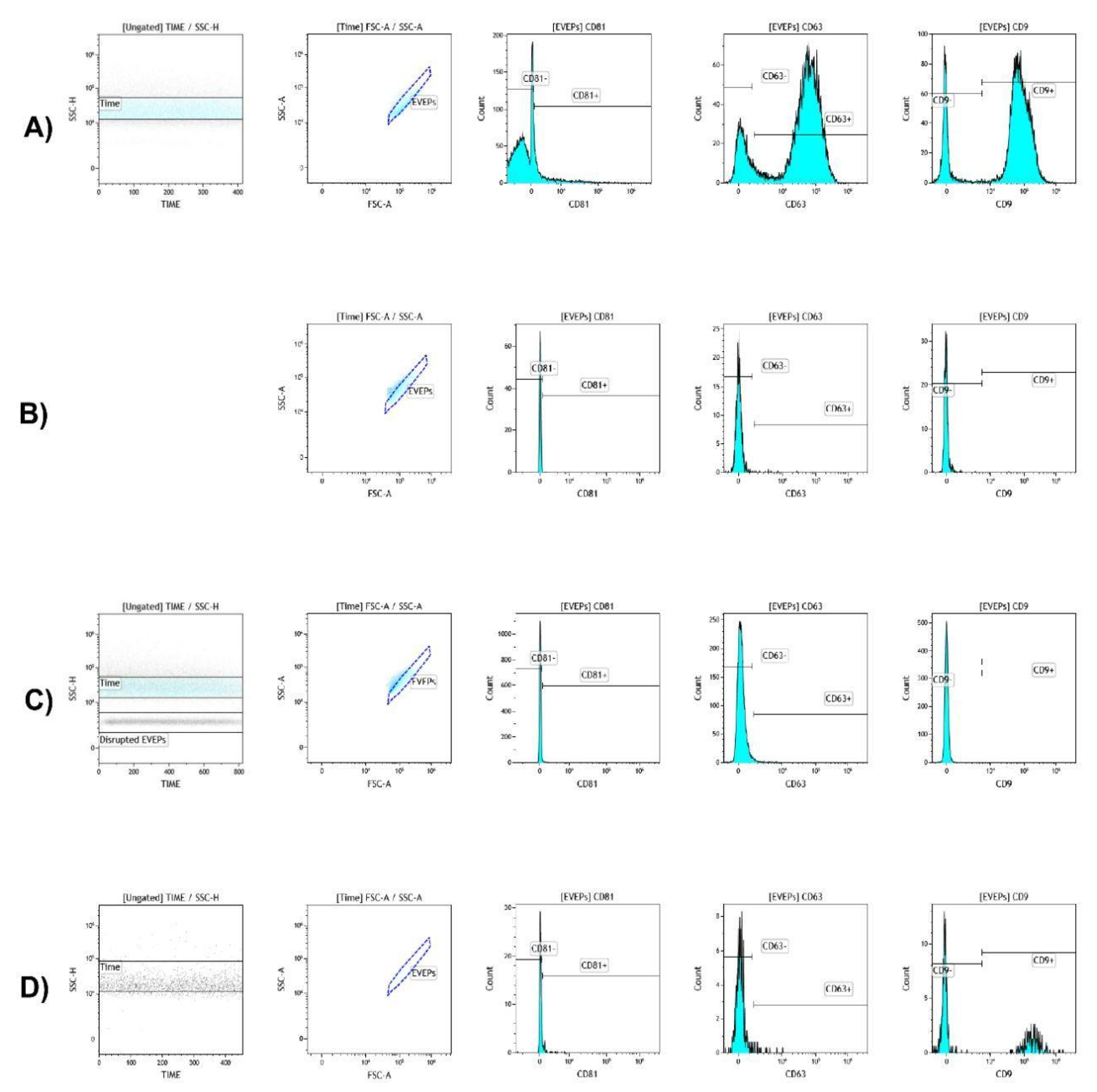
3.2. Standardization of EVEPs Analysis by Flow Cytometry
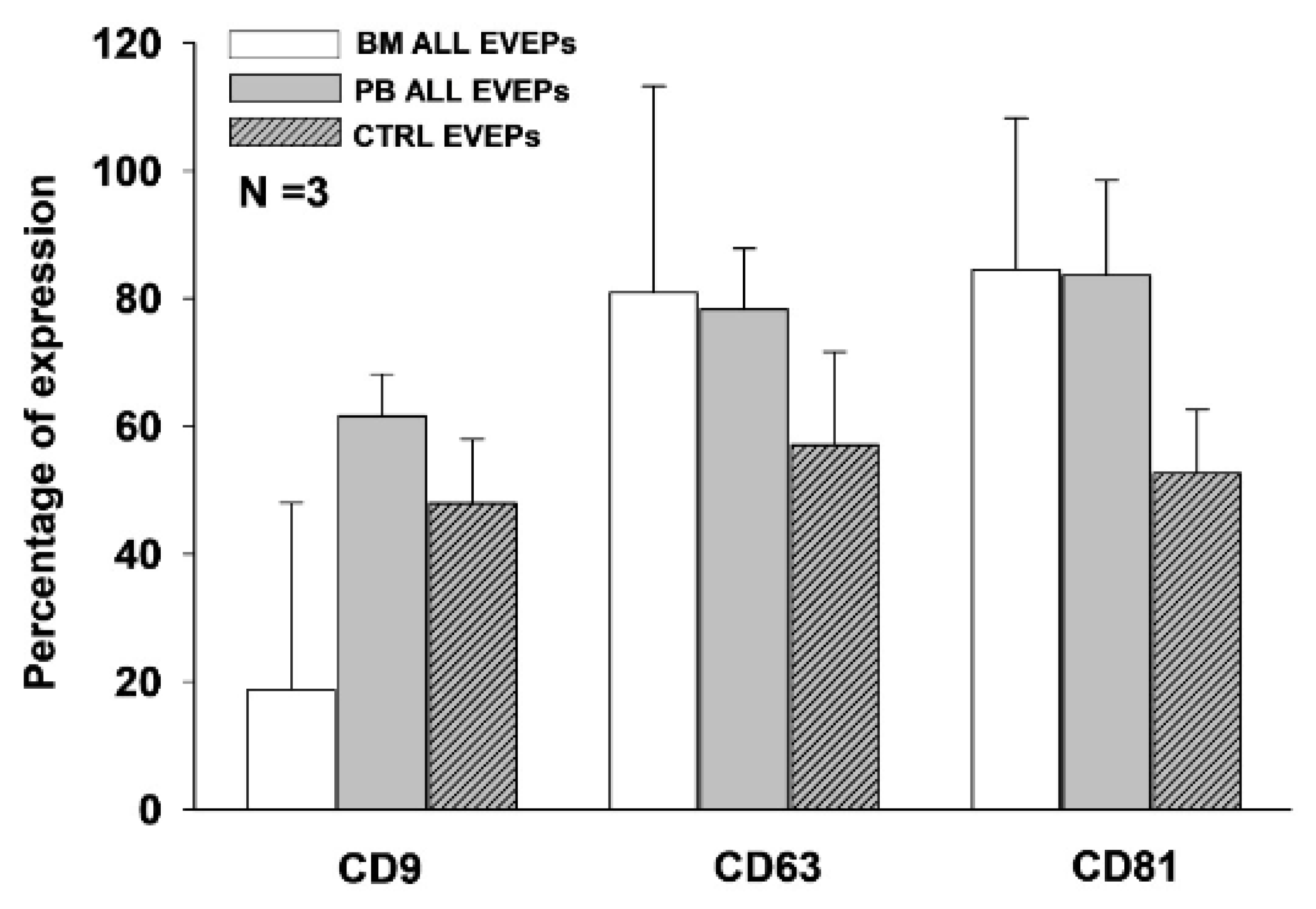
3.3. Comparison of the Expression of Tetraspanins Associated with Extracellular Vesicles Between Patients with B-ALL and Controls
3.4. Immunophenotype of EVEPs and Comparison with the Clinical Immunophenotype
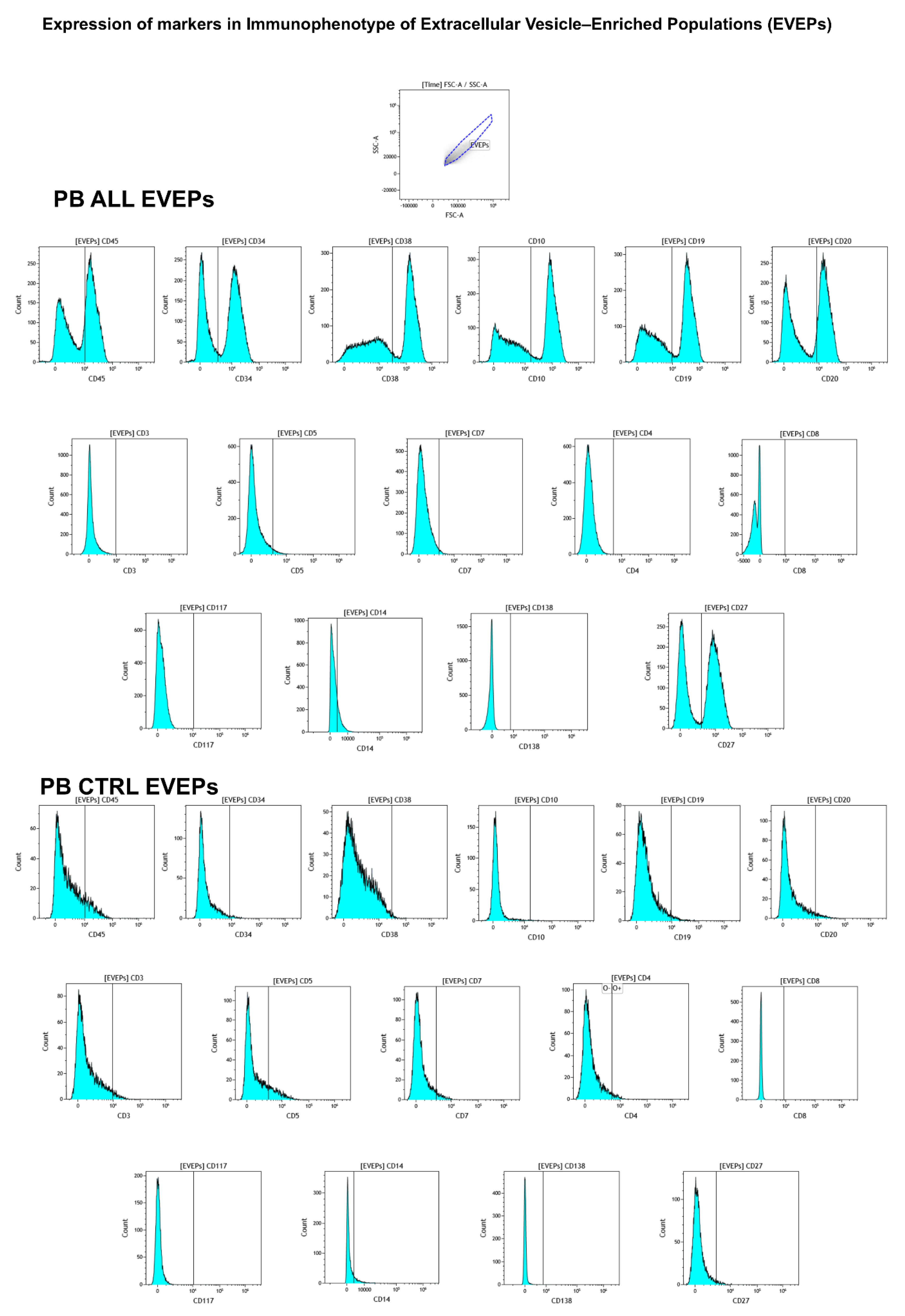
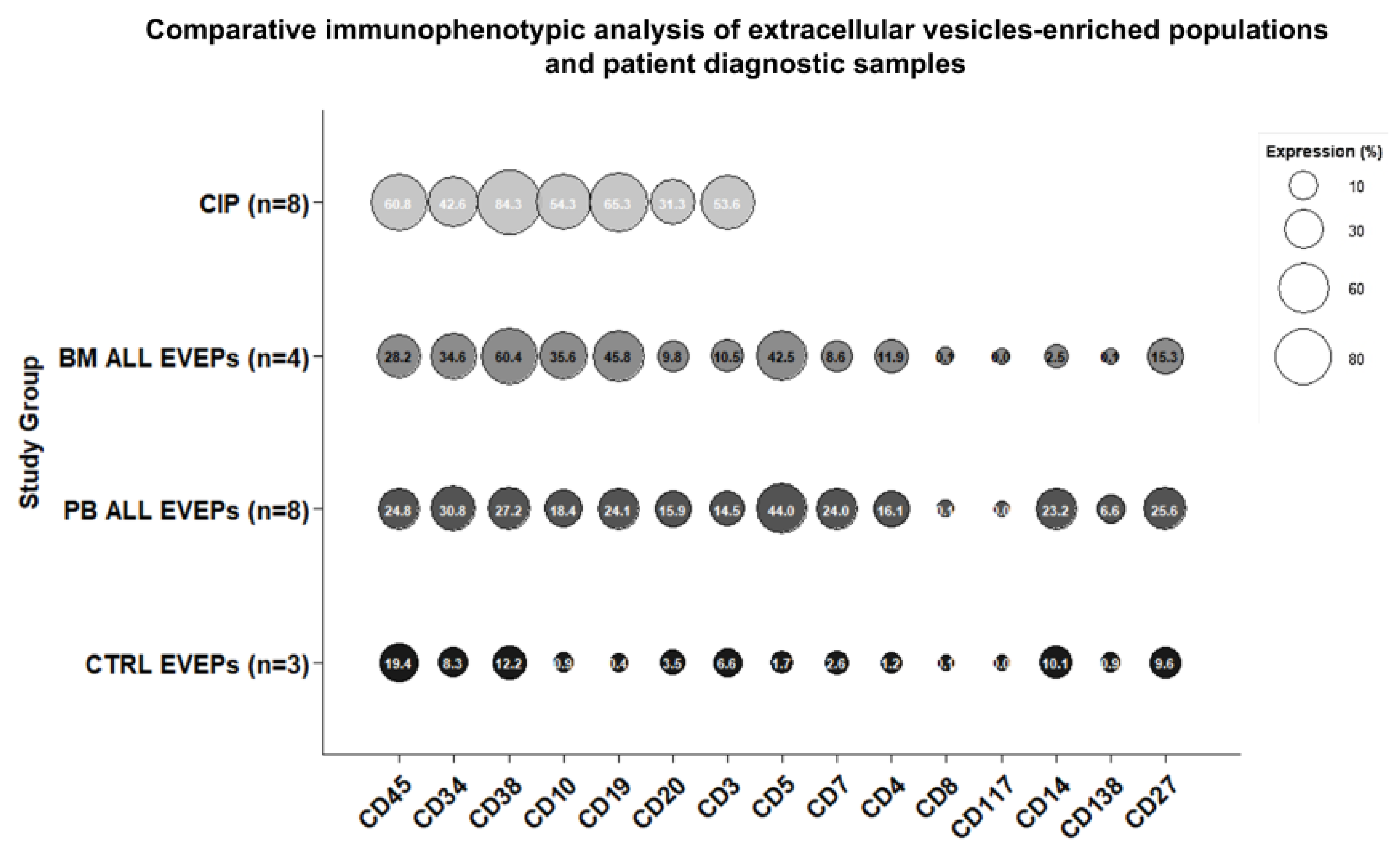
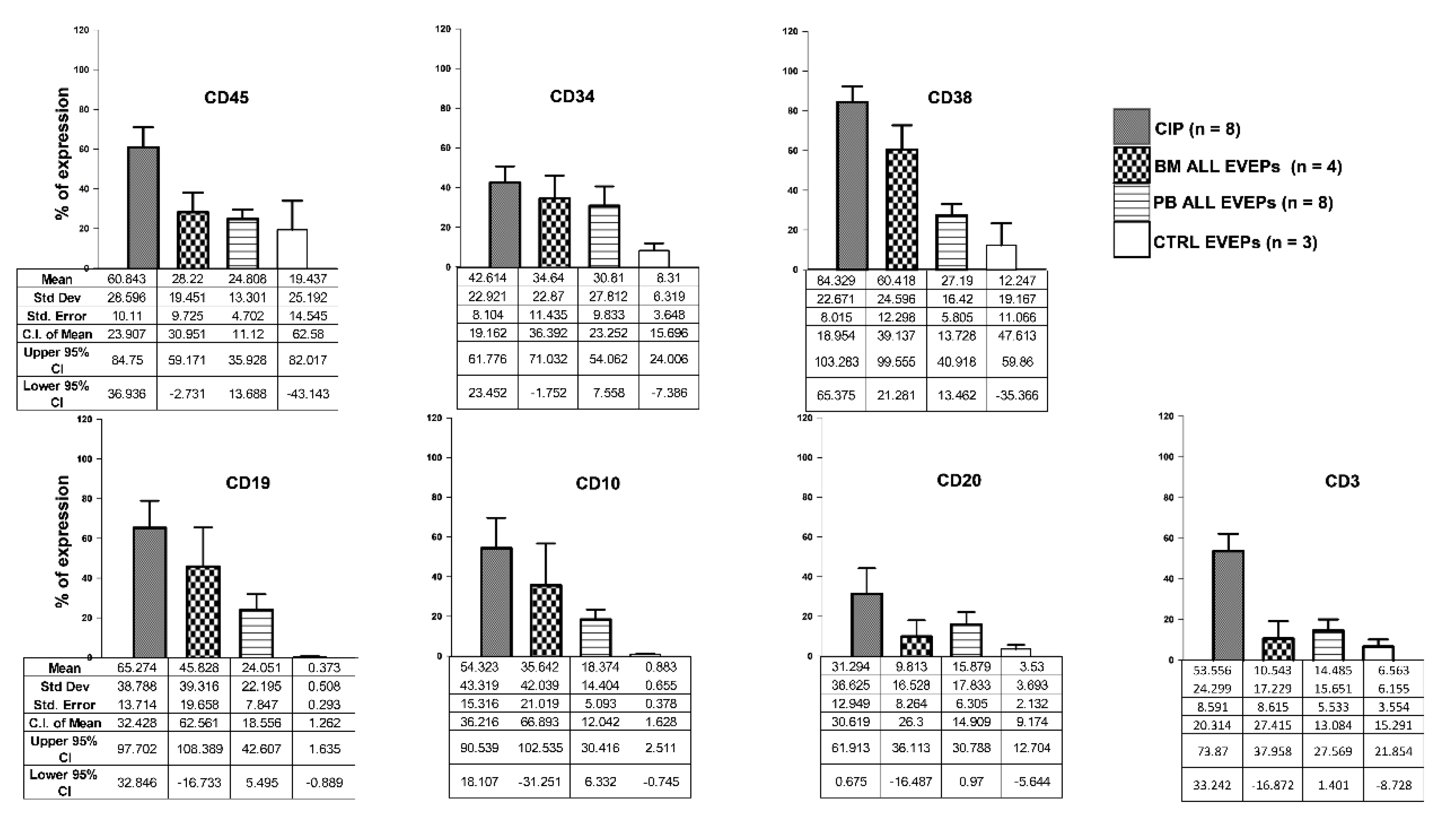
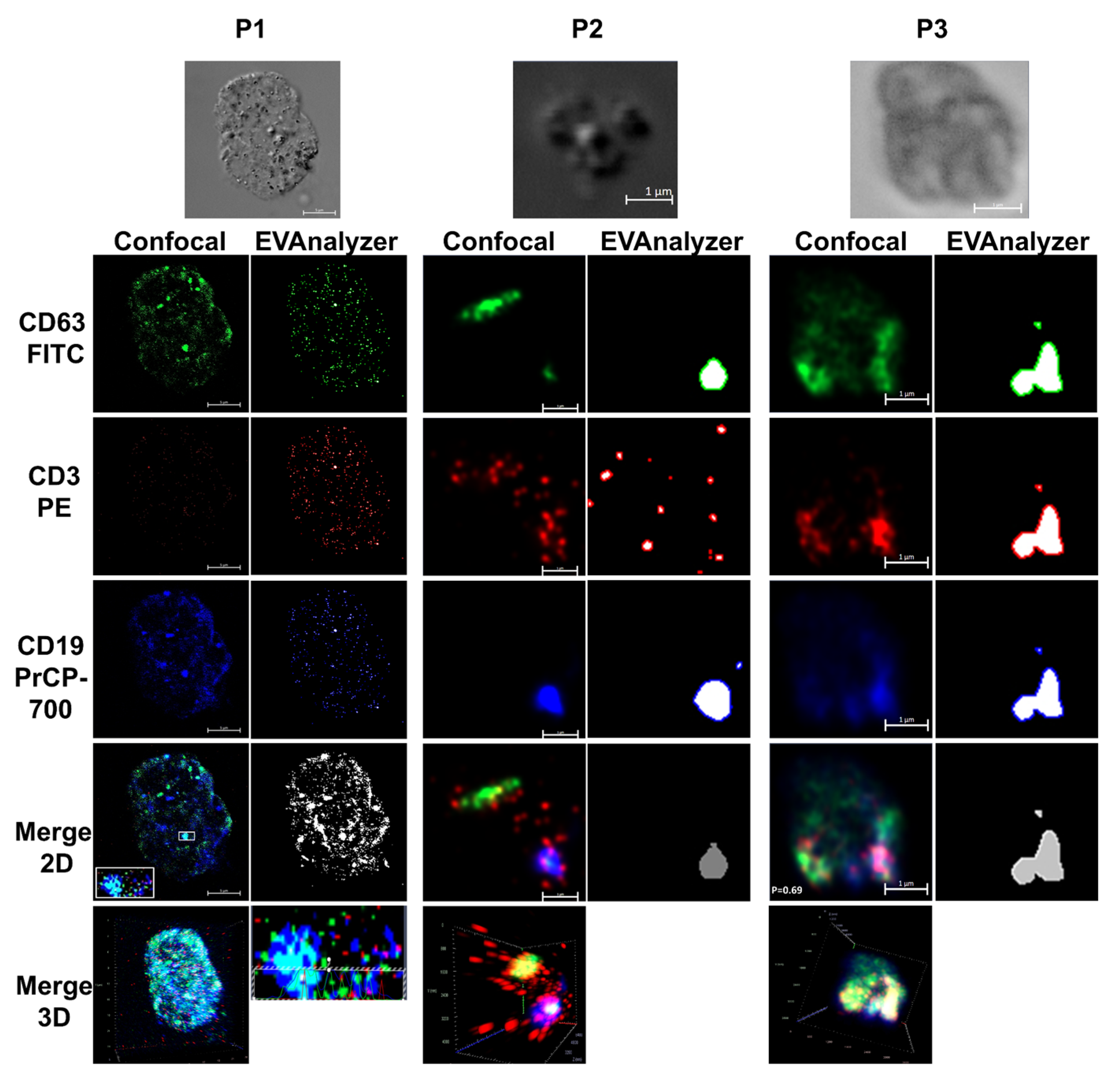
3.5. Identification of CD3⁺ CD19⁺ CD63⁺ Aggregates Using Confocal Microscopy
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ALL | Acute lymphoblastic leukemia |
| ASF | Signal amplification factor |
| B-ALL | B-lineage acute lymphoblastic leukemia |
| BM | Bone marrow |
| BCR-ABL1 | Breakpoint cluster region–Abelson murine leukemia 1 |
| BSA | Bovine serum albumin |
| CD | Cluster of differentiation |
| CIP | Clinical immunophenotype |
| EVEPs | Extracellular vesicle–enriched populations |
| EVs | Extracellular vesicles |
| FSC | Forward scatter |
| LDL | Low-density lipoprotein |
| MHC | Major histocompatibility complex |
| MISEV | Minimal Information for Studies of Extracellular Vesicles |
| PB | Peripheral blood |
| SSC | Side scatter |
| SSC-H | Side scatter height |
| TCR | T-cell receptor |
| VLDL | Very low-density lipoprotein |
References
- Sedgwick, A. E.; D’Souza-Schorey, C. The Biology of Extracellular Microvesicles. Traffic 2018, 19, 319–327. [Google Scholar] [CrossRef]
- Nguyen, P. H. D.; Le, A. H.; Pek, J. S. Q.; Pham, T. T.; Jayasinghe, M. K.; Do, D. V.; Phung, C. D.; Le, M. T. N. Extracellular Vesicles and Lipoproteins - Smart Messengers of Blood Cells in the Circulation. Journal of extracellular biology 2022, 1, e49. [Google Scholar] [CrossRef]
- Doyle, L. M.; Wang, M. Z. Overview of Extracellular Vesicles, Their Origin, Composition, Purpose, and Methods for Exosome Isolation and Analysis. Cells 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Meldolesi, J. Exosomes and Ectosomes in Intercellular Communication. Current Biology 2018, 28, R435–R444. [Google Scholar] [CrossRef] [PubMed]
- Cocucci, E.; Meldolesi, J. Ectosomes and Exosomes: Shedding the Confusion between Extracellular Vesicles. Trends Cell Biol. 2015, 25, 364–372. [Google Scholar] [CrossRef] [PubMed]
- Kalluri, R.; McAndrews, K. M. The Role of Extracellular Vesicles in Cancer. Cell 2023, 186, 1610–1626. [Google Scholar] [CrossRef]
- Rezaie, J.; Ahmadi, M.; Ravanbakhsh, R.; Mojarad, B.; Mahbubfam, S.; Shaban, S. A.; Shadi, K.; Berenjabad, N. J.; Etemadi, T. Tumor-Derived Extracellular Vesicles: The Metastatic Organotropism Drivers. Life Sci. 2022, 289, 120216. [Google Scholar] [CrossRef]
- Jeppesen, D. K.; Zhang, Q.; Franklin, J. L.; Coffey, R. J. Extracellular Vesicles and Nanoparticles: Emerging Complexities. Trends Cell Biol. 2023, 33, 667–681. [Google Scholar] [CrossRef]
- Ratajczak, M. Z.; Ratajczak, J. Leukemogenesis Occurs in a Microenvironment Enriched by Extracellular Microvesicles/Exosomes: Recent Discoveries and Questions to Be Answered. Leukemia 2024, 38, 692–698. [Google Scholar] [CrossRef]
- Samii, A.; Razmkhah, F. Transformation of Hematopoietic Stem and Progenitor Cells by Leukemia Extracellular Vesicles: A Step Toward Leukemogenesis. Stem Cell Rev. Rep. 2020, 16, 1081–1091. [Google Scholar] [CrossRef]
- Yamashita, M.; Dellorusso, P. V.; Olson, O. C.; Passegué, E. Dysregulated Haematopoietic Stem Cell Behaviour in Myeloid Leukaemogenesis. Nat. Rev. Cancer 2020, 20, 365–382. [Google Scholar] [CrossRef] [PubMed]
- Johnsen, K. B.; Gudbergsson, J. M.; Andresen, T. L.; Simonsen, J. B. What Is the Blood Concentration of Extracellular Vesicles? Implications for the Use of Extracellular Vesicles as Blood-Borne Biomarkers of Cancer. Biochimica et Biophysica Acta (BBA) - Reviews on Cancer 2019, 1871, 109–116. [Google Scholar] [CrossRef] [PubMed]
- Lucien, F.; Gustafson, D.; Lenassi, M.; Li, B.; Teske, J. J.; Boilard, E.; von Hohenberg, K. C.; Falcón-Perez, J. M.; Gualerzi, A.; Reale, A.; et al. MIBlood-EV: Minimal Information to Enhance the Quality and Reproducibility of Blood Extracellular Vesicle Research. J. Extracell. Vesicles 2023, 12. [Google Scholar] [CrossRef] [PubMed]
- Vu, L. T.; Peng, B.; Zhang, D. X.; Ma, V.; Mathey-Andrews, C. A.; Lam, C. K.; Kiomourtzis, T.; Jin, J.; McReynolds, L.; Huang, L.; et al. Tumor-secreted Extracellular Vesicles Promote the Activation of Cancer-associated Fibroblasts via the Transfer of MicroRNA-125b. J. Extracell. Vesicles 2019, 8. [Google Scholar] [CrossRef]
- Bebelman, M. P.; Janssen, E.; Pegtel, D. M.; Crudden, C. The Forces Driving Cancer Extracellular Vesicle Secretion. Neoplasia 2021, 23, 149–157. [Google Scholar] [CrossRef]
- Dilsiz, N. Role of Exosomes and Exosomal MicroRNAs in Cancer. Future Sci. OA 2020, 6. [Google Scholar] [CrossRef]
- Urabe, F.; Kosaka, N.; Ito, K.; Kimura, T.; Egawa, S.; Ochiya, T. Extracellular Vesicles as Biomarkers and Therapeutic Targets for Cancer. American Journal of Physiology-Cell Physiology 2020, 318, C29–C39. [Google Scholar] [CrossRef]
- Costa-Silva, B.; Aiello, N. M.; Ocean, A. J.; Singh, S.; Zhang, H.; Thakur, B. K.; Becker, A.; Hoshino, A.; Mark, M. T.; Molina, H.; et al. Pancreatic Cancer Exosomes Initiate Pre-Metastatic Niche Formation in the Liver. Nat. Cell Biol. 2015, 17, 816–826. [Google Scholar] [CrossRef]
- Duffield, A. S.; Mullighan, C. G.; Borowitz, M. J. International Consensus Classification of Acute Lymphoblastic Leukemia/Lymphoma. Virchows Archiv 2023, 482, 11–26. [Google Scholar] [CrossRef]
- Arber, D. A.; Orazi, A.; Hasserjian, R. P.; Borowitz, M. J.; Calvo, K. R.; Kvasnicka, H. M.; Wang, S. A.; Bagg, A.; Barbui, T.; Branford, S.; et al. International Consensus Classification of Myeloid Neoplasms and Acute Leukemias: Integrating Morphologic, Clinical, and Genomic Data. Blood 2022, 140, 1200–1228. [Google Scholar] [CrossRef]
- Harris, M. H.; Czuchlewski, D. R.; Arber, D. A.; Czader, M. Genetic Testing in the Diagnosis and Biology of Acute Leukemia. Am. J. Clin. Pathol. 2019, 152, 322–346. [Google Scholar] [CrossRef] [PubMed]
- Estey, E. H. Acute Myeloid Leukemia: 2019 Update on Risk-Stratification and Management. Am. J. Hematol. 2018, 93, 1267–1291. [Google Scholar] [CrossRef] [PubMed]
- De Kouchkovsky, I.; Abdul-Hay, M. Acute Myeloid Leukemia: A Comprehensive Review and 2016 Update. Blood Cancer J. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Malard, F.; Mohty, M. Acute Lymphoblastic Leukaemia. The Lancet 2020, 395, 1146–1162. [Google Scholar] [CrossRef] [PubMed]
- Polyatskin, I. L.; Artemyeva, A. S.; Krivolapov, Yu. A. Revised WHO Classification of Tumors of Hematopoietic and Lymphoid Tissues, 2017 (4th Edition):Lymphoid Tumors. Arkh. Patol. 2019, 81, 59. [Google Scholar] [CrossRef] [PubMed]
- Miljkovic-Licina, M.; Arraud, N.; Zahra, A. D.; Ropraz, P.; Matthes, T. Quantification and Phenotypic Characterization of Extracellular Vesicles from Patients with Acute Myeloid and B-Cell Lymphoblastic Leukemia. Cancers (Basel). 2021, 14, 56. [Google Scholar] [CrossRef]
- Kumar, M. A.; Baba, S. K.; Sadida, H. Q.; Marzooqi, S. Al.; Jerobin, J.; Altemani, F. H.; Algehainy, N.; Alanazi, M. A.; Abou-Samra, A.-B.; Kumar, R.; et al. Extracellular Vesicles as Tools and Targets in Therapy for Diseases. Signal Transduct. Target. Ther. 2024, 9, 27. [Google Scholar] [CrossRef]
- Longjohn, M. N.; Hudson, J.-A. B. J.; Peña-Castillo, L.; Cormier, R. P. J.; Hannay, B.; Chacko, S.; Lewis, S. M.; Moorehead, P. C.; Christian, S. L. Extracellular Vesicle Small RNA Cargo Discriminates Non-Cancer Donors from Pediatric B-Lymphoblastic Leukemia Patients. Front. Oncol. 2023, 13. [Google Scholar] [CrossRef]
- Trino, S.; Lamorte, D.; Caivano, A.; De Luca, L.; Sgambato, A.; Laurenzana, I. Clinical Relevance of Extracellular Vesicles in Hematological Neoplasms: From Liquid Biopsy to Cell Biopsy. Leukemia 2021, 35, 661–678. [Google Scholar] [CrossRef]
- Théry, C.; Witwer, K. W.; Aikawa, E.; Alcaraz, M. J.; Anderson, J. D.; Andriantsitohaina, R.; Antoniou, A.; Arab, T.; Archer, F.; Atkin-Smith, G. K.; et al. Minimal Information for Studies of Extracellular Vesicles 2018 (MISEV2018): A Position Statement of the International Society for Extracellular Vesicles and Update of the MISEV2014 Guidelines. J. Extracell. Vesicles 2018, 7. [Google Scholar] [CrossRef]
- Menck, K.; Bleckmann, A.; Schulz, M.; Ries, L.; Binder, C. Isolation and Characterization of Microvesicles from Peripheral Blood. Journal of Visualized Experiments 2017, No. 119. [Google Scholar] [CrossRef]
- Parolini, I.; Federici, C.; Raggi, C.; Lugini, L.; Palleschi, S.; De Milito, A.; Coscia, C.; Iessi, E.; Logozzi, M.; Molinari, A.; et al. Microenvironmental PH Is a Key Factor for Exosome Traffic in Tumor Cells. Journal of Biological Chemistry 2009, 284, 34211–34222. [Google Scholar] [CrossRef] [PubMed]
- Welsh, J. A.; Goberdhan, D. C. I.; O’Driscoll, L.; Buzas, E. I.; Blenkiron, C.; Bussolati, B.; Cai, H.; Di Vizio, D.; Driedonks, T. A. P.; Erdbrügger, U.; et al. Minimal Information for Studies of Extracellular Vesicles (MISEV2023): From Basic to Advanced Approaches. J. Extracell. Vesicles 2024, 13. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Ye, S.; Guo, X.; Li, W.; Xia, Y.; Wen, X.; Yu, J.; Jia, Y.; Liu, X.; Guo, Y.; et al. Discovery and Characteristics of B Cell-like T Cells: A Potential Novel Tumor Immune Marker? Immunol. Lett. 2020, 220, 44–50. [Google Scholar] [CrossRef]
- Sódar, B. W.; Kittel, Á.; Pálóczi, K.; Vukman, K. V; Osteikoetxea, X.; Szabó-Taylor, K.; Németh, A.; Sperlágh, B.; Baranyai, T.; Giricz, Z.; et al. Low-Density Lipoprotein Mimics Blood Plasma-Derived Exosomes and Microvesicles during Isolation and Detection. Sci. Rep. 2016, 6, 24316. [Google Scholar] [CrossRef]
- Jimenez, D. E.; Tahir, M.; Faheem, M.; Alves, W. B. dos S.; Correa, B. de L.; de Andrade, G. R.; Larsen, M. R.; de Oliveira, G. P.; Pereira, R. W. Comparison of Four Purification Methods on Serum Extracellular Vesicle Recovery, Size Distribution, and Proteomics. Proteomes 2023, 11, 23. [Google Scholar] [CrossRef]
- Japp, A. S.; Meng, W.; Rosenfeld, A. M.; Perry, D. J.; Thirawatananond, P.; Bacher, R. L.; Liu, C.; Gardner, J. S.; Atkinson, M. A.; Kaestner, K. H.; et al. TCR+/BCR+ Dual-Expressing Cells and Their Associated Public BCR Clonotype Are Not Enriched in Type 1 Diabetes. Cell 2021, 184, 827–839.e14. [Google Scholar] [CrossRef]
- zhang, Y.; Guo, C.; Zhou, Y.; Zhang, W.; Zhu, Z.; Wang, W.; Wan, Y. A Biphenotypic Lymphocyte Subset Displays Both T- and B-Cell Functionalities. Commun. Biol. 2024, 7, 28. [Google Scholar] [CrossRef] [PubMed]
- Buzás, E. I.; Tóth, E. Á.; Sódar, B. W.; Szabó-Taylor, K. É. Molecular Interactions at the Surface of Extracellular Vesicles. Semin. Immunopathol. 2018, 40, 453–464. [Google Scholar] [CrossRef] [PubMed]
- Tóth, E. Á.; Turiák, L.; Visnovitz, T.; Cserép, C.; Mázló, A.; Sódar, B. W.; Försönits, A. I.; Petővári, G.; Sebestyén, A.; Komlósi, Z.; et al. Formation of a Protein Corona on the Surface of Extracellular Vesicles in Blood Plasma. J. Extracell. Vesicles 2021, 10, e12140. [Google Scholar] [CrossRef]
- Alexander, T. B.; Gu, Z.; Iacobucci, I.; Dickerson, K.; Choi, J. K.; Xu, B.; Payne-Turner, D.; Yoshihara, H.; Loh, M. L.; Horan, J.; et al. The Genetic Basis and Cell of Origin of Mixed Phenotype Acute Leukaemia. Nature 2018, 562, 373–379. [Google Scholar] [CrossRef] [PubMed]
- Johnson, S. M.; Dempsey, C.; Parker, C.; Mironov, A.; Bradley, H.; Saha, V. Acute Lymphoblastic Leukaemia Cells Produce Large Extracellular Vesicles Containing Organelles and an Active Cytoskeleton. J. Extracell. Vesicles 2017, 6. [Google Scholar] [CrossRef] [PubMed]
- Al-Nedawi, K.; Meehan, B.; Micallef, J.; Lhotak, V.; May, L.; Guha, A.; Rak, J. Intercellular Transfer of the Oncogenic Receptor EGFRvIII by Microvesicles Derived from Tumour Cells. Nat. Cell Biol. 2008, 10, 619–624. [Google Scholar] [CrossRef] [PubMed]
- Abdulmawjood, B.; Roma-Rodrigues, C.; Fernandes, A. R.; Baptista, P. V. Liquid Biopsies in Myeloid Malignancies. Cancer Drug Resist. 2019, 2, 1044–1061. [Google Scholar] [CrossRef]
- Arber, DA; Orazi, A; Hasserjian, R; et al. The 2016 Revision to the World Health Organization Classification of Myeloid Neoplasms and Acute Leukemia. Blood;Blood 2016, 127((20)), 2391-2405, Blood, 2016, 128 (3), 462–463. [Google Scholar] [CrossRef]
- Sierra-López, F.; Iglesias-Vazquez, V.; Baylon-Pacheco, L.; Ríos-Castro, E.; Osorio-Trujillo, J. C.; Lagunes-Guillén, A.; Chávez-Munguía, B.; Hernández, S. B.; Acosta-Altamirano, G.; Talamás-Rohana, P.; et al. A Fraction of Escherichia Coli Bacteria Induces an Increase in the Secretion of Extracellular Vesicle Polydispersity in Macrophages: Possible Involvement of Secreted EVs in the Diagnosis of COVID-19 with Bacterial Coinfections. Int. J. Mol. Sci. 2025, 26, 3741. [Google Scholar] [CrossRef]
- Karami Fath, M.; Azami, J.; Jaafari, N.; Akbari Oryani, M.; Jafari, N.; Karim poor, A.; Azargoonjahromi, A.; Nabi-Afjadi, M.; Payandeh, Z.; Zalpoor, H.; et al. Exosome Application in Treatment and Diagnosis of B-Cell Disorders: Leukemias, Multiple Sclerosis, and Arthritis Rheumatoid. Cell. Mol. Biol. Lett. 2022, 27, 74. [Google Scholar] [CrossRef]
- Ratajczak, J.; Wysoczynski, M.; Hayek, F.; Janowska-Wieczorek, A.; Ratajczak, M. Z. Membrane-Derived Microvesicles: Important and Underappreciated Mediators of Cell-to-Cell Communication. Leukemia 2006, 20, 1487–1495. [Google Scholar] [CrossRef]
- Caivano, A.; Laurenzana, I.; De Luca, L.; La Rocca, F.; Simeon, V.; Trino, S.; D’Auria, F.; Traficante, A.; Maietti, M.; Izzo, T.; et al. High Serum Levels of Extracellular Vesicles Expressing Malignancy-Related Markers Are Released in Patients with Various Types of Hematological Neoplastic Disorders. Tumor Biology 2015, 36, 9739–9752. [Google Scholar] [CrossRef]
- Yang, H.-Q.; Qiu, F.-Q.; Jin, K. E.; Jiang, N.-G.; Zhang, L. I. High Plasma Levels of Oxidatively Modified Low-Density Lipoproteins Are Associated with the Suppressed Expression of Immunomodulatory Molecules in Patients with Hematological Malignancies. Exp. Ther. Med. 2015, 9, 2394–2400. [Google Scholar] [CrossRef]
- Kwaan, H.; Magalhães Rego, E. Role of Microparticles in the Hemostatic Dysfunction in Acute Promyelocytic Leukemia. Semin. Thromb. Hemost. 2010, 36, 917–924. [Google Scholar] [CrossRef]
- Lozano-Andrés, E.; Enciso-Martinez, A.; Gijsbers, A.; Libregts, S. F. W. M.; Pinheiro, C.; Van Niel, G.; Hendrix, A.; Peters, P. J.; Otto, C.; Arkesteijn, G. J. A.; et al. Physical Association of Low Density Lipoprotein Particles and Extracellular Vesicles Unveiled by Single Particle Analysis; 1 September 2022. [Google Scholar] [CrossRef]
- Morel, S.; Leahy, J.; Fournier, M.; Lamarche, B.; Garofalo, C.; Grimard, G.; Poulain, F.; Delvin, E.; Laverdière, C.; Krajinovic, M.; et al. Lipid and Lipoprotein Abnormalities in Acute Lymphoblastic Leukemia Survivors. J. Lipid Res. 2017, 58, 982–993. [Google Scholar] [CrossRef]
- Malhotra, J.; Tonorezos, E. S.; Rozenberg, M.; Vega, G. L.; Sklar, C. A.; Chou, J.; Moskowitz, C. S.; Eshelman-Kent, D. A.; Janiszewski, P.; Ross, R.; et al. Atherogenic Low Density Lipoprotein Phenotype in Long-Term Survivors of Childhood Acute Lymphoblastic Leukemia. J. Lipid Res. 2012, 53, 2747–2754. [Google Scholar] [CrossRef] [PubMed]
- Welsh, J. A.; Van Der Pol, E.; Arkesteijn, G. J. A.; Bremer, M.; Brisson, A.; Coumans, F.; Dignat-George, F.; Duggan, E.; Ghiran, I.; Giebel, B.; et al. MIFlowCyt-EV: A Framework for Standardized Reporting of Extracellular Vesicle Flow Cytometry Experiments. J. Extracell. Vesicles 2020, 9. [Google Scholar] [CrossRef]
- Li, L.; Görgens, A.; Mussack, V.; Pepeldjiyska, E.; Hartz, A. S.; Rank, A.; Schmohl, J.; Krämer, D.; El Andaloussi, S.; Pfaffl, M. W.; et al. Description and Optimization of a Multiplex Bead-Based Flow Cytometry Method (MBFCM) to Characterize Extracellular Vesicles in Serum Samples from Patients with Hematological Malignancies. Cancer Gene Ther. 2022, 29, 1600–1615. [Google Scholar] [CrossRef] [PubMed]
- Zhong, X.; Ma, H. Targeting CD38 for Acute Leukemia. Front. Oncol. 2022, 12. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Z.; Wu, D.; Lin, S.; Li, P. CD34 and CD38 Are Prognostic Biomarkers for Acute B Lymphoblastic Leukemia. Biomark. Res. 2016, 4, 23. [Google Scholar] [CrossRef]
- Eveillard, M.; Floc’h, V.; Robillard, N.; Debord, C.; Wuilleme, S.; Garand, R.; Rialland, F.; Thomas, C.; Peterlin, P.; Guillaume, T.; et al. CD38 Expression in B-Lineage Acute Lymphoblastic Leukemia, a Possible Target for Immunotherapy. Blood 2016, 128, 5268–5268. [Google Scholar] [CrossRef]
- Magalhães-Gama, F.; Malheiros Araújo Silvestrini, M.; Neves, J. C. F.; Araújo, N. D.; Alves-Hanna, F. S.; Kerr, M. W. A.; Carvalho, M. P. S. S.; Tarragô, A. M.; Soares Pontes, G.; Martins-Filho, O. A.; et al. Exploring Cell-Derived Extracellular Vesicles in Peripheral Blood and Bone Marrow of B-Cell Acute Lymphoblastic Leukemia Pediatric Patients: Proof-of-Concept Study. Front. Immunol. 2024, 15. [Google Scholar] [CrossRef]
- Auber, M.; Svenningsen, P. An Estimate of Extracellular Vesicle Secretion Rates of Human Blood Cells. Journal of Extracellular Biology 2022, 1. [Google Scholar] [CrossRef]
- Wahlgren, J.; Karlson, T. D. L.; Glader, P.; Telemo, E.; Valadi, H. Activated Human T Cells Secrete Exosomes That Participate in IL-2 Mediated Immune Response Signaling. PLoS One 2012, 7, e49723. [Google Scholar] [CrossRef]
- Griesel, L.; Kaleja, P.; Tholey, A.; Lettau, M.; Janssen, O. Comparative Analysis of Extracellular Vesicles from Cytotoxic CD8+ Aβ T Cells and Γδ T Cells. Cells 2024, 13, 1745. [Google Scholar] [CrossRef]
- Lu, C.-W.; Lo, Y.-H.; Chen, C.-H.; Lin, C.-Y.; Tsai, C.-H.; Chen, P.-J.; Yang, Y.-F.; Wang, C.-H.; Tan, C.-H.; Hou, M.-F.; et al. VLDL and LDL, but Not HDL, Promote Breast Cancer Cell Proliferation, Metastasis and Angiogenesis. Cancer Lett. 2017, 388, 130–138. [Google Scholar] [CrossRef] [PubMed]
- Oguro, H. The Roles of Cholesterol and Its Metabolites in Normal and Malignant Hematopoiesis. Front. Endocrinol. (Lausanne). 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Goossens, P.; Rodriguez-Vita, J.; Etzerodt, A.; Masse, M.; Rastoin, O.; Gouirand, V.; Ulas, T.; Papantonopoulou, O.; Van Eck, M.; Auphan-Anezin, N.; et al. Membrane Cholesterol Efflux Drives Tumor-Associated Macrophage Reprogramming and Tumor Progression. Cell Metab. 2019, 29, 1376–1389.e4. [Google Scholar] [CrossRef] [PubMed]
- Karmacharya, M.; Kumar, S.; Cho, Y.-K. Tuning the Extracellular Vesicles Membrane through Fusion for Biomedical Applications. J. Funct. Biomater. 2023, 14, 117. [Google Scholar] [CrossRef]
- Morandi, M. I.; Busko, P.; Ozer-Partuk, E.; Khan, S.; Zarfati, G.; Elbaz-Alon, Y.; Abou Karam, P.; Napso Shogan, T.; Ginini, L.; Gil, Z.; et al. Extracellular Vesicle Fusion Visualized by Cryo-Electron Microscopy. PNAS Nexus 2022, 1. [Google Scholar] [CrossRef]
- Li, Q.; Wang, J.; Zhang, M.; Tang, Y.; Lu, H. Discovery of CD3+ CD19 +cells, a Novel Lymphocyte Subset with a Potential Role in Human Immunodeficiency Virus- Mycobacterium Tuberculosis Coinfection, Using Mass Cytometry. Clin. Transl. Med. 2021, 11. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




