Submitted:
02 March 2026
Posted:
03 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
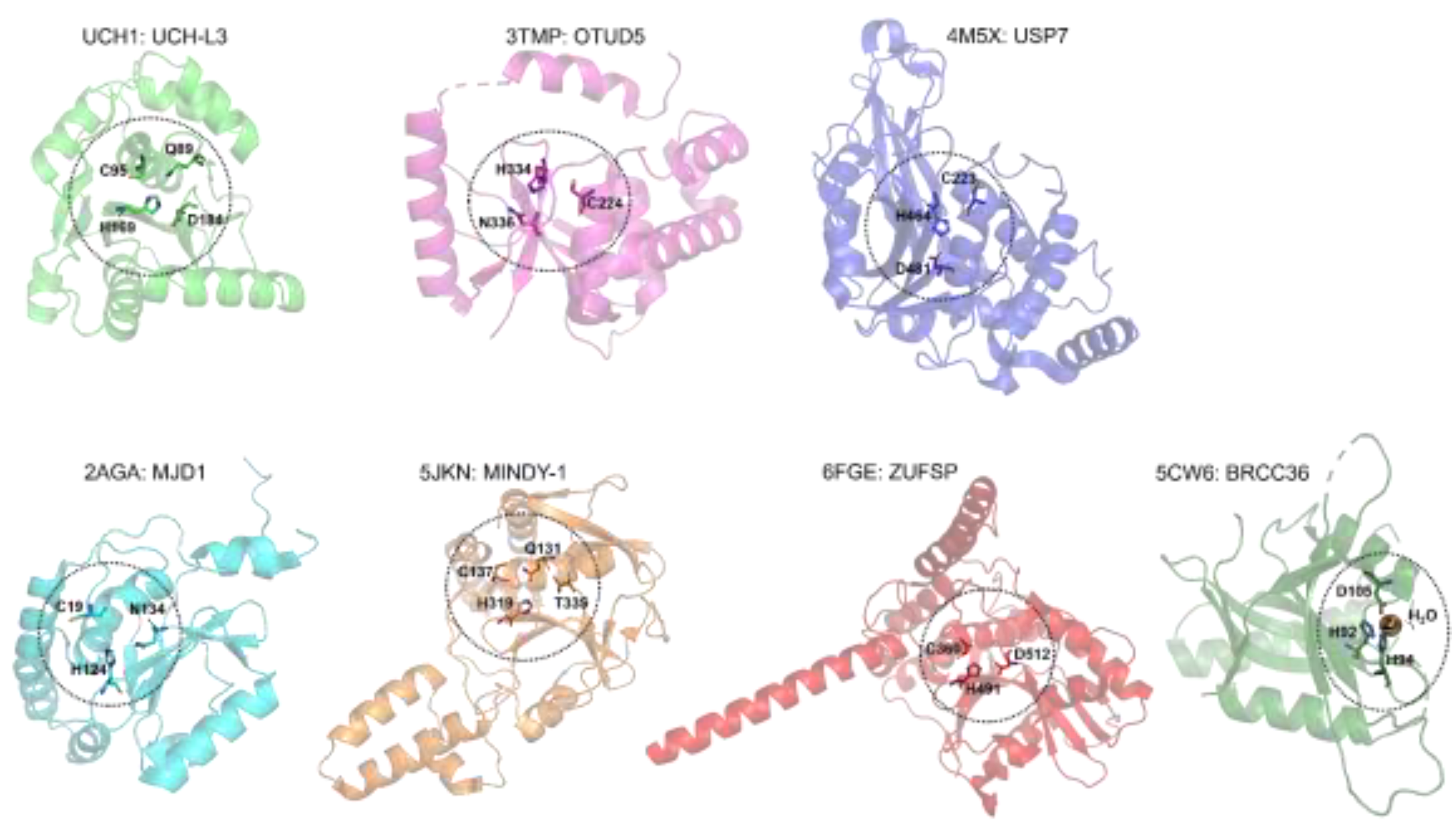
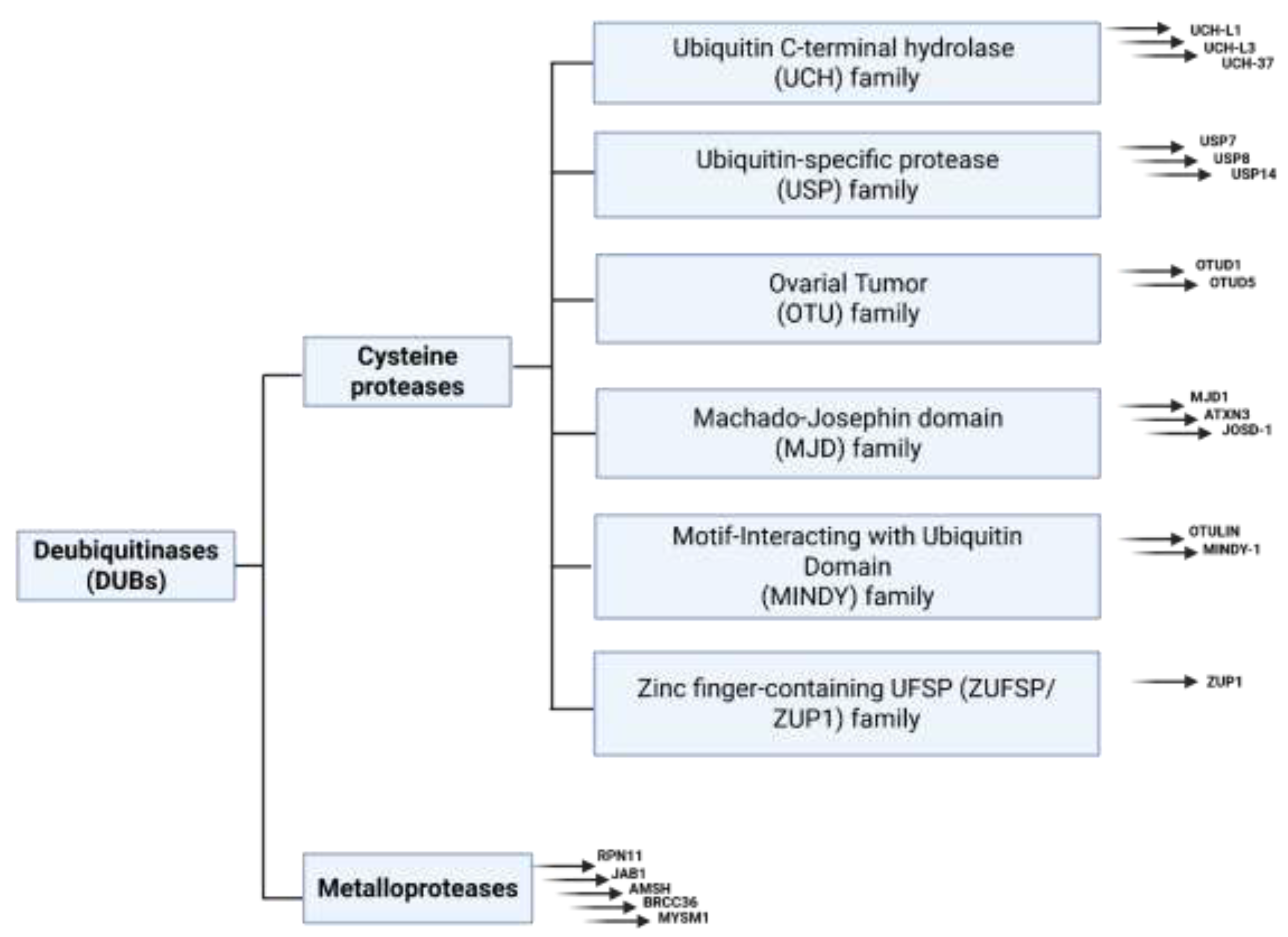
A. Cysteine Protease DUBs:
2. Physiological Functions of Deubiquitinating Enzymes (DUBs)
3. Dysregulation of DUBs in Human Diseases
4. Therapeutic Targeting of DUBs
5. Challenges in Targeting DUBs
Funding
Conflicts of Interest
References
- Hershko, A.; Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 1998, 67, 425–479. [Google Scholar] [CrossRef]
- Ciechanover, A. The ubiquitin-proteasome pathway: On protein death and cell life. EMBO J. 1998, 17, 7151–7160. [Google Scholar] [CrossRef]
- Goldberg, A.L. Protein degradation and protection against misfolded or damaged proteins. Nature 2003, 426, 895–899. [Google Scholar] [CrossRef]
- Pickart, C.M.; Eddins, M.J. Ubiquitin: Structures, functions, mechanisms. Biochim. Biophys. Acta Mol. Cell Res. 2004, 1695, 55–72. [Google Scholar] [CrossRef]
- Mani, A.; Gelmann, E.P. The ubiquitin-proteasome pathway and its role in cancer. J. Clin. Oncol. 2005, 23, 4776–4789. [Google Scholar] [CrossRef]
- Mukhopadhyay, D.; Riezman, H. Proteasome-independent functions of ubiquitin in endocytosis and signaling. Science 2007, 315, 201–205. [Google Scholar] [CrossRef]
- Rock, K.L.; Goldberg, A.L. Degradation of cell proteins and the generation of MHC class I-presented peptides. Annu. Rev. Immunol. 1999, 17, 739–779. [Google Scholar] [CrossRef]
- Kloetzel, P.M. Antigen processing by the proteasome. Nat. Rev. Mol. Cell Biol. 2001, 2, 179–187. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, K.D. Ubiquitin-dependent signaling: The role of ubiquitination in the response of cells to their environment. J. Nutr. 1999, 129, 1933–1936. [Google Scholar] [CrossRef]
- Schulman, B.A.; Harper, J.W. Ubiquitin-like protein activation by E1 enzymes: The apex for downstream signalling pathways. Nat. Rev. Mol. Cell Biol. 2009, 10, 319–331. [Google Scholar] [CrossRef] [PubMed]
- Ye, Y.; Rape, M. Building ubiquitin chains: E2 enzymes at work. Nat. Rev. Mol. Cell Biol. 2009, 10, 755–764. [Google Scholar] [CrossRef] [PubMed]
- Deshaies, R.J.; Joazeiro, C.A.P. RING domain E3 ubiquitin ligases. Annu. Rev. Biochem. 2009, 78, 399–434. [Google Scholar] [CrossRef] [PubMed]
- Varshavsky, A. The N-end rule pathway of protein degradation. Genes Cells 1997, 2, 13–28. [Google Scholar] [CrossRef] [PubMed]
- Glickman, M.H.; Ciechanover, A. The ubiquitin-proteasome proteolytic pathway: Destruction for the sake of construction. Physiol. Rev. 2002, 82, 373–428. [Google Scholar] [CrossRef]
- Finley, D. Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annu. Rev. Biochem. 2009, 78, 477–513. [Google Scholar] [CrossRef]
- Voges, D.; Zwickl, P.; Baumeister, W. The 26S proteasome: A molecular machine designed for controlled proteolysis. Annu. Rev. Biochem. 1999, 68, 1015–1068. [Google Scholar] [CrossRef]
- Tanaka, K. The proteasome: Overview of structure and functions. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 2009, 85, 12–36. [Google Scholar] [CrossRef] [PubMed]
- Ravid, T.; Hochstrasser, M. Diversity of degradation signals in the ubiquitin-proteasome system. Nat. Rev. Mol. Cell Biol. 2008, 9, 679–690. [Google Scholar] [CrossRef]
- Nalepa, G.; Rolfe, M.; Harper, J.W. Drug discovery in the ubiquitin-proteasome system. Nat. Rev. Drug Discov. 2006, 5, 596–613. [Google Scholar] [CrossRef]
- Hoeller, D.; Hecker, C.-M.; Dikic, I. Ubiquitin and ubiquitin-like proteins in cancer pathogenesis. Nat. Rev. Cancer 2006, 6, 776–788. [Google Scholar] [CrossRef]
- Bedford, L.; Lowe, J.; Dick, L.R.; Mayer, R.J.; Brownell, J.E. Ubiquitin-like protein conjugation and the ubiquitin-proteasome system as drug targets. Nat. Rev. Drug Discov. 2011, 10, 29–46. [Google Scholar] [CrossRef]
- Rubinsztein, D.C. The roles of intracellular protein-degradation pathways in neurodegeneration. Nature 2006, 443, 780–786. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, A.L.; Ciechanover, A. Targeting proteins for destruction by the ubiquitin system: Implications for human pathobiology. Annu. Rev. Pharmacol. Toxicol. 2009, 49, 73–96. [Google Scholar] [CrossRef]
- Amerik, A.Y.; Hochstrasser, M. Mechanism and function of deubiquitinating enzymes. Biochim. Biophys. Acta 2004, 1695, 189–207. [Google Scholar] [CrossRef] [PubMed]
- Kabra, A.; Li, Y. Conformational Dynamics of Deubiquitinase A and Functional Implications. Biochemistry 2021, 60, 201–209. [Google Scholar] [CrossRef]
- Wilkinson, K.D. DUBs at a glance. J. Cell Sci. 2009, 122, 2325–2329. [Google Scholar] [CrossRef] [PubMed]
- Komander, D.; Clague, M.J.; Urbé, S. Breaking the chains: Structure and function of the deubiquitinases. Nat. Rev. Mol. Cell Biol. 2009, 10, 550–563. [Google Scholar] [CrossRef]
- Kabra, A.; et al. Structural characterization of peptidyl-tRNA hydrolase from Mycobacterium smegmatis by NMR spectroscopy. Biochim. Biophys. Acta 2016, 1864, 1304–1314. [Google Scholar] [CrossRef]
- Kabra, A.; et al. Unraveling the stereochemical and dynamic aspects of the catalytic site of bacterial peptidyl-tRNA hydrolase. RNA 2017, 23, 202–216. [Google Scholar] [CrossRef]
- Shahid, S.; et al. Role of methionine 71 in substrate recognition and structural integrity of bacterial peptidyl-tRNA hydrolase. Biochim. Biophys. Acta Proteins Proteom. 2018, 1866, 865–874. [Google Scholar] [CrossRef]
- Mundra, S.; Kabra, A. Unveiling the Druggable Landscape of Bacterial Peptidyl tRNA Hydrolase: Insights into Structure, Function, and Therapeutic Potential. Biomolecules 2024, 14. [Google Scholar] [CrossRef]
- Snyder, N.A.; Silva, G.M. Deubiquitinating enzymes (DUBs): Regulation, homeostasis, and oxidative stress response. J. Biol. Chem. 2021, 297, 101077. [Google Scholar] [CrossRef]
- Lange, S.M.; Armstrong, L.A.; Kulathu, Y. Deubiquitinases: From mechanisms to their inhibition by small molecules. Mol. Cell 2022, 82, 15–29. [Google Scholar] [CrossRef]
- Estavoyer, B.; et al. Mechanisms orchestrating the enzymatic activity and cellular functions of deubiquitinases. J. Biol. Chem. 2022, 298, 102198. [Google Scholar] [CrossRef] [PubMed]
- Ren, J.; et al. Deubiquitylating Enzymes in Cancer and Immunity. Adv. Sci. (Weinh). 2023, 10, e2303807. [Google Scholar] [CrossRef] [PubMed]
- Sahtoe, D.D.; Sixma, T.K. Layers of DUB regulation. Trends Biochem. Sci. 2015, 40, 456–467. [Google Scholar] [CrossRef] [PubMed]
- Snyder, N.A.; Silva, G.M. Deubiquitinating enzymes (DUBs): Regulation, homeostasis, and oxidative stress response. J. Biol. Chem. 2021, 297, 101077. [Google Scholar] [CrossRef]
- Cotto-Rios, X.M.; Békés, M.; Chapman, J.; Ueberheide, B.; Huang, T.T. Deubiquitinases as a signaling target of oxidative stress. Cell Rep. 2012, 2, 1475–1484. [Google Scholar] [CrossRef]
- Kumar, P.; Kumar, P.; Mandal, D.; Velayutham, R. The emerging role of Deubiquitinases (DUBs) in parasites: A foresight review. Front. Cell. Infect. Microbiol. 2022, 12, 985178. [Google Scholar] [CrossRef]
- Suresh, H.G.; Pascoe, N.; Andrews, B. The structure and function of deubiquitinases: Lessons from budding yeast. Open Biol. 2020, 10, 200279. [Google Scholar] [CrossRef]
- Hafez, N.; Modather El-Awadly, Z.; Arafa, R.K. UCH-L3 structure and function: Insights about a promising drug target. Eur. J. Med. Chem. 2022, 227, 113970. [Google Scholar] [CrossRef]
- McGouran, J.F.; Gaertner, S.R.; Altun, M.; Kramer, H.B.; Kessler, B.M. Deubiquitinating enzyme specificity for ubiquitin chain topology profiled by di-ubiquitin activity probes. Chem. Biol. 2013, 20, 1447–1455. [Google Scholar] [CrossRef]
- Hewings, D.S.; Flygare, J.A.; Bogyo, M.; Wertz, I.E. Activity-based probes for the ubiquitin conjugation-deconjugation machinery: New chemistries, new tools, and new insights. FEBS J. 2017, 284, 1555–1576. [Google Scholar] [CrossRef]
- Puri, S.; Hsu, S.-T.D. Functional dynamics of human ubiquitin C-terminal hydrolases. Frontiers in Biophysics 2024, 2. [Google Scholar] [CrossRef]
- Porchietto, E.; et al. UCH-L1 in Alzheimer’s Disease: A Crucial Player in Dementia-Associated Mechanisms. Int. J. Mol. Sci. 2025, 26. [Google Scholar] [CrossRef]
- Mi, Z.; Graham, S.H. Role of UCHL1 in the pathogenesis of neurodegenerative diseases and brain injury. Ageing Res. Rev. 2023, 86, 101856. [Google Scholar] [CrossRef]
- Setsuie, R.; Wada, K. The functions of UCH-L1 and its relation to neurodegenerative diseases. Neurochem. Int. 2007, 51, 105–111. [Google Scholar] [CrossRef]
- Jacobson, A.D.; MacFadden, A.; Wu, Z.; Peng, J.; Liu, C.-W. Autoregulation of the 26S proteasome by in situ ubiquitination. Mol. Biol. Cell 2014, 25, 1824–1835. [Google Scholar] [CrossRef]
- VanderLinden, R.T.; et al. Structural Basis for the Activation and Inhibition of the UCH37 Deubiquitylase. Mol. Cell 2016, 61, 487. [Google Scholar] [CrossRef] [PubMed]
- Yao, T.; et al. Distinct modes of regulation of the Uch37 deubiquitinating enzyme in the proteasome and in the Ino80 chromatin-remodeling complex. Mol. Cell 2008, 31, 909–917. [Google Scholar] [CrossRef] [PubMed]
- Gersch, M.; et al. Distinct USP25 and USP28 Oligomerization States Regulate Deubiquitinating Activity. Mol. Cell 2019, 74, 436–451.e7. [Google Scholar] [CrossRef]
- Maurer, S.K.; et al. Ubiquitin-specific protease 11 structure in complex with an engineered substrate mimetic reveals a molecular feature for deubiquitination selectivity. J. Biol. Chem. 2023, 299, 105300. [Google Scholar] [CrossRef] [PubMed]
- Paudel, P.; et al. Crystal structure and activity-based labeling reveal the mechanisms for linkage-specific substrate recognition by deubiquitinase USP9X. Proceedings of the National Academy of Sciences 2019, 116, 7288–7297. [Google Scholar] [CrossRef] [PubMed]
- Ward, S.J.; et al. The structure of the deubiquitinase USP15 reveals a misaligned catalytic triad and an open ubiquitin-binding channel. J. Biol. Chem. 2018, 293, 17362–17374. [Google Scholar] [CrossRef] [PubMed]
- Devi, U.; Shukla, P.K. The structural, functional, and regulatory insight of deubiquitinating enzyme—USP22. Int. J. Biol. Macromol. 2025, 318, 145164. [Google Scholar] [CrossRef]
- Clague, M.J.; et al. Deubiquitylases From Genes to Organism. Physiol. Rev. 2013, 93, 1289–1315. [Google Scholar] [CrossRef]
- Wu, Y.; et al. Deubiquitinating enzymes in parkinson’s disease: Molecular mechanisms and therapeutic potential. Mol. Med. 2025, 31, 329. [Google Scholar] [CrossRef]
- Pozhidaeva, A.; Bezsonova, I. USP7: Structure, substrate specificity, and inhibition. DNA Repair (Amst). 2019, 76, 30–39. [Google Scholar] [CrossRef]
- Valles, G.J.; Bezsonova, I.; Woodgate, R.; Ashton, N.W. USP7 Is a Master Regulator of Genome Stability. Front. Cell Dev. Biol. 2020, 8, 717. [Google Scholar] [CrossRef]
- Song, Y.; et al. Ubiquitin-Specific Protease 7 (USP7) as a Promising Therapeutic Target for Drug Discovery: From Mechanisms to Therapies. J. Med. Chem. 2025, 68, 7914–7931. [Google Scholar] [CrossRef]
- Berlin, I.; Higginbotham, K.M.; Dise, R.S.; Sierra, M.I.; Nash, P.D. The deubiquitinating enzyme USP8 promotes trafficking and degradation of the chemokine receptor 4 at the sorting endosome. J. Biol. Chem. 2010, 285, 37895–37908. [Google Scholar] [CrossRef] [PubMed]
- Dufner, A.; Knobeloch, K.-P. Ubiquitin-specific protease 8 (USP8/UBPy): A prototypic multidomain deubiquitinating enzyme with pleiotropic functions. Biochem. Soc. Trans. 2019, 47, 1867–1879. [Google Scholar] [CrossRef]
- Srinivasan, V.; et al. USP14 is crucial for proteostasis regulation and α-synuclein degradation in human SH-SY5Y dopaminergic cells. Heliyon 2025, 11, e42031. [Google Scholar] [CrossRef]
- Wang, F.; Ning, S.; Yu, B.; Wang, Y. USP14: Structure, Function, and Target Inhibition. Front. Pharmacol. 2021, 12, 801328. [Google Scholar] [CrossRef]
- Oikawa, D.; et al. OTUD1 deubiquitinase regulates NF-κB- and KEAP1-mediated inflammatory responses and reactive oxygen species-associated cell death pathways. Cell Death Dis. 2022, 13, 694. [Google Scholar] [CrossRef]
- Fu, L.; Lu, K.; Jiao, Q.; Chen, X.; Jia, F. The Regulation and Double-Edged Roles of the Deubiquitinase OTUD5. Cells 2023, 12. [Google Scholar] [CrossRef]
- Li, F.; et al. The deubiquitinase OTUD5 regulates Ku80 stability and non-homologous end joining. Cell. Mol. Life Sci. 2019, 76, 3861–3873. [Google Scholar] [CrossRef]
- Kabra, A.; Benson, C.A.; Li, Y. Backbone 1H, 13C, and 15N resonance assignments of deubiquitinase A in non-phosphorylated and phosphorylated forms. Biomol. NMR Assign. 2019, 13, 37–42. [Google Scholar] [CrossRef]
- Kabra, A.; Rumpa, E.; Li, Y. Observation of Arginine Side-Chain Motions Coupled to the Global Conformational Exchange Process in Deubiquitinase A. ACS Omega 2022, 7, 9936–9943. [Google Scholar] [CrossRef] [PubMed]
- Kabra, A.; Li, Y. Conformational Dynamics of Deubiquitinase A and Functional Implications. Biochemistry 2021, 60, 201–209. [Google Scholar] [CrossRef] [PubMed]
- Kabra, A.; Rumpa, E.; Li, Y. Modulation of conformational equilibrium by phosphorylation underlies the activation of deubiquitinase A. Journal of Biological Chemistry 2020, 295, 3945–3951. [Google Scholar] [CrossRef]
- Lork, M.; Verhelst, K.; Beyaert, R. CYLD, A20 and OTULIN deubiquitinases in NF-κB signaling and cell death: So similar, yet so different. Cell Death Differ. 2017, 24, 1172–1183. [Google Scholar] [CrossRef]
- Weinelt, N.; van Wijk, S.J.L. Ubiquitin-dependent and -independent functions of OTULIN in cell fate control and beyond. Cell Death Differ. 2021, 28, 493–504. [Google Scholar] [CrossRef]
- Bettencourt, C.; Lima, M. Machado-Joseph Disease: From first descriptions to new perspectives. Orphanet J. Rare Dis. 2011, 6, 35. [Google Scholar] [CrossRef]
- Zeng, C.; et al. Machado-Joseph Deubiquitinases: From Cellular Functions to Potential Therapy Targets. Front. Pharmacol. 2020, 11, 1311. [Google Scholar] [CrossRef]
- Weeks, S.D.; Grasty, K.C.; Hernandez-Cuebas, L.; Loll, P.J. Crystal structure of a Josephin-ubiquitin complex: Evolutionary restraints on ataxin-3 deubiquitinating activity. J. Biol. Chem. 2011, 286, 4555–4565. [Google Scholar] [CrossRef]
- Grasty, K.C.; Weeks, S.D.; Loll, P.J. Structural insights into the activity and regulation of human Josephin-2. J. Struct. Biol. X 2019, 3, 100011. [Google Scholar] [CrossRef]
- Yang, H.; et al. Aggregation of polyglutamine-expanded ataxin-3 sequesters its specific interacting partners into inclusions: Implication in a loss-of-function pathology. Sci. Rep. 2014, 4, 6410. [Google Scholar] [CrossRef] [PubMed]
- Seki, T.; et al. JosD1, a membrane-targeted deubiquitinating enzyme, is activated by ubiquitination and regulates membrane dynamics, cell motility, and endocytosis. J. Biol. Chem. 2013, 288, 17145–17155. [Google Scholar] [CrossRef]
- Abdul Rehman, S.A.; et al. MINDY-1 Is a Member of an Evolutionarily Conserved and Structurally Distinct New Family of Deubiquitinating Enzymes. Mol. Cell 2016, 63, 146–155. [Google Scholar] [CrossRef] [PubMed]
- Abdul Rehman, S.A.; et al. Mechanism of activation and regulation of deubiquitinase activity in MINDY1 and MINDY2. Mol. Cell 2021, 81, 4176–4190.e6. [Google Scholar] [CrossRef] [PubMed]
- Haahr, P.; et al. ZUFSP Deubiquitylates K63-Linked Polyubiquitin Chains to Promote Genome Stability. Mol. Cell 2018, 70, 165–174.e6. [Google Scholar] [CrossRef] [PubMed]
- Hermanns, T.; Pichlo, C.; Baumann, U.; Hofmann, K. A structural basis for the diverse linkage specificities within the ZUFSP deubiquitinase family. Nat. Commun. 2022, 13, 401. [Google Scholar] [CrossRef] [PubMed]
- Kwasna, D.; et al. Discovery and Characterization of ZUFSP/ZUP1, a Distinct Deubiquitinase Class Important for Genome Stability. Mol. Cell 2018, 70, 150–164.e6. [Google Scholar] [CrossRef]
- Hameed, D.S.; et al. Development of Ubiquitin-Based Probe for Metalloprotease Deubiquitinases. Angew. Chem. Int. Ed Engl. 2019, 58, 14477–14482. [Google Scholar] [CrossRef]
- Pan, X.; et al. Structural and Functional Basis of JAMM Deubiquitinating Enzymes in Disease. Biomolecules 2022, 12. [Google Scholar] [CrossRef]
- Guo, Y.; et al. Structural and functional characterization of ubiquitin variant inhibitors for the JAMM-family deubiquitinases STAMBP and STAMBPL1. Journal of Biological Chemistry 2021, 297, 101107. [Google Scholar] [CrossRef]
- Shrestha, R.K.; et al. Insights into the mechanism of deubiquitination by JAMM deubiquitinases from cocrystal structures of the enzyme with the substrate and product. Biochemistry 2014, 53, 3199–3217. [Google Scholar] [CrossRef]
- Spataro, V.; Buetti-Dinh, A. POH1/Rpn11/PSMD14: A journey from basic research in fission yeast to a prognostic marker and a druggable target in cancer cells. Br. J. Cancer 2022, 127, 788–799. [Google Scholar] [CrossRef]
- Htet, Z.M.; Dong, K.C.; Martin, A. The deubiquitinase Rpn11 functions as an allosteric ubiquitin sensor to promote substrate engagement by the 26S proteasome. Cell Rep. 2025, 44, 115736. [Google Scholar] [CrossRef]
- Echalier, A.; et al. Insights into the regulation of the human COP9 signalosome catalytic subunit, CSN5/Jab1. Proceedings of the National Academy of Sciences 2013, 110, 1273–1278. [Google Scholar] [CrossRef]
- Muromoto, R.; et al. Jun activation domain-binding protein 1 (JAB1) is required for the optimal response to interferons. J. Biol. Chem. 2013, 288, 30969–30979. [Google Scholar] [CrossRef]
- McCullough, J.; Clague, M.J.; Urbé, S. AMSH is an endosome-associated ubiquitin isopeptidase. J. Cell Biol. 2004, 166, 487–492. [Google Scholar] [CrossRef]
- Agromayor, M.; Martin-Serrano, J. Interaction of AMSH with ESCRT-III and Deubiquitination of Endosomal Cargo. Journal of Biological Chemistry 2006, 281, 23083–23091. [Google Scholar] [CrossRef] [PubMed]
- Ng, H.-M.; Wei, L.; Lan, L.; Huen, M.S.Y. The Lys63-deubiquitylating Enzyme BRCC36 Limits DNA Break Processing and Repair. J. Biol. Chem. 2016, 291, 16197–16207. [Google Scholar] [CrossRef] [PubMed]
- Rabl, J.; et al. Structural Basis of BRCC36 Function in DNA Repair and Immune Regulation. Mol. Cell 2019, 75, 483–497.e9. [Google Scholar] [CrossRef]
- Jiang, X.-X.; et al. Epigenetic Regulation of Antibody Responses by the Histone H2A Deubiquitinase MYSM1. Sci. Rep. 2015, 5, 13755. [Google Scholar] [CrossRef] [PubMed]
- Nandakumar, V.; Chou, Y.; Zang, L.; Huang, X.F.; Chen, S.-Y. Epigenetic control of natural killer cell maturation by histone H2A deubiquitinase, MYSM1. Proc. Natl. Acad. Sci. USA 2013, 110, E3927–E3936. [Google Scholar] [CrossRef]
- Nijman, S.M.B.; et al. The Deubiquitinating Enzyme USP1 Regulates the Fanconi Anemia Pathway. Mol. Cell 2005, 17, 331–339. [Google Scholar] [CrossRef]
- Lee, B.-C.; Miyata, M.; Lim, J.H.; Li, J.-D. Deubiquitinase CYLD acts as a negative regulator for bacterium NTHi-induced inflammation by suppressing K63-linked ubiquitination of MyD88. Proceedings of the National Academy of Sciences 2016, 113. [Google Scholar] [CrossRef]
- Sun, S.-C. CYLD: A tumor suppressor deubiquitinase regulating NF-kappaB activation and diverse biological processes. Cell Death Differ. 2010, 17, 25–34. [Google Scholar] [CrossRef]
- Ning, S.; Pagano, J.S. The A20 Deubiquitinase Activity Negatively Regulates LMP1 Activation of IRF7. J. Virol. 2010, 84, 6130–6138. [Google Scholar] [CrossRef]
- Ketscher, L.; et al. Selective inactivation of USP18 isopeptidase activity in vivo enhances ISG15 conjugation and viral resistance. Proceedings of the National Academy of Sciences 2015, 112, 1577–1582. [Google Scholar] [CrossRef] [PubMed]
- Elliott, P.R.; et al. Molecular Basis and Regulation of OTULIN-LUBAC Interaction. Mol. Cell 2014, 54, 335–348. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Guo, Y.; Wang, Q. USP21 accelerates the proliferation and glycolysis of esophageal cancer cells by regulating the STAT3/FOXO1 pathway. Tissue Cell 2022, 79, 101916. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.-X.; et al. The nucleolar ubiquitin-specific protease USP36 deubiquitinates and stabilizes c-Myc. Proceedings of the National Academy of Sciences 2015, 112, 3734–3739. [Google Scholar] [CrossRef]
- Ashkenazi, A.; et al. Polyglutamine tracts regulate beclin 1-dependent autophagy. Nature 2017, 545, 108–111. [Google Scholar] [CrossRef]
- Bridges, C.R.; et al. USP9X deubiquitylating enzyme maintains RAPTOR protein levels, mTORC1 signalling and proliferation in neural progenitors. Sci. Rep. 2017, 7, 391. [Google Scholar] [CrossRef]
- Herhaus, L.; Al-Salihi, M.; Macartney, T.; Weidlich, S.; Sapkota, G.P. OTUB1 enhances TGFβ signalling by inhibiting the ubiquitylation and degradation of active SMAD2/3. Nat. Commun. 2013, 4, 2519. [Google Scholar] [CrossRef]
- Qi, S.-M.; et al. Targeting USP7-Mediated Deubiquitination of MDM2/MDMX-p53 Pathway for Cancer Therapy: Are We There Yet? Front. Cell Dev. Biol. 2020, 8. [Google Scholar] [CrossRef]
- Popov, N.; et al. The ubiquitin-specific protease USP28 is required for MYC stability. Nat. Cell Biol. 2007, 9, 765–774. [Google Scholar] [CrossRef] [PubMed]
- Luo, G.; Hu, N.; Xia, X.; Zhou, J.; Ye, C. RPN11 deubiquitinase promotes proliferation and migration of breast cancer cells. Mol. Med. Rep. 2017, 16, 331–338. [Google Scholar] [CrossRef]
- Rybarikova, M.; et al. Gene editing for Spinocerebellar ataxia type 3 taking advantage of the human ATXN3L paralog as replacement gene. Gene Ther. 2025, 32, 462–474. [Google Scholar] [CrossRef] [PubMed]
- Durcan, T.M.; et al. USP8 regulates mitophagy by removing K6-linked ubiquitin conjugates from parkin. EMBO J. 2014, 33, 2473–2491. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Liu, J.; Wen, J.; Sun, Y.; He, M. Exploring the ubiquitination regulatory network: Opening new perspectives for rheumatoid arthritis therapy. Ann. Med. 2025, 57. [Google Scholar] [CrossRef]
- Watanabe, T.; Kudo, M. Roles of Deubiquitinases OTUD3 and OTUD5 in Inflammatory Bowel Diseases. Int. J. Mol. Sci. 2025, 26, 9924. [Google Scholar] [CrossRef]
- Wang, N.; et al. USP16 drives psoriasis progression by deubiquitinating and stabilizing NLRP3 in keratinocytes. JCI Insight 2026. [Google Scholar] [CrossRef]
- Wu, Y.; He, X.; Huang, N.; Yu, J.; Shao, B. A20: A master regulator of arthritis. Arthritis Res. Ther. 2020, 22, 220. [Google Scholar] [CrossRef]
- Yu, H.; Lin, L.; Zhang, Z.; Zhang, H.; Hu, H. Targeting NF-κB pathway for the therapy of diseases: Mechanism and clinical study. Signal Transduct. Target. Ther. 2020, 5, 209. [Google Scholar] [CrossRef]
- Cao, D.; et al. The SARS-CoV-2 papain-like protease suppresses type I interferon responses by deubiquitinating STING. Sci. Signal. 2023, 16. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; et al. SARS-CoV-2 papain-like protease plays multiple roles in regulating cellular proteins in the endoplasmic reticulum. Journal of Biological Chemistry 2023, 299, 105346. [Google Scholar] [CrossRef]
- Bodda, C.; et al. HSV1 VP1-2 deubiquitinates STING to block type I interferon expression and promote brain infection. Journal of Experimental Medicine 2020, 217. [Google Scholar] [CrossRef]
- Gao, W.; et al. Specific Deubiquitinating Enzymes Promote Host Restriction Factors Against HIV/SIV Viruses. Front. Immunol. 2021, 12. [Google Scholar] [CrossRef]
- Molusky, M.M.; Li, S.; Ma, D.; Yu, L.; Lin, J.D. Ubiquitin-Specific Protease 2 Regulates Hepatic Gluconeogenesis and Diurnal Glucose Metabolism Through 11β-Hydroxysteroid Dehydrogenase 1. Diabetes 2012, 61, 1025–1035. [Google Scholar] [CrossRef] [PubMed]
- Kitamura, H. Ubiquitin-Specific Proteases (USPs) and Metabolic Disorders. Int. J. Mol. Sci. 2023, 24, 3219. [Google Scholar] [CrossRef]
- Li, X.; et al. Ubiquitin Specific Protease 9X Regulates the Activation of ARK5 and Promotes Progression of Fibrotic Remodeling. JACC Basic Transl. Sci. 2025, 10, 101255. [Google Scholar] [CrossRef] [PubMed]
- Schauer, N.J.; et al. Selective USP7 inhibition elicits cancer cell killing through a p53-dependent mechanism. Sci. Rep. 2020, 10, 5324. [Google Scholar] [CrossRef]
- Nie, L.; et al. USP7 substrates identified by proteomics analysis reveal the specificity of USP7. Genes Dev. 2022, 36, 1016–1030. [Google Scholar] [CrossRef]
- Okarmus, J.; et al. USP30 inhibition induces mitophagy and reduces oxidative stress in parkin-deficient human neurons. Cell Death Dis. 2024, 15, 52. [Google Scholar] [CrossRef]
- Huang, Z.; Tan, Y. The Potential of Cylindromatosis (CYLD) as a Therapeutic Target in Oxidative Stress-Associated Pathologies: A Comprehensive Evaluation. Int. J. Mol. Sci. 2023, 24. [Google Scholar] [CrossRef]
- Mathis, B.J.; Lai, Y.; Qu, C.; Janicki, J.S.; Cui, T. CYLD-mediated signaling and diseases. Curr. Drug Targets 2015, 16, 284–294. [Google Scholar] [CrossRef]
- Yamanaka, S.; et al. Subquinocin, a small molecule inhibitor of CYLD and USP-family deubiquitinating enzymes, promotes NF-κB signaling. Biochem. Biophys. Res. Commun. 2020, 524, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; et al. Capzimin is a potent and specific inhibitor of proteasome isopeptidase Rpn11. Nat. Chem. Biol. 2017, 13, 486–493. [Google Scholar] [CrossRef]
- Perez, C.; et al. Discovery of an Inhibitor of the Proteasome Subunit Rpn11. J. Med. Chem. 2017, 60, 1343–1361. [Google Scholar] [CrossRef]
- Boselli, M.; et al. An inhibitor of the proteasomal deubiquitinating enzyme USP14 induces tau elimination in cultured neurons. Journal of Biological Chemistry 2017, 292, 19209–19225. [Google Scholar] [CrossRef]
- Wang, Y.; et al. Small molecule inhibitors reveal allosteric regulation of USP14 via steric blockade. Cell Res. 2018, 28, 1186–1194. [Google Scholar] [CrossRef]
- Jaen Maisonet, I.; et al. Small-molecule allosteric activator of ubiquitin-specific protease 7 (USP7). Proceedings of the National Academy of Sciences 2025, 122. [Google Scholar] [CrossRef]
- Wang, X.; Liu, N.; Li, N.; Lu, S.; Chai, Z. Mechanistic Insights into the Mechanism of Allosteric Inhibition of Ubiquitin-Specific Protease 7 (USP7). Biomolecules 2025, 15. [Google Scholar] [CrossRef] [PubMed]
- Cremer, A.; Stegmaier, K. Targeting DUBs to degrade oncogenic proteins. Br. J. Cancer 2020, 122, 1121–1123. [Google Scholar] [CrossRef]
- Murgai, A.; et al. Targeting the deubiquitinase USP7 for degradation with PROTACs. Chemical Communications 2022, 58, 8858–8861. [Google Scholar] [CrossRef]
- Ma, X.b.; et al. Synthesis and identification of a novel selective USP7 degrader that inhibits the migration of upper gastrointestinal cancer cells without affecting proliferation. Eur. J. Med. Chem. 2026, 304, 118488. [Google Scholar] [CrossRef]
- Diefenbacher, M.E.; et al. The deubiquitinase USP28 controls intestinal homeostasis and promotes colorectal cancer. Journal of Clinical Investigation 2014, 124, 3407–3418. [Google Scholar] [CrossRef]
- Wang, J.; et al. The deubiquitinase USP28 stabilizes the expression of RecQ family helicases and maintains the viability of triple negative breast cancer cells. J. Biol. Chem. 2022, 298, 101443. [Google Scholar] [CrossRef]
- Ma, Z.; Zhou, M.; Chen, H.; Shen, Q.; Zhou, J. Deubiquitinase-Targeting Chimeras (DUBTACs) as a Potential Paradigm-Shifting Drug Discovery Approach. J. Med. Chem. 2025, 68, 6897–6915. [Google Scholar] [CrossRef] [PubMed]
- Henning, N.J.; et al. Deubiquitinase-targeting chimeras for targeted protein stabilization. Nat. Chem. Biol. 2022, 18, 412–421. [Google Scholar] [CrossRef]
- Liu, J.; et al. USP7-Based Deubiquitinase-Targeting Chimeras Stabilize AMPK. J. Am. Chem. Soc. 2024. [Google Scholar] [CrossRef]
- Wang, Z.; et al. USP28-Based Deubiquitinase-Targeting Chimeras for Cancer Treatment. J. Am. Chem. Soc. 2025, 147, 13754–13763. [Google Scholar] [CrossRef]
- Chen, L.; et al. Deubiquitinase-Targeting Chimeras Mediated Stabilization of Tumor Suppressive E3 Ligase Proteins as a Strategy for Cancer Therapy. J. Am. Chem. Soc. 2025, 147, 29875–29883. [Google Scholar] [CrossRef]
- Ma, Z.; Zhou, M.; Chen, H.; Shen, Q.; Zhou, J. Deubiquitinase-Targeting Chimeras (DUBTACs) as a Potential Paradigm-Shifting Drug Discovery Approach. J. Med. Chem. 2025, 68, 6897–6915. [Google Scholar] [CrossRef]
- Opferman, J.T.; Green, D.R. DUB-le Trouble for Cell Survival. Cancer Cell 2010, 17, 117–119. [Google Scholar] [CrossRef] [PubMed]
- HU, H.; et al. Reduced ubiquitin-specific protease 9X expression induced by RNA interference inhibits the bioactivity of hepatocellular carcinoma cells. Oncol. Lett. 2015, 10, 268–272. [Google Scholar] [CrossRef]
- D’Arcy, P.; et al. Inhibition of proteasome deubiquitinating activity as a new cancer therapy. Nat. Med. 2011, 17, 1636–1640. [Google Scholar] [CrossRef]
- Wang, X.; et al. The proteasome deubiquitinase inhibitor VLX1570 shows selectivity for ubiquitin-specific protease-14 and induces apoptosis of multiple myeloma cells. Sci. Rep. 2016, 6, 26979. [Google Scholar] [CrossRef]
- Wang, H.; et al. USP28 and USP25 are downregulated by Vismodegib in vitro and in colorectal cancer cell lines. FEBS J. 2021, 288, 1325–1342. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Wu, Y.; Yao, M.; Chen, Z.; Li, Q. The other side of the coin: Protein deubiquitination by Ubiquitin-Specific Protease 1 in cancer progression and therapy. Future Med. Chem. 2025, 17, 329–345. [Google Scholar] [CrossRef] [PubMed]
- Pauzaite, T.; Nathan, J.A. A closer look at the role of deubiquitinating enzymes in the Hypoxia Inducible Factor pathway. Biochem. Soc. Trans. 2024, 52, 2253–2265. [Google Scholar] [CrossRef] [PubMed]
- Guo, D.; Peng, Y.; Yu, Y. The proteostasis paradox: From systemic collapse in aging to pathway-specific addiction in prostate cancer. Front. Cell Dev. Biol. 2026, 14. [Google Scholar] [CrossRef]
- Liang, Q.; et al. A selective USP1–UAF1 inhibitor links deubiquitination to DNA damage responses. Nat. Chem. Biol. 2014, 10, 298–304. [Google Scholar] [CrossRef]
| Cellular Process | Specific DUBs | Known Substrates/Targets |
| Protein Quality Control | USP7, USP14, UCH-L1, USP9X, ZUFSP | Misfolded proteins, proteasome components, Chaperones |
| DNA Damage Repair | OTUB1, Ataxin-3 (ATXN3), USP28 | DNA repair factors (e.g., FANCD2, PCNA), chromatin-associated proteins, replication stress regulators |
| Immune Responses | AMSH, USP18, USP30, CYLD, OTULIN | Immune signaling proteins (e.g., TRAF6, STING, NF-κB components), cytokine receptors, interferon pathway proteins |
| Transcriptional Regulation | USP22, USP7, USP9X, USP21 | Transcription factors (e.g., c-Myc, p53), histones, RNA Pol II, chromatin modifiers |
| Membrane Traffic | AMSH, USP8, USP30 | Endocytic vesicles, membrane receptors (e.g., EGFR), mitophagy regulators, vesicle trafficking proteins |
| Cell Cycle & Apoptosis | USP7, USP9X, CYLD, USP28 | Cyclins, checkpoint proteins (e.g., p53, MDM2), apoptotic regulators (Bcl-2 family) |
| Signal Transduction | USP7, USP21, CYLD, OTULIN | Kinases, ubiquitin ligases, NF-κB, Wnt/β-catenin pathway proteins, MAPK signaling components |
| Autophagy & Mitophagy | USP30, USP10, USP15, USP13 | Parkin substrates, mitochondrial proteins, autophagy regulators (LC3, Beclin1) |
| Disease | Dysregulated DUBs | Mechanisms and Consequences |
| Cancer | USP7, USP14, USP22, UCH-L1, USP9X, CYLD, OTUD7B | Altered protein stability, cell proliferation, impaired apoptosis, Oncogene stabilization, modulation of NF-κB and p53 pathway |
| Neurodegenerative Disorders | Ataxin-3 (ATXN3), USP14, USP7, USP30, USP9X | Aggregation of misfolded proteins, impaired proteostasis. Defective mitophagy, neuronal death |
| Inflammatory Diseases | AMSH, USP18, USP30, CYLD, OTULIN | Dysregulation of immune signaling pathways, abnormal cytokine production, chronic inflammation |
| Viral Infections | USP7, USP14, USP30, USP21 | Promoting Viral replication, immune evasion, inhibition of host antiviral response, modulation of interferon signaling |
| Cardiovascular Diseases | USP10, USP2, USP20 | Regulation of cardiac hypertrophy, endothelial dysfunction, inflammation, and atherosclerosis via modulation of NF-κB and MAPK pathways |
| Metabolic Disorders | USP2, USP14, USP19 | Dysregulation of lipid metabolism, glucose homeostasis, insulin signaling, contributing to obesity, diabetes, and metabolic syndrome |
| Fibrotic Diseases | USP4, USP15, CYLD | Enhanced TGF-β signaling, fibroblast activation, extracellular matrix accumulation, promoting organ fibrosis (liver, lung, kidney) |
| Inhibitor | Target DUBs | Therapeutic potential |
| IU1 | USP14 | Neurogenerative diseases |
| Capzimin | RPN11 | Cancer therapy |
| VLX1570 | USP14 | Multiple myeloma |
| G9 | USP7 | Cancer therapy |
| P22077 | USP7 | Cancer therapy |
| WP1130 | USP5, USP14, USP9x, UCH37 | Multiple Myeloma |
| b-AP15 | USP14, UCHL5 | Cancer therapy |
| FT671 | USP7 | Cancer therapy |
| PR-619 | Multiple DUBs (Broad spectrum DUB inhibitor) | Neurogenerative diseases |
| GSK2643943A | USP30 | Neurogenerative diseases |
| USP7-1 | USP7 | Cancer therapy |
| USP30-2 | USP30 | Mitochondrial disorders & Parkinsons disease |
| USP8i | USP8 | Cushing’s disease, cancer |
| DUB-IN-1 | Multiple DUBs (Broad spectrum DUB inhibitor) | Cancer therapy |
| USP9x-1 | USP9X | Cancer therapy |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





