Submitted:
27 February 2026
Posted:
02 March 2026
You are already at the latest version
Abstract
Keywords:
Africa as a Frontier for Ancient Pathogen Genomics
Ancient Pathogens - Revising Origins, Hosts and Spread
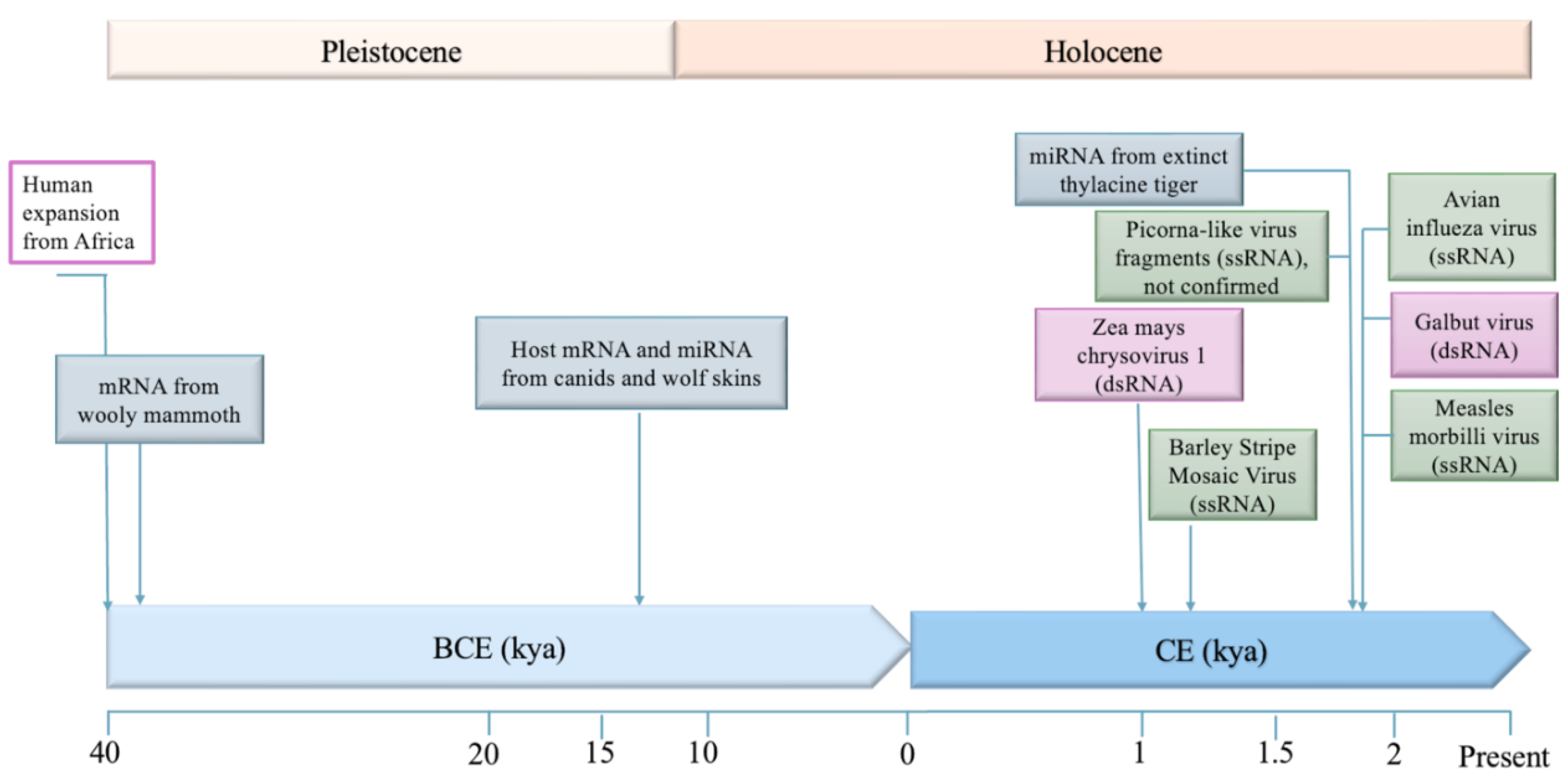
Recovering Ancient RNA Viruses - Challenges and Promise
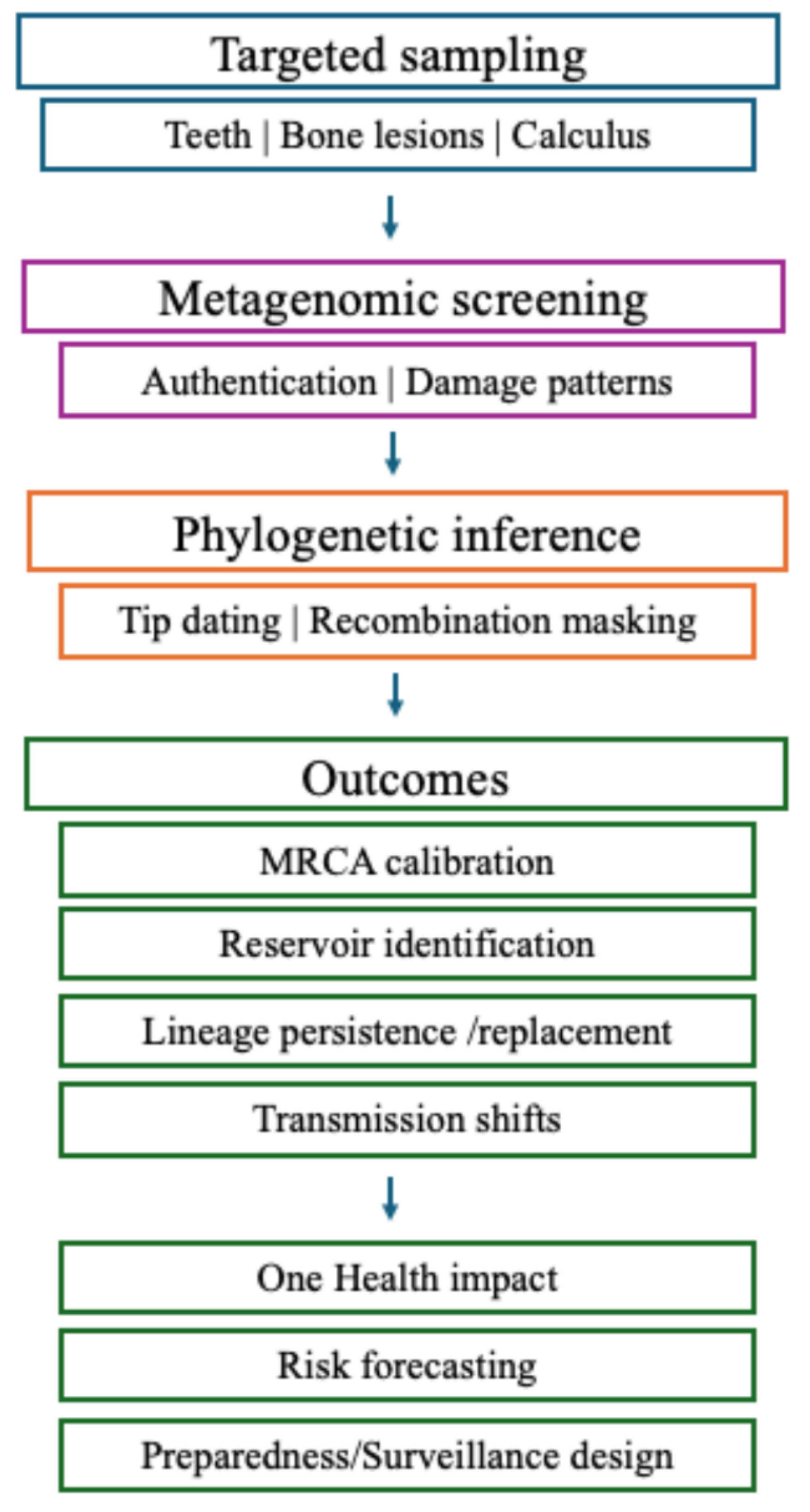
Ancient Pathogen Genomics in Africa - Current Limitations and Insights into Disease Landscape
Future aDNA Research Perspectives on Africa’s Disease History
Conclusion
Acknowledgments
Declaration of interests
References
- Avila-Arcos, M.C.; et al. Going local with ancient DNA: A review of human histories from regional perspectives. Science 2023, 382, 53–58. [Google Scholar] [CrossRef]
- Sikora, M.; et al. The spatiotemporal distribution of human pathogens in ancient Eurasia. Nature 2025, 643, 1011–1019. [Google Scholar] [CrossRef] [PubMed]
- Blevins, K.E.; et al. Ancient DNA insights into diverse pathogens and their hosts. Nature Reviews Genetics 2026, 27, 96–111. [Google Scholar] [CrossRef] [PubMed]
- Jones, K.E.; et al. Global trends in emerging infectious diseases. Nature 2008, 451, 990–993. [Google Scholar] [CrossRef] [PubMed]
- Wolfe, N.D.; et al. Origins of major human infectious diseases. Nature 2007, 447, 279–283. [Google Scholar] [CrossRef]
- Bos, K.I.; et al. A draft genome of Yersinia pestis from victims of the Black Death. Nature 2011, 478, 506–510. [Google Scholar] [CrossRef]
- Spyrou, M.A.; et al. Ancient pathogen genomics as an emerging tool for infectious disease research. Nature Reviews Genetics 2019, 20, 323–340. [Google Scholar] [CrossRef]
- Key, F.M.; et al. Mining metagenomic data sets for ancient DNA: recommended protocols for authentication. Trends in Genetics 2017, 33, 508–520. [Google Scholar] [CrossRef]
- Philips, A.; et al. Comprehensive analysis of microorganisms accompanying human archaeological remains. GigaScience 2017, 6, gix044. [Google Scholar] [CrossRef]
- Rifkin, R.F.; et al. Rickettsia felis DNA recovered from a child who lived in southern Africa 2000 years ago. Communications biology 2023, 6, 240. [Google Scholar] [CrossRef]
- Kerner, G.; et al. Ancient DNA as a tool for medical research. Nature medicine 2023, 29, 1048–1051. [Google Scholar] [CrossRef]
- Warinner, C.; et al. A robust framework for microbial archaeology. Annual review of genomics and human genetics 2017, 18, 321–356. [Google Scholar] [CrossRef] [PubMed]
- Donoghue, H.D. Insights gained from ancient biomolecules into past and present tuberculosis—a personal perspective. International Journal of Infectious Diseases 2017, 56, 176–180. [Google Scholar] [CrossRef] [PubMed]
- Swali, P.; et al. Yersinia pestis genomes reveal plague in Britain 4000 years ago. Nature Communications 2023, 14, 2930. [Google Scholar] [CrossRef]
- Mühlemann, B.; et al. Ancient hepatitis B viruses from the Bronze Age to the Medieval period. Nature 2018, 557, 418–423. [Google Scholar] [CrossRef] [PubMed]
- Kocher, A.; et al. Ten millennia of hepatitis B virus evolution. Science 2021, 374, 182–188. [Google Scholar] [CrossRef]
- Guellil, M.; et al. Ancient herpes simplex 1 genomes reveal recent viral structure in Eurasia. Science Advances 2022, 8, eabo4435. [Google Scholar] [CrossRef]
- Gelabert Xirinachs, P.; et al. Malaria was a weak selective force in ancient Europeans. Scientific reports 2017, 7, 1377. [Google Scholar] [CrossRef]
- Marciniak, S.; et al. Plasmodium falciparum malaria in 1st–2nd century CE southern Italy. Current Biology 2016, 26, R1220–R1222. [Google Scholar] [CrossRef]
- Guzmán-Solís, A.A.; et al. Ancient viral genomes reveal introduction of human pathogenic viruses into Mexico during the transatlantic slave trade. Elife 2021, 10, e68612. [Google Scholar] [CrossRef]
- Vågene, Å.J.; et al. Salmonella enterica genomes from victims of a major sixteenth-century epidemic in Mexico. Nature ecology & evolution 2018, 2, 520–528. [Google Scholar]
- Bos, K.I.; et al. Pre-Columbian mycobacterial genomes reveal seals as a source of New World human tuberculosis. Nature 2014, 514, 494–497. [Google Scholar] [CrossRef]
- Spyrou, M.A.; et al. Analysis of 3800-year-old Yersinia pestis genomes suggests Bronze Age origin for bubonic plague. Nature communications 2018, 9, 2234. [Google Scholar] [CrossRef]
- Kahila Bar-Gal, G.; et al. Tracing hepatitis B virus to the 16th century in a Korean mummy. Hepatology 2012, 56, 1671–1680. [Google Scholar] [CrossRef]
- Spyrou, M.A.; et al. Phylogeography of the second plague pandemic revealed through analysis of historical Yersinia pestis genomes. Nature communications 2019, 10, 4470. [Google Scholar] [CrossRef] [PubMed]
- Rascovan, N.; et al. Emergence and spread of basal lineages of Yersinia pestis during the Neolithic decline. Cell 2019, 176, 295–305. [Google Scholar] [CrossRef] [PubMed]
- Seersholm, F.V.; et al. Repeated plague infections across six generations of Neolithic Farmers. Nature 2024, 632, 114–121. [Google Scholar] [CrossRef]
- Macleod, R.; et al. Lethal Plague Outbreaks in Lake Baikal Hunter–gatherers 5500 Years Ago. bioRxiv 2024. [Google Scholar] [CrossRef]
- Valtueña, A.A.; et al. The Stone Age plague and its persistence in Eurasia. Current biology 2017, 27, 3683–3691. [Google Scholar] [CrossRef]
- Wagner, D.M.; et al. Yersinia pestis and the Plague of Justinian 541–543 AD: a genomic analysis. The Lancet Infectious Diseases 2014, 14, 319–326. [Google Scholar] [CrossRef]
- Spyrou, M.A.; et al. The source of the Black Death in fourteenth-century central Eurasia. Nature 2022, 606, 718–724. [Google Scholar] [CrossRef]
- Spyrou, M.A.; et al. Historical Y. pestis genomes reveal the European Black Death as the source of ancient and modern plague pandemics. Cell host & microbe 2016, 19, 874–881. [Google Scholar]
- Vågene, Å.J.; et al. Geographically dispersed zoonotic tuberculosis in pre-contact South American human populations. Nature communications 2022, 13, 1195. [Google Scholar] [CrossRef]
- Comas, I.; et al. Out-of-Africa migration and Neolithic coexpansion of Mycobacterium tuberculosis with modern humans. Nature genetics 2013, 45, 1176–1182. [Google Scholar] [CrossRef]
- Sabin, S.; et al. A seventeenth-century Mycobacterium tuberculosis genome supports a Neolithic emergence of the Mycobacterium tuberculosis complex. Genome biology 2020, 21, 201. [Google Scholar] [CrossRef] [PubMed]
- Michel, M.; et al. Ancient Plasmodium genomes shed light on the history of human malaria. Nature 2024, 631, 125–133. [Google Scholar] [CrossRef]
- Liu, W.; et al. Origin of the human malaria parasite Plasmodium falciparum in gorillas. Nature 2010, 467, 420–425. [Google Scholar] [CrossRef] [PubMed]
- Michel, M.; et al. Ancient Plasmodium genomes shed light on the history of human malaria. Nature 2024, 631, 125–133. [Google Scholar] [CrossRef]
- Loy, D.E.; et al. Out of Africa: origins and evolution of the human malaria parasites Plasmodium falciparum and Plasmodium vivax. International journal for parasitology 2017, 47, 87–97. [Google Scholar] [CrossRef]
- Mu, J.; et al. Host switch leads to emergence of Plasmodium vivax malaria in humans. Molecular biology and evolution 2005, 22, 1686–1693. [Google Scholar] [CrossRef] [PubMed]
- Jongwutiwes, S.; et al. Mitochondrial genome sequences support ancient population expansion in Plasmodium vivax. Molecular biology and evolution 2005, 22, 1733–1739. [Google Scholar] [CrossRef]
- Loy, D.E.; et al. Evolutionary history of human Plasmodium vivax revealed by genome-wide analyses of related ape parasites. Proceedings of the National Academy of Sciences 2018, 115, E8450–E8459. [Google Scholar] [CrossRef] [PubMed]
- Neukamm, J.; et al. 2000-year-old pathogen genomes reconstructed from metagenomic analysis of Egyptian mummified individuals. BMC biology 2020, 18, 108. [Google Scholar] [CrossRef]
- Andernach, I.E.; et al. Slave trade and hepatitis B virus genotypes and subgenotypes in Haiti and Africa. Emerging Infectious Diseases 2009, 15, 1222. [Google Scholar] [CrossRef] [PubMed]
- Quintero, A.; et al. Molecular epidemiology of hepatitis B virus in Afro-Venezuelan populations. Archives of virology 2002, 147, 1829–1836. [Google Scholar] [CrossRef]
- Brichler, S.; et al. African, Amerindian and European hepatitis B virus strains circulate on the Caribbean Island of Martinique. Journal of General Virology 2013, 94, 2318–2329. [Google Scholar] [CrossRef] [PubMed]
- Alvarado-Mora, M.V.; et al. Phylogenetic analysis of complete genome sequences of hepatitis B virus from an Afro-Colombian community: presence of HBV F3/A1 recombinant strain. Virology Journal 2012, 9, 244. [Google Scholar] [CrossRef]
- Forni, D.; et al. Recent out-of-Africa migration of human herpes simplex viruses. Molecular Biology and Evolution 2020, 37, 1259–1271. [Google Scholar] [CrossRef]
- Duffy, S.; et al. Rates of evolutionary change in viruses: patterns and determinants. Nature Reviews Genetics 2008, 9, 267–276. [Google Scholar] [CrossRef]
- Firth, C.; et al. Using time-structured data to estimate evolutionary rates of double-stranded DNA viruses. Molecular biology and evolution 2010, 27, 2038–2051. [Google Scholar] [CrossRef]
- Duchêne, S.; et al. Genome-scale rates of evolutionary change in bacteria. Microbial genomics 2016, 2, e000094. [Google Scholar] [CrossRef]
- Katzourakis, A.; Gifford, R.J. Endogenous viral elements in animal genomes. PLoS genetics 2010, 6, e1001191. [Google Scholar] [CrossRef] [PubMed]
- Aiewsakun, P.; Katzourakis, A. Endogenous viruses: Connecting recent and ancient viral evolution. Virology 2015, 479, 26–37. [Google Scholar] [CrossRef]
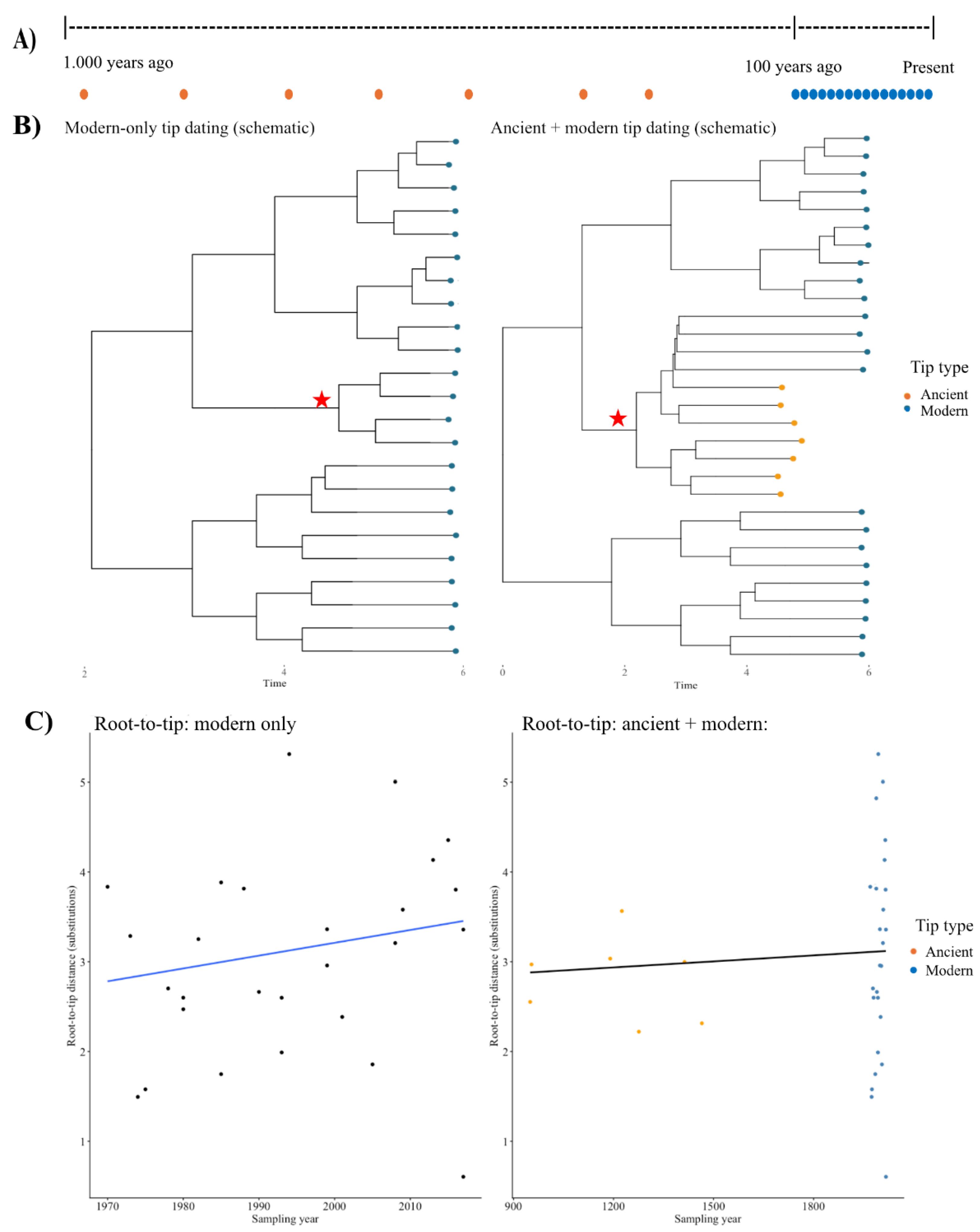
- Ho, S.Y.; et al. Time dependency of molecular rate estimates and systematic overestimation of recent divergence times. Molecular biology and evolution 2005, 22, 1561–1568. [Google Scholar] [CrossRef] [PubMed]
- Aiewsakun, P.; Katzourakis, A. Time-dependent rate phenomenon in viruses. Journal of virology 2016, 90, 7184–7195. [Google Scholar] [CrossRef]
- Duchêne, S.; et al. Analyses of evolutionary dynamics in viruses are hindered by a time-dependent bias in rate estimates. Proceedings of the Royal Society B: Biological Sciences 2014, 281. [Google Scholar] [CrossRef]
- Bromham, L.; et al. Bayesian molecular dating: opening up the black box. Biological Reviews 2018, 93, 1165–1191. [Google Scholar] [CrossRef]
- Menardo, F.; et al. The molecular clock of Mycobacterium tuberculosis. PLoS pathogens 2019, 15, e1008067. [Google Scholar] [CrossRef] [PubMed]
- Mármol-Sánchez, E.; et al. Historical RNA expression profiles from the extinct Tasmanian tiger. Genome research 2023, 33, 1299–1316. [Google Scholar] [CrossRef]
- Smith, O.; et al. A complete ancient RNA genome: identification, reconstruction and evolutionary history of archaeological Barley Stripe Mosaic Virus. Scientific reports 2014, 4, 4003. [Google Scholar] [CrossRef]
- Smith, O.; et al. Ancient RNA from Late Pleistocene permafrost and historical canids shows tissue-specific transcriptome survival. PLoS biology 2019, 17, e3000166. [Google Scholar] [CrossRef]
- Fromm, B.; et al. Ancient microRNA profiles of 14,300-yr-old canid samples confirm taxonomic origin and provide glimpses into tissue-specific gene regulation from the Pleistocene. RNA 2021, 27, 324–334. [Google Scholar] [CrossRef] [PubMed]
- Peyambari, M.; et al. A 1,000-year-old RNA virus. Journal of virology 2019, 93, 10–1128. [Google Scholar] [CrossRef] [PubMed]
- Keene, A.H.; Stenglein, M.D. Viral metagenomics of 100-year-old museum specimens highlights the long-term stability of RNA. bioRxiv 2024. [Google Scholar] [CrossRef]
- Mármol-Sánchez, E.; et al. Ancient RNA expression profiles from the extinct woolly mammoth. Cell 2025, 189, 52–69. [Google Scholar] [CrossRef]
- Guzmán-Solís, A.A.; et al. A glimpse into the past: What ancient viral genomes reveal about human history. Annual Review of Virology 2023, 10, 49–75. [Google Scholar] [CrossRef]
- Düx, A.; et al. Measles virus and rinderpest virus divergence dated to the rise of large cities. Science (New York, NY) 2020, 368, 1367. [Google Scholar] [CrossRef] [PubMed]
- Gad, Y.Z.; et al. Insights from ancient DNA analysis of Egyptian human mummies: clues to disease and kinship. Human Molecular Genetics 2021, 30, R24–R28. [Google Scholar] [CrossRef]
- Lipson, M.; et al. Ancient DNA and deep population structure in sub-Saharan African foragers. Nature 2022, 603, 290–296. [Google Scholar] [CrossRef]
- Brielle, E.S.; et al. Entwined African and Asian genetic roots of medieval peoples of the Swahili coast. Nature 2023, 615, 866–873. [Google Scholar] [CrossRef]
- Zink, A.R.; et al. Molecular history of tuberculosis from ancient mummies and skeletons. International Journal of Osteoarchaeology 2007, 17, 380–391. [Google Scholar] [CrossRef]
- Crubézy, E.; et al. Identification of Mycobacterium DNA in an Egyptian Pott’s disease of 5400 years old. Comptes Rendus de l’Académie des Sciences-Series III-Sciences de la Vie 1998, 321, 941–951. [Google Scholar] [CrossRef] [PubMed]
- Lalremruata, A.; et al. Molecular identification of falciparum malaria and human tuberculosis co-infections in mummies from the Fayum depression (Lower Egypt). PloS one 2013, 8, e60307. [Google Scholar] [CrossRef] [PubMed]
- Nerlich, A.G.; et al. Plasmodium falciparum in ancient Egypt. Emerging infectious diseases 2008, 14, 1317. [Google Scholar] [CrossRef] [PubMed]
- Khairat, R.; et al. First insights into the metagenome of Egyptian mummies using next-generation sequencing. Journal of applied genetics 2013, 54, 309–325. [Google Scholar] [CrossRef]
- Lu, J.; Salzberg, S.L. Removing contaminants from databases of draft genomes. PLoS computational biology 2018, 14, e1006277. [Google Scholar] [CrossRef]
- Matheson, C.D.; et al. Molecular confirmation of Schistosoma and family relationship in two ancient Egyptian mummies. Verlag Dr. Friedrich Pfeil 2014, 39–47. [Google Scholar]
- Zink, A.R.; et al. Leishmaniasis in ancient Egypt and upper Nubia. Emerging infectious diseases 2006, 12, 1616. [Google Scholar] [CrossRef]
- L’Hôte, L.; et al. An 8000 years old genome reveals the Neolithic origin of the zoonosis Brucella melitensis. Nature Communications 2024, 15, 6132. [Google Scholar] [CrossRef]
- Majander, K.; et al. Ancient Bacterial Genomes Reveal a High Diversity of Treponema pallidum Strains in Early Modern Europe. Current Biology 2020, 30, 3788–3803.e10. [Google Scholar] [CrossRef]
- Angelakis, E.; et al. Rickettsia felis: the complex journey of an emergent human pathogen. Trends in Parasitology 2016, 32, 554–564. [Google Scholar] [CrossRef]
- Sarris, P. The Justinianic plague: origins and effects. Continuity and change 2002, 17, 169–182. [Google Scholar] [CrossRef]
- Esquivel Gomez, L.R.; et al. Phylogenetic analysis of the origin and spread of plague in Madagascar. PLoS Neglected Tropical Diseases 2023, 17, e0010362. [Google Scholar] [CrossRef]
- Rakotobe Harimanana, R.; et al. Bioecology of fleas with a focus on the plague vector Xenopsylla Brasiliensis in Mandritsara district, Northern Madagascar. Scientific Reports 2025, 15, 24297. [Google Scholar] [CrossRef]
- Mediannikov, O.; et al. Common epidemiology of Rickettsia felis infection and malaria, Africa. Emerging infectious diseases 2013, 19, 1775. [Google Scholar] [CrossRef] [PubMed]
- Maina, A.N.; et al. Rickettsia felis infection in febrile patients, western Kenya, 2007–2010. Emerging infectious diseases 2012, 18, 328. [Google Scholar] [CrossRef] [PubMed]
- Jakobsson, M.; et al. Homo sapiens-specific evolution unveiled by ancient southern African genomes. Nature 2025. [Google Scholar] [CrossRef] [PubMed]
- Phillipson, D.W. African archaeology; Cambridge University Press: Cambridge, UK, 2005. [Google Scholar]
- Li, S.; et al. Genetic variation reveals large-scale population expansion and migration during the expansion of Bantu-speaking peoples. Proceedings of the Royal Society B: Biological Sciences 2014, 281, 20141448. [Google Scholar] [CrossRef]
| Pathogen (genome type) | Disease | Method | Sampling material | Location | Reference |
|---|---|---|---|---|---|
| Mycobacterium tuberculosis complex (dsDNA) | Tuberculosis | PCR | Mummified tissue/bones | Egypt | [71,72,73] |
| Mycobacterium leprae (dsDNA) | Leprae | NGS | Mummified tissue/bones | Egypt | [43] |
| HBV (partially dsDNA) | Hepatitis B | NGS | Mummified tissue/bones | Egypt | [43] |
| Plasmodium falciparum (dsDNA) | Malaria | PCR and NGS | Mummified tissue/bones | Egypt | [73,74,75] |
| Toxoplasma gondii (dsDNA) | Toxoplasmosis | PCR | Embalmed heads | Egypt | [75] |
| Schistosoma mansoni / S. haematobium (dsDNA) | Schistosomiasis | PCR | Canopic liver | Egypt | [77] |
| Leishmania donovani (dsDNA) | Leishmaniasis | PCR | Mummified tissue/bones | Egypt | [78] |
| Rickettsia felis (dsDNA) | Rickettsiosis | NGS | Petrous bone | South Africa | [10] |
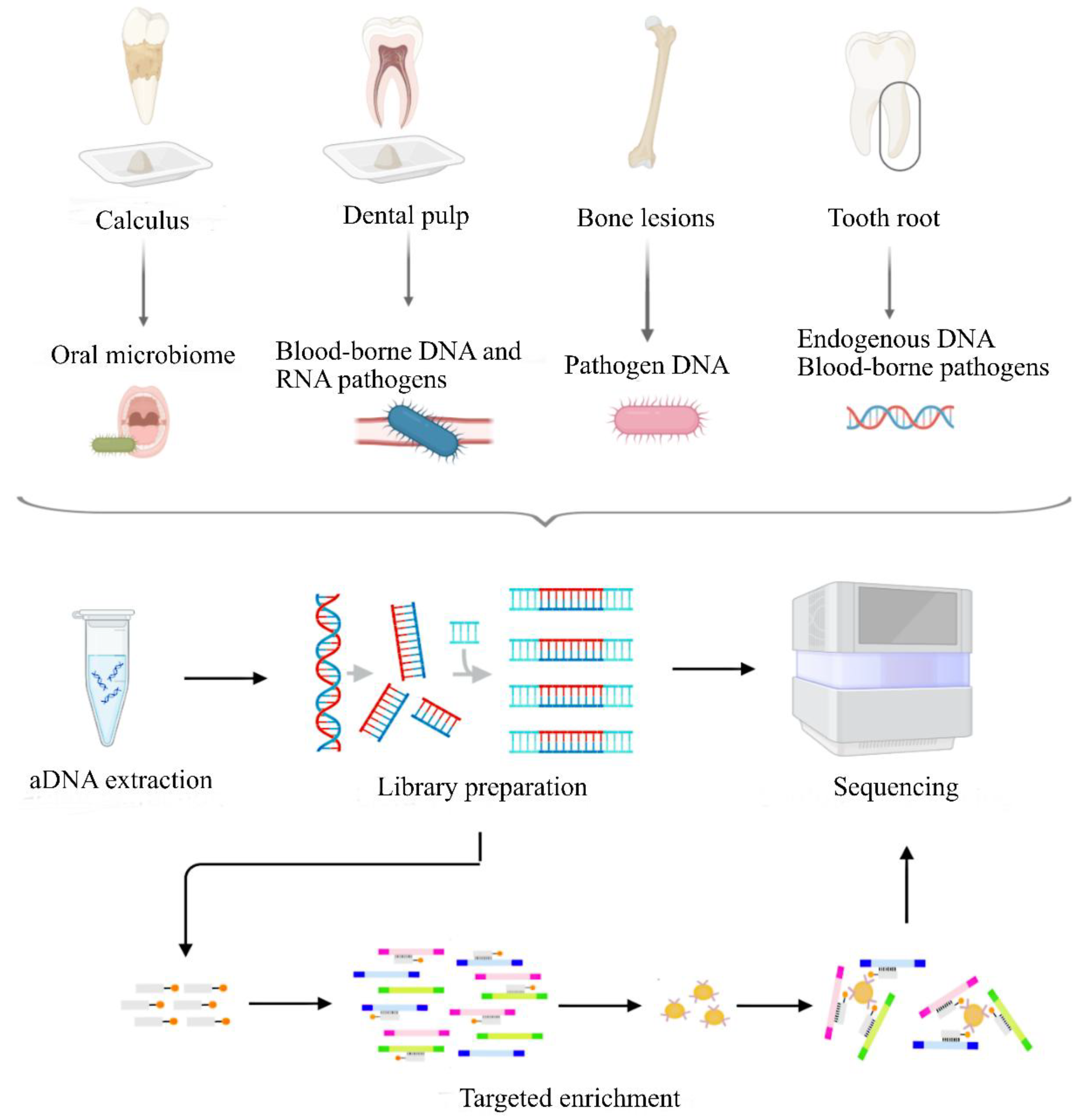
| Parasites | Bacteria | DNA viruses | RNA viruses | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Blood borne | Oral-fecal | Mucosal | Blood borne | Oral-fecal | Mucosal | Blood borne | Oral-fecal | Mucosal | Blood borne | Oral-fecal | Mucosal | |
| Teeth | ●● | ○ | ● | ●●● | ● | ●● | ●● | ○ | ● | ○ | ○ | ○ |
| Bone lesions | ● | ○ | ○ | ●●● | ○ | ● | ● | ○ | ○ | ○ | ○ | ○ |
| Dental calculus | ● | ●●● | ●● | ●●● | ●●● | ●●● | ● | ● | ● | ○ | ○ | ○ |
| Sediment (skeletal remains associated) | ● | ● | ○ | ●● | ● | ● | ● | ● | ○ | ○ | ○ | ○ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





