Submitted:
19 February 2026
Posted:
03 March 2026
You are already at the latest version
Abstract
Keywords:
Highlights
- MEMS captures ecosystem SOC and NEE dynamics across diverse U.S. grazing lands
- The Bayesian framework used enabled the quantification of full model uncertainty
- Interpretation of predicted grazing outcomes can be limited without uncertainty quantification
- Quantifying model and measurement uncertainty is key for model-data comparison
1. Introduction
2. Material and Methods
2.1. The MEMS Model
2.2. Data Used for Modeling
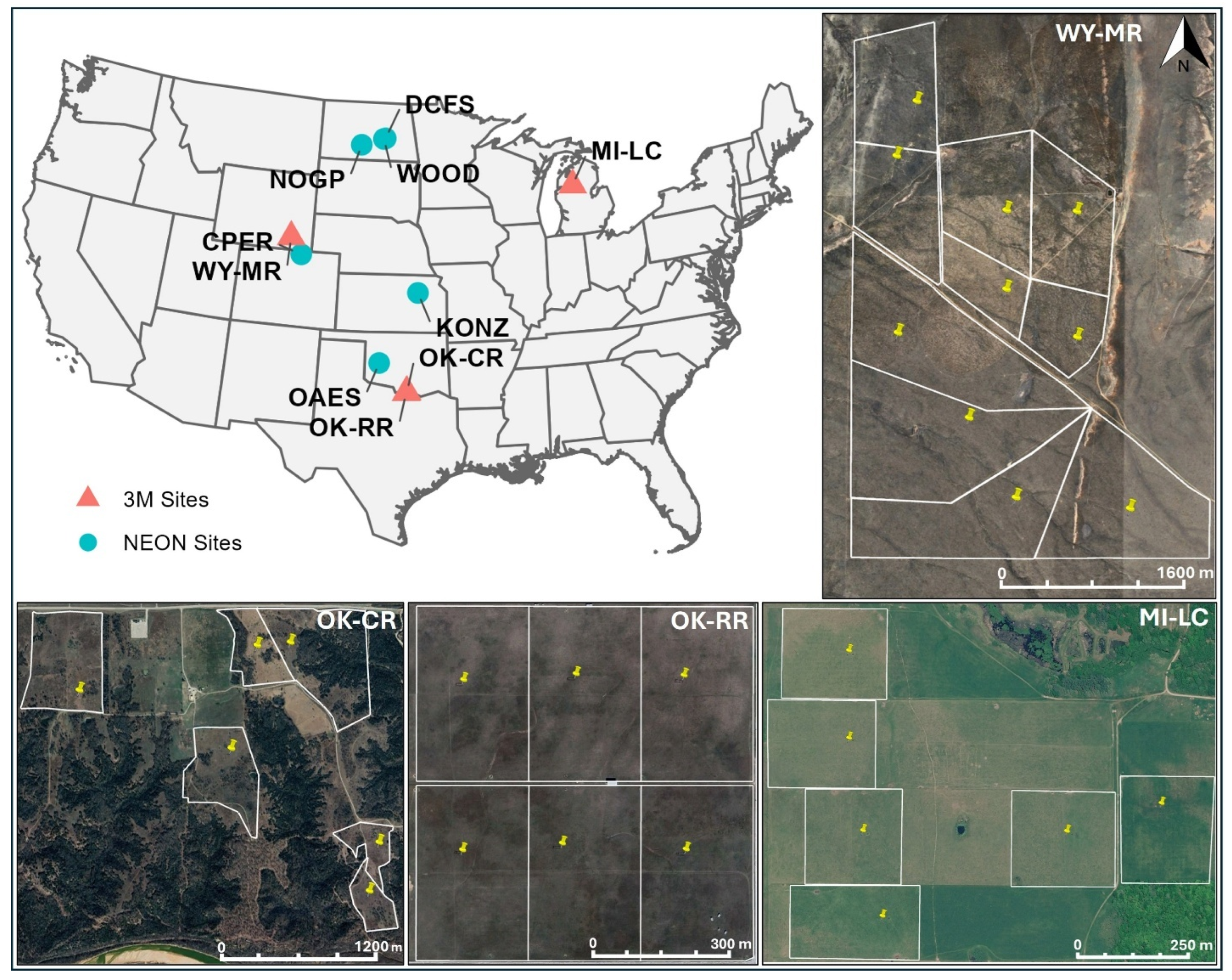
2.3. NEON Sites
2.4. 3M Experimental Sites
2.4.1. Flux Tower Set-up and Measurements
2.4.2. Plant Aboveground Biomass and Leaf Area Index Measured Data
2.4.3. Soil Sampling and Analysis
2.4.4. Soil Water Content Measurements
2.5. Plant Parameterization
2.6. Bayesian Framework: Model Calibration and Uncertainty Quantification
2.7. Model Set-up
2.7.1. NEON Sites
2.7.2. 3M Experimental Sites
2.8. Statistical Analysis
3. Results
3.1. Plant Parameterization for the NEON and 3M Experimental Sites
3.2. Model Performance on Soil Temperature and Soil Water Content at the 3M Experimental Sites
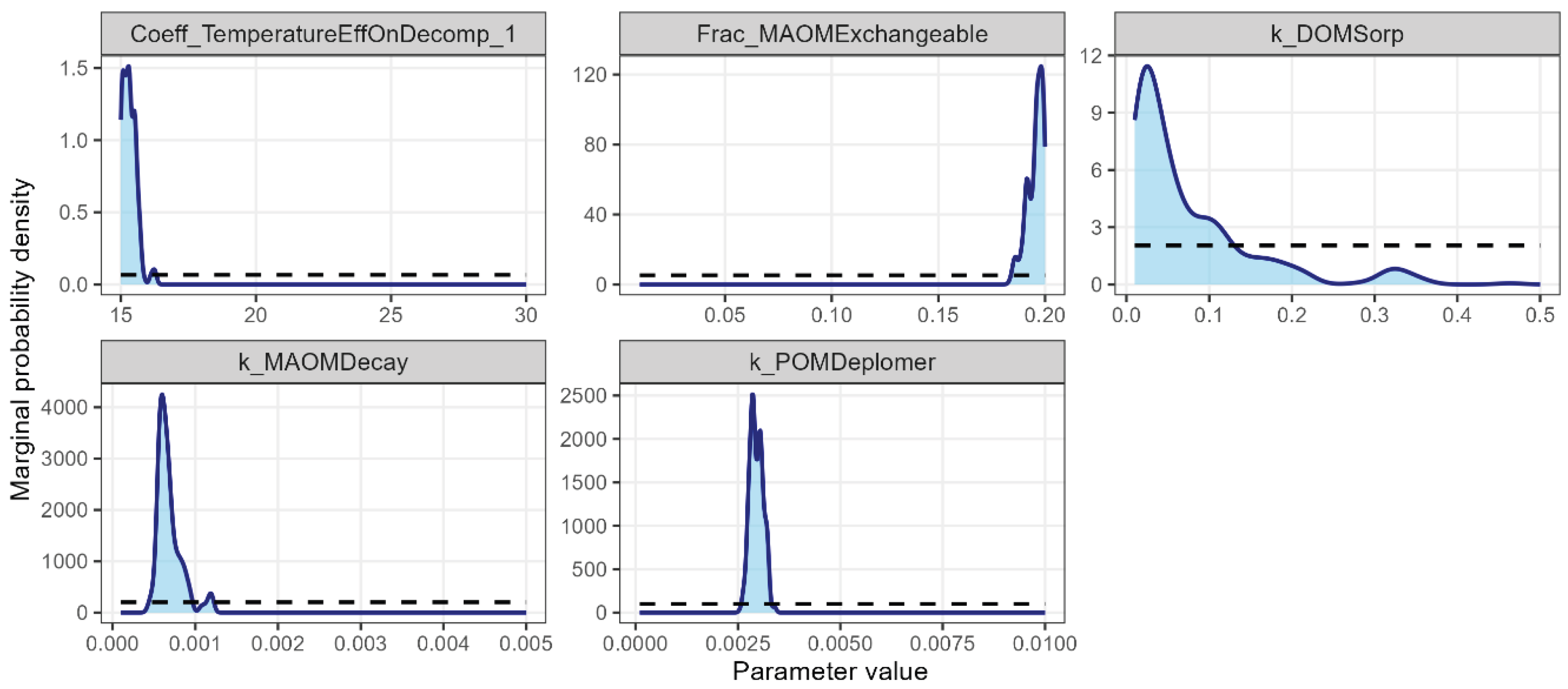
3.3. Bayesian Calibration Using the NEON Sites
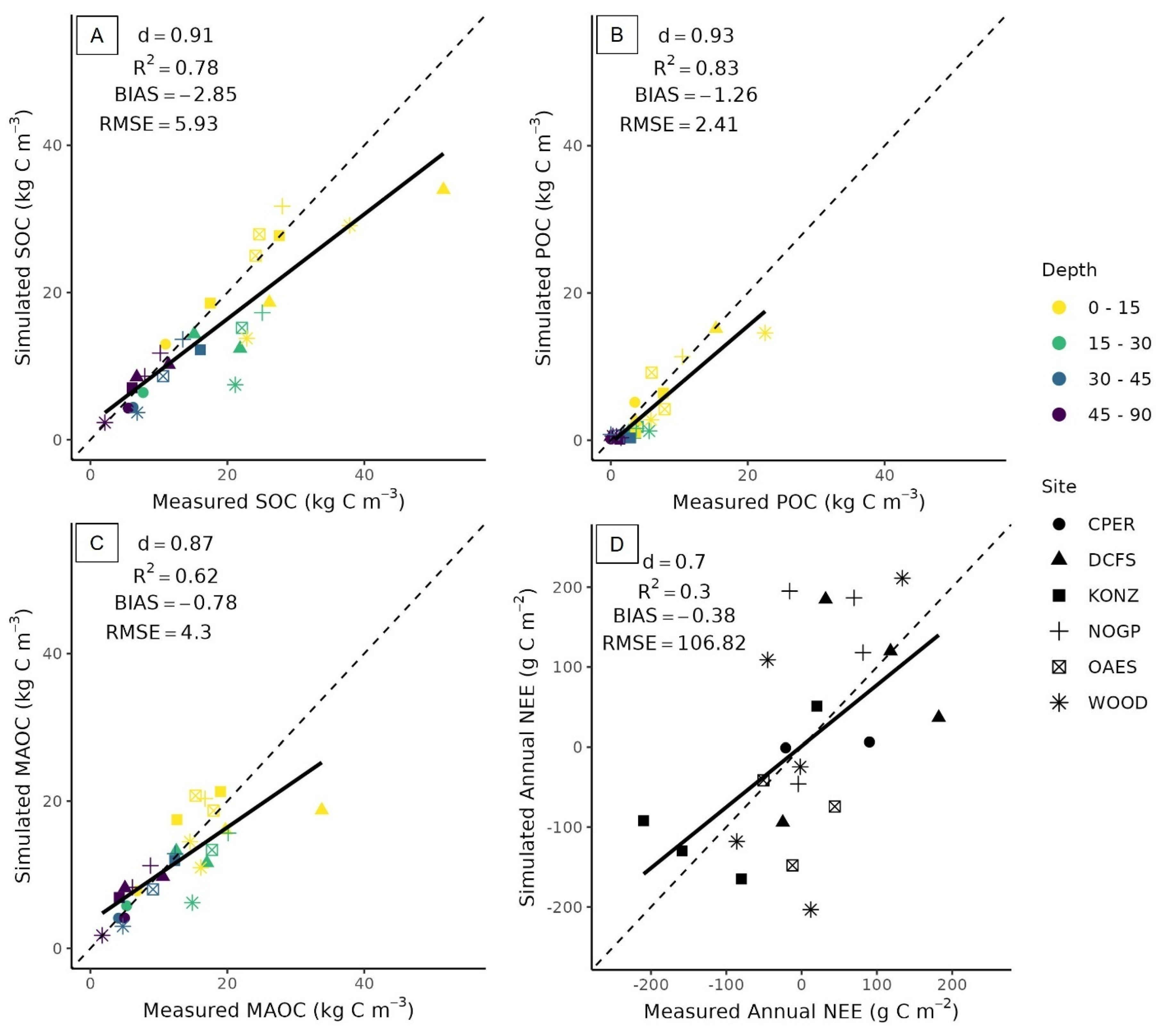
3.4. Model Evaluation
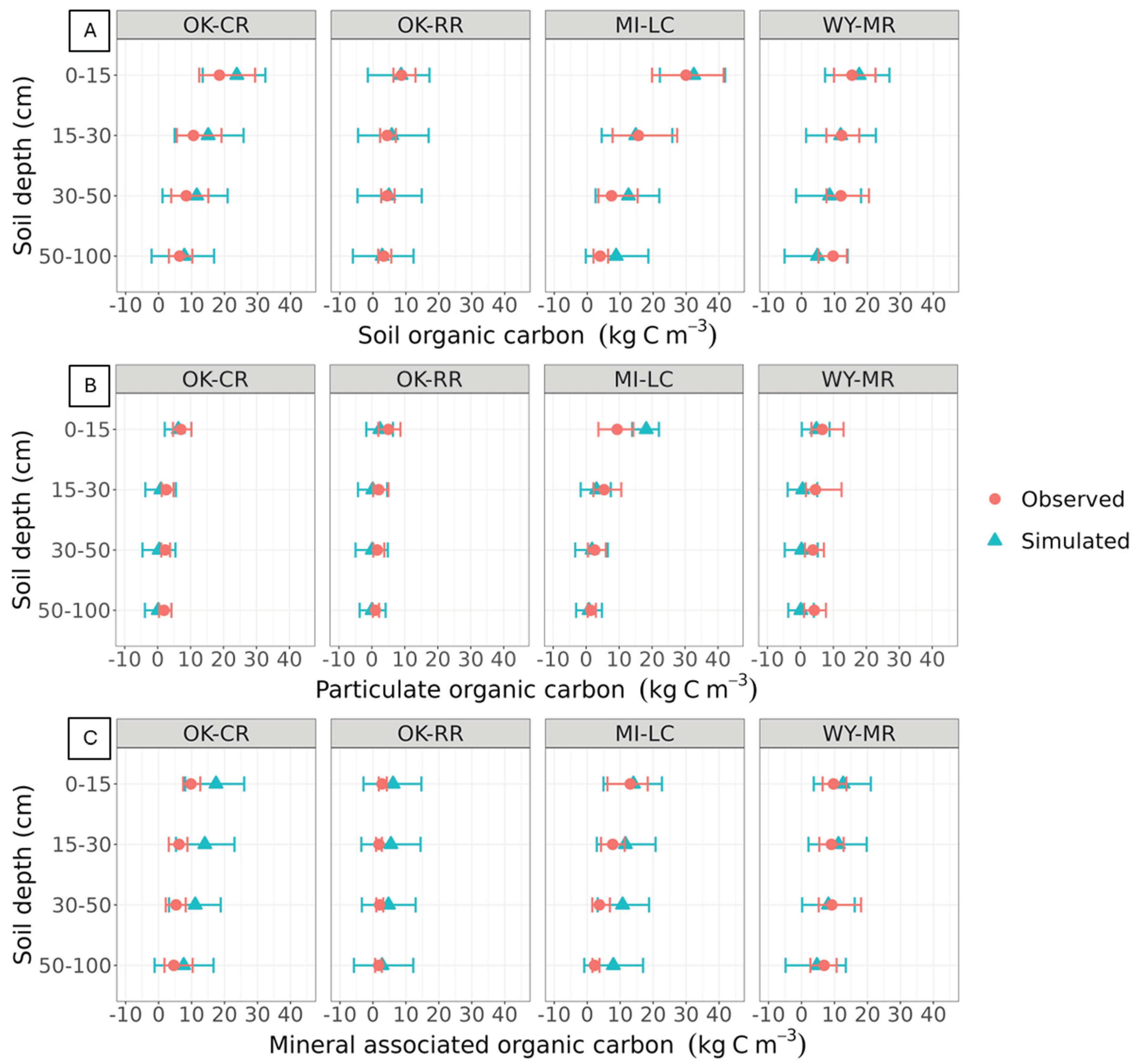
3.4.1. Soil Organic Carbon Dynamics at the 3M Experimental Sites
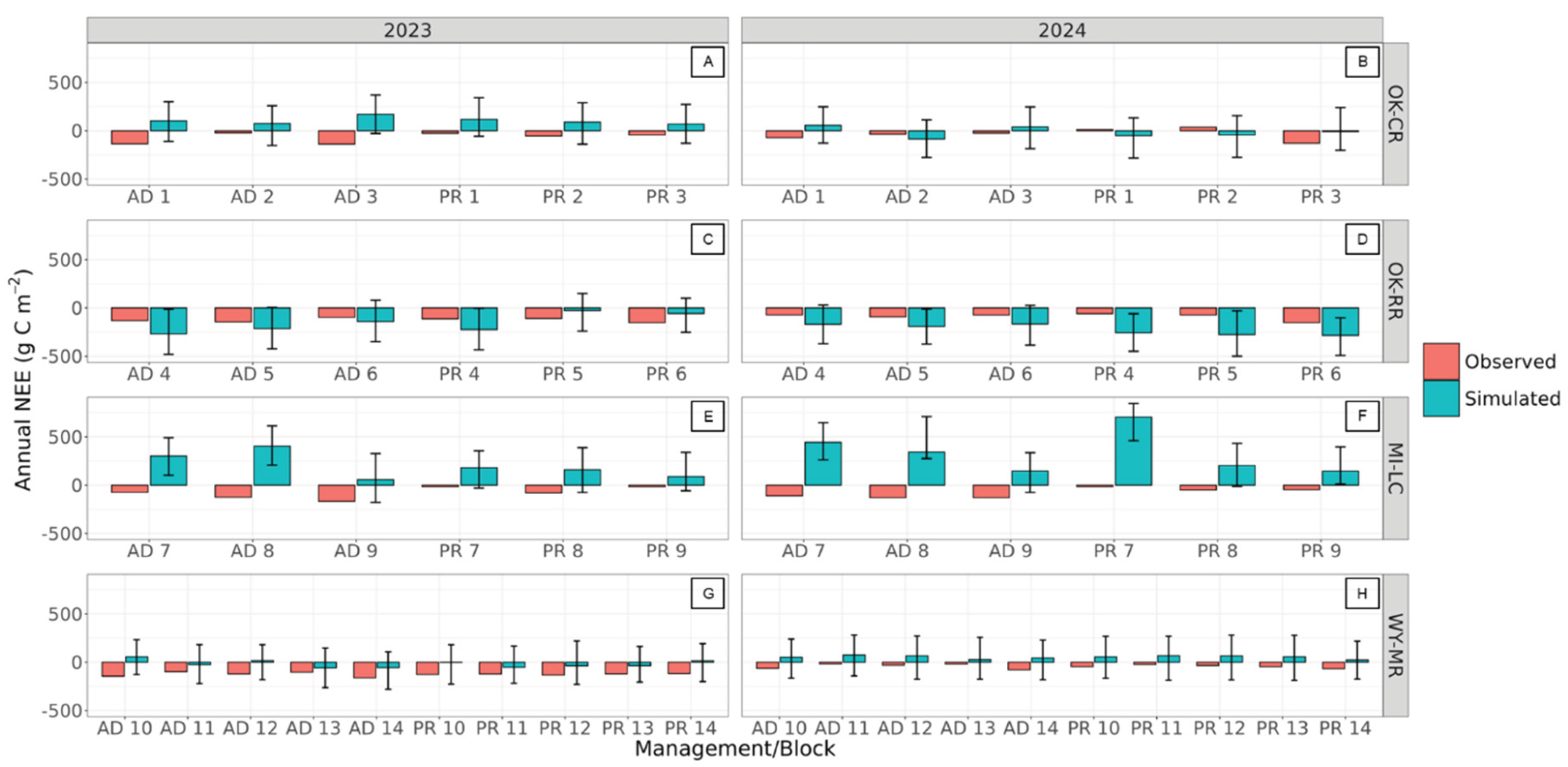
3.4.2. Ecosystem Fluxes at the 3M Experimental Sites
4. Discussion
4.1. Model Performance Representing Plant Growth Dynamics at NEON and 3M Experimental Sites
4.2. Soil Temperature and Soil Water Content Dynamics at 3M Experimental Sites
4.3. Model Calibration Using the NEON Sites
4.4. Model Performance Representing Soil Organic Carbon Dynamics at 3M Experimental Sites
4.5. Model Performance on Gross Plant Productivity and Ecosystem Respiration at 3M Experimental Sites
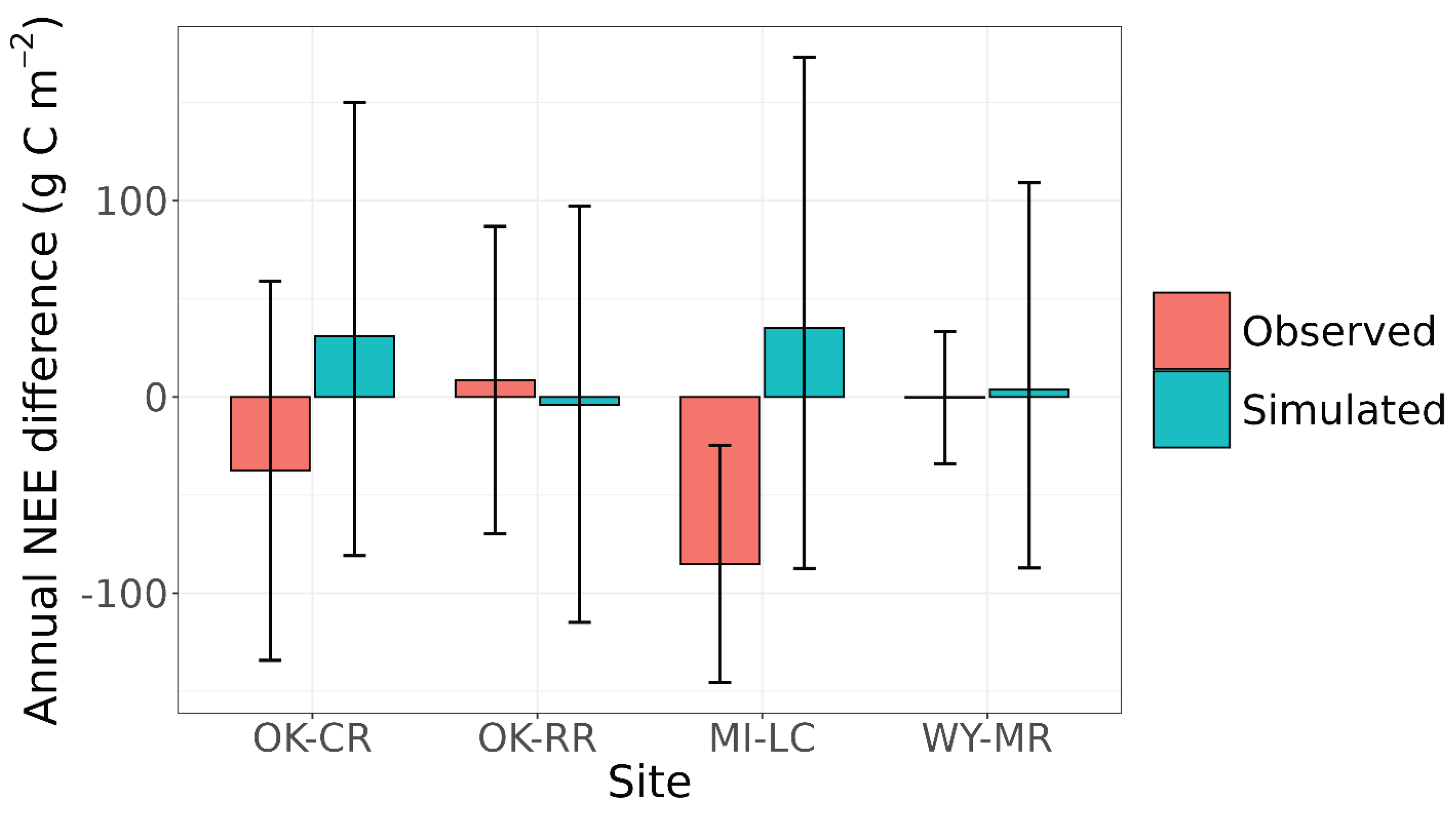
4.6. Net Ecosystem Exchange at 3M Experimental Sites
5. Conclusions
Author Contributions
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Panmao, Z. Climate Change 2022: Mitigation of Climate Change; Contribution of Working Group III to the Sixth Assessment Report of the Intergovernmental Panel on Climate Change; IPCC Sixth Assessment Report; IPCC: Geneva, Switzerland, 2022. [CrossRef]
- Zhang, C.; Dong, Q.; Chu, H.; Shi, J.; Li, S.; Wang, Y.; Yang, X. Grassland Community Composition Response to Grazing Intensity Under Different Grazing Regimes. Rangel. Ecol. Manag. 2018, 71, 196–204. [CrossRef]
- Ren, S.; Terrer, C.; Li, J.; Cao, Y.; Yang, S.; Liu, D. Historical impacts of grazing on carbon stocks and climate mitigation opportunities. Nat. Clim. Chang. 2024, 14, 380–386. [CrossRef]
- Sanderman, J.; Hengl, T.; Fiske, G.J. Soil carbon debt of 12,000 years of human land use. Proc. Natl. Acad. Sci. 2017, 114, 9575–9580. [CrossRef]
- Lal, R. Digging deeper: A holistic perspective of factors affecting soil organic carbon sequestration in agroecosystems. Glob. Chang. Biol. 2018, 24, 3285–3301. [CrossRef]
- FAO. Land statistics 2001–2022 – Global, regional and country trends. FAOSTAT Analytical Briefs, No. 88. Rome. [Internet]. . Available from: https://openknowledge.fao.org/handle/20.500.14283/cd1484en.
- Bai, Y.; Cotrufo, M.F. Grassland soil carbon sequestration: Current understanding, challenges, and solutions. Science 2022, 377, 603–608. [CrossRef]
- McSherry, M.E.; Ritchie, M.E. Effects of grazing on grassland soil carbon: a global review. Glob. Chang. Biol. 2013, 19, 1347–1357. [CrossRef]
- Paustian, K.; Lehmann, J.; Ogle, S.; Reay, D.; Robertson, G.P.; Smith, P. Climate-smart soils. Nature 2016, 532, 49–57. [CrossRef]
- Bradford, M.A.; Kuebbing, S.E.; Polussa, A.; Sanderman, J.; Oldfield, E.E. Upstream data need to prove soil carbon as a climate solution. Nat. Clim. Chang. 2025, 15, 1013–1016. [CrossRef]
- Dong M, de Kroon H. Plasticity in Morphology and Biomass Allocation in Cynodon dactylon, a Grass Species Forming Stolons and Rhizomes. Oikos [Internet]. 70(1), 99 (1994). Available from: https://www.jstor.org/stable/3545704?origin=crossref.
- Turner, C.L.; Seastedt, T.R.; Dyer, M.I. Maximization of Aboveground Grassland Production: The Role of Defoliation Frequency, Intensity, and History. Ecol. Appl. 1993, 3, 175–186. [CrossRef]
- Milchunas, D.G.; Varnamkhasti, A.S.; Lauenroth, W.K.; Goetz, H. Forage quality in relation to long-term grazing history, current-year defoliation, and water resource. Oecologia 1995, 101, 366–374. [CrossRef]
- Pavlů, V.; Hejcman, M.; Pavlů, L.; Gaisler, J.; Nežerková, P. Effect of continuous grazing on forage quality, quantity and animal performance. Agric. Ecosyst. Environ. 2006, 113, 349–355. [CrossRef]
- Stanley, P.L.; Wilson, C.; Patterson, E.; Machmuller, M.B.; Cotrufo, M.F. Ruminating on soil carbon: Applying current understanding to inform grazing management. Glob. Chang. Biol. 2024, 30, e17223. [CrossRef]
- Chang, J.F.; Viovy, N.; Vuichard, N.; Ciais, P.; Wang, T.; Cozic, A.; Lardy, R.; Graux, A.-I.; Klumpp, K.; Martin, R.; et al. Incorporating grassland management in ORCHIDEE: model description and evaluation at 11 eddy-covariance sites in Europe. Geosci. Model Dev. 2013, 6, 2165–2181. [CrossRef]
- Lawton, D.; Leahy, P.; Kiely, G.; Byrne, K.A.; Calanca, P. Modeling of net ecosystem exchange and its components for a humid grassland ecosystem. J. Geophys. Res. Biogeosciences 2006, 111. [CrossRef]
- Takeda, N.; Rowlings, D.; Parton, W.; Grace, L.; Day, K.; Nguyen, T.; Grace, P. Soil carbon sequestration potential in subtropical grasslands estimated by DayCent-CABBI. Soil Sci. Soc. Am. J. 2025, 89. [CrossRef]
- E Campbell, E.; Paustian, K. Current developments in soil organic matter modeling and the expansion of model applications: a review. Environ. Res. Lett. 2015, 10, 123004. [CrossRef]
- Zhang Y, Santos RS, Hamilton EK, et al. Modeling the Effects of Livestock Rotation Frequency on Forage Production and Soil Carbon Using a Spatially Explicit Grazing Distribution [Internet]. (2026). Available from: https://www.preprints.org/manuscript/202602.0923/v1.
- Le Noë, J.; Manzoni, S.; Abramoff, R.; Bölscher, T.; Bruni, E.; Cardinael, R.; Ciais, P.; Chenu, C.; Clivot, H.; Derrien, D.; et al. Soil organic carbon models need independent time-series validation for reliable prediction. Commun. Earth Environ. 2023, 4, 1–8. [CrossRef]
- Stanley P, Kuhl A, Rowntree J, Maciel I. Merging Rigor and Relevance in Grazingland Research: A Comprehensive Social-Ecological Monitoring Approach. Under review. (2026).
- Baldocchi, D. Measuring fluxes of trace gases and energy between ecosystems and the atmosphere – the state and future of the eddy covariance method. Glob. Chang. Biol. 2014, 20, 3600–3609. [CrossRef]
- Eugster, W.; Merbold, L. Eddy covariance for quantifying trace gas fluxes from soils. SOIL 2015, 1, 187–205. [CrossRef]
- Baldocchi, D.D. How eddy covariance flux measurements have contributed to our understanding of Global Change Biology. Glob. Chang. Biol. 2019, 26, 242–260. [CrossRef]
- Oldfield, E.E.; Lavallee, J.M.; Blesh, J.; Bradford, M.A.; Cameron-Harp, M.; Cotrufo, M.F.; Eagle, A.J.; Eash, L.; Even, R.J.; Kuebbing, S.E.; et al. Greenhouse gas mitigation on croplands: clarifying the debate on knowns, unknowns and risks to move forward with effective management interventions. Carbon Manag. 2024, 15. [CrossRef]
- Gottschalk, P.; Wattenbach, M.; Neftel, A.; Fuhrer, J.; Jones, M.; Lanigan, G.; Davis, P.; Campbell, C.; Soussana, J.-F.; Smith, P. The role of measurement uncertainties for the simulation of grassland net ecosystem exchange (NEE) in Europe. Agric. Ecosyst. Environ. 2007, 121, 175–185. [CrossRef]
- Vuichard, N.; Soussana, J.; Ciais, P.; Viovy, N.; Ammann, C.; Calanca, P.; Clifton-Brown, J.; Fuhrer, J.; Jones, M.; Martin, C. Estimating the greenhouse gas fluxes of European grasslands with a process-based model: 1. Model evaluation from in situ measurements. Glob. Biogeochem. Cycles 2007, 21. [CrossRef]
- Zhang, Y.; Lavallee, J.M.; Robertson, A.D.; Even, R.; Ogle, S.M.; Paustian, K.; Cotrufo, M.F. Simulating measurable ecosystem carbon and nitrogen dynamics with the mechanistically defined MEMS 2.0 model. Biogeosciences 2021, 18, 3147–3171. [CrossRef]
- Hu, T.; Malone, S.L.; Rumpel, C.; Chabbi, A. Maximizing soil organic carbon stocks through optimal ploughing and renewal strategies in (Ley) grassland. Commun. Earth Environ. 2024, 5, 1–10. [CrossRef]
- Puche, N.; Senapati, N.; Flechard, C.R.; Klumpp, K.; Kirschbaum, M.U.; Chabbi, A. Modeling Carbon and Water Fluxes of Managed Grasslands: Comparing Flux Variability and Net Carbon Budgets between Grazed and Mowed Systems. Agronomy 2019, 9, 183. [CrossRef]
- Sándor, R.; Barcza, Z.; Hidy, D.; Lellei-Kovács, E.; Ma, S.; Bellocchi, G. Modelling of grassland fluxes in Europe: Evaluation of two biogeochemical models. Agric. Ecosyst. Environ. 2016, 215, 1–19. [CrossRef]
- Thentu, T.L.; Forster, D.; Li, Y.; Virkajärvi, P.; Harrison, M.T.; Mitra, B.; Deng, J.; Korhonen, P.; Shurpali, N. DNDC modelling of greenhouse gas exchange from a boreal legume grassland under organic and mineral nitrogen management. J. Environ. Manag. 2025, 390, 126344. [CrossRef]
- Ogle, S.M.; Breidt, F.J.; Easter, M.; Williams, S.; Killian, K.; Paustian, K. Scale and uncertainty in modeled soil organic carbon stock changes for US croplands using a process-based model. Glob. Chang. Biol. 2010, 16, 810–822. [CrossRef]
- Gurung, R.B.; Ogle, S.M.; Breidt, F.J.; Williams, S.A.; Parton, W.J. Bayesian calibration of the DayCent ecosystem model to simulate soil organic carbon dynamics and reduce model uncertainty. Geoderma 2020, 376. [CrossRef]
- Mathers, C.; Black, C.K.; Segal, B.D.; Gurung, R.B.; Zhang, Y.; Easter, M.J.; Williams, S.; Motew, M.; Campbell, E.E.; Brummitt, C.D.; et al. Validating DayCent-CR for cropland soil carbon offset reporting at a national scale. Geoderma 2023, 438. [CrossRef]
- Ogle, S.M.; Breidt, F.J.; Easter, M.; Williams, S.; Paustian, K. An empirically based approach for estimating uncertainty associated with modelling carbon sequestration in soils. Ecol. Model. 2007, 205, 453–463. [CrossRef]
- Zhang, Y.; Arabi, M.; Paustian, K. Analysis of parameter uncertainty in model simulations of irrigated and rainfed agroecosystems. Environ. Model. Softw. 2020, 126. [CrossRef]
- van Oijen, M. Bayesian Methods for Quantifying and Reducing Uncertainty and Error in Forest Models. Curr. For. Rep. 2017, 3, 269–280. [CrossRef]
- Van Oijen, M.; Rougier, J.; Smith, R. Bayesian calibration of process-based forest models: bridging the gap between models and data. Tree Physiol. 2005, 25, 915–927. [CrossRef]
- Lu, D.; Ricciuto, D.; Walker, A.; Safta, C.; Munger, W. Bayesian calibration of terrestrial ecosystem models: a study of advanced Markov chain Monte Carlo methods. Biogeosciences 2017, 14, 4295–4314. [CrossRef]
- Hinckley, E.S.; Bonan, G.B.; Bowen, G.J.; Colman, B.P.; Duffy, P.A.; Goodale, C.L.; Houlton, B.Z.; Marín-Spiotta, E.; Ogle, K.; Ollinger, S.V.; et al. The soil and plant biogeochemistry sampling design for The National Ecological Observatory Network. Ecosphere 2016, 7, e01234. [CrossRef]
- Rocci, K.S.; Cotrufo, M.F.; Ernakovich, J.; Foster, E.; Frey, S.; Georgiou, K.; Grandy, A.S.; Malhotra, A.; Reich, P.B.; Schlerman, E.P.; et al. Bridging 20 Years of Soil Organic Matter Frameworks: Empirical Support, Model Representation, and Next Steps. J. Geophys. Res. Biogeosciences 2024, 129. [CrossRef]
- Cotrufo, M.F.; Soong, J.L.; Horton, A.J.; Campbell, E.E.; Haddix, M.L.; Wall, D.H.; Parton, W.J. Formation of soil organic matter via biochemical and physical pathways of litter mass loss. Nat. Geosci. 2015, 8, 776–779. [CrossRef]
- Liang, C.; Schimel, J.P.; Jastrow, J.D. The importance of anabolism in microbial control over soil carbon storage. Nat. Microbiol. 2017, 2, 17105. [CrossRef]
- Sokol, N.W.; Sanderman, J.; Bradford, M.A. Pathways of mineral-associated soil organic matter formation: Integrating the role of plant carbon source, chemistry, and point of entry. Glob. Chang. Biol. 2018, 25, 12–24. [CrossRef]
- Zhang, Y.; King, A.E.; Hamilton, E.; Cotrufo, M.F. Representing cropping systems with the MEMS 2 ecosystem model. Agron. J. 2024, 116, 2328–2345. [CrossRef]
- Santos, R.S.; Hamilton, E.K.; Stanley, P.L.; Paustian, K.; Cotrufo, M.F.; Zhang, Y. Simulating adaptive grazing management on soil organic carbon in the Southeast U.S.A. using MEMS 2. J. Environ. Manag. 2024, 365, 121657. [CrossRef]
- Santos, R.S.; Hamilton, E.K.; Stanley, P.L.; Paustian, K.; Cotrufo, M.F.; Zhang, Y. Simulating adaptive grazing management on soil organic carbon in the Southeast U.S.A. using MEMS 2. J. Environ. Manag. 2024, 365, 121657. [CrossRef]
- Running S, Mu Q, Zhao M. MODIS/Terra Gross Primary Productivity 8-Day L4 Global 500m SIN Grid V061 [Data set]. NASA Land Processes Distributed Active Archive Center. (2021).
- NEON. NEON Eddy-Covariance Tower Data (DP4.00200.001) for sites US-xDC, US-xAE, US-xCP, US-xKZ, US-xNG, and US-xWD. Dataset accessed via AmeriFlux Portal - Lawrence Berkeley National Laboratory (https://ameriflux.lbl.gov). [Internet]. . Available from: https://ameriflux.lbl.gov.
- Novick, K.; Biederman, J.; Desai, A.; Litvak, M.; Moore, D.; Scott, R.; Torn, M. The AmeriFlux network: A coalition of the willing. Agric. For. Meteorol. 2018, 249, 444–456. [CrossRef]
- Pastorello G, Trotta C, Canfora E, et al. The FLUXNET2015 dataset and the ONEFlux processing pipeline for eddy covariance data. Sci. Data [Internet]. 7(1), 225 (2020). Available from: https://www.nature.com/articles/s41597-020-0534-3.
- Kormann, R.; Meixner, F.X. An Analytical Footprint Model For Non-Neutral Stratification. Boundary-Layer Meteorol. 2001, 99, 207–224. [CrossRef]
- Hersbach, H.; Bell, B.; Berrisford, P.; Hirahara, S.; Horányi, A.; Muñoz-Sabater, J.; Nicolas, J.; Peubey, C.; Radu, R.; Schepers, D.; et al. The ERA5 global reanalysis. Q. J. R. Meteorol. Soc. 2020, 146, 1999–2049. [CrossRef]
- Spencer, S.; Ogle, S.M.; Breidt, F.J.; Goebel, J.J.; Paustian, K. Designing a national soil carbon monitoring network to support climate change policy: a case example for US agricultural lands. Greenh. Gas Meas. Manag. 2011, 1, 167–178. [CrossRef]
- Poeplau, C.; Vos, C.; Don, A. Soil organic carbon stocks are systematically overestimated by misuse of the parameters bulk density and rock fragment content. SOIL 2017, 3, 61–66. [CrossRef]
- Leuthold SJ, Haddix ML, Lavallee J, Cotrufo MF. Physical fractionation techniques [Internet]. In: Reference Module in Earth Systems and Environmental Sciences, Elsevier, 1–13 (2022). Available from: https://linkinghub.elsevier.com/retrieve/pii/B9780128229743000677.
- Samson, M.; Chantigny, M.H.; Vanasse, A.; Menasseri-Aubry, S.; Angers, D.A. Coarse mineral-associated organic matter is a pivotal fraction for SOM formation and is sensitive to the quality of organic inputs. Soil Biol. Biochem. 2020, 149. [CrossRef]
- Cotrufo, M.F.; Haddix, M.L.; Mullen, J.L.; Zhang, Y.; McKay, J.K. Deepening Root Inputs: Potential Soil Carbon Accrual From Breeding for Deeper Rooted Maize. Glob. Chang. Biol. 2024, 30, e17591. [CrossRef]
- NEON. Soil physical and chemical properties, Megapit (DP1.00096.001), RELEASE-2025. Dataset accessed from https://data.neonscience.org/data-products/DP1.00096.001/RELEASE-2025 on October 10, 2025. [Internet]. Available from: https://data.neonscience.org/data-products/DP1.00096.001/RELEASE-2025. [CrossRef]
- Vrugt JA, Ter Braak CJF. DREAM (D) : an adaptive Markov Chain Monte Carlo simulation algorithm to solve discrete, noncontinuous, and combinatorial posterior parameter estimation problems. Hydrol. Earth Syst. Sci. [Internet]. 15(12), 3701–3713 (2011). Available from: https://hess.copernicus.org/articles/15/3701/2011/.
- Bates D, Mächler M, Bolker B, Walker S. Fitting Linear Mixed-Effects Models Using lme4. J. Stat. Softw. [Internet]. 67(1) (2015). Available from: http://www.jstatsoft.org/v67/i01/.
- Guillaume J, Andrews F. dream: DiffeRential Evolution Adaptive Metropolis. R package version 0.4-2. URL http://CRAN.R-project.org/package=dream [Internet]. CRAN: Contributed Packages (2012). Available from: http://CRAN.R-project.org/package=dream.
- R Core Team. R: A Language and Environment for Statistical Computing_. R Foundation for Statistical Computing. at https://www.r-project.org/ (2024).
- Gelman A, Rubin DB. Inference from Iterative Simulation Using Multiple Sequences. Statistical Science [Internet]. 7(4) (1992). Available from: https://projecteuclid.org/journals/statistical-science/volume-7/issue-4/Inference-from-Iterative-Simulation-Using-Multiple-Sequences/10.1214/ss/1177011136.full.
- NEON. Soil physical and chemical properties, distributed initial characterization (DP1.10047.001), RELEASE-2025. Dataset accessed from https://data.neonscience.org/data-products/DP1.10047.001/RELEASE-2025 on October 10, 2025. [CrossRef]
- Abatzoglou, J.T. Development of gridded surface meteorological data for ecological applications and modelling. Int. J. Climatol. 2013, 33, 121–131. [CrossRef]
- Guyette, R.P.; Stambaugh, M.C.; Dey, D.C.; Muzika, R.-M. Predicting Fire Frequency with Chemistry and Climate. Ecosystems 2012, 15, 322–335. [CrossRef]
- Knapp, A.K.; Briggs, J.M.; Collins, S.L.; Hartnett, D.C.; Johnson, L.C.; Towne, E.G. The Keystone Role of Bison in North American Tallgrass PrairieBison increase habitat heterogeneity and alter a broad array of plant, community, and ecosystem processes. BioScience 1999, 49, 39–50. [CrossRef]
- NEON. Site management and event reporting (DP1.10111.001), RELEASE-2025. Dataset accessed from https://data.neonscience.org/data-products/DP1.10111.001/RELEASE-2025 on October 10, 2025. [CrossRef]
- Beaudette, D.; Skovlin, J.; Roecker, S.; Brown, A. soilDB: Soil Database Interface. R package version 2.8.13. (2025). Available from: https://CRAN.R-project.org/package=soilDB.
- Schaap, M.G.; Leij, F.J.; van Genuchten, M.T. rosetta: a computer program for estimating soil hydraulic parameters with hierarchical pedotransfer functions. J. Hydrol. 2001, 251, 163–176. [CrossRef]
- Romero-Ruiz A, Rivero MJ, Milne A, et al. Grazing livestock move by Lévy walks: Implications for soil health and environment. J. Environ. Manage. 345 (2023).
- Google. Google Maps Satellite Imagery [Data set]. [Internet]. (2025). Available from: https://www.google.com/maps.
- M., S.E.; J., J.; B., S.; M., D.; C., K. Above-ground resource use increases with plant species richness in experimental grassland ecosystems. Funct. Ecol. 2000, 14, 326–337. [CrossRef]
- Riedo, M.; Grub, A.; Rosset, M.; Fuhrer, J. A pasture simulation model for dry matter production, and fluxes of carbon, nitrogen, water and energy. Ecol. Model. 1998, 105, 141–183. [CrossRef]
- Van Oijen, M.; Barcza, Z.; Confalonieri, R.; Korhonen, P.; Kröel-Dulay, G.; Lellei-Kovács, E.; Louarn, G.; Louault, F.; Martin, R.; Moulin, T.; et al. Incorporating Biodiversity into Biogeochemistry Models to Improve Prediction of Ecosystem Services in Temperate Grasslands: Review and Roadmap. Agronomy 2020, 10, 259. [CrossRef]
- Kipling, R.P.; Virkajärvi, P.; Breitsameter, L.; Curnel, Y.; De Swaef, T.; Gustavsson, A.-M.; Hennart, S.; Höglind, M.; Järvenranta, K.; Minet, J.; et al. Key challenges and priorities for modelling European grasslands under climate change. Sci. Total. Environ. 2016, 566-567, 851–864. [CrossRef]
- Parton, W.J.; Hartman, M.; Ojima, D.; Schimel, D. DAYCENT and its land surface submodel: description and testing. Glob. Planet. Chang. 1998, 19, 35–48. [CrossRef]
- Maurer, G.E.; Bowling, D.R. Seasonal snowpack characteristics influence soil temperature and water content at multiple scales in interior western U.S. mountain ecosystems. Water Resour. Res. 2014, 50, 5216–5234. [CrossRef]
- Pulina, A.; Lai, R.; Salis, L.; Seddaiu, G.; Roggero, P.P.; Bellocchi, G. Modelling pasture production and soil temperature, water and carbon fluxes in Mediterranean grassland systems with the Pasture Simulation model. Grass Forage Sci. 2017, 73, 272–283. [CrossRef]
- Wu, L.; Liu, H.; Liang, B.; Zhu, X.; Cao, J.; Wang, Q.; Jiang, L.; Cressey, E.L.; Quine, T.A. A process-based model reveals the restoration gap of degraded grasslands in Inner Mongolian steppe. Sci. Total. Environ. 2022, 806, 151324. [CrossRef]
- Rowley, M.C.; Grand, S.; Verrecchia, É.P. Calcium-mediated stabilisation of soil organic carbon. Biogeochemistry 2018, 137, 27–49. [CrossRef]
- Glaser, B.; Amelung, W. Pyrogenic carbon in native grassland soils along a climosequence in North America. Glob. Biogeochem. Cycles 2003, 17. [CrossRef]
- Cotrufo, M.F.; Lavallee, J.M. Soil organic matter formation, persistence, and functioning: A synthesis of current understanding to inform its conservation and regeneration. In: Advances in Agronomy, Academic Press Inc., 1–66 (2022).
- Poeplau, C.; Don, A.; Six, J.; Kaiser, M.; Benbi, D.; Chenu, C.; Cotrufo, M.F.; Derrien, D.; Gioacchini, P.; Grand, S.; et al. Isolating organic carbon fractions with varying turnover rates in temperate agricultural soils – A comprehensive method comparison. Soil Biol. Biochem. 2018, 125, 10–26. [CrossRef]
- Manzoni, S.; Schimel, J.P. Advances in modelling soil microbial dynamics. Soil Biol. Biochem. 2024, 197. [CrossRef]
- Stanley, P.; Spertus, J.; Chiartas, J.; Stark, P.B.; Bowles, T. Valid inferences about soil carbon in heterogeneous landscapes. Geoderma 2023, 430. [CrossRef]
- Hansen, P.M.; Even, R.; King, A.E.; Lavallee, J.; Schipanski, M.; Cotrufo, M.F. Distinct, direct and climate-mediated environmental controls on global particulate and mineral-associated organic carbon storage. Glob. Chang. Biol. 2023, 30, e17080. [CrossRef]
- Lavallee, J.M.; Haddix, M.L.; Swan, A.; Hoover, J.D.; Cotrufo, M.F. Using aridity as an overarching factor to advance understanding of soil organic carbon storage at the continental scale. Biogeochemistry 2025, 168, 1–25. [CrossRef]
- Oikawa, P.; Sturtevant, C.; Knox, S.; Verfaillie, J.; Huang, Y.; Baldocchi, D. Revisiting the partitioning of net ecosystem exchange of CO2 into photosynthesis and respiration with simultaneous flux measurements of 13CO2 and CO2, soil respiration and a biophysical model, CANVEG. Agric. For. Meteorol. 2017, 234-235, 149–163. [CrossRef]
- Reichstein, M.; Falge, E.; Baldocchi, D.; Papale, D.; Aubinet, M.; Berbigier, P.; Bernhofer, C.; Buchmann, N.; Gilmanov, T.; Granier, A.; et al. On the separation of net ecosystem exchange into assimilation and ecosystem respiration: review and improved algorithm. Glob. Chang. Biol. 2005, 11, 1424–1439. [CrossRef]
- Lei, L.; Xia, J.; Li, X.; Huang, K.; Zhang, A.; Chen, S.; Weng, E.; Luo, Y.; Wan, S. Water response of ecosystem respiration regulates future projection of net ecosystem productivity in a semiarid grassland. Agric. For. Meteorol. 2018, 252, 175–191. [CrossRef]
- Kirschbaum, M.U.; Puche, N.J.; Giltrap, D.L.; Liáng, L.L.; Chabbi, A. Combining eddy covariance measurements with process-based modelling to enhance understanding of carbon exchange rates of dairy pastures. Sci. Total. Environ. 2020, 745, 140917. [CrossRef]
- van Ramshorst, J.G.V.; Knohl, A.; Callejas-Rodelas, J.Á.; Clement, R.; Hill, T.C.; Siebicke, L.; Markwitz, C. Lower-cost eddy covariance for CO2 and H2O fluxes over grassland and agroforestry. 2024, 17, 6047–6071. [CrossRef]
- Schmidt, A.; Hanson, C.; Chan, W.S.; Law, B.E. Empirical assessment of uncertainties of meteorological parameters and turbulent fluxes in the AmeriFlux network. J. Geophys. Res. Biogeosciences 2012, 117. [CrossRef]
- Liu, Y.; Stoy, P.; Chu, H.; Hollinger, D.Y.; Ollinger, S.V.; Ouimette, A.P.; Durden, D.J.; Sturtevant, C.; Lucas, B.; Richardson, A.D. A tale of two towers: comparing NEON and AmeriFlux data streams at Bartlett Experimental Forest. Agric. For. Meteorol. 2025, 378. [CrossRef]
- Falvo, G.; Zhang, Y.; Abraha, M.; Mosier, S.; Su, Y.; Lei, C.; Chen, J.; Cotrufo, M.F.; Robertson, G.P. Combining Eddy Covariance Towers, Field Measurements, and the MEMS 2 Ecosystem Model Improves Confidence in the Climate Impacts of Bioenergy With Carbon Capture and Storage. GCB Bioenergy 2025, 17. [CrossRef]
- Oren, R.; Hsieh, C.; Stoy, P.; Albertson, J.; Mccarthy, H.R.; Harrell, P.; Katul, G.G. Estimating the uncertainty in annual net ecosystem carbon exchange: spatial variation in turbulent fluxes and sampling errors in eddy-covariance measurements. Glob. Chang. Biol. 2006, 12, 883–896. [CrossRef]
- Ammann, C.; Flechard, C.; Leifeld, J.; Neftel, A.; Fuhrer, J. The carbon budget of newly established temperate grassland depends on management intensity. Agric. Ecosyst. Environ. 2007, 121, 5–20. [CrossRef]
| Site name | Location | Altitude (m) | Koppen's climate classification | Dominant plant | # of blocks | Grazing management | Plot size (ha) | Paddock size (ha)* | Average plot resting period (day) | Length of grazing event (day) | Stocking rate (AU ha-1) |
| OK-CR | Oklahoma - Coffey Ranch | 250 | Humid subtropical (Cfa) | Warm-season grasses (e.g., Schizachyrium scoparium), but it also includes legumes and shrubs | 3 | PR | 11.5 | - | 104 | 1-13 | 1.9-6.3 |
| AD | 7.4 | 0.4-1.5 | 170 | 5-17 | 5-8.4 | ||||||
| OK-RR | Oklahoma - Red River Grazing Facility | 226 | Humid subtropical (Cfa) | Warm-season grasses, particularly bermudagrass (Cynodon dactylon), with some presence of annuals and forbs | 3 | PR | 9.7 | - | 76 | 2-11 | 4.3 |
| AD | 9.7 | 0.97-1.94 | 178 | 1 | 20.5-46.7 | ||||||
| MI-LC | Michigan - Lake City Research Center | 340 | Humid continental (Dfb) | Cool-season grasses (e.g., Dactylis glomerta, Poa pratensis, Bromus inermis) | 3 | PR | 4 | - | 81 | 2-16 | 7.8 |
| AD | 4 | 0.2-2 | 153 | 0.2-1 | 64-425 | ||||||
| WY-MR | Wyoming - McGuire Ranch | 2190 | Arid cold steppe (BSk) | Cool-season grasses (e.g., Pascopyrum smithii, Koeleria macrantha, Poa fendleriana) and sagebrush | 5 | PR | 38.8-148.5 | - | 308 | 70-77 | 0.3 |
| AD | 38.8-148.5 | - | 353 | 8-21 | 0.5-3.5 |
| Dataset | Site name | Variable | Statistical metric | |||
| d | R2 | BIAS | RMSE | |||
| NEON | CPER | GPP (g C m-2 8-day sum) | 0.89 | 0.70 | -2.62 | 5.46 |
| DCFS | GPP (g C m-2 8-day sum) | 0.91 | 0.86 | -6.49 | 11.17 | |
| KONZ | GPP (g C m-2 8-day sum) | 0.95 | 0.86 | 2.18 | 9.94 | |
| NOGP | GPP (g C m-2 8-day sum) | 0.84 | 0.79 | -6.54 | 9.93 | |
| OAES | GPP (g C m-2 8-day sum) | 0.85 | 0.67 | 3.82 | 9.78 | |
| WOOD | GPP (g C m-2 8-day sum) | 0.95 | 0.82 | -1.82 | 9.03 | |
| 3M project | OK-CR | ABG (Mg ha-1) | 0.67 | 0.21 | 0.17 | 0.99 |
| LAI (m2 m-2) | 0.79 | 0.41 | -0.08 | 0.69 | ||
| OK-RR | ABG (Mg ha-1) | 0.64 | 0.13 | 0.07 | 0.71 | |
| LAI (m2 m-2) | 0.63 | 0.14 | 0.09 | 0.53 | ||
| MI-LC | ABG (Mg ha-1) | 0.59 | 0.29 | 0.88 | 1.61 | |
| LAI (m2 m-2) | 0.58 | 0.12 | -0.39 | 1.75 | ||
| WY-MR | ABG (Mg ha-1) | 0.50 | 0.05 | 0.04 | 0.15 | |
| LAI (m2 m-2) | - | - | - | - | ||
| Location | Statistical metric | General model performance | Model evaluation | ||||||||
| Soil temperature10 cm | SWC15 cm | SWC40 cm | SWC60 cm | SOC | POC | MAOC | NEE | GPP | ER | ||
| ºC | cm3 cm-3 | kg C m-3 | g C m-2 week-1 | ||||||||
| OK-CR | d | 0.95 | 0.90 | 0.92 | 0.91 | 0.88 | 0.90 | 0.47 | 0.80 | 0.94 | 0.94 |
| R2 | 0.95 | 0.83 | 0.74 | 0.71 | 0.98 | 1.00 | 0.83 | 0.47 | 0.82 | 0.80 | |
| BIAS | -2.76 | 0.03 | 0.01 | -0.01 | 3.54 | -1.48 | 6.01 | 1.82 | -2.48 | -0.66 | |
| RMSE | 3.72 | 0.06 | 0.04 | 0.04 | 3.87 | 1.54 | 6.35 | 7.57 | 10.23 | 7.11 | |
| OK-RR | Soil temperature10 cm | SWC15 cm | SWC40 cm | SWC60 cm | SOC | POC | MAOC | NEE | GPP | ER | |
| d | 0.94 | 0.86 | 0.86 | 0.86 | 0.96 | 0.65 | 0.25 | 0.68 | 0.80 | 0.73 | |
| R2 | 0.93 | 0.78 | 0.79 | 0.74 | 0.88 | 0.97 | 0.50 | 0.25 | 0.52 | 0.65 | |
| BIAS | -3.84 | -0.04 | -0.03 | -0.01 | 0.33 | -1.71 | 2.66 | -1.60 | -5.76 | -7.36 | |
| RMSE | 4.53 | 0.05 | 0.05 | 0.05 | 0.79 | 1.80 | 2.85 | 7.71 | 11.80 | 9.67 | |
| MI-LC | Soil temperature10 cm | SWC15 cm | SWC40 cm | SWC60 cm | SOC | POC | MAOC | NEE | GPP | ER | |
| d | 0.95 | 0.76 | 0.83 | 0.87 | 0.96 | 0.80 | 0.68 | 0.79 | 0.86 | 0.84 | |
| R2 | 0.94 | 0.54 | 0.54 | 0.59 | 0.95 | 0.86 | 0.88 | 0.54 | 0.68 | 0.79 | |
| BIAS | -2.92 | 0.04 | 0.02 | -0.01 | 2.90 | 1.30 | 4.39 | 6.51 | 3.02 | 9.54 | |
| RMSE | 3.52 | 0.06 | 0.05 | 0.05 | 3.75 | 4.60 | 4.94 | 13.51 | 18.80 | 15.81 | |
| WY-MR | Soil temperature10 cm | SWC10 cm | SWC20 cm | SWC30 cm | SOC | POC | MAOC | NEE | GPP | ER | |
| d | 0.98 | 0.60 | 0.66 | 0.77 | 0.79 | 0.42 | 0.72 | 0.79 | 0.93 | 0.86 | |
| R2 | 0.94 | 0.26 | 0.38 | 0.48 | 0.95 | 0.95 | 0.81 | 0.59 | 0.78 | 0.70 | |
| BIAS | -1.57 | -0.05 | -0.05 | -0.02 | -1.53 | -3.29 | 0.51 | 1.91 | -0.45 | 1.46 | |
| RMSE | 2.83 | 0.07 | 0.06 | 0.04 | 3.14 | 3.41 | 2.20 | 3.82 | 3.70 | 3.08 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.





