Submitted:
27 February 2026
Posted:
02 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
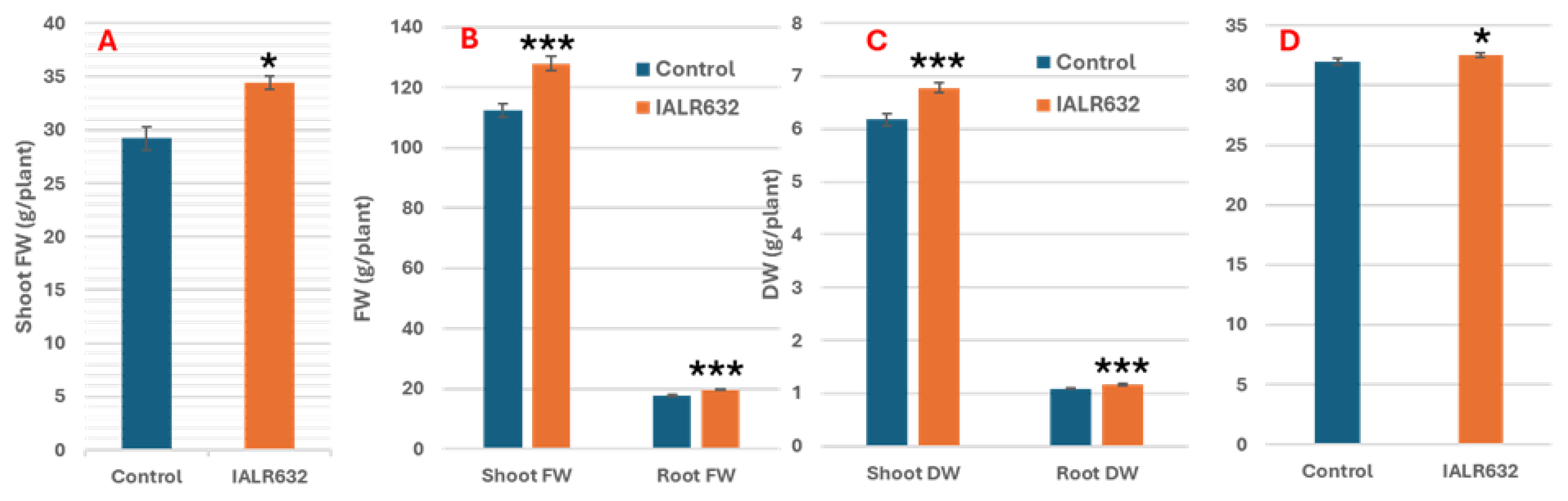
2.1. Lettuce Growth Promotion by IALR632
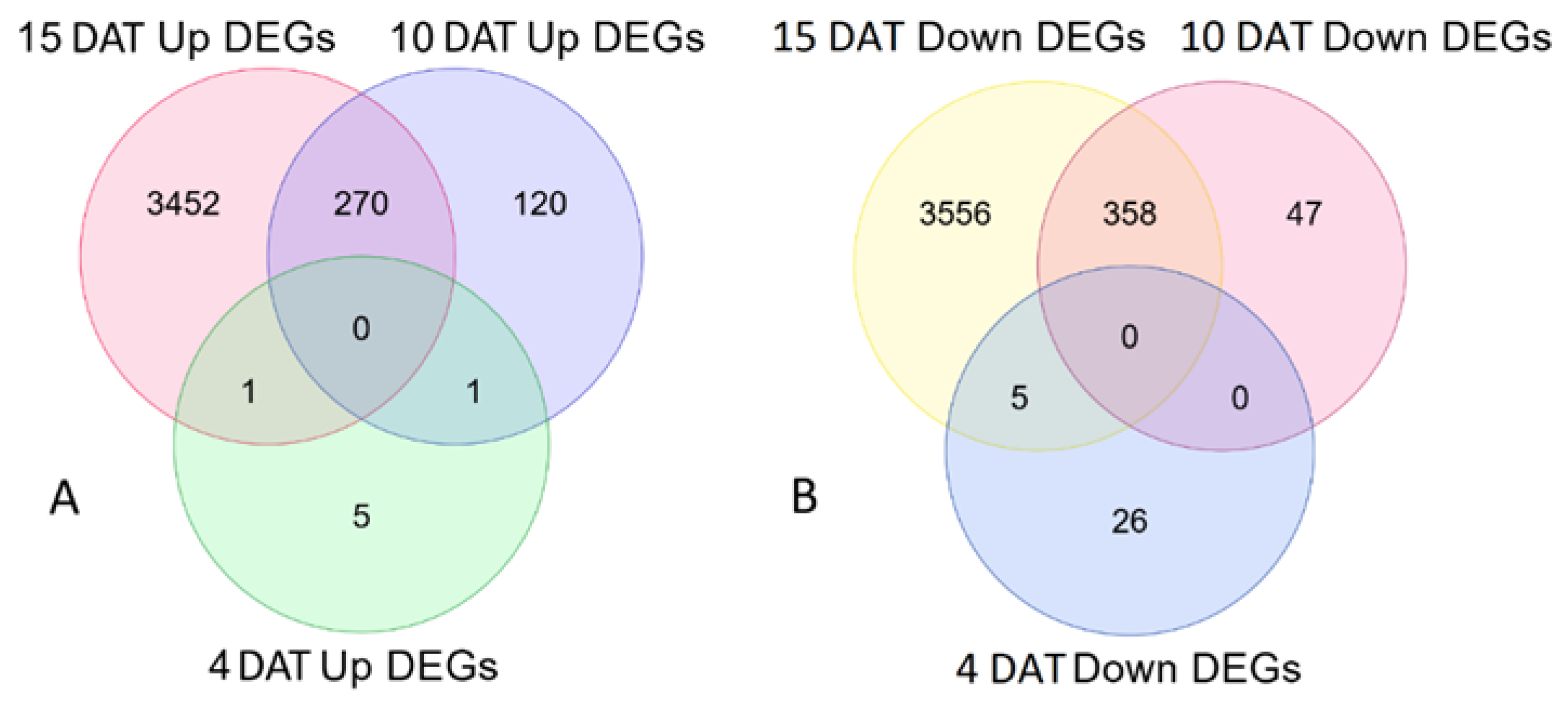
2.2. Lettuce Transcriptome Analyses
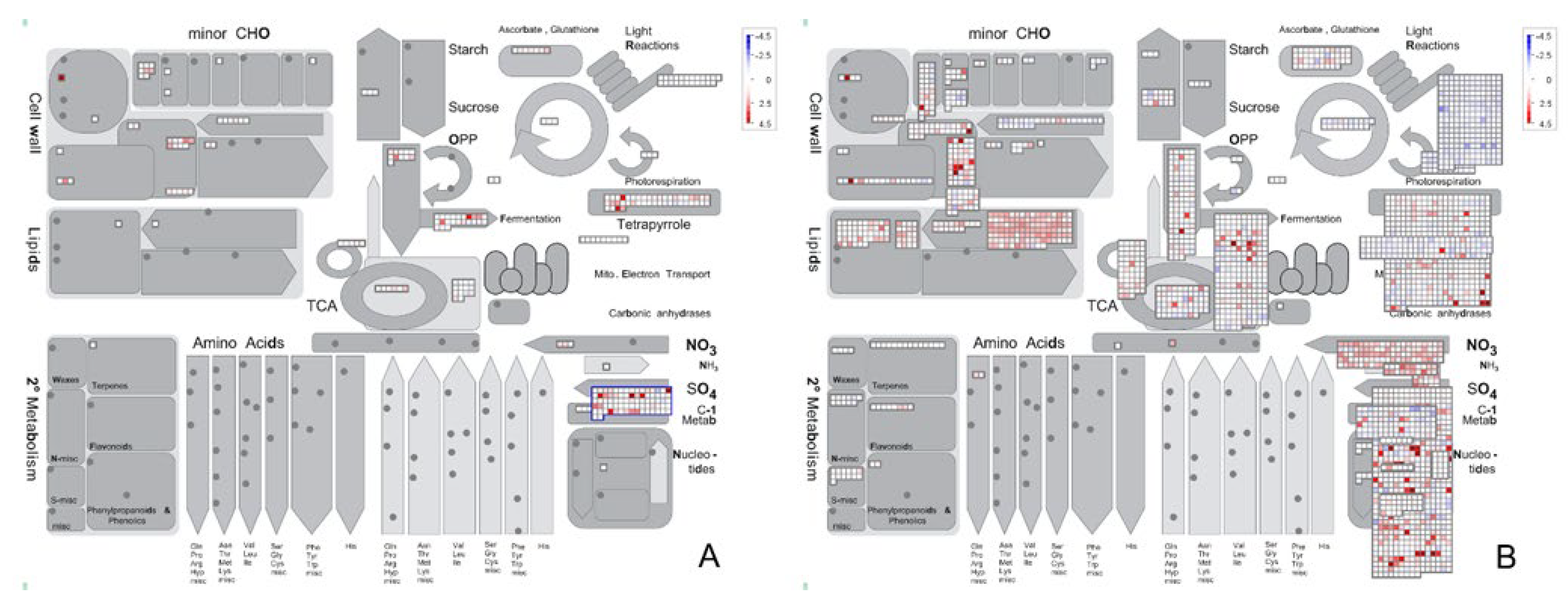
2.3. Metabolic Pathway Changes
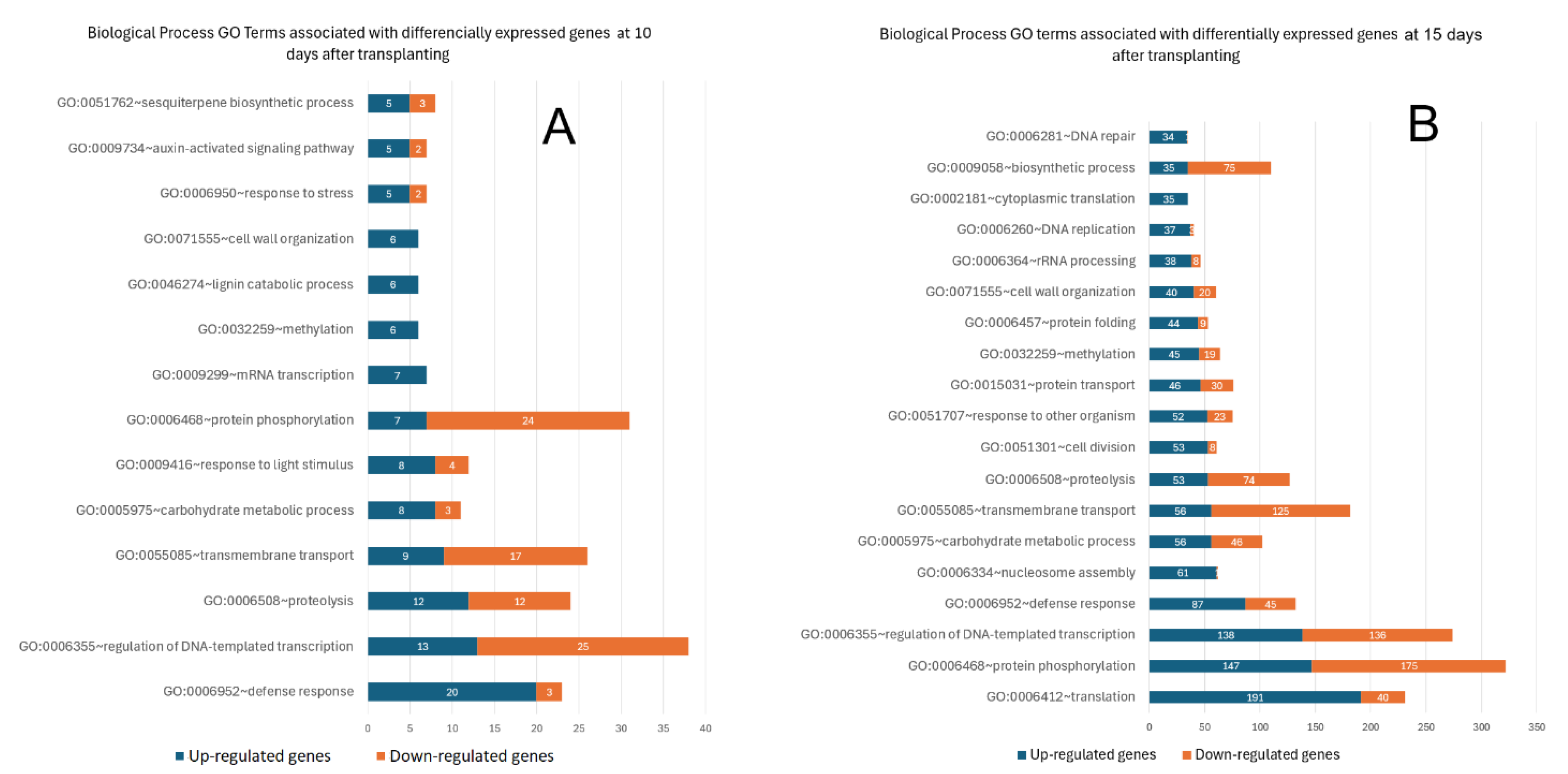
2.4. Biological Process
2.5. Growth and Stress-Related Genes
2.5. Root Microbiome Analysis
2.5.1. Sequencing Data Quality and Filtering
2.5.2. Operational Taxonomic Unit (OTU) Clustering and Abundance
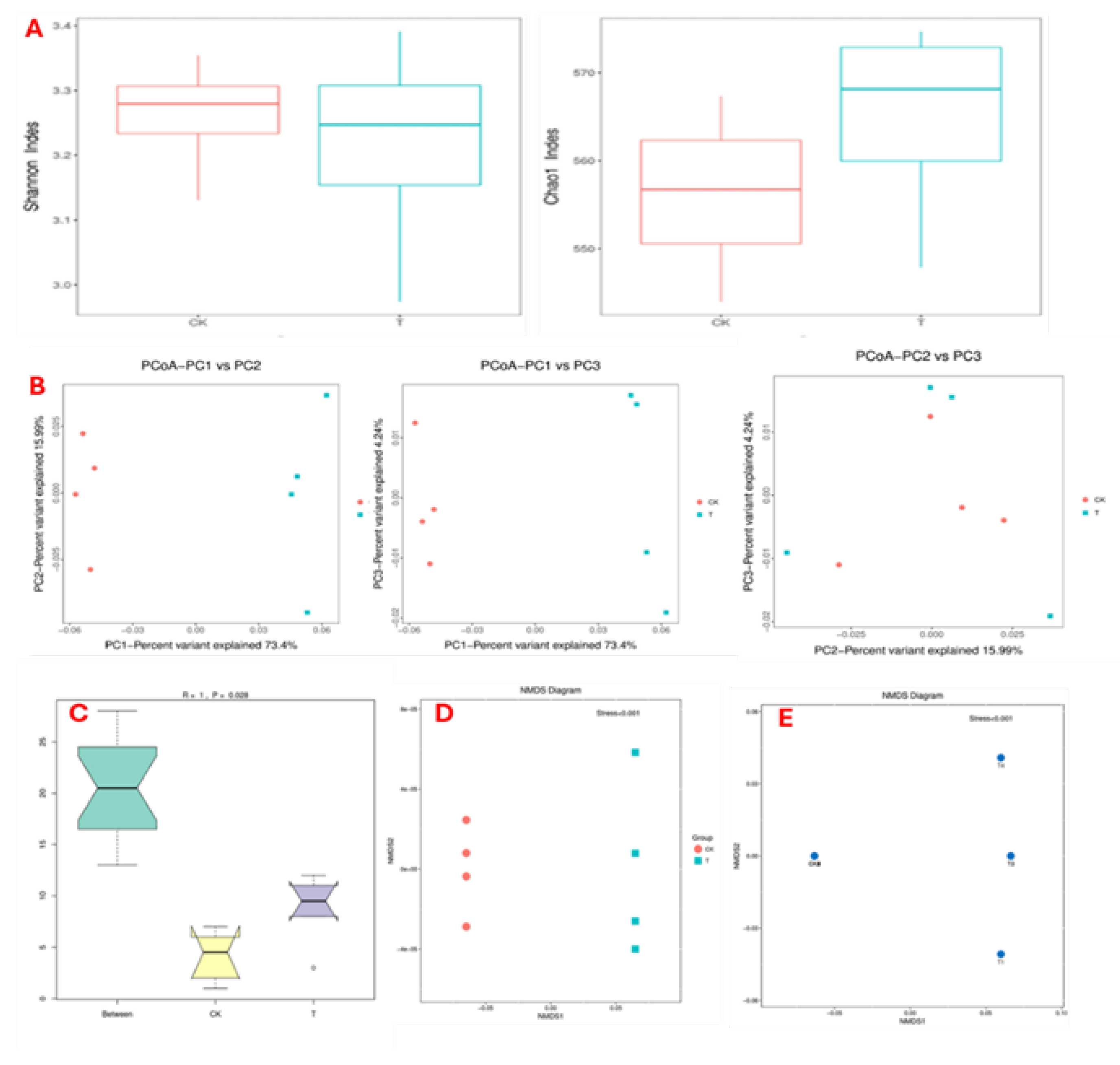
2.5.3. Bacterial Community Diversity
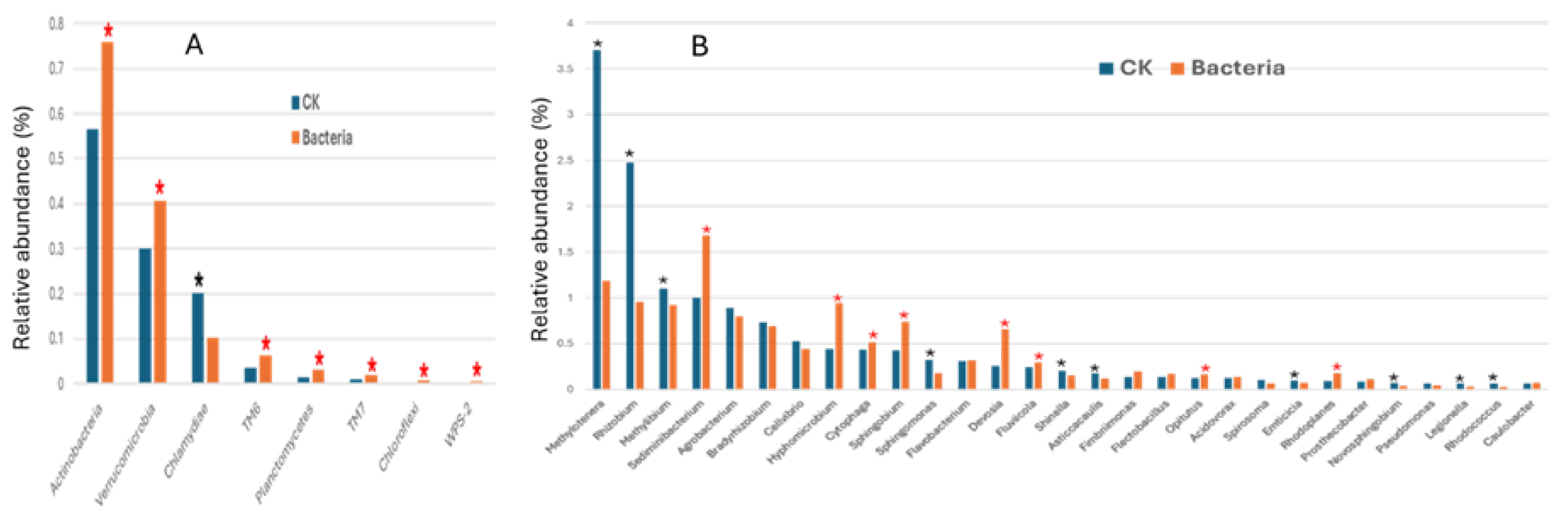
2.5.4. Taxonomic Composition at Phylum and Genus Levels
3. Discussion
3.1. Lettuce Gene Expression Profiling
3.2. Root Microbiome Shifts in Response to IALR632 Inoculation
4. Materials and Methods
4.1. Lettuce Seeds and Germination
4.2. Bacterial Strain and Culture
4.3. Bacterial Inoculation and Plant Transplant
4.4. Greenhouse NFT System and Growth Measurements
4.5. RNA Sampling and Extraction
4.6. RNA Sequencing and Transcriptome Analysis
4.7. MapMan Analysis
4.8. DAVID Analysis
4.9. Root Sampling and DNA Isolation
4.10. Library Construction and Miseq Sequencing
4.11. Bioinformatics Analysis Workflow
4.12. OTU Analysis and Species Annotation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ACC | 1-aminocyclopropane-1-carboxylic acid |
| ANOSIM | Analysis of similarities |
| ACC | 1-aminocyclopropane-1-carboxylic acid |
| ANOSIM | Analysis of similarities |
| BP | Biological process |
| CEA | Controlled environment agriculture |
| CFU | Colony forming unit |
| DAT | Days after transplanting |
| DAVID | Database for Annotation, Visualization, and Integrated Discovery |
| DEG | Differentially expressed genes |
| DET | Differentially expressed transcript |
| DW | Dry weight |
| ERF | Ethylene-responsive transcription factor |
| FW | Fresh weight |
| GO | Gene ontology |
| LB | Lennox broth |
| NFT | Nutrient film technique |
| NMDS | Non-metric multidimensional scaling |
| OD | Optical density |
| OPP | Oxidative pentose phosphate |
| OTU | Operational taxonomic unit |
| PCoA | Principal Coordinates Analysis |
| PGPB | Plant growth-promoting bacteria |
| PGPR | PGPR: Plant growth-promoting rhizobacteria |
| RMTA | Read mapping and transcript assembly |
| SPAD | Soil plant analysis development |
| TCA | Tricarboxylic acid cycle |
References
- Lee, S.; Lee, J. Beneficial bacteria and fungi in hydroponic systems: Types and characteristics of hydroponic food production methods. Sci Hortic 2015, 206–215. [Google Scholar] [CrossRef]
- Dhawi, F. The role of plant growth-promoting microorganisms (PGPMs) and their feasibility in hydroponics and vertical farming. Metabolites 2023, 13, 247. [Google Scholar] [CrossRef]
- Moncada, A.; Miceli, A.; Vetrano, F. Use of plant growth-promoting rhizobacteria (PGPR) and organic fertilization for soilless cultivation of basil. Sci Hortic 2021, 275, 109733. [Google Scholar] [CrossRef]
- Putra, A. M.; Anastasya, N. A.; Rachmawati, S. W.; Yusnawan, E.; Syib`li, M. A.; Trianti, I.; Setiawan, A.; Aini, L.Q. Growth performance and metabolic changes in lettuce inoculated with plant growth promoting bacteria in a hydroponic system. Sci Hortic 2024, 327, 112868. [Google Scholar] [CrossRef]
- Aini, N.; Yamika, W. S. D.; Ulum, B. Effect of nutrient concentration, PGPR and AMF on plant growth, yield and nutrient uptake of hydroponic lettuce. Int J Agric Biol 2019, 21, 175–183. [Google Scholar] [CrossRef]
- Sheridan, C.; Depuydt, P.; De Ro, M.; Petit, C.; Van Gysegem, E.; Delaere, P.; Dixon, M.; Stasiak, M.; Aciksöz, S.B.; Frossard, E.; Paradiso, R. Microbial community dynamics and response to plant growth-promoting microorganisms in the rhizosphere of four common food crops cultivated in hydroponics. Microb Ecol 2017, 73(2), 378–393. [Google Scholar] [CrossRef] [PubMed]
- Thomas, B. O.; Lechner, S. L.; Ross, H. C.; Joris, B. R.; Glick, B. R.; Stegelmeier, A. A. Friends and foes: Bacteria of the hydroponic plant microbiome. Plants 2024, 13, 3069. [Google Scholar] [CrossRef]
- Balthazar, C.; Joly, D. L.; Filion, M. Exploiting beneficial Pseudomonas spp. for cannabis production. Front Microbiol 2022, 12, 833172. [Google Scholar] [CrossRef]
- Alattas, H.; Glick, B. R.; Murphy, D. V.; Scott, C. Harnessing Pseudomonas spp. for sustainable plant crop protection. Front Microbiol 2024, 15, 1485197. [Google Scholar] [CrossRef]
- Lee, S. W.; Ahn, I. P.; Sim, S. Y.; Lee, S. Y.; Seo, M. W.; Kim, S.; Park, S.Y.; Lee, Y.H.; Kang, S. Pseudomonas sp. LSW25R, antagonistic to plant pathogens, promoted plant growth, and reduced blossom-end rot of tomato fruits in a hydroponic system. Eur J Plant Pathol 2010, 126(1), 1–11. [Google Scholar] [CrossRef]
- Lee, S.; An, R.; Grewal, P.; Yu, Z.; Borherova, Z.; Lee, J. High-performing windowfarm hydroponic system: Transcriptomes of fresh produce and microbial communities in response to beneficial bacterial treatment. MPMI 2016, 29(12), 965–976. [Google Scholar] [CrossRef]
- Koprivova, A.; Kopriva, S. Molecular mechanisms of regulation of sulfate assimilation: First steps on a long road. Front Plant Sci 2014, 5, 589. [Google Scholar] [CrossRef]
- Lee, J.; Heo, J.; Lee, E. S.; Kang, H. Y.; Warsadiharja, S. M.; Kim, K.; Kim, K.; Chae, Y.; Pervitasari, A.N.; Priatama, R.A.; Lee, Y.K.; Kim, I.H. Molecular and phenotypic insights into sulfur’s role in enhancing tomato plant growth, stress tolerance, and productivity. Sci Rep 2025, 15, 32773. [Google Scholar] [CrossRef]
- Larkin, R. M. Tetrapyrrole signaling in plants. Front Plant Sci 2016, 7, 1586. [Google Scholar] [CrossRef]
- Terry, M. J.; Smith, A. G. A model for tetrapyrrole synthesis as the primary mechanism for plastid-to-nucleus signaling during chloroplast biogenesis. Front Plant Sci 2013, 4, 14. [Google Scholar] [CrossRef] [PubMed]
- Hou, Q.; Ufer, G.; Bartels, D. Lipid signalling in plant responses to abiotic stress. Plant Cell Environ 2016, 39, 1029–1048. [Google Scholar] [CrossRef] [PubMed]
- Mei, C.; Chretien, R. L.; Amaradasa, B. S.; He, Y.; Turner, A.; Lowman, S. Characterization of phosphate solubilizing bacterial endophytes and plant growth promotion in vitro and in greenhouse. Microorganisms 2021, 9, 1935. [Google Scholar] [CrossRef]
- Mei, C.; Zhou, D.; Chretien, R. L.; Turner, A.; Hou, G.; Evans, M. R.; Lowman, S. A potential application of Pseudomonas psychrotolerans IALR632 for lettuce growth promotion in hydroponics. Microorganisms 2023, 11, 376. [Google Scholar] [CrossRef]
- Pan, L.; Cai, B. Phosphate-solubilizing bacteria: advances in their physiology, molecular mechanisms and microbial community effects. Microorganisms 2023, 11, 2904. [Google Scholar] [CrossRef]
- Varga, T.; Hixson, K. K.; Ahkami, A. H.; Sher, A. W.; Barnes, M. E.; Chu, R. K.; Battu, A.K.; Nicora, C.D.; Winkler, T.E.; Reno, L.R.; Fakra, S.C. Endophyte-promoted phosphorus solubilization in Populus. Front Plant Sci 2020, 11, 567918. [Google Scholar] [CrossRef] [PubMed]
- Glick, B. R.; Todorovic, B.; Czarny, J.; Cheng, Z.; Duan, J.; McConkey, B. Promotion of plant growth by bacterial ACC deaminase. Crit. Rev. Plant Sci 2007, 26, 227–242. [Google Scholar] [CrossRef]
- Shahid, M.; Singh, U. B.; Khan, M. S.; Singh, P.; Kumar, R.; Singh, R. N.; Kumar, A.; Singh, H.V. Bacterial ACC deaminase: Insights into enzymology, biochemistry, genetics, and potential role in amelioration of environmental stress in crop plants. Front Microbiol 2023, 14, 1132770. [Google Scholar] [CrossRef]
- Aloo, B. N.; Dessureault-Rompré, J.; Tripathi, V.; Nyongesa, B. O.; Were, B. A. Signaling and crosstalk of rhizobacterial and plant hormones that mediate abiotic stress tolerance in plants. Front Microbiol 2023, 14, 1171104. [Google Scholar] [CrossRef] [PubMed]
- Pieterse, C. M. J.; Zamioudis, C.; Berendsen, R. L.; Weller, D. M.; Van Wees, S. C. M.; Bakker, P. A. H. M. Induced systemic resistance by beneficial microbes. Annl Rev Phytopathol 2014, 52, 347–375. [Google Scholar] [CrossRef]
- Renganathan, P.; Gaysina, L. A.; Rueda-Puente, E. O. Microbial biofertilizers for salinity stress mitigation in hydroponic systems. Curr Issues Mol Biol 2025, 47(12), 1029. [Google Scholar] [CrossRef] [PubMed]
- Abou Jaoudé, R.; Luziatelli, F.; Ficca, A. G.; Ruzzi, M. Effect of plant growth-promoting rhizobacteria synthetic consortium on growth, yield, and metabolic profile of lettuce (Lactuca sativa L.) grown under suboptimal nutrient regime. Horticulturae 2025, 11, 64. [Google Scholar] [CrossRef]
- Yacoub, A.; Renault, D.; Haidar, R.; Boulisset, F.; Letousey, P.; Guyoneaud, R.; Attard, E.; Rey, P. Impact of the biocontrol product, Esquive® WP, on the indigenous grapevine wood microbiome after a 6-year application period. J Fungi 2024, 10, 566. [Google Scholar] [CrossRef]
- Merchant, N.; Lyons, E.; Goff, S.; Vaughn, M.; Ware, D.; Micklos, D.; Antin, P. The iPlant collaborative: cyberinfrastructure for enabling data to discovery for the life sciences. PLoS Biol 2016, 14, e1002342. [Google Scholar] [CrossRef]
- Williams, J. CyVerse for reproducible research: RNA-seq analysis. In Methods in Molecular Biology; Humana Press Inc.: New York, NY; Springer US, 2022; Vol. 2443, pp. pp 57–79. [Google Scholar] [CrossRef]
- Peri, S.; Roberts, S.; Kreko, I. R.; McHan, L. B.; Naron, A.; Ram, A.; Murphy, R.L.; Lyons, E.; Gregory, B.D.; Devisetty, U.K.; Nelson, A.D. Read mapping and transcript assembly: a scalable and high-throughput workflow for the processing and analysis of ribonucleic acid sequencing data. Front Genet 2020, 10, 1361. [Google Scholar] [CrossRef]
- Kim, D.; Paggi, J. M.; Park, C.; Bennett, C.; Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotechnol 2019, 37, 907–915. [Google Scholar] [CrossRef]
- Love, M. I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Thimm, O.; Bläsing, O.; Gibon, Y.; Nagel, A.; Meyer, S.; Krüger, P.; Selbig, J.; Müller, L.A.; Rhee, S.Y.; Stitt, M. MAPMAN: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J 2004, 37(6), 914–939. [Google Scholar] [CrossRef] [PubMed]
- Nouri, M. Z.; Moumeni, A.; Komatsu, S. Abiotic stresses: Insight into gene regulation and protein expression in photosynthetic pathways of plants. Int J Mol Sci 2015, 16, 20392–20416. [Google Scholar] [CrossRef]
- Ahmed, W.; Wang, Y.; Ji, W.; Liu, S.; Zhou, S.; Pan, J.; Pan, J.; Li, Z.; Wang, F.; Wang, X. Unraveling the mechanism of the endophytic bacterial strain pseudomonas oryzihabitans GDW1 in enhancing tomato plant growth through modulation of the host transcriptome and bacteriome. Int J Mol Sci 2025, 26, 1922. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



