Submitted:
27 February 2026
Posted:
28 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Developmental Pharmacology: The Biological Barrier
2.1. The Cytochrome P450 “Isoform Switch”
2.2. Renal Immaturity and Pharmacokinetic Variability
2.3. Pharmacogenomic Considerations
| Domain | Neonatal feature | Design / evaluation implication | Example |
|---|---|---|---|
| Absorption | Variable gastric pH, motility, and feeding patterns | Oral bioavailability is unpredictable; prefer exposure-guided strategies | Feeding intolerance drugs |
| Distribution | High total body water; low fat stores; lower albumin in preterm infants | Larger Vd for hydrophilic drugs; higher free fraction for highly bound drugs | Aminoglycosides; cisapride protein binding |
| Metabolism | CYP3A7 predominance early; low CYP3A4/UGT activity at birth | High vulnerability to DDIs; metabolite profiles differ from adults | CYP3A inhibitors increasing cisapride exposure |
| Excretion | Rapid maturation of GFR and tubular transport | Clearance changes over days–weeks; dosing must be dynamic | Vancomycin MIPD improves target attainment |
| Electrophysiology | Limited repolarization reserve and co-morbidity burden | Safety margins for hERG blockade may be smaller; require developmental safety assays | Torsadogenic risk signals |
3. The Cisapride Paradigm: A Forensic Case Study
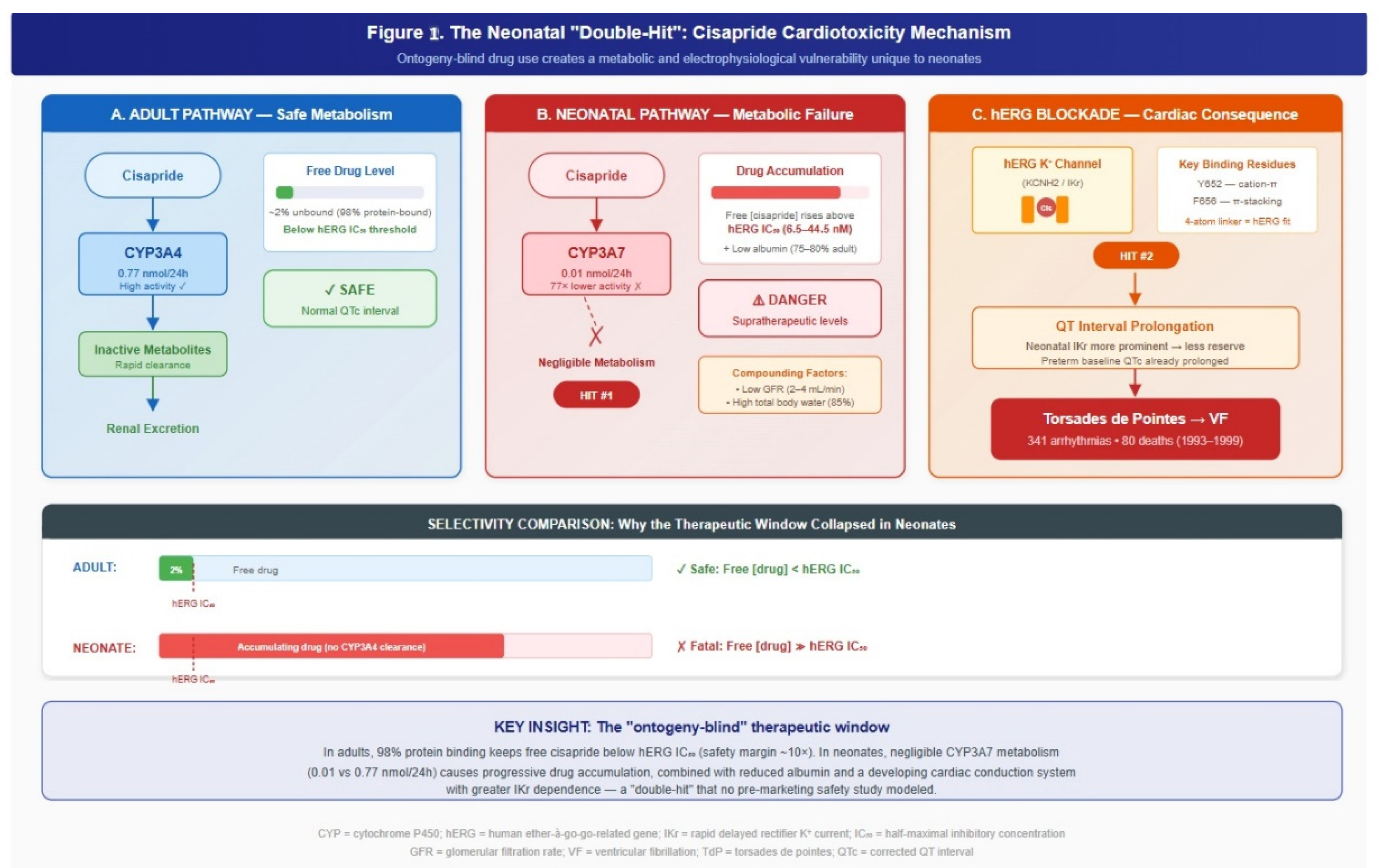
3.1. Mechanism of Action and Cardiotoxicity
3.2. Developmental Vulnerability: CYP3A-Mediated Exposure and the Neonatal “Double-Hit”
4. The Artificial Intelligence Landscape: Tools for Discovery and Safety
4.1. AlphaFold 3: Protein-Ligand Structure Prediction and Molecular Docking
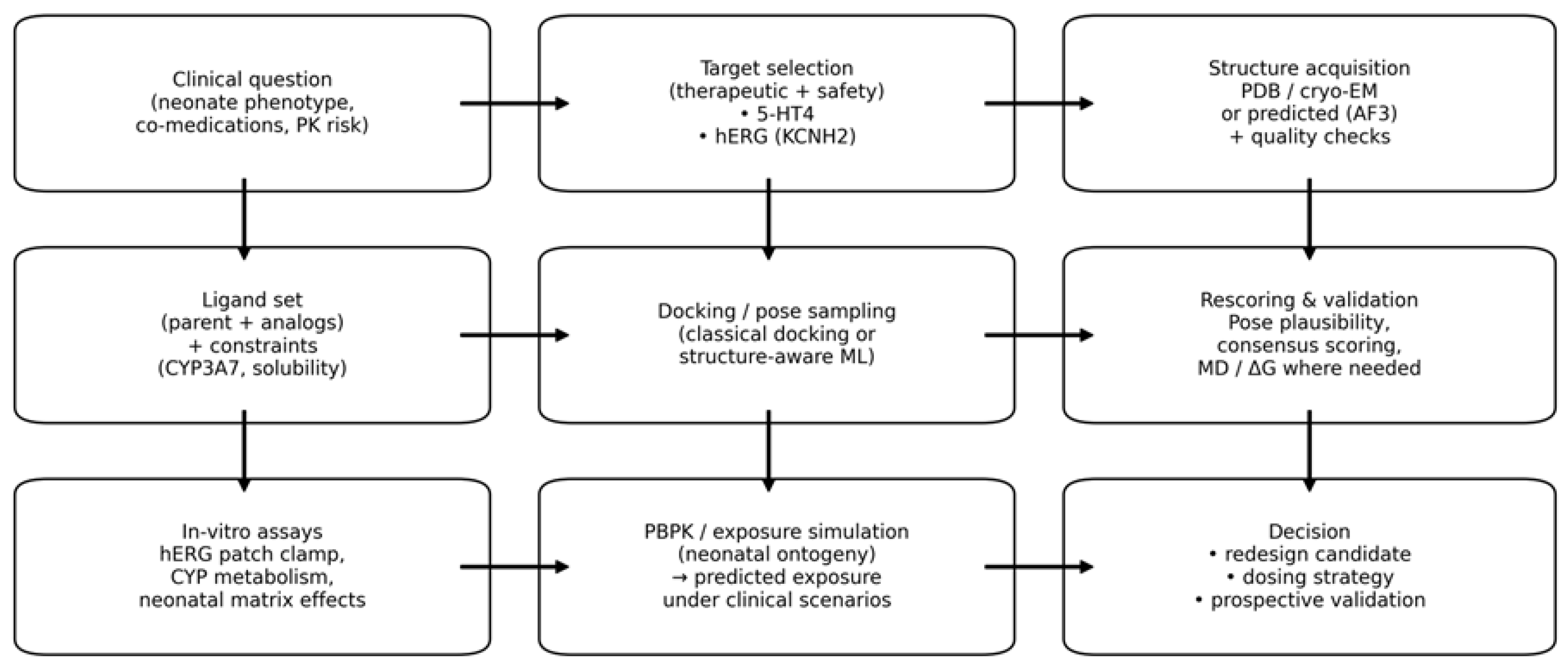
4.2. Large Language Models and Neuro-Symbolic Constraints in Chemistry
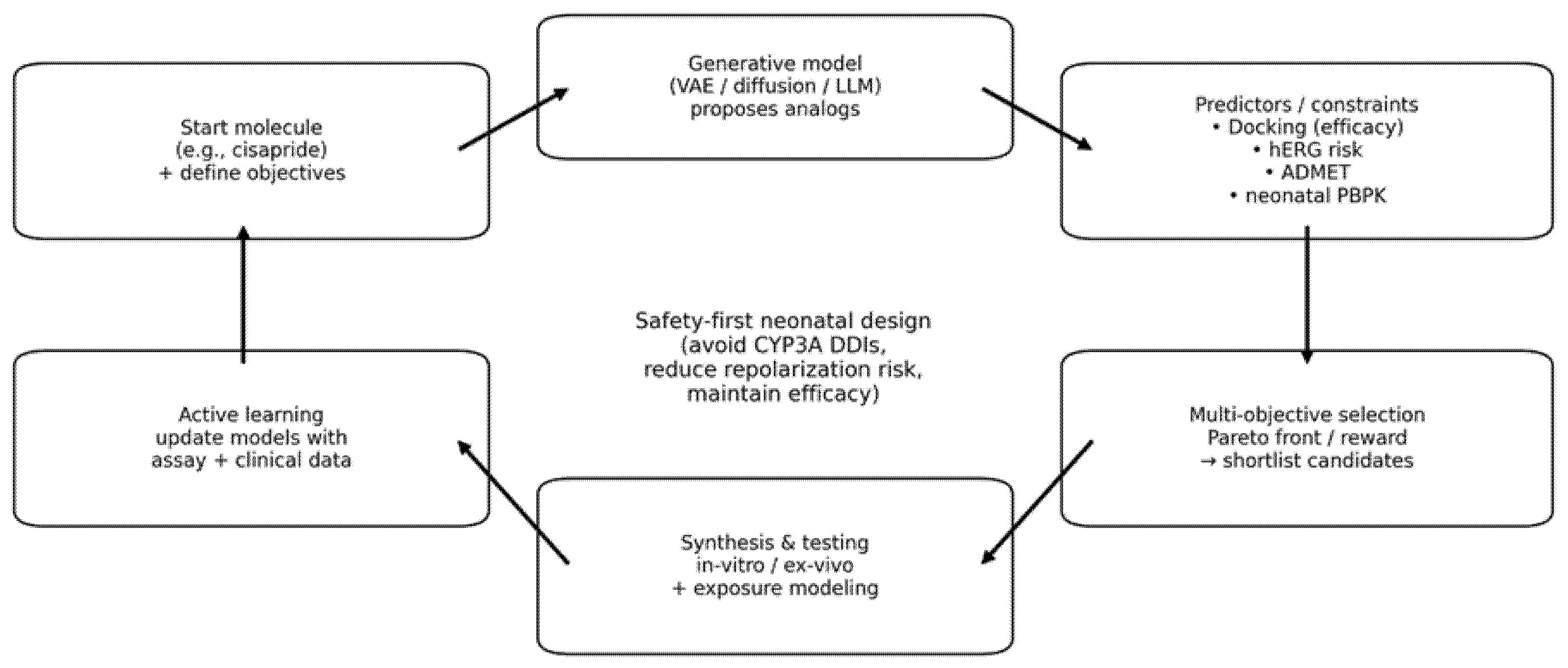
4.3. World Foundation Models: Simulating the Biological System
5. Generative Redesign: A Cisapride 2.0 Conceptual Framework
5.1. Structure–Activity Relationships and the Optimization Objective
5.2. The Generative Workflow
5.3. Validation: Prucalopride and Mosapride as Proof-of-Concept
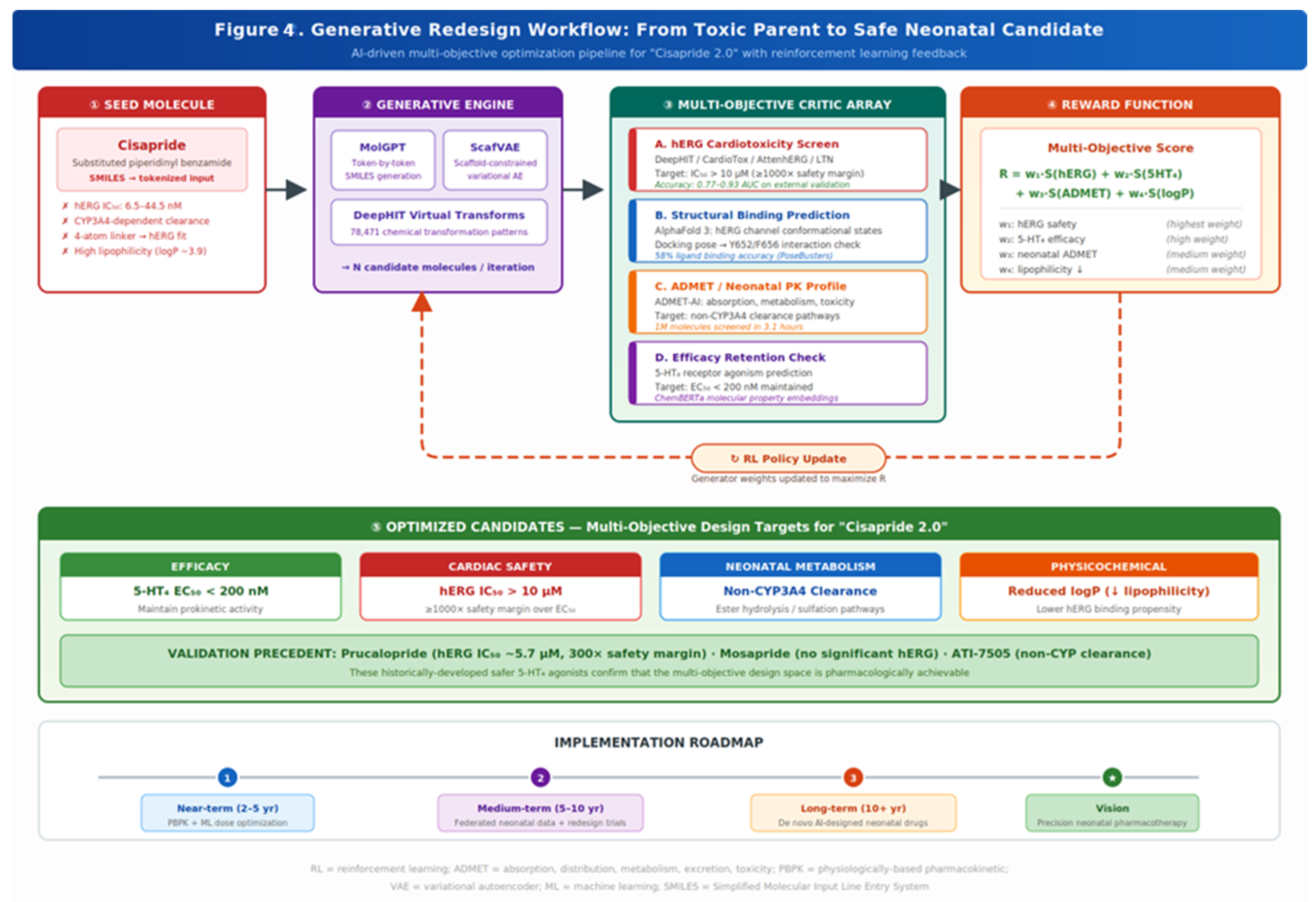
6. Implementation Barriers and Strategic Roadmap
6.1. Data Scarcity and the Neonatal Training Gap
6.2. Regulatory Framework Immaturity
6.3. A Phased Development Roadmap
6.4. Ethical and Equity Considerations
7. Conclusions
Abbreviations.
| 5-HT4 | 5-Hydroxytryptamine Receptor Type 4 |
| ADMET | Absorption, Distribution, Metabolism, Excretion, and Toxicity |
| AF3 | AlphaFold 3 |
| BPCA | Best Pharmaceuticals for Children Act |
| CDER | Center for Drug Evaluation and Research |
| CYP | Cytochrome P450 |
| EC50 | Half-Maximal Effective Concentration |
| GFR | Glomerular Filtration Rate |
| hERG | Human Ether-à-go-go-Related Gene |
| IC50 | Half-Maximal Inhibitory Concentration |
| IKr | Rapid Delayed Rectifier Potassium Current |
| LLM | Large Language Model |
| NICU | Neonatal Intensive Care Unit |
| PBPK | Physiologically-Based Pharmacokinetic |
| PK | Pharmacokinetic |
| PREA | Pediatric Research Equity Act |
| QTc | Corrected QT Interval |
| RL | Reinforcement Learning |
| RMSD | Root Mean Square Deviation |
| SMILES | Simplified Molecular Input Line Entry System |
| TDC | Therapeutics Data Commons |
| TRL | Technology Readiness Level |
| UGT | UDP-Glucuronosyltransferase |
| WFM | World Foundation Model |
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Shirkey, H. Therapeutic orphans. J. Pediatr. 1968, 72, 119–120. [Google Scholar] [CrossRef]
- Costa, H.T.M.L.; Costa, T.X.; Martins, R.R.; Oliveira, A.G. Use of off-label and unlicensed medicines in neonatal intensive care. PLoS ONE 2018, 13, e0204427. [Google Scholar] [CrossRef]
- Nir-Neuman, H.; Abu-Kishk, I.; Toledano, M.; et al. Unlicensed and Off-Label Medication Use in Pediatric and Neonatal Intensive Care Units: No Change Over a Decade. Adv. Ther. 2018, 35, 1122–1132. [Google Scholar] [CrossRef] [PubMed]
- Horen, B.; Montastruc, J.L.; Lapeyre-Mestre, M. Adverse drug reactions and off-label drug use in paediatric outpatients. Br. J. Clin. Pharmacol. 2002, 54, 665–670. [Google Scholar] [CrossRef] [PubMed]
- Kaushal, R.; Bates, D.W.; Landrigan, C.; et al. Medication errors and adverse drug events in pediatric inpatients. JAMA 2001, 285, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.H.S.; Lipshultz, S.E. Caffeine and clinical outcomes in premature neonates. Children 2019, 6, 118. [Google Scholar] [CrossRef]
- U.S. Food and Drug Administration. Pediatric Labeling Changes. Available online: https://www.fda.gov/science-research/pediatrics/pediatric-labeling-changes (accessed on 13 February 2026).
- Laughon, M.M.; Avant, D.; Tripathi, N.; et al. Drug labeling and exposure in neonates. JAMA Pediatr. 2014, 168, 130–136. [Google Scholar] [CrossRef]
- Stevens, J.C. New perspectives on the impact of cytochrome P450 3A expression for pediatric pharmacology. Drug Discov. Today 2006, 11, 440–445. [Google Scholar] [CrossRef]
- Hines, R.N. Ontogeny of human hepatic cytochromes P450. J. Biochem. Mol. Toxicol. 2007, 21, 169–175. [Google Scholar] [CrossRef]
- de Wildt, S.N.; Kearns, G.L.; Leeder, J.S.; et al. Cytochrome P450 3A: Ontogeny and drug disposition. Clin. Pharmacokinet. 1999, 37, 485–505. [Google Scholar] [CrossRef]
- Bhatt, D.K.; Mehrotra, A.; Gaedigk, A.; et al. Age- and Genotype-Dependent Variability in the Protein Abundance and Activity of Six Major Uridine Diphosphate-Glucuronosyltransferases in Human Liver. Clin. Pharmacol. Ther. 2019, 105, 131–141. [Google Scholar] [CrossRef]
- Knibbe, C.A.J.; Krekels, E.H.J.; van den Anker, J.N.; et al. Morphine Glucuronidation in Preterm Neonates, Infants and Children Younger than 3 Years. Clin. Pharmacokinet. 2009, 48, 371–385. [Google Scholar] [CrossRef]
- van den Anker, J.; Reed, M.D.; Allegaert, K.; et al. Developmental changes in pharmacokinetics and pharmacodynamics. J. Clin. Pharmacol. 2018, 58, S10–S25. [Google Scholar] [CrossRef] [PubMed]
- Sherwin, C.M.; Svahn, S.; Van der Linden, A.; et al. Individualised dosing of amikacin in neonates: a pharmacokinetic/pharmacodynamic analysis. Eur. J. Clin. Pharmacol. 2009, 65, 705–713. [Google Scholar] [CrossRef] [PubMed]
- Kalamees, R.; Soeorg, H.; Ilmoja, M.; et al. Prospective validation of a model-informed precision dosing tool for vancomycin treatment in neonates. Antimicrob. Agents Chemother. 2024, 68, e01591-23. [Google Scholar] [CrossRef] [PubMed]
- Gallaway, K.A.; Cann, K.; Oetting, K.; et al. The potential impact of preemptive pharmacogenetic genotyping in the neonatal intensive care unit. J. Pediatr. 2023, 259, 113489. [Google Scholar] [CrossRef]
- McDermott, J.H.; Mahaveer, A.; James, R.A.; et al. Rapid point-of-care genotyping to avoid aminoglycoside-induced ototoxicity in neonatal intensive care. JAMA Pediatr. 2022, 176, 486–492. [Google Scholar] [CrossRef]
- Quigley, E.M.M. Cisapride: What can we learn from the rise and fall of a prokinetic? J. Dig. Dis. 2011, 12, 147–156. [Google Scholar] [CrossRef]
- Mohammad, S.; Zhou, Z.; Gong, Q.; et al. Blockage of the HERG human cardiac K+ channel by the gastrointestinal prokinetic agent cisapride. Am. J. Physiol. 1997, 273, H2534–H2538. [Google Scholar] [CrossRef]
- Redfern, W.S.; Carlsson, L.; Davis, A.S.; et al. Relationships between preclinical cardiac electrophysiology, clinical QT interval prolongation and torsade de pointes for a broad range of drugs. Cardiovasc. Res. 2003, 58, 32–45. [Google Scholar] [CrossRef]
- Wysowski, D.K.; Corken, A.; Gallo-Torres, H.; et al. Postmarketing reports of QT prolongation and ventricular arrhythmia in association with cisapride and Food and Drug Administration regulatory actions. Am. J. Gastroenterol. 2001, 96, 1698–1703. [Google Scholar] [CrossRef] [PubMed]
- Treluyer, J.M.; Rey, E.; Sonnier, M.; et al. Evidence of impaired cisapride metabolism in neonates. Br. J. Clin. Pharmacol. 2001, 52, 419–425. [Google Scholar] [CrossRef] [PubMed]
- Pearce, R.E.; Gotschall, R.R.; Kearns, G.L.; Leeder, J.S. Cytochrome P450 involvement in the biotransformation of cisapride and racemic norcisapride in vitro: Differential activity of individual human CYP3A isoforms. Drug Metab. Dispos. 2001, 29, 1548–1554. [Google Scholar] [PubMed]
- Kearns, G.L.; Robinson, P.K.; Wilson, J.T.; et al. Cisapride disposition in neonates and infants: In vivo reflection of cytochrome P450 3A4 ontogeny. Clin. Pharmacol. Ther. 2003, 74, 312–325. [Google Scholar] [CrossRef]
- Cordeiro, J.M.; Panama, B.K.; Goodrow, R.; et al. Developmental changes in expression and biophysics of ion channels in the canine ventricle. J. Mol. Cell. Cardiol. 2013, 65, 34–45. [Google Scholar] [CrossRef]
- Vandenplas, Y.; Benatar, A.; Cools, F.; et al. Efficacy and tolerability of cisapride in children. Paediatr. Drugs 2001, 3, 559–573. [Google Scholar] [CrossRef]
- Roden, D.M. Drug-induced prolongation of the QT interval. N. Engl. J. Med. 2004, 350, 1013–1022. [Google Scholar] [CrossRef]
- Barnett, C.P.; Omari, T.; Davidson, G.P.; et al. Effect of cisapride on gastric emptying in premature infants with feed intolerance. J. Paediatr. Child Health 2001, 37, 559–563. [Google Scholar] [CrossRef] [PubMed]
- Abramson, J.; Adler, J.; Dunger, J.; et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 2024, 630, 493–500. [Google Scholar] [CrossRef]
- Google DeepMind. AlphaFold 3 predicts the structure and interactions of all of life’s molecules. Available online: https://blog.google/technology/ai/google-deepmind-isomorphic-alphafold-3-ai-model/ (accessed on 13 February 2026).
- Kryshtafovych, A.; Schwede, T.; Topf, M.; et al. Critical assessment of methods of protein structure prediction (CASP)—Round XV. Proteins 2023, 91, 1539–1549. [Google Scholar] [CrossRef]
- Ngo, K.; Yang, P.C.; Yarov-Yarovoy, V.; Clancy, C.E.; Vorobyov, I. Harnessing AlphaFold to Reveal hERG Channel Conformational State Secrets. eLife 2025, 13, RP104901. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Lin, H.; Alade, A.A.; et al. AlphaFold3 in Drug Discovery: A Comprehensive Assessment of Strengths and Limitations. bioRxiv 2025. [Google Scholar] [CrossRef]
- Krokidis, M.G.; Koumadorakis, D.E.; Lazaros, K.; et al. AlphaFold 3: An overview of applications and performance insights. Int. J. Mol. Sci. 2025, 26, 3671. [Google Scholar] [CrossRef] [PubMed]
- Isomorphic Labs. Partnerships with Eli Lilly and Novartis. Available online: https://www.isomorphiclabs.com/ (accessed on 13 February 2026).
- Chithrananda, S.; Grand, G.; Ramsundar, B. ChemBERTa: Large-scale self-supervised pretraining for molecular property prediction. arXiv 2020, arXiv:2010.09885. [Google Scholar]
- Hossain, D.; Al Abir, F.; Chen, J.Y. hERG-LTN: A New Paradigm in hERG Cardiotoxicity Assessment Using Neuro-Symbolic and Generative AI Embedding Approach. bioRxiv 2025. [Google Scholar] [CrossRef]
- Bagal, V.; Aggarwal, R.; Vinod, P.K.; et al. MolGPT: Molecular Generation Using a Transformer-Decoder Model. J. Chem. Inf. Model. 2022, 62, 2064–2076. [Google Scholar] [CrossRef]
- Swanson, K.; Walters, P.; Sellers, B. ADMET-AI: A machine learning ADMET platform for evaluation of large-scale chemical libraries. Bioinformatics 2024, 40, btae416. [Google Scholar] [CrossRef]
- Yang, T.; Ding, X.; McMichael, E.; et al. AttenhERG: A reliable and interpretable graph neural network framework for predicting hERG channel blockers. J. Cheminform. 2024, 16, 95. [Google Scholar] [CrossRef]
- Richardson, P.; Griffin, I.; Tucker, C.; et al. Baricitinib as potential treatment for 2019-nCoV acute respiratory disease. Lancet 2020, 395, e30–e31. [Google Scholar] [CrossRef]
- Xu, Z.; Ren, F.; Wang, P.; et al. A generative AI-discovered TNIK inhibitor for idiopathic pulmonary fibrosis: A randomized phase 2a trial. Nat. Med. 2025, 31, 2602–2610. [Google Scholar] [CrossRef]
- Zhavoronkov, A.; Ivanenkov, Y.A.; Aliper, A.; et al. Deep learning enables rapid identification of potent DDR1 kinase inhibitors. Nat. Biotechnol. 2019, 37, 1038–1040. [Google Scholar] [CrossRef]
- NVIDIA. World Models Definition. NVIDIA Glossary. Available online: https://www.nvidia.com/en-us/glossary/world-models/ (accessed on 13 February 2026).
- NVIDIA. Cosmos: Physical AI World Foundation Models. Available online: https://www.nvidia.com/en-us/ai/cosmos/ (accessed on 13 February 2026).
- Noetik, AI. OCTO—World Models as Simulators of Patient Biology. Available online: https://www.noetik.ai/octo (accessed on 13 February 2026).
- Cui, H.; Wang, C.; Maan, H.; et al. scGPT: Toward building a foundation model for single-cell multi-omics using generative AI. Nat. Methods 2024, 21, 1470–1480. [Google Scholar] [CrossRef] [PubMed]
- Velez-Arce, A.; Huang, K.; Li, X.L.; et al. TDC-2: Multimodal Foundation for Therapeutic Science. bioRxiv 2024. [Google Scholar] [CrossRef]
- Fermini, B.; Fossa, A.A. The impact of drug-induced QT interval prolongation on drug discovery and development. Nat. Rev. Drug Discov. 2003, 2, 439–447. [Google Scholar] [CrossRef] [PubMed]
- Mitcheson, J.S.; Chen, J.; Lin, M.; et al. A structural basis for drug-induced long QT syndrome. Proc. Natl. Acad. Sci. U.S.A 2000, 97, 12329–12333. [Google Scholar] [CrossRef]
- Mouchlis, V.D.; Afantitis, A.; Serra, A.; et al. Advances in de novo drug design: From conventional to machine learning methods. Int. J. Mol. Sci. 2021, 22, 1676. [Google Scholar] [CrossRef]
- Ryu, J.Y.; Lee, M.Y.; Lee, J.H.; et al. DeepHIT: a deep learning framework for prediction of hERG-induced cardiotoxicity. Bioinformatics 2020, 36, 3049–3055. [Google Scholar] [CrossRef]
- Dong, T.; You, L.; Chen, CY-C..; et al. Multi-objective drug design with a scaffold-aware variational autoencoder. Chem. Sci. 2025, 16, 13352–13367. [Google Scholar] [CrossRef]
- Camilleri, M.; Kerstens, R.; Rykx, A.; et al. A placebo-controlled trial of prucalopride for severe chronic constipation. N. Engl. J. Med. 2008, 358, 2344–2354. [Google Scholar] [CrossRef]
- Carlsson, L.; Amos, G.J.; Andersson, B.; et al. Electrophysiological Characterization of the Prokinetic Agents Cisapride and Mosapride in Vivo and in Vitro: Implications for Proarrhythmic Potential? J. Pharmacol. Exp. Ther. 1997, 282, 220–227. [Google Scholar] [CrossRef] [PubMed]
- Tack, J.; Camilleri, M.; Chang, L.; et al. Systematic Review: Cardiovascular Safety Profile of 5-HT4 Agonists Developed for Gastrointestinal Disorders. Aliment. Pharmacol. Ther. 2012, 35, 745–767. [Google Scholar] [CrossRef]
- Camilleri, M.; Vazquez-Roque, M.I.; Burton, D.; et al. Pharmacodynamic effects of a novel prokinetic 5-HT receptor agonist, ATI-7505, in humans. Neurogastroenterol. Motil. 2007, 19, 30–38. [Google Scholar] [CrossRef]
- Allegaert, K.; van de Velde, M.; van den Anker, J. Neonatal clinical pharmacology. Paediatr. Anaesth. 2014, 24, 30–38. [Google Scholar] [CrossRef]
- Allegaert, K.; Yalcin, N.; Flint, R.B.; et al. Neonatal Clinical Pharmacology. Children 2024, 11, 1102. [Google Scholar] [CrossRef]
- U.S. Food and Drug Administration. Considerations for the Use of Artificial Intelligence to Support Regulatory Decision-Making for Drug and Biological Products. Draft Guidance for Industry. Available online: https://www.fda.gov/regulatory-information/search-fda-guidance-documents (accessed on 13 February 2026).
- U.S. Food and Drug Administration. Artificial Intelligence for Drug Development. Available online: https://www.fda.gov/about-fda/center-drug-evaluation-and-research-cder/artificial-intelligence-drug-development (accessed on 13 February 2026).
- Conte, L.; Decembrino, N.; Arribas, C.; et al. Leveraging Artificial Intelligence for Decision Support in Neonatal and Pediatric Pharmacotherapy: A Scoping Review. Semin. Fetal Neonatal Med. 2025, 19, 101691. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




