1. Introduction
The turbot (
Scophthalmus maximus) is the farmed flatfish with higher production worldwide (~ 72,000 tonnes) being Spain and PR China, the principal producers. Controlling reproduction and sex ratio is one of the main challenges for turbot producers along with increasing growth rate and disease resistance [
1]. This teleost is a gonochoristic species, in which each mature individual possesses only ovaries or testes, and exhibits the highest sexual dimorphism among marine fish both in growth rate and sexual maturity [
2,
3]. In turbot females sexual maturity occurs about at a year earlier than in males, increasing somatic growth and reducing susceptibility to diseases and mortality, which have aroused the interest of the aquaculture industry in obtaining all-female populations [
2].
Sexual development occurs in a similar way across teleost fishes and, as in all vertebrates, relies on two concatenated processes, sex determination (SD) and sex differentiation. SD occurs early during embryogenesis when sex is established by the determination of the developmental pathway of the bipotential gonadal primordium which is identical in males and females. Then sex differentiation involves the forming of the testis or the ovary throughout development from the undifferentiated bipotential gonadal primordium [
4].
In contrast to mammals and birds, which shows a common fixed sex determining molecular machinery, teleost fishes display notably different sexual development mechanisms, even among closely related species [
5]. In fact, in certain cases SD and sex differentiation is species-specific and can be regulated by a gene network, by environmental factors (temperature, population density, hypoxia, pH and social interaction [
6]), or by both [
4,
5,
7]. In the turbot, sex is considered as a quantitative threshold trait, where genetic and environmental factors, as the temperature at which the embryos develop, are involved at the beginning of sex differentiation for determining the fate of the undifferentiated primordium [
8,
9,
10,
11]. Moreover, size and growth of the turbot specimens could be critical elements that strongly influence SD [
9] as it was reported in other teleost species such as the European sea bass [
12].
As it was mentioned above, SD mechanisms are very variable among teleosts so many sex-related genes have been identified in these fishes [
5], including the turbot [
1,
8,
9,
10,
13], although the SD master gene can vary even among close species. Studies carried out in
S. maximus have identified a ZZ/ZW sex chromosome system with limited environmental influence based on the sex ratio obtained in progenies from hormonally sex-reversed parents [
14]. These results were confirmed by Martínez et al. [
15] who further identified a major sex-related Quantitative Trait Loci (QTL) at linkage group (LG) 5. After identification of several candidate genes at LG5 using the whole genome sequence of this species [
16] and a genome-wide association study [
10]
Sox2 has been proposed as the SD gene in the sexual development of the turbot.
Understanding how sexual development occurs in the turbot and the genes involved in this process is an important challenge that requires studies following different approaches. Analyses with RT-PCR, qPCR, microarray, and transcriptome sequencing have revealed the expression of SD and sex differentiation key genes in the turbot throughout gonadal development [
9,
13,
17,
18,
19,
20,
21]. Moreover, qPCR analyses of genes related to primordial germ cells development have reported signs of sex differentiation in postmetamorphic turbot larvae (75-90 days post fertilization (dpf), 5-6 cm length) [
9]. However, in the same study using the classical haematoxylin and eosin (H&E) stain morphological gonad differences between sexes were not detected before 105 dpf (7,2 cm length, [
9]). More recent histological studies by H&E stain in turbot reported that the ovaries and testis could be morphologically distinguished in earlier stages of development [
11,
21].
Although numerous molecular analyses (i.e., by RT-PCR, qPCR, microarray, transcriptome sequencing) about gene expression patterns in teleosts gonads have been performed, histological studies are scarce despite identifying the cell types where sex-related genes are expressed represents essential complementary information to understand the process of gonadal development. To the best of our knowledge, only
Vasa [
17,
22,
23] and
Dnd [
18,
23] expression have been studied by in situ hybridization (ISH) in differentiated testis and ovary of juvenile and adult turbot. We have performed ISH on differentiated ovary and testis paraffin sections (SISH) of juvenile and adult turbot to evidence the tissue-specific and cellular sites of the expression of
Vasa,
Cyp19a1a,
Foxl2,
Amh,
Sox2,
Sox17, and
Sox19. The reason for studying the expression patterns of these genes is that they are involved in sexual development in turbot and have been previously characterized in our lab by molecular analyses in turbot gonads [
9,
10,
13,
15,
16,
24,
25].
The aim of our research was to use SISH with two goals (1) to determine the tissue specific expression of Vasa, Cyp19a1a, Foxl2, Amh, Sox2, Sox17, and Sox19 in turbot gonads, i.e., cell types where these genes were expressed, and (2) to confirm on tissue sections the data observed in previous molecular analysis. Thus, our results showed that SISH, which gives spatial information on RNA in a tissue sample, and previous qPCR analyses, which quantifies the average expression level within the entire sample, provide complementary information for understanding gene expression profiles in the gonads of the turbot. Moreover, we have compared our results with similar studies performed in other teleost species to find out if there were differences in the expression patterns of the genes studied here between male and female specimens or through the gonad development.
2. Materials and Methods
2.1. Experimental Animals and Tissue Preparation
Turbot (Scophthalmus maximus) of different developmental stages were provided by the Instituto Español de Oceanografía (Vigo, Spain). Six turbot 130 dpf (three males and three females; 10-13 cm length, 21-37 g weight), five 180 dpf (two males and three females; 12-15.1 cm length; 37.9-62.8 g weight) and six adults (2 years post fertilization (ypf); three males and three females; 25-31 cm length; 280-550 g weight) were used for the in situ hybridization study. Gonads are differentiated at every stages studied, although they are mature only in male adult specimens.
All specimens were deeply anesthetized in a 0.5% solution of MS-222 (tricaine methane sulfonate; Sigma-Aldrich, St. Louis, MO) in seawater and then perfused transcardially with Ringer’s solution containing 0.1% procaine (Sigma-Aldrich), followed by perfusion with cold 4% paraformaldehyde in 0.1 M phosphate-buffered saline pH 7.4 (PBS) and finally sacrificed. The gonads were then removed from each individual and immersed in the same fixative for 13-16 hours at 4 °C. After several PBS washes, the gonads were dehydrated in methanol at 4 °C and then embedded in paraffin wax. All buffers used in the experiments were made in RNA free distilled water. Parallel transverse serial sections (10-14 mm thick) were cut on a rotary microtome and mounted on Superfrost Plus glass slides (Menzel, Braunschweig, Germany).
To facilitate identification of the anatomy and cell types in the testis and the ovary parallel gonad sections were stained with hematoxylin-eosin (H&E) following standard protocols.
2.2. Riboprobe Synthesis
Total RNA was isolated from gonads using Trizol reagent (Invitrogen, Carlsbad, CA) and reverse transcribed to cDNA with AffinityScript Multiple Temperature cDNA Synthesis Kit (Agilent Technologies, Santa Clara, CA) following supplier recommendations. Several genes involved in sex determination and differentiation (
Vasa,
Amh,
Cyp19a1a,
Foxl2,
Sox2,
Sox17, and
Sox19) were amplified from cDNA with the primers detailed in
Table 1. These primers were designed prior to the publication of the turbot genome from the sequences in public data base. Gene sequences from different fish species were used. Finally, it was verified by sequencing if the primers pair amplified the right gene. PCR was carried out in a volume of 50 μl: 75 ng of cDNA, 20 pmol of each primer, 0.2 mM of each dNTP, 1× PCR reaction buffer, and 2.5 units of GreenTaq DNA polymerase (GenScript, Leiden, The Netherlands), in a MyCycler Thermal cycler (Bio-Rad, Hercules, CA) as follows: initial denaturation at 94 °C for 3 min; 30 cycles including 94 °C during 30 s, 50 s at their specific annealing temperature between 59-60 °C, and an extension time dependent on amplicon size (about 1 kb/min) at 72 °C; and a final extension step at 72 °C for 7 min. The PCR products were separated on 1% agarose gels, stained with SYBR gold (Invitrogen) and purified using DNA Clean & Concentrator kit (Zymo Research, Irvine, CA) for sequencing following the manufacturer’s protocol.
These DNA fragments were cloned in a pGEM-T vector (Promega, Madison, WI)), purified with QIAprep Spin Miniprep Kit (Qiagen, Hilden, Germany) and sequenced following the ABI Prism BigDye Terminator v3.1 Cycle Sequencing Kit protocol on an ABI 3730xl Genetic Analyzer (Applied Biosystems, Foster city, CA) with the SP6 and T7 universal primers (SP6: ATTTAGGTGACACTATAG, T7: TAATACGACTCACTATAGGG). The orientation of the insert was analyzed by the homology sequence analysis through BLASTn (Basic Local Alignment Search Tool Nucleotide). Sense and antisense digoxygenin-UTP-labeled riboprobes were synthesized by means of SP6 and T7 RNA polymerase (Promega) following the manufacturer’s protocol and stored at -80 °C.
2.3. In Situ Hybridization
In situ hybridization on tissue sections (SISH) was performed following standard protocols. Gonad sections were treated with proteinase K (Roche; 10 mg/ml) for 30 min at room temperature and hybridized separately with sense and antisense probes overnight at 65 °C in a humid chamber.
The posthybridization washes included RNAse A treatment (Invitrogen; 100 mg/ml, 30 min at 37 ºC), and then, the samples were incubated overnight with the alkaline phosphatase-coupled anti-digoxigenin antibody (1:2000, Roche Applied Science, Penzberg, Germany) at room temperature in a humid chamber. The color reaction was performed in the presence of BM-Purple (Roche) until the signal was clearly visible. The sections were dehydrated and coverslipped.
No positive signals were detected in the control hybridization sections with sense probes in none of the analyzed genes.
2.4. Image Acquisition and Analysis
ISH images were acquired with an Olympus BX51 microscope fitted with an Olympus DP71 color digital camera. For presentation, bright-field microscopy images were sized and optimized for brightness and contrast with Photoshop CS2 (Adobe, San Jose, CA) and plate photomontage and lettering were also done with Photoshop CS2.
3. Results and Discussion
In this work, we used H&E stain to study the gonadal morphology of juvenile and adult turbot (
Scophthalmus maximus) and performed SISH on paraffin sections of differentiated testis and ovary to analyze from a histological point of view the tissue-specific expression patterns of some genes involved in sexual development in this flatfish species. Thus, we investigated the expression patterns of
Vasa, Cyp19a1a, Foxl2, Amh,
Sox2, Sox17, and
Sox19 genes, all of which are involved in the sex differentiation of teleost fishes, including the turbot, as qPCR, microarray and transcriptomic analyses have previously proven [
9,
13]. The most noticeable of our histological study was that genes studied were expressed in male and female germ cells and that our results support on tissue sections previous data on gene expression analyses in the turbot gonads. These results proved the usefulness of in situ hybridization as a useful complement for transcriptomic and/or proteomic evaluations to better understand the molecular mechanisms of gonad differentiation in teleost fishes.
3.1. Histology of the Differentiated Turbot Gonads
With the aim of better identifying the positive in situ hybridization regions and cell types in the gonads of the turbot, H&E stain were performed on paraffin sections to study the anatomy of the ovary and testis of juveniles (130 and 180 dpf) and adults (2 years post fertilization (ypf)) specimens. As mentioned in the introduction, Zhao et al. [
21] established by H&E stain on early development turbot stages the ovary differentiation in premetamorphic larvae (60 days post hatching (dph), 3,5 cm length) while testis remained undifferentiated until 90dph (6 cm length). However, a recent study also by H&E stain reported an earlier differentiation of the turbot gonads, i.e., the ovary was already distinguished at 45 dph (3,7 cm length) with both the ovarian cavity and the germinal epithelium clearly defined [
11]. However, the testis remained undifferentiated until 65dph when the number of germ cells increased through mitosis and the spermatogonial cysts were formed in the edge of the gonads [
11]. As total length of turbot specimens initiating gonadal sex differentiation was consistent in the different studies, Meng et al. [
11] proposed that body size or length rather than age (measured as dph or dpf) is basic in determining the timing of sex differentiation in this flatfish species. Anyway, all data obtained until now show a delay in testicular differentiation compared to ovarian development in turbot and, moreover, confirm that juvenile and adult turbot specimens used in our study present already differentiated gonads.
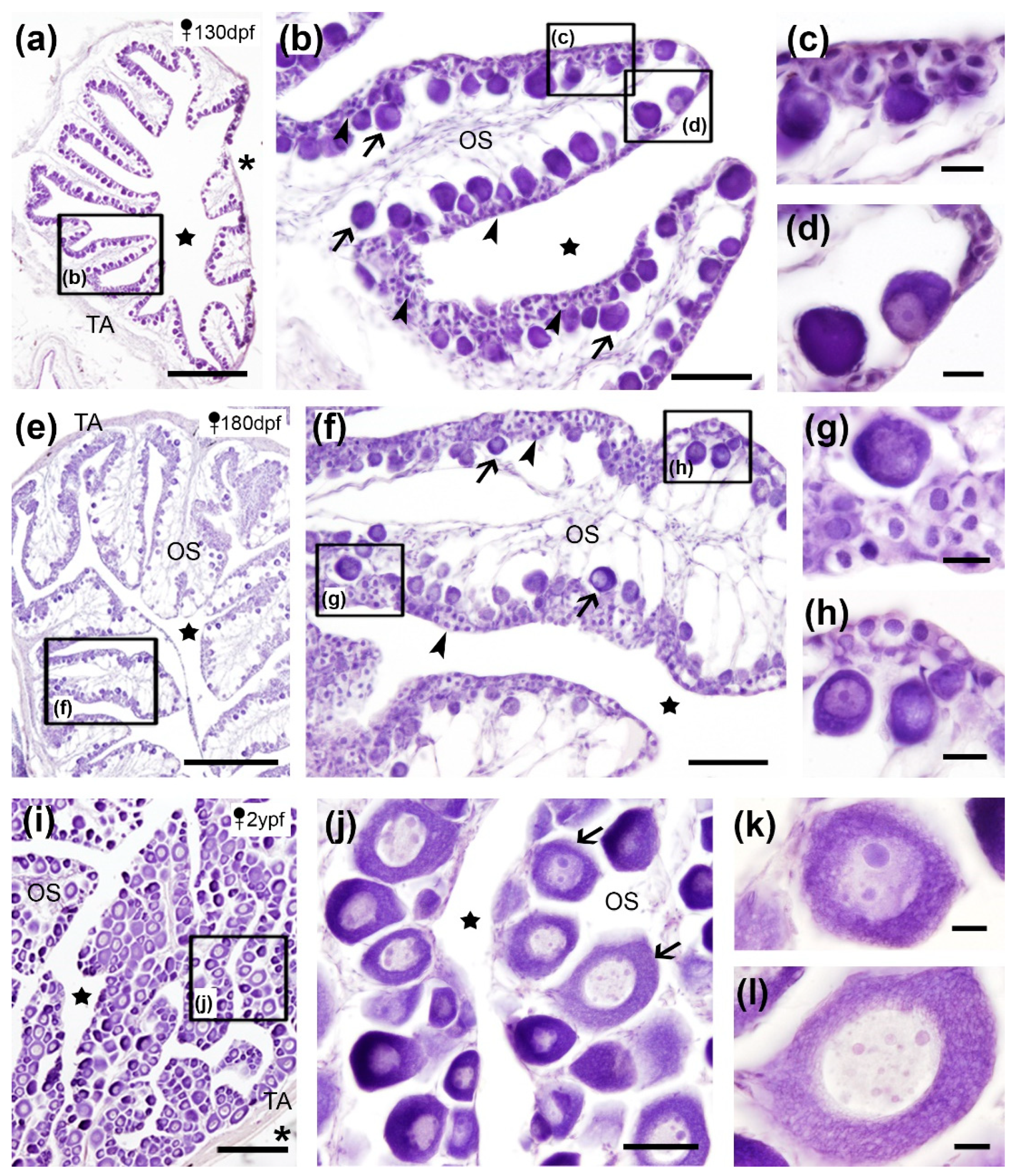
Our results showed that the turbot ovary was surrounded by a layer of connective condensed tissue, the tunica albuginea, and presented a hollow sac-like morphology with an ovarian cavity into which numerous ovigerous lamellae extended (
Figure 1). These lamellae consisted of ovarian stroma bordered by germinal epithelium supporting somatic (epithelial and follicular cells) and germ cells (oogonia and oocytes) (
Figure 1). The ovigerous lamellae at 130 dpf (
Figure 1a-d) and 180 dpf (
Figure 1e-h) stages contained abundant oogonia (9-13 µm in diameter) which showed a basophile nucleus, and numerous oocytes of different size (14-37 µm in diameter) in different steps of the primary growth stage. Primary oocytes displayed a densely basophile cytoplasm and one nucleolus but occasionally multiple nucleoli in the nucleus (
Figure 1b-d and f-h). In the ovary of 2 ypf specimens, still sexually immature females, a small amount of oogonia was observed while there was an evident increase in the number and size (50-115 µm in diameter) of primary oocytes (
Figure 1i-l), as previously described in immature ovaries of turbot specimens [
26]. At this stage, the primary oocytes showed an evident nucleus with irregularly distributed multiple nucleoli (
Figure 1j-l) and a basophile cytoplasm during the earlier stages which turned more acidophilic and showed a more apparent Balbiani body (a non-membrane bound compartment packed with RNA, mitochondria and other organelles) as development progressed (
Figure 1k-l).
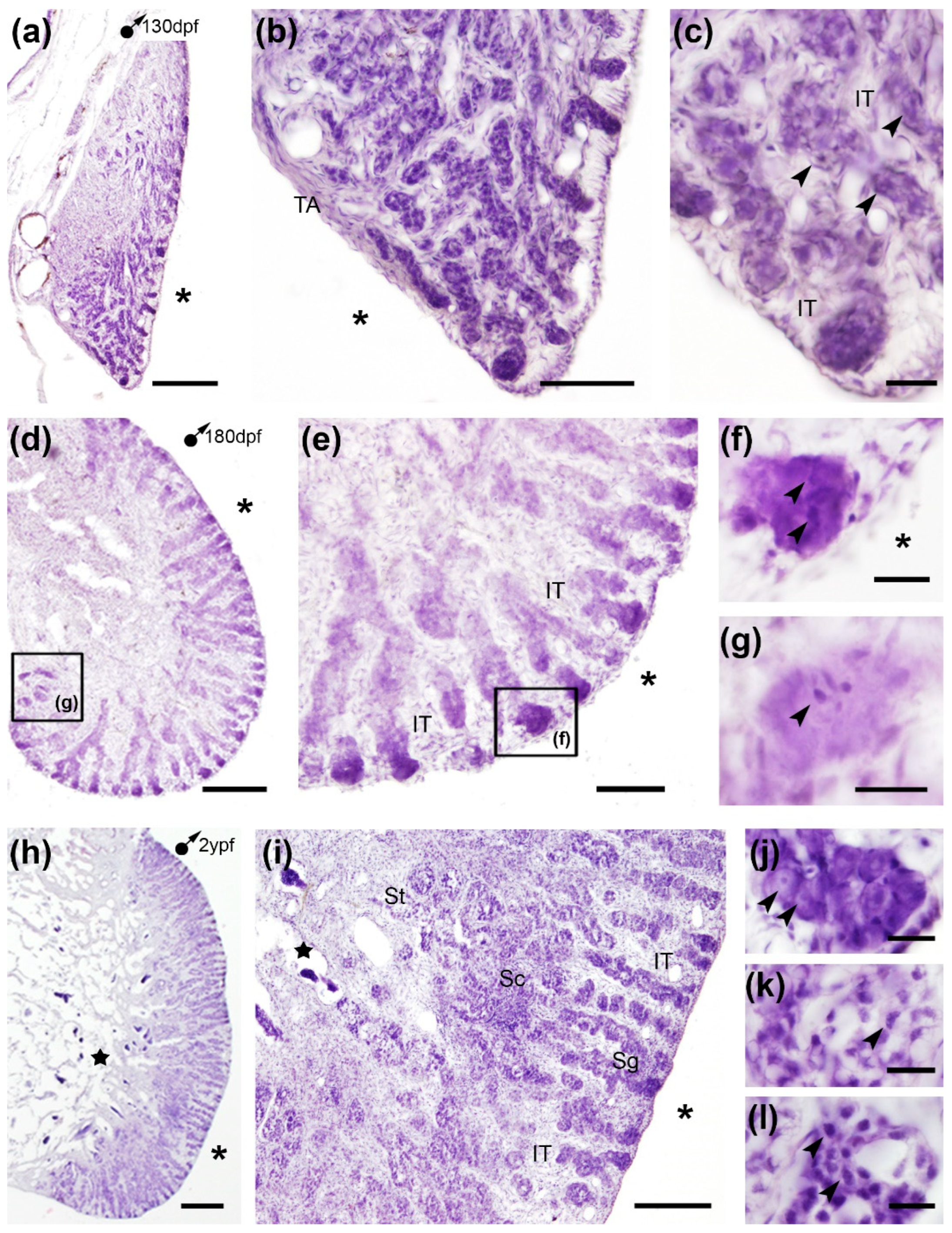
The turbot testis displayed an elongated morphology surrounded by the tunica albuginea and a lobular inner structure formed of germ cells and somatic cells comprising the germinal and the interstitial compartments (
Figure 2). The germinal compartment was formed of seminiferous or testicular lobules that extended to the periphery of the testis with germinal cells and somatic epithelial cells (Sertoli cells) along their wall, while the interstitial compartment surrounding the testicular lobules was composed of connective tissue and Leydig cells (
Figure 2a, d and h). In juveniles abundant spermatogonia were arranged in cysts throughout the length of the lobules and showed a basophilic nucleus (
Figure 2b-c and e-g). The testis of 2 ypf specimens, already sexually mature males, showed a similar histology to that previously described in mature testes of turbot specimens [
23,
26]. Thus, we observed seminiferous lobules more numerous than in juveniles which run parallel with spermatogonia around the edge of the gonad (
Figure 2i-j), cysts of primary spermatocytes, some of them in leptotene/zygotene throughout the length of the lobules (
Figure 2i and k), and spermatids near the lumen of the efferent ducts where spermiation will occur (
Figure 2i and l).
3.2. Histological Analysis of the Expression Pattern of Genes Involved in Sexual Development in Turbot
As mentioned in the introduction, the genetic cascade underlying the process of SD and gonad differentiation present a high diversity among teleosts, and even between closely related species, both in the master SD genes that trigger the male or female pathways and in the relative gene positions and their functions in testicular and ovarian differentiation [
4,
5,
8,
27]. Moreover, diverse expression patterns of the sex-related genes were observed between male and females of different teleost species over the course of development [
5,
7,
28,
29,
30,
31], including the turbot [
9,
13,
19,
20,
21].
Regarding the expression of the genes studied in this work, previous data based on qPCR analysis in turbot gonads showed that
Vasa,
Amh and
Cyp19a1a were the first sexual differentially expressed genes with a higher expression of
Vasa and
Cyp19a1a in females and
Amh in males [
9].
Vasa,
Amh and
Cyp19a1a increased their expression from 75 to 90 dpf stages (5-6 cm length), as well as
Foxl2, which showed a significantly higher expression in turbot females after 90 dpf. Thus, 90 dpf was proposed as the development stage representing the onset of sex differentiation in the turbot based in molecular signs [
9] although as it was mentioned in the introduction, more recent data showed morphological gonad differences at earlier developmental stages [
11,
21].
Finally, we have compared our results on tissue-expression pattern of genes involved in the sex differentiation of turbot with those obtained in other teleost species to try to elucidate possible histological or morphological differences which could underlay in gonad differentiation in this group of vertebrates.
3.2.1. Vasa
The identification of
Vasa as a specific germ cell marker in the gonads of a multitude of vertebrate species, including fishes [
32], has proved its conserved an essential role in germ cell development. Studies on this gene have facilitated the progress in knowing germ cell development and evolution of its localization in the animal kingdom.
Vasa is the most characterized germ cell marker in teleostean species, including turbot, mainly by RT-PCR and/or qPCR analysis, but also by ISH or immunohistochemistry (IHC) techniques [
32]. In fact, qPCR analyses in the turbot showed that
Vasa expression is the first molecular signs (along with that of
gsdf and
tdrd1 gene expression) of sex differentiation which started to increase between 75-90 dpf [
9].
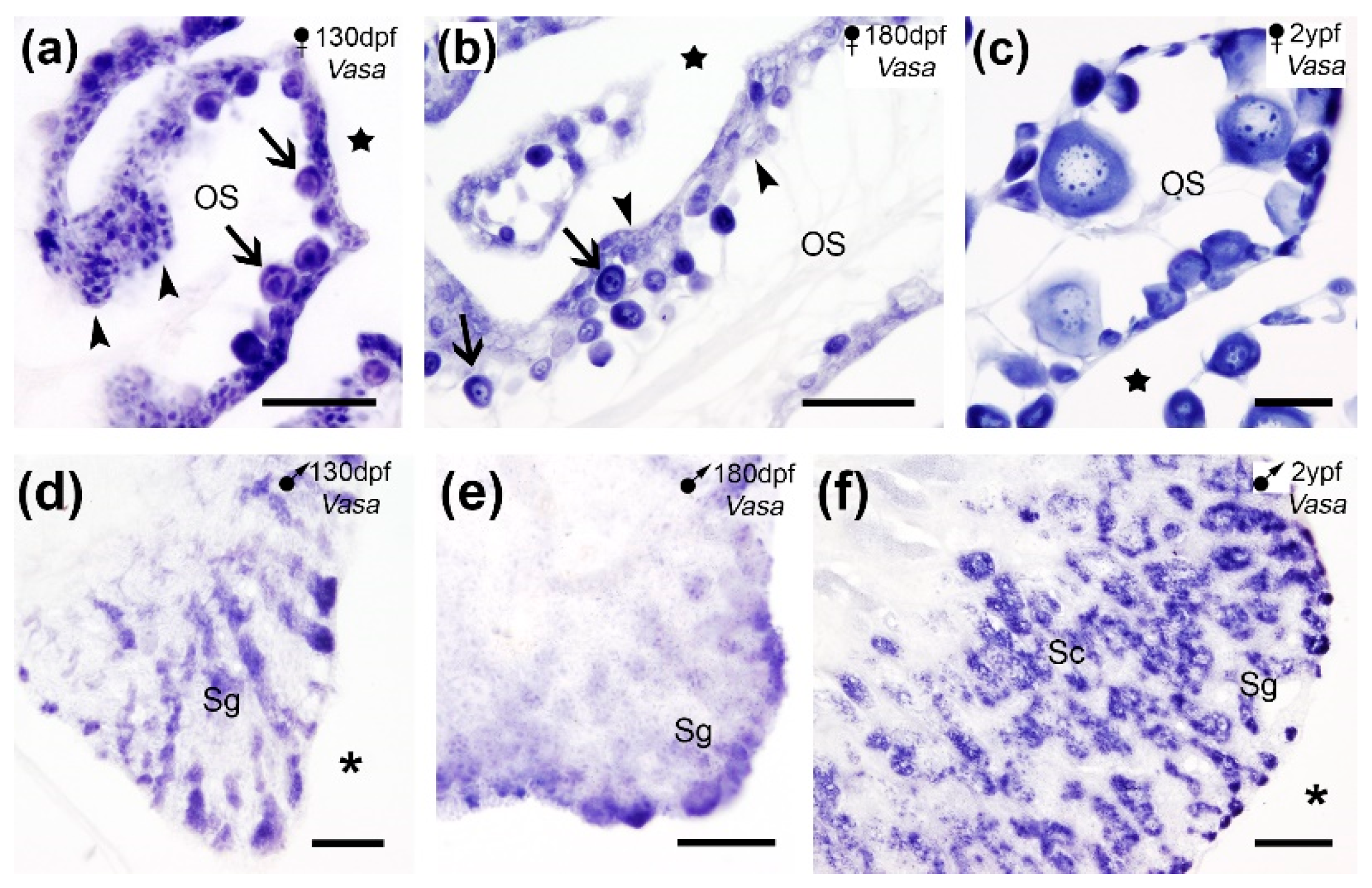
Our SISH results showed
Vasa expression in germ cells on paraffin sections of turbot gonads. In the ovary,
Vasa mRNA was detected in oogonia and the ooplasm of the primary oocytes (
Figure 3a-c). In the testis
Vasa expression was observed in spermatogonia and spermatocytes (
Figure 3d-f) but no expression was founded in spermatids, indicating a decline in its expression with the testis differentiation. Our results aligned with those previously reported by SISH and/or IHC in germ cells of differentiated gonads of the same species [
17,
22,
23] and in other teleosts [
30,
32,
33,
34,
35,
36,
37,
38,
39]. Thus,
Vasa is expressed in turbot primordial germ cells where it primarily participates in early gonadal differentiation [
17,
40] and remains active in differentiated gonads to regulate the development of germ cells [22, present results].
In addition, qPCR and transcriptome analyses showed a dimorphic sex-biased
Vasa expression with higher expression in females from the onset of gonadal differentiation (90 dpf) onwards [
9,
20]. Nevertheless, turbot adults showed the highest
Vasa expression levels in testes (3.5-fold more than ovary) probably due to the delay in primordial germ cells proliferation in males that occurs in this species [
22]. However, our SISH results did not support this female-biased expression pattern maybe due to detection of different transcripts of
Vasa homologues previously reported in the turbot [
17,
22]. Another reason for
Vasa sex-biased expression could be environmental factors which alter gene expression [
6] as it was reported in medaka which showed a decrease in
Vasa expression under hypoxia which leading to a male-biased population [
41].
3.2.2. Cyp19a1a
Cyp19a1a gene encodes for aromatase, a key enzyme in ovary differentiation, necessary for the starting and maintenance of the ovarian fate throughout development since it catalyzes the transformation of androgens into estrogens and its repression leads to testicular development [
42].
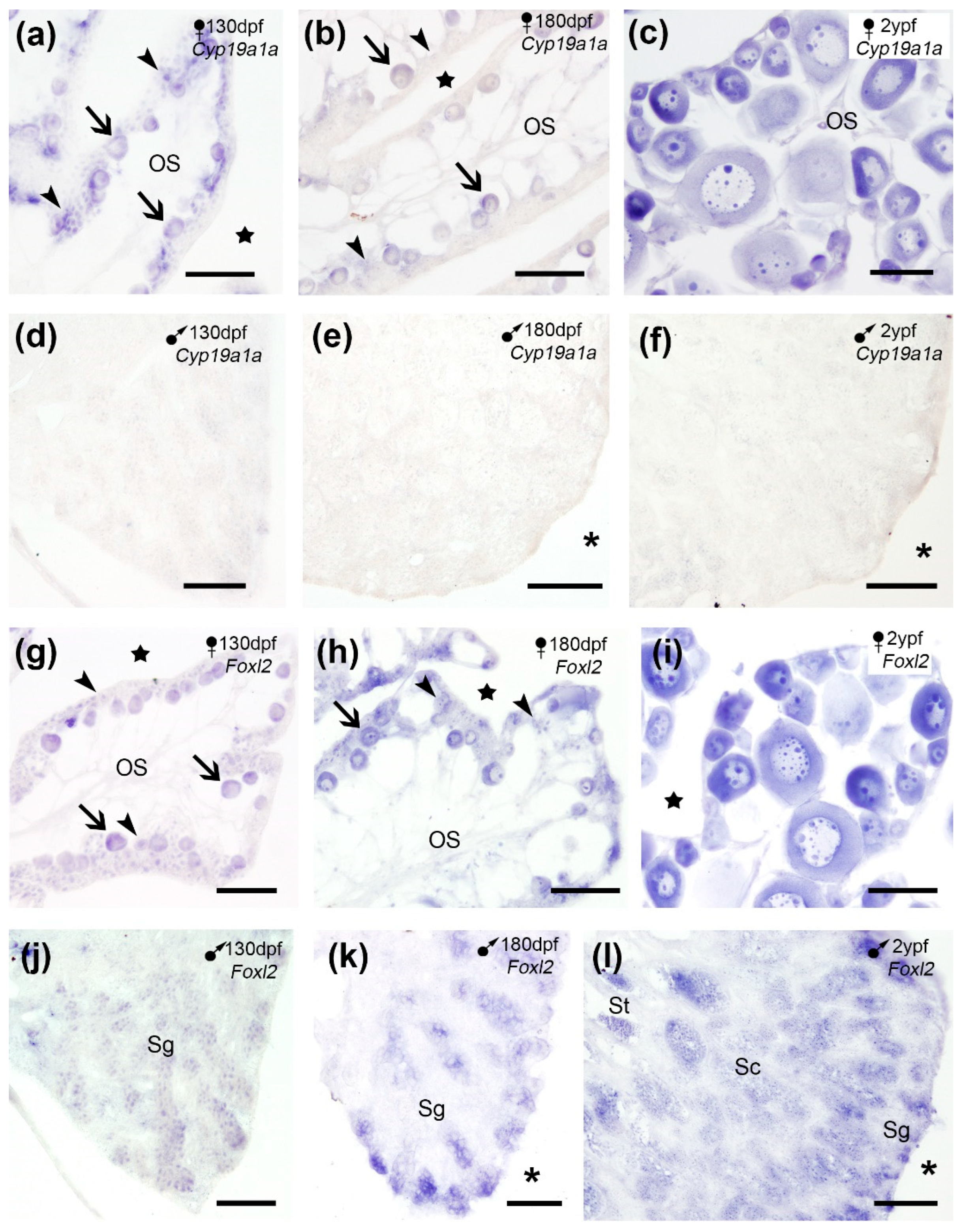
In our study,
Cyp19a1a mRNA was detected in oogonia and the ooplasm of the primary oocytes in juveniles (
Figure 4a-b) which increased in adults (
Figure 4c), while no
Cyp19a1a expression was observed in the turbot testis (
Figure 4d-f). These results were in concordance with previous qPCR and transcriptomic analyses in turbot which reported a sex-dimorphic
Cyp19a1a expression which allows discriminating males and females from 105 dpf until 435 dpf [
9,
13] and maintained in adults [
20]. Thus,
Cyp19a1a is a female-biased gene in turbot suggesting its key role in female differentiation and reproduction [9,13,20,present results].
The cellular localizations of
Cyp191a transcripts were also investigated using SISH or IHC in the ovary of the killifish [
43], the medaka [
44], the catfish [
45]) and the zebrafish [
46], where they were detected in the primary oocytes suggesting a contribution of germ cells in the estrogen synthesis in some teleosts species, including the turbot [present results]. Moreover, numerous molecular analyses have revealed
Cyp191a expression in the ovary of many other fish species so this gene have been considered as a reliable early marker of ovarian differentiation in these vertebrates [
6].
Cyp19a1a is not expressed in the turbot males [present results] but its expression was detected by RT-PCR and/or SISH and IHC in the testis of other teleostean species [
7,
43,
46,
47,
48] so
Cyp19a1a seems to be an indispensable gene not only for female but also for male sex differentiation in some fish. Therefore, data obtained until now about the sex steroid pathway indicate that it is not conserved in fish, supporting the notion of flexibility in sexual determination and differentiation across species.
3.2.3. Foxl2
Foxl2 gene encodes the Forkhead Box (Fox) protein L2, a transcription factor involved and considered a good marker of ovarian differentiation which is highly conserved in vertebrates [
49].
Our results in the turbot ovary showed
Foxl2 expression in oogonia and the ooplasm of primary oocytes in juveniles (
Figure 4g-h) which increased in adults (
Figure 4i). In the testes of juveniles
Foxl2 signal was detected in spermatogonia preferably located close to tunica albuginea (
Figure 4j-k), and in spermatogonia, spermatocytes and spermatids in adults (
Figure 4j).
Foxl2 expression in ovarian germ cells was also previously reported by IHC in the ovary of some teleots such as the zebrafish [
46], the Atlantic salmon [
48] and two medaka species [
29,
50] while its expression in testicular germ cells were only described in the zebrafish [
46] and the Atlantic salmon [
48]. On the other hand, SISH or IHC have revealed a
Foxl2 expression restricted to somatic ovarian (follicular/granulosa) cells in other teleostean species [
45,
51,
52]. These data suggest a different role for this gene in the regulation of gonad differentiation depending on the species and evidence the need of more research in
Foxl2 characterization in fish to clarify its functions in developing and adult gonads.
In the turbot ovary,
Foxl2 showed a similar expression to
Cyp19a1a [present results] in accordance with previous qPCR analyses which showed a high
Foxl2 expression in turbot females from 105 dpf onwards positively correlated with
Cyp19a1a [
9]. In fact,
Foxl2 has been described as a direct transcriptional activator of
Cyp19a1a in teleosts [
6,
7], including the turbot [
9,
13]. However, although both genes are co-expressed during turbot sex differentiation until 90 dpf in both sexes,
Cyp19a1a expression decays at later stages in males while
Foxl2 expression maintains through development [
9,
13,
20] according to our SISH observations in juveniles and adult turbot. These results suggest other role for
Foxl2 not related to the activation of aromatase in turbot [
9].
3.2.4. Anti-Müllerian Hormone (Amh)
Anti-Müllerian hormone (AMH) is mainly known by inducing the regression of Mullerian ducts, a characteristic feature of the female urogenital system in tetrapods [
53]. Although Müllerian ducts are lacking in fishes, AMH homologs have been identified in many species of teleosts, indicating a different role for the
Amh gene which acts a master-key gene of SD in some fish species [
5,
54]. Studies in turbot [
9] and other teleosts species [
54,
55,
56] have pointed to the inhibitory effect of
Amh gene in proliferation of germ cells whose number may be crucial for gonadal fate in fish. In fact, a low cell number correlates with male development at least in the zebrafish and the medaka [
57,
58] and the same could happen in the turbot gonad due to the negative regulation of germ cell proliferation by
Amh expression. On the contrary, high number of germ cells seems to be linked with ovarian differentiation, and proliferative germ cells might trigger
Cyp19a1a expression in the somatic cells of the gonad [
42,
57,
58].
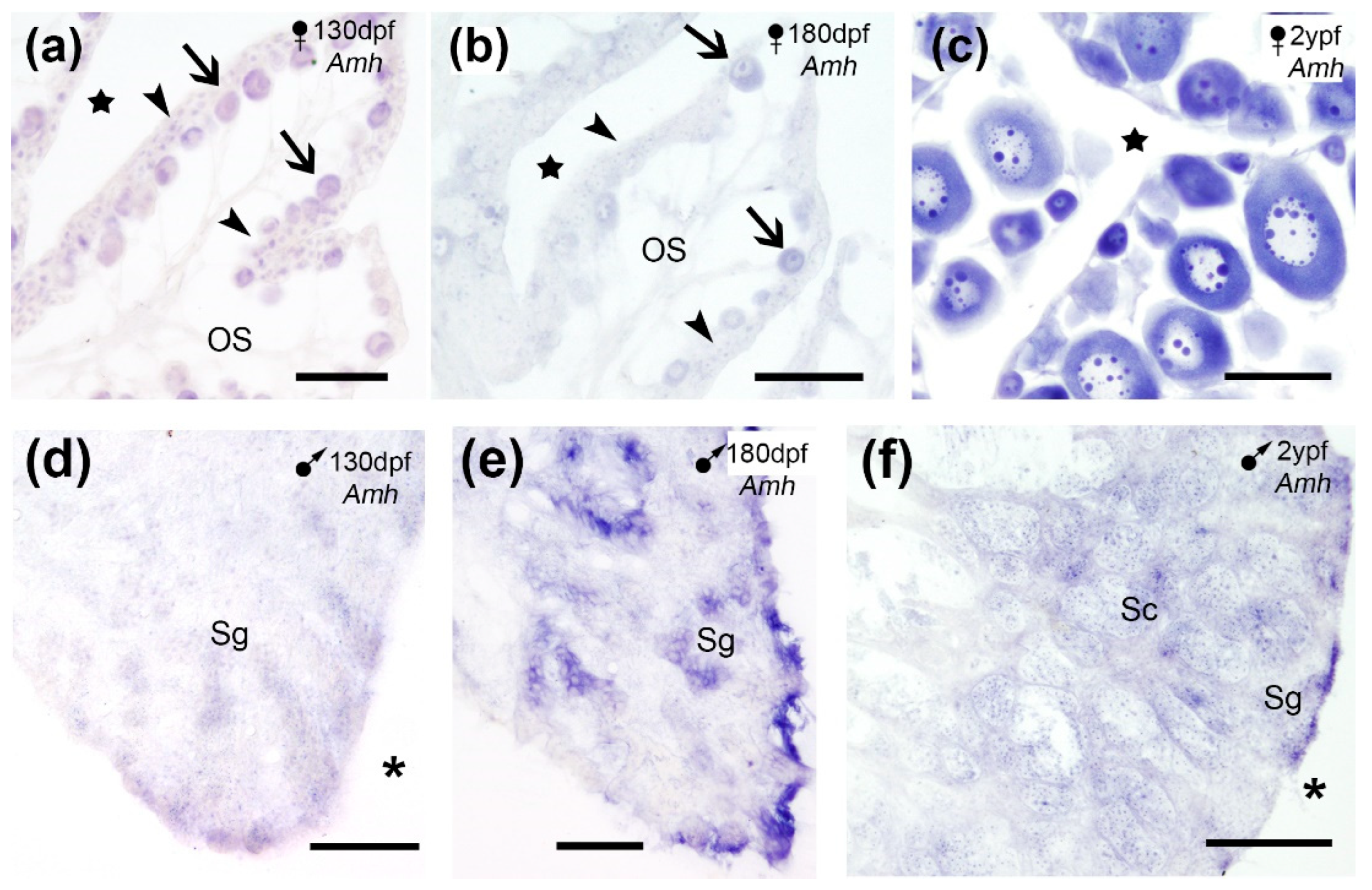
Considered as a gonad specific factor for male sex differentiation, our results in turbot gonads showed
Amh RNAm signal in both sexes. In the ovary a weak
Amh expression was detected in oogonia and the ooplasm of primary oocytes in juveniles (
Figure 5a-b) which increased in adults (
Figure 5c), like
Cyp19a1a and
Foxl2 expressions described above [present results]. In the testis,
Amh mRNA was revealed in spermatogonia at the edge of the testes in juveniles (
Figure 5d-e) and spermatogonia and spermatocytes in adults (
Figure 5f), like
Foxl2 expression in turbot males [present results]. SISH revealed
Amh expression in differentiated turbot ovary and testis in consistence with studies by qPCR which showed
Amh expression from 90 to 135dpf, higher in males [
9]. These results suggested that
Amh has a role in turbot gonad differentiation with functions in both sexes.
However, in other teleosts species SISH has shown
Amh expression restricted to ovarian and testicular somatic cells [
54,
55,
59,
60,
61,
62]. Thus, our results in turbot gonads deviate from this pattern, showing
Amh expression in both male and female germ cells, as previously reported by IHC for the first time in a fish - in the ovary and testis of the Atlantic salmon [
48] - and later in the ovary of the Nile tilapia [
55] where somatic cells also exhibited
Amh expression. Therefore,
Amh expression is not restricted to the somatic cells in all fish suggesting a role for this gene (and likely also for
Foxl2 and
Cyp19a1a) on germ cells probably related with regulation of the transcription and activity of specific genes in the surrounding cells through secretory signaling as it was proposed for the Atlantic salmon [
48].
Therefore,
Cyp19a1a,
Foxl2, and
Amh expression were detected by SISH in germ cells of differentiated turbot gonads in accordance with that observed by IHC in the Atlantic salmon [
48] and with previous qPCR analyses which showed the expression of these three genes in turbot gonads [
9]. On the other hand, many molecular and histological analyses on
Cyp19a1a,
Foxl2, and
Amh in teleost gonads [
27,
30,
54,
56,
61] showed that their expression can vary depending on the species, developmental stage, sex, or temperature proving the huge variety of mechanisms sexual development present in fish.
3.2.5. Sox Genes
We have performed SISH to detect the
Sox2,
Sox17 and
Sox19 expression in the turbot differentiated gonads previously reported by qPCR and microarray in undifferentiated and differentiated gonads of this species [
9].
The
Sox gene family encodes a group of conserved transcription factors related to the mammalian testis-determining factor
sry (sex determining region Y) and they are developmental regulators of multiple processes in vertebrates, including sex determination and reproduction in fish [
63]. Sox genes are classified based upon their HMG box sequences into subgroups A to K found throughout the animal kingdom [
64] most of them identified in teleosts (except soxA, soxG and sox J subgroups) in which Sox genes play essential roles in fish embryogenesis and ontogenesis [
63].
Sox2 and
Sox19 belong to the B1 subgroup and
Sox17 to the F group.
3.2.5.1. B1 Sox Genes: Sox2 and Sox19
In turbot,
Sox2 gene deserved special attention since it is located within the turbot main sex determination region [
24] and it has been proposed as a candidate SD in this species [
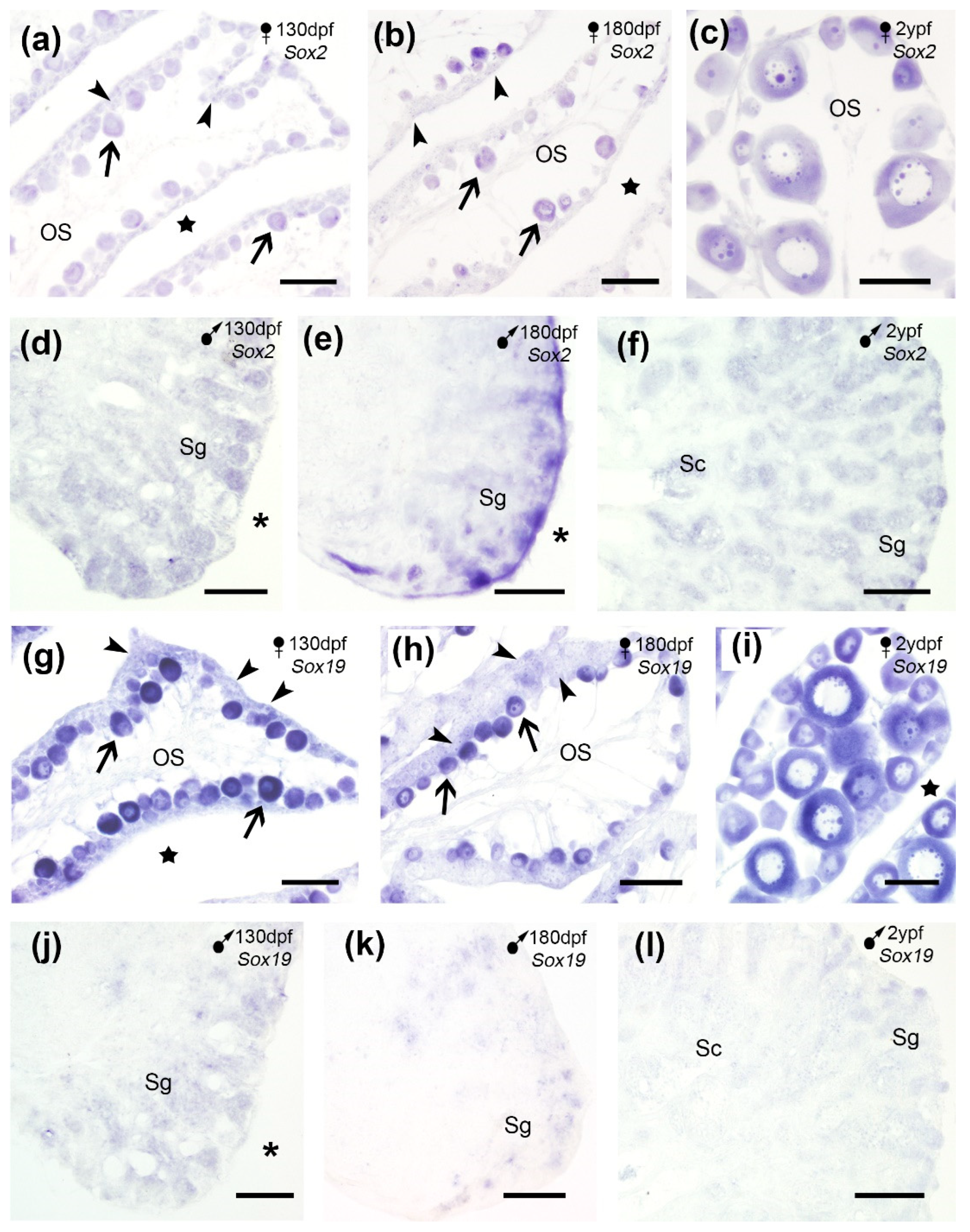
15]. Our results in the turbot ovary showed a weak
Sox2 expression in the oogonia which increased along the primary oocyte development (
Figure 6a-c) while in testis,
Sox2 was exclusively expressed in spermatogonia-containing cysts, close to the tunica albuginea in juveniles and adults (
Figure 6d-f), suggesting effects during the proliferative stage. These observations were in accordance with previous reported in qPCR and microarray analyses where no differences were observed in
Sox2 expression between turbot males and females [
9]. As far as we know, there are no
Sox2 expression studies by SIHS or IHC in fish differentiated gonads but qRT-PCR analyses in the ovary or testes of the rohu carp [
65], the channel catfish [
66], and the Japanese flounder [
67] suggesting a role for
Sox2 in gonadogenesis of fish.
On the other hand, a strong
Sox19 expression, a fish-specific gene, was detected by SISH in oogonia and in the ooplasm of the primary oocytes of juvenile and adult turbot (
Figure 6g-i) while in testis its expression was much weaker (
Figure 6j-l). Thus,
Sox19 can be considered as a female-biased gene in the turbot due higher expression levels were observed in the ovary which is maintained until adulthood suggesting a role in ovarian differentiation. Accordingly, previous qPCR analyses in turbot gonads reported an increase of
Sox19 transcription from 75 to 90 dpf in both sexes, but only in females from 105 dpf to 135 dpf [
9], a dimorphic expression which is maintained at 400 dpf [
13] and adult specimens [
20].
To our knowledge, there are no histological studies on
Sox19 expression in fish except for the common carp where
Sox19 protein was observed in somatic (i.e., granulosa) cells of postvitellogenic oocytes suggesting a role in late ovarian differentiation in this species [
68]. Moreover, RT-PCR analyses showed that this gene is more expressed in adult ovary than in the testis in the common carp [
68] and in the European sea bass [
69] and playing a role in the differentiation of the ovary but not the testis.
Furthermore,
Sox2 and
Sox19 have a crucial role in neurogenesis and nervous system development in vertebrates, including fish [
70]. In fact,
Sox2 and
Sox19 present a similar expression pattern by SISH in the developing and adult turbot forebrain which suggest similar functions in brain development in this species [
71]. Moreover,
Sox2 expression in subependymal cells of the preoptic and hypothalamic regions could indicate a role for this gene in the control of reproduction, onset of puberty and sexual maturation in turbot [
71].
Sox2 and
Sox19 expression were also reported in the brain of other teleostean species, so their involvement in the regulation of the reproductive system through the brain-pituitary-gonadal axis in these vertebrates [
63,
67,
69,
70,
72,
73]. A recent study by histological methods reported a distinct sex difference in the turbot brain during the gonadogenesis with a total brain volume and a pituitary gland size larger in mature males [
26] as it was observed in the brain of other teleosts species [
70]. However, how the brain regulates sex determination and differentiation in teleosts is still needs to be unraveled [
7].
Therefore,
Sox2 and
Sox19, as other SoxB1 genes (i.e.,
Sox1 and
Sox3), play an essential regulatory role in gonadal development, sex determination and differentiation, and in nervous system development of fish [
61,
68].
3.2.5.2. F Sox Genes: Sox17
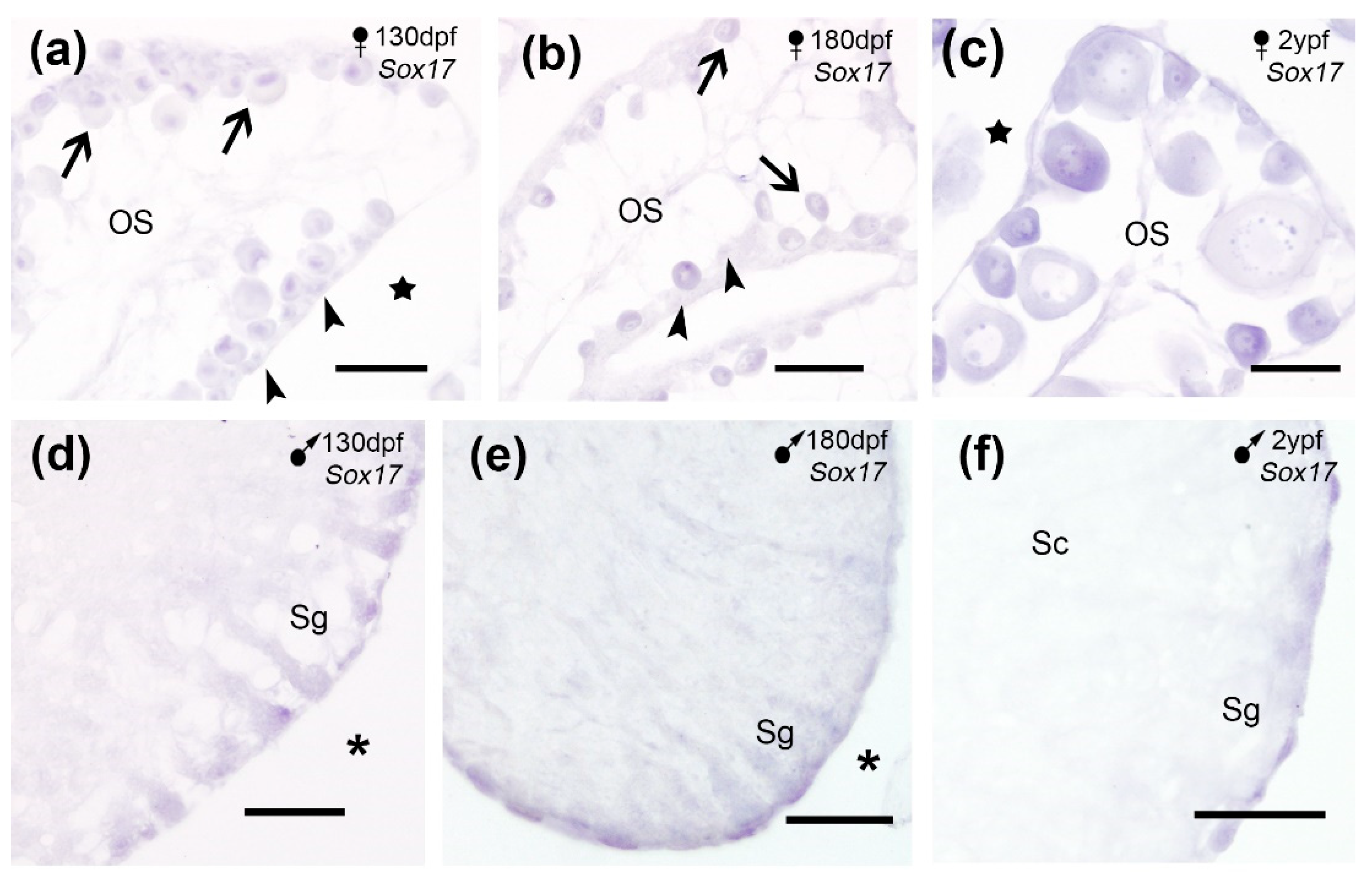
Sox17 showed a very weak no sexual dimorphic expression in juveniles and adult turbot gonads, which was detected in oogonia and the ooplasm of the primary oocytes (
Figure 7a-c) and in spermatogonia in the testis (
Figure 7d-f). In accordance with our SISH observations, transcriptomic studies reported a weak
Sox17 expression in ovary and testis of the adult turbot [
20]. In fact, qPCR and microarray analysis of
Sox17 expression in turbot reported a peak of expression around the onset of sex differentiation (90 dpf) in both sexes concomitantly with a raise in the expression of other germ cell markers such as
Vasa [
9,
13]. These data suggest some function for this gene, previously mapped in sex-related QTL [
25], in early gonadogenesis and germ cell proliferation [
9].
Although as far as we know, there are no
Sox17 expression studies by SISH in fish differentiated gonads except for the rice eel where
Sox17 mRNA was detected in the germinal lamellae of ovaries (but not in the oocytes), ovotestis and testes as well as in developing germ cells of the testis during sex change from male to female which this species undergoes along its life [
74]. Thus, most of
Sox17 expression data in teleost gonads were obtained by RT-PCR which detected
Sox17 transcripts in the gonads of the Japanese flounder [
67], the Nile tilapia [
72], the channel catfish [
73], the rice eel [
74], the sea bass [
75], the medaka [
76], and the Yellow River carp [
77]. Although all these data was obtained by molecular analyses and avoid attribute a specific function to
Sox17 in fish reproduction, allow knowing that this gene is expressed in the testis and ovary of fish, suggesting its important role in sex differentiation of these vertebrates [
63,
70].
To our knowledge there is no still data on the
Sox17 expression in the turbot brain, but
Sox17 transcripts were detected in the brain of the Japanese flounder [
67], the rice eel [
74], the sea bass [
75], and the Yellow River carp [
77] suggesting that SoxF genes may be involved in the regulation of the nervous system as it was pointed for SoxB1 genes.
4. Conclusions
In the present study, several genes were analyzed for the first time by in situ hybridization (ISH) in turbot gonads at different developmental stages. We performed ISH on differentiated ovaries and testes paraffin sections to know the cell/tissue specific expression of sex differentiation and gonadal development related genes in this species. Vasa, Foxl2, Amh, Sox2 and Sox17 expression were observed in both ovary and testis while Cyp19a1a and Sox19 expression was ovary specific, suggesting a role for these genes in gonad development or reproductive functions in turbot. Moreover, ISH results support previous data on these genes expression by molecular analyses in the turbot gonads and highlight the cellular types, i.e., germ cells, where they are expressed. Therefore, we consider ISH provides complementary information to the functional studies with transcriptomic or proteomic techniques so gives spatial information on RNA in a tissue sample, i.e., cellular localization, to better understand the molecular mechanisms of sex differentiation in vertebrates. However, further research in gonadal sections of early developmental turbot stages is needed to gain knowledge on the gonadal development in this flatfish species from a histological point of view. Finally, comparison with the results obtained in differentiated gonads of other teleosts species showed a similar tissue-specific gene expression but also revealed the notable variety of sexual development and differentiation molecular mechanism in fish.
Author Contributions
Conceptualization, A.V.; methodology, F.A and X.T.; supervision of the data collection and research activity execution, F.A and A.V.; analysis and interpretation of the data F.A and A.V.; writing-original draft and designing of the figures, F.A. and V.A; writing, review and editing, F.A., P.M. and A.V; funding acquisition, A.V. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the government of Xunta de Galicia, grant number 10MMA200027PR, and by Spanish Economy and Competitiveness Ministry-FEDER, grant number BFU2014-58631-P.
Institutional Review Board Statement
The animal study protocol was approved by the Ethics Committee of the University of Santiago de Compostela and all procedures were in accordance with the guidelines established by the European Union Directive (2010/63/UE) and by the Spanish Royal Decree 118/2021 for animal experiments.
Data Availability Statement
The original contributions presented in this study are included in the article. Further inquiries can be directed to the corresponding author.
Acknowledgments
We thank the Instituto Español de Oceanografía de Vigo for generously providing the fish samples.
Conflicts of Interest
The authors declare no conflicts of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
References
- Robledo, D.; Hermida, M.; Rubiolo, J.A.; Fernández, C.; Blanco, A.; Bouza, C.; Martínez, P. Integrating genomic resources of flatfish (Pleuronectiformes) to boost aquaculture production. Comp. Biochem. Physiol. D: Genomics Proteomics 2017, 21, 41–45. [Google Scholar] [CrossRef]
- Imsland, A.K.; Folkvord, A.; Grung, G.L.; Stefansson, S.O.; Taranger, G.L. Sexual dimorphism in growth and maturation of turbot Scophthalmus maximus (Rafinesque 1810). Aquac. Res. 1997, 28, 101–114. [Google Scholar] [CrossRef]
- Piferrer, F.; Cal, R.; Gómez, C.; Álvarez-Blázquez, B.; Castro, J.; Martínez, P. Induction of gynogenesis in the turbot (Scophthalmus maximus): effects of UV irradiation on sperm motility the Hertwig effect and viability during the first 6 months of age. Aquaculture 2004, 238, 403–419. [Google Scholar] [CrossRef]
- Nagahama, Y.; Chakraborty, T.; Paul-Prasanth, B.; Ohta, K.; Nakamura, M. Sex determination, gonadal sex differentiation, and plasticity in vertebrate species. Physiol Rev. 2021, 101(3), 1237–1308. [Google Scholar] [CrossRef] [PubMed]
- Kitano, J.; Ansai, S.; Takehana, Y.; Yamamoto, Y. Diversity and convergence of sex-determination mechanisms in teleost fish. Annu. Rev. Anim. Biosci. 2024, 12, 233–259. [Google Scholar] [CrossRef]
- Yu, Y.; Chen, M.; Shen, Z.G. Molecular biological, physiological, cytological, and epigenetic mechanisms of environmental sex differentiation in teleosts: A systematic review. Ecotoxicol. Environ. Saf. 2023, 267, 115654. [Google Scholar] [CrossRef] [PubMed]
- Rajendiran, P.; Jaafar, F.; Kar, S.; Sudhakumari, C.; Senthilkumaran, B.; Parhar, I.S. Sex determination and differentiation in Teleost: Roles of genetics, environment, and brain. Biology (Basel) 2021, 10(10), 973. [Google Scholar] [CrossRef]
- Martínez, P.; Viñas, A.M.; Sánchez, L.; Díaz, N.; Ribas, L.; Piferrer, F. Genetic architecture of sex determination in fish: applications to sex ratio control in aquaculture. Front. Genet. 2014, 5, 340. [Google Scholar] [CrossRef]
- Robledo, D.; Ribas, L.; Cal, R.; Sánchez, L.; Piferrer, F.; Martínez, P.; Viñas, A. Gene expression analysis at the onset of sex differentiation in turbot (Scophthalmus maximus). BMC genomics 2015, 16, 973. [Google Scholar] [CrossRef]
- Martínez, P.; Robledo, D.; Taboada, X.; Blanco, A.; Moser, M.; Maroso, F.; Hermida, M.; Gómez-Tato, A.; Álvarez-Blázquez, B.; Cabaleiro, S.; Piferrer, F.; Bouza, C.; Lien, S.; Viñas, A. A genome-wide association study, supported by a new chromosome-level genome assembly, suggests Sox2 as a main driver of the undifferentiated ZZ/ZW sex determination of turbot (Scophthalmus maximus). Genomics 2021, 113(4), 1705–1718. [Google Scholar] [CrossRef]
- Meng, Z.; Cheng, L.; Hu, P.; Liu, X.; Zhang, H.; Liu, B.; Yang, Z. Histological chronology and thermosensitive period of gonadal sex differentiation in turbot (Scophthalmus maximus) 2026. Aquac. Rep. 2026, 46, 103307. [Google Scholar] [CrossRef]
- Vandeputte, M.; Dupont-Nivet, M.; Chavanne, H.; Chatain, B. A polygenic hypothesis for sex determination in the European sea bass Dicentrarchus labrax. Genetics 2007, 176, 1049–1057. [Google Scholar] [CrossRef]
- Ribas, L.; Robledo, D.; Gómez-Tato, A.; Viñas, A.; Martínez, P.; Piferrer, F. Comprehensive transcriptomic analysis of the process of gonadal sex differentiation in the turbot (Scophthalmus maximus). Mol. Cell Endocrinol. 2016, 422, 132–149. [Google Scholar] [CrossRef]
- Haffray, P.; Lebegue, E.; Jeu, S.; Guennoc, M.; Guiguen, Y.; Baroiller, J.F.; Fostier, A. Genetic determination and temperature effects on turbot Scophthalmus maximus sex differentiation: An investigation using steroid sex-inverted males and females. Aquaculture 2009, 294, 30–36. [Google Scholar] [CrossRef]
- Martínez, P.; Bouza, C.; Hermida, M.; Fernández, J.; Toro, M.A.; Vera, M.; Pardo, B.; Millán, A.; Fernández, C.; Vilas, R.; Viñas, A.; Sánchez, L.; Felip, A.; Piferrer, F.; Ferreiro, I.; Cabaleiro, S. Identification of the major sex-determining region of turbot (Scophthalmus maximus). Genetics 2009, 183, 1443–1452. [Google Scholar] [CrossRef]
- Figueras, A.; Robledo, D.; Corvelo, A.; Hermida, M.; Pereiro, P.; Rubiolo, J.A.; Gómez-Garrido, J.; Carreté, L.; Bello, X.; Gut, M.; Gut, I.; Marcet-Houben, M.; Forn-Cuní, G.; Galán, B.; García, J.L.; Abal-Fabeiro, J.L.; Pardo, B.G.; Taboada, X.; Fernández, C.; Vlasova, A.; Hermoso-Pulido, A.; Guigó, R.; Álvarez-Dios, J.A.; Gómez-Tato, A.; Viñas, A.; Maside, X.; Gabaldón, T.; Novoa, B.; Bouza, C.; Alioto, T.; Martínez, P. Whole genome sequencing of turbot (Scophthalmus maximus; Pleuronectiformes): A fish adapted to demersal life. DNA Res. 2016, 23(3), 181–192. [Google Scholar] [CrossRef]
- Lin, F.; Xu, S.; Ma, D.; Xiao, Z.; Zhao, C.; Xiao, Y.; Chi, L.; Liu, Q.; Li, J. Germ line specific expression of a Vasa homologue gene in turbot (Scophthalmus maximus): Evidence for Vasa localization at cleavage furrows in euteleostei. Mol. Reprod. Dev. 2012, 79, 803–813. [Google Scholar] [CrossRef]
- Lin, F.; Zhao, C.Y.; Xu, S.H.; Ma, D.Y.; Xiao, Z.Z.; Xiao, Y.S.; Xu, C.A.; Liu, Q.H.; Li, J. Germline-specific and sexually dimorphic expression of a dead end gene homologue in turbot (Scophthalmus maximus). Theriogenology 2013, 80, 665–672. [Google Scholar] [CrossRef]
- Hu, Y.; Huang, M.; Wang, W.; Guan, J.; Kong, J. Characterization of gonadal transcriptomes from the turbot (Scophthalmus maximus). Genome 2016, 59(1), 1–10. [Google Scholar] [CrossRef] [PubMed]
- Ma, D.; Ma, A.; Huang, Z.; Wang, G.; Wang, T.; Xia, D.; Ma, B. Transcriptome analysis for identification of genes related to gonad differentiation growth immune response and marker discovery in the turbot (Scophthalmus maximus). PloS One 2016, 11, e0149414. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Xu, S.; Liu, Y.; Wang, Y.; Liu, Q.; Li, J. Gonadogenesis analysis and sex differentiation in cultured turbot (Scophthalmus maximus). Fish Physiol. Biochem. 2017, 43, 265–278. [Google Scholar] [CrossRef]
- Pacchiarini, T.; Sarasquete, C.; Cabrita, E. Development of interspecies testicular germ-cell transplantation in flatfish. Reprod. Fertil. Dev. 2014, 26(5), 690–702. [Google Scholar] [CrossRef]
- Zhao, C.; Liu, Q.; Xu, S.; Xiao, Y.; Wang, W.; Yang, J.; Yang, Y.; Wang, Y.; Song, Z.; Li, J. Identification of type A spermatogonia in turbot (Scophthalmus maximus) using a new cell-surface marker of lymphocyte antigen 75 (ly75/CD205). Theriogenology 2018, 113, 137–145. [Google Scholar] [CrossRef]
- Taboada, X.; Hermida, M.; Pardo, B.G.; Vera, M.; Piferrer, F.; Viñas, A.; Bouza, C.; Martínez, P. Fine mapping and evolution of the major sex determining region in turbot (Scophthalmus maximus). G3 2014, 4, 1871–1880. [Google Scholar] [CrossRef]
- Viñas, A.; Taboada, X.; Vale, L.; Robledo, D.; Hermida, M.; Vera, M.; Martínez, P. Mapping of DNA sex-specific markers and genes related to sex differentiation in turbot (Scophthalmus maximus). Mar. Biotechnol. 2012, 14, 655–663. [Google Scholar] [CrossRef]
- Zhao, C.; Chi, L.; Xiao, Y.; Li, B.; Lu, Y.; Cui, Y.; Waang, W.; Li, J. Morphological and histological changes in the brain of turbot (Scophthalmus maximus) with gonadal development. Acta Oceanol. Sin. 2022, 41, 115–122. [Google Scholar] [CrossRef]
- Heule, C.; Salzburger, W.; Böhne, A. Genetics of sexual development: an evolutionary playground for fish. Genetics 2014, 196, 579–591. [Google Scholar] [CrossRef]
- Hale, M.C.; Xu, P.; Scardina, J.; Wheeler, P.A.; Thorgaard, G.H.; Nichols, K.M. Differential gene expression in male and female rainbow trout embryos prior to the onset of gross morphological differentiation of the gonads. BMC Genomics 2011, 12, 404. [Google Scholar] [CrossRef] [PubMed]
- Herpin, A.; Adolfi, M.C.; Nicol, B.; Hinzmann, M.; Schmidt, C.; Klughammer, J.; Engel, M.; Tanaka, M.; Guiguen, Y.; Schartl, M. Divergent expression regulation of gonad development genes in medaka shows incomplete conservation of the downstream regulatory network of vertebrate sex determination. Mol. Biol. Evol. 2013, 30, 2328–2346. [Google Scholar] [CrossRef]
- Li, C.; Li, Y.; Qin, C.; Yu, C.; Hu, J.; Guo, C.; Wang, Y. Determination of the timing of early gonadal differentiation in silver pomfret, Pampus argenteus. Anim. Reprod. Sci. 2024, 261, 107373. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wu, Y.; Zhang, Y.; Zhang, J.; Chu, P.; Chen, K.; Liu, H.; Luo, Q.; Fei, S.; Zhao, J.; Ou, M. Histological observations and transcriptome analyses reveal the dynamic changes in the gonads of the blotched snakehead (Channa maculata) during sex differentiation and gametogenesis. Biol. Sex. Differ. 2024, 15(1), 70. [Google Scholar] [CrossRef]
- Begum, S.; Gnanasree, S.M.; Anusha, N.; Senthilkumaran, B. Germ cell markers in fishes- A review. Aquac. Fish. 2022, 7, 540–552. [Google Scholar] [CrossRef]
- Cao, M.; Yang, Y.; Xu, H.; Duan, J.; Cheng, N.; Wang, J.; Hu, W.; Zhao, H. Germ cell specific expression of Vasa in rare minnow, Gobiocypris rarus. Comp. Biochem. Physiol. A 2012, 162, 163–170. [Google Scholar] [CrossRef]
- Haugen, T.; Almeida, F.F.; Andersson, E.; Bogerd, J.; Male, R.; Skaar, K.S.; Taranger, G.L. Sex differentiation in Atlantic cod (Gadus morhua L): morphological and gene expression studies. Reprod. Biol. Endocrinol. 2012, 10, 47. [Google Scholar] [CrossRef]
- Pacchiarini, T.; Cross, I.; Leite, R.B.; Gavaia, P.; Ortiz-Delgado, J.B.; Pousão-Ferreira, P.; Rebordinos, L.; Sarasquete, C.; Cabrita, E. Solea senegalensis Vasa transcripts: molecular characterisation, tissue distribution and developmental expression profiles. Reprod. Fertil. Dev. 2013, 25(4), 646–660. [Google Scholar] [CrossRef]
- Úbeda-Manzanaro, M.; Rebordinos, L.; Sarasquete, C. Cloning and characterization of Vasa gene expression pattern in adults of the Lusitanian toadfish Halobatrachus didactylus. Aquatic Biology 2014, 21, 37–46. [Google Scholar] [CrossRef]
- Chen, X.; Zhu, Y.; Zhu, T.; Song, P.; Guo, J.; Zhong, Y.; Gui, L.; Li, M. Vasa identifies germ cells in embryos and gonads of Oryzias celebensis. Gene 2022, 823, 146369. [Google Scholar] [CrossRef]
- Wang, M.; Ding, H.; Wu, S.; Wang, M.; Wei, C.; Wang, B.; Bao, Z.; Hu, J. Vasa is a potential germ cell marker in leopard coral grouper (Plectropomus leopardus). Genes 2022, 13(6), 1077. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Yan, L.; Zhao, C.; Zhang, B.; Zhang, B.; Qiu, L. Genetic characterization of primordial germ cells in spotted sea bass (Lateolabrax maculatus). Genes (Basel) 2025, 16(9), 1012. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Wang, X.; Liu, Q.; Xu, S.; Zhao, H.; Han, M.; Wang, Y.; Song, Z.; Li, J. Visualization of turbot (Scophthalmus maximus) primordial germ cells in vivo using fluorescent protein mediated by the 3’ untranslated region of nanos3 or vasa gene. Mar. Biotechnol. (NY) 2019, 21(5), 671–682. [Google Scholar] [CrossRef]
- Cheung, C.H.Y.; Chiu, J.M.Y.; Wu, R.S.S. Hypoxia turns genotypic female medaka fish into phenotypic males. Ecotoxicology 2014, 23, 1260–1269. [Google Scholar] [CrossRef]
- Guiguen, Y.; Fostier, A.; Piferrer, F.; Chang, C.F. Ovarian aromatase and estrogens: a pivotal role for gonadal sex differentiation and sex change in fish. Gen. Comp. Endocrinol. 2010, 165, 352–366. [Google Scholar] [CrossRef]
- Dong, W.; Willett, K.L. Local expression of CYP19A1 and CYP19A2 in developing and adult killifish (Fundulus heteroclitus). Gen. Comp. Endocrinol. 2008, 155(2), 307–317. [Google Scholar] [CrossRef]
- Park, J.W.; Tompsett, A.; Zhang, X.; Newsted, J.L.; Jones, P.D.; Au, D.; Kong, R.; Wu, R.S.S.; Giesi, J.P.; Hecker, M. Fluorescence in situ hybridization techniques (FISH) to detect changes in CYP19a gene expression of Japanese medaka (Oryzias latipes). Toxicol. Appl. Pharmacol. 2008, 232, 226–235. [Google Scholar] [CrossRef]
- Raghuveer, K.; Senthilkumaran, B.; Sudhakumari, C.C.; Sridevi, P.; Rajakumar, A.; Singh, R.; Murugananthkumar, R.; Majumdar, K.C. Dimorphic expression of various transcription factor and steroidogenic enzyme genes during gonadal ontogeny in the air-breathing catfish Clarias gariepinus. Sex. Dev. 2011, 5, 213–223. [Google Scholar] [CrossRef] [PubMed]
- Caulier, M.; Brion, F.; Chadili, E; Turies, C.; Piccini, B; Porcher, J.M.; Guiguen, Y.; Hinfray, N. Localization of steroidogenic enzymes and Foxl2a in the gonads of mature zebrafish (Danio rerio). Comp. Biochem. Physiol. A Mol. Integr. Physiol. 2015, 188, 96–106. [Google Scholar] [CrossRef]
- Böhne, A.; Heule, C.; Boileau, N.; Salzburger, W. Expression and sequence evolution of aromatase Cyp19a1a and other sexual development genes in East African cichlid fishes. Mol. Biol. Evol. 2013, 10, 2268–2285. [Google Scholar] [CrossRef]
- von Schalburg, K.R.; Gowen, B.E.; Rondeau, E.B.; Johnson, N.W.; Minkley, D.R.; Leong, J.S.; Davidson, W.S.; Koop, B.F. Sex-specific expression, synthesis and localization of aromatase regulators in one-year-old Atlantic salmon ovaries and testes. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 2013, 164(4), 236–246. [Google Scholar] [CrossRef]
- Bertho, S.; Pasquier, J.; Pan, Q.; Le Trionnaire, G.; Bobe, J.; Postlethwait, J.H.; Pailhoux, E.; Schartl, M.; Herpin, A.; Guiguen, Y. Foxl2 and its relatives are evolutionary conserved players in gonadal sex differentiation. Sex. Dev. 2016, 10, 111–129. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Zhang, Y.; Zhong, Y.; Guo, J.; Mengyue, L.; Gui, L.; Li, M. Molecular identification and expression analysis of foxl2 and sox9b in Oryzias celebensis. Aquac. Fish. 2021, 6(5), 471–478. [Google Scholar] [CrossRef]
- Wang, D.D.; Zhang, G.R.; Wei, K.J.; Ji, W.; Gardner, J.P.; Yang, R.B.; Chen, K.C. Molecular identification and expression of the Foxl2 gene during gonadal sex differentiation in northern snakehead Channa argus. Fish Physiol. Biochem. 2015, 41, 1419–1433. [Google Scholar] [CrossRef]
- Shu, C.; Wang, L.; Zou, C.; Tan, X.; Zou, Y.; Kong, L.; Wu, Z.; Wu, Q.; Wang, L.; Wang, G.; Li, Z.; You, F. Function of Foxl2 and Dmrt1 proteins during gonadal differentiation in the olive flounder Paralichthys olivaceus. Int. J. Biol. Macromol. 2022, 215, 141–154. [Google Scholar] [CrossRef] [PubMed]
- Adolfi, M.C.; Nakajima, R.T.; Nobrega, R.H.; Schartl, M. Intersex, hermaphroditism, and gonadal plasticity in vertebrates: Evolution of the Mullerian duct and Amh/Amhr2 signaling. Annu. Rev. Anim. Biosci 2019, 7, 149–172. [Google Scholar] [CrossRef] [PubMed]
- Pfennig, F.; Standke, A.; Gutzeit, H.O. The role of Amh signaling in teleost fish-multiple functions not restricted to the gonads. Gen. Comp. Endocrinol. 2015, 223, 87–107. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.Y.; Xiao, H.S.; Jie, M.M.; Dai, S.F.; Wu, X.; Li, M.H.; Wang, D.S. Amh regulate female folliculogenesis and fertility in a dose-dependent manner through Amhr2 in Nile tilapia. Mol. Cell. Endocrinol. 2020, 499, 110593. [Google Scholar] [CrossRef]
- Wu, K.; Yue, Y.; Zhou, L.; Zhang, Z.; Shan, H.; He, H.; Ge, W. Disrupting Amh and androgen signaling reveals their distinct roles in zebrafish gonadal differentiation and gametogenesis. Commun. Biol. 2025, 8, 371. [Google Scholar] [CrossRef]
- Tanaka, M.; Saito, D.; Morinaga, C.; Kurokawa, H. Cross talk between germ cells and gonadal somatic cells is critical for sex differentiation of the gonads in the teleost fish medaka (Oryzias latipes). Dev. Growth Differ. 2008, 50, 273–278. [Google Scholar] [CrossRef] [PubMed]
- Nagabhushana, A.; Mishra, R.K. Finding clues to the riddle of sex determination in zebrafish. J. Biosci. 2016, 41, 145–155. [Google Scholar] [CrossRef]
- Rodríguez-Marí, A.; Yan, Y.L.; BreMiller, R.A.; Wilson, C.; Cañestro, C.; Postlethwait, J.H. Characterization and expression pattern of zebrafish Anti-Müllerian hormone (Amh) relative to sox9a, sox9b and Cyp19a1a during gonad development. Gene Expr. Patterns 2005, 5, 655–667. [Google Scholar] [CrossRef]
- Vizziano, D.; Randuineau, G.; Baron, D.; Cauty, C.; Guiguen, Y. Characterization of early molecular sex differentiation in rainbow trout Oncorhynchus mykiss. Dev. Dyn. 2007, 236, 2198–2206. [Google Scholar] [CrossRef]
- Yamaguchi, T.; Kitano, T. Amh/Amhr2 signaling causes masculinization by inhibiting estrogen synthesis during gonadal sex differentiation in Japanese flounder (Paralichthys olivaceus). Int. J. Mol. Sci. 2023, 24(3), 2480. [Google Scholar] [CrossRef]
- Liu, H.; Wang, Y.; Guo, Q.; Tian, X.; Ma, X.; Zhang, M.; Shi, X.; Ma, W.; Zhang, J.; Xu, R.; Li, X.; Kong, X. Identification of Amh and its potential signaling pathway involved in the ovary development in the largemouth bass (Micropterus salmoides). J. Steroid Biochem. Mol. Biol. 2025, 253, 106811. [Google Scholar] [CrossRef]
- Anitha, A.; Senthilkumaran, B. Role of sox family genes in teleostean reproduction-an overview. Reprod. Breed. 2021, 1(1), 22–31. [Google Scholar] [CrossRef]
- Bowles, J.; Schepers, G.; Koopman, P. Phylogeny of the SOX family of developmental transcription factors based on sequence and structural indicators. Dev. Biol. 2000, 227, 239–255. [Google Scholar] [CrossRef]
- Patra, S.K.; Chakrapani, V.; Panda, R.P.; Mohapatra, C.; Jayasankar, P.; Barman, H.K. First evidence of molecular characterization of rohu carp Sox2 gene being expressed in proliferating spermatogonial cells. Theriogenology 2015, 84, 268–276. [Google Scholar] [CrossRef] [PubMed]
- Shang, M.; Su, B.; Lipke, E.A.; Perera, D.A.; Li, C.; Qin, Z.; Li, Y.; Dunn, D.A.; Cek, S.; Peatman, E.; Dunham, R.A. Spermatogonial stem cells specific marker identification in channel catfish Ictalurus punctatus and blue catfish I. furcatus. Fish Physiol. Biochem. 2015, 41, 1545–1556. [Google Scholar] [CrossRef]
- Yu, H.; Du, X.; Li, X.; Qu, J.; Zhu, H.; Zhang, Q.; Wang, X. Genome-wide identification and transcriptome-based expression analysis of sox gene family in the Japanese flounder Paralichthys olivaceus. J. Oceanol. Limnol. 2018, 36, 1731–1745. [Google Scholar] [CrossRef]
- Anitha, A.; Senthilkumaran, B. sox19 regulates ovarian steroidogenesis in common carp. J. Steroid. Biochem. Mol. Biol. 2022, 217, 106044. [Google Scholar] [CrossRef] [PubMed]
- Navarro-Martín, L.; Galay-Burgos, M.; Piferrer, F.; Sweeney, G. Characterisation and expression during sex differentiation of Sox19 from the sea bass Dicentrarchus labrax. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 2012, 163, 316–323. [Google Scholar] [CrossRef]
- Hu, Y.; Wang, B.; Du, H. A review on sox genes in fish. Rev. Aquac. 2021, 13, 1986–2003. [Google Scholar] [CrossRef]
- Taboada, X.; Viñas, A.; Adrio, F. Comparative expression patterns of Sox2 and Sox19 genes in the forebrain of developing and adult turbot (Scophthalmus maximus). J. Comp. Neurol. 2018, 526(5), 899–919. [Google Scholar] [CrossRef]
- Wei, L.; Yang, C.; Tao, T.; Wang, D. Genome-wide identification and transcriptome based expression profiling of the sox gene family in the Nile tilapia (Oreochromis niloticus). Int. J. Mol. Sci. 2016, 17, 270. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Chen, X.; Wang, M.; Zhang, W.; Pan, J.; Qin, Q.; Zhong, L.; Shao, J.; Sun, M.; Jiang, H.; Bian, W. Genome-wide identification, phylogeny and expressional profile of the sox gene family in channel catfish (Ictalurus punctatus). Comp. Biochem. Physiol. Genom. Proteonomics 2018, 28, 17–26. [Google Scholar] [CrossRef]
- Wang, L.; Cheng, H.; Xia, L.; Guo, Y.; Huang, X.; Zhou, R. Molecular cloning and expression of sox17 in gonads during sex reversal in the rice field eel, a teleost fish with a characteristic of natural sex transformation. Biochem. Biophys. Res. Commun. 2003, 303, 452–457. [Google Scholar] [CrossRef]
- Navarro-Martín, L.; Galay-Burgos, M.; Sweeney, G.; Piferrer, F. Different Sox17 transcripts during sex differentiation in sea bass Dicentrarchus labrax. Mol. Cell Endocrinol. 2009, 299, 240–251. [Google Scholar] [CrossRef]
- Cui, J.; Shen, X.; Zhao, H.; Nagahama, Y. Genome-wide analysis of Sox genes in medaka (Oryzias latipes) and their expression pattern in embryonic development. Cytogenet. Genome Res. 2011, 134, 283–294. [Google Scholar] [CrossRef] [PubMed]
- Liang, T.; Jia, Y.; Zhang, R.; Du, Q.; Chang, Z. Identification, molecular characterization and analysis of the expression pattern of SoxF subgroup genes the Yellow River carp, Cyprinus carpio. J. Genet. 2018, 97(1), 157–172. [Google Scholar] [CrossRef] [PubMed]
Figure 1.
Cross sections of ovaries of juveniles and adult turbots stained with hematoxylin-eosin. (a-h) In juveniles, abundant oogonia (arrowheads) and primary oocytes (arrows) were observed (details in c-d and g-h). (i-l) In adults, numerous and larger primary oocytes were observed (arrows, details in k and l). TA, tunica albuginea; OS, ovarian stroma; stars, ovarian cavity; asterisks, celomic cavity. Squares in low magnification images (a-b; e-f; i) indicate the location of details (c-d; g-h; j). Scale bars: 500 µm (a,e,i); 100 µm (b,f,j); 20 µm (c-d, g-h, k-l).
Figure 1.
Cross sections of ovaries of juveniles and adult turbots stained with hematoxylin-eosin. (a-h) In juveniles, abundant oogonia (arrowheads) and primary oocytes (arrows) were observed (details in c-d and g-h). (i-l) In adults, numerous and larger primary oocytes were observed (arrows, details in k and l). TA, tunica albuginea; OS, ovarian stroma; stars, ovarian cavity; asterisks, celomic cavity. Squares in low magnification images (a-b; e-f; i) indicate the location of details (c-d; g-h; j). Scale bars: 500 µm (a,e,i); 100 µm (b,f,j); 20 µm (c-d, g-h, k-l).
Figure 2.
Cross sections of testis of juveniles and adult turbots stained with hematoxylin-eosin. (a-g) In juveniles, abundant spermatogonia (arrowheads) were observed. (h-l) In adults, spermatogonia (i: Sg, j: arrowheads), spermatocytes (i: Sc, k: arrowhead), and spermatids (i: St, l: arrowheads) were observed. IT, interstitial tissue; TA, tunica albuginea; stars, lumen of the efferent ducts; asterisks, celomic cavity. Squares in low magnification images (d-e) indicate the location of details (f-g). Scale bars: 500 µm (a, d, h); 200 µm (i); 100 µm (b, e); 25 µm; (c, f-g); 20 µm (j-l).
Figure 2.
Cross sections of testis of juveniles and adult turbots stained with hematoxylin-eosin. (a-g) In juveniles, abundant spermatogonia (arrowheads) were observed. (h-l) In adults, spermatogonia (i: Sg, j: arrowheads), spermatocytes (i: Sc, k: arrowhead), and spermatids (i: St, l: arrowheads) were observed. IT, interstitial tissue; TA, tunica albuginea; stars, lumen of the efferent ducts; asterisks, celomic cavity. Squares in low magnification images (d-e) indicate the location of details (f-g). Scale bars: 500 µm (a, d, h); 200 µm (i); 100 µm (b, e); 25 µm; (c, f-g); 20 µm (j-l).
Figure 3.
Vasa expression in cross sections of the turbot gonads. (a-c) Vasa expression in oogonia (arrowheads) and primary oocytes (arrows) in juveniles (a-b) and adults (c). (d-f) Vasa expression in spermatogonia (Sg) in juveniles (d-e) and in spermatogonia and spermatocytes (Sc) in adults (f). OS, ovarian stroma; stars, ovarian cavity; asterisks, celomic cavity. Scale bars: 100 µm (a-e); 200 µm (f).
Figure 3.
Vasa expression in cross sections of the turbot gonads. (a-c) Vasa expression in oogonia (arrowheads) and primary oocytes (arrows) in juveniles (a-b) and adults (c). (d-f) Vasa expression in spermatogonia (Sg) in juveniles (d-e) and in spermatogonia and spermatocytes (Sc) in adults (f). OS, ovarian stroma; stars, ovarian cavity; asterisks, celomic cavity. Scale bars: 100 µm (a-e); 200 µm (f).
Figure 4.
Cyp19a1a (a-f) and Foxl2 (g-l) expression in cross sections of the turbot gonads. (a-c) Cyp19a1a expression in oogonia (arrowheads) and primary oocytes (arrows) in juveniles (a-b) and adults (c). (d-f) No Cyp19a1a expression was observed at any developmental stage in males. (g-l) Foxl2 expression in oogonia (arrowheads) and primary oocytes (arrows) in juveniles (g-h) and adults (i). (j-l) Foxl2 expression in spermatogonia (Sg) in juveniles (j-k) and in Sg, spermatocytes (Sc) and spermatids (St) in adults (l). OS, ovarian stroma; stars, ovarian cavity; asterisks, celomic cavity. Scale bars: 100 µm (a-d, g-k); 200 µm (e-f, l).
Figure 4.
Cyp19a1a (a-f) and Foxl2 (g-l) expression in cross sections of the turbot gonads. (a-c) Cyp19a1a expression in oogonia (arrowheads) and primary oocytes (arrows) in juveniles (a-b) and adults (c). (d-f) No Cyp19a1a expression was observed at any developmental stage in males. (g-l) Foxl2 expression in oogonia (arrowheads) and primary oocytes (arrows) in juveniles (g-h) and adults (i). (j-l) Foxl2 expression in spermatogonia (Sg) in juveniles (j-k) and in Sg, spermatocytes (Sc) and spermatids (St) in adults (l). OS, ovarian stroma; stars, ovarian cavity; asterisks, celomic cavity. Scale bars: 100 µm (a-d, g-k); 200 µm (e-f, l).
Figure 5.
Amh expression in cross sections of the turbot gonads. (a-c) Amh expression in oogonia (arrowheads) and the ooplasm of the primary oocytes (arrows) in juveniles (a-b) and adults (c). (d-f) Amh expression in spermatogonia (Sg) in juveniles (d-e), and in Sg and spermatocytes (Sc) in adults (f). OS, ovarian stroma; stars, ovarian cavity; asterisks, celomic cavity. Scale bars: 100 µm (a-e); 200 µm (f).
Figure 5.
Amh expression in cross sections of the turbot gonads. (a-c) Amh expression in oogonia (arrowheads) and the ooplasm of the primary oocytes (arrows) in juveniles (a-b) and adults (c). (d-f) Amh expression in spermatogonia (Sg) in juveniles (d-e), and in Sg and spermatocytes (Sc) in adults (f). OS, ovarian stroma; stars, ovarian cavity; asterisks, celomic cavity. Scale bars: 100 µm (a-e); 200 µm (f).
Figure 6.
Sox2 (a-f) and Sox19 (g-l) expression in cross sections of the turbot gonads. (a-c) Sox2 expression in oogonia (arrowheads) and primary oocytes (arrows) in juveniles (a-b) and adults (c). (d-f) In the testis, Sox2 expression was observed in spermatogonia (Sg) in juveniles (d-e) and Sg and spermatocytes (Sc) in adults (f). (g-i) Sox19 expression in oogonia (arrowheads) and primary oocytes (arrows) in juveniles (g-h) and adults (i). (j-l) Weak Sox19 expression in spermatogonia of juveniles (j-k) and adults (l). OS, ovarian stroma; stars, ovarian cavity; asterisks, celomic cavity. Scale bars: 100 µm (a-e, g-k); 200 µm (f, l).
Figure 6.
Sox2 (a-f) and Sox19 (g-l) expression in cross sections of the turbot gonads. (a-c) Sox2 expression in oogonia (arrowheads) and primary oocytes (arrows) in juveniles (a-b) and adults (c). (d-f) In the testis, Sox2 expression was observed in spermatogonia (Sg) in juveniles (d-e) and Sg and spermatocytes (Sc) in adults (f). (g-i) Sox19 expression in oogonia (arrowheads) and primary oocytes (arrows) in juveniles (g-h) and adults (i). (j-l) Weak Sox19 expression in spermatogonia of juveniles (j-k) and adults (l). OS, ovarian stroma; stars, ovarian cavity; asterisks, celomic cavity. Scale bars: 100 µm (a-e, g-k); 200 µm (f, l).
Figure 7.
Sox17 expression in cross sections of the turbot gonads. (a-c) Sox17 expression in oogonia (arrowheads) and primary oocytes (arrows) in juveniles (a-b) and adults (c). (d-f) Sox17 expression was observed in spermatogonia (Sg) in juveniles (d-e) and adults (f). OS, ovarian stroma; stars, ovarian cavity; asterisks, celomic cavity. Scale bars: 100 µm (a-e); 200 µm (f).
Figure 7.
Sox17 expression in cross sections of the turbot gonads. (a-c) Sox17 expression in oogonia (arrowheads) and primary oocytes (arrows) in juveniles (a-b) and adults (c). (d-f) Sox17 expression was observed in spermatogonia (Sg) in juveniles (d-e) and adults (f). OS, ovarian stroma; stars, ovarian cavity; asterisks, celomic cavity. Scale bars: 100 µm (a-e); 200 µm (f).
Table 1.
Primers used for cDNA amplification in riboprobe synthesis.
Table 1.
Primers used for cDNA amplification in riboprobe synthesis.
| GENE |
PRIMERS |
AMPLICON SIZE (pb) |
| Vasa |
F: GAGCCTGAAGCCATCATTGT
R: ACGTTCTCCTGGCACATCAACG |
634 |
| Amh |
F: CTGGGCGTATCCTCTAACGA
R: CATCCGCACTCCTTTGCTAT |
693 |
| Cyp19a |
F: GGAGGTTTGTGTCTCCTCCA
R: GAACTCCGTGCGGTGCATGC |
715 |
| Foxl2 |
F: AGAAGCGGCTCACGCTGTC
R: TTTGGTCTCGTGCTCCCAGT |
712 |
| Sox2 |
F: CTGTTGGACTACTTAACGTTGGA
R: CAGATGAAAAGTGGGAGACG |
729 |
| Sox17 |
F: GAGCAGATGCACCACTCTGA
R: GCAAAAAGTATAAAAACACCGTCA |
539 |
| Sox19 |
F: TGTGCCTTGCCTTTTGATTT
R: GGTTTGCTGGAGATTTAGTCTGA |
600 |
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).