3. Learnings from an Extreme Standard Curve
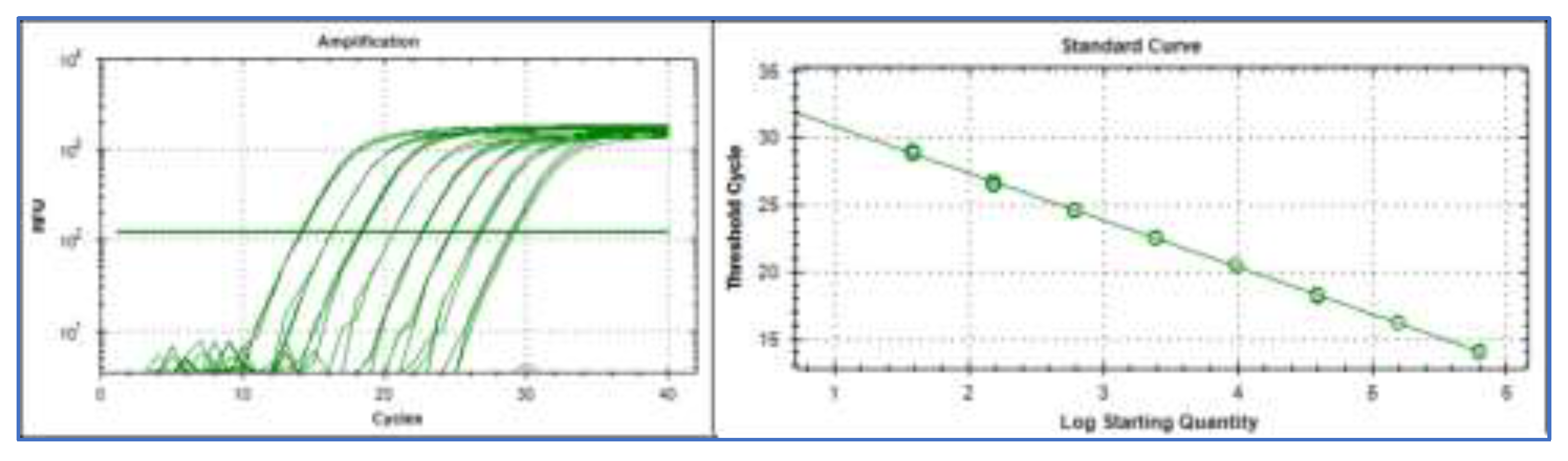
We begin by examining an extreme standard curve constructed using a very large number of replicates per concentration and small concentration increments. This example is included as a deliberately idealised, illustrative case to highlight fundamental properties of qPCR behavior that are not readily visible in routine standard curves. It represents a validation experiment for the Reference Sequence (
Figure 1) [
7] performed using the IntelliQube instrument that uses very small reaction volumes, enabling high throughput [
8]. The example data with tutorials are available [
10].
3.1. Learnings from an Extreme Standard Curve
The Reference Sequences were originally proposed as a cost-efficient strategy to correct for background genomic DNA (gDNA) in gene expression analysis.
7 The Reference Sequences are species-specific, and each targets a conserved, non-transcribed genomic sequence present in exactly one copy per haploid genome. This allows for accurate quantification of the gDNA background in complementary DNA (cDNA) samples [
7].
Standard samples were prepared from a stock solution of purified synthetic DNA containing a single Reference Sequence per double-stranded molecule. The concentration of the stock solution was determined using dPCR with the reference sequence assay. Solutions covering the range of 2048 down to 1 target molecule on average per reaction were prepared using 2-fold dilution steps. At the lowest concentration, and for the non-template control, 128 replicates were assessed. Higher concentrations were replicated 64 times and the highest 32 times.
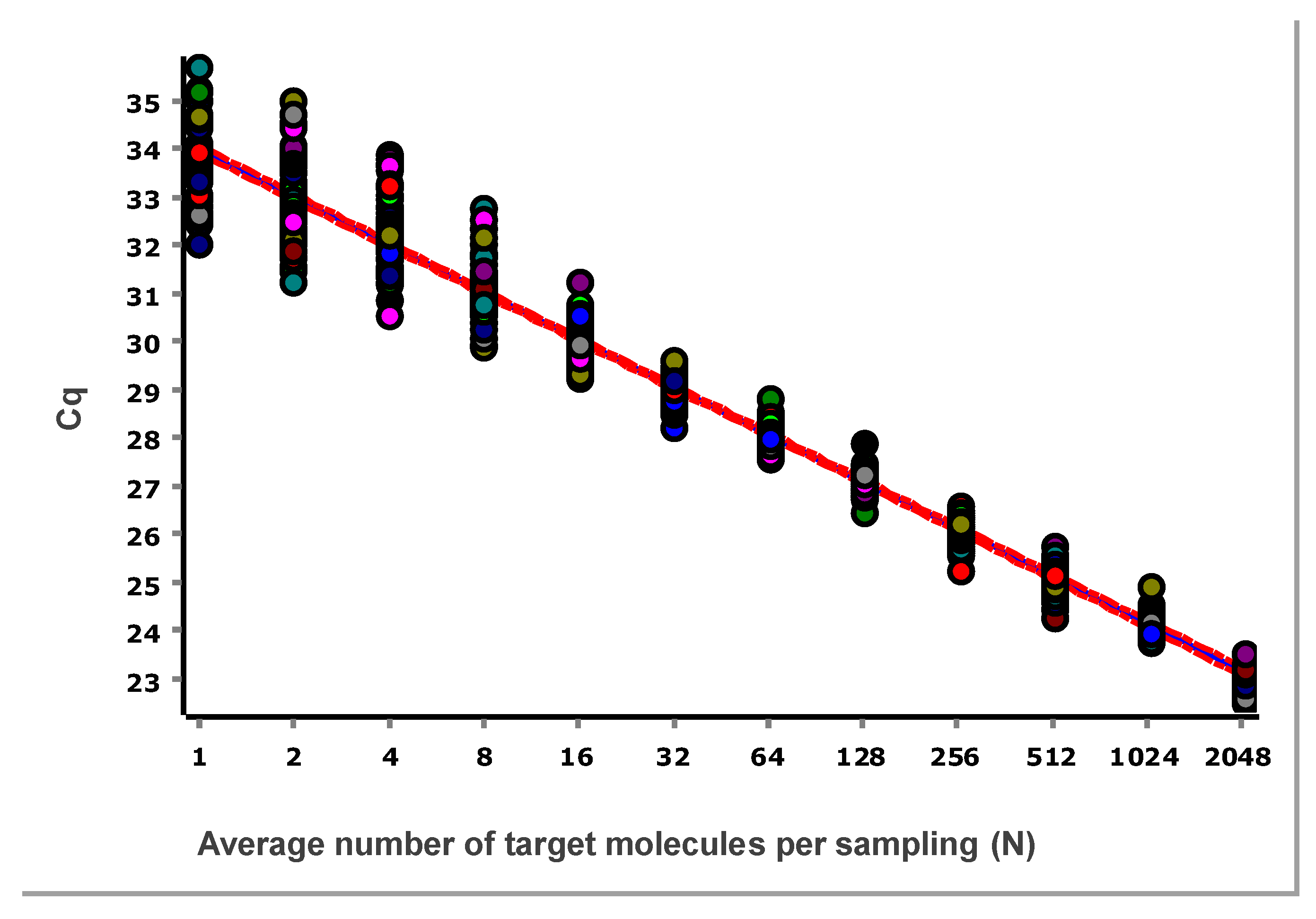
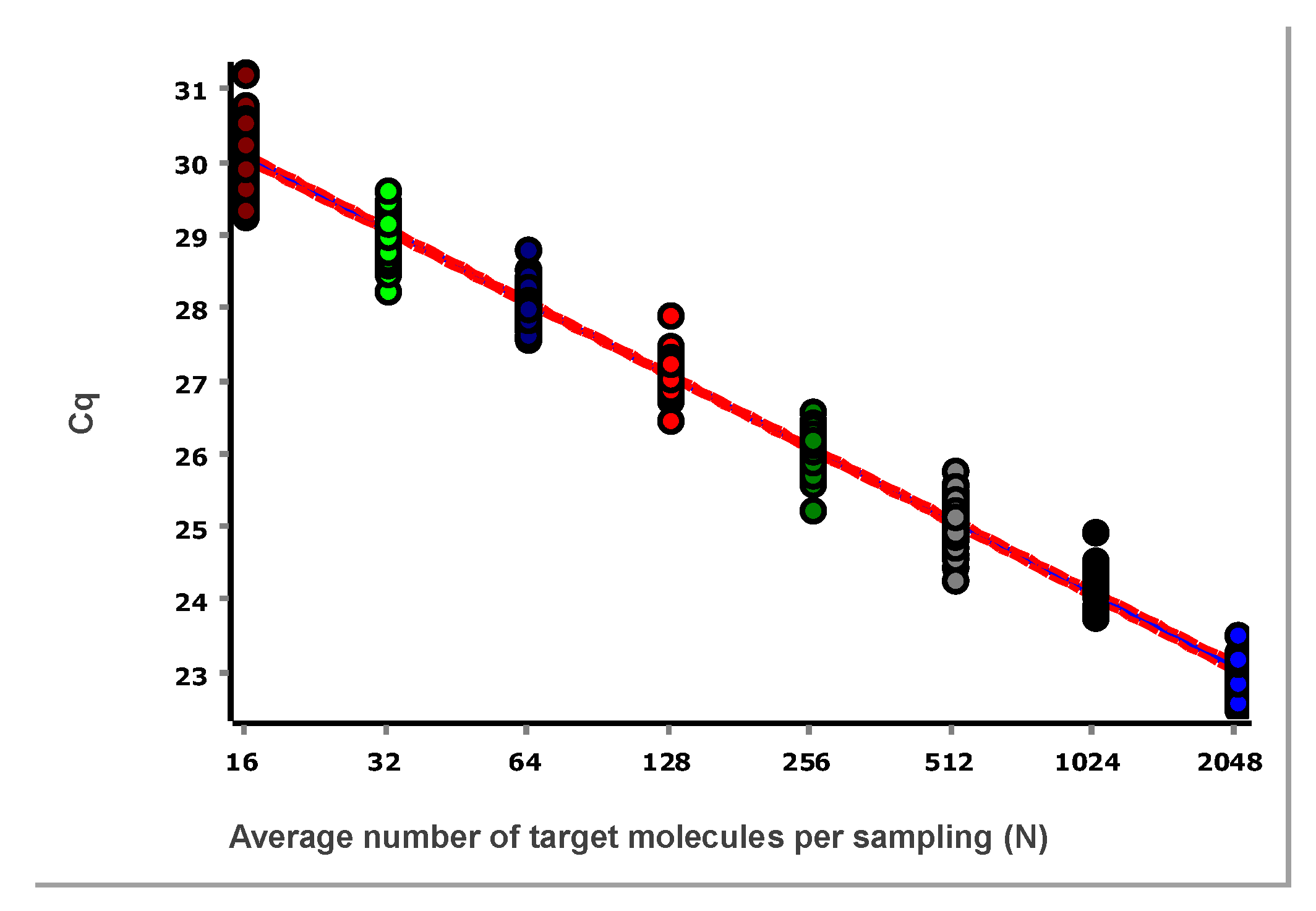
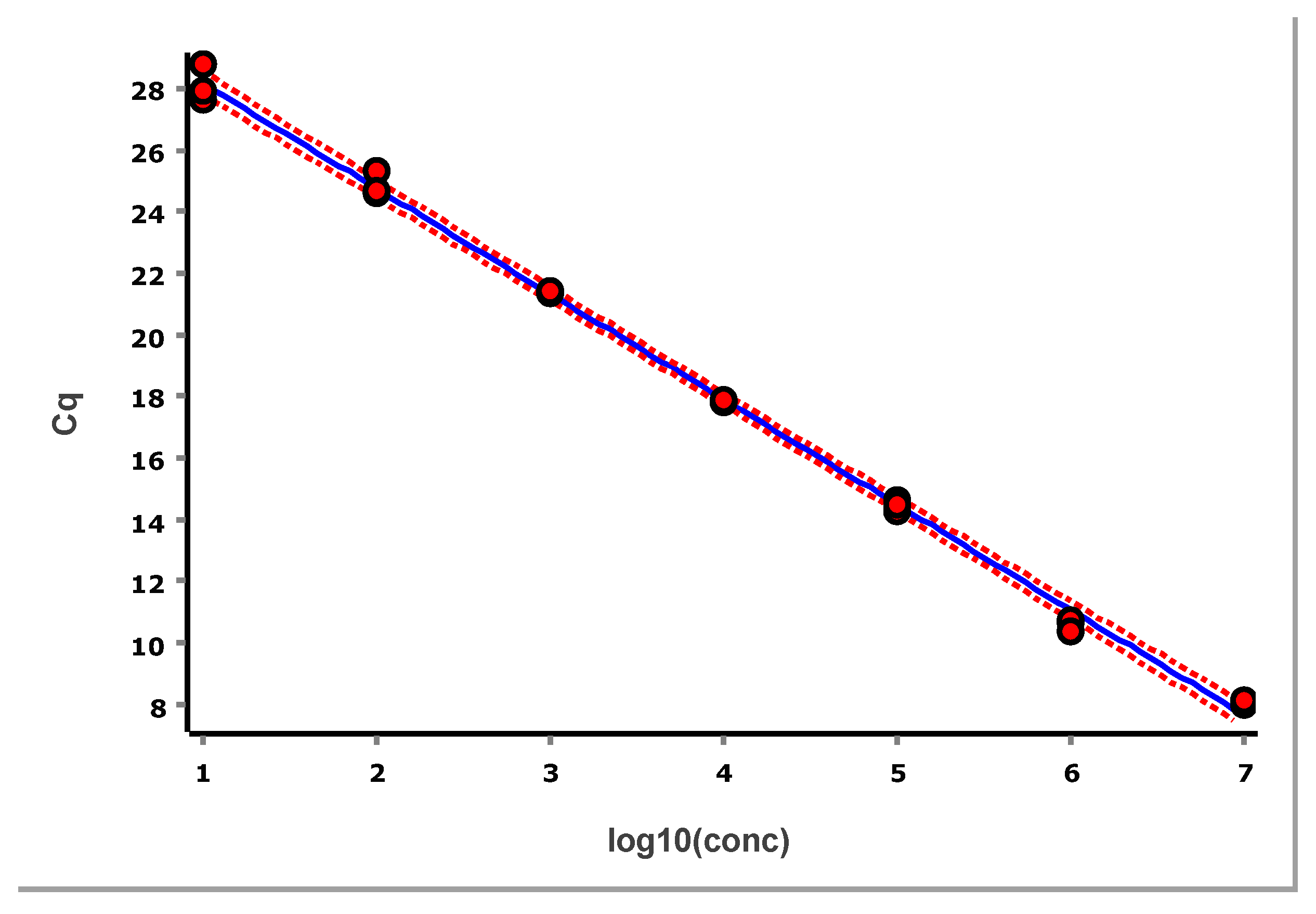
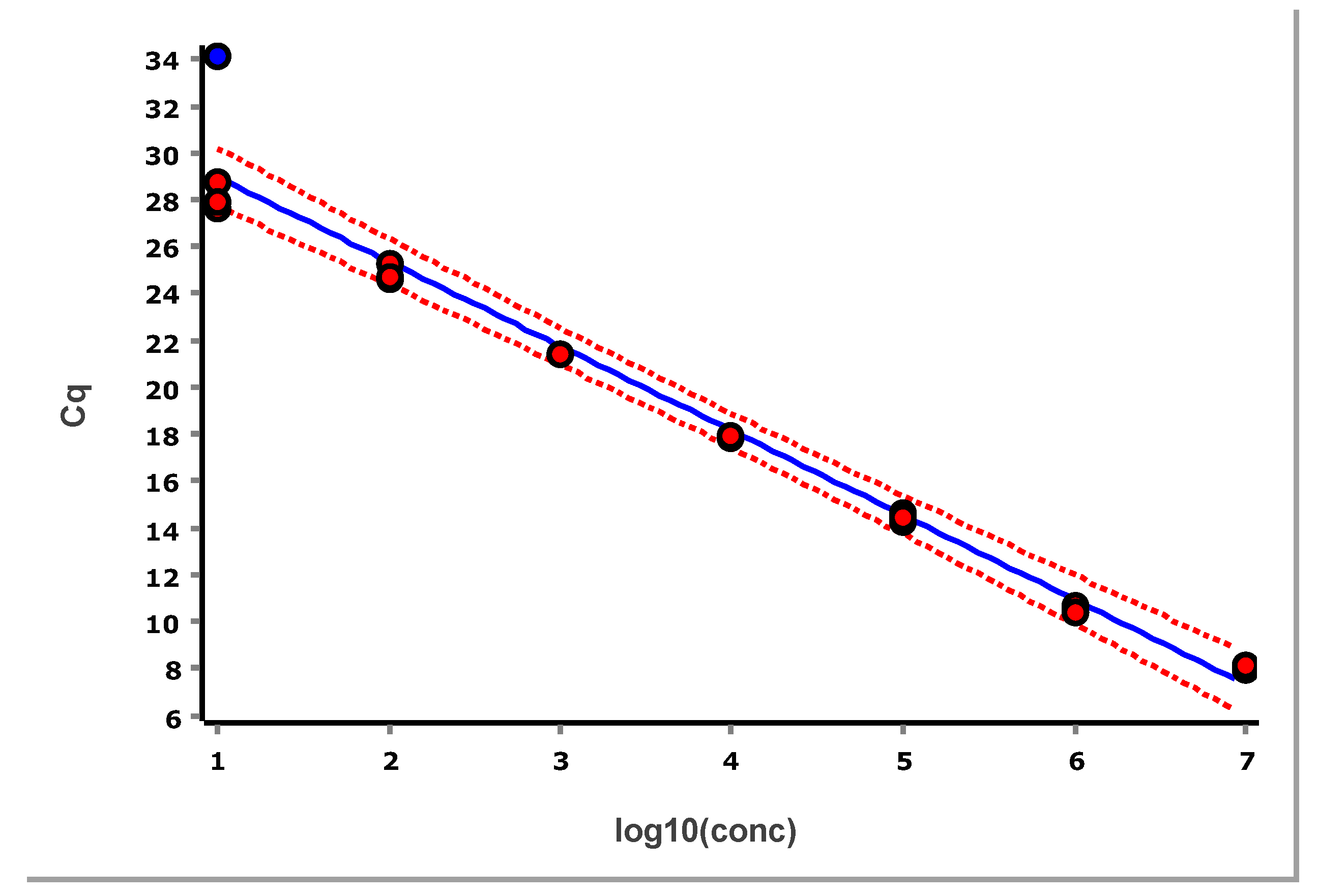
The data, after removing some obviously technically failed reactions, fitted to a straight line using linear regression (ordinary least squares fit, OLS), are shown in
Figure 3.
3.2. Residual Plot
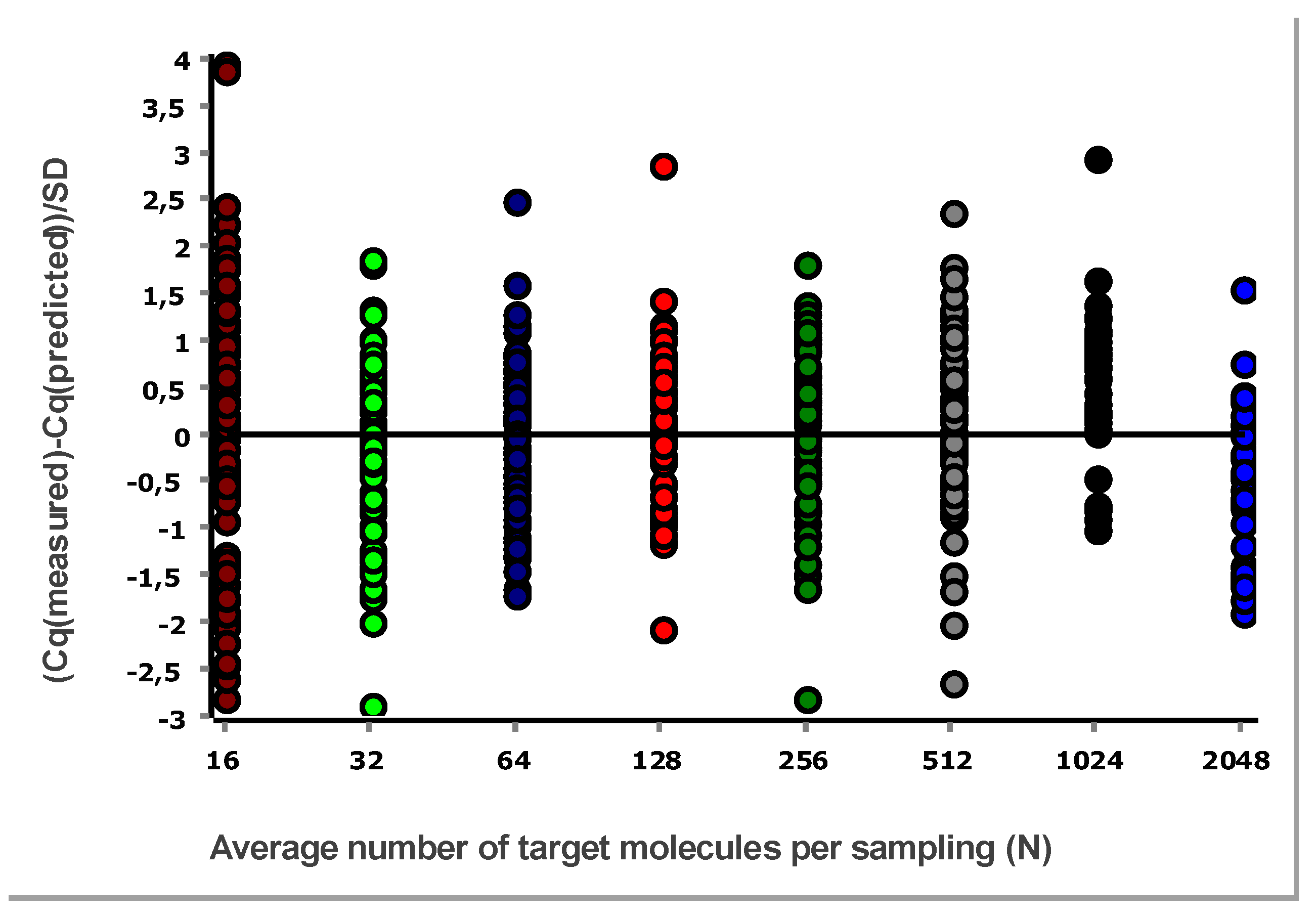
A general good practice is to study the difference between Cq values and those predicted by the linear regression, i.e., the residual. This is done using a residuals plot. Either the combined data at all levels (different concentrations) are considered, as in
Figure 4, or per each level separately [
11].
From these illustrative data with the very large number of replicates, we see in
Figure 3 and
Figure 4 that at lower N, the variation, which can be quantified as the standard deviation (SD), becomes higher. This violates one of the assumptions of the OLS criterion. Either weighted least squares should be applied, or, as we will do below, only samples above the lower limit of quantification (LLOQ) will be considered.
3.3. Relative Standard Deviation and Limit of Quantification
The variation across replicates can be quantified by means of the SD, where SD can be calculated either for the measured Cq-values or for concentrations derived from the Cq-values. These SDs are very different because of the difference in scale. While concentrations are in a linear scale, Cq-values are in a log scale. For comparison with other bioanalytical methods, performance parameters are preferably presented in linear scale.
SD, like the mean, increases with the expected number of target molecules per sample. For comparison across concentrations, the relative standard deviation (RSD), also referred to as the coefficient of variation, is the most widely used measure of imprecision. In a linear scale, RSD is obtained from the SD of the Cq values as depicted in Equation 2 (where E is the PCR efficiency):
8
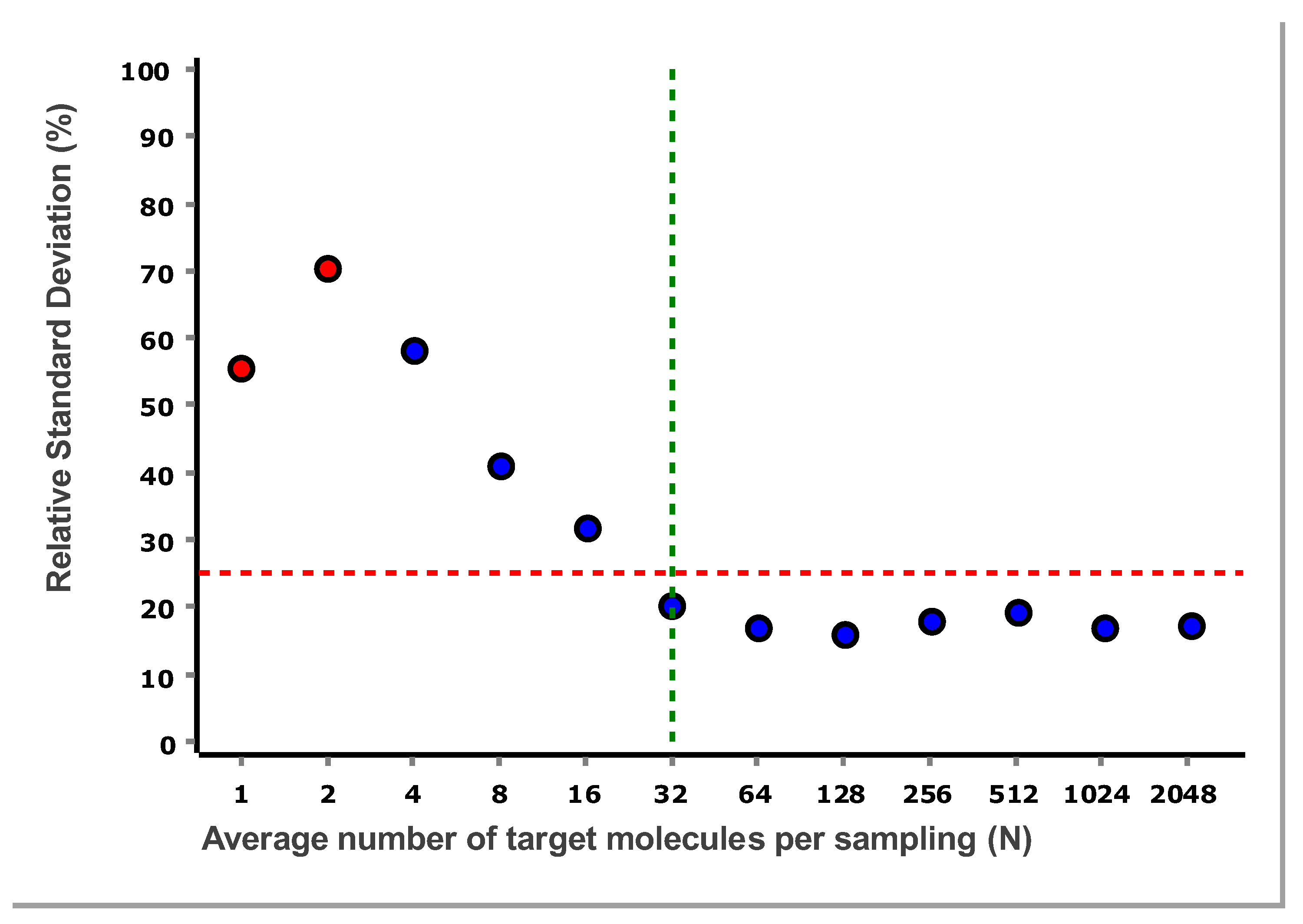
The RSD for the data in
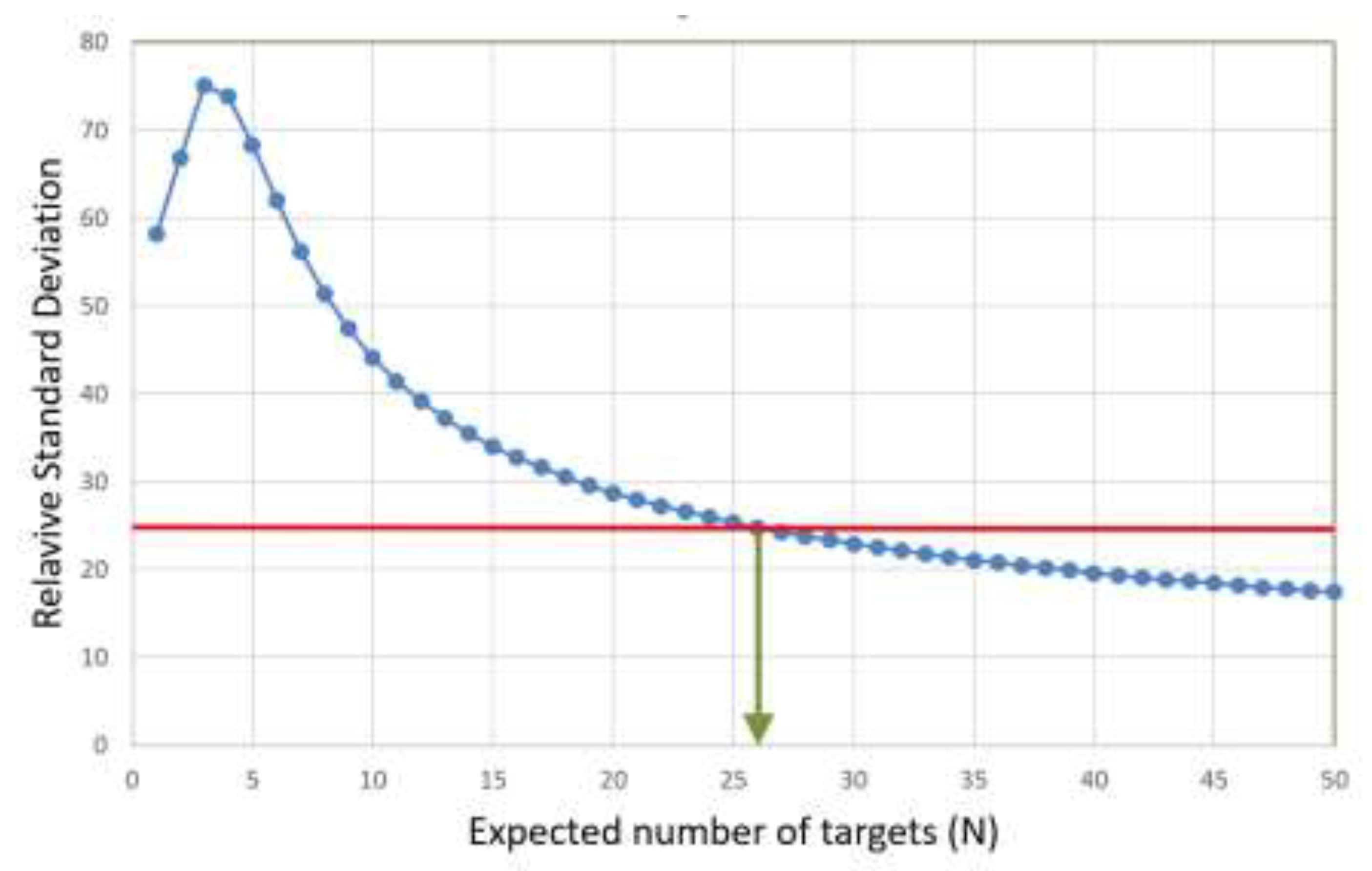
Figure 4, is plotted as a function of the number of target molecules per sample in
Figure 5.
The RSD levels off at 32 molecules and stabilizes, which is coherent with what can be seen in
Figure 4. At these concentrations, various sources of imprecision, such as sample handling, pipetting, and measurement error, become significant. Although these data were generated with the rather unique IntelliQube instrument, repeatability is in line with previous reports using standard qPCR instrumentation [
12].
For bioanalytical methods, we need to estimate important performance parameters. The two most important are the limit of detection (LOD) and LOQ. LOD is the lowest analyte concentration likely to be reliably distinguished from the blank and at which detection is feasible, while LOQ is the limiting concentration at which the analyte cannot only be reliably detected but at which some predefined goals for bias and imprecision are met [
13]. The LOQ may be equivalent to the LOD but is usually higher.
For qPCR, there is no single universal recommendation for LOQ. It depends on the assay, sample type, matrix, and purpose. But to be in line with other bioanalytical methods, a maximum RSD of 25 % in linear scale is commonly used. Sometimes the RSD increases towards very low and also towards very high concentrations. In those cases, we refer to the lower limit of quantification (LLOQ) and the upper limit of quantification (ULOQ). The LLOQ is then the lowest concentration where RSD ≤ 25 %, and ULOQ is the highest concentration where RSD ≤ 25 %.
In
Figure 5 we see that 32 target molecules and higher per sample meet this criterion. Thus, the LLOQ of the reference assay, when analyzing purified gDNA standard using the IntelliQube qPCR instrument, is 32 molecules. There is no ULOQ within the concentration range studied.
3.4. Sampling Uncertainty
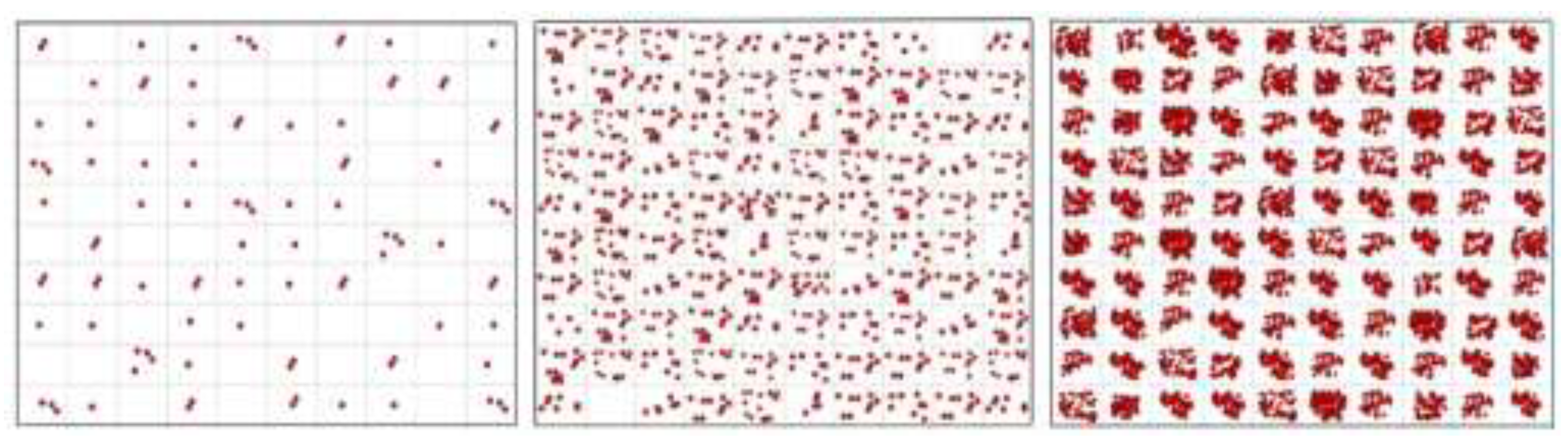
PCR, using an optimized assay, can amplify a single target molecule as routinely demonstrated in dPCR. If we take a cell that contains a single DNA and place it into a qPCR tube so we are sure it is there, add PCR reagents and an optimized assay for a target sequence in the DNA, we will detect it.
For most samples, however, we don’t control the number of target molecules. Consider a 1 mL homogeneous sample containing 1,000 lysed cells. From this solution, transfer 1 L, which is 1/1000th of the volume, to a tube for analysis with qPCR targeting the DNA. The sampling can, of course, contain one DNA molecule. However, it may also contain two, perhaps three, or even more DNA molecules. These cases will produce a product, and the PCR will be positive. But the sampling (pipetted sample aliquot) may contain no DNA, in which case the PCR will be negative. Since the original sample did contain DNA, this is a false negative.
Stochastic variation in the number of targets across sampling replicates gives rise to sampling uncertainty. Sampling uncertainty becomes important when few targets are expected (small N).
The probability that a sampling contains x target molecules,
, is given by the Poisson distribution shown in Equation 3:
where N is the expectation value, which is equivalent to the average number of target molecules per sampled volume, x is the actual number of targets in a particular sampling, and ! indicates the factorial of the number (x! = 1 × 2 × 3 × … × x).
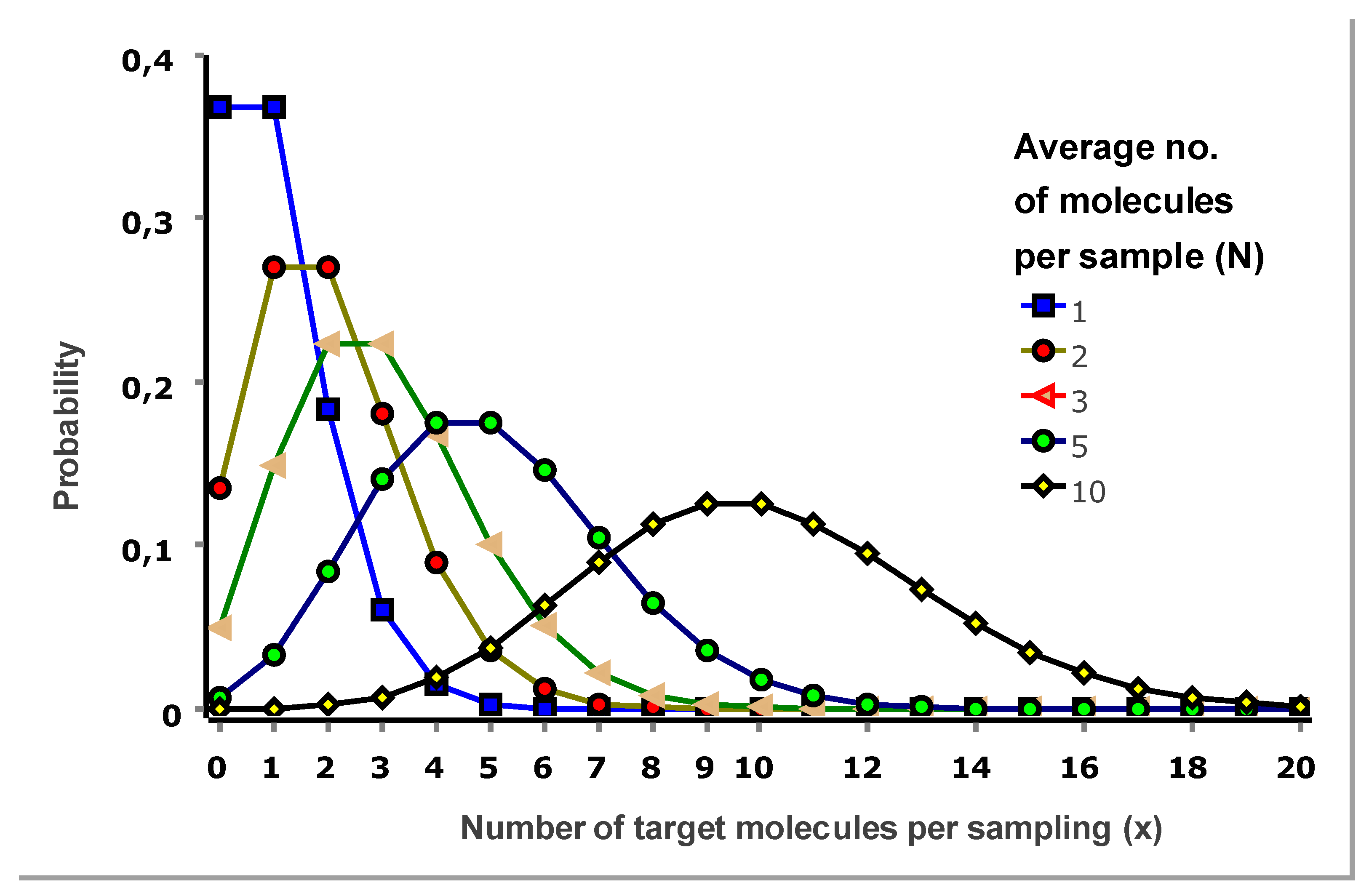
Figure 6 shows Poisson distributions for selected expectation values.
Let us inspect the Poisson distributions in
Figure 6. For an expectation value of one molecule per sampling (N=1), the probability that a sampling contains one molecule (x=1) is 37 %. Probability is 18 %, it contains two target molecules (x=2); 6 % it contains three (x=3), and 1.5 % it contains four (x=4). There is also a relevant 37 % probability that a sampling is negative (x=0).
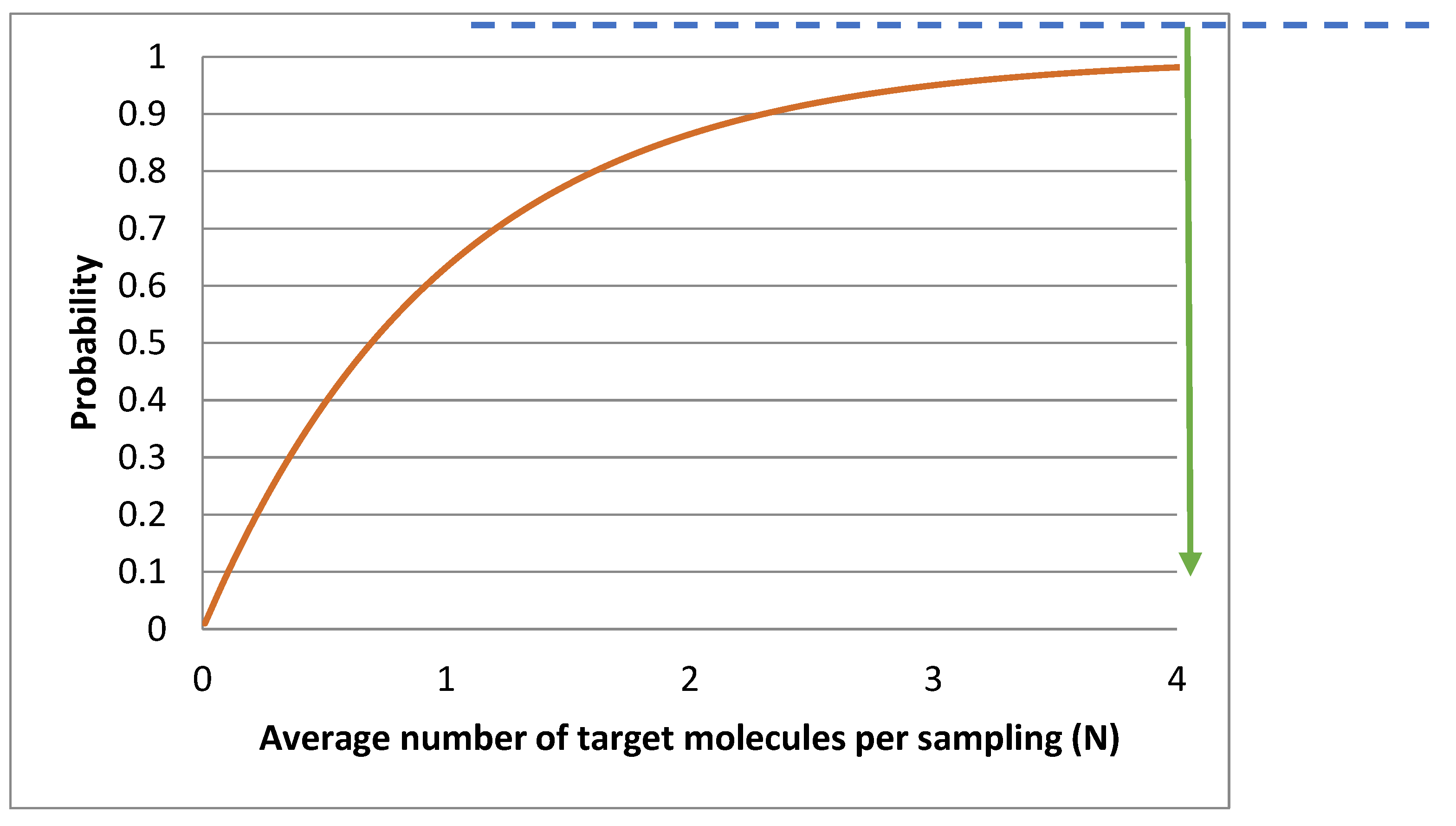
Figure 7 shows the probability that a sampling is positive P(x>0), as a function of the average number of target molecules per sampling (N).
3.5. Limit of Detection (LOD)
When analyzing test samples, we want to be reasonably confident that we obtain a positive test result when the sample is expected to contain even very few target molecules per analyzed volume (N). The challenge here is often to decide when the signal obtained cannot be statistically confounded with the signal from a negative sample, i.e., matrix. While buffer only shall not produce a positive PCR, a negative sample may contain similar nucleic acids that amplify. Such a background signal must then be taken into account, limiting the LOD.
LOD can be limited by many factors related to the sampling and processing of the samples, instrumental noise, quality of reagents, assay performance, etc. Under ideal conditions, when limited by the sampling uncertainty, LOD can be predicted. Working at a 95 % confidence level means at least 95 % of replicate samplings shall be qPCR positive when the sample contains targets. In
Figure 7 the dashed line indicates that an average of 3 molecules per analyzed volume (N) is required for a sampling to be positive in 95 % of cases. Hence, the theoretical LOD is N=3 molecules when sampling uncertainty dominates. This theoretical limit is independent of the analytical method.
In the standard curve in
Figure 3 and its corresponding residuals plot in
Figure 4 we see the spread of replicates increases towards lower concentrations, which is due to increasing sampling uncertainty. The effect is visualized and quantified in
Figure 5.
The mean Cq of the replicates at the lowest concentrations (N≤2), where some samplings are negative, is lower than that predicted by the linear regression (
Figure 3,
Figure 4). This is because the average is calculated only for the positive samples that have Cq values, which introduces bias. Negative samplings are also the reason RSD is lower for N=1 than for N=2 (
Figure 5).
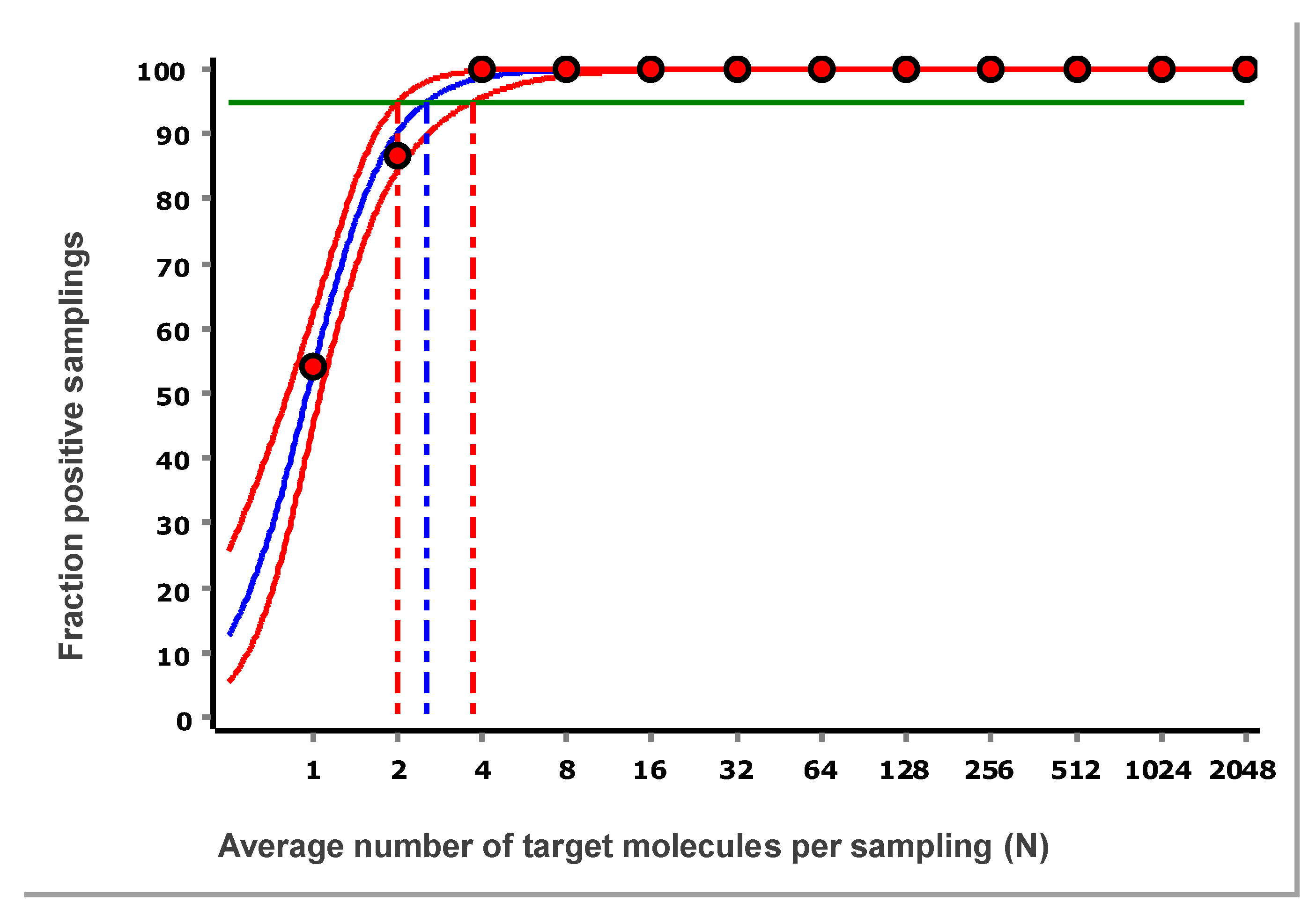
The fraction of samplings that were positive as a function of N for the data in
Figure 3 is shown in
Figure 8. The data are fitted with a sigmoidal function to interpolate the expectation value at 95 % probability for samplings to be positive. The confidence area of the fitted curve is also estimated, from which the confidence range of the LOD is obtained [
8].
For the reference assay, targeting purified synthetic DNA measured using the IntelliQube, the estimated LOD at 95 % confidence is 2.6 target molecules per sampling, with a 95 % confidence interval (CI) of 2.0–3.7 target molecules. The CI encompasses the theoretical limit of three molecules, and we conclude that the reference assay, when applied to purified synthetic targets, reaches this limit. Switching to dPCR will not improve the sensitivity, since it is limited by the sampling uncertainty. To achieve higher sensitivity, larger sampling volumes should be used, or the sample may be concentrated.
It should be noted that the approaches currently used to estimate LOD and LOQ are based on definitions from 1975 and 1980, respectively. These have issues. LOD can be calculated differently depending on the setup, particularly how the ‘blank’ is defined, and suffers from a high false negative rate (often around 50 %), and the LOQ threshold, here set at 25 %, is arbitrary. In 1993, IUPAC, ISO, and the European Union initiated harmonizing criteria and agreed on new LOD and LOQ definitions that are derived from the straight line fit and error propagation [
14,
15,
16]. This solves the issue of high false rates and the ambiguous LOQ [
17]. A general-purpose introduction is found elsewhere [
18]. When these new definitions will be introduced into PCR analytics remains to be seen.
3.6. Expected Imprecision
With the Poisson model (Equation 3), the expected imprecision in logarithmic scale due to sampling uncertainty expressed as either SD or RSD can be calculated using (Equation 2).
Figure 9 shows the predicted RSD as a function of the expected number of target molecules per sampling (N).
From
Figure 9 it follows that under conditions when other contributions than sampling uncertainty to imprecision are negligible, RSD reaches 25 %, which is a common criterion for LOQ, around N=26 target molecules per sampling. This is the theoretical LOQ at 25 % RSD.
For the reference assay in
Figure 5 RSD drops below 25 % at N = 32 target molecules, which is consistent with the theoretical expectations.
3.7. Variance PCR for Absolute Quantification
Note the resemblance of the theoretical curve in
Figure 9 with the experimentally determined RSD of the reference assay in
Figure 5. This suggests that when sampling uncertainty dominates variation across sampling aliquots, the expected number of target molecules per sampling can be predicted. The concept of this approach, which we term variance PCR (vPCR), is illustrated in
Figure 10. This is intended as a conceptual extension to illuminate the fundamental concept of the limit of quantification.
While the targets in the samplings are clearly observed to vary when the average is one target molecule per sampling, one hardly notices variation among samplings at N = 100 molecules per sampling. Although SD increases with N, the RSD decreases. This dependence was, precisely, presented in
Figure 9.
An apparent ambiguity is that the same RSD is obtained at two different expectation values (N), one at each side of the maximum RSD (
Figure 9). This, however, is not an issue. At low concentration, the drop in RSD is due to some replicates becoming negative, while at high concentration, all replicates are positive.
A comparison between experimentally measured RSD (
Figure 5) and theoretically predicted RSD assuming sampling uncertainty only (
Figure 9) is provided in
Table 1.
Given that measurements are performed on a logarithmic scale, the agreement between the predicted average number of targets per sampling derived from the measured RSD and the expected number of targets based on the dilutions is remarkably good. For all samples, the predicted values exceed the expected values somewhat, which may indicate that the expectation values based on dilutions of the stock solution, whose concentration was determined by dPCR, could be slightly underestimated.
At conditions where sampling uncertainty dominates, N can be determined directly from the SD (or RSD) of the Cq values of replicate samplings. This represents an alternative approach to absolute quantification that differs fundamentally from dPCR.
A potential application of vPCR is as a complement to dPCR, particularly on dPCR platforms capable of real-time fluorescence detection that generate Cq values. At low target loading, where a significant fraction of reaction partitions (which are equivalent to replicate samplings) are negative, standard dPCR readout provides optimal precision. At higher target loading, when most or all partitions are positive and classical dPCR loses resolving power, target concentration can instead be estimated by the vPCR principle based on the SD of the Cq values. Combining these approaches would dramatically expand the dynamic range of absolute quantification achievable with dPCR platforms.
3.8. Dynamic Range
The dynamic range of a qPCR method extends from LLOQ to ULOQ, defined as the concentration range over which the RSD remains below a specified threshold,
14 like 25% in our example. For the reference assay, RSD does not exceed 25% at high concentrations.
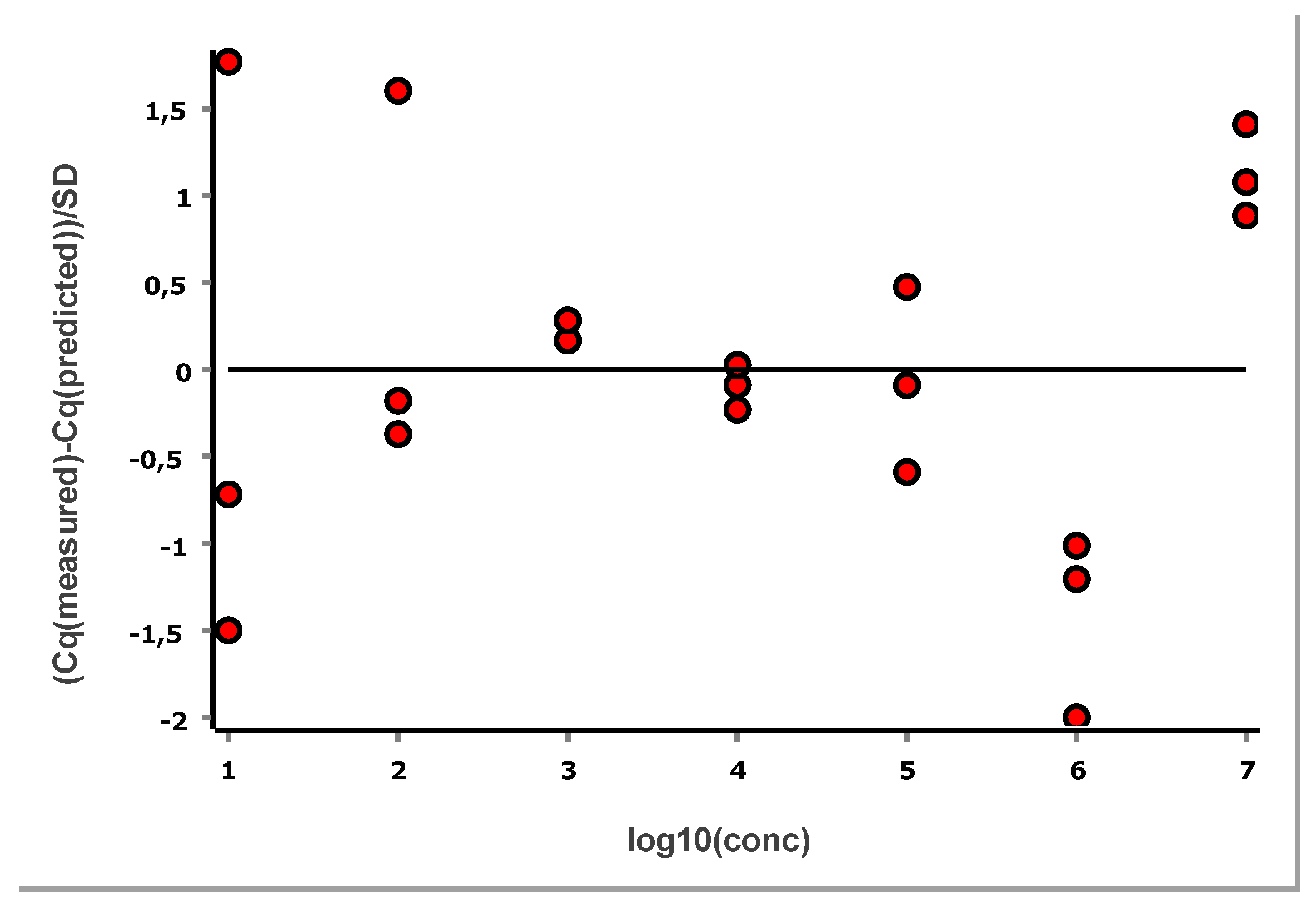
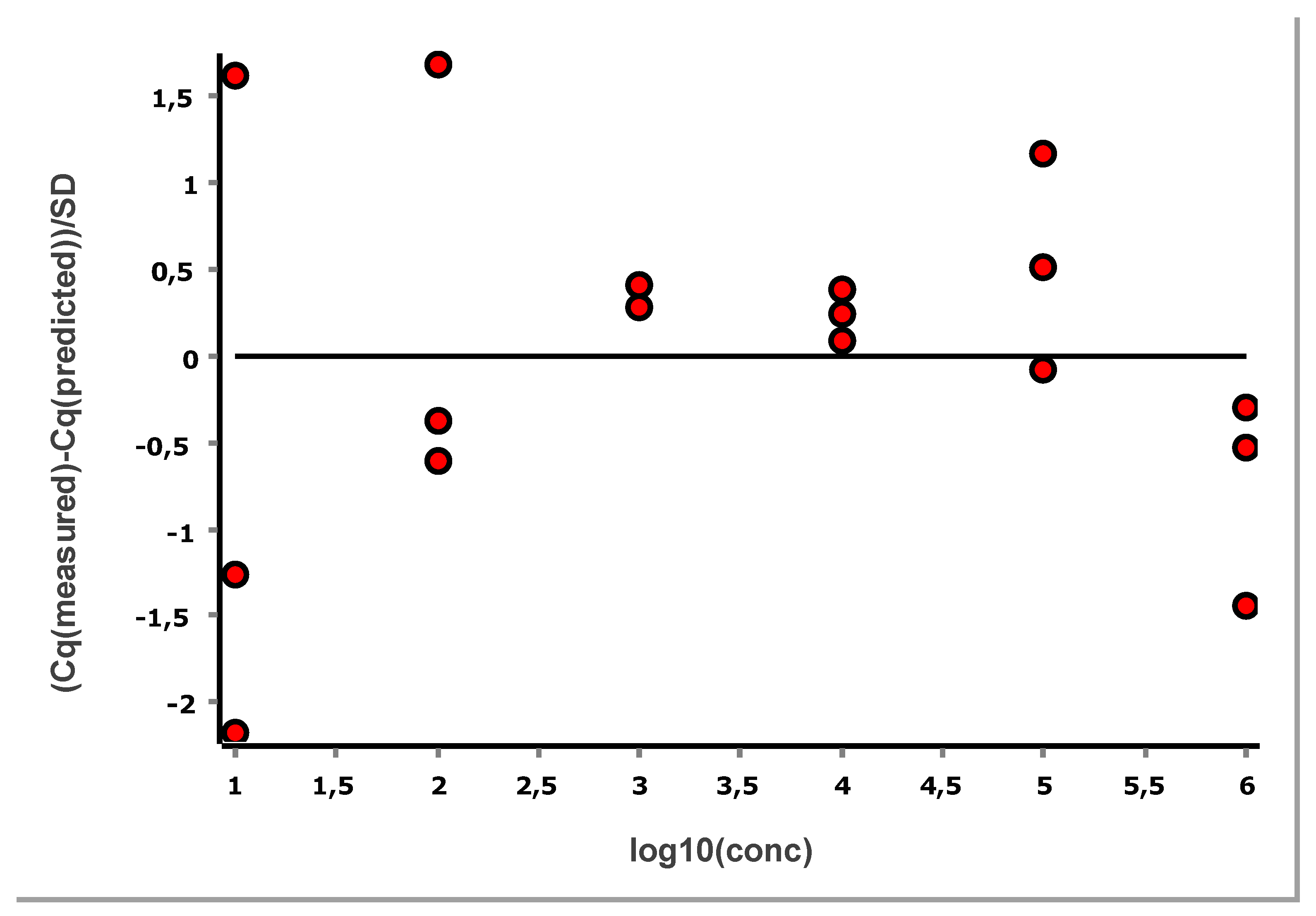
Figure 11 shows the standard curve above the LLOQ, and
Figure 12 shows the corresponding standardized residuals, i.e., residuals scaled by the SD, which is a suitable means for testing for the presence of outliers. Considering each level separately, data points outside ± 3 are considered to have an outlying behaviour. Alternatively, the traditional Grubbs’ test can be applied to the overall set of residuals, which assumes they follow a normal distribution as required also for the OLS [
19].
Within this range, replicate variability is effectively independent of concentration. This is referred to as homoscedasticity in statistics and is an assumption behind linear regression when based on the standard least squares criterion.
3.9. PCR Efficiency
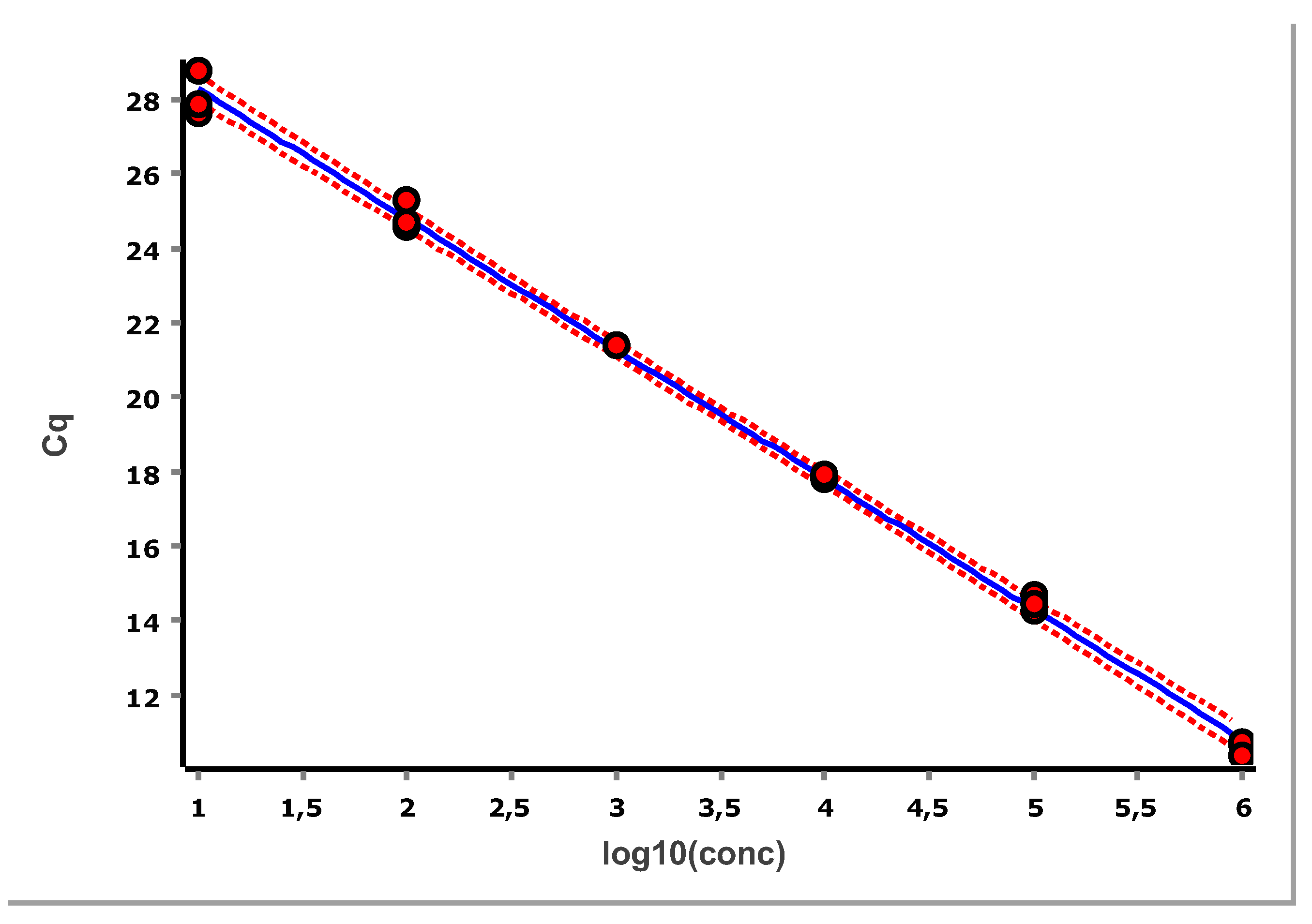
Linear regression of the data in
Figure 11 yields the slope and intercept, along with the Working-Hotelling 95 % confidence band, shown with red dashed lines. The confidence region is derived from error propagation and, for this example, is exceedingly small, which is a positive and direct consequence of the large number of replicates. The PCR efficiency (E) is estimated as in Equation 4 [
6]:
The standard error (SE) of the mean of the PCR efficiency can be estimated using Equation 5 [
6]:
for which the Student’s 95 % CI, considering the
t factor at 95 % confidence level and n-2 degrees of freedom, with n being the total number of samples used in the curve, is calculated using Equation 6 [
20]:
For the data in
Figure 11 we obtain:
|
|
−3.33 [-3.37, -3.29] |
|
|
34.1 [34.0, 34.2] |
|
|
0.996 [0.978, 1.014] |
3.10. Real-Life Standard Curves
Typical standard curves have fewer replicates. Still, the standard samples shall be independent and handled the same way as future test samples, including preanalytics [
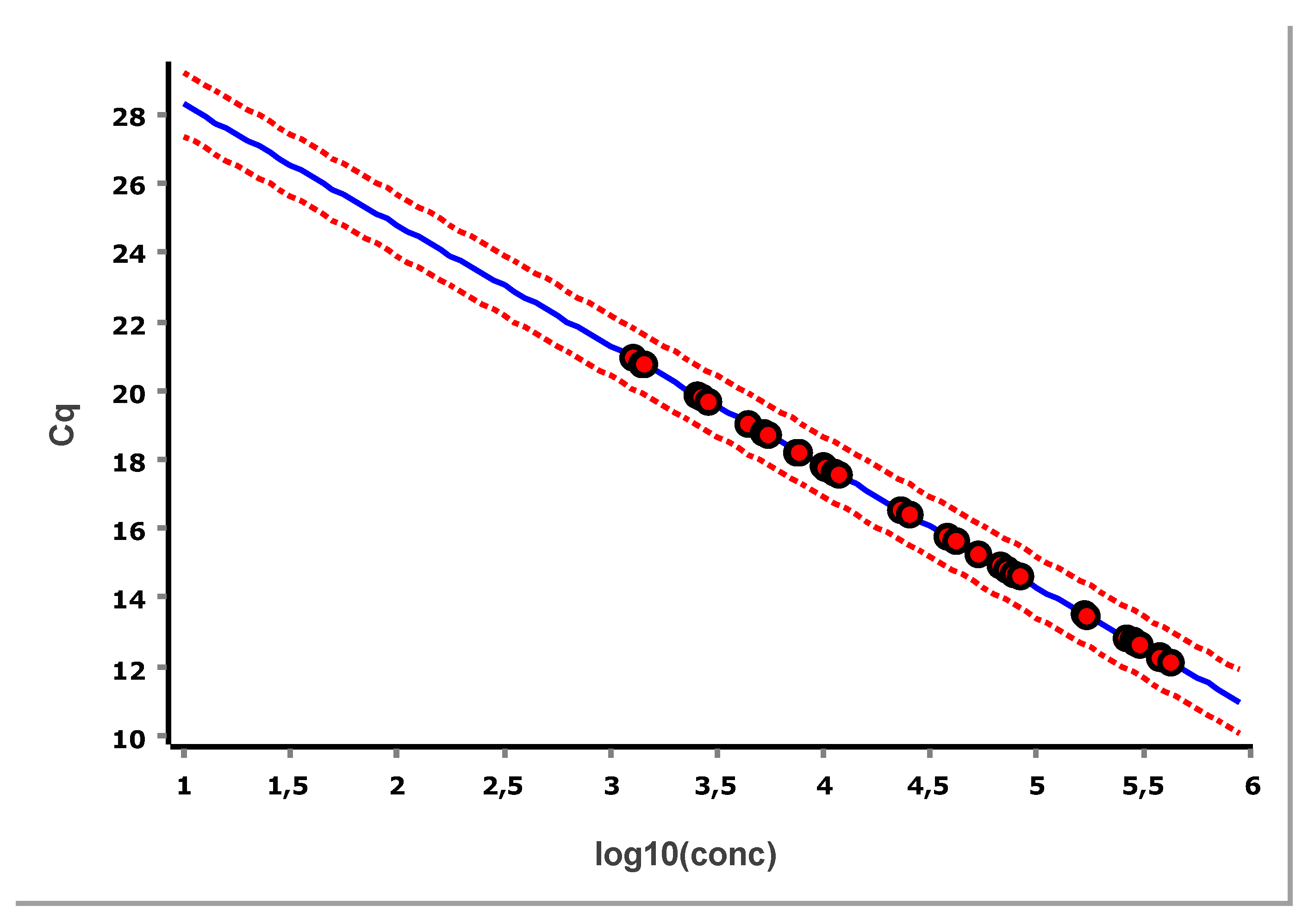
10]. To get the example in
Figure 13 standard samples were measured in triplicate in dilution steps of ten covering six logs in concentration.
The data are fitted using OLS, which yields the following regression parameter estimates along with their 95 % CIs:
|
|
−3.42 [-3.50, -3.34] |
|
|
31.58 [31.21, 31.96] |
|
|
0.960 [0.928, 0.992] |
The PCR efficiency, estimated as 96 ± 3 %, is well within acceptable limits. Also shown in
Figure 13 is the Working–Hotelling area illustrating the precision of the fit. The standardized residuals plot is shown in
Figure 14.
3.11. Validation of the Linear Dynamic Range
While the dynamic range extends from LLOQ to ULOQ, the linear range, which is usually narrower [
21], corresponds to the interval where Cq is proportional to log(N). The linear range (Equation 7) can be assessed by fitting the data to higher-order polynomial models and testing whether the
k1 and
k2 terms in Equations 8 and 9 have statistical significance:
13
The second-order polynomial accounts for curvature at one end of the concentration range, while the third-order model takes into account curvature at both ends. An F-test is used to determine whether the inclusion of higher-order terms significantly improves the fit compared with the linear model. We will refer to this as ‘polynomial test’ [
13,
22].
The full dataset spanning six logs, when fitted by OLS regression, fails the polynomial test for the overall linear dynamic range, i.e., the distribution of the standards does not follow a straight line. This result may initially appear surprising, as no other indicators suggest a problem. However, closer inspection of the residuals plot reveals that all three replicates at the highest concentration exhibit positive residuals, while all three replicates at the second-highest concentration exhibit negative residuals. This systematic pattern indicates curvature at the upper end of the range.
After removing the three highest-concentration samples, the revised standard curve is shown in
Figure 15.
Fit by linear regression yields the following parameters:
|
|
−3.50 [-3.59, -3.41] |
|
|
31.80 [31.44, 32.16] |
|
|
0.930 [0.897, 0.964] |
The corresponding standardized residuals plot is shown in
Figure 16.
Although a slight curvature at high concentration may still be observed, it is not statistically significant. The data now pass the polynomial test, and we conclude the method exhibits a linear dynamic range of five orders of magnitude. The PCR efficiency estimate is somewhat reduced to 93 ± 3 % but remains within acceptable limits.
The polynomial test is very sensitive to deviations and must be used thoughtfully. When the standard samples fall very precisely on a straight line, the test may indicate deviations that are purely noise dependent and may suggest removing good data points. We therefore recommend considering also the “allowable deviation from linearity”, as stated elsewhere [
13]. If it is below 20 %, we would typically accept the standard curve.
Deviations from linearity at high concentrations can arise from several factors, including inhibition in undiluted samples, errors in baseline subtraction when fluorescence accumulates at very early cycles, or incorrect dilutions. In many cases, the exact cause remains unidentified. If uncorrected, the too high a Cq value of the most concentrated sample causes a tilt of the fitted straight line, reducing its slope. A reduced slope implies a higher PCR efficiency, which is incorrect. The impact can be large due to the lever effect and may result in PCR efficiency estimates above 100 %.
The true PCR efficiency cannot exceed 100 %, which corresponds to copying all molecules in the sample every amplification cycle. An estimate, though, may be higher due to experimental error and imprecision. This is reflected by the CI of the PCR efficiency estimate, which then should encompass 100 %.
If the lower confidence range of the PCR efficiency estimate is above 100 % the result should not be accepted as, most likely, concentrations outside the linear dynamic range are included, causing an artificial increase in the efficiency estimate.
3.12. Impact of outliers
In
Figure 17 we have included an outlier sample in the standard curve in
Figure 13 for illustrative purposes.
The outlier is readily identified with a statistical test like the Grubbs and would normally be removed. However, here we keep it to illustrate its impact.
The linear regression yields the following estimates (95 % CI):
|
|
-3.61 [-3.88, -3.33 ] |
|
|
32.57 [31.34, 33.79] |
|
|
0.894 [0.800, 0.988] |
The estimated PCR efficiency is 89.4 %, close to the commonly acceptance threshold of 90%. This illustrates that PCR efficiency alone is not a sensitive indicator of problems in standard curve data.
In contrast, the CI of the PCR efficiency is highly informative. Although 89.4 % is the point estimate, the 95 % confidence range spans from 80 % to 99 %, which is excessively wide and provides little certainty about the actual PCR efficiency. Indeed, the width of the CI is a more meaningful quality indicator than the point estimate itself.
When applying the polynomial test to study the goodness of the linear fit, the presence of the outlier prevents the detection of the deviation from linearity that was clearly visible in
Figure 17. This occurs because the outlier introduces substantial imprecision and biases the least squares fit to itself. This not only reduces the precision of the estimated PCR efficiency, but also affects the entire linear regression, as reflected by the very broad Working–Hotelling confidence band. Due to this imprecision, the triplicate samples at the second-highest and highest concentrations are no longer significantly below and above the fitted regression line, and the linear range would erroneously be accepted.
This example highlights the critical importance of carefully assessing reliable standard data when constructing qPCR standard curves. A single outlier that is not removed can have a profound impact, particularly if it is at an extreme low or high concentration, on regression results and lead to misleading conclusions.
3.13. Prediction of Concentrations of Test Samples
Figure 18 uses the standard curve established in
Figure 15 to predict the concentrations of test samples.
For accurate prediction, standard samples used to construct the standard curve must be handled exactly in the same manner as the test samples and must have exactly the same matrix effects.4 This ensures that the same sources of confounding variation are maintained. For example, if the test samples were analyzed using technical replicates that were subsequently averaged, the standard samples must be analyzed and averaged in the same way.
Test samples may be available as biological replicates. In such cases, two approaches are possible. The biological replicates can be analyzed separately, yielding independent readouts that may be averaged if a single estimate is desired. Alternatively, the Cq values of the biological replicates can be averaged before estimation, resulting in a single concentration estimate. In the statistical analysis, it is important to account for this averaging, as it reduces variation and therefore improves precision.
The concentration in logarithmic scale of a test sample is estimated by applying Equation 10 and considering the measured Cq and the slope (
k) and intercept (
m) of the standard curve:
The standard curve is typically validated on each sample run using at least two quality control samples at high and low concentrations, with a preference for three (high, medium, and low concentrations). Intra and inter-assay accuracy of +/- 1 cycle (corresponding to 50 to 100 % relative error (RE) in linear scale) is a common validation criterion [
4].
The Working–Hotelling prediction band is wider than the corresponding confidence band for the fit because the former includes three sources of uncertainty. In addition to the imprecision of the slope (k) and the imprecision of the intercept (m), the imprecision of the measured Cq of the test sample also contributes to the imprecision of predictions.
The standard error, or uncertainty, of the estimated concentration in logarithmic scale of a test sample is given by Equation 11 [
23,24].
The equation looks complex, although it can be implemented readily in a spreadsheet. It is worth looking into the different parameters it contains to understand its essentials.
is the standard error of the least squares fit or the average error of the standard curve. It reflects the average spread of data in the residuals plot in
Figure 16. The lower the
, the better the fit of the experimental points to the line.
b is the number of test sample replicates. The more replicates, the higher is the precision of the predictions. When measurement error dominates, precision scales with the square root of b.
n is the total number of standard samples used to construct the standard curve. n = the number of levels ✕ the number of replicates (or samplings) at each level. For method development, a minimum of nine levels is recommended in at least four replicates [
13].
is the difference between the measured Cq and the average Cq of all the standard samples, squared. Its effect is that precision is highest at the center of the standard curve and decreases towards the edges, as reflected by the Working-Hotelling band.
is the sum of squared differences between the standards’ concentrations and the mean concentration of all the standards in logarithmic scale. This is the analytical measuring interval of the standard curve and should be within the linear dynamic range. The wider the interval, the greater the precision of the predicted concentrations.
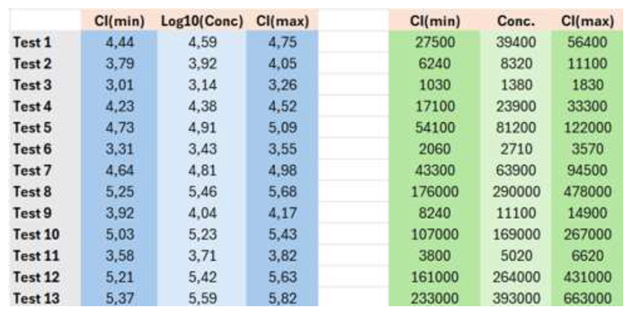
From the SE of the interpolated value, the CI is obtained considering Equation 12:
This is on a logarithmic scale, and the CI is symmetric around the point estimate (
Table 2).
For example, the estimated concentration of sample 5, with the corresponding 95 % CI, is .
Note the CI is symmetrical around the point estimate, and the relative error is about 7.3 % (=100*(5.09-4.73)/4.91). In a linear scale, the CI is asymmetric. The best estimate is , which has an RE of 83.5 % (=100*(121900-54100)/81200).