1. Introduction: Depression, Unmet Needs, and the Search for Blood Biomarkers
Clinical depression affects hundreds of millions of people worldwide and remains a leading cause of disability [
1]. MDD is frequently accompanied by anxiety disorders and is a major contributor to suicide risk. Despite therapeutic advances, remission rates remain limited and decline with successive treatments, underscoring the need for objective biomarkers to monitor treatment outcomes and identify targetable pathways for intervention. Here, we consider intron retention (IR) as a regulatory RNA-processing readout that can both report and potentially mediate stress adaptation.
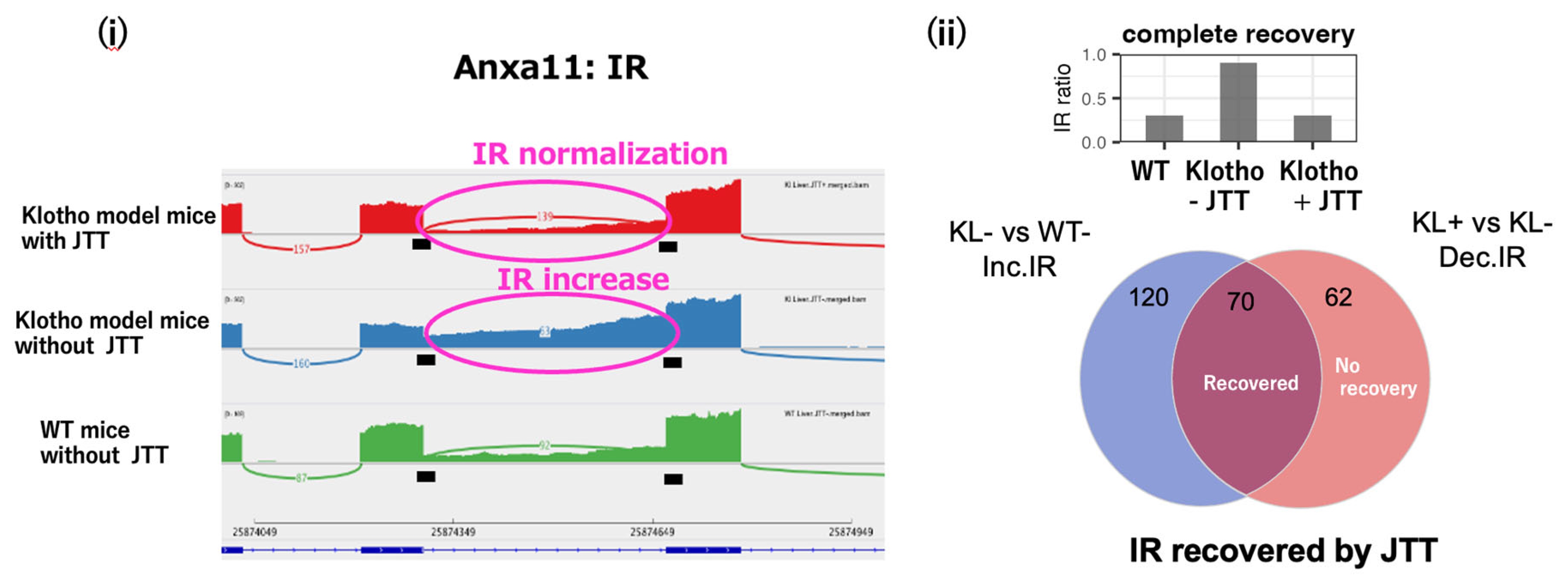
A motivating observation is that IR can behave as a reversible, drug-responsive state marker rather than mere splicing noise. In a pre-symptomatic, starvation-like state in klotho mice, retained introns accumulated across organs, and administration of the traditional multi-herbal medicine juzentaihoto (JTT) restored many IR events toward a healthy pattern (
Figure 1) [
2]. Importantly, the Venn diagram in
Figure 1(ii) highlights that approximately 70 IR-affected genes show recovery toward the control pattern, implying that a shared upstream control mechanism can coordinately tune IR across diverse transcripts. Notably, JTT (and the related Kampo formula ninjinyoeito) also alleviates depressive-like behaviors and normalizes hippocampal neuroinflammatory and genome-maintenance transcriptional programs in a depression-like aging model (SAMP8) [
3]. Independent work in SAMP8 mice further links depressive-like behavior to hippocampal neuroinflammation and shows that Kampo formulas can mitigate these emotional/inflammatory disturbances [
4]. Consistent with this mood disorder-relevant context, hippocampal RNA-seq in SAMP8 mice has also documented stress-associated IR accumulation in homeostasis “sensor” genes, with restoration toward the healthy state by Kampo medicines [
5]. Together, these observations suggest that IR is embedded in a controllable homeostatic layer and set the stage for the IR-Homeostat model proposed later in this review.
Blood-based biomarker development has been motivated by feasibility and by growing evidence that peripheral inflammation correlates with depression incidence. Meta-analyses have reported elevated peripheral inflammatory markers—most robustly IL-6 and CRP—in subsets of patients with major depression, although consistency varies across cytokines and study designs [
6,
7,
8]. Preclinical aging models also support this inflammation–mood link. Specifically, SAMP8 mice exhibit depression-like behaviors accompanied by hippocampal neuroinflammation, and these changes can be buffered by Kampo formulas [
4]. In clinical settings where inflammatory changes precede depressive symptoms, the hypothesis that inflammation contributes to depression has gained traction, supporting a framework in which blood is not merely a surrogate tissue but may capture upstream pathogenic drivers. Importantly, inflammatory signaling can regulate not only transcriptional abundance but also RNA-processing states, making IR/DI a plausible interface where upstream homeostatic inputs are converted into output control.
Against this backdrop, we treat IR/DI as a mechanistically interpretable RNA-processing layer for blood-based biomarker discovery. Building on the intervention-responsive IR-normalization paradigm in aging/stress models (
Figure 1) [
2,
9], we hypothesize that IR constitutes an early and reversible stress readout with utility for mood disorder biomarker development. We therefore organize this review around the idea that IR/DI acts as a post-transcriptional “throttle” on effective gene output, culminating in an IR-Homeostat model that links sensed deviations from physiological set points to switchable IR/DI regulation.
2. Why Differential Gene Expression Struggled as a Universal Blood Biomarker (Up to ~2016): Context Sensitivity and Cohort Instability
By the early-to-mid 2010s, many studies attempted to derive blood-based diagnostic signatures for MDD using genome-wide differential expression analyses, predominantly with microarray platforms. A representative example is the report by Hori et al., which noted highly variable findings across prior MDD transcriptome studies and combined discovery with candidate gene and pathway/network analyses in medication-free outpatients [
10].
Whole-blood gene expression is an exceptionally sensitive, genome-wide, multi-gene state readout. Expression abundance can be shifted by infection and subclinical inflammation, sleep and circadian timing, diet, smoking and BMI, psychosocial stressors, and medication history. Technical and analytical factors further destabilize DEG rankings; batch effects can produce apparent group differences if not detected and corrected [
11].
A particularly important issue is cellular composition heterogeneity. Whole blood and PBMC samples are mixtures of leukocyte subsets, whose proportions vary across individuals and can shift with stress and inflammation. Because many transcripts are cell-type enriched, mixture changes can masquerade as DEGs, reducing transferability across cohorts [
12,
13].
These features explain why DEG panels have struggled to become reliable, universal blood biomarkers. Specifically, the measurement target (steady-state mRNA abundance) is intrinsically state- and composition-sensitive, and it is further perturbed by technical variation. This motivates shifting biomarker emphasis from abundance endpoints to more regulated control processes.
3. The Bullmore Turning Point: Peripheral Inflammation as a Causal Driver Legitimatizes Blood Sampling
Bullmore’s “inflamed mind” framework proposed that, for a substantial subset of patients, peripheral inflammation can act upstream of depressive symptoms, providing a plausible causal route from systemic immune activation to altered mood and cognition [
14]. If this framework is correct, blood becomes a legitimate compartment in which disease-relevant upstream drivers can be measured.
Crucially, this physiological framing does not exclude psychological causation. Negative life events, chronic adversity, and other psychosocial stressors can be conceptualized as upstream inputs that are translated into measurable physiological load through well-established pathways (e.g., neuroendocrine/autonomic signaling, sleep disruption, and downstream immune–metabolic changes). In this sense, blood-based markers capture the physiological implementation of psychosocial stress rather than the subjective experience itself. This provides a natural context to position RNA-processing readouts, such as IR/DI, as candidate state variables: they may reflect how psychosocial and biological stressors jointly shift upstream regulatory states and how those states normalize during recovery.
Evidence that extends beyond association strengthens this framework. Childhood IL-6/CRP levels predict later depression risk in population-based longitudinal data [
15]. Immune activation during IFN-α treatment is associated with subsequent depressive symptom trajectories [
16]. Stratified intervention data suggest that baseline inflammatory status moderates antidepressant response to anti-inflammatory therapy in TRD [
17].
Although these findings strengthen the rationale for blood sampling, they do not resolve DEG instability. The translational task becomes identifying a mechanistically anchored readout that resists context dependence—an argument that motivates RNA-processing regulation, particularly IR/DI.
4. The “Reproducibility Crisis” Does Not Imply Misconduct: A Framework That Explains DEG Instability
The reproducibility debate has largely framed irreproducibility as a structural and methodological challenge rather than a phenomenon attributable primarily to misconduct. Begley and Ellis argued that raising experimental standards is essential because influential preclinical results often fail to replicate under independent verification [
18]. A
Nature survey similarly reported that many researchers experience replication difficulties and perceive reproducibility as a major problem [
19].
This framing is directly relevant to psychiatric biomarkers: failure to replicate can emerge from the interaction between a sensitive measurement target and heterogeneous human biology. Blood transcriptomics is intrinsically vulnerable because expression abundance is reactive to multiple state variables and strongly influenced by leukocyte composition. In addition, it is susceptible to technical variation (e.g., batch effects) [
11,
12,
13].
A more promising strategy is to focus on molecular layers that represent regulated control processes rather than final abundance endpoints. This motivates emphasizing IR/DI as a candidate biomarker layer.
5. Intron Retention as a Stress-Responsive RNA-Processing “Throttle”: IncIR (Brake) and DecIR (Accelerator)
IR is a form of alternative splicing in which introns remain within transcripts that would otherwise be fully spliced. IR is increasingly recognized as a conserved and regulated mode of gene control [
20,
21,
22,
23,
24]. Retained introns can regulate gene output through NMD in some contexts and through nuclear detention/retention, which prevents translation-competent mRNA from reaching the cytoplasm [
24].
We conceptualize IR as a throttle controlling the effective supply of mature, export-competent (fully spliced; nuclear export-competent) mRNA. Here, IncIR and DecIR denote the direction of IR change relative to the matched baseline (homeostatic) state. IncIR indicates a stress-associated increase in intron-containing precursor/DI transcripts (higher IR), whereas DecIR indicates a stress-associated decrease (lower IR). In this framework, IncIR functions as a brake, whereas DecIR functions as an accelerator. This model aligns with the view that IR can regulate gene output via nuclear detention and/or decay routes and with observations that IR signatures can normalize during recovery [
2,
9,
24].
Detained introns (DIs) are an operationally defined, mechanistically prominent subset of intron-retention events: introns retained in nuclear polyadenylated transcripts with substantial half-lives, often not primarily targeted to NMD [
25]. Because DI status is defined by nuclear enrichment/retention, it is formally established using subcellular fractionation or comparable assays; most bulk blood RNA-seq datasets (including those discussed here) do not separate nuclear and cytoplasmic compartments. Accordingly, throughout this review we use “IR” as an umbrella term that may encompass DI-like nuclear-retained introns as well as NMD-linked retained introns, and we use “IR/DI” to emphasize the shared switchable control logic across these contexts. Consistent with this view, DI splicing can change rapidly in response to signaling perturbation [
25]. In neurons, activity-dependent signals can trigger rapid splicing/export/ribosome loading, releasing mature mRNAs within minutes of stimulation [
26]. Together, DI biology and rapid IR switching support IR/DI as a switchable, reversible control layer.
6. Mechanistic Exemplars: Stimuli Switch IR/DI States to Control Output Kinetics and Homeostasis
Across systems, this IR/DI control logic is repurposed for distinct physiological needs, and the direction of the switch (DecIR-like release vs IncIR-like detention) is itself informative. In immune effector control, cytokine synergy provides a particularly clean example. Here, IL-12 primes IFNG transcription yet leaves a substantial fraction of intron-containing transcripts, and IL-2–NF-κB signaling acts as a required secondary signal (“permission”) that drives productive processing to mature mRNA, rapidly boosting IFNγ output (DecIR-like acceleration). In metabolic homeostasis, DI switching implements feedback control. Specifically, OGT intron detention buffers O-GlcNAc under high-load conditions (IncIR-like brake) and is relieved when output must be restored (DecIR-like release), whereas SAM depletion promotes intron removal to increase MAT2A output. In fast neural programs, neuronal activity-triggered intron excision releases pre-existing transcripts on a minute-scale (e.g., within ~15 min), without waiting for new transcription (elongation). Together, these cases argue that IR/DI switching is not pathway-specific noise but a regulated mechanism that tunes output kinetics and prevents overshoot or conflict by gating translation-competent mRNA availability in response to defined inputs [
26,
28,
29,
30,
31]. Representative mechanistic exemplars across systems are summarized in
Table 1.
Representative examples across immune, metabolic, and neuronal systems illustrate how defined inputs (e.g., cytokine co-stimulation, O-GlcNAc or SAM availability, neuronal activity, and differentiation cues) trigger switch-like intron retention/detention (IR/DI) control at specific loci in key regulatory genes. In the IR-Homeostat framework, IncIR/DI detention acts as a reversible “brake”, whereas DecIR/DI release/excision acts as an “accelerator”, thereby modulating the supply of translation-competent mRNA and tuning protein output kinetics. Columns list the input/trigger, biological system, target gene/locus, IR/DI switching mechanism, functional output consequence, and key references.
Collectively, these examples support the central translational claim of this review: IR/DI captures regulated switching at control nodes, which may generalize more robustly across cohorts than steady-state abundance endpoints that are highly sensitive to environment and cell composition (
Section 2,
Section 3 and
Section 4).
Expanded exemplar (adult neurogenic niche). González-Iglesias et al. demonstrate that intron detention can operate as an on/off switch that prevents conflicts between antagonistic transcriptional programs in adult neural stem cells (NSCs). Specifically, transcripts from differentiation genes are detained (intron-retained, nuclear-enriched) in NSCs, whereas stemness-related transcripts are preferentially processed and exported. Upon differentiation cues, this balance switches, and m6A-dependent release of intron detention enables coordinated nuclear export and rapid output activation [
27]. Importantly, their genome-wide analysis shows that IR/DI patterns separate early differentiation time points more clearly than DEGs (Figure 6 in [
27]), consistent with IR as an early-state readout. In Figure 7 of [
27], they provide representative examples of two opposite DI/IR programs that map naturally onto our IncIR/DecIR framework. Here, Cluster 1 follows a DecIR-like pattern (detention in NSCs, splicing upon differentiation), exemplified by Chd5, Sox6, Atl1, and Camk2a, whereas Cluster 2 follows an IncIR-like pattern (productive processing in NSCs, increased detention upon differentiation), exemplified by Kat2a, Lgr5, Fancc, and Ptprv. These paired examples provide a particularly clear demonstration that IR/DI switching can implement homeostatic or fate-decision control by gating the supply of translation-competent mRNA.
Together, the exemplars in
Table 1 illustrate why IR/DI can function as a robust state readout. By modulating the availability of translation-competent mRNA, IR/DI provides a reversible RNA-processing control layer that can reshape protein output kinetics without requiring large de novo transcriptional remodeling.
7. Practical Implications: A Layered Biomarker Strategy Centered on IR/DI
Why overlap matters. A practical method to judge whether a molecular signature is likely to generalize is to ask whether it recurs across independent cohorts. Because steady-state abundance is highly sensitive to sampling context and cell-composition differences (
Section 2,
Section 3 and
Section 4), DEG lists often show limited gene-level overlap between studies. In a military analogy of immunity where macrophages are stationed like “border guards or centurions” across tissues and lymphocytes are more like “generals” (Bullmore [
14], pp. 28, 34), DEGs resemble rank-and-file soldiers—numerous downstream effectors whose abundance shifts with local “battlefield” conditions. In contrast, IR/DI highlights higher-level command decisions (a “commander-in-chief” layer) that gate the supply of translation-competent mRNA at pre-wired control loci [
9]. If this model is correct, IR gene sets are expected to converge more strongly across cohorts than DEGs. We summarize this intuition schematically in
Figure 2.
Empirical example. In our cross-cohort comparison (Figure 8D in Okada et al. [
9]), IR gene sets (IncIR+DecIR) from a Japanese cohort at an early, subthreshold depressive stage overlapped strongly with two independent MDD cohorts: a Chinese case–control study (Zhang et al. [
39]) and a European treatment-resistant depression cohort enriched for ketamine nonresponders (Cathomas et al. [
34]). Despite differences in ancestry and clinical stage, pairwise IR overlaps were enriched 2.5- to 3.2-fold over random expectation (p < 8.6×10^-12 to < 1.1×10^-19), and 15 IR genes were shared across all three datasets. In contrast, the corresponding DEG comparison (Figure 8E in Okada et al. [
9]) yielded only two genes shared across all three studies and fold-enrichment values close to or below 1 (0.6–0.8), consistent with near-random overlap.
Together, these features make IR/DI readouts better positioned than DEGs for subtype stratification and monitoring changes within individuals over time. IR/DI reports upstream, reversible RNA-processing “throttle” decisions and shows stronger cross-cohort reproducibility (e.g., Figure 8D–E in Okada et al. [
9]). These findings align with frameworks proposing an immune-activated/inflammation-linked subtype of depression and mechanistic exemplars of regulated IR/DI control [
2,
6,
7,
8,
9,
14,
15,
16,
17,
18,
19,
20,
21,
22,
23,
24,
25,
26,
28,
29,
30,
31].
Although blood cell composition can influence bulk transcriptomic outputs, the strong cross-cohort reproducibility of IR gene sets across ancestries and across PBMC versus whole blood suggests that the IR layer highlighted here is relatively robust to such variation. Nevertheless, composition-aware checks (CBC/differential or deconvolution) remain advisable for clinical deployment.
IR complements (rather than replaces) existing depression biomarker readouts. Existing blood biomarkers for depression include peripheral inflammatory markers, such as CRP and IL-6, which index inflammatory load in subsets of patients [
6,
7,
8]. These markers are clinically practical and can help define immune-activated subtypes. However, they are nonspecific and do not report upstream RNA-processing states. We therefore position IR/DI as a complementary biomarker layer: inflammatory markers quantify immune load, DEGs summarize downstream abundance outputs, and IR/DI reports regulated RNA-processing throttle states that are reversible and exhibit stronger cross-cohort reproducibility than DEG lists [
9,
34,
39].
In practice, a layered strategy can integrate these readouts, rather than relying on a single measure. For example, combining CRP/IL-6 with IR/DI state signatures may help distinguish high-load immune activation states from other stress programs and may improve longitudinal monitoring of treatment engagement and recovery, whereas DEG panels can still provide pathway-specific effector information when composition and context are controlled [
11,
12,
13].
Table 2 summarizes the complementary roles, strengths, and key confounds of IR/DI, DEGs, and inflammatory markers.
In practice, an IR-centered workflow involves the following: (i) defining biological stress axes (e.g., inflammation and metabolic/oxidative stress); (ii) quantifying IR/DI throttle states at sentinel loci; (iii) enforcing robustness by requiring cross-cohort overlap and within-individual longitudinal normalization; (iv) connecting loci to mechanism via exemplars; and (v) translating the minimal set into targeted assays suitable for clinical use [
2,
6,
7,
8,
9,
11,
12,
13,
25,
26,
28,
29,
30,
31,
32,
33].
8. Ketamine: Responder/Nonresponder Stratification and Pharmacodynamic Profiling
Cathomas et al. generated whole-blood RNA-seq profiles in TRD patients before and after ketamine, enabling interrogation of response heterogeneity [
34]. Building on these data, our IR-based reanalysis indicated that nonresponders were characterized by an elevated viral infection/innate immune activation state that emerged prominently through an IR-centric pathway structure [
35]. This supports reframing nonresponse not as constitutional resistance, but as a subgroup with an immune activation burden that may prevent clinical response thresholds from being reached within the response window [
35].
IR can also function as a pharmacodynamic readout. Subsets of IR loci showed restoration toward healthy IR states after ketamine treatment regardless of responder/nonresponder classification, suggesting that IR can decompose drug-induced molecular normalization from clinical outcome divergence [
35]. Thus, IR can simultaneously report current state, baseline response-limiting biology, and the direction/magnitude of drug-induced normalization [
34,
35].
9. Evolutionary Perspective: Introns as Units Regulating Homeostasis
The concept of homeostasis has been foundational in physiology for more than a century; however, its molecular implementation remains surprisingly under-specified in most discussions of gene regulation. In many contexts, we describe “homeostatic maintenance” as if it were an inherent property of cells, but it remains unclear whether a general molecular layer exists that actively senses deviations from optimal set points and restores balance through a defined regulatory circuit. Here, we propose that intron retention/detained introns (IR/DIs) may constitute such a mechanism: a switchable RNA-processing layer that converts homeostatic inputs into controlled changes in the supply of translation-competent mRNA, thereby buffering or accelerating protein output to stabilize cellular state.
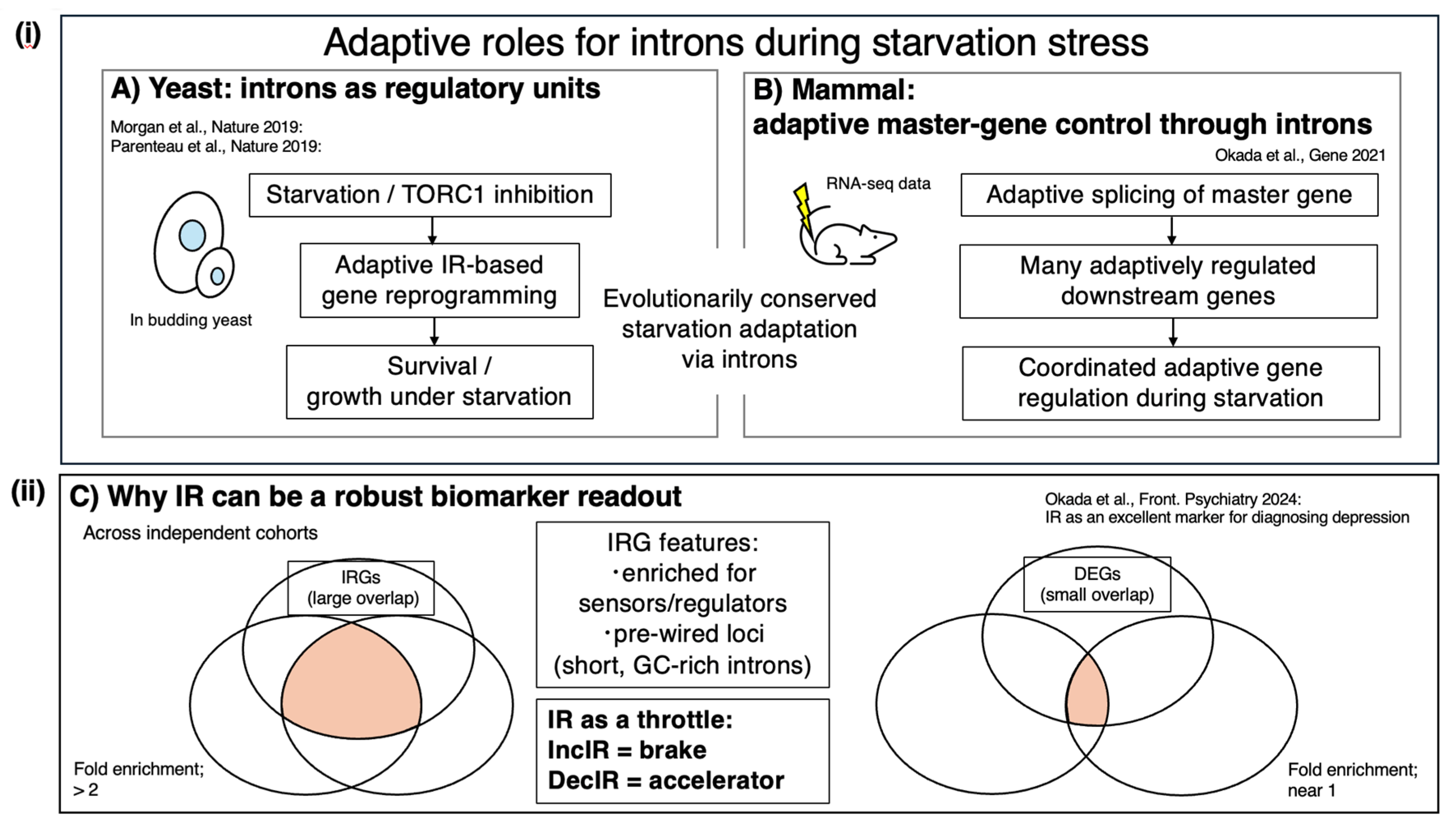
The IR-Homeostat hypothesis is consistent with the idea that introns can function as regulatory units across eukaryotes. In yeast, Parenteau et al. reported that introns mediate cellular responses to starvation, with systematic intron perturbations affecting fitness under nutrient limitation [
36]. Starvation is among the most fundamental adaptive pressures, and our motivating mammalian example—the klotho mouse model depicted in
Figure 1—shows a starvation-like metabolic signature, as evidenced by elevated ketone bodies, together with widespread IR accumulation that can be driven back toward a healthy pattern by intervention [
2]. This supports the view that intron-based buffering of gene output may be evolutionarily conserved. In the same issue, Morgan et al. showed that excised linear introns can accumulate and regulate growth in yeast, demonstrating that intron sequences can retain biological activity even after splicing [
37]. Together, these studies suggest functional diversification of intron-centered control. Specifically, introns can act in cis as switchable detention/retention elements that tune mRNA output during nutrient stress, whereas excised intron RNAs can act in trans to modulate growth programs, potentially converging on nutrient-sensing circuits (e.g., TOR/mTOR). This evolutionary context frames the conceptual comparison shown in
Figure 3, which highlights intron-based regulation as an ancient strategy and motivates its extension to mammalian IR/DI control.
Dobzhansky’s dictum—“Nothing in biology makes sense except in the light of evolution” [
38]—helps connect these observations. The starvation-responsive intron program in yeast [
36] and the starvation-like, ketone-associated IncIR accumulation in klotho mice (
Figure 1) [
2] can be viewed as evolutionarily linked manifestations of intron-based homeostatic control.
Importantly, IR/DI processes are not expected to involve all introns equally. In the IR-Homeostat framework, stress-responsive IncIR/DecIR events are expected to concentrate in a limited subset of cis-tunable “regulatory introns” (i.e., genetically specified control knobs) rather than reflecting random splicing noise. We return to the supporting evidence and factor-dependent intron subclasses in
Section 10 [
24,
32,
33,
40].
These findings suggest that intron-based regulation is not a mammal-specific curiosity, but an evolutionarily conserved strategy for encoding environmental and homeostatic information. In mammals, retained/detained introns can be switched in response to defined inputs (
Table 1), thereby tuning the availability of translation-competent mRNA and protein output kinetics; under this view, the IR signatures we observe in stress and recovery may represent the visible tip of a conserved intron-centered homeostatic control system.
Viewed through this evolutionary lens, intron-based regulation provides a coherent evolutionary backbone for the IR-Homeostat model.
10. The IR-Homeostat Model: Intron Retention/Detention as a Switchable Layer of Homeostatic Control
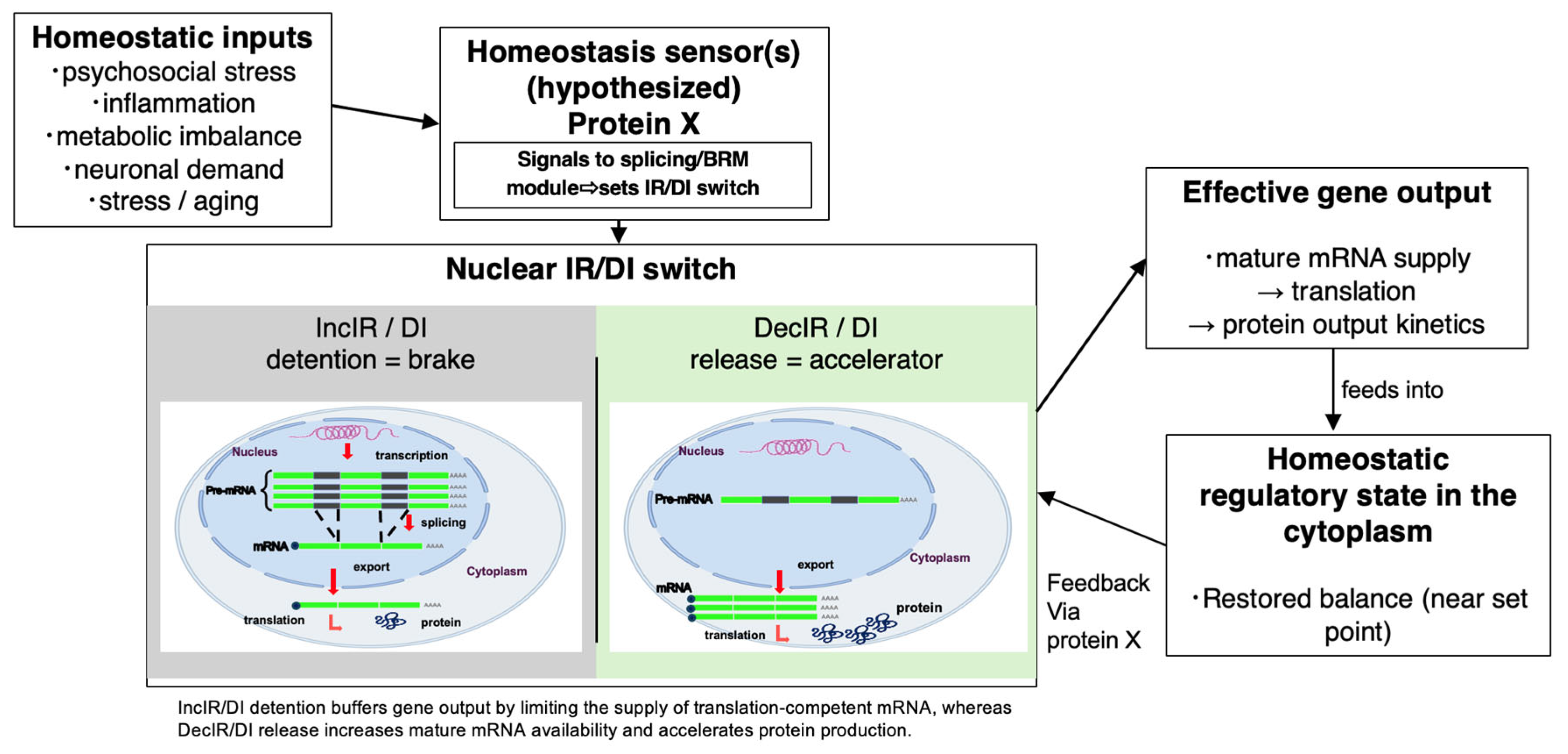
IR-Homeostat model. We propose the “Intron Retention Homeostat” (IR-Homeostat) model (
Figure 3), in which cells sense deviations from physiological set points, such as inflammatory load, nutrient imbalance, or time-critical demand, and translate this information into switchable nuclear RNA-processing states (IR/DI) that control the effective supply of export- and translation-competent mRNA. In this framework, IncIR/DI detention functions as a reversible brake that buffers output under stress, whereas DecIR-like release accelerates mature mRNA production when output must increase.
Mechanistic logic. As illustrated by the exemplars in
Table 1 (
Section 6), homeostatic control can be implemented at the level of mRNA availability. Selected transcripts are held as intron-containing nuclear RNAs and can be rapidly converted into mature, export-competent mRNAs through regulated splicing and release. This gating layer can reshape protein-output kinetics without large de novo transcriptional remodeling, and it can prevent overshoot or conflicts between antagonistic programs by limiting the cytoplasmic mRNA pool until an appropriate context-confirming second input (“permission” signal) arrives (e.g., cytokine co-stimulation, metabolic sufficiency cues, or neuronal activity).
Motivation and evidence. The model is motivated by the drug-responsive normalization of widespread IR patterns in a pre-symptomatic, stress-like state (
Figure 1) [
2] and by mechanistic exemplars showing stimulus-dependent IR/DI switching across immune, metabolic, and neuronal systems (
Table 1, including intron detention-mediated fate switching in the adult neurogenic niche [
27]). Importantly, this control is not expected to involve all introns. IR/DI-prone loci often display distinctive cis-architecture (e.g., short and/or GC-rich introns with suboptimal splice signals), suggesting that “tunable” introns are genetically specified regulatory units rather than stochastic splicing failures. Consistent with this idea, specific classes of human short introns can be spliced in a factor-dependent manner (SPF45/RBM17, with SAP30BP as a cooperative component), demonstrating that alternative splice-factor logic exists for distinct intron subclasses [
33,
40].
Testable predictions. Several concrete predictions follow from the IR-Homeostat framework. (i) If IR/DI encodes a conserved homeostatic control layer, independent cohorts and diverse stressors should repeatedly converge on a core set of tunable IR loci, with consistent IncIR/DecIR directionality as stress load increases or resolves. (ii) In time-resolved perturbations, IR/DI switching should occur earlier than—or partially decouple from—steady-state abundance changes, thereby better tracking output kinetics for selected pathways. (iii) The cis-architecture of tunable introns (length, GC content, weak splice signals, and decoy elements) should predict switchability, such that engineering these features alters detention/release dynamics. (iv) Interventions that restore physiology should drive IR signatures back toward a healthy pattern even when DEG endpoints remain heterogeneous. The next section outlines a practical roadmap to identify the mediators of these throttle decisions and to translate robust IR/DI loci into clinically deployable assays.
11. Clinical Translation
From a translational standpoint, the IR-Homeostat view shifts the emphasis from one-shot, cohort-specific DEG lists toward within-individual longitudinal monitoring and subtype stratification using regulated IR/DI switches. Discovery can rely on genome-wide RNA-seq, whereas clinical deployment could use targeted assays to measure a small panel of sentinel IR/DI events (e.g., junction-/intron-specific RT-qPCR, ddPCR, or targeted amplicon sequencing). Key practical considerations include standardized pre-analytics (blood collection and RNA stabilization/QC), event-level normalization/reporting, and attention to cell composition effects, enabling cost-effective pharmacodynamic monitoring in clinical cohorts.
12. Limitations and Future Directions: From Biomarker Phenomenology to Mechanistic Control
Several limitations and near-term priorities shape the path from descriptive blood signatures to a mechanistic, clinically usable IR/DI framework:
IR calling remains sensitive to read depth, annotation, and thresholding. Because intronic signal is relatively sparse in typical poly(A) RNA-seq, sequencing depths that are adequate for DEG analysis can be underpowered for robust IR/DI detection. Splicing-oriented evaluations suggest performance becomes relatively stable beyond approximately 40-60 million reads per sample, whereas comprehensive splice-event detection can require more than 100 million reads—a key caveat when re-using DEG-centric public datasets. Consensus standards for IR-aware QC, normalization, and longitudinal comparability are required [
32,
41,
42,
43].
Bulk blood conflates immune subsets; composition effects remain relevant and must be addressed using sorting/single-cell approaches and composition-aware modeling [
12,
13].
Although mechanistic exemplars exist (
Table 1), we still lack a mood disorder-specific map that connects clinically relevant inputs (e.g., inflammatory, metabolic, or stress-related cues) to the specific mediators and IR/DI “switch” loci they engage. Perturb-and-measure experiments and CRISPR-based genetic screens provide a practical roadmap to build this causal chain [
25,
26,
28,
29,
30,
31].
Prospective cohorts with repeated measures are needed to test response prediction, relapse risk, and individual-level normalization [
2,
9]. Ketamine datasets illustrate that IR can stratify nonresponders by immune activation burden and capture drug-induced normalization, supporting evaluation of IR as both predictive and pharmacodynamic biomarker [
34,
35].
Different stressors may recruit overlapping but non-identical IR modules; defining core versus context-specific IR programs is essential for generalization [
9,
32].
Model systems that show reversible, drug-responsive IR shifts provide tractable tests for mechanisms. As shown in the klotho model (
Figure 1) [
2], IR accumulation in a pre-symptomatic, starvation-like state can be selectively normalized by juzentaihoto. Complementing this blood-centric framework, CNS-focused datasets (e.g., hippocampal RNA-seq in depression-like SAMP8 mice) and Kampo interventions offer a tractable entry point to test IR/DI predictions in the brain [
3].
13. Concluding Remarks
Across diverse biological systems, IR/DI behaves as a regulated, switchable RNA-processing layer that gates the supply of translation-competent mRNA and thereby tunes output kinetics. In mood disorder research, this property offers a practical advantage. Specifically, IR-based signatures appear more robust to cohort and sampling variation than steady-state abundance endpoints, supporting cross-cohort generalization, subtype stratification, and longitudinal monitoring.
We propose the IR-Homeostat model as a unifying framework that links mechanistic exemplars of detention/release to clinically meaningful “state load” axes, such as inflammatory and metabolic stress. Testing this framework will require standardized IR quantification, cell type-resolved profiling, time-resolved perturbations, and genetic dissection of the regulators that couple signaling inputs to intron switch states.
If successful, an IR-centered strategy could provide both interpretable biomarkers and mechanistic entry points for intervention, enabling separation of molecular normalization from symptom-based thresholds and helping to map how homeostatic stress adaptation is implemented at the RNA-processing level.
Author Contributions
Conceptualization, N.O.; methodology, K.O.;data curation, A.M.; writing—original draft preparation, N.O.; writing—review and editing, N.O.; supervision, A.N.; project administration, Y.K.; funding acquisition, N.O. All authors have read and agreed to the published version of the manuscript.
Funding
The research was supported financially by Tsumura & Co.
Data Availability Statement
The RNA-seq data analyzed shown in
Figure 1 are publicly available in the DDBJ Sequence Read Archive (DRA) under BioProject PRJDB7898 (run accessions: DRR167982–DRR167990; DRR259222–DRR259248). These datasets were originally generated and reported in ref. 2 (Okada et al., Gene 2021, 794:145752). No new sequencing data were generated in this study.
Conflicts of Interest
N.O., A.M., and K.O. were supported by a research grant from Tsumura & Co. A.N. is employed by Tsumura & Co. The remaining author declares that the research was conducted in the absence of conflict.
Abbreviations
| APEX2 |
engineered ascorbate peroxidase 2 |
| CLK |
CDC-like kinase |
| CRP |
C-reactive protein |
| DEG |
differentially expressed gene(s) |
| DI |
detained intron(s) |
| IFNγ |
interferon gamma |
| IFNG |
interferon gamma (gene) |
| IL-6 |
interleukin 6 |
| IR |
intron retention |
| JTT |
juzentaihoto |
| m6A |
N6-methyladenosine |
| MDD |
major depressive disorder |
| METTL16 |
methyltransferase-like 16 |
| NMD |
nonsense-mediated decay |
| NF-κB |
nuclear factor kappa B |
| NK |
natural killer |
| NSC |
neural stem cell |
| O-GlcNAc |
O-linked β-N-acetylglucosamine |
| OGA |
O-GlcNAcase |
| OGT |
O-GlcNAc transferase |
| PBMC |
peripheral blood mononuclear cell(s) |
| RNA-FISH |
RNA fluorescence in situ hybridization |
| SAM |
S-adenosylmethionine |
| SAP30BP |
SAP30 binding protein |
| SF3B1 |
splicing factor 3B subunit 1 |
| SON |
SON DNA/RNA-binding protein |
| SPF45/RBM17 |
RNA-binding motif protein 17 |
| SRRM2 |
serine/arginine repetitive matrix 2 |
| TNF |
tumor necrosis factor |
| TRD |
treatment-resistant depression |
| U2AF |
U2 small nuclear RNA auxiliary factor |
References
- World Health Organization. Depression and other common mental disorders: global health estimates. Geneva: World Health Organization; 2017.
- Okada N, Oshima K, Iwasaki Y, Maruko A, Matsumura K, Iioka E, et al. Intron retention as a new pre-symptomatic marker of aging and its recovery to the normal state by a traditional Japanese multi-herbal medicine. Gene. 2021;794:145752. [CrossRef]
- Maruko A, Ito N, Oshima K, Nishi A, Kobayashi Y, Okada N. Distinct neuroprotective and anti-inflammatory effects of Kampo formulas ninjinyoeito and juzentaihoto in depression-like SAMP8 mice. Front Pharmacol. 2025;16:1600176. [CrossRef]
- Ito N, Maruko A, Oshima K, Yoshida M, Honma K, Sugiyama C, Nagai T, Kobayashi Y, Odaguchi H, Okada N. Kampo formulas alleviate aging-related emotional disturbances and neuroinflammation in male senescence-accelerated mouse prone 8 mice. Aging (Albany NY). 2022;14(1):109-142. [CrossRef]
- Vu TD, Ito N, Oshima K, Maruko A, Nishi A, Mizoguchi K, Odaguchi H, Kobayashi Y, Okada N. Intron retention is a stress response in sensor genes and is restored by Japanese herbal medicines: A basis for future clinical applications. Gene. 2022;830:146496. [CrossRef]
- Howren MB, Lamkin DM, Suls J. Associations of depression with C-reactive protein, IL-1, and IL-6: a meta-analysis. Psychosom Med. 2009;71(2):171-186. [CrossRef]
- Dowlati Y, Herrmann N, Swardfager W, Liu H, Sham L, Reim EK, et al. A meta-analysis of cytokines in major depression. Biol Psychiatry. 2010;67(5):446-457. [CrossRef]
- Haapakoski R, Mathieu J, Ebmeier KP, Alenius H, Kivimaki M. Cumulative meta-analysis of interleukins 6 and 1beta, tumour necrosis factor alpha and C-reactive protein in patients with major depressive disorder. Brain Behav Immun. 2015;49:206-215. [CrossRef]
- Okada N, Oshima K, Maruko A, Sekine M, Ito N, Wakasugi A, et al. Intron retention as an excellent marker for diagnosing depression and for discovering new potential pathways for drug intervention. Front Psychiatry. 2024;15:1450708. [CrossRef]
- Hori H, Sasayama D, Teraishi T, Yamamoto N, Nakamura S, Ota M, et al. Blood-based gene expression signatures of medication-free outpatients with major depressive disorder: integrative genome-wide and candidate gene analyses. Sci Rep. 2016;6:18776. [CrossRef]
- Leek JT, Scharpf RB, Bravo HC, Simcha D, Langmead B, Johnson WE, et al. Tackling the widespread and critical impact of batch effects in high-throughput data. Nat Rev Genet. 2010;11(10):733-739. [CrossRef]
- Jaffe AE, Irizarry RA. Accounting for cellular heterogeneity is critical in epigenome-wide association studies. Genome Biol. 2014;15(2):R31. [CrossRef]
- Houseman EA, Accomando WP, Koestler DC, Christensen BC, Marsit CJ, Nelson HH, et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinformatics. 2012;13:86. [CrossRef]
- Bullmore E. The Inflamed Mind: A Radical New Approach to Depression. London: Short Books; 2018.
- Khandaker GM, Pearson RM, Zammit S, Lewis G, Jones PB. Association of serum interleukin 6 and C-reactive protein in childhood with depression and psychosis in young adult life: a population-based longitudinal study. JAMA Psychiatry. 2014;71(10):1121-1128. [CrossRef]
- Prather AA, Rabinovitz M, Pollock BG, Lotrich FE. Cytokine-induced depression during IFN-alpha treatment: the role of IL-6 and sleep quality. Brain Behav Immun. 2009;23(8):1109-1116. [CrossRef]
- Raison CL, Rutherford RE, Woolwine BJ, Shuo C, Schettler P, Drake DF, et al. A randomized controlled trial of the tumor necrosis factor antagonist infliximab for treatment-resistant depression: the role of baseline inflammatory biomarkers. JAMA Psychiatry. 2013;70(1):31-41. [CrossRef]
- Begley CG, Ellis LM. Drug development: Raise standards for preclinical cancer research. Nature. 2012;483(7391):531-533. [CrossRef]
- Baker M. 1,500 scientists lift the lid on reproducibility. Nature. 2016;533(7604):452-454. [CrossRef]
- Wong JJL, Ritchie W, Ebner OA, Selbach M, Wong JWH, Huang Y, et al. Orchestrated intron retention regulates normal granulocyte differentiation. Cell. 2013;154(3):583-595. [CrossRef]
- Pimentel H, Parra M, Gee SL, Mohandas N, Pachter L, Conboy JG. A dynamic intron retention program enriched in RNA processing genes regulates gene expression during terminal erythropoiesis. Nucleic Acids Res. 2016;44(2):838-851. [CrossRef]
- Naro C, Jolly A, Di Persio S, Bielli P, Setterblad N, Alberdi AJ, et al. An orchestrated intron retention program in meiosis controls timely usage of transcripts during germ cell differentiation. Dev Cell. 2017;41(1):82-93.e4. [CrossRef]
- Ullrich S, Guigó R. Dynamic changes in intron retention are tightly associated with regulation of splicing factors and proliferative activity during B-cell development. Nucleic Acids Res. 2020;48(3):1327-1340. [CrossRef]
- Wong JJL, Schmitz U. Intron retention: importance, challenges, and opportunities. Trends Genet. 2022;38(8):789-792. [CrossRef]
- Boutz PL, Bhutkar A, Sharp PA. Detained introns are a novel, widespread class of post-transcriptionally spliced introns. Genes Dev. 2015;29(1):63-80. [CrossRef]
- Mauger O, Lemoine F, Scheiffele P. Targeted intron retention and excision for rapid gene regulation in response to neuronal activity. Neuron. 2016;92(6):1266-1278. [CrossRef]
- González-Iglesias A, Arcas A, Domingo-Muelas A, Mancini E, Galcerán J, Valcárcel J, Fariñas I, Nieto MA. Intron detention tightly regulates the stemness/differentiation switch in the adult neurogenic niche. Nat Commun. 2024, correction in Nat Commun. 2024, 15, 3458. [CrossRef]
- Van Gelder RD, Gokhale NS, Genoyer E, Omelia DS, Anderson SK, Young HA, et al. Interleukin-2-mediated NF-kB-dependent mRNA splicing modulates interferon gamma protein production. EMBO Rep. 2025;26(1):16-35. [CrossRef]
- Govindan A, Conrad NK. SFSWAP is a negative regulator of OGT intron detention and global pre-mRNA splicing. Elife. 2024;13:RP104439. [CrossRef]
- Tan ZW, Fei G, Paulo JA, Bellaousov S, Martin SES, Duveau DY, et al. O-GlcNAc regulates gene expression by controlling detained intron splicing. Nucleic Acids Res. 2020;48(10):5656-5669. [CrossRef]
- Pendleton KE, Chen B, Liu K, Hunter OV, Xie Y, Tu BP, Conrad NK. The U6 snRNA m6A methyltransferase METTL16 regulates SAM synthetase intron retention. Cell. 2017;169(5):824-835.e14. [CrossRef]
- Monteuuis G, Wong JJL, Bailey CG, Schmitz U, Rasko JEJ. The changing paradigm of intron retention: regulation, ramifications and recipes. Nucleic Acids Res. 2019;47(22):11497-11513. [CrossRef]
- Fukumura K, Yoshimoto R, Sperotto L, et al. SPF45/RBM17-dependent, but not U2AF-dependent, splicing in a distinct subset of human short introns. Nat Commun. 2021;12:4910. [CrossRef]
- Cathomas F, Bevilacqua L, Ramakrishnan A, Kronman H, Costi S, Schneider M, et al. Whole blood transcriptional signatures associated with rapid antidepressant response to ketamine in patients with treatment resistant depression. Transl Psychiatry. 2022;12(1):12. [CrossRef]
- Okada N, Oshima K, Maruko A, Kobayashi Y. Intron retention: a novel method for evaluating the response to ketamine in patients with treatment-resistant depression. npj Ment Health Res. 2025;4(1):44. [CrossRef]
- Parenteau J, Maignon L, Berthoumieux M, Catala M, Gagnon V, Abou Elela S. Introns are mediators of cell response to starvation. Nature. 2019;565(7741):612-617. [CrossRef]
- Morgan JT, Fink GR, Bartel DP. Excised linear introns regulate growth in yeast. Nature. 2019;565(7741):606-611. [CrossRef]
- Dobzhansky T. Nothing in Biology Makes Sense except in the Light of Evolution. Am Biol Teach. 1973;35(3):125-129. [CrossRef]
- Zhang D, Ji Y, Chen X, Chen R, Wei Y, Peng Q, et al. Peripheral blood circular RNAs as a biomarker for major depressive disorder and prediction of possible pathways. Front Neurosci. 2022;16:844422. [CrossRef]
- Fukumura K, Sperotto L, Seuss S, et al. SAP30BP interacts with RBM17/SPF45 to promote splicing in a subset of human short introns. Cell Rep. 2023;42(12):113534. [CrossRef]
- Shen S, Park JW, Lu ZX, Lin L, Henry MD, Wu YN, Zhou Q, Xing Y. rMATS: Robust and flexible detection of differential alternative splicing from replicate RNA-Seq data. Proc Natl Acad Sci U S A. 2014;111(51):E5593-E5601. [CrossRef]
- Mehmood A, Laiho A, Venalainen MS, McGlinchey AJ, Wang N, Elo LL. Systematic evaluation of differential splicing tools for RNA-seq studies. Briefings in Bioinformatics. 2020;21(6):2052-2065. [CrossRef]
- Hackl LM, Neuhaus F, Ameling S, Volker U, Baumbach J, Tsoy O. Detection of alternative splicing: deep sequencing or deep learning? Briefings in Bioinformatics. 2026;27(1):bbaf705. [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).