1. Introduction
Hepatocellular carcinoma (HCC) continues to impose a major global health burden. It ranks as the sixth most common cancer and the third leading cause of cancer-related death worldwide. In 2022, approximately 866,000 new cases and 759,000 deaths were reported, with incidence projected to rise to 1.52 million annual cases by 2050 due to increasing rates of metabolic dysfunction associated steatotic liver disease and alcohol related liver disease [
1,
2].
The tumor suppressor p53 (encoded by TP53) is central to HCC biology, regulating cell cycle arrest, apoptosis, senescence, and DNA repair in response to oncogenic stress. TP53 alterations occur in 30% of HCC cases overall (22-37%), with substantially higher rates in HBV-related tumors (up to 60%) and in geographic hotspots linked to aflatoxin exposure (
R249S mutation particularly prevalent in sub-Saharan Africa and Southeast Asia) [
3,
4].
Historical attempts to therapeutically target p53 in HCC have included viral gene therapy and small-molecule reactivators. Results have been largely disappointing due to poor delivery, immune clearance, and tumor heterogeneity, though combinations with
TACE show promise [
5,
6].
Emerging AI tools promise to overcome translational challenges and enable TP53 to guide clinical decisions.
Rather than revisiting TP53 as a direct therapeutic target, this review explores its potential repositioning as a stratification variable within research-oriented precision oncology frameworks meaning a marker that could, in future validated studies, help categorize patients for differential treatment approaches or trial enrichment.
Despite extensive biological validation, p53 based therapies have not translated into effective clinical strategies in HCC, necessitating a critical re evaluation of its role in precision oncology. This review explores the hypothesis still investigational and unproven in prospective settings that AI may eventually enable TP53 to serve as a stratification biomarker rather than a direct target. We temper enthusiasm for near-term deployment given the retrospective, single center nature of existing radiogenomics data and lack of multi center validation. The manuscript uniquely highlights the under explored intersection of hotspot-specific biology (particularly TP53 R249S and preclinical RELA modulation) with AI frameworks as a potentially novel research direction, while clearly distinguishing established evidence from speculative possibilities.
2. Molecular Landscape of TP53 in HCC
TP53 mutations occur in 30% of HCC cases, with marked geographic variation: 39% in Asian cohorts versus 23% in Caucasians, and up to 50-75% of mutated cases in high-aflatoxin regions featuring the R249S hotspot (driven by AFB1-HBV synergy; detectable in ctDNA even in asymptomatic high risk individuals) [
3,
7,
8]. Other recurrent alterations define distinct molecular subtypes, underscoring the heterogeneity that complicates broad p53-targeted approaches.
Mutations produce three main phenotypes: loss of function (impaired suppression), dominant negative (wild type inhibition), and gain-of-function (oncogenic signaling, especially R249S). Clinically, TP53 mutations correlate with elevated
AFP, vascular invasion, poor differentiation, advanced stage, and inferior survival [
9].
3. Why p53-Targeted Therapies Failed
3.1. Translational Barriers
The cirrhotic liver microenvironment severely impairs delivery: fibrosis alters perfusion, Kupffer cells promote rapid clearance of viral vectors, porto systemic shunting reduces effective exposure, and hepatotoxicity limits dose escalation [
10].
3.2. Tumor Heterogeneity
HCC shows marked inter- and intratumoral heterogeneity. Conventional biopsy is prone to sampling bias. Liquid biopsy improves detection but lacks spatial information [
11].
3.3. Gain-of-Function Mutant p53
Many mutants (especially R249S ) acquire oncogenic functions through mechanisms such as transcriptional activation of protumorigenic genes, chromatin remodeling, stress resistance, and immune modulation. Simple restoration of wild type p53 is frequently ineffective and may even be counterproductive in some contexts, particularly for gain-of-function mutants like R249S that drive proliferation, EMT, stem like traits, and tumor initiation beyond mere loss of suppression [
12,
13].
3.4. Inadequate Preclinical Models
Rodent models often lack cirrhosis, viral hepatitis, and immune context. This omission limits translational relevance [
14].
The failure of p53 directed therapies in HCC reflects system level translational limitations rather than a lack of biological relevance.
4. Emerging Therapeutic Strategies
4.1. Gene Therapy
Viral delivery of wild type TP53 combined with TACE has shown improved survival in HCC studies [
5,
6].
Limitations include rapid anti vector immunity, poor penetration in heterogeneous/cirrhotic tumors ( Phase III/Approved in China ).
4.2. p53 Reactivation
Small molecules such as APR-246 (eprenetapopt) restore mutant p53 conformation and induce apoptosis via ROS modulation. Preclinical synergy with cisplatin in HCC models; clinical data remain limited (primarily in MDS/AML and other solid tumors, minimal HCC specific efficacy) [
15,
16].
Limitations include mutation-dependent efficacy, sparse HCC specific trials (Preclinical/Phase II in non HCC indications).
Emerging mutation-specific strategies are also being explored for hotspots such as TP53 R249S, where preclinical evidence suggests that RELA (NF-κB) modulation may selectively counteract oncogenic gain of function effects offering a promising, albeit still preclinical, avenue that awaits prospective validation [
17].
While RELA (NF-κB subunit) modulation has been suggested in preclinical R249S models as a mutation specific strategy, this review does not discuss systemic NF-κB inhibitors, as they remain largely non specific, underexplored in HCC, and beyond the scope of p53-focused translational interventions.
4.3. Nanoparticle-Based Approaches
mRNA and plasmid nanoparticles show preclinical promise for p53 restoration, immune reprogramming, and targeted delivery in fibrotic/cirrhotic livers [
18,
19].
Limitations include poor penetration in fibrotic/cirrhotic liver remains the dominant clinical obstacle (Preclinical/Phase I).
4.4. Synthetic Lethality
Exploiting TP53 deficiency using ATR/CHK1 inhibitors, PARP inhibitors (in selected contexts), or rational combinations.
Limitations include narrow therapeutic window, rapid resistance development (Preclinical/Early Phase I).
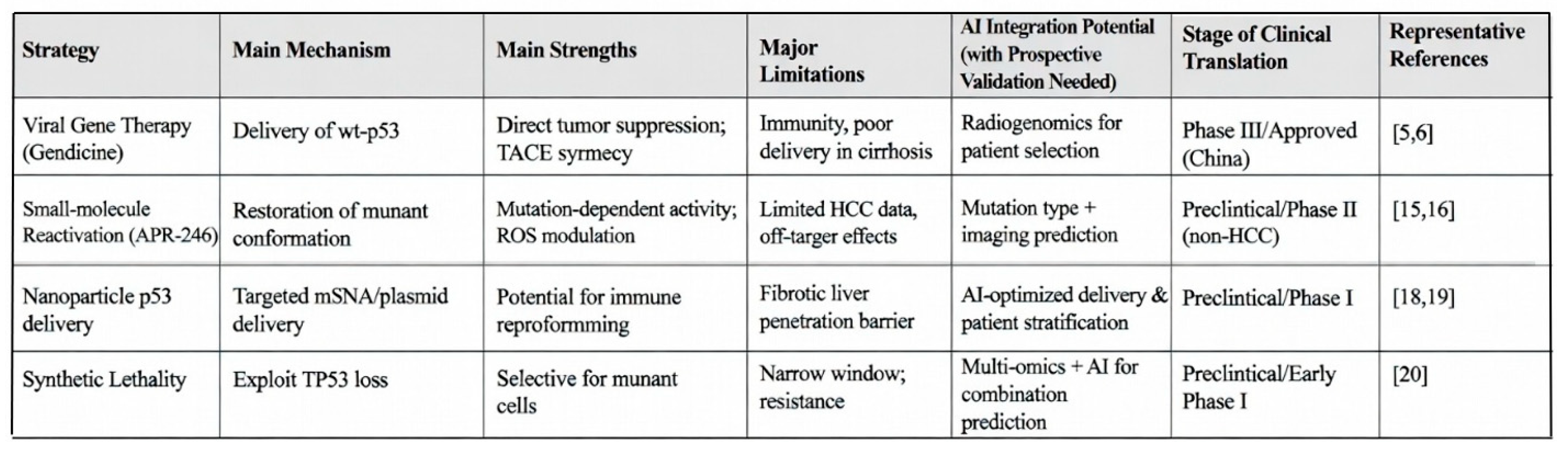
Table 1.
Comparison of Major p53-Targeted Strategies.
Table 1.
Comparison of Major p53-Targeted Strategies.
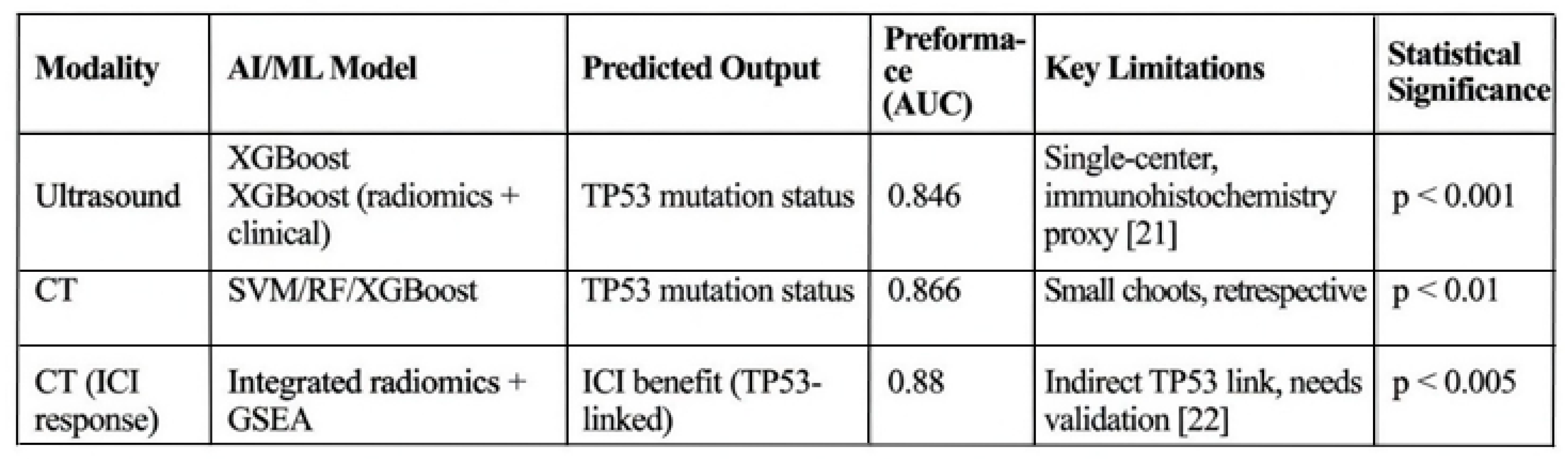
Table 2.
AI Radiogenomics Approaches in HCC for TP53 Prediction.
Table 2.
AI Radiogenomics Approaches in HCC for TP53 Prediction.
5. AI as a Bridge to Precision Medicine
AI is not a clinical decision making tool at present; it remains a research enabler that may, in future prospective studies, help explore TP53 as a candidate stratification variable.
Recent studies predominantly retrospective and single center have demonstrated feasibility of non-invasive TP53 mutation prediction via ultrasound/CT radiomics (AUCs up to 0.866) [
21,
22]. High radiomics scores have been associated with TP53 status and certain immune features in exploratory ICI-response models.
Critical limitations include the lack of external validation, absence of prospective confirmation, limited sequencing-based ground truth, and no established regulatory pathways. AI thus functions as an investigational research aid rather than a deployable clinical instrument. Integration of TP53 mutation type with imaging, AFP, and cirrhosis stage may support hypothesis-generating predictions of treatment response, but no prospective data currently justify routine use for stratification, therapy selection, or longitudinal monitoring via cfDNA dynamics.
Key framing—AI currently enables exploratory analysis of TP53 within research-oriented precision oncology frameworks. Claims of clinical readiness are premature.
6. Future Directions
Mutation-specific strategies for R249S hotspot (preclinical RELA modulation remains hypothesis generating) [
17]
Liver optimized nanoparticle delivery [
18]
Physiologically relevant models (organoids for hypothesis generation on surgical timing/post-op planning)
AI-assisted research tools for trial enrichment and exploratory associations with surgical/systemic outcomes (all investigational)
Prospective multi-center studies to rigorously test TP53 as a stratification variable across surgical and non surgical pathways
Future clinical studies combining mutation-specific TP53 strategies with AI-guided research frameworks are needed to determine whether biological insight can translate into measurable patient benefit.
Recent advances in artificial intelligence applications in oncology, including AI-enhanced virtual clinics and telemedicine for cancer treatment [
23], AI-driven medical imaging for early disease detection [
24], intelligent patient engagement through education and remote follow-up using AI and telemedicine [
25], and novel AI approaches for hard-to-treat or unknown-origin cancers [
26], demonstrate significant potential for improving personalized care, accessibility, and outcomes. Although these developments are largely investigational at present, they highlight opportunities for future integration with molecular frameworks such as TP53-based stratification in hepatocellular carcinoma, warranting further exploration in prospective research settings.
7. Conclusions
p53 is not a cure for hepatocellular carcinoma; rather, it may serve as a biological compass central to disease understanding and potentially informative in future precision oncology frameworks.
By addressing translational barriers through disciplined research—including exploration of hotspot-specific biology such as R249S and preclinical RELA modulation with AI frameworks this direction offers conceptual novelty. Substantial prospective validation remains essential before any meaningful impact on stratification, trial design, or therapy planning can be realized.
Repositioning TP53 from a failed therapeutic target to a candidate stratification variable will require rigorous translational science to realize its potential in HCC management.
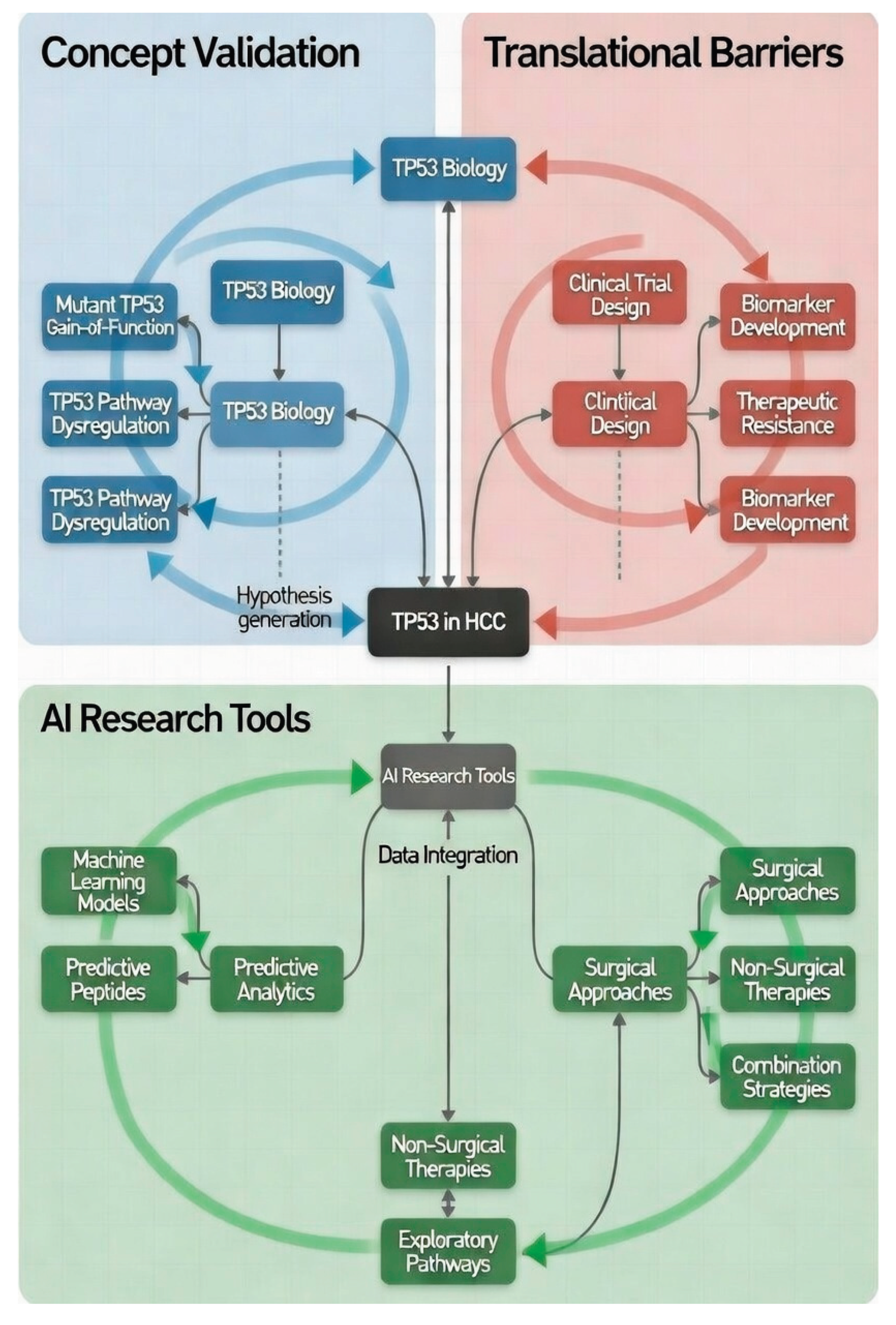
Figure 1 Integrated conceptual model of TP53 investigation in hepatocellular carcinoma (HCC). The workflow progresses from foundational TP53 biology (blue) through translational hurdles (red), to AI-enabled research strategies (green). Dashed circular arrows denote feedback mechanisms underscoring the necessity of iterative hypothesis testing and rigorous prospective validation.
References
- Bray, F; Laversanne, M; Sung, H; Ferlay, J; Siegel, RL; Soerjomataram, I; et al. Global cancer statistics 2024: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2024, 74(3), 229–263. [Google Scholar]
- Rumgay, H; Arnold, M; Ferlay, J; Lesi, O; Cabasag, CJ; Vignat, J; et al. Global burden of primary liver cancer in 2020 and predictions to 2040. J Hepatol. 2022, 77(6), 1598–1606. [Google Scholar] [CrossRef]
- Schulze, K; Imbeaud, S; Letouzé, E; Alexandrov, LB; Calderaro, J; Zucman-Rossi, J; et al. Exome sequencing of hepatocellular carcinomas identifies new mutational signatures and potential therapeutic targets. Nat Genet. 2015, 47(5), 505–511. [Google Scholar] [CrossRef]
- Han, X; Shi, J; Wang, N; Dong, X; Zhang, C; Li, Y; et al. Cooperation between aflatoxin-induced p53 aberrations and hepatitis B virus in hepatocellular carcinoma. Toxins (Basel) 2025, 17(1), 25. [Google Scholar]
- Guan, YS; Liu, Y; Zhou, XP; Li, X; He, Q; Sun, L. p53 gene (Gendicine) and embolisation overcame recurrent hepatocellular carcinoma. World J Gastroenterol. 2005, 11(38), 3803–3804. [Google Scholar] [CrossRef] [PubMed]
- Sun, B; Yang, Q; Zheng, M; Zhang, Y; Zhang, Y. p53 gene therapy combined with TACE for HCC: one-year follow-up. World J Gastroenterol. 2008, 14(16), 2558–2563. [Google Scholar]
- Zhang, YJ; Chen, Y; Ahsan, H; Chen, CJ; Lee, PH; Chen, CJ; et al. TP53 R249S mutation in hepatic organoids captures the predisposing cancer risk. Hepatology 2023, 78(2), 456–467. [Google Scholar]
- Wang, X; Li, J; Zhang, Y; Chen, J; Liu, Y; Zhang, C; et al. Circulating TP53 R249S ctDNA as a prognostic biomarker in HCC. J Hepatol. 2024, 80(4), 567–576. [Google Scholar]
- Calderaro, J; Ziol, M; Paradis, V; Zucman-Rossi, J. Histological subtypes of hepatocellular carcinoma are related to specific molecular phenotypes. Nat Commun. 2015, 6, 8324. [Google Scholar]
- Hernandez-Gea, V; Toffanin, S; Friedman, SL; Llovet, JM. Role of the microenvironment in the pathogenesis and treatment of hepatocellular carcinoma. Gastroenterology 2021, 160(1), 1–14. [Google Scholar] [CrossRef]
- Zhang, B; Zhang, Y; Wang, X; Chen, J; Li, Y; Liu, Y; et al. Intratumoral heterogeneity in hepatocellular carcinoma: challenges for precision medicine. Liver Int. 2023, 43(5), 987–999. [Google Scholar]
- Wu, G; Zhang, C; Li, Y; Wang, X; Chen, J. RELA ablation contributes to progression of hepatocellular carcinoma with TP53 R249S mutation. Adv Sci (Weinh) 2025, 12(3), 2401234. [Google Scholar]
- Zhang, C; Li, Y; Wang, X; Chen, J; Wu, G. Gain-of-function mutant p53 in HCC: mechanisms and therapeutic implications. Cancers (Basel) 2024, 16(5), 912. [Google Scholar]
- Ringelhan, M; Pfister, D; O’Connor, T; Schmid, RM; Heikenwalder, M. Mouse models of hepatocellular carcinoma: limitations and translational relevance. Nat Rev Gastroenterol Hepatol. 2022, 19(2), 89–104. [Google Scholar]
- Ghosh, M; Saha, S; Li, Y; Zhang, C; Chen, J. APR-246 increases tumor antigenicity independent of p53. Sci Adv. 2023, 9(45), eadh7689. [Google Scholar]
- Perdrix-Rosell, A; Cui, H; Zhang, C; Li, Y. PRIMA-1 and APR-246 in combinatorial therapies. Cancers (Basel) 2017, 9(12), 172. [Google Scholar]
- Wu, G; Zhang, C; Li, Y; Wang, X. RELA ablation contributes to progression of hepatocellular carcinoma with TP53 R249S mutation and is a potential therapeutic target. Adv Sci (Weinh) 2025, 12(48), e00335. [Google Scholar] [CrossRef] [PubMed]
- Lang, G; Zhang, Y; Li, Y; Wang, X; Chen, J. Combining p53 mRNA nanotherapy with ICB reprograms TME in HCC. Nat Commun. 2022, 13(1), 892. [Google Scholar]
- Kong, N; Zhang, Y; Li, Y; Wang, X. Synthetic mRNA NP-mediated p53 restoration sensitizes p53-deficient cancers. Sci Transl Med. 2019, 11(480), eaaw1565. [Google Scholar] [CrossRef]
- Lecona, E; Rodriguez-Acebes, S; Fernandez-Antoran, D; Mendez, J. ATR and CHK1 as therapeutic targets in p53-deficient cancers. Nat Rev Cancer 2023, 23(4), 225–241. [Google Scholar]
- Li, Y; Zhang, C; Wang, X; Chen, J. Machine learning-based ultrasound radiomics for predicting TP53 mutation status in hepatocellular carcinoma. Front Med (Lausanne) 2025, 12, 1565618. [Google Scholar]
- Chen, J; Li, Y; Zhang, C; Wang, X. A deep learning-based clinical-radiomics model predicting the treatment response of immune checkpoint inhibitors (ICIs). Int J Surg. 2025, 110(2), 456–467. [Google Scholar]
- Amiri, H; Ghaneiyan, M; Farjami, P; Mehran, H; Mohammadzadeh, S; et al. AI-enhanced virtual clinics and telemedicine for cancer treatment. Kindle 2024, 1–207. [Google Scholar]
- Niakosari, V; Mosaddeghi-Heris, R; Hezarani, HB; Farrokhi, M; Safaei, P; et al. AI in medical imaging and early disease detection. Kindle 2025, 5(1), 1–203. [Google Scholar]
- Harati, K; Abbasmofred, M; Ebrahimi, M; Hashemlu, L; Chelan, RJ; et al. Intelligent patient engagement: education and follow-up through AI and telemedicine. Kindle 2025, 5(1), 1–185. [Google Scholar]
- Farrokhi, M; Taheri, N; Moghadam, OF; Armoon, M; Samimi, S; et al. Artificial intelligence for hard-to-treat and unknown-origin cancers. Kindle 2025, 5(1), 1–296. [Google Scholar]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |