Submitted:
26 February 2026
Posted:
27 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Growth Conditions
2.2. Preparation of Activated Carbon-Amended Substrate
2.3. Preparation of Saline–Alkali Soil Microbial Inoculum and Substrate Inoculation
2.4. Experimental Design and Salt/Silicon Treatments
2.5. Sampling Scheme
2.6. Growth Traits and Survival Assessment
2.7. Soil Physicochemical Properties (EC, pH, and Soil Ions)
2.8. Determination of Na⁺ and K⁺ in Plant Tissues
2.9. Oxidative Stress Markers (MDA, H₂O₂, and O₂·⁻)
2.10. Rhizosphere Microbiome Sampling, 16S rRNA Sequencing, and Bioinformatics
2.11. RNA Extraction, Library Construction, RNA-Seq Data Processing, and Heatmap Visualization
2.12. Statistical Analysis
3. Results
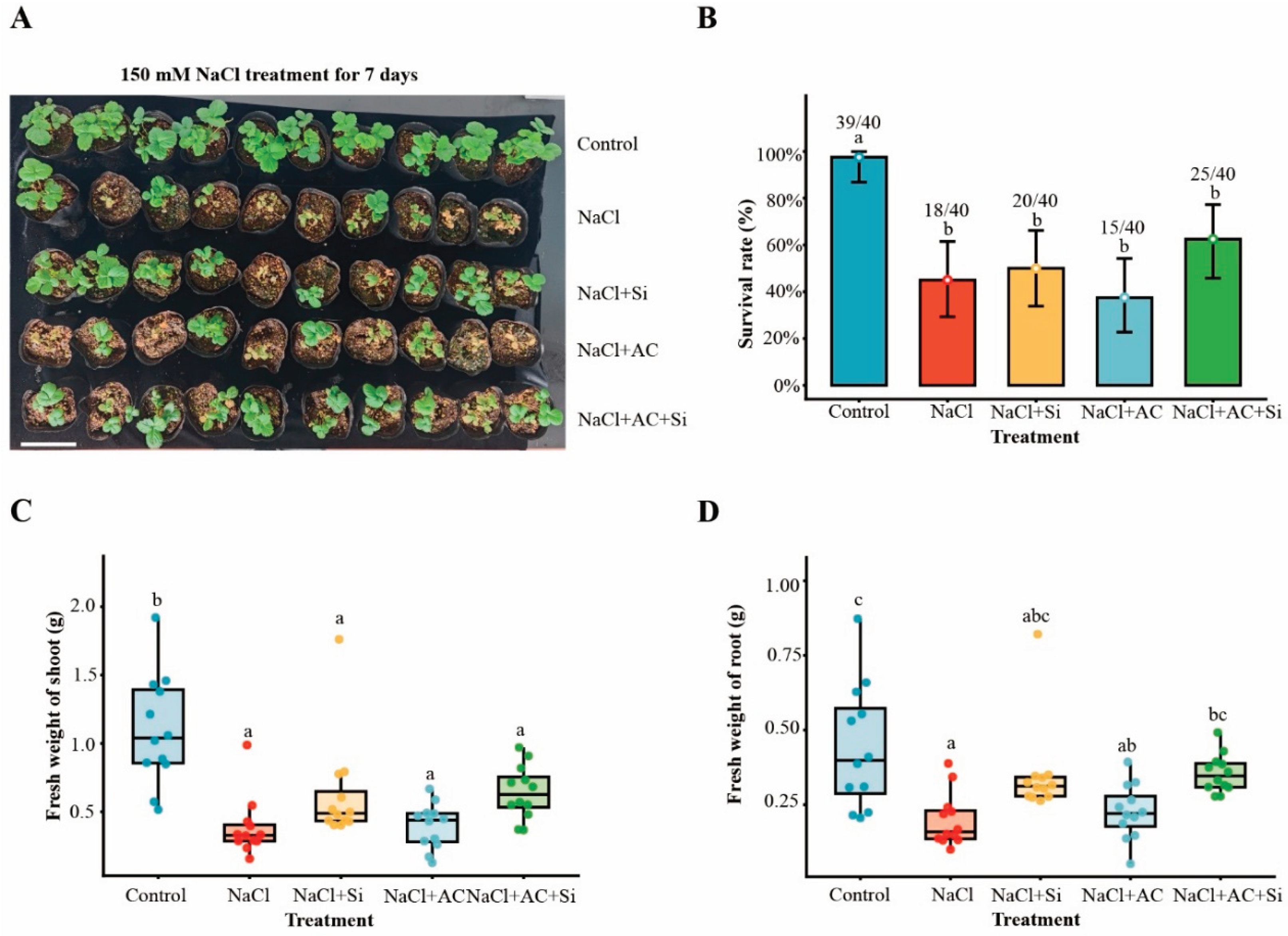
3.1. Co-Application of Activated Carbon and Silicon Mitigated Salt Injury and Improved Plant Performance
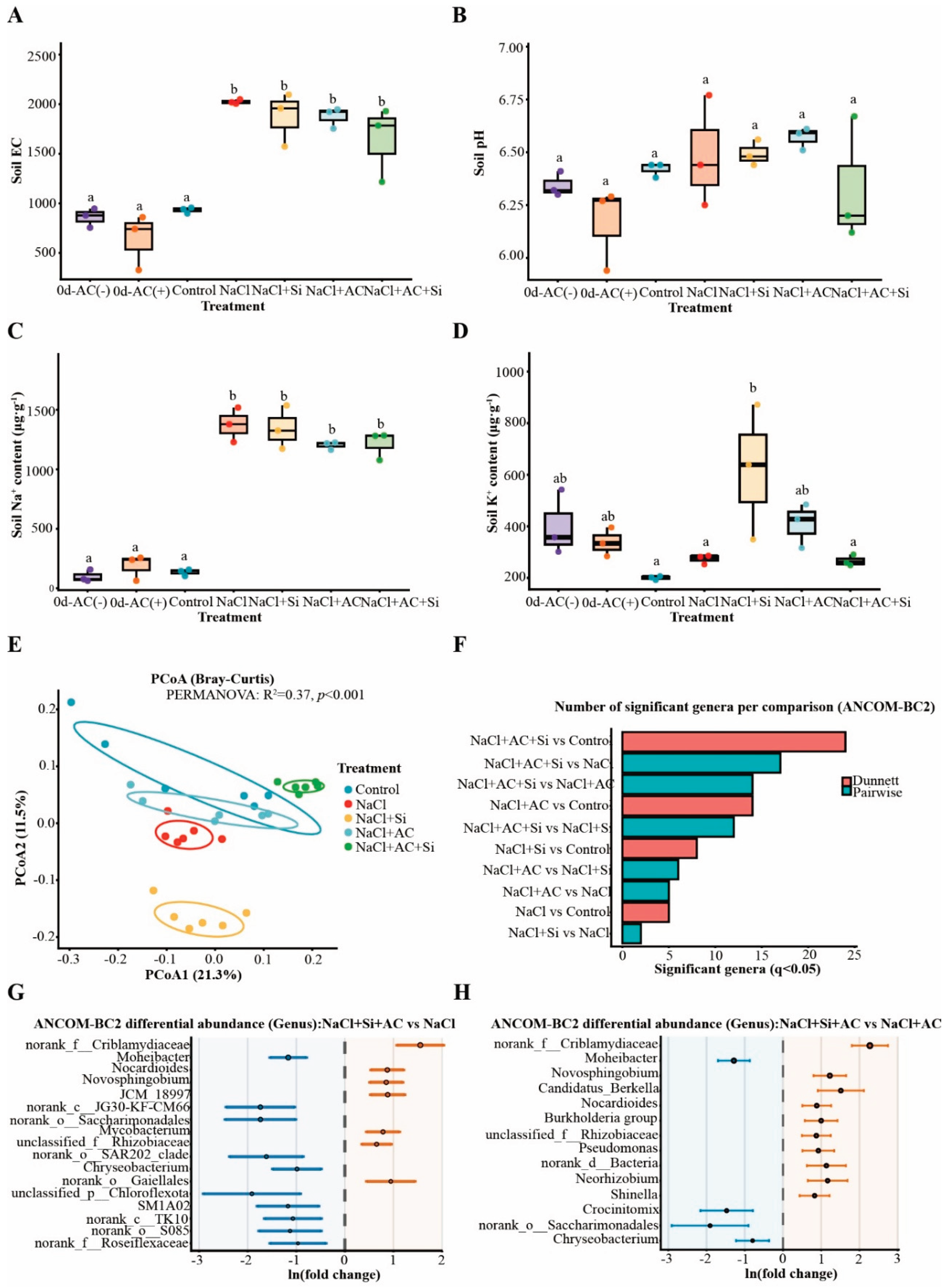
3.2. AC+Si Modulated Soil Properties and Reshaped Rhizosphere Bacterial Communities Under Salinity
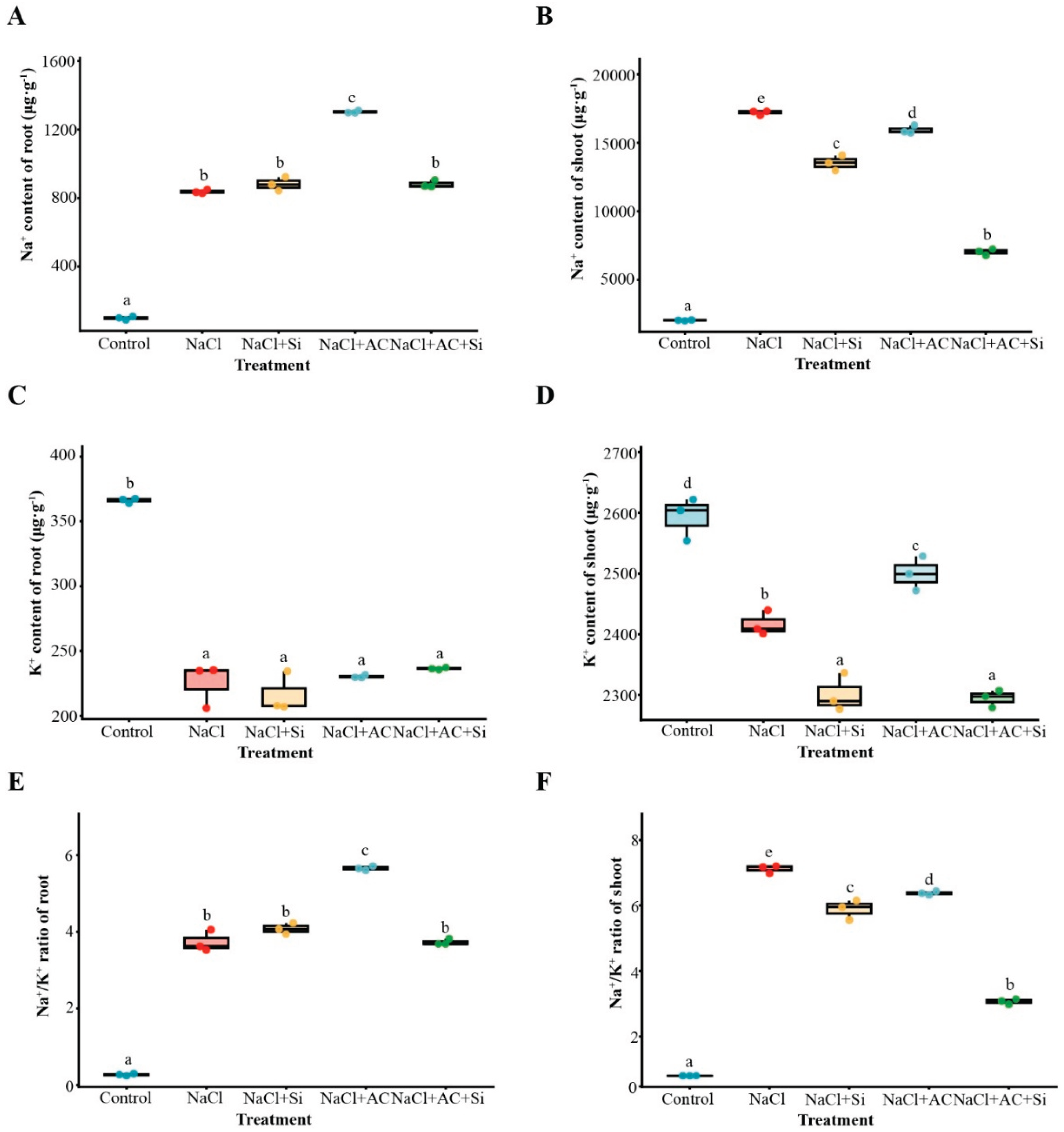
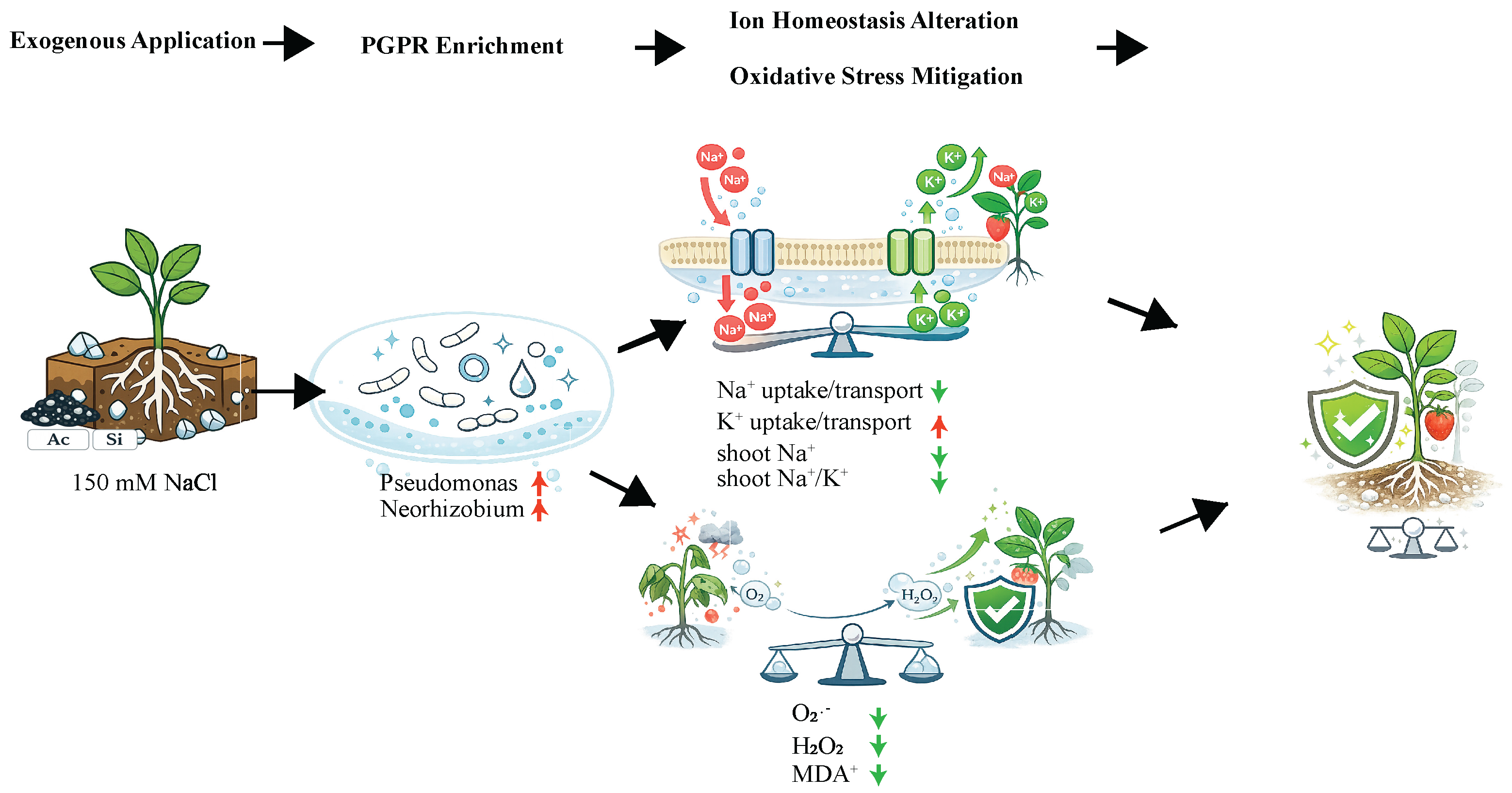
3.3. AC+Si Improved Ion Homeostasis Primarily by Reducing Shoot Na⁺ Accumulation and Lowering the Shoot Na⁺/K⁺ Ratio
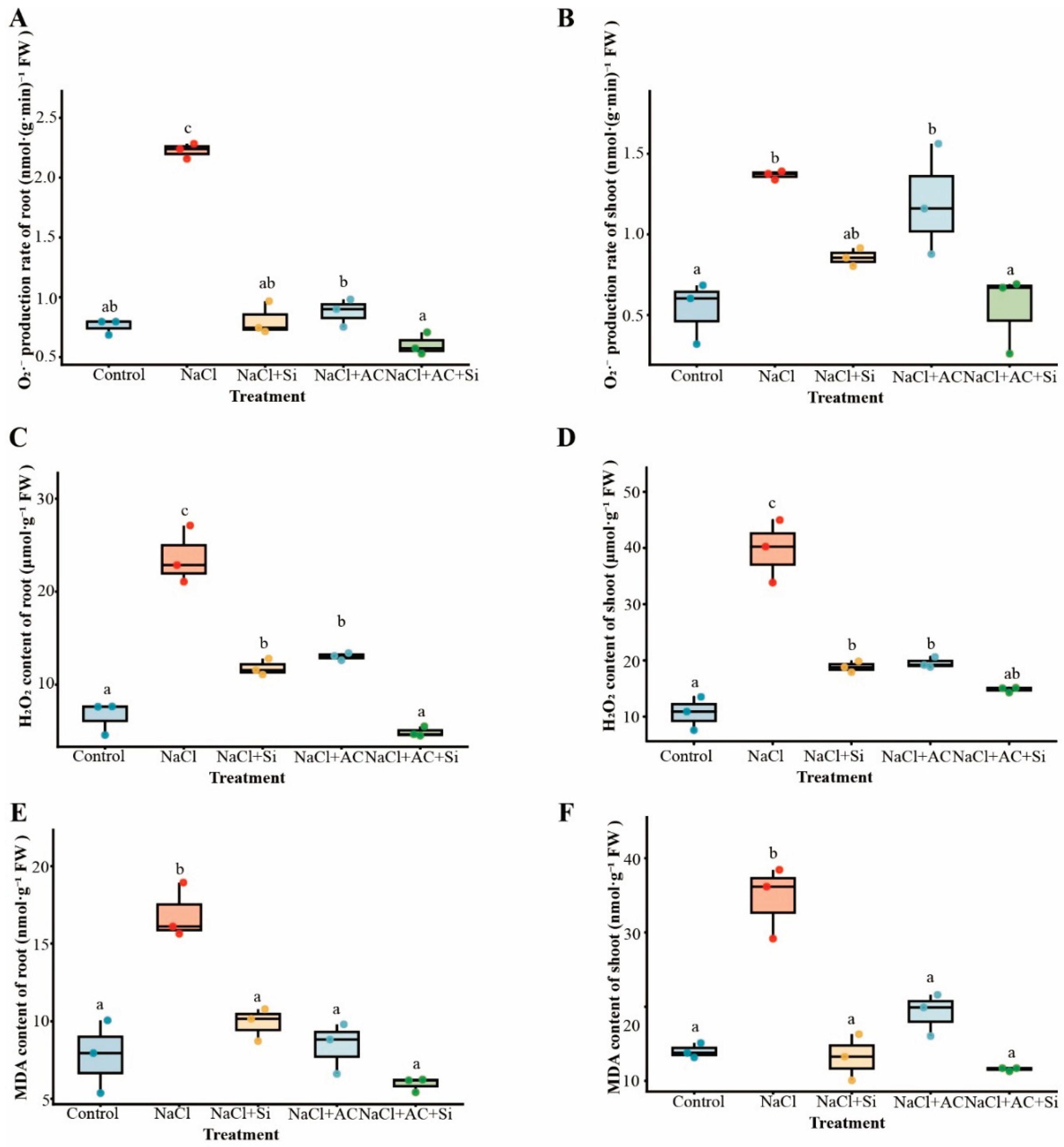
3.4. AC+Si Attenuated Salt-Induced Oxidative Damage in Roots and Shoots
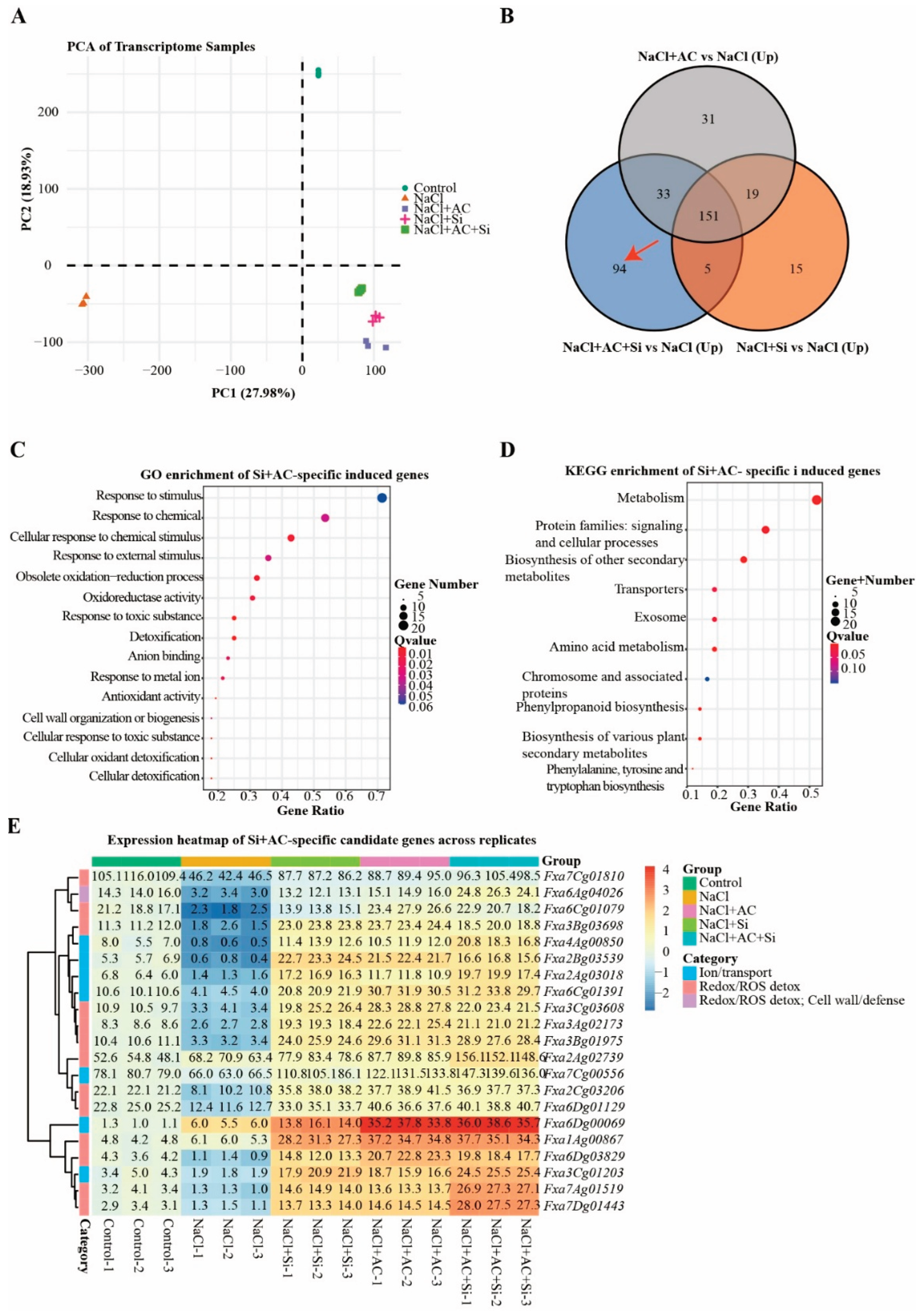
3.5. Transcriptome Profiling Supported Coordinated Induction of Ion-Transport and Redox/ROS-Mitigation Programs Under AC+Si
4. Discussion
4.1. A Non-Additive Response: Si Helps, AC Alone Is Weak, and Si+AC Is Most Effective
4.2. Ion Homeostasis and Redox Buffering: A Coherent Physiological Explanation
4.3. Rhizosphere Restructuring as an “Amplifier” of Plant Stress Tolerance
4.4. Transcriptome Insights and Agronomic Relevance
5. Conclusions
Supplementary Materials
Author Contributions
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Munns, R.; Tester, M. Mechanisms of salinity tolerance. Annu Rev Plant Biol 2008, 59, 651–681. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.K. Abiotic Stress Signaling and Responses in Plants. Cell 2016, 167, 313–324. [Google Scholar] [CrossRef]
- Negrao, S.; Schmockel, S.M.; Tester, M. Evaluating physiological responses of plants to salinity stress. Ann Bot 2017, 119, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Zhu, J.; Gong, Z.; Zhu, J.K. Abiotic stress responses in plants. Nat Rev Genet 2022, 23, 104–119. [Google Scholar] [CrossRef]
- Li, S.; Chang, L.; Sun, R.; Dong, J.; Zhong, C.; Gao, Y.; Zhang, H.; Wei, L.; Wei, Y.; Zhang, Y.; et al. Combined transcriptomic and metabolomic analysis reveals a role for adenosine triphosphate-binding cassette transporters and cell wall remodeling in response to salt stress in strawberry. Front Plant Sci 2022, 13, 996765. [Google Scholar] [CrossRef]
- Zahedi, S.M.; Abdelrahman, M.; Hosseini, M.S.; Hoveizeh, N.F.; Tran, L.P. Alleviation of the effect of salinity on growth and yield of strawberry by foliar spray of selenium-nanoparticles. Environ Pollut 2019, 253, 246–258. [Google Scholar] [CrossRef]
- Christou, A.; Manganaris, G.A.; Papadopoulos, I.; Fotopoulos, V. Hydrogen sulfide induces systemic tolerance to salinity and non-ionic osmotic stress in strawberry plants through modification of reactive species biosynthesis and transcriptional regulation of multiple defence pathways. J Exp Bot 2013, 64, 1953–1966. [Google Scholar] [CrossRef]
- Baral, A. Strawberries under salt stress: ALA and ROS to the rescue. Physiol Plant 2019, 167, 2–4. [Google Scholar] [CrossRef] [PubMed]
- Galli, V.; Messias, R.S.; Guzman, F.; Perin, E.C.; Margis, R.; Rombaldi, C.V. Transcriptome analysis of strawberry (Fragaria x ananassa) fruits under osmotic stresses and identification of genes related to ascorbic acid pathway. Physiol Plant 2019, 166, 979–995. [Google Scholar] [CrossRef] [PubMed]
- Rizwan, M.; Ali, S.; Ibrahim, M.; Farid, M.; Adrees, M.; Bharwana, S.A.; Zia-Ur-Rehman, M.; Qayyum, M.F.; Abbas, F. Mechanisms of silicon-mediated alleviation of drought and salt stress in plants: a review. Environ Sci Pollut Res Int 2015, 22, 15416–15431. [Google Scholar] [CrossRef] [PubMed]
- Rios, J.J.; Martinez-Ballesta, M.C.; Ruiz, J.M.; Blasco, B.; Carvajal, M. Silicon-mediated Improvement in Plant Salinity Tolerance: The Role of Aquaporins. Front Plant Sci 2017, 8, 948. [Google Scholar] [CrossRef] [PubMed]
- Kurowska, M.M. Aquaporins in Cereals-Important Players in Maintaining Cell Homeostasis under Abiotic Stress. Genes (Basel) 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.X.; Xu, X.B.; Hu, Y.H.; Han, W.H.; Yin, J.L.; Li, H.L.; Gong, H.J. Silicon improves salt tolerance by increasing root water uptake in Cucumis sativus L. Plant Cell Rep 2015, 34, 1629–1646. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.; Rizwan, M.; Qayyum, M.F.; Ok, Y.S.; Ibrahim, M.; Riaz, M.; Arif, M.S.; Hafeez, F.; Al-Wabel, M.I.; Shahzad, A.N. Biochar soil amendment on alleviation of drought and salt stress in plants: a critical review. Environ Sci Pollut Res Int 2017, 24, 12700–12712. [Google Scholar] [CrossRef] [PubMed]
- Poveda, J.; Martinez-Gomez, A.; Fenoll, C.; Escobar, C. The Use of Biochar for Plant Pathogen Control. Phytopathology 2021, 111, 1490–1499. [Google Scholar] [CrossRef]
- Berendsen, R.L.; Pieterse, C.M.; Bakker, P.A. The rhizosphere microbiome and plant health. Trends Plant Sci 2012, 17, 478–486. [Google Scholar] [CrossRef] [PubMed]
- Edwards, J.; Johnson, C.; Santos-Medellin, C.; Lurie, E.; Podishetty, N.K.; Bhatnagar, S.; Eisen, J.A.; Sundaresan, V. Structure, variation, and assembly of the root-associated microbiomes of rice. Proc Natl Acad Sci U S A 2015, 112, E911-920. [Google Scholar] [CrossRef]
- Pieterse, C.M.; Zamioudis, C.; Berendsen, R.L.; Weller, D.M.; Van Wees, S.C.; Bakker, P.A. Induced systemic resistance by beneficial microbes. Annu Rev Phytopathol 2014, 52, 347–375. [Google Scholar] [CrossRef]
- Bulgarelli, D.; Schlaeppi, K.; Spaepen, S.; Ver Loren van Themaat, E.; Schulze-Lefert, P. Structure and functions of the bacterial microbiota of plants. Annu Rev Plant Biol 2013, 64, 807–838. [Google Scholar] [CrossRef] [PubMed]
- Souza, R.; Ambrosini, A.; Passaglia, L.M. Plant growth-promoting bacteria as inoculants in agricultural soils. Genet Mol Biol 2015, 38, 401–419. [Google Scholar] [CrossRef]
- Glick, B.R. Plant growth-promoting bacteria: mechanisms and applications. Scientifica (Cairo) 2012, 2012, 963401. [Google Scholar] [CrossRef]
- Orozco-Mosqueda, M.D.C.; Glick, B.R.; Santoyo, G. ACC deaminase in plant growth-promoting bacteria (PGPB): An efficient mechanism to counter salt stress in crops. Microbiol Res 2020, 235, 126439. [Google Scholar] [CrossRef]
- Glick, B.R. Modulation of plant ethylene levels by the bacterial enzyme ACC deaminase. FEMS Microbiol Lett 2005, 251, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Glick, B.R. Bacteria with ACC deaminase can promote plant growth and help to feed the world. Microbiol Res 2014, 169, 30–39. [Google Scholar] [CrossRef]
- Gamalero, E.; Glick, B.R. Recent Advances in Bacterial Amelioration of Plant Drought and Salt Stress. Biology (Basel) 2022, 11. [Google Scholar] [CrossRef]
- Malik, L.; Sanaullah, M.; Mahmood, F.; Hussain, S.; Siddique, M.H.; Anwar, F.; Shahzad, T. Unlocking the potential of co-applied biochar and plant growth-promoting rhizobacteria (PGPR) for sustainable agriculture under stress conditions. Chem Biol Technol Agric 2022, 9, 58. [Google Scholar] [CrossRef] [PubMed]
- Tu, C.; Wei, J.; Guan, F.; Liu, Y.; Sun, Y.; Luo, Y. Biochar and bacteria inoculated biochar enhanced Cd and Cu immobilization and enzymatic activity in a polluted soil. Environ Int 2020, 137, 105576. [Google Scholar] [CrossRef] [PubMed]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 2019, 37, 852–857. [Google Scholar] [CrossRef] [PubMed]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 2016, 13, 581–583. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glockner, F.O. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 2013, 41, D590–596. [Google Scholar] [CrossRef] [PubMed]
- McMurdie, P.J.; Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 2013, 8, e61217. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.; Peddada, S.D. Analysis of compositions of microbiomes with bias correction. Nat Commun 2020, 11, 3514. [Google Scholar] [CrossRef]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Peng, Y.; Liu, L.; Li, G.; Zhao, X.; Wang, X.; Cao, S.; Muyle, A.; Zhou, Y.; Zhou, H. Phased gap-free genome assembly of octoploid cultivated strawberry illustrates the genetic and epigenetic divergence among subgenomes. Hortic Res 2024, 11, uhad252. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: a fast spliced aligner with low memory requirements. Nat Methods 2015, 12, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; Genome Project Data Processing, S. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.C.; Mendell, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol 2015, 33, 290–295. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 2014, 15, 550. [Google Scholar] [CrossRef]
- Chen, C.; Wu, Y.; Li, J.; Wang, X.; Zeng, Z.; Xu, J.; Liu, Y.; Feng, J.; Chen, H.; He, Y.; et al. TBtools-II: A "one for all, all for one" bioinformatics platform for biological big-data mining. Mol Plant 2023, 16, 1733–1742. [Google Scholar] [CrossRef] [PubMed]
- Hothorn, T.; Bretz, F.; Westfall, P. Simultaneous inference in general parametric models. Biom J 2008, 50, 346–363. [Google Scholar] [CrossRef]
- Hasegawa, P.M.; Bressan, R.A.; Zhu, J.K.; Bohnert, H.J. Plant Cellular and Molecular Responses to High Salinity. Annu Rev Plant Physiol Plant Mol Biol 2000, 51, 463–499. [Google Scholar] [CrossRef]
- Zhu, J.K. Plant salt tolerance. Trends Plant Sci 2001, 6, 66–71. [Google Scholar] [CrossRef]
- Ma, J.F.; Yamaji, N. Silicon uptake and accumulation in higher plants. Trends Plant Sci 2006, 11, 392–397. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.X.; Gong, H.J.; Yin, J.L. Role of Silicon in Mediating Salt Tolerance in Plants: A Review. Plants (Basel) 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Coskun, D.; Deshmukh, R.; Sonah, H.; Menzies, J.G.; Reynolds, O.; Ma, J.F.; Kronzucker, H.J.; Belanger, R.R. The controversies of silicon's role in plant biology. New Phytol 2019, 221, 67–85. [Google Scholar] [CrossRef]
- Wu, Y.; Wang, X.; Zhang, L.; Zheng, Y.; Liu, X.; Zhang, Y. The critical role of biochar to mitigate the adverse impacts of drought and salinity stress in plants. Front Plant Sci 2023, 14, 1163451. [Google Scholar] [CrossRef]
- Apel, K.; Hirt, H. Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 2004, 55, 373–399. [Google Scholar] [CrossRef]
- Foyer, C.H.; Noctor, G. Redox homeostasis and antioxidant signaling: a metabolic interface between stress perception and physiological responses. Plant Cell 2005, 17, 1866–1875. [Google Scholar] [CrossRef] [PubMed]
- Gill, S.S.; Tuteja, N. Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 2010, 48, 909–930. [Google Scholar] [CrossRef]
- Mendes, R.; Garbeva, P.; Raaijmakers, J.M. The rhizosphere microbiome: significance of plant beneficial, plant pathogenic, and human pathogenic microorganisms. FEMS Microbiol Rev 2013, 37, 634–663. [Google Scholar] [CrossRef]
- Yang, J.; Kloepper, J.W.; Ryu, C.M. Rhizosphere bacteria help plants tolerate abiotic stress. Trends Plant Sci 2009, 14, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Etesami, H.; Beattie, G.A. Mining Halophytes for Plant Growth-Promoting Halotolerant Bacteria to Enhance the Salinity Tolerance of Non-halophytic Crops. Front Microbiol 2018, 9, 148. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



