Submitted:
25 February 2026
Posted:
28 February 2026
You are already at the latest version
Abstract
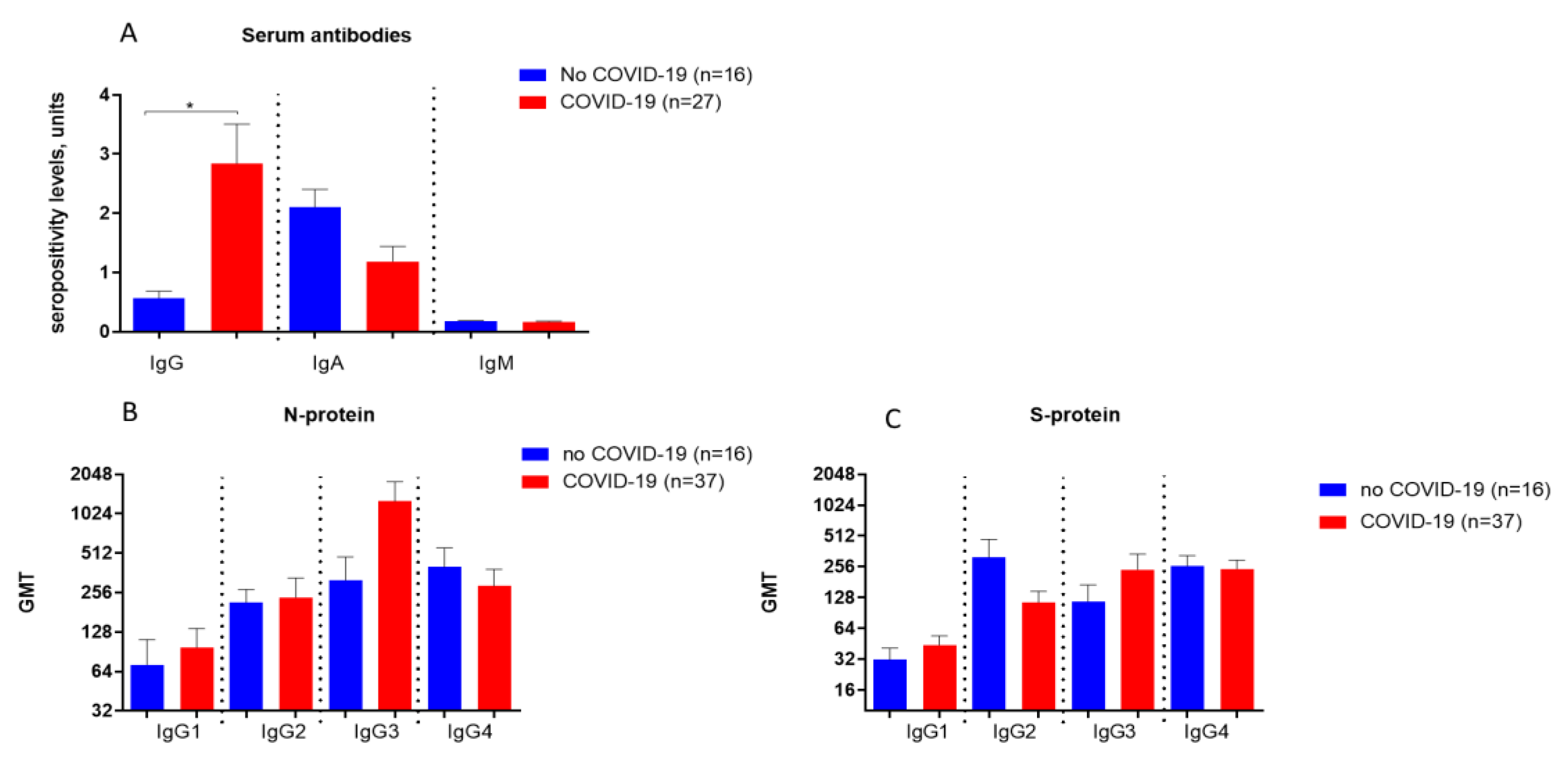
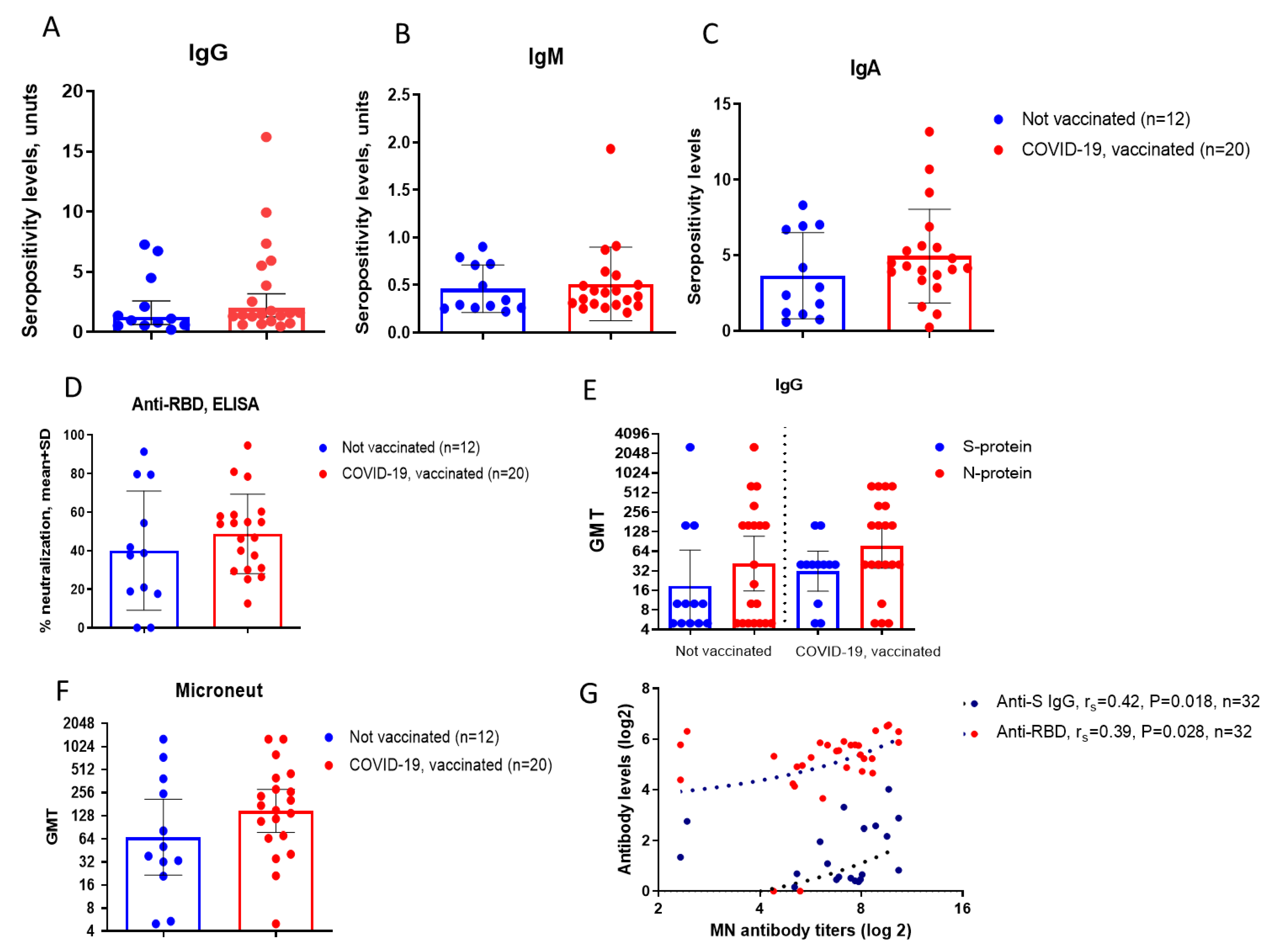
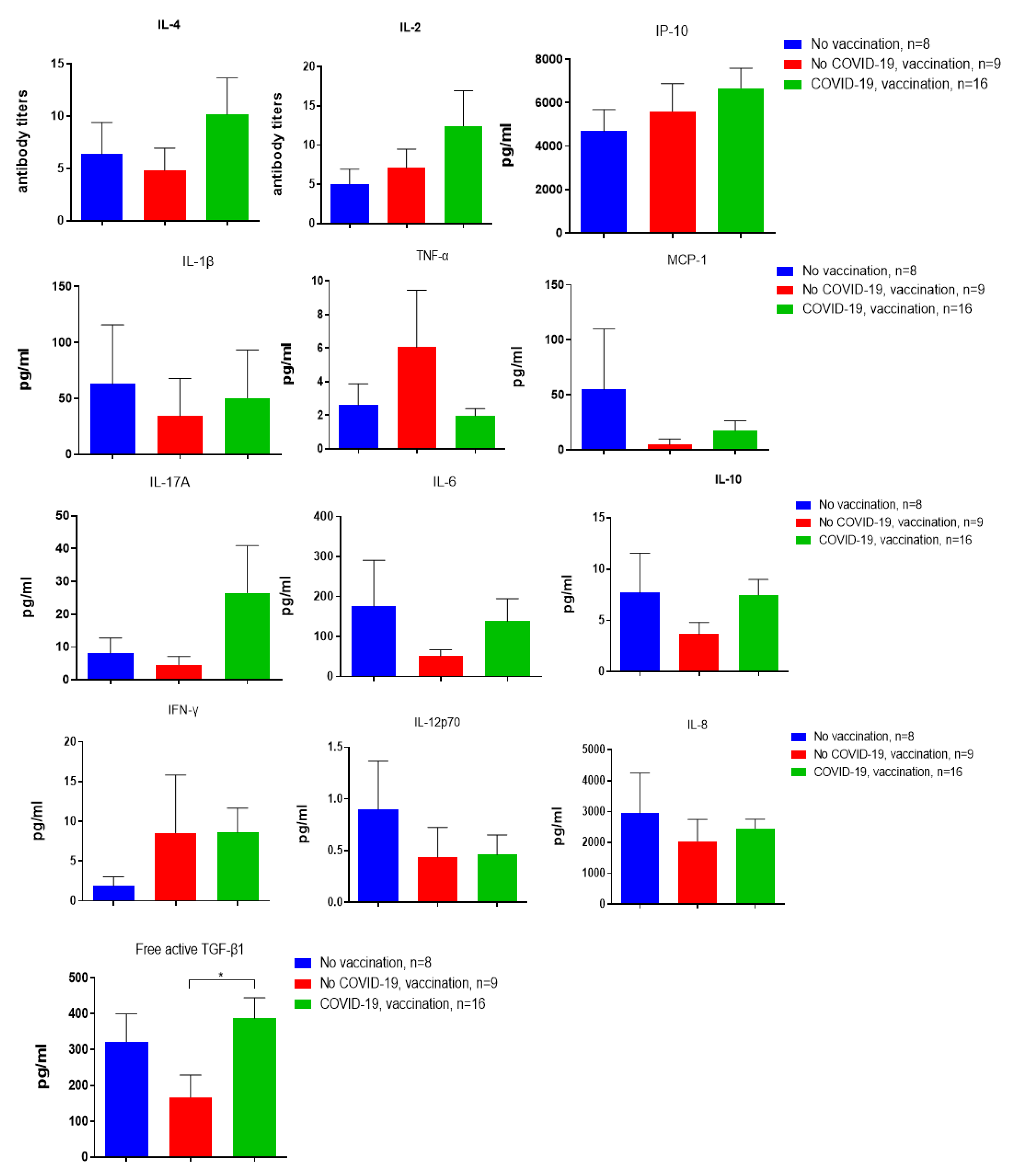
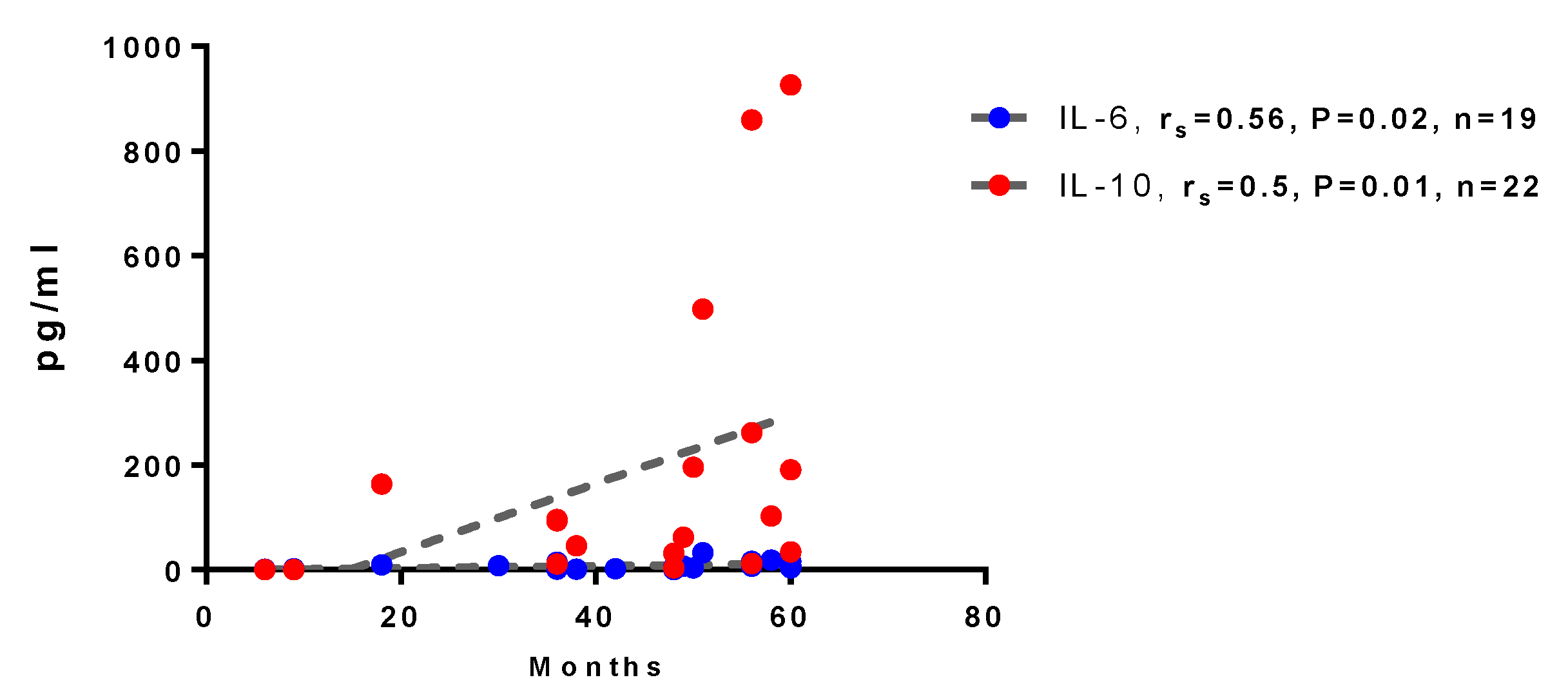
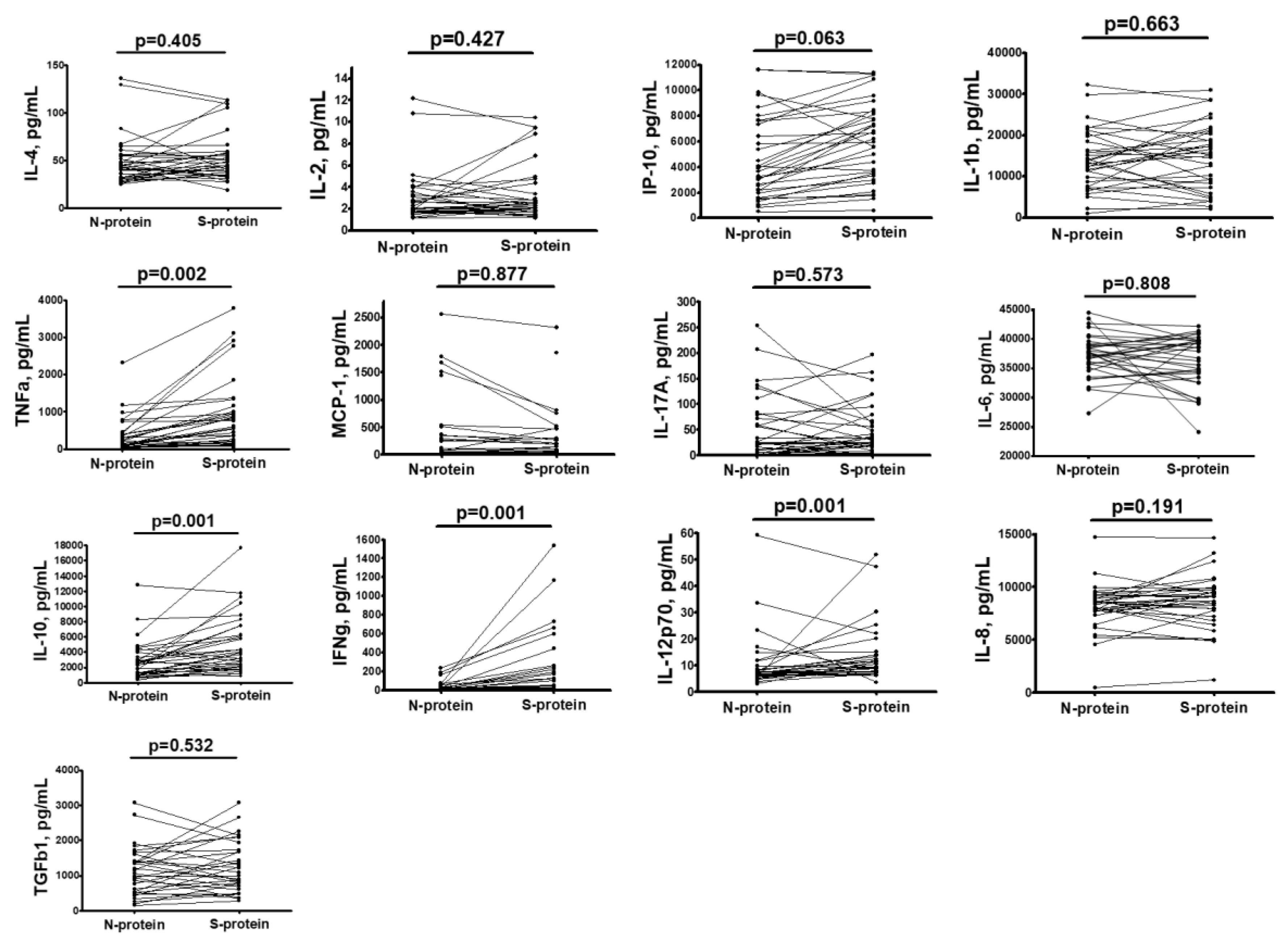
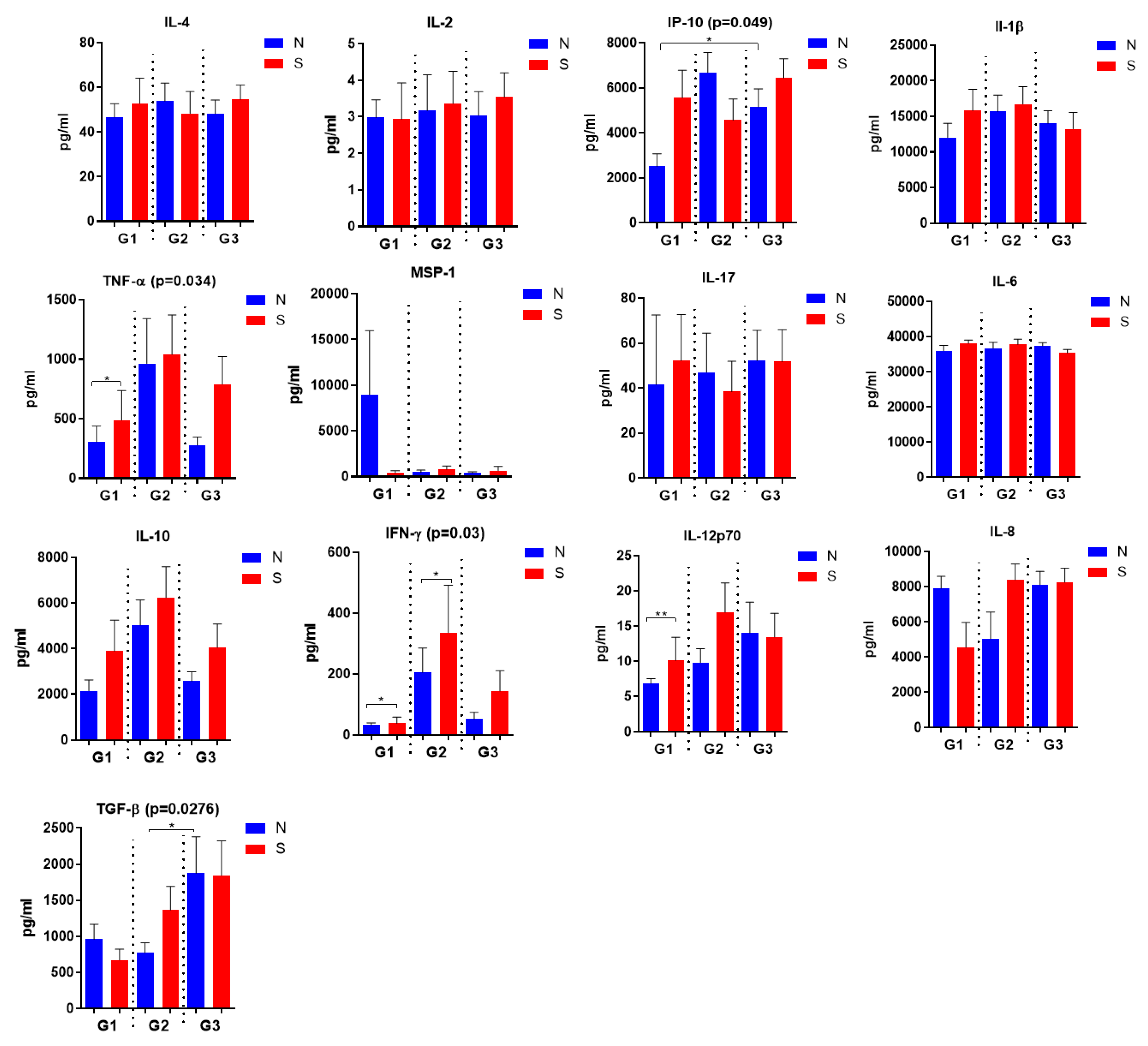
Background/Objective: This study Investigates long-term immune responses (up to 4–4.5 years) to SARS-CoV-2 in individuals with: Past COVID-19 infection, Vaccination, Combined exposure, with focus on immune reactivity to recombinant spike (S) and nucleocapsid (N) proteins. Materials and methods: Serum antibody responses were assessed up to 4–4.5 years after infection or immunization, including virus-specific IgG, IgA, IgM, and neutralizing antibodies. Cellular immunity was evaluated by phenotypic and functional analysis of peripheral blood mononuclear cells (PBMCs), including memory T-helper and cytotoxic T-cell subsets, as well as cytokine production following in vitro stimulation with recombinant SARS-CoV-2 proteins. Multiplex cytokine profiling was used to characterize effector and regulatory immune responses. Results: Virus-specific IgG antibodies persisted for several years after SARS-CoV-2 exposure, with anti-RBD IgG showing the strongest correlation with virus-neutralizing activity, whereas antibodies to the N- protein primarily reflected prior infection. Vaccinated individuals exhibited a distinct immunoglobulin profile characterized by a higher prevalence of IgA. No IgM detected suggesting the detected immune responses reflect immunological memory rather than active infection. PBMCs from individuals with combined COVID-19 and vaccination history demonstrated enhanced responsiveness and more memory T cells. Hybrid immunity (infection and vaccination) provides stronger and broader immune responses.. Stimulation with S- protein induces stronger cytokine production (IFN-γ, TNF-α, IL-12p70) than N- protein. Regulatory cytokines (IL-10, TGF-β) also elevated which suggests immune regulation rather than chronic inflammation.Conclusion: SARS-CoV-2 infection and vaccination induce persistent humoral and cellular immunity. Neutralizing activity correlates only with anti-RBD and anti-S IgG. Future research should focus on long-term effects, hybrid immunity, and optimizing other vaccine types, in addition to Adenovector vaccines, such as recombinant antigen-based vaccines.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Reagents
2.2. Bioinformatics Analysis
2.3. Patients
2.4. Vaccine
2.5. Obtaining Blood Serum Samples
2.6. Determination of the Level of Antibodies to SARS-CoV-2
2.7. Concurrent Microneutralization Assay
2.8. Microneutralization (MN) Assay in Cell Culture
2.9. Recombinant Proteins of SARS-CoV-2
2.10. Enzyme-Linked Immunosorbent Assay (ELISA) for the Determination of IgG Subclasses in Blood Serum Using Recombinant Peptides
2.11. Obtaining and Incubating PBMCs
2.12. Stimulation of PBMC with Recombinant SARS-CoV-2 Virus Proteins
2.13. The T Cell Antigen-Specific Immune Response
2.14. Determination of Cytokine Concentrations
2.15. Statistical Analysis
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| COVID-19 | coronavirus disease 19 |
| IFN-γ | interferon-γ |
| IL | interleukin |
| MCP-1 | monocyte chemotactic protein |
| Me | medians |
| N- protein | Nucleocapsid coronavirus protein |
| Q1; Q3 | lower and upper quartiles |
| PBMCs | peripheral mononuclear cells |
| SARS-CoV-2 | severe acute respiratory syndrome coronavirus 2 |
| S- protein | Spike coronavirus protein |
| TGF-β | transforming growth factor-beta |
| TNF-α | tumor necrosis factor alpha |
References
- Kovalenko, A.; Ryabchevskaya, E.; Evtushenko, E.; Nikitin, N.; Karpova, O. Recombinant protein vaccines against human betacoronaviruses: strategies, approaches and progress. International Journal of Molecular Sciences 2023, 24(2), 1701. [Google Scholar] [CrossRef]
- Asadbeigi, A.; Razavirad, A.; Khajehahmadi, F.; Eghtedari, M.; Shirkoohi, R. Expanding SARS-CoV-2 antigen targets beyond spike using a baculovirus expression system. Scientific Reports 2026. [Google Scholar] [CrossRef]
- Claro, F.; Silva, D.; Bogado, J. A. P.; Rangel, H. R.; de Waard, J. H. Lasting SARS-CoV-2 specific IgG Antibody response in health care workers from Venezuela, 6 months after vaccination with Sputnik V. International Journal of Infectious Diseases 2022, 122, 850–854. [Google Scholar] [CrossRef] [PubMed]
- Sette, A.; Crotty, S. Immunological memory to SARS-CoV-2 infection and COVID-19 vaccines. Immunological reviews 2022, 310(1), 27–46. [Google Scholar] [CrossRef]
- Cohen, K. W.; Linderman, S. L.; Moodie, Z.; Czartoski, J.; Lai, L.; Mantus, G.; McElrath, M. J. Longitudinal analysis shows durable and broad immune memory after SARS-CoV-2 infection with persisting antibody responses and memory B and T cells. Cell Reports Medicine 2021, 2(7). [Google Scholar] [CrossRef]
- Yang, Y.; Du, L. SARS-CoV-2 spike protein: a key target for eliciting persistent neutralizing antibodies. Signal transduction and targeted therapy 2021, 6(1), 95. [Google Scholar] [CrossRef]
- Chen, B.; Farzan, M.; Choe, H. SARS-CoV-2 spike protein: structure, viral entry and variants. Nature Reviews Microbiology 2025, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Magalhaes, B. D. A. P.; Medeiros Minasi, J.; Lobato, R. C.; Lemos, L. C.; de Britto, L. S.; Barros, R. M.; da Hora, V. P. Globally approved vaccines for COVID-19: a systematic review. Brazilian Journal of Microbiology 2025, 56(1), 511–527. [Google Scholar] [CrossRef] [PubMed]
- Edalat, F.; Khakpour, N.; Heli, H.; Letafati, A.; Ramezani, A.; Hosseini, S. Y.; Moattari, A. Immunological mechanisms of the nucleocapsid protein in COVID-19. Scientific Reports 2024, 14(1), 3711. [Google Scholar] [CrossRef]
- López-Muñoz, A. D.; Kosik, I.; Holly, J.; Yewdell, J. W. Cell surface SARS-CoV-2 nucleocapsid protein modulates innate and adaptive immunity. Science Advances 2022, 8(31), eabp9770. [Google Scholar] [CrossRef]
- Duysburgh, E.; Mortgat, L.; Barbezange, C.; Dierick, K.; Fischer, N.; Heyndrickx, L.; Desombere, I. Persistence of IgG response to SARS-CoV-2. The Lancet Infectious Diseases 2021, 21(2), 163–164. [Google Scholar] [CrossRef] [PubMed]
- Feng, C.; Shi, J.; Fan, Q.; Wang, Y.; Huang, H.; Chen, F.; Li, F. Protective humoral and cellular immune responses to SARS-CoV-2 persist up to 1 year after recovery. Nature communications 2021, 12(1), 4984. [Google Scholar] [CrossRef] [PubMed]
- Matei, L.; Chivu-Economescu, M.; Dragu, L. D.; Grancea, C.; Bleotu, C.; Hrișcă, R.; Ruţă, S. M. Long-term durability and variant-specific modulation of SARS-CoV-2 humoral and cellular immunity over two years. International Journal of Molecular Sciences 2025, 26(16), 8106. [Google Scholar] [CrossRef]
- Okonechnikov, K.; Golosova, O.; Fursov, M.; Ugene Team. Unipro UGENE: a unified bioinformatics toolkit. Bioinformatics 2012, 28(8), 1166–1167. [Google Scholar] [CrossRef] [PubMed]
- Andreatta, M; Nielsen, M. Gapped sequence alignment using artificial neural networks: application to the MHC class I system. Bioinformatics 2016, 32(4), 511–7. [Google Scholar] [CrossRef]
- Nielsen, M; Lundegaard, C; Worning, P; Lauemoller, SL; Lamberth, K; Buus, S; Brunak, S; Lund, O. Reliable prediction of T-cell epitopes using neural networks with novel sequence representations. Protein Sci 2003, 12, 1007–17. [Google Scholar] [CrossRef]
- Abramson, J.; Adler, J.; Dunger, J.; Evans, R.; Green, T.; Pritzel, A.; Jumper, J. M. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 2024, 630(8016), 493–500. [Google Scholar] [CrossRef]
- Barchuk, A.; Bulina, A.; Cherkashin, M.; Berezina, N.; Rakova, T.; Kuplevatskaya, D.; Okhotin, A. Gam-COVID-Vac, EpiVacCorona, and CoviVac effectiveness against lung injury during Delta and Omicron variant surges in St. Petersburg, Russia: a test-negative case–control study. Respiratory research 2022, 23(1), 276. [Google Scholar] [CrossRef]
- Matyushenko, V.; Isakova-Sivak, I.; Kudryavtsev, I.; Goshina, A.; Chistyakova, A.; Stepanova, E.; Rudenko, L. Detection of IFNγ-secreting CD4+ and CD8+ memory T cells in COVID-19 convalescents after stimulation of peripheral blood mononuclear cells with live SARS-CoV-2. Viruses 2021, 13(8), 1490. [Google Scholar] [CrossRef]
- Rak, A.; Bazhenova, E.; Prokopenko, P.; Matyushenko, V.; Orshanskaya, Y.; Sivak, K. V.; Isakova-Sivak, I. Protective Potential and Functional Role of Antibodies Against SARS-CoV-2 Nucleocapsid Protein. Antibodies 2025, 14(2), 45. [Google Scholar] [CrossRef]
- Gupalova, T. V.; Bormotova, E. A.; Leontieva, G. F.; Kramskaya, T. A.; Desheva, Y. A.; Suvorov, A. N. Production of Recombinant Nucleocapsid Protein SARSN1 and Its Use for Assessing the Immune Response in Mice after Vaccination with a Live Probiotic Vaccine Containing the Coronavirus Nucleocapsid Protein. Bulletin of Experimental Biology and Medicine 2025, 179(3), 322–325. [Google Scholar] [CrossRef]
- Leontieva, G.; Kramskaya, T.; Gupalova, T.; Bormotova, E.; Desheva, Y.; Korzhevsky, D.; Suvorov, A. Comparative Efficacy of Parenteral and Mucosal Recombinant Probiotic Vaccines Against SARS-CoV-2 and S. pneumoniae Infections in Animal Models. Vaccines 2024, 12(10), 1195. [Google Scholar] [CrossRef]
- Desheva, Y.; Muruzheva, Z.; Tumashova, O.; Shvedova, T.; Kudar, P.; Matveev, G.; Suvorov, A. Cytokine profiles and neurological manifestations in post-COVID syndrome. Exploration of Immunology 2025, 5, 1003184. [Google Scholar] [CrossRef]
- Naĭkhin, A. N.; Rekstin, A. R.; Barantseva, I. B.; Donina, S. A.; Grigor'eva, E. P.; Kiseleva, I. V.; Rudenko, L. G. Immune response to live influenza vaccine. Vestnik Rossiiskoi Akademii Meditsinskikh Nauk 2002, (12), 24–28. [Google Scholar]
- Kuklina, E. M. T lymphocytes as targets for SARS-CoV-2. Biochemistry (Moscow) 2022, 87(6), 566–576. [Google Scholar] [CrossRef]
- Andreatta, M.; Nielsen, M. Gapped sequence alignment using artificial neural networks: application to the MHC class I system. Bioinformatics 2016, 32(4), 511–517. [Google Scholar] [CrossRef] [PubMed]
- Delgado, J. F.; Vidal-Pla, M.; Moya, M. C.; Espasa, M.; Casabella, A.; Seda, M.; Peña, P. SARS-CoV-2 spike protein vaccine-induced immune imprinting reduces nucleocapsid protein antibody response in SARS-CoV-2 infection. Journal of Immunology Research 2022, 2022(1), 8287087. [Google Scholar] [CrossRef]
- Gaeta, A.; Angeloni, A.; Napoli, A.; Pucci, B.; Cinti, L.; Roberto, P.; Anastasi, E. Anti-N SARS-CoV-2 assays for evaluation of natural viral infection. Journal of Immunological Methods 2023, 518, 113486. [Google Scholar] [CrossRef] [PubMed]
- Turner, J. S.; Kim, W.; Kalaidina, E.; Goss, C. W.; Rauseo, A. M.; Schmitz, A. J.; Ellebedy, A. H. SARS-CoV-2 infection induces long-lived bone marrow plasma cells in humans. Nature 2021, 595(7867), 421–425. [Google Scholar] [CrossRef]
- Liu, J.; Lian, R.; Zhang, G.; Hou, B.; Wang, C.; Dong, J.; Ye, T. Changes in serum virus-specific IgM/IgG antibody in asymptomatic and discharged patients with reoccurring positive COVID-19 nucleic acid test (RPNAT). Annals of Medicine 2021, 53(1), 34–42. [Google Scholar] [CrossRef] [PubMed]
- Moss, P. The T cell immune response against SARS-CoV-2. Nature immunology 2022, 23(2), 186–193. [Google Scholar] [CrossRef]
- Zivancevic-Simonovic, S.; Jovanovic, D.; Cupurdija, V.; Milosevic-Djordjevic, O.; Stanojevic, M.; Marinkovic, M.; Mihaljevic, O. Cytokine producing ability of peripheral blood cells from COVID-19 patients after unspecific in vitro stimulation. Inflammation research 2022, 71(3), 331–341. [Google Scholar] [CrossRef]
- Borowska, M.; Winiarska, H.; Dworacka, M.; Wesołowska, A.; Dworacki, G.; Mikołajczak, P. Ł. The effect of homocysteine on the secretion of IL-1β, IL-6, IL-10, IL-12 and RANTES by peripheral blood mononuclear cells—an in vitro study. Molecules 2021, 26(21), 6671. [Google Scholar] [CrossRef] [PubMed]
- Gupta, G.; Shareef, I.; Tomar, S.; Kumar, M. S. N.; Pandey, S.; Sarda, R.; Sinha, S. Th1/Th2/Th17 cytokine profile among different stages of COVID-19 infection. National Academy Science Letters 2022, 45(4), 363–369. [Google Scholar] [CrossRef]
- Savchenko, A. A.; Kudryavtsev, I. V.; Isakov, D. V.; Sadowski, I. S.; Belenyuk, V. D.; Borisov, A. G. Recombinant human interleukin-2 corrects NK cell phenotype and functional activity in patients with post-COVID syndrome. Pharmaceuticals 2023, 16(4), 537. [Google Scholar] [CrossRef] [PubMed]
- Miteva, L.; Stanilova, S. The combined effect of interleukin (IL)-10 and IL-12 polymorphisms on induced cytokine production. Human immunology 2008, 69(9), 562–566. [Google Scholar] [CrossRef]
- Alfaro, E.; Casitas, R.; Díaz-García, E.; García-Tovar, S.; Galera, R.; Torres-Vargas, M.; Cubillos-Zapata, C. TGF-β1 overexpression in severe COVID-19 survivors and its implications for early-phase fibrotic abnormalities and long-term functional impairment. Frontiers in Immunology 2024, 15, 1401015. [Google Scholar] [CrossRef] [PubMed]
- Ferreira-Gomes, Marta; et al. SARS-CoV-2 in severe COVID-19 induces a TGF-β-dominated chronic immune response that does not target itself. Nature communications 2021, 12.1, 1961. [Google Scholar] [CrossRef]
- Tao, T.; Tian, L.; Ke, J.; Zhang, C.; Li, M.; Xu, X.; Fan, H. Antibody-dependent enhancement of coronaviruses. International Journal of Biological Sciences 2025, 21(4), 1686. [Google Scholar] [CrossRef]
- Boris, Nikolaev. Nano- and Microemulsions in Biomedicine: From Theory to Practice. Pharmaceutics 2023, 15(7), 1989. [Google Scholar] [CrossRef]
| Name | Analysis of epitopes compatible with human MHC | Spatial structure |
|---|---|---|
| SARSN1 |  |
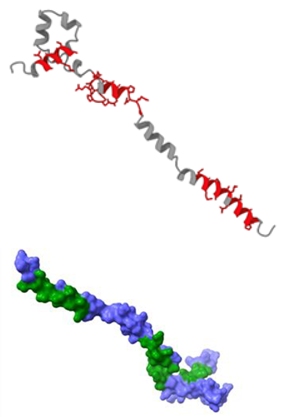 |
| S-SARS "XBB.1.5" |  |
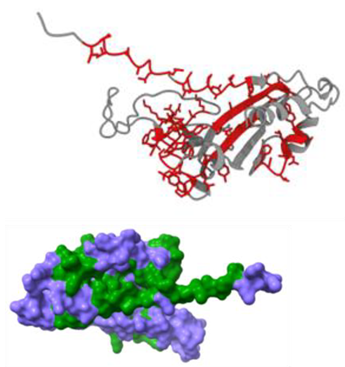 |
| No. | Epitope (location in the amino acid sequence of a protein) | Allele |
|---|---|---|
| 1 | LPTAAAGLL (5-13) | HLA-B0702 |
| 2 | LAAQPAMAM (15-23) | HLA-B0702 |
| 3 | TRNPANNAA (45-53) | HLA-B3901 |
| 4 | LQLPQGTTL (56-64) | HLA-B3901 |
| 5 | SSMKLAAAL (98-106) | HLA-B3901 |
| No. | Epitope (location in the amino acid sequence of a protein) | Allele |
|---|---|---|
| 1 | LPTAAAGLL (5-13) | HLA-B0702 |
| 2 | LAAQPAMAM (15-23) | HLA-B0702 |
| 3 | YSVLNSTSF (33-41) | HLA-B5801, HLA-B1501 |
| 4 | FTNVYADSF (59-67) | HLA-B5801 |
| 5 | VSGNYNYLY (112-120) | HLA-A0101 |
| 6 | NYNYLYRLF (115-123) | HLA-A2402 |
| 7 | YRLFRKSNL (120-128) | HLA-B2705 |
| 8 | RLFRKSNLK (121-129) | HLA-A0301 |
| 9 | QSYGFQPTY (160-170) | HLA-B5801 |
| 10 | YQPYRVVVL (172-180) | HLA-B3901 |
| 11 | PYRVVVLSF (174-182) | HLA-A2402 |
| 12 | TNLKLAAAL (198-206) | HLA-B3901 |
| Parametrs | Categories | |
|---|---|---|
| 1st cohort – (n=43) | 2nd cohortт - (n=32) | |
| Age, Me (Q25; Q75) | 62.50 (46.75; 76.00) | 61.00 (24.00; 67.50) |
| Males | 17 (39.5%) | 5 (15.6%) |
| Females | 26 (60.5%) | 27 (84.4%) |
| No COVID-19 | 16 (37.2%) | 4 (12.5%) |
| COVID-19 | 27 (62.8%) | 28 (87.5%) |
| Time from onset of illness, months Me (Q25; Q75) |
48.00 (36.00; 56.00) | 13.00 (6.00; 19.50) |
| No vaccination | 10 (23.3%) | 12 |
| After vaccination | 33 (78.7%) | 20 |
| Cell populations | Cell subpopulations | Group | |||||
| Not vaccinated (Group1) | No COVID-19, Vaccinated (Group 2) |
COVID-19, vaccinated (Group 3) |
|||||
| # in group | |||||||
| 10 | 14 | 19 | |||||
| SARS-Cov-2 proteins | |||||||
| N | S | N | S | N | S | ||
| General pool of T-helper memory cells (CD3+CD4+CD45RA-) | IFNg+ | 4(40%) | 7(70%) | 8(57%) | 5(36%) | 13(68%) | 13(68%) |
| IL-2+ | 5(50%) | 7(70%) | 7(50%) | 9(64%) | 12(63%) | 15(79%) | |
| TNFa+ | 6(60%) | 6(60%)1 | 10(71%) | 12(86%) | 15(79%) | 18(95%) | |
| IFN+TNF+ | 5(50%) | 4(40%) | 5(36%) | 3(21%) | 9(47%) | 13(68%) | |
| polyfunc | 2(20%) | 2(20%)2 | 3(21%) | 4(29%)3 | 7(37%) | 14(74%) | |
| T-helper cells of central memory (CD45RA-CCR7+) | IFNg+ | 1(10%)5 | 7(70%) | 2(14%) | 5(36%) | 6(32%) | 7(37%) |
| IL-2+ | 5(50%) | 6(60%) | 5(36%) | 8(57%) | 11(58%) | 10(53%) | |
| TNFa+ | 6(60%) | 6(60%) | 9(64%) | 10(71%) | 15(79%) | 16(84%) | |
| IFN+TNF+ | 3(30%) | 1(10%) | 1(7%) | 1(7%) | 3(16%) | 4(21%) | |
| polyfunc | 0(0%) | 0(0%) | 0(0%) | 0(0%) | 3(16%) | 3(16%) | |
| Effector memory T-helper cells (CD45RA-CCR7-) | IFNg+ | 6(60%) | 5(50%) | 9(64%) | 7(50%) | 12(63%) | 13(68%) |
| IL-2+ | 4(40%) | 5(50%) | 7(50%) | 7(50%) | 11(58%) | 16(84%) | |
| TNFa+ | 6(60%) | 6(60%) | 10(7%) | 11(79%) | 15(79%) | 17(90%) | |
| IFN+TNF+ | 5(50%) | 4(40%) | 4(29%) | 3(21%) | 8(42%) | 11(58%) | |
- 1
- The number of S-protein responders in Group 1 was lower than in Group 3 (P<0.05, Fisher's exact test)
- 1
- The number of S-protein responders in Group 1 was lower than in Group 3 (P<0.05, Fisher's exact test)
- 1
- The number of S-protein responders in Group 1 was lower than in Group 3 (P<0.05, Fisher's exact test)
- 1
- The number of N-protein responders was lower than S-protein responders (P<0.05, Fisher's exact test)
- 1
- The number of N-protein responders was lower than S-protein responders (P<0.01, Fisher's exact test)
| Cell populations | Cell subpopulations | Group | |||||
| Not vaccinated (Group1) |
No COVID-19, Vaccinated (Group 2) | COVID-19, vaccinated (group 3) | |||||
| # in group | |||||||
| 10 | 14 | 19 | |||||
| SARS-Cov-2 proteins | |||||||
| N | S | N | S | N | S | ||
| Total pool of cytotoxic CD8+ memory T cells (CD3+CD8+CD45RA-) | IFNg+ | 2(20%) | 4(40%) | 5(36%) | 4(29%) | 7(37%) | 3(16%) |
| IL-2+ | 3(30%) | 3(30%) | 4(29%) | 4(29%) | 9(47%) | 9(47%) | |
| TNFa+ | 6(60%) | 5(50%)6 | 13(93%) | 11(79%) | 16(84%) | 17(90%) | |
| IFN+TNF+ | 0(0%) | 1(10%) | 1(7%) | 0(0%) | 1(5%) | 1(5%) | |
| polyfunc | 0(0%) | 0(0%) | 0(0%) | 0(0%) | 0(0%) | 1(5%) | |
| Сytotoxic CD8+ T cells of central memory (CD45RA-CCR7+), | IFNg+ | 1(10%) | 1(10%) | 1(7%) | 1(7%) | 3(16%) | 1(5%) |
| IL-2+ | 4(40%) | 3(30%) | 2(14%) | 4(29%) | 4(21%) | 6(32%) | |
| TNFa+ | 6(60%) | 5(50%) | 6(43%) | 4(29%) | 11(58%) | 10(53%) | |
| IFN+TNF+ | 0(0%) | 0(0%) | 0(0%) | 0(0%) | 0(0%) | 0(0%) | |
| polyfunc | 0(0%) | 0(0%) | 0(0%) | 0(0%) | 0(0%) | 0(0%) | |
| Сytotoxic CD8+ T cells of effector memory (CD45RA-CCR7-) | IFNg+ | 2(20%) | 4(40%) | 5(36%) | 5(36%) | 5(26%) | 2(11%) |
| IL-2+ | 2(20%) | 2(20%) | 4(29%) | 4(29%) | 8(42%) | 9(47%) | |
| TNFa+ | 5(50%) | 5(50%) | 11(79%) | 9(64%) | 13(68%) | 14(74%) | |
| IFN+TNF+ | 3(30%) | 3(30%) | 7(50%) | 7(50%) | 9(47%) | 9(47%) | |
| polyfunc | 0(0%) | 0(0%) | 0(0%) | 0(0%) | 0(0%) | 0(0%) | |
- 1
- The number of patients who responded to S-protein stimulation in Group 1 was lower than in Group 3 (P<0.05, Fisher's exact test).
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




