3.1. Phylogenetic and Diversity Analysis of Culturable Fungal Strains
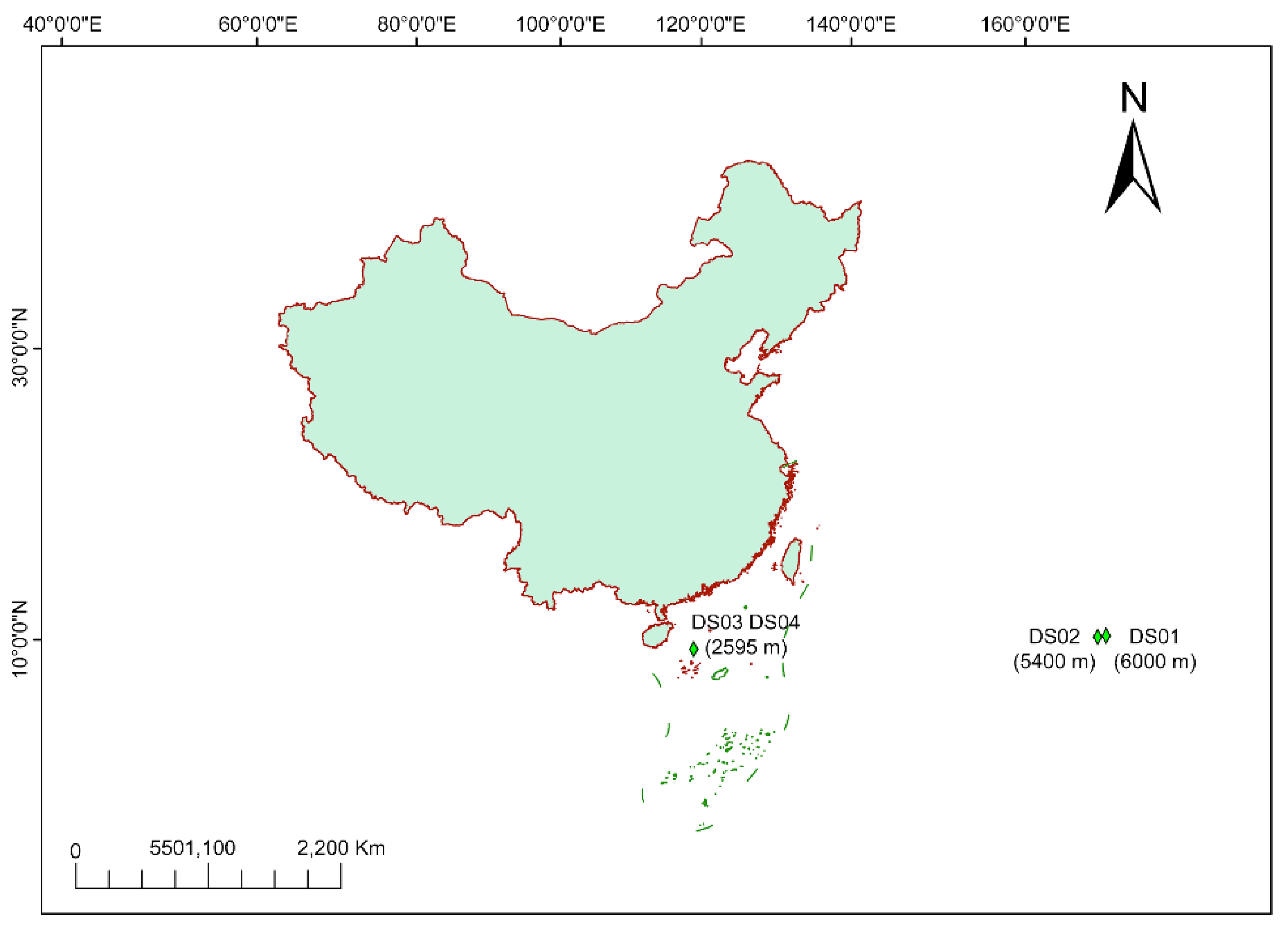
A total of 159 culturable fungal strains were isolated from four deep-sea sediment samples (
Supplementary Table S3) using 12 distinct isolation protocols (
Supplementary Table S2). Taxonomic status was determined by sequencing the ITS region, which identified 128, 30, and 1 strains as Ascomycota, Basidiomycota, and Mucoromycota, respectively. The ITS sequences of 125 strains were highly similar (95%–100%) to known species belonging to 34 genera. However, similarity matches of 34 strains (listed in
Supplementary Table S3) were <95%, a threshold commonly used to identify species, indicating their potential as novel taxonomic units.
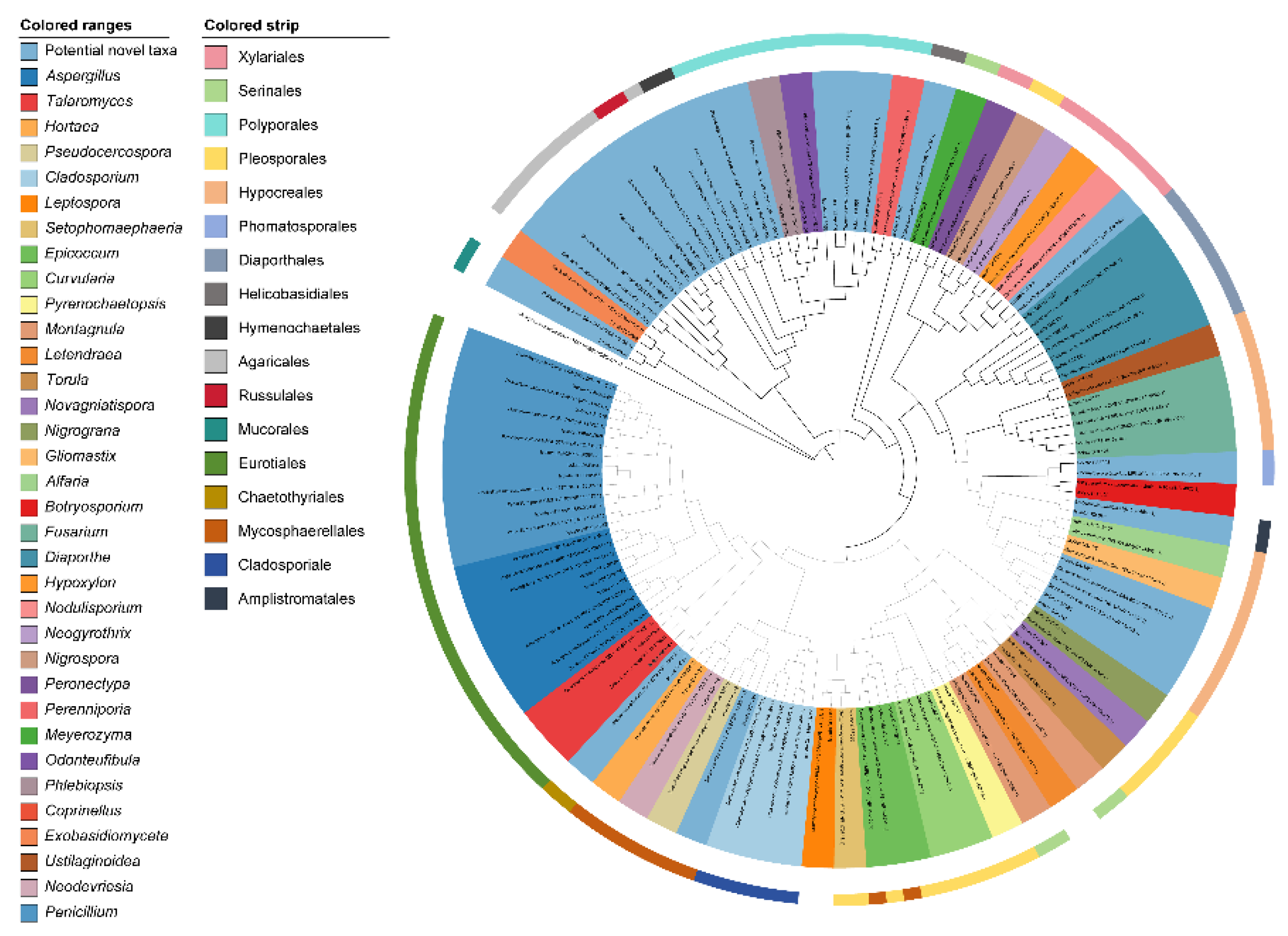
We selected 76 representative isolates for phylogenetic analysis based on combined ITS sequence and phenotypic data, generating the neighbor-joining tree shown in
Figure 2. The 76 isolates belonged to the following 35 genera:
Alfaria,
Aspergillus,
Botryosporium,
Cladosporium,
Coprinellus,
Curvularia,
Diaporthe,
Epicoccum,
Exobasidiomycetidae,
Fusarium, Gliomastix,
Hortaea,
Hypoxylon,
Leptospora,
Letendraea,
Meyerozyma,
Montagnula,
Neodevriesia,
Neodidymella,
Neogyrothrix,
Neovaginatispora,
Nigrograna,
Nigrospora,
Nodulisporium,
Odontoefibula,
Penicillium,
Perenniporia,
Peroneutypa,
Phlebiopsis, Pseudocercospora,
Pyrenochaetopsis,
Setophaeosphaeria,
Talaromyces,
Torula, and
Ustilaginoidea. These were distributed across 17 orders: Agaricales, Amplistromatales, Chaetothyriales, Cladosporiales, Diaporthales, Eurotiales, Helicobasidiales, Hymenochaetales, Hypocreales, Mycosphaerellales, Mucorales, Pleosporales, Polyporales, Russulales, Serinales, Xylariales, and Phomatosporales. Additionally, two, one, and one strains belonged to the class Dothideomycetes (order uncertain), class Exobasidiomycetes (order uncertain), and x Ascomycota (class and order uncertain), respectively. The dominant genera were
Cladosporium (41 strains, 25.79%), followed by
Penicillium (19 strains, 11.95%), and
Aspergillus (15 strains, 9.43%) (
Table 1).
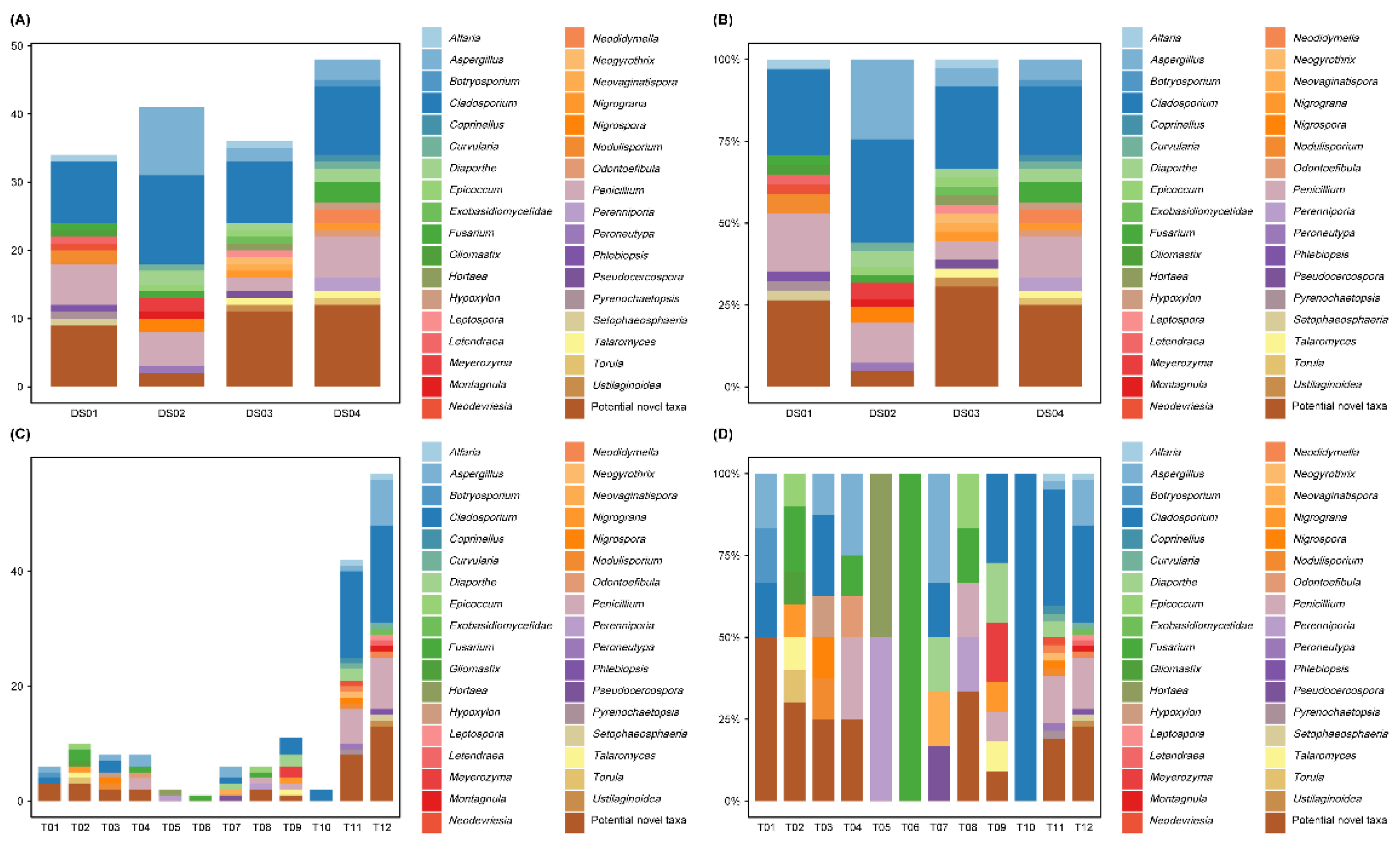
The distribution of the 159 isolates across the four sampling sites is shown in
Figure 3A and 3B. Site DS04 yielded the highest number and diversity of isolates (48 strains, 15 genera, 30.19%), followed by DS02 (41 strains, 11 genera, 25.79%), DS03 (36 strains, 15 genera, 22.64%), and DS01 (34 strains, 11 genera, 21.38%). Heatmap analysis (
Supplementary Figure S1A) showed
Cladosporium as the dominant (20.83%–31.71% of isolates) genus across all sites, followed by
Penicillium (5.56%–17.65%).
Aspergillus was isolated from all sites except DS01 and was particularly abundant at DS02, constituting 24.39% of its community. We identified 20 rare genera (defined as comprising <1% of total abundance), which were mostly detected at DS03 (7 genera), and least detected at DS02 (2 genera). Sites DS02 and DS04 had the fewest and highest proportion of potentially novel taxa, respectively.
Fungal recovery varied significantly with the isolation protocol, as shown in
Figure 3C and 3D. Among the 12 treatment combinations, T12 yielded the highest number and diversity of isolates, accounting for 35.85% of the total (57 strains, 13 genera), followed by protocol T11 (42 strains, 14 genera, 26.42%). Specifically, the "stamping method" combined with either GPY or Fungi No. II medium was the most effective strategy, collectively accounting for 99 strains (62.26% of the total). The 10⁻¹ dilution (T1-T4) yielded 32 strains (20.13%, 14 genera), whereas the 10⁻² dilution (T5-T8) yielded 15 strains (9.43%, 12 genera), consistent with the dilution effect. The 10⁻¹ dilution on MEA (T2) yielded the highest number of isolates (10 strains, 6 genera); however, the differences among the different media were not significant. In contrast, at the 10⁻² dilution, Fungus No. II medium (T7) and Martin’s agar (T8) grew significantly more isolates (6 strains each) than GPY (T5) or MEA (T6). Pretreatment with EDTA (T9 and T10) yielded slightly more isolates, but this improvement was not statistically significant (P > 0.05) (
Supplementary Figure S1B).
3.2. Alpha and Beta Diversity of Fungal Communities
Comparison of alpha and beta diversity indices of fungal communities across different sites and isolation methods revealed significant differences in alpha diversity among the four sampling sites (
Table 2). The values of Shannon and Simpson indices indicate the highest species richness and diversity (H’ = 3.589, 1–D = 0.9699) at site DS04, followed by DS03 (H’ = 3.066, 1–D = 0.9302). The evenness index values for DS01, DS03, and DS04 are high (>0.97), with a value of 1.131 for DS04, indicating a relatively balanced abundance distribution, whereas the evenness at DS02 was lower (0.8143). Values of all indices consistently suggest lower community diversity at DS02. Notably, the Chao1 richness estimator was substantially higher than the observed species count for all sites. Results of PCoA based on Bray-Curtis dissimilarity reveal clear differences in community composition (beta diversity) among sites (
Supplementary Figure S2A). The first two PCoA axes explain 71.45% of the total variation. The plot shows four distinct, nonoverlapping clusters corresponding to each site. Sites DS01 and DS03 show a similar clustering pattern, while DS02 and DS04 form distinct clusters separated from the others.
Alpha diversity also varied with the different isolation methods (
Table 3). The observed number of genera ranged from 1 (T06) to 30 (T12), with the latter yielding the highest diversity (Taxa = 30, H’ = 3.242, 1–D = 0.9292, Evenness = 0.8526, Chao1 = 71.26). Methods using 10⁻² diluted soil suspensions yielded the lowest diversity. The evenness index values are close to or slightly above 1 for most dilution-based methods (T01-T10), reflecting an extremely even distribution of recovered taxa, likely a consequence of the limited number of isolates per method. In contrast, we observed lower evenness in T11 and T12 (0.7411 and 0.8526, respectively), suggesting the dominance of a few taxa. Notably, we found higher Chao1 values than the observed count for all methods, most dramatically for T11 (229.1 Chao1 vs. 24 observed), indicating that even the most effective isolation strategy captured only a fraction of the presumed fungal diversity. Results of PCoA of beta diversity reveal that the isolation method was the primary factor shaping the recoverable fungal community composition (
Supplementary Figure S2B). The first two axes explained 55.22% of the variation, with the plot showing two distinct clusters: T01-T10, i.e., samples that underwent dilution-based methods forming a relatively tight cluster, whereas those from stamping methods (T11 and T12) were clearly separated and distant from the former. This result indicates that the stamping and dilution methods accessed distinct subsets of the fungal community.
3.3. In Vitro Antagonism Assessed by Dual-Culture Assay
The 23 representative fungal isolates screened for antagonism against phytopathogenes were selected based on a dereplication analysis of ITS sequences to capture phylogenetic diversity. We focused on
Penicillium and
Aspergillus because they are well-documented as prolific producers of a broad spectrum of antimicrobial metabolites [
2,
15,
16]. Nineteen of these isolates (82.6%) exhibited antagonistic activity against at least one of the five phytopathogens tested, confirming our hypothesis that deep-sea fungi are a reservoir of potential biocontrol agents with a broad spectrum of antifungal activity (
Table 4,
Figure 4).
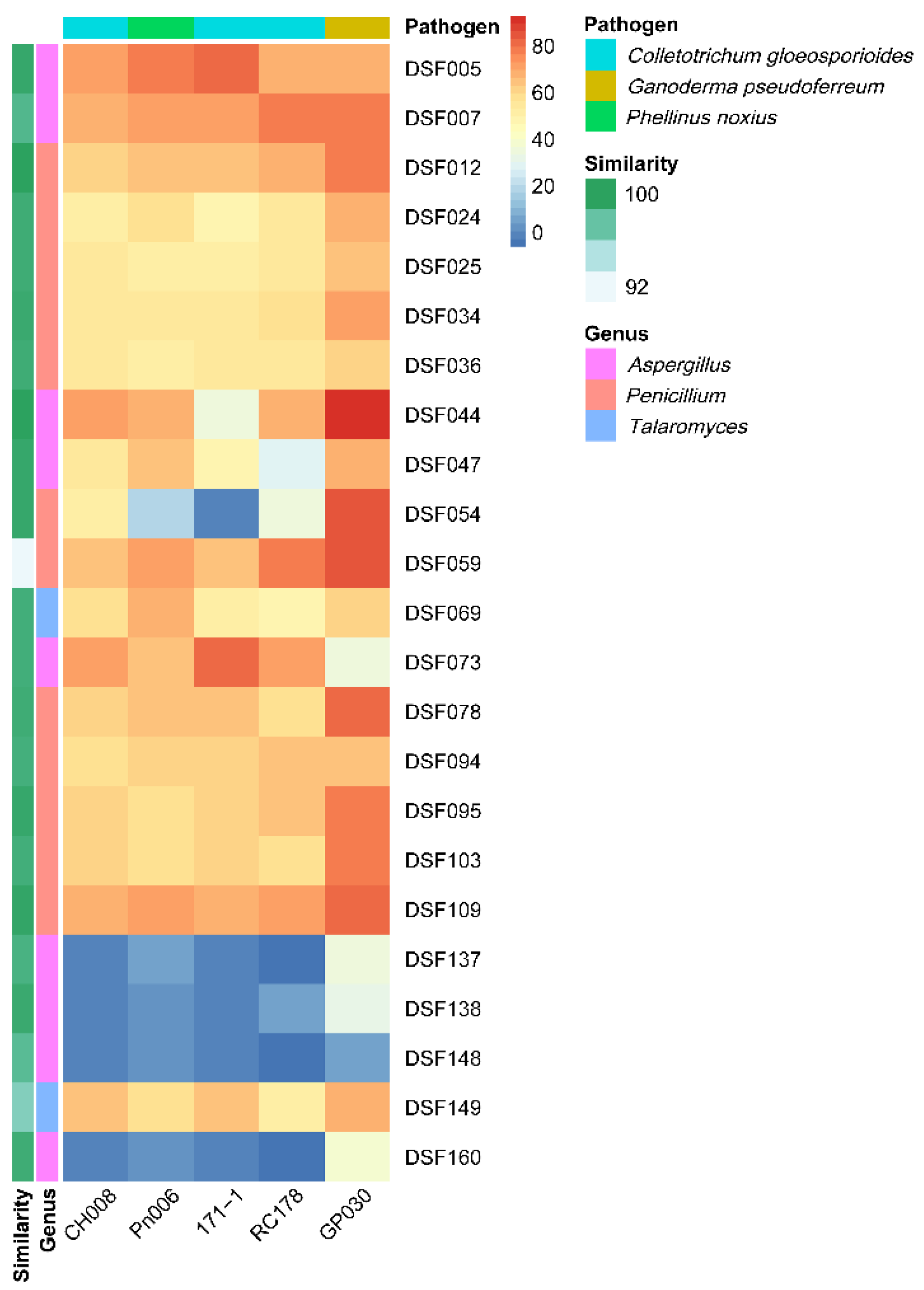
The antagonistic activity of isolates varied considerably according to the pathogen. The three strains of C. gloeosporioides were inhibited at rates that ranged from 50.2 ± 5.43% to 70.59 ± 0% for CH008; from 34.51 ± 0.68% to 81.96 ± 3.4% for 171-1; and from 3.75 ± 4.33% to 79.17 ± 9.02% for RC178. The basidiomycete pathogens were inhibited at rates that ranged from 0.39 ± 0.68% to 78.82 ± 2.04% against P. noxius Pn006, and from 3.93 ± 0% to a maximum inhibition of 92.72 ± 0% against G. pseudoferreum GP030.
We identified a subset of highly effective antagonists: Isolates DSF005, DSF007, DSF044, DSF059, DSF073, and DSF109 consistently performed well against multiple pathogens. For instance, DSF073 showed the highest inhibition(81.96% ± 3.4%) against
C. gloeosporioides 171-1, whereas DSF044 exhibited the strongest overall activity (92.72% ± 0%) against
G. pseudoferreum GP030. DSF005 demonstrated potent broad-spectrum activity, achieving high inhibition rates against
P. noxius Pn006 (78.82% ± 2.04%) and several
C. gloeosporioides strains (
Figure 5A). These isolates form a robust subset of promising candidates for subsequent, mode-of-action-focused assays.
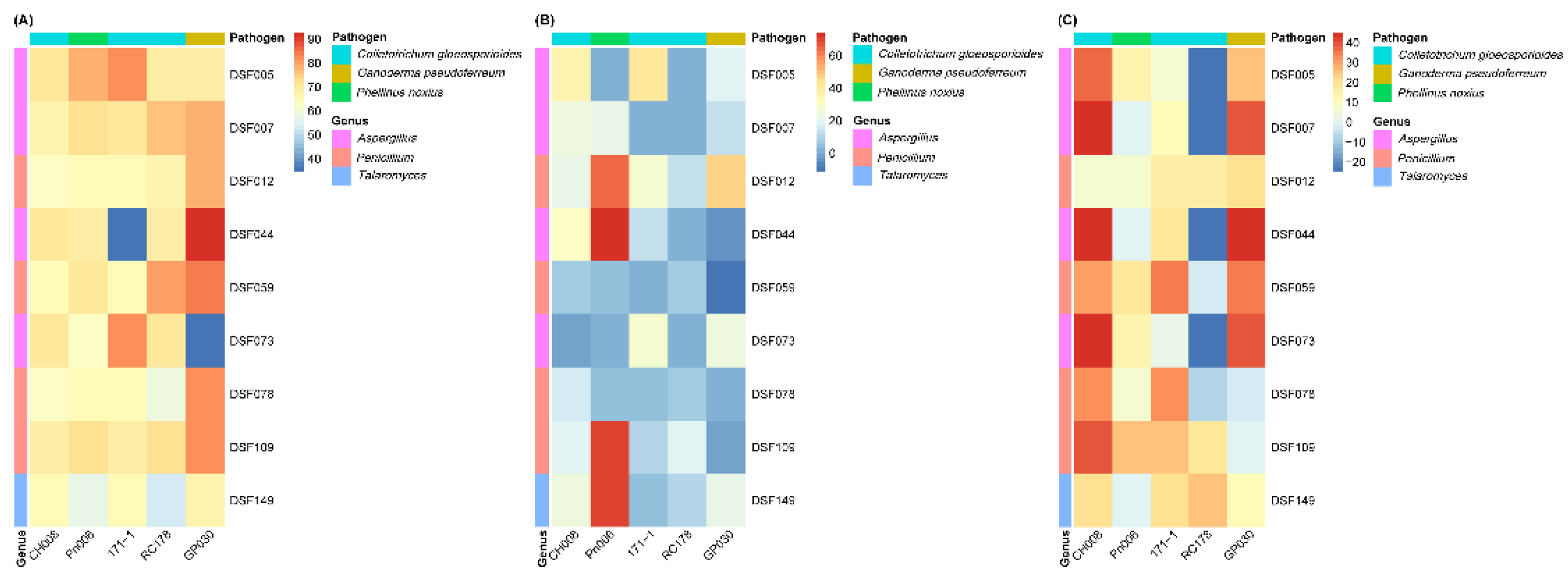
3.4. Antifungal Activity of Fungal Culture Filtrates
To determine the contribution of diffusible metabolites in the inhibition of phytopathogens, we evaluated the antifungal activities of sterile culture filtrates from nine fungal strains (DSF005, DSF007, DSF012, DSF044, DSF059, DSF073, DSF078, DSF109, and DSF149). These strains were selected based on the antagonistic spectrum observed in the dual-culture assay and their taxonomic classification. The results (
Table 5,
Figure 5B) reveal pronounced heterogeneity in inhibition mediated by diffusible metabolites.
C. gloeosporioides CH008 was most strongly inhibited by the culture filtrates of DSF005 and DSF044 at inhibition rates of 36.95 ± 1.39% and 30.92±1.39%, respectively. Several isolates, including DSF149, DSF007, and DSF012, showed moderate inhibition (22.49 ± 2.51%-24.9±3.03%), whereas DSF059, DSF078, and DSF109 displayed weak inhibition (<18%). Notably, the culture filtrate of DSF073 did not inhibit the mycelial growth of CH008.
P. noxius Pn006 was strongly inhibited by the culture filtrates of multiple isolates. Strong inhibition (>70%) was observed for DSF044 (74.12% ± 4.08%), DSF149 (70.59% ± 1.18%), and DSF109 (70.20% ± 11.18%), and DSF012 (63.53% ± 4.24%). Other isolates exhibited weak or no inhibitory effects against this pathogen.
C. gloeosporioides 171-1 was most strongly inhibited by DSF005 (40.83% ± 22.09%), with DSF012 and DSF073 showing moderate activity (~25%). C. gloeosporioides RC178 was inhibited by DSF109 at a rate of 18.43% ± 4.75%, followed by DSF012 and DSF149.
Among the tested isolates, DSF012 displayed the highest inhibitory activity against G. pseudoferreum GP030 (46.77% ± 4.27%), whereas DSF073 and DSF149 showed moderate inhibition. The remaining isolates exhibited little to no inhibitory effect and in some cases appeared to promote mycelial growth.
Overall, these results demonstrate that antifungal activity mediated by culture filtrates was highly pathogen-dependent and did not necessarily correlate with antagonistic performance observed in the dual-culture assay. These findings suggest that antifungal activity of the isolates is not fully explained by diffusible metabolite production.
3.5. Antifungal Activity of Fungal Volatile Organic Compounds
The antifungal activity of VOCs produced by the nine selected deep-sea fungal isolates varied according to both isolate and pathogen (
Table 6,
Figure 5C). Overall, VOC-mediated effects differed substantially from those observed in both dual-culture and culture filtrate assays, indicating a distinct and independent mode of antagonism.
C. gloeosporioides CH008 was most strongly inhibited by VOCs from DSF007, DSF044, and DSF073 at rates of 44.18 ± 1.39%, 44.18 ± 1.39%, and 44.98 ± 0.70%, respectively. Isolates DSF005, DSF059, DSF078, and DSF109 exhibited notable inhibition rates ranging from 30.12 ± 9.39% to 39.76 ± 4.78%. VOCs from DSF012 and DSF149 were weakly inhibitory at rates of 6.02 ± 2.41% and 20.48 ± 3.61%, respectively.
P. noxius Pn006 was modestly inhibited by VOCs of DSF109 at a rate of 24.31 ± 4.45%, followed by DSF059 at a rate of 18.04 ± 4.13%. VOCs of isolates DSF005, DSF012, DSF073, and DSF078 were weakly inhibitory at rates ranging from 6.67 ± 1.36% to 13.73 ± 3.40%, whereas VOCs of DSF007, DSF044, and DSF149 failed to inhibit phytopathogens under the tested conditions.
A distinct inhibition pattern was observed for C. gloeosporioides 171-1: VOCs from DSF059 and DSF078 suppressed mycelial growth at the highest rates of 33.74 ± 8.99% and 32.52 ± 7.62%, respectively. Moderate inhibition was observed for DSF109 at a rate of 24.39 ± 16.91%, whereas DSF012, DSF044, and DSF149 exhibited intermediate inhibition rates ranging from 16.67 ± 14.75% to 19.51 ± 7.32%. The remaining isolates displayed limited inhibition.
Notably, VOCs exerted both inhibitory and stimulatory effects, depending on the pathogen. For example, C. gloeosporioides RC178 was inhibited by VOCs from DSF149 at a rate of 24.51 ± 5.94%, whereas VOCs from DSF005, DSF007, DSF044, and DSF073 significantly promoted mycelial growth by approximately 25%. Similarly, the growth of G. pseudoferreum GP030 was weakly promoted by VOCs from DSF078 and DSF109. Conversely, VOCs from DSF044 strongly inhibited G. pseudoferreum GP030, achieving a maximum inhibition of 45.00 ± 14.74%.
Collectively, these results demonstrate that fungal VOCs can both inhibit and stimulate growth of phytopathogens, underscoring the ecological complexity of VOC-driven fungal antagonism. Therefore, it is necessary to evaluate multiple antagonistic mechanisms when screening deep-sea fungi for biocontrol applications.