Submitted:
19 February 2026
Posted:
26 February 2026
You are already at the latest version
Abstract
Haiti is a Caribbean country of about 11 million people with a high burden of mosquito-transmitted disease and limited vector control, thereby making effective operational mosquito control of high import. Previous studies have examined vector-borne disease burden and insecticide resistance markers in Haitian Aedes and Anopheles mosquitoes but not Culex species. In this study, we examined collections of Culex quinquefasciatus from 12 locations in northern and southern Haiti for the presence of markers of insecticide resistance (using a variety of target site mutations and biochemical assays) and pathogens (using a deep sequencing microbiome workflow). The metagenome analysis identified Wolbachia, Rhabdoviridae and Plasmodium infection in all sample pools at relatively high levels along with less frequent findings of other potential pathogens. Resistance marker examination identified variable frequencies of knockdown resistance and acetylcholinesterase resistance mutations, as well as variation in resistance-associated enzymatic activities in these populations, which indicate that insecticide resistance to the primary pyrethroid and organophosphate insecticides is likely. Though there was variation between Culex mosquito populations and no clear activity pattern, enzymatic activity was significantly higher in the southern sites compared to the northern sites. Similar findings in Cx. quinquefasciatus populations in other locations in the Americas strongly suggest that vector control with pyrethroid and organophosphate adulticides may be of limited efficacy.
Keywords:
1. Introduction
2. Materials and Methods
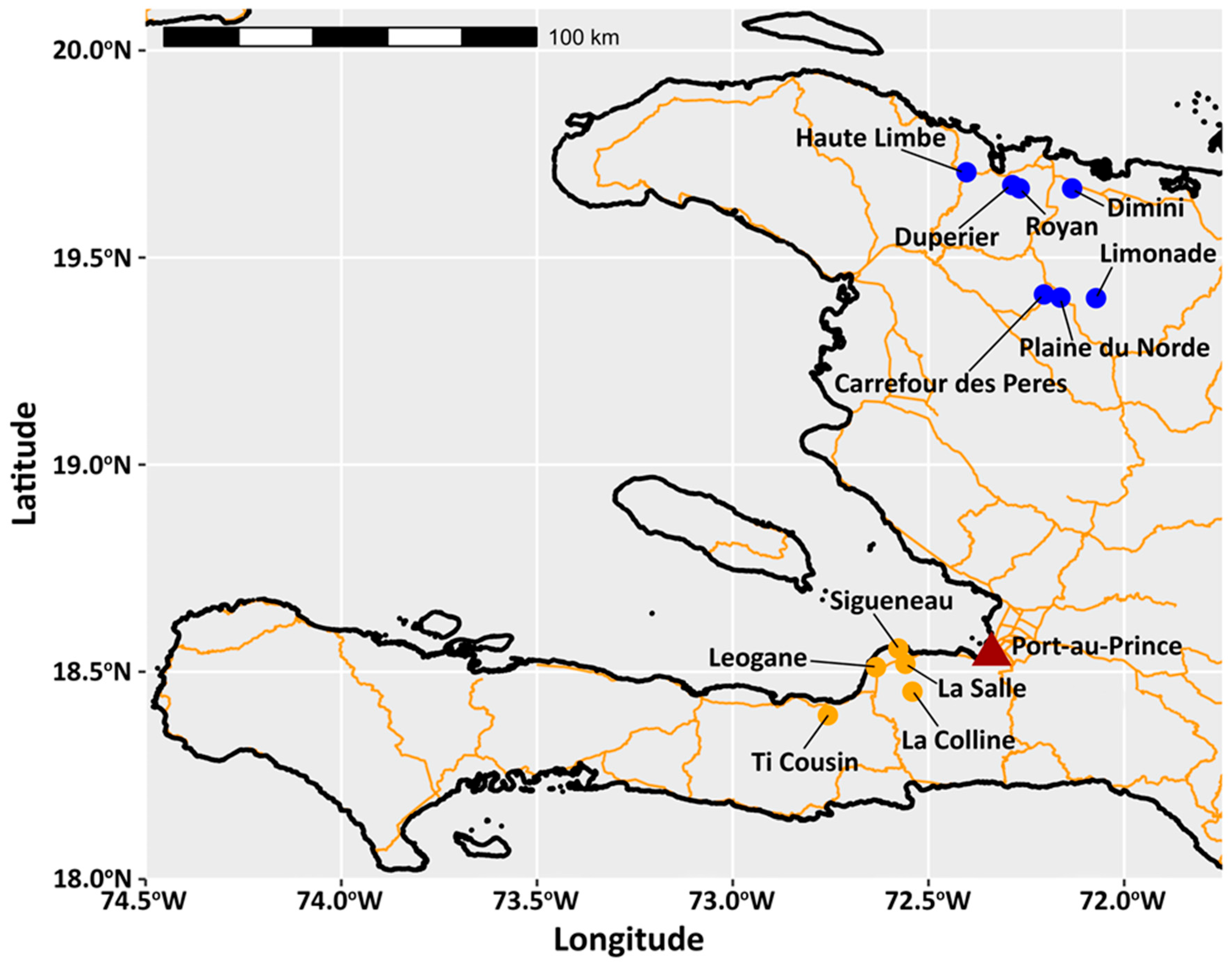
2.1. Arthropod Surveillance Collection Procedures
2.2. Control Strain
2.3. Sample Homogenization, Pooling and RNA Purification
2.4. Speciation, Knockdown Resistance and Acetylcholinesterase Mutation Detection Assays
2.5. Metabolic Resistance Assays
2.5.1. Bradford Protein Assay
2.5.2. Cytochrome P450 Assay
2.5.3. Glutathione S-Transferase (GST) Activity Assay
2.5.4. α-Carboxylesterase Activity Assay
2.5.5. β-Carboxylesterase Activity Assay
2.6. Nanopore Sequencing and Bioinformatics
3. Results
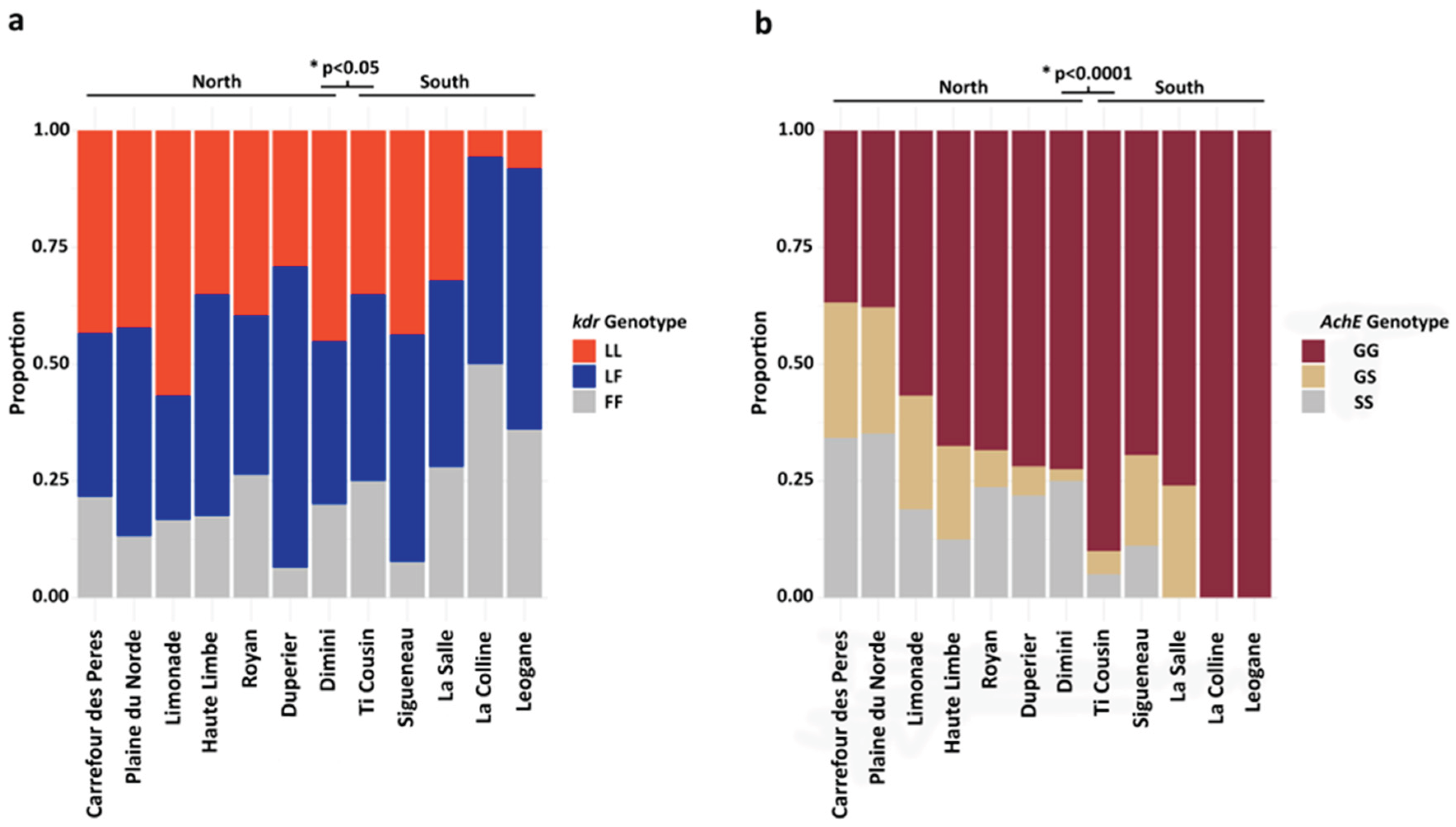
3.1. Assessment of Knockdown and Acetylcholinesterase Target Site Mutations
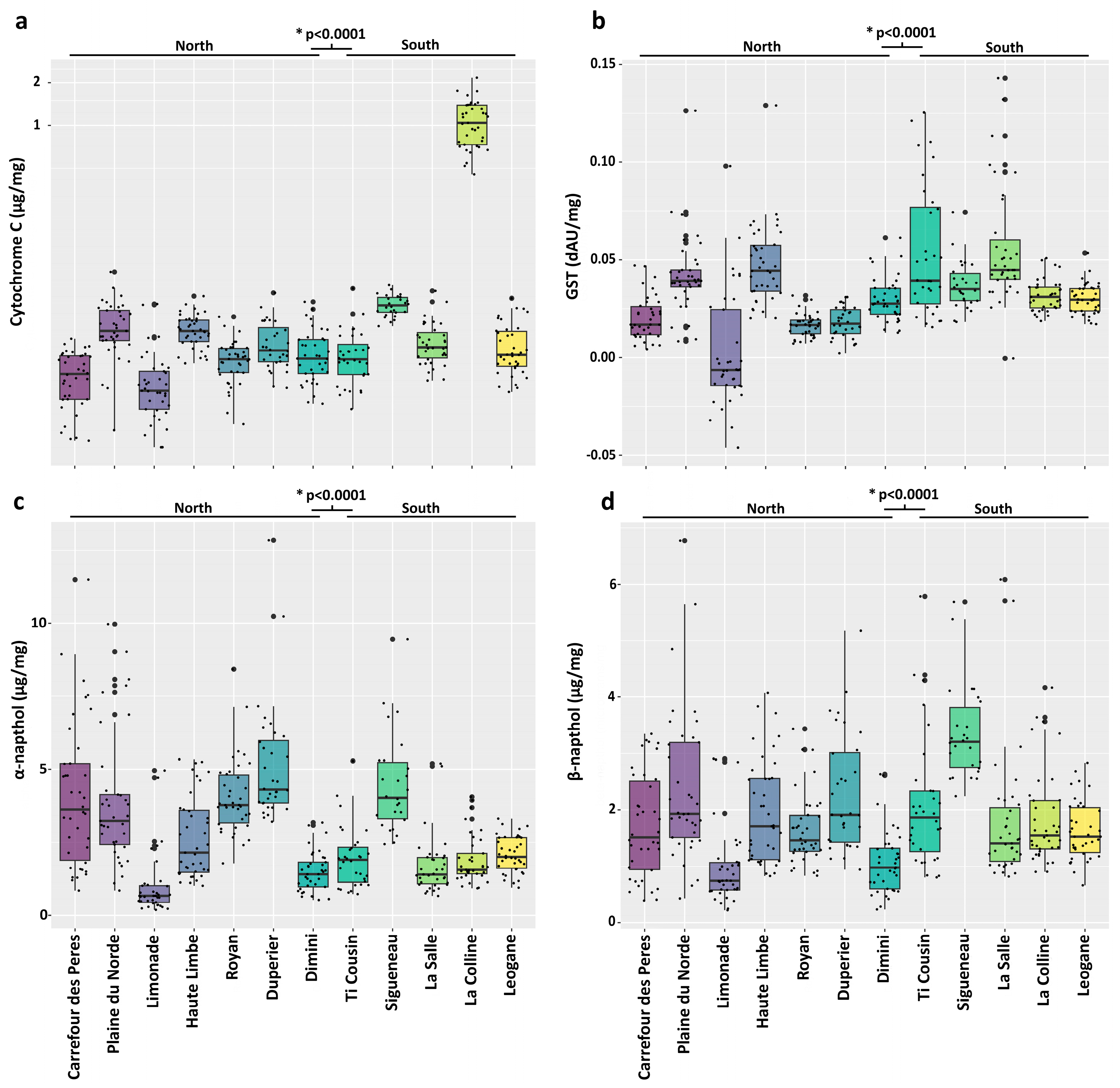
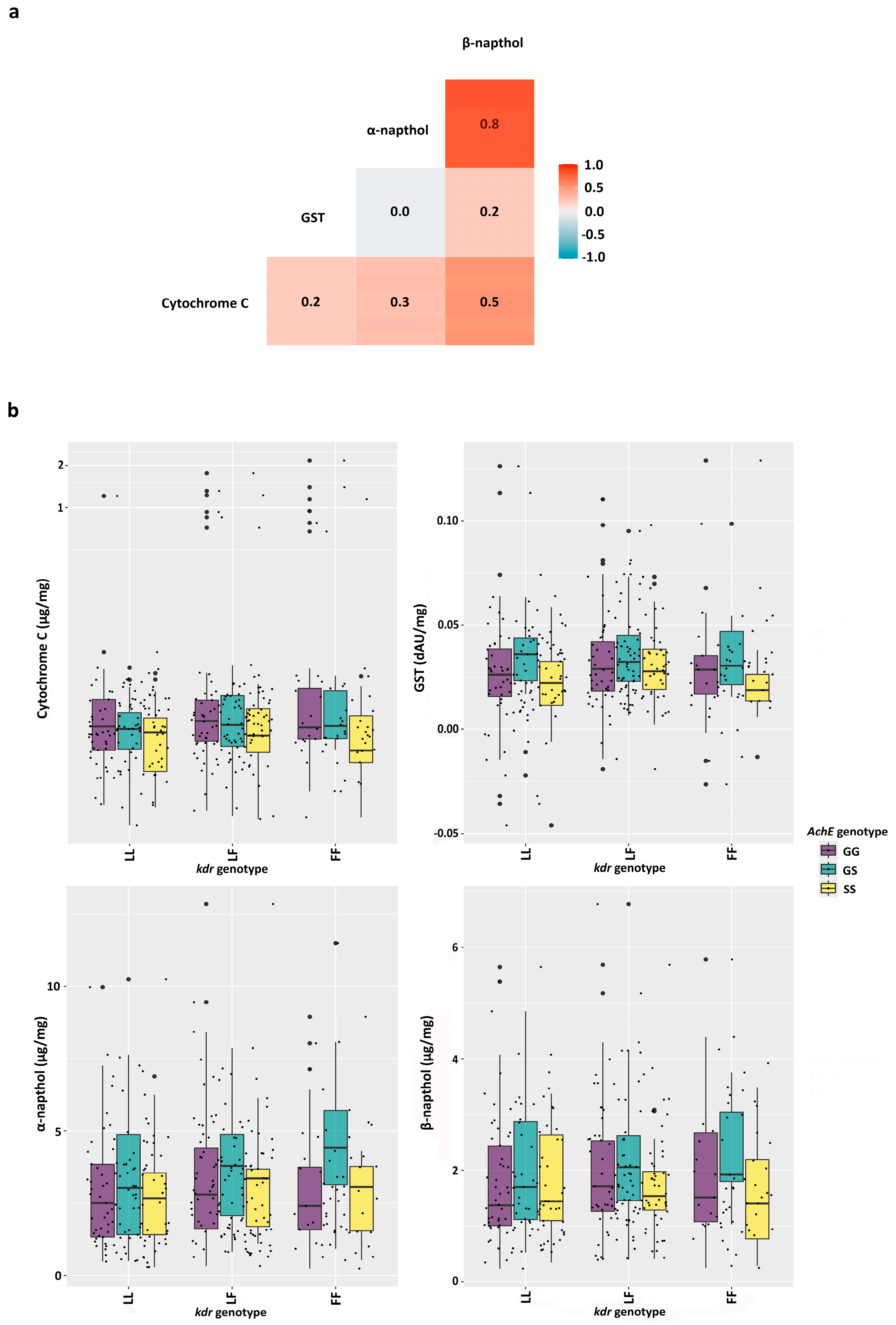
3.2. Metabolic Resistance Assays
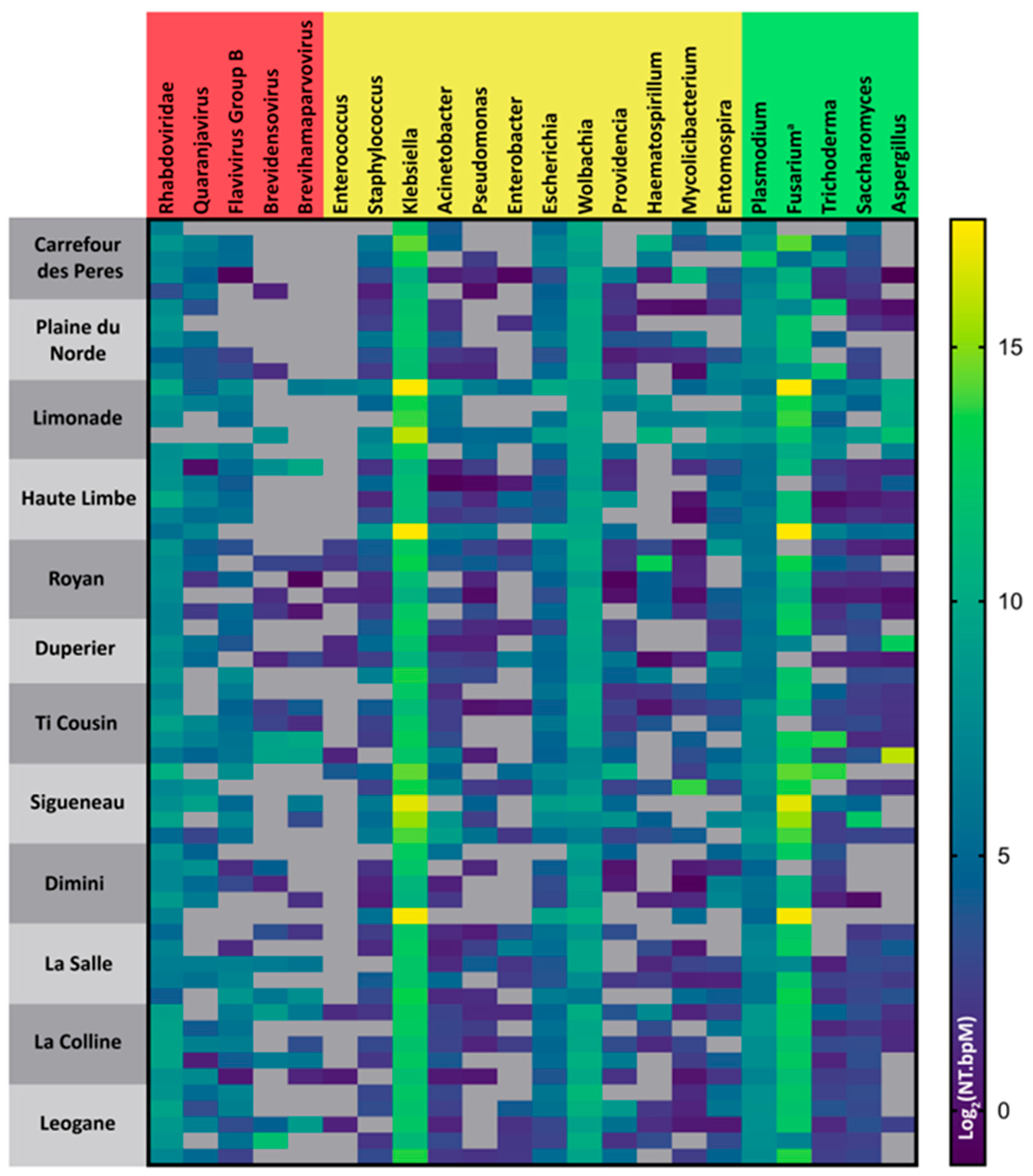
3.2. Microbiome Analysis
4. Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| WNV kdr AchE NFW MCA Tm |
West Nile virus knock down resistance acetylcholinesterase nuclease free water melt curve assay melting temperature |
| dscDNA | double stranded cDNA |
| Ct | cycle threshold |
| GST | glutathione S transferase |
| CytC | cytochrome C oxidase |
| NT.bpm | NT/NR database matching bases per million |
References
- Beau de Rochars, M.V.; Milord, M.D.; St Jean, Y.; Desormeaux, A.M.; Dorvil, J.J.; Lafontant, J.G.; Addiss, D.G.; Streit, T.G. Geographic distribution of lymphatic filariasis in Haiti. Am J Trop Med Hyg 2004, 71, 598–601. [Google Scholar] [CrossRef]
- Drexler, N.; Washington, C.H.; Lovegrove, M.; Grady, C.; Milord, M.D.; Streit, T.; Lammie, P. Secondary mapping of lymphatic filariasis in Haiti-definition of transmission foci in low-prevalence settings. PLoS Negl Trop Dis 2012, 6, e1807. [Google Scholar] [CrossRef]
- Kent, R.J.; Crabtree, M.B.; Miller, B.R. Transmission of West Nile virus by Culex quinquefasciatus say infected with Culex Flavivirus Izabal. PLoS Negl Trop Dis 2010, 4, e671. [Google Scholar] [CrossRef] [PubMed]
- Crockett, R.K.; Burkhalter, K.; Mead, D.; Kelly, R.; Brown, J.; Varnado, W.; Roy, A.; Horiuchi, K.; Biggerstaff, B.J.; Miller, B.; et al. Culex flavivirus and West Nile virus in Culex quinquefasciatus populations in the southeastern United States. J Med Entomol 2012, 49, 165–174. [Google Scholar] [CrossRef] [PubMed]
- Diaz, L.A.; Flores, F.S.; Beranek, M.; Rivarola, M.E.; Almiron, W.R.; Contigiani, M.S. Transmission of endemic St Louis encephalitis virus strains by local Culex quinquefasciatus populations in Cordoba, Argentina. Trans R Soc Trop Med Hyg 2013, 107, 332–334. [Google Scholar] [CrossRef] [PubMed]
- Richards, S.L.; Lord, C.C.; Pesko, K.; Tabachnick, W.J. Environmental and biological factors influencing Culex pipiens quinquefasciatus Say (Diptera: Culicidae) vector competence for Saint Louis encephalitis virus. Am J Trop Med Hyg 2009, 81, 264–272. [Google Scholar] [CrossRef]
- Mitchell, C.J.; Gubler, D.J.; Monath, T.P. Variation in infectivity of Saint Louis encephalitis viral strains for Culex pipiens quinquefasciatus (Diptera: Culicidae). J Med Entomol 1983, 20, 526–533. [Google Scholar] [CrossRef]
- Sang, R.; Kioko, E.; Lutomiah, J.; Warigia, M.; Ochieng, C.; O'Guinn, M.; Lee, J.S.; Koka, H.; Godsey, M.; Hoel, D.; et al. Rift Valley fever virus epidemic in Kenya, 2006/2007: the entomologic investigations. Am J Trop Med Hyg 2010, 83, 28–37. [Google Scholar] [CrossRef]
- Ben-Chetrit, E.; Schwartz, E. Vector-borne diseases in Haiti: a review. Travel Med Infect Dis 2015, 13, 150–158. [Google Scholar] [CrossRef]
- Beatty, M.E.; Hunsperger, E.; Long, E.; Schurch, J.; Jain, S.; Colindres, R.; Lerebours, G.; Bernard, Y.M.; Dobbins, J.G.; Brown, M.; et al. Mosquitoborne infections after Hurricane Jeanne, Haiti, 2004. Emerg Infect Dis 2007, 13, 308–310. [Google Scholar] [CrossRef]
- Verma, M.; Phartyal, R.; Bhatt, A. Introduction to Flaviviruses and Their Global Prevalence. In Human Viruses: Diseases, Treatments and Vaccines: The New Insights; Ahmad, S.I., Ed.; Springer International Publishing: Cham, 2021; pp. 411–439. [Google Scholar]
- Scott, J.G.; Yoshimizu, M.H.; Kasai, S. Pyrethroid resistance in Culex pipiens mosquitoes. Pestic Biochem Physiol 2015, 120, 68–76. [Google Scholar] [CrossRef]
- Burgess, E.R., IV; Lopez, K.; Irwin, P.; Jaeger, C.P.; Estep, A.S. Assessing pyrethroid resistance status in the Culex pipiens complex (Diptera: Culicidae) from the northwest suburbs of Chicago, Illinois using Cox regression of bottle bioassays and other detection tools. PLoS One 2022, 17, e0268205. [Google Scholar] [CrossRef] [PubMed]
- Mathews, G.; Derraik, J.G.B.; Walker, M.; Knox, R.; Barraclough, R.K. Morphological variation in invasive mosquito Culex quinquefasciatus Say (Diptera: Culicidae) larvae from an urban site in Auckland, New Zealand. N Z J Zool 2017, 44, 342–353. [Google Scholar] [CrossRef]
- Weill, M.; Fort, P.; Berthomieu, A.; Dubois, M.P.; Pasteur, N.; Raymond, M. A novel acetylcholinesterase gene in mosquitoes codes for the insecticide target and is non-homologous to the ace gene in Drosophila. R Soc Lond B Biol Sci 2002, 269, 2007–2016. [Google Scholar] [CrossRef] [PubMed]
- Lopes, R.P.; Lima, J.B.P.; Martins, A.J. Insecticide resistance in Culex quinquefasciatus Say, 1823 in Brazil: a review. Parasit Vectors 2019, 12, 591. [Google Scholar] [CrossRef]
- Chandrasiri, P.; Fernando, S.D.; De Silva, B. Insecticide resistance and molecular characterization of knockdown resistance (kdr) in Culex quinquefasciatus mosquitoes in Sri Lanka. J Vector Ecol 2020, 45, 204–210. [Google Scholar] [CrossRef]
- McAllister, J.C.; Godsey, M.S.; Scott, M.L. Pyrethroid resistance in Aedes aegypti and Aedes albopictus from Port-au-Prince, Haiti. J Vector Ecol 2012, 37, 325–332. [Google Scholar] [CrossRef]
- Estep, A.S.; Sanscrainte, N.D.; Okech, B.A. Aedes aegypti Knockdown Resistance Mutations and Dengue Virus Infection in Haiti. J Am Mosq Control Assoc 2024, 40, 102–108. [Google Scholar] [CrossRef]
- Delannay, C.; Goindin, D.; Kellaou, K.; Ramdini, C.; Gustave, J.; Vega-Rua, A. Multiple insecticide resistance in Culex quinquefasciatus populations from Guadeloupe (French West Indies) and associated mechanisms. PLoS One 2018, 13, e0199615. [Google Scholar] [CrossRef]
- Nchoutpouen, E.; Talipouo, A.; Djiappi-Tchamen, B.; Djamouko-Djonkam, L.; Kopya, E.; Ngadjeu, C.S.; Doumbe-Belisse, P.; Awono-Ambene, P.; Kekeunou, S.; Wondji, C.S.; et al. Culex species diversity, susceptibility to insecticides and role as potential vector of Lymphatic filariasis in the city of Yaounde, Cameroon. PLoS Negl Trop Dis 2019, 13, e0007229. [Google Scholar] [CrossRef]
- Ukpai, O.; Ekedo, C. Insecticide susceptibility status of Culex quinquefasciatus [Diptera: Culicidae] in Umudike, Ikwuano LGA Abia State, Nigeria. Int J Mosq Res 2019, 6, 114–118. [Google Scholar]
- Davis, E.L.; Prada, J.; Reimer, L.J.; Hollingsworth, T.D. Modelling the Impact of Vector Control on Lymphatic Filariasis Programs: Current Approaches and Limitations. Clin Infect Dis 2021, 72, S152–S157. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. _R: A language and environment for statistical computing_. R foundation for statistical computing, Vienna, Austria, 2024; Available online: https://www.R-project.org/.
- Reinert, J.F.; Kaiser, P.E.; Seawright, J.A. Analysis of the Anopheles (Anopheles) quadrimaculatus complex of sibling species (Diptera: Culicidae) using morphological, cytological, molecular, genetic, biochemical, and ecological techniques in an integrated approach. J Am Mosq Control Assoc 1997, 13 Suppl, 1–102. [Google Scholar] [PubMed]
- McCall, P.J.; Eaton, G. Olfactory memory in the mosquito Culex quinquefasciatus. Med Vet Entomol 2001, 15, 197–203. [Google Scholar] [CrossRef]
- Pridgeon, J.W.; Pereira, R.M.; Becnel, J.J.; Allan, S.A.; Clark, G.G.; Linthicum, K.J. Susceptibility of Aedes aegypti, Culex quinquefasciatus Say, and Anopheles quadrimaculatus Say to 19 pesticides with different modes of action. J Med Entomol 2008, 45, 82–87. [Google Scholar] [CrossRef]
- Estep, A.S.; Sanscrainte, N.D.; Stuck, J.; Unlu, I.; Prasauskas, A.; Mundis, S.J.; Cotter, N.; Romero-Weaver, A.L.; Fedirko, T.J.; Kendziorski, N.L.; et al. The L1014F knockdown resistance mutation is not a strong correlate of phenotypic resistance to pyrethroids in Florida populations of Culex quinquefasciatus. Insects 2024, 15. [Google Scholar] [CrossRef]
- Unlu, I.; Buckner, E.A.; Medina, J.; Vasquez, C.; Cabrera, A.; Romero-Weaver, A.L.; Ramirez, D.; Kendziorski, N.L.; Kosinski, K.J.; Fedirko, T.J.; et al. Insecticide resistance of Miami-Dade Culex quinquefasciatus populations and initial field efficacy of a new resistance-breaking adulticide formulation. PLoS One 2024, 19, e0296046. [Google Scholar] [CrossRef]
- Kolde, R. pheatmap: Pretty Heatmaps. Available online: https://CRAN.R-project.org/package=pheatmap.
- Boyd, A.; Won, K.Y.; McClintock, S.K.; Donovan, C.V.; Laney, S.J.; Williams, S.A.; Pilotte, N.; Streit, T.G.; Beau de Rochars, M.V.; Lammie, P.J. A community-based study of factors associated with continuing transmission of lymphatic filariasis in Leogane, Haiti. PLoS Negl Trop Dis 2010, 4, e640. [Google Scholar] [CrossRef]
- Liu, J.; Wang, Y.; Liu, P.; Yu, X.; Tan, A.; Zeng, J.; Li, L.; Qiu, X. Detection of Target Site Mutations in the Acetylcholinesterase and Voltage-Gated Sodium Channel in Field Populations of Culex quinquefasciatus and Cx. tritaeniorhynchus From Southern Sichuan Region of China. J Am Mosq Control Assoc 2023, 39, 57–60. [Google Scholar] [CrossRef]
- Wang, L.; Soto, A.; Remue, L.; Rosales Rosas, A.L.; De Coninck, L.; Verwimp, S.; Bouckaert, J.; Vanwinkel, M.; Matthijnssens, J.; Delang, L. First Report of Mutations Associated With Pyrethroid (L1014F) and Organophosphate (G119S) Resistance in Belgian Culex (Diptera: Culicidae) Mosquitoes. J Med Entomol 2022, 59, 2072–2079. [Google Scholar] [CrossRef]
- Zhao, M.; Dong, Y.; Ran, X.; Wu, Z.; Guo, X.; Zhang, Y.; Xing, D.; Yan, T.; Wang, G.; Zhu, X.; et al. Point mutations associated with organophosphate and carbamate resistance in Chinese strains of Culex pipiens quinquefasciatus (Diptera: Culicidae). PLoS One 2014, 9, e95260. [Google Scholar] [CrossRef] [PubMed]
- Talipouo, A.; Mavridis, K.; Nchoutpouen, E.; Djiappi-Tchamen, B.; Fotakis, E.A.; Kopya, E.; Bamou, R.; Kekeunou, S.; Awono-Ambene, P.; Balabanidou, V.; et al. High insecticide resistance mediated by different mechanisms in Culex quinquefasciatus populations from the city of Yaounde, Cameroon. Sci Rep 2021, 11, 7322. [Google Scholar] [CrossRef] [PubMed]
- Davis, A.P.; Grondin, C.J.; Johnson, R.J.; Sciaky, D.; McMorran, R.; Wiegers, J.; Wiegers, T.C.; Mattingly, C.J. The Comparative Toxicogenomics Database: update 2019. Nucleic Acids Res 2019, 47, D948–D954. [Google Scholar] [CrossRef] [PubMed]
- Waters, M.D.; Fostel, J.M. Toxicogenomics and systems toxicology: aims and prospects. Nat Rev Genet 2004, 5, 936–948. [Google Scholar] [CrossRef]
- Pirmohamed, M. Pharmacogenomics: current status and future perspectives. Nat Rev Genet 2023, 24, 350–362. [Google Scholar] [CrossRef]
- Relling, M.V.; Evans, W.E. Pharmacogenomics in the clinic. Nature 2015, 526, 343–350. [Google Scholar] [CrossRef]
- Donnelly, M.J.; Corbel, V.; Weetman, D.; Wilding, C.S.; Williamson, M.S.; Black, W.C. Does kdr genotype predict insecticide-resistance phenotype in mosquitoes? Trends Parasitol 2009, 25, 213–219. [Google Scholar] [CrossRef]
- Estep, A.S.; Sanscrainte, N.D.; Waits, C.M.; Bernard, S.J.; Lloyd, A.M.; Lucas, K.J.; Buckner, E.A.; Vaidyanathan, R.; Morreale, R.; Conti, L.A.; et al. Quantification of permethrin resistance and kdr alleles in Florida strains of Aedes aegypti (L.) and Aedes albopictus (Skuse). PLoS Negl Trop Dis 2018, 12, e0006544. [Google Scholar] [CrossRef]
- Estep, A.S.; Sanscrainte, N.D.; Farooq, M.; Lucas, K.J.; Heinig, R.L.; Norris, E.J.; Becnel, J.J. Impact of Aedes aegypti V1016I and F1534C knockdown resistance genotypes on operational interventions. Sci Rep 2025, 15, 10146. [Google Scholar] [CrossRef]
| Primer name | Sequence |
| kdr_1014F [13] | TTCACGCTGGAATACTCACGACA |
| kdr_1014L [13] | GGGCGGCGGGCAGGGCGGCGGGGGCGGGGTTCACGCTGGAATACTCACGACTA |
| kdr_1014S [13] | AGCGCGGAGCGCGGTTCACGCTGGAATACTCACGACTG |
| kdr_1014r [13] | GGATCGAATCCATGTGGGACTGCAT |
| AchE_2340S | CCGGCAGGCCGACGGCGACGACTGTGGATCTTCGGGGTTA |
| AchE_2340G | CTGTGGATCTTCGGGGGTG |
| AchE_2362_r | GTGGTCGTACACGTCCAGCG |
| Cxq/n_Cxq | GCGGGCAGGGCGGCGGGGGCGGGGGGAGCTCCAGATATGGCCTTT |
| Cxq/n_Cxn | GGAGCTCCTGATATAGCTTTC |
| Cxq/n_r | ATGAAGGAGGTAGTATTCAAAAACTTAT |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




