1. Introduction
The lens epithelium-derived growth factor p75 (LEDGF/p75) is a chromatin-associated protein that plays a central role in transcriptional regulation and viral integration through its N-terminal PWWP domain. This domain mediates chromatin engagement by recognizing di- and trimethylated lysine 36 on histone H3 (H3K36me2/3) in conjunction with interactions with nucleosomal DNA. Beyond its physiological role, LEDGF/p75 is co-opted by HIV-1 integrase to direct viral genome integration into actively transcribed regions [
1,
2,
3,
4]. Dysregulation of LEDGF/p75 binding partners, such as MLL fusion proteins, has been implicated in oncogenic transcriptional programs [
5,
6]. As such, the LEDGF PWWP domain has emerged as a target of interest for epigenetic drug discovery [
7,
8].
Structural studies have shown that the LEDGF PWWP domain engages chromatin by recognizing H3K36me2/3-modified histones, forming specific interactions with the methylated lysine, while also interacting with nucleosomal DNA [
9,
10,
11]. These multivalent contacts are critical for chromatin targeting [
11,
12,
13]. Small-molecule ligands designed to target the H3K36me2/3 pocket typically exhibit only low micromolar affinity and attempts to extend binding into adjacent regions have met with limited success [
14]. Together, these observations highlight the need for alternative molecular tools to probe the functional surfaces of the PWWP domain.
Single-domain antibodies, also known as nanobodies (Nbs), offer unique advantages as high-affinity molecular probes for challenging protein surfaces. Their comparatively small size, high solubility, and generally long complementarity-determining region 3 (CDR3) loops enable them to recognize concave, shallow, or cryptic epitopes that are often poorly tractable by both conventional antibodies and small-molecule ligands [
15,
16,
17,
18]. Although Nbs are not generally suited for intracellular therapeutic delivery [
19], they are valuable as structural chaperones, conformational stabilizers, and functional inhibitors in biochemical and cellular studies [
18,
20].
Here, we report on the generation and characterization of a panel of Nbs targeting the LEDGF PWWP domain. Using animal immunization followed by phage display selection, we identified a diverse set of Nbs which bind the LEDGF PWWP domain with nanomolar affinity. We also characterized their solution properties and cross-reactivity with the closely related HRP-2 PWWP domain. While co-crystal structures of Nb-PWWP complexes could not be obtained, one Nb was found to inhibit the interaction between full-length LEDGF/p75 and H3K36me3-modified nucleosomes in vitro. The developed Nbs are convenient tools for probing the LEDGF PWWP domain and provide a foundation for mechanistic functional studies and structure-guided ligand development.
2. Materials and Methods
2.1. Protein Constructs
The human LEDGF PWWP domain (UniProt: O75475, residues 1-110) was cloned in a pET16b vector encoding an N-terminal His10-tag. A Ser-to-Cys mutation at position 62 (S62C) was introduced using Quik-Change site-directed mutagenesis to enable site-specific biotinylation on the face opposite the H3K36me2/3 pocket. Primers were manually designed (forward: TT CCT TAC TGCGAA AAT AAG GAA AAG TAT GGC AAA C, reverse: TT ATT TTC GCA GTA AGG AAA TAT ATC CTT TGG TC) and ordered from Integrated DNA Technologies (Leuven, Belgium).
A shorter LEDGF PWWP construct (residues 1-90) was cloned into a pETSUK2 vector encoding an N-terminal His
6-tag followed by the SUMO domain as described previously [
26].
Human HRP-2 PWWP (UniProt: Q7Z4V5, residues 1-93) and its C64S mutant were prepared as described previously [
14].
After Nb generation, selection and sequencing (section 4.4), nine representative Nb sequences were subcloned from the His6-FLAG3 tag vector into the pETSUK2 vector using the following primers: forward, GCGAACAGATTGGTGGTGGTGAGGTGCAATTGGTGGAGTCT; and reverse, TTGTTAGCAGAAGCTTATTATGAGGAGACGGTGACCTGG.
FLAG-LEDGF/p75 was expressed from pCPnat-FLAG
3-LEDGF/p75 in
E. coli BL21 Star (DE3). Cultures were grown in LB medium containing 100 µg/mL ampicillin at 37 °C until to OD
600 around 0.6, induced with 0.5 mM IPTG, and incubated for 3-4 h at 29 °C before harvesting. All other constructs were overexpressed in heat-shock transformed
E. coli Rosetta
TM 2 (DE3) pLysS competent cells (Merck) using auto-induction in ZYP-5052 medium [
27,
28]. Cells were harvested by centrifugation and stored at -80 °C.
2.2. Protein Purification
Unless otherwise stated, proteins were purified using a four-step protocol consisting of IMAC, tag cleavage followed by sIMAC, ion-exchange chromatography (IEX) and SEC. For the LEDGF PWWP (residues 1-110) construct and the S62C mutant, the His
10-tag was retained and the sIMAC step omitted. For HRP-2 PWWP and the LEDGF PWWP S62C mutant, 10 mM DTT was included in all IEX and SEC buffers. Reducing agents were omitted during Nb purification, and the IEX step was skipped; instead, Nb samples were dialyzed directly into SEC buffer. Buffer compositions are listed in
Table S3.
Cell pellets were thawed on ice and resuspended in low-imidazole buffer (50 mM sodium phosphate pH 7.5, 250 mM NaCl, 12.5 mM imidazole pH 7.5 and 5 mM b-mercaptoethanol), supplemented with SigmaFast protease inhibitor cocktail (1 tablet/100 mL; Merck) and CryonaseTM Cold-Active Nuclease (1 U/mL; Takara Bio Europe SAS). Cells were lysed by sonication and clarified by centrifugation.
The supernatant was loaded onto a His-Select® Nickel Affinity Gel resin (Merck) column pre-equilibrated with low-imidazole buffer. The column was subsequently washed with low-imidazole buffer supplemented with 0.1% v/v TritonTM X-100, and the target protein was eluted with high-imidazole buffer (50 mM sodium phosphate pH 7.5, 250 mM NaCl, 500 mM imidazole pH 7.5 and 5 mM b-mercaptoethanol). For His6-SUMO tagged constructs, tag cleavage was performed overnight with SUMO Hydrolase 7K (1:500) during dialysis (3 kDa) against low-imidazole buffer, followed by sIMAC to remove the cleaved tag. For constructs subjected to IEX, samples were dialyzed into a low-salt buffer supplemented with 10% w/v glycerol. All subsequent chromatographic steps were carried out on an ÄktaTM Pure system with UV detector (GE Healthcare). IEX was performed on HiTrap® SP HP or Q HP columns (5 mL; Cytiva), equilibrated in low-salt buffer, and proteins were eluted using a linear NaCl gradient with increasing salt concentration. Protein fractions were pooled, concentrated using Amicon® Ultra filters (3 kDa cut-off) (Merck Millipore). SEC was performed on a Superdex 75 16/60 GL column (GE Healthcare) equilibrated with SEC buffer. Protein fractions were concentrated, flash-frozen in liquid nitrogen, and stored at -80 °C.
FLAG-LEDGF/p75 was purified using heparin affinity chromatography followed by SEC. Pellets were resuspended in 500 mM NaCl, 30 mM Tris-HCl pH 7.4, 1 mM DTT, cOmplete protease inhibitor cocktail (Roche) and DNase I (Roche) for lysis by sonication. The lysate was clarified by centrifugation and filtered through a 0.22 µm syringe filter before loading onto a 5 mL HiTrap Heparin HP column equilibrated in 150 mM NaCl, 30 mM Tris-HCl pH 7.0, 1 mM DTT. Bound protein was eluted with a linear salt gradient (150 mM-2 M NaCl). Peak fractions were pooled and further purified by SEC on a Superose™ 6 10/300 GL column equilibrated in 150 mM NaCl, 30 mM Tris-HCl pH 7.4, 1 mM DTT. Fractions containing FLAG-LEDGF/p75 were supplemented with 10% (v/v) glycerol, flash-frozen, and stored at -80 °C.
For phage display and the Nb selection AlphaScreen, the LEDGF PWWP S62C mutant was biotinylated using the EZ-LinkTM Maleimide Protein Labeling Kit (Thermo Scientific).
2.3. Immunization and Nanobody Library Preparation
One llama and one alpaca were immunized with recombinant LEDGF PWWP domain (residues 1-110). Animals received four injections at two-week intervals. Nb libraries were generated as described previously [
29]. Peripheral blood mononuclear cells were isolated, total RNA was extracted, and first-strand cDNA synthesized using oligo(dT) primers. Nb-encoding sequences were amplified by nested PCR and cloned into an in-house phagemid vector. Recombinant phagemids were transformed into
E. coli TG1 cells, and Nb-displaying phages were produced using M13KO7 helper phage.
2.4. Nanobody Selection
Two rounds of phage display panning were performed using 100 nM biotinylated LEDGF PWWP. Streptavidin-coated Pierce™ beads were used in the first round and Dynabeads™ in the second. Following enrichment, Nb sequences were subcloned into an expression vector encoding an OmpA signal peptide and a C-terminal FLAG3-His₆ tag and expressed in E. coli TG1.
Periplasmic extracts were screened by AlphaScreen using 2 nM biotinylated LEDGF PWWP and anti-FLAG acceptor beads (20 µg/mL). After incubation, streptavidin donor beads were added, and signals measured on an EnVision plate reader. Clones producing signals ≥3-fold above background were sequenced and clustered (≥91% identity), yielding nine clusters (
Figure S1).
2.5. Binding Studies
2.5.1. Size-Exclusion Chromatography
SEC binding studies were performed on a Superdex 200 10/300 GL column (GE Healthcare) equilibrated in 50 mM Tris-HCl pH 7.0 and 150 mM NaCl. Each Nb was injected individually (2 mg/mL). LEDGF PWWP (residues 1-90) was injected separately at a concentration corresponding to a 1:1.1 molar ratio relative to the Nb. For complex analysis, LEDGF PWWP and the respective Nbs were pre-mixed at these final concentrations to maintain a 1:1.1 molar ratio.
2.5.2. Surface Plasmon Resonance
SPR was used to characterize the molecular interaction between the Nbs and LEDGF using a BiacoreTM 8K instrument (Cytiva, Uppsala, Sweden). LEDGF PWWP (residues 1-90) was covalently coupled on the surface of eight flow cells of a CM5 Series S sensor chip (Cytiva) using standard amine coupling and a running buffer composed of 10 mM HEPES pH 7.4, 150 mM NaCl, 3 mM EDTA and 0.05% Tween 20. First, carboxymethylated groups of the chip were activated for 7 min with a mixture of 200 mM 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide hydrochloride (EDC) and 50 mM N-hydroxysuccinimide (NHS) at 10 µL/min. Then, 2 µg/mL of LEDGF PWWP diluted in 10 mM HEPES pH 7 was injected for 6 min at 10 µL/min. The remaining activated groups were blocked with 1 M ethanolamine-HCl pH 8.5 for 7 min at 10 µL/min. The obtained LEDGF PWWP immobilization levels were about 200 RU. Eight reference flow cells without LEDGF PWWP were used as a control for non-specific binding and refractive index changes (i.e. one reference flow cell per LEDGF PWWP flow cell). All binding experiments were performed at 25 °C in 50 mM Tris-HCl pH 7.4 supplemented with 150 mM NaCl and 0.05% Tween 20. Three-fold serial dilutions (12.3, 37.0, 111.1, 333.3, and 1000 nM) of Nbs were sequentially injected from low to high concentration for 120 sec at a flow rate of 30 μL/min in one single cycle. The dissociation was monitored for 15 min and finally the surface was regenerated with 50 mM NaOH. Several buffer blanks were used for double referencing. The equilibrium dissociation constant KD were calculated after fitting the experimental data to the 1:1 Langmuir interaction model using the Biacore Insight Evaluation Software (version 6.0). All SPR experiments were performed in triplicate.
The interaction between the Nbs and the HRP-2 PWWP domain was analyzed using the same protocol, with immobilization levels ranging from 300 to 800 RU.
2.6. Protein Crystallization
Initial crystallization screening of the Nbs and their complexes with LEDGF PWWP (1-90) or HRP-2 PWWP C64S mutant (prepared in 1:1 molar ratio) was carried out using commercial crystallization kits (Molecular Dimensions, Hampton Research, Qiagen, and Rigaku) at 4 °C and 20 °C. Screening was performed using the sitting-drop method in Swissci 96-Well 2-drop plates (Hampton Research), with 75 µL reservoir solution per well. Two drops per condition were set up, consisting of either 150 nL + 150 nL or 200 nL + 100 nL protein and crystallization solution respectively using a Mosquito robot (SPT Labtech, Hertfordshire, UK). The individual protein concentrations ranged from 5 to 12 mg/mL. For crystallization of the complexes, the Nb and the LEDGF PWWP domain or the HRP-2 PWWP C64S mutant domain were mixed at a 1:1 molar ratio at low concentration, incubated to allow complex formation, and then concentrated together to the desired final concentration. Plates were sealed, stored and imaged by Rock Imager (Formulatrix, Bedford, MA, USA).
Optimizations of crystallization hits were performed using either sitting-drop with the same setup as initial screening or the hanging-drop vapor diffusion method in 24-well XRLTM plates (Molecular Dimensions). Hanging-drop crystallization involved 2 µL drops and 500 µL reservoir solution. NbC03 crystals were grown at 20 °C by sitting-drop vapor diffusion. Drops consisted of 150 nL protein solution (6.7 mg/mL) + 150 nL crystallization solution (1.5 M ammonium sulfate, 12% glycerol and 0.1 M Tris-HCl pH 8.5). NbH10 crystals were grown at 20 °C by sitting-drop vapor diffusion. NbH10 and LEDGF PWWP domain were mixed in a 1:1 molar ratio (12 mg/mL NbH10 final concentration). Drops consisted of 200 nL protein mixture + 100 nL crystallization solution (1 M succinic acid, 1% PEG MME 2000 and 0.1 M HEPES pH 7.0).
2.7. X-Ray Crystallography
Crystals were harvested, cryoprotected using crystallization solution supplemented with 10-20% ethylene glycol, and cryo-cooled in liquid nitrogen. X-ray data were collected at beamlines ID30A-3 [
30] and ID30B [
31] of the European Synchrotron Radiation Facility (ESRF) at 100 K using an X-ray wavelength close to 1 Å. Data were processed using autoPROC with STARANISO option [
32,
33] and phased by molecular replacement using PHASER [
34] and AlphaFold3 models of the Nbs [
35]. Structures were fitted in COOT [
36], refined in REFMAC5 [
37] and Phenix [
38], and analyzed using PyMOL (Version 3.0, Schrödinger, LLC). Crystallographic statistics are in
Table S1. Crystal interfaces were analyzed using the PISA software [
23].
2.8. AlphaFold3 Modeling of the Nanobody-LEDGF PWWP Complexes
The LEDGF PWWP domain and Nb sequences were submitted as paired inputs to the AlphaFold3 web server to predict the structure of the 1:1 complexes. Modeling was performed using default parameters, and the resulting models were evaluated using the ipTM score to assess the confidence of the predicted binding interface.
2.9. AlphaScreen Inhibition Assay
AlphaScreen in vitro inhibition assays were performed in a 384-well OptiPlate (PerkinElmer) with a final volume of 25 µL. Nbs were diluted in SEC buffer, while FLAG-LEDGF/p75 and reagents were diluted in AlphaScreen buffer (25 mM Tris-HCl pH 7.4, 150 mM NaCl, 1 mM MgCl2, 1 mM DTT, 0.1% Tween 20 (Thermo Fisher Scientific), and 0.1% bovine serum albumin (Sigma-Aldrich)). FLAG-LEDGF/p75 and varying Nb concentrations were pre-incubated for 30 minutes at 4 °C, followed by addition of H3K36me3-modified nucleosomes and incubation for 1 hour at 4 °C. Optimal concentrations determined by cross-titration were 10 nM FLAG-LEDGF/p75 and 5 nM nucleosomes. Donor and acceptor beads (10 µg/mL each; PerkinElmer) were then added, and plates were incubated for 1h at room temperature before reading on an Envision Xcite Multilabel Reader (PerkinElmer). Counts were plotted, and a non-linear regression sigmoidal curve fit was obtained using GraphPad Prism 10.6.1 (San Diego, CA, USA).
3. Results and Discussion
3.1. Generation and Selection of LEDGF PWWP-Specific Nanobodies
One llama and one alpaca were immunized with recombinant human LEDGF PWWP domain, and immune phage display libraries were generated from VHH repertoires. Following two rounds of panning against the antigen, 94 individual clones were screened for binding to the human LEDGF PWWP domain using periplasmic extracts in an AlphaScreen assay. Twenty-eight clones produced signals at least three-fold above the background. Sequencing yielded twelve distinct Nb sequences. Clustering analysis (≥91% sequence identity cut-off) revealed one cluster comprising three sequences (NbA03, NbB03 and NbG01) differing mainly in their CDRs, another cluster comprising two sequences (NbA02 and NbC02) which differ in one framework residue, and seven further singlets (
Figure S1).
3.2. Expression, Purification, and Solution Behavior of Selected Nanobodies
One representative Nb from each cluster (
Figure S1) was selected for large-scale expression as a His₆-SUMO fusion protein. After expression, NbA03 exhibited reduced solubility and partial precipitation after tag removal and was excluded from further analysis. The remaining eight Nbs were expressed at sufficient levels and purified using subtractive IMAC (sIMAC) with tag cleavage. In particular, NbH10 could be purified to >90% purity using sIMAC (
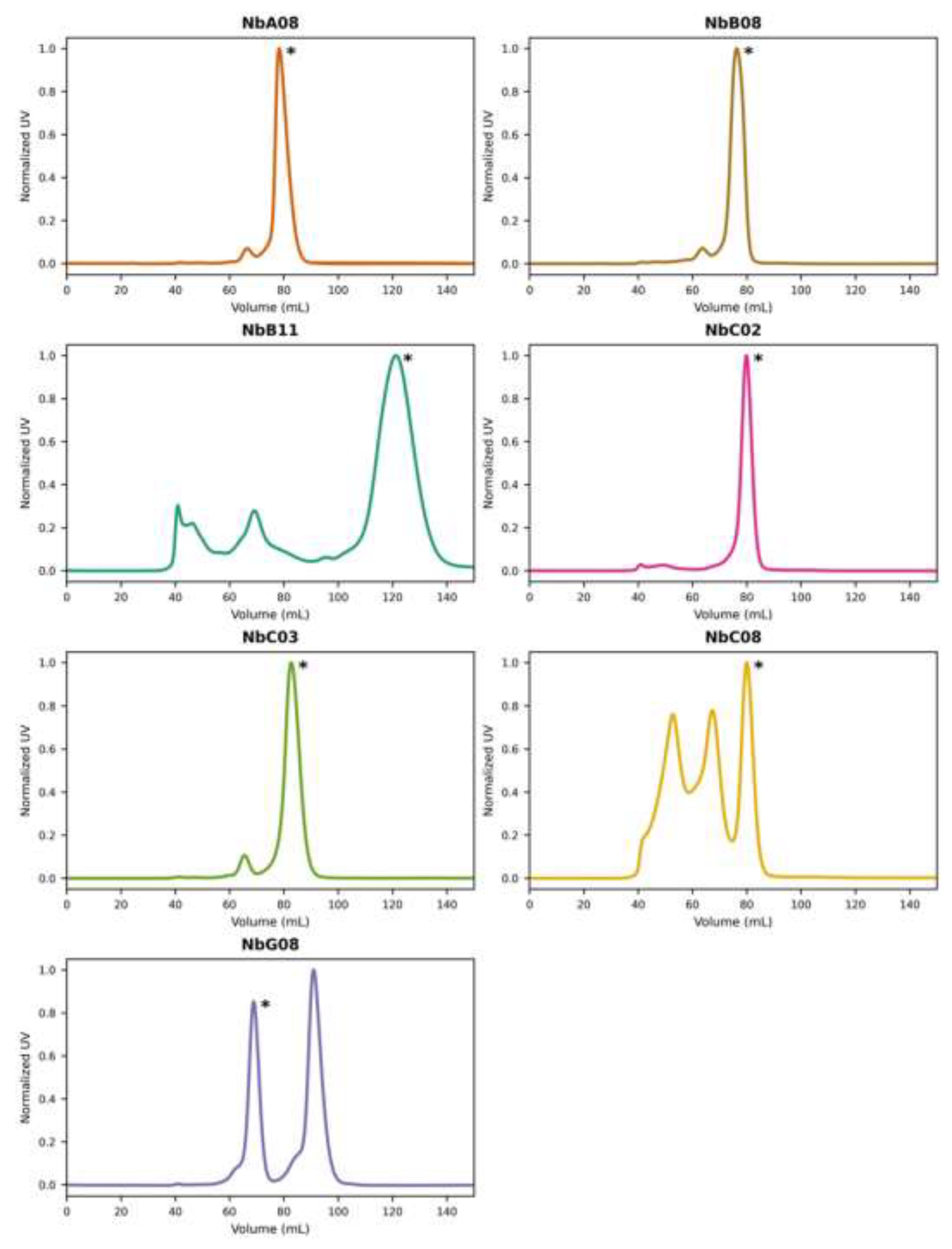
Figure S2A) but displayed limited solubility and was used as such for downstream assays. The remaining seven Nbs were further purified by SEC (
Figure 1 and
Figure S2B). SEC profiles for NbA08, NbB08, NbC02 and NbC03 revealed a dominant main peak at around 80 mL elution, presumably corresponding to a monomer. In contrast, NbB11, NbC08 and NbG08 displayed multiple SEC peaks. Judging by SDS-PAGE, all these peaks contained the target Nb, suggesting the presence of higher oligomeric species. For NbB11, the elution between 40 and 50 mL corresponded to impurities rather than the Nb. For NbG08, the presumed monomeric fraction precipitated after purification, requiring the use of the higher-oligomer fraction for subsequent experiments.
3.3. High-Affinity Binding of Nanobodies to the LEDGF PWWP Domain
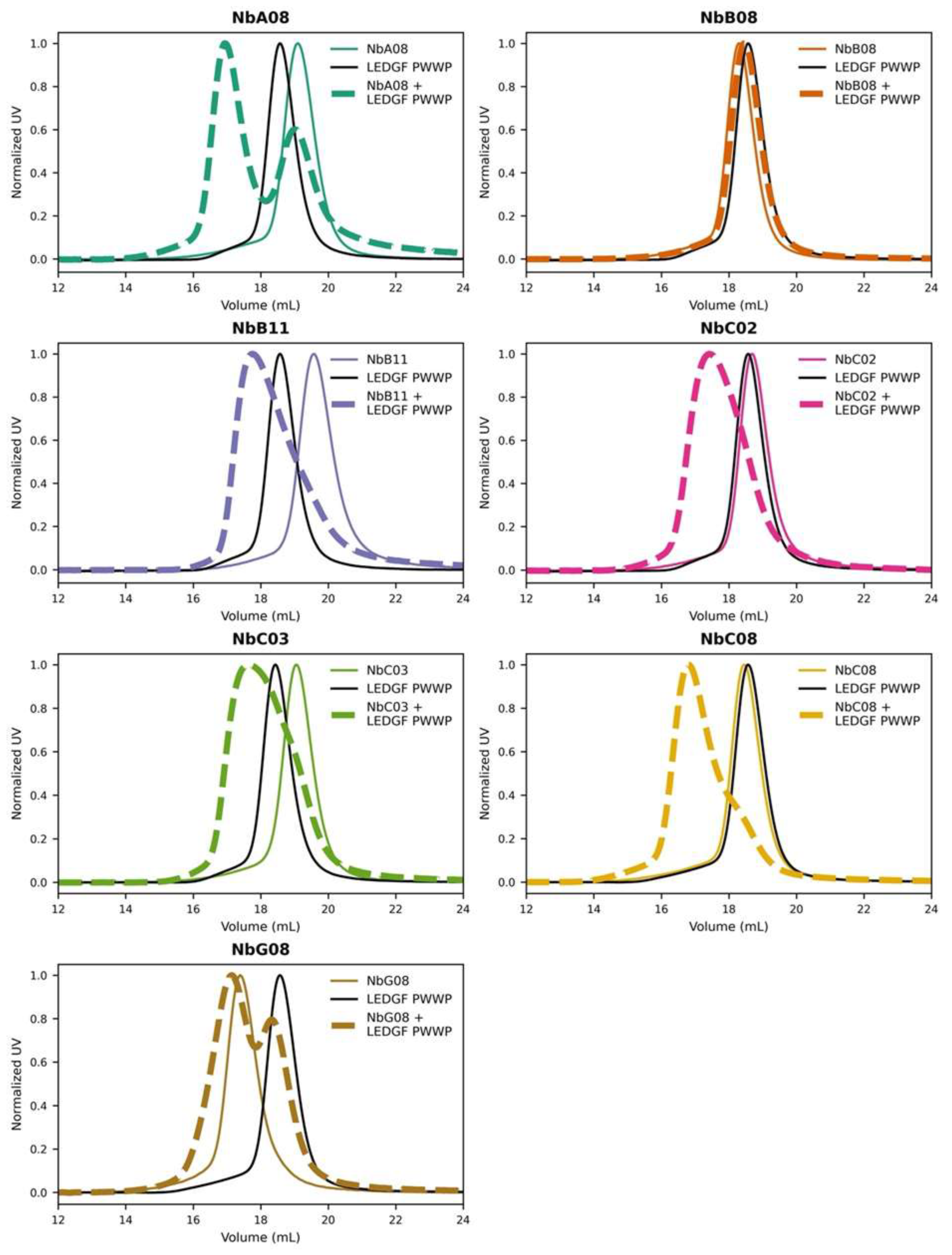
Towards an initial test of the binding, each Nb was mixed with LEDGF PWWP at a 1.1:1 molar ratio and analyzed on a Superdex 200 column. Four Nbs (NbB11, NbC02, NbC03, NbC08) revealed a single major elution peak located left of the elution of either individual protein, pointing to the complex formation (
Figure 2). NbA08 and NbG08 both showed considerable elution at the left-hand side but also individual components, which suggested partial complex formation. Finally, NbB08 did not exhibit any detectable complex under these conditions.
To quantify binding affinities, surface plasmon resonance (SPR) experiments were performed with immobilized LEDGF PWWP as ligand and Nbs as analytes. In agreement with SEC, NbB08 showed no measurable binding. The remaining seven Nbs bound to LEDGF PWWP with equilibrium dissociation constants (KD) ranging from 46 to 292 nM when fitted using a 1:1 interaction model (
Table 1). For several Nbs, including NbB11, NbC02 and NbH10, the fits showed some deviations from this model (
Figure S3), possibly caused by mass transport limitations, oligomerization, or protein heterogeneity on the sensor surface. At the same time, this model was consistent with the 1:1 complex formation observed in SEC.
3.4. Cross-Reactivity with the HRP-2 PWWP Domain
We additionally studied the interaction of the Nbs with the PWWP domain of hepatoma-derived growth factor-related protein 2 (HRP-2) which is a paralog of LEDGF. While the two domains have an overall sequence identity of 79%, their H3K36me2/3 pockets and their surroundings are nearly identical (
Figure S4). Our SPR experiments revealed that NbB11, NbC03, NbC08, NbG08 and NbH10 bound HRP-2 PWWP with affinities close to those observed for LEDGF PWWP (
Table 1). We therefore hypothesize that all these Nbs bind to the conserved regions. Given that the H3K36me2/3 pocket and its borders correspond to a major conserved patch, it is highly plausible that these Nbs are binding at or near the pocket, but additional data were needed.
In contrast, NbA08 and NbC02 showed substantially reduced affinities for HRP-2, suggesting their binding to LEDGF-specific surface features such as present outside the conserved H3K36me2/3 pocket and nucleosomal DNA binding surface (
Figure S4). Of note, as NbA08 and NbC02 recognize distinct epitopes between the two paralogs, these Nbs may be suitable for applications requiring LEDGF selectivity.
3.5. Structural Characterization of Nanobodies
Extensive crystallization trials were performed for the eight purified Nbs alone, for equimolar mixes with LEDGF PWWP domain, and for equimolar mixtures with HRP-2 PWWP. In two cases, NbC03 and NbH10-LEDGF PWWP mix, diffraction quality crystals could be obtained (
Table S1).
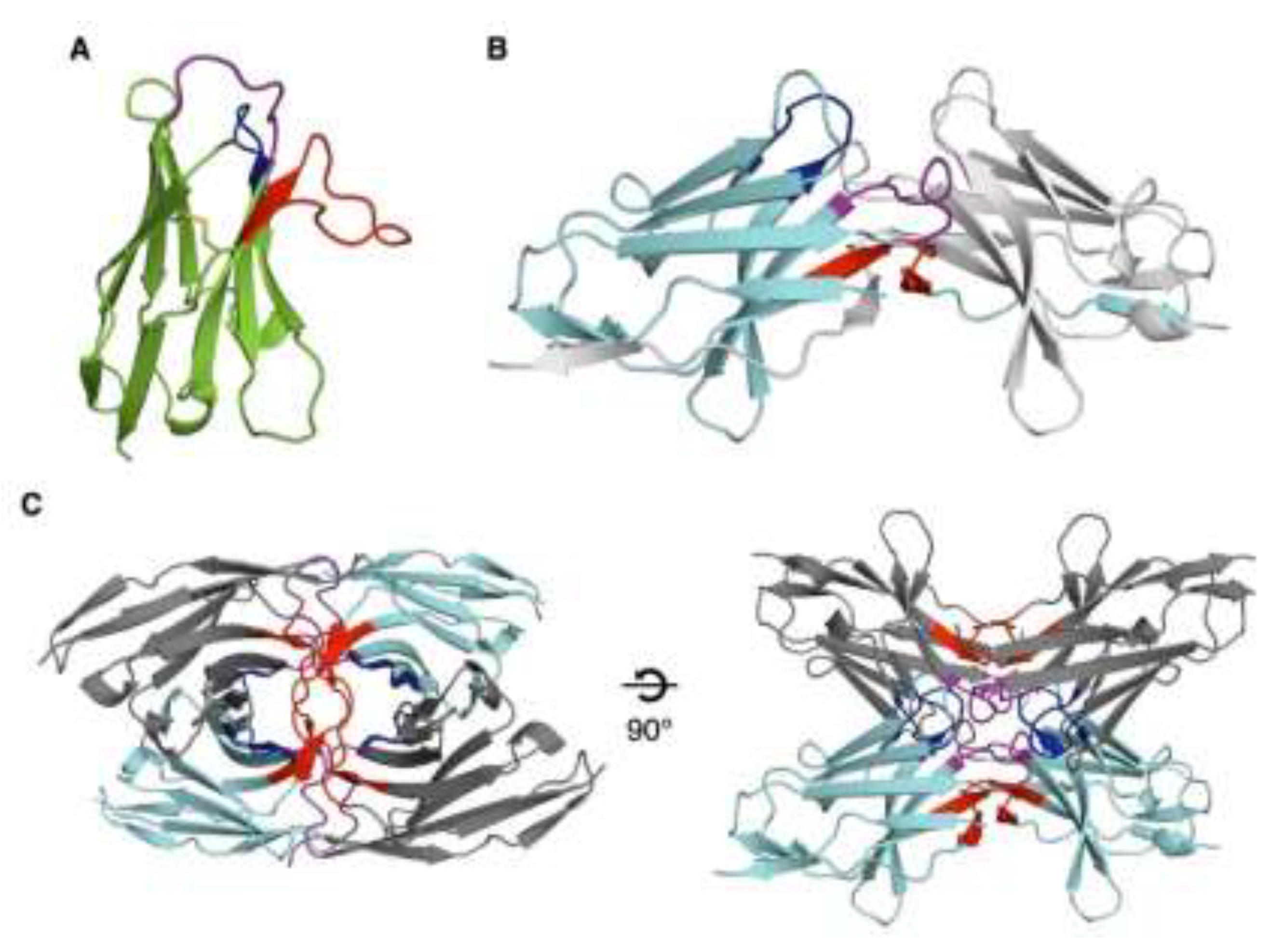
Crystals of NbC03 diffracted to 2.9 Å resolution. The asymmetric unit contained two Nb molecules, each adopting a canonical immunoglobulin fold stabilized by a conserved disulfide bond between Cys22 and Cys96 (
Figure 3A). The CDR1, CDR2, and CDR3 loops comprise eight, seven, and fourteen residues, respectively, confirming correct folding and typical Nb architecture.
Crystals grown from the NbH10-LEDGF PWWP mixture diffracted to 2.3 Å resolution, however structure determination revealed the presence of NbH10 alone. Two NbH10 molecules were present per asymmetric unit, each forming a domain-swapped dimer through exchange of the CDR3 loop and terminal b-strand via a two-fold crystallographic symmetry axis (
Figure 3B). Domain swapping has previously been reported for Nbs, both as a crystallization artifact [
21]—often promoted by high protein concentration, ionic strength, or short CDR3 loops—and, in some cases, as a functionally relevant assembly capable of antigen binding [
22]. The software PISA [
23] identified large interfaces both within the domain-swapped dimer and between the two chains in the asymmetric unit. The latter interface is mediated by both CDR1 and CDR2. All interfaces were predicted to be energetically favorable (
Table S2). Collectively, these intersubunit interactions yield a highly stable tetramer with 222 symmetry (
Figure 3C).
Given the lack of co-crystal structures, Nb-PWWP complexes were also modeled using AlphaFold3. However, all generated models exhibited low interface confidence scores (interface predicted template modeling score (ipTM) < 0.3), consistent with current limitations of structure prediction for antigen-recognizing protein-protein interfaces lacking strong co-evolutionary signals [
24].
3.6. NbH10 but not NbA08 Inhibits the LEDGF/p75-H3K36me3 Nucleosome Interaction In Vitro
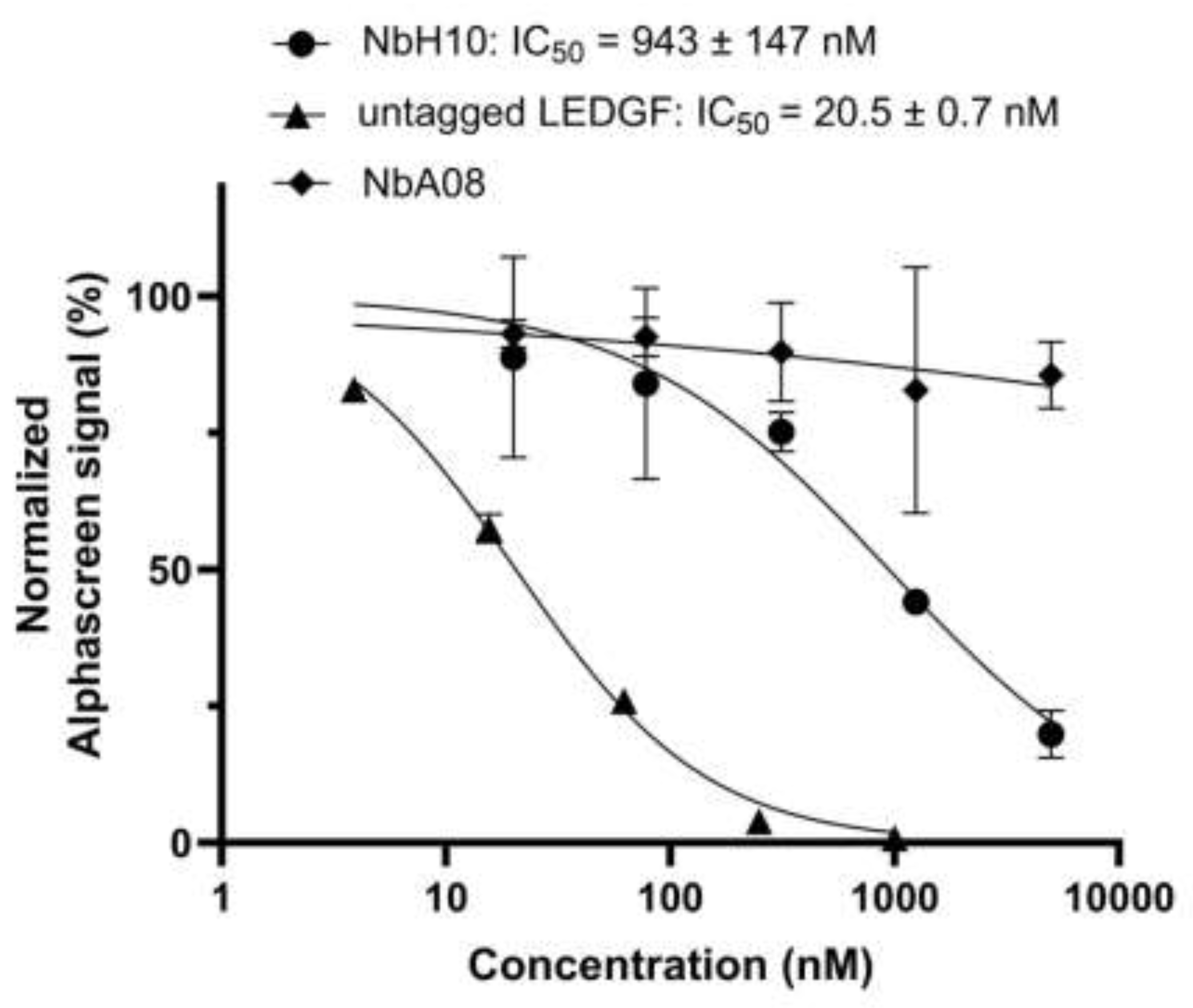
The functional impact of Nb binding was assessed using an AlphaScreen assay measuring the interaction between full-length LEDGF/p75 and H3K36me3-modified nucleosomes. Here, NbH10 produced a dose-dependent inhibition of this interaction with an IC
50 of 0.94 ± 0.15 mM (Figure 5). This demonstrates that NbH10 engages a functionally relevant surface of the PWWP domain involved in chromatin recognition. Given that this interface is fully conserved between LEDGF and HRP-2, this observation is well in line with the very similar affinities of ~100 nM measured for this Nb against both LEDGF and HRP-2 PWWP domains (
Table 1).
At the same time, the AlphaScreen assay revealed that NbA08 did not inhibit the interaction between full-length LEDGF and H3K36me3-modified nucleosomes. This observation is interesting given its high affinity (43 nM) for the isolated LEDGF PWWP domain, but a hundredfold lesser affinity for the HRP-2 domain (
Table 1). Taken together, these data make us conclude that NbA08 does not interfere with the nucleosome binding site but attaches to a distinct, non-functional surface.
Together, these results indicate that high-affinity binding alone is insufficient for functional inhibition and that NbH10 likely targets conserved determinants critical for chromatin engagement.
Figure 4.
NbH10 dose-dependently inhibits the LEDGF/p75-nucleosome interaction. AlphaScreen assays were performed using 10 nM FLAG-LEDGF/p75 and 5 nM biotinylated H3K36me3 nucleosome. Increasing concentrations of NbH10, NbA08, or untagged LEDGF/p75 were added to compete with FLAG-LEDGF/p75 for nucleosome binding. The normalized AlphaScreen signal (%) is plotted against inhibitor concentration on a logarithmic scale. Curves represent nonlinear regression fits (four-parameter logistic). Data points show mean ± standard deviation of three independent experiments.
Figure 4.
NbH10 dose-dependently inhibits the LEDGF/p75-nucleosome interaction. AlphaScreen assays were performed using 10 nM FLAG-LEDGF/p75 and 5 nM biotinylated H3K36me3 nucleosome. Increasing concentrations of NbH10, NbA08, or untagged LEDGF/p75 were added to compete with FLAG-LEDGF/p75 for nucleosome binding. The normalized AlphaScreen signal (%) is plotted against inhibitor concentration on a logarithmic scale. Curves represent nonlinear regression fits (four-parameter logistic). Data points show mean ± standard deviation of three independent experiments.
4. Conclusions
This study presents the first systematic generation and characterization of Nbs targeting the PWWP domain of LEDGF/p75. We show that as many as seven identified Nbs bind the isolated PWWP domain with affinity ranging between 43 and 292 nM, and also display paralog-selective properties. Our results demonstrate that this chromatin reader domain is readily targetable by Nbs, overcoming the previously encountered challenges with developing high-affinity small-molecule ligands [
14,
25]. Importantly, NbH10 was validated as a functional, submicromolar inhibitor of the LEDGF/p75-H3K36me3 nucleosome interaction
in vitro, which is also corroborated by its measured affinity to both the LEDGF and HRP-2 PWWP domains.
In summary, our study establishes a proof-of-principle that Nbs can interfere with PWWP-mediated chromatin engagement. Collectively, the Nb panel described here provides valuable molecular tools for probing LEDGF function. These results offer a foundation for future epitope-mapping, structural, and cellular studies targeting PWWP-dependent disease mechanisms.
Supplementary Materials
The following supporting information can be downloaded at the website of this paper posted on Preprints.org. Figure S1: Alignment of unique nanobody sequences identified and clustered. Figure S2: SDS-PAGES of nanobody samples. Figure S3: Representative SPR sensograms and theoretical binding profiles of the nanobodies over immobilized LEDGF PWWP and HRP-2 PWWP. Figure S4: Comparison of the PWWP domain sequences. Table S1: Crystallographic statistics. Table S2: Crystal interface analysis for the NbH10 crystals. Table S3: Buffers used for ion-exchange chromatography and size-exclusion chromatography of different protein constructs.
Author Contributions
T.V.: Conceptualization, Investigation, Formal analysis, Visualization, Funding acquisition, Writing – original draft. S.J.: Investigation, Writing – review and editing. T.Z.A.: Investigation, Funding acquisition. P.V.M.: Investigation, Funding acquisition. S.N.: Investigation. D.S.: Supervision. M.D.: Supervision, Writing – review and editing. Z.D.: Supervision, Funding acquisition, Writing – review and editing. S.V.S.: Conceptualization, Supervision, Funding acquisition, Writing – review and editing. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by grants G053822N (to S.V.S.) and I007220N (to S.V.S. and Z.D) from the Research Foundation Flanders (FWO), and by a grant C32/18/038 (to Z.D. and S.V.S.) from KU Leuven. FWO further granted doctoral fellowships to T.V. (1SC9822N, 1SC9824N), T.Z.A. (1S11223N), and P.V.M. (1SF0324N).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
X-ray diffraction data and atomic coordinates for NbC03 and NbH10 were deposited to the Protein Data Bank under the accession code 9TZZ and 9U00, respectively.
Acknowledgments
We are grateful to Noelia Gesteira Perez, Tom Jaspers and Flor Laenen for preparing the phage library, and to Dr. Zhongyao Zhang for help with the AlphaScreen assay. We also thank the staff of beamlines ID30A-3 and ID30B (European Synchrotron Radiation Facility, France) for their assistance with X-ray data collection.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| H3K36me2/3 |
di-or trimethylated Lys36 of histone H3 |
| IEX |
Ion-exchange chromatography |
| IMAC |
Immobilized metal affinity chromatography |
| KD
|
Dissociation constant |
| LEDGF/p75 |
Lens epithelium-derived growth factor p75 |
| Nbs |
Nanobodies |
| PWWP domain |
Pro-Trp-Trp-Pro domain |
| SEC |
Size-exclusion chromatography |
| sIMAC |
Subtractive IMAC |
| SPR |
Surface plasmon resonance |
| SUMO |
Small ubiquitin-like modifier |
References
- Cherepanov, P.; Ambrosio, A.L.B.; Rahman, S.; Ellenberger, T.; Engelman, A. Structural Basis for the Recognition between HIV-1 Integrase and Transcriptional Coactivator P75. Proc Natl Acad Sci U S A 2005, 102, 17308–17313. [Google Scholar] [CrossRef]
- Llano, M.; Saenz, D.T.; Meehan, A.; Wongthida, P.; Peretz, M.; Walker, W.H.; Teo, W.; Poeschla, E.M. An Essential Role for LEDGF/P75 in HIV Integration. Science 2006, 314, 461–464. [Google Scholar] [CrossRef] [PubMed]
- Busschots, K.; Voet, A.; De Maeyer, M.; Rain, J.-C.; Emiliani, S.; Benarous, R.; Desender, L.; Debyser, Z.; Christ, F. Identification of the LEDGF/P75 Binding Site in HIV-1 Integrase. Journal of Molecular Biology 2007, 365, 1480–1492. [Google Scholar] [CrossRef] [PubMed]
- Engelman, A.; Cherepanov, P. The Lentiviral Integrase Binding Protein LEDGF/P75 and HIV-1 Replication. PLoS Pathog 2008, 4, e1000046. [Google Scholar] [CrossRef]
- El Ashkar, S.; Schwaller, J.; Pieters, T.; Goossens, S.; Demeulemeester, J.; Christ, F.; Van Belle, S.; Juge, S.; Boeckx, N.; Engelman, A.; et al. LEDGF/P75 Is Dispensable for Hematopoiesis but Essential for MLL-Rearranged Leukemogenesis. Blood 2018, 131, 95–107. [Google Scholar] [CrossRef]
- Van Belle, S.; El Ashkar, S.; Čermáková, K.; Matthijssens, F.; Goossens, S.; Canella, A.; Hodges, C.H.; Christ, F.; De Rijck, J.; Van Vlierberghe, P.; et al. Unlike Its Paralog LEDGF/P75, HRP-2 Is Dispensable for MLL-R Leukemogenesis but Important for Leukemic Cell Survival. Cells 2021, 10, 192. [Google Scholar] [CrossRef]
- Cermakova, K.; Weydert, C.; Christ, F.; De Rijck, J.; Debyser, Z. Lessons Learned: HIV Points the Way Towards Precision Treatment of Mixed-Lineage Leukemia. Trends in Pharmacological Sciences 2016, 37, 660–671. [Google Scholar] [CrossRef]
- Blokken, J.; De Rijck, J.; Christ, F.; Debyser, Z. Protein–Protein and Protein–Chromatin Interactions of LEDGF/P75 as Novel Drug Targets. Drug Discov Today Technol 2017, 24, 25–31. [Google Scholar] [CrossRef]
- Wu, H.; Zeng, H.; Lam, R.; Tempel, W.; Amaya, M.F.; Xu, C.; Dombrovski, L.; Qiu, W.; Wang, Y.; Min, J. Structural and Histone Binding Ability Characterizations of Human PWWP Domains. PLoS One 2011, 6, e18919. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Farnung, L.; Dienemann, C.; Cramer, P. Structure of H3K36-Methylated Nucleosome–PWWP Complex Reveals Multivalent Cross-Gyre Binding. Nat Struct Mol Biol 2020, 27, 8–13. [Google Scholar] [CrossRef]
- Koutná, E.; Lux, V.; Kouba, T.; Škerlová, J.; Nováček, J.; Srb, P.; Hexnerová, R.; Šváchová, H.; Kukačka, Z.; Novák, P.; et al. Multivalency of Nucleosome Recognition by LEDGF. Nucleic Acids Res 2023, 51, 10011–10025. [Google Scholar] [CrossRef]
- Eidahl, J.O.; Crowe, B.L.; North, J.A.; McKee, C.J.; Shkriabai, N.; Feng, L.; Plumb, M.; Graham, R.L.; Gorelick, R.J.; Hess, S.; et al. Structural Basis for High-Affinity Binding of LEDGF PWWP to Mononucleosomes. Nucleic Acids Research 2013, 41, 3924–3936. [Google Scholar] [CrossRef] [PubMed]
- van Nuland, R.; van Schaik, F.M.; Simonis, M.; van Heesch, S.; Cuppen, E.; Boelens, R.; Timmers, H.M.; van Ingen, H. Nucleosomal DNA Binding Drives the Recognition of H3K36-Methylated Nucleosomes by the PSIP1-PWWP Domain. Epigenetics Chromatin 2013, 6, 12. [Google Scholar] [CrossRef]
- Vantieghem, T.; Aslam, N.A.; Osipov, E.M.; Akele, M.; Van Belle, S.; Beelen, S.; Drexler, M.; Paulovcakova, T.; Lux, V.; Fearon, D.; et al. Rational Fragment-Based Design of Compounds Targeting the PWWP Domain of the HRP Family. European Journal of Medicinal Chemistry 2024, 280, 116960. [Google Scholar] [CrossRef]
- Hamers-Casterman, C.; Atarhouch, T.; Muyldermans, S.; Robinson, G.; Hammers, C.; Songa, E.B.; Bendahman, N.; Hammers, R. Naturally Occurring Antibodies Devoid of Light Chains. Nature 1993, 363, 446–448. [Google Scholar] [CrossRef]
- De Genst, E.; Silence, K.; Decanniere, K.; Conrath, K.; Loris, R.; Kinne, J.; Muyldermans, S.; Wyns, L. Molecular Basis for the Preferential Cleft Recognition by Dromedary Heavy-Chain Antibodies. Proc. Natl. Acad. Sci. U.S.A. 2006, 103, 4586–4591. [Google Scholar] [CrossRef]
- Muyldermans, S. Nanobodies: Natural Single-Domain Antibodies. Annu. Rev. Biochem. 2013, 82, 775–797. [Google Scholar] [CrossRef]
- Pardon, E.; Laeremans, T.; Triest, S.; Rasmussen, S.G.F.; Wohlkönig, A.; Ruf, A.; Muyldermans, S.; Hol, W.G.J.; Kobilka, B.K.; Steyaert, J. A General Protocol for the Generation of Nanobodies for Structural Biology. Nat Protoc 2014, 9, 674–693. [Google Scholar] [CrossRef] [PubMed]
- Silva-Pilipich, N.; Smerdou, C.; Vanrell, L. A Small Virus to Deliver Small Antibodies: New Targeted Therapies Based on AAV Delivery of Nanobodies. Microorganisms 2021, 9, 1956. [Google Scholar] [CrossRef] [PubMed]
- Dingus, J.G.; Tang, J.C.; Amamoto, R.; Wallick, G.K.; Cepko, C.L. A General Approach for Stabilizing Nanobodies for Intracellular Expression. eLife 2022, 11, e68253. [Google Scholar] [CrossRef]
- Spinelli, S.; Desmyter, A.; Frenken, L.; Verrips, T.; Tegoni, M.; Cambillau, C. Domain Swapping of a Llama VHH Domain Builds a Crystal-wide Β-sheet Structure. FEBS Letters 2004, 564, 35–40. [Google Scholar] [CrossRef]
- Gallant, J.P.; Hicks, D.; Shi, K.; Moeller, N.H.; Hoppe, B.; Lake, E.W.; Baehr, C.; Pravetoni, M.; Aihara, H.; LeBeau, A.M. Identification and Biophysical Characterization of a Novel Domain-Swapped Camelid Antibody Specific for Fentanyl. Journal of Biological Chemistry 2024, 300, 107502. [Google Scholar] [CrossRef]
- Krissinel, E.; Henrick, K. Inference of Macromolecular Assemblies from Crystalline State. Journal of Molecular Biology 2007, 372, 774–797. [Google Scholar] [CrossRef]
- McCoy, K.M.; Ackerman, M.E.; Grigoryan, G. A Comparison of Antibody–Antigen Complex Sequence-to-structure Prediction Methods and Their Systematic Biases. Protein Science 2024, 33, e5127. [Google Scholar] [CrossRef] [PubMed]
- Shell, D.J.; Rectenwald, J.M.; Buttery, P.H.; Johnson, R.L.; Foley, C.A.; Guduru, S.K.R.; Uguen, M.; Rubiano, J.S.; Zhang, X.; Li, F.; et al. Discovery of Hit Compounds for Methyl-Lysine Reader Proteins from a Target Class DNA-Encoded Library. SLAS Discov 2022, 27, 428–439. [Google Scholar] [CrossRef] [PubMed]
- Weeks, S.D.; Drinker, M.; Loll, P.J. Ligation Independent Cloning Vectors for Expression of SUMO Fusions. Protein Expr Purif 2007, 53, 40–50. [Google Scholar] [CrossRef]
- Studier, F.W. Protein Production by Auto-Induction in High-Density Shaking Cultures. Prot Expr Purif 2005, 41, 207–234. [Google Scholar] [CrossRef]
- Studier, F.W. Stable Expression Clones and Auto-Induction for Protein Production in E. Coli. Methods Mol Biol 2014, 1091, 17–32. [Google Scholar] [CrossRef] [PubMed]
- Wouters, Y.; Jaspers, T.; De Strooper, B.; Dewilde, M. Identification and in Vivo Characterization of a Brain-Penetrating Nanobody. Fluids Barriers CNS 2020, 17, 62. [Google Scholar] [CrossRef] [PubMed]
- Von Stetten, D.; Carpentier, P.; Flot, D.; Beteva, A.; Caserotto, H.; Dobias, F.; Guijarro, M.; Giraud, T.; Lentini, M.; McSweeney, S.; et al. ID30A-3 (MASSIF-3) – a Beamline for Macromolecular Crystallography at the ESRF with a Small Intense Beam. J Synchrotron Rad 2020, 27, 844–851. [Google Scholar] [CrossRef] [PubMed]
- McCarthy, A.A.; Barrett, R.; Beteva, A.; Caserotto, H.; Dobias, F.; Felisaz, F.; Giraud, T.; Guijarro, M.; Janocha, R.; Khadrouche, A.; et al. ID30B – a Versatile Beamline for Macromolecular Crystallography Experiments at the ESRF. J Synchrotron Rad 2018, 25, 1249–1260. [Google Scholar] [CrossRef] [PubMed]
- Vonrhein, C.; Flensburg, C.; Keller, P.; Sharff, A.; Smart, O.; Paciorek, W.; Womack, T.; Bricogne, G. Data Processing and Analysis with the autoPROC Toolbox. Acta Crystallogr D Biol Crystallogr 2011, 67, 293–302. [Google Scholar] [CrossRef] [PubMed]
- Vonrhein, C.; Tickle, I.J.; Flensburg, C.; Keller, P.; Paciorek, W.; Sharff, A.; Bricogne, G. Advances in Automated Data Analysis and Processing within autoPROC, Combined with Improved Characterisation, Mitigation and Visualisation of the Anisotropy of Diffraction Limits Using STARANISO. Acta Crystallogr A Found Adv 2018, 74, a360–a360. [Google Scholar] [CrossRef]
- McCoy, A.J. Solving Structures of Protein Complexes by Molecular Replacement with Phaser. Acta Crystallogr D Biol Crystallogr 2007, 63, 32–41. [Google Scholar] [CrossRef]
- Abramson, J.; Adler, J.; Dunger, J.; Evans, R.; Green, T.; Pritzel, A.; Ronneberger, O.; Willmore, L.; Ballard, A.J.; Bambrick, J.; et al. Accurate Structure Prediction of Biomolecular Interactions with AlphaFold 3. Nature 2024, 630, 493–500. [Google Scholar] [CrossRef]
- Emsley, P.; Cowtan, K. Coot: Model-Building Tools for Molecular Graphics. Acta Crystallogr D Biol Crystallogr 2004, 60, 2126–2132. [Google Scholar] [CrossRef]
- Murshudov, G.N.; Skubák, P.; Lebedev, A.A.; Pannu, N.S.; Steiner, R.A.; Nicholls, R.A.; Winn, M.D.; Long, F.; Vagin, A.A. REFMAC5 for the Refinement of Macromolecular Crystal Structures. Acta Crystallogr D Biol Crystallogr 2011, 67, 355–367. [Google Scholar] [CrossRef]
- Liebschner, D.; Afonine, P.V.; Baker, M.L.; Bunkóczi, G.; Chen, V.B.; Croll, T.I.; Hintze, B.; Hung, L.-W.; Jain, S.; McCoy, A.J.; et al. Macromolecular Structure Determination Using X-Rays, Neutrons and Electrons: Recent Developments in Phenix. Acta Crystallogr D Struct Biol 2019, 75, 861–877. [Google Scholar] [CrossRef]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).