Submitted:
23 February 2026
Posted:
25 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
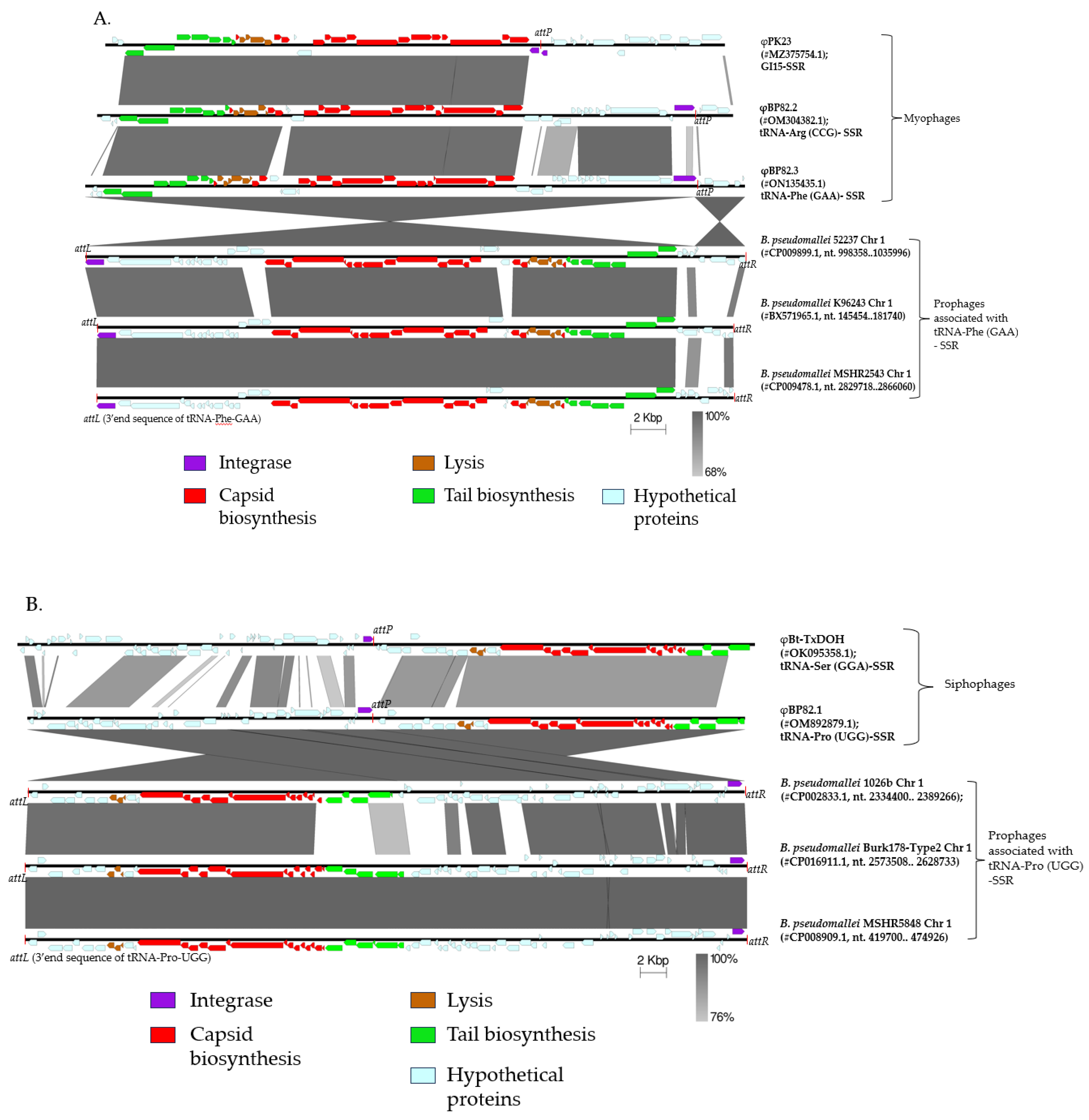
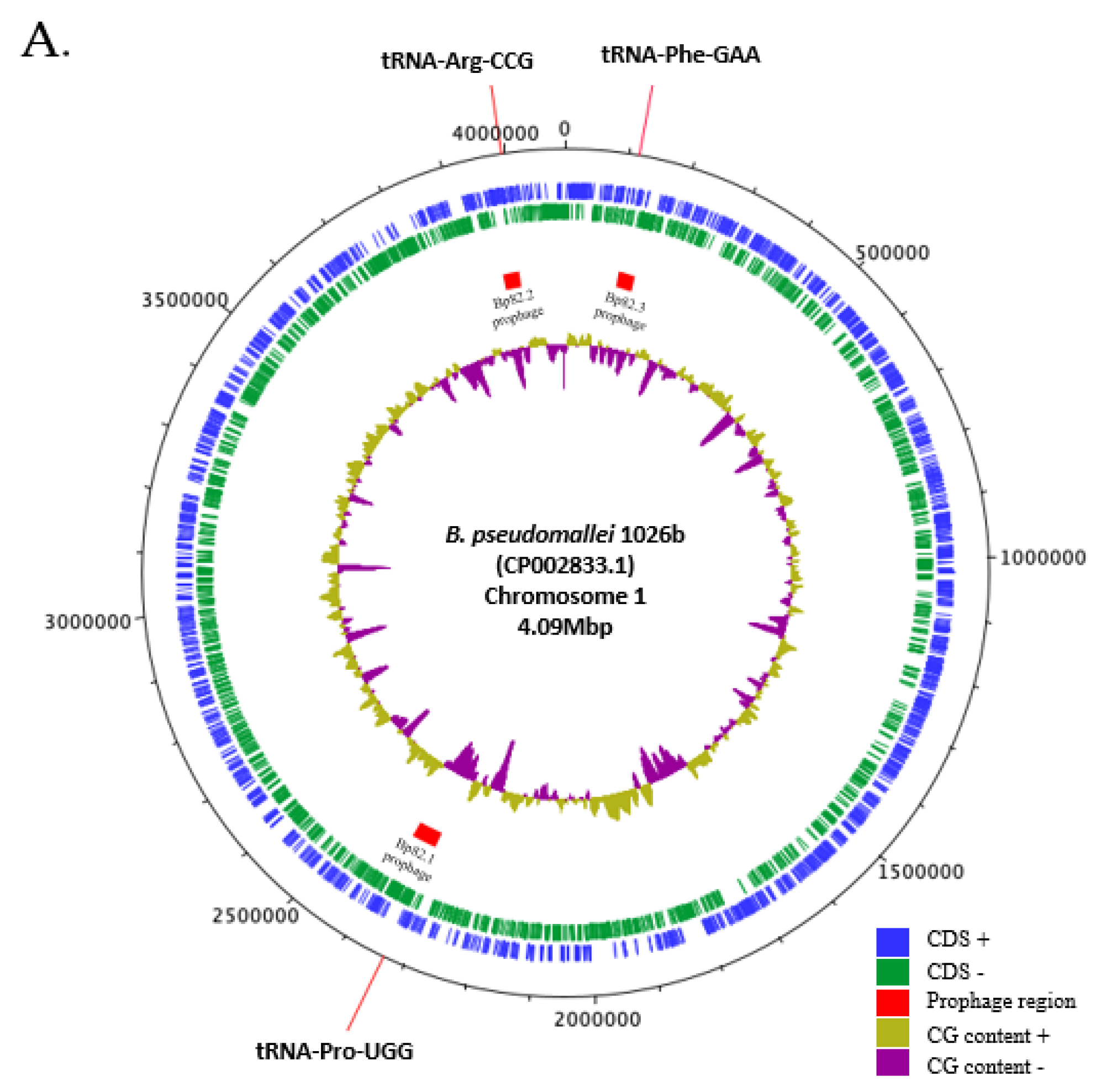
2.1. Genomic Analysis of B. pseudomallei Prophages
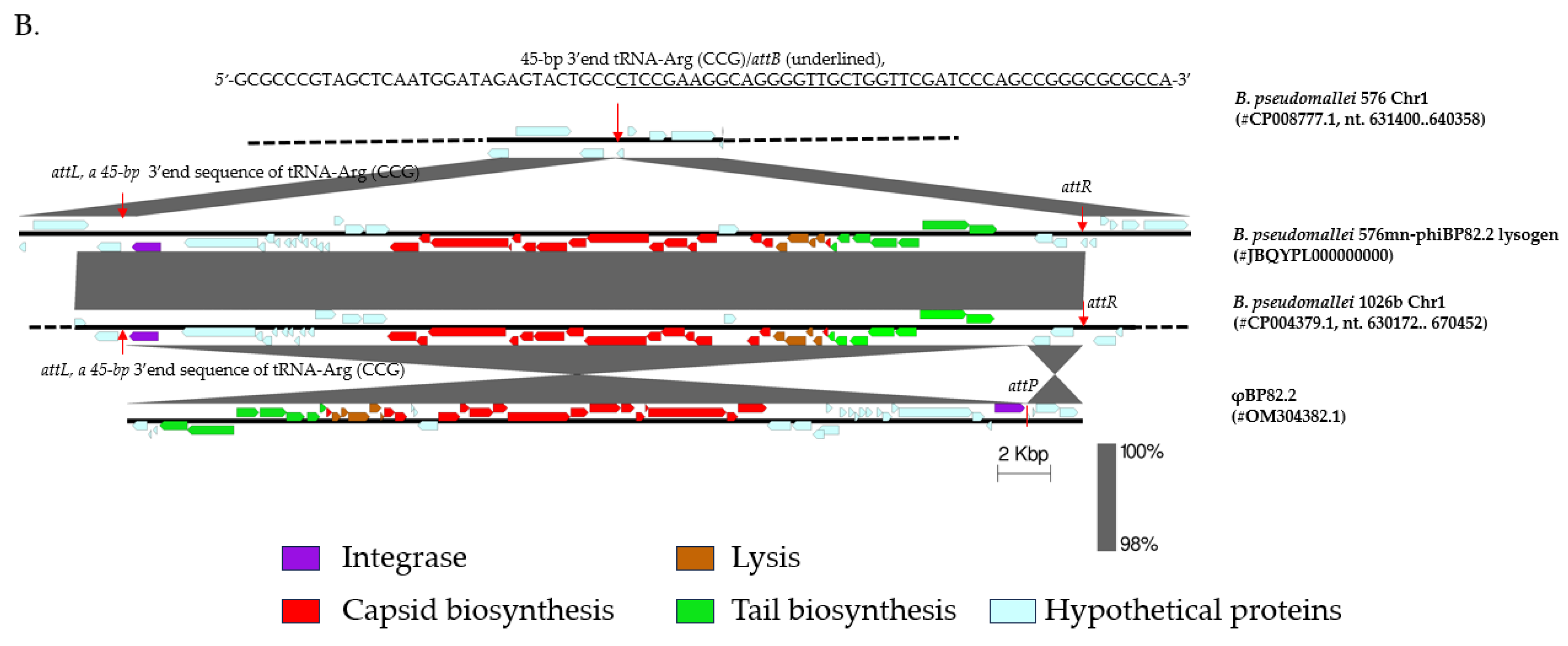
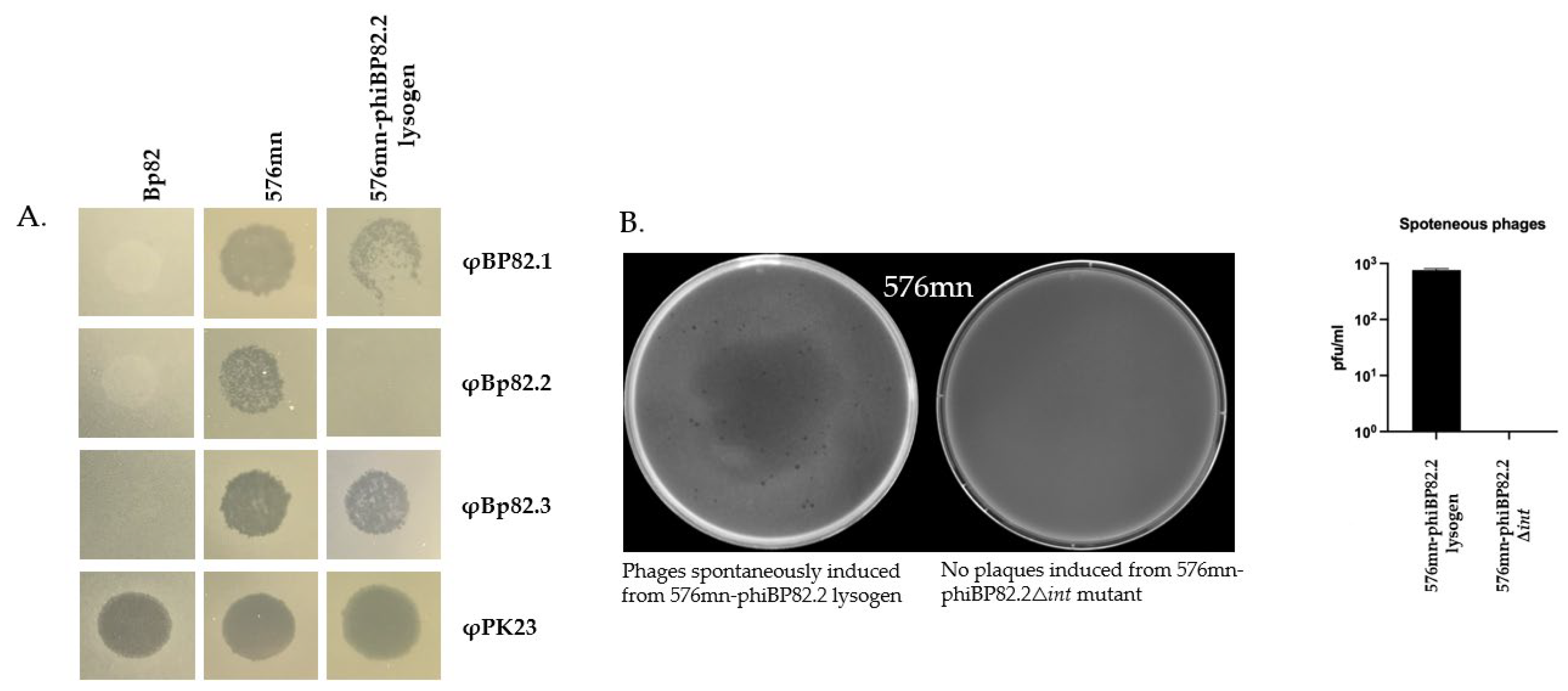
2.2. Isolation of a Lysogenic Clone of B. pseudomallei 576mn Following φBP82.2 Infection
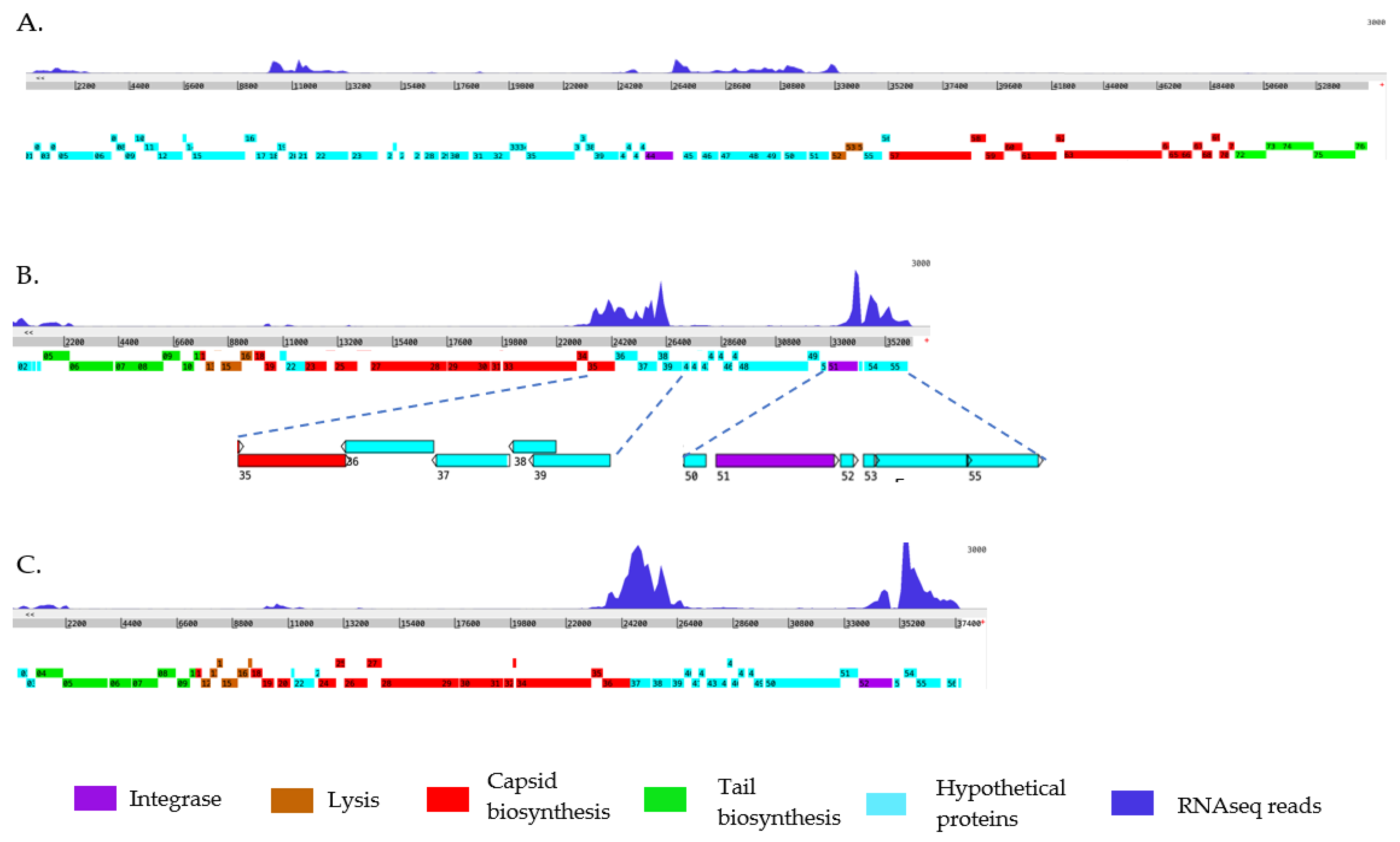
2.3. Expression Profile of the Prophages in B. pseudomallei Bp82 During Normal Exponential Growth
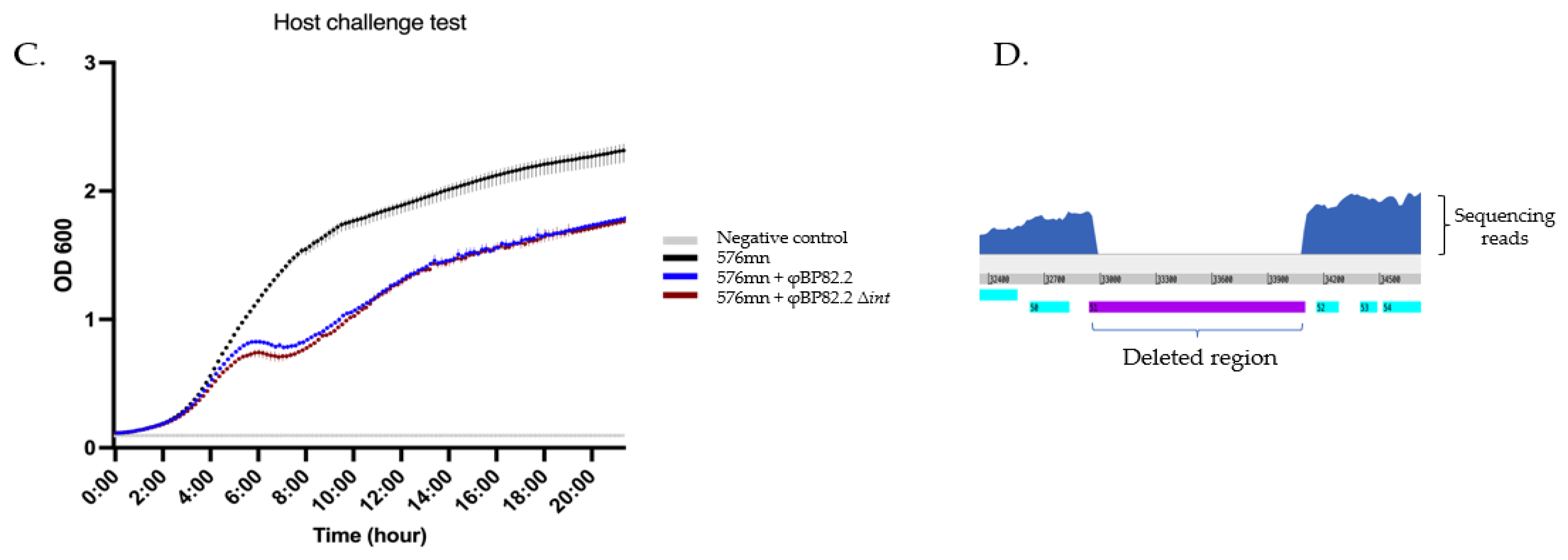
2.4. Deletion of the Integrase Gene gp51 in the φBP82.2 Prophage Resulted in the Loss of Phage Excision
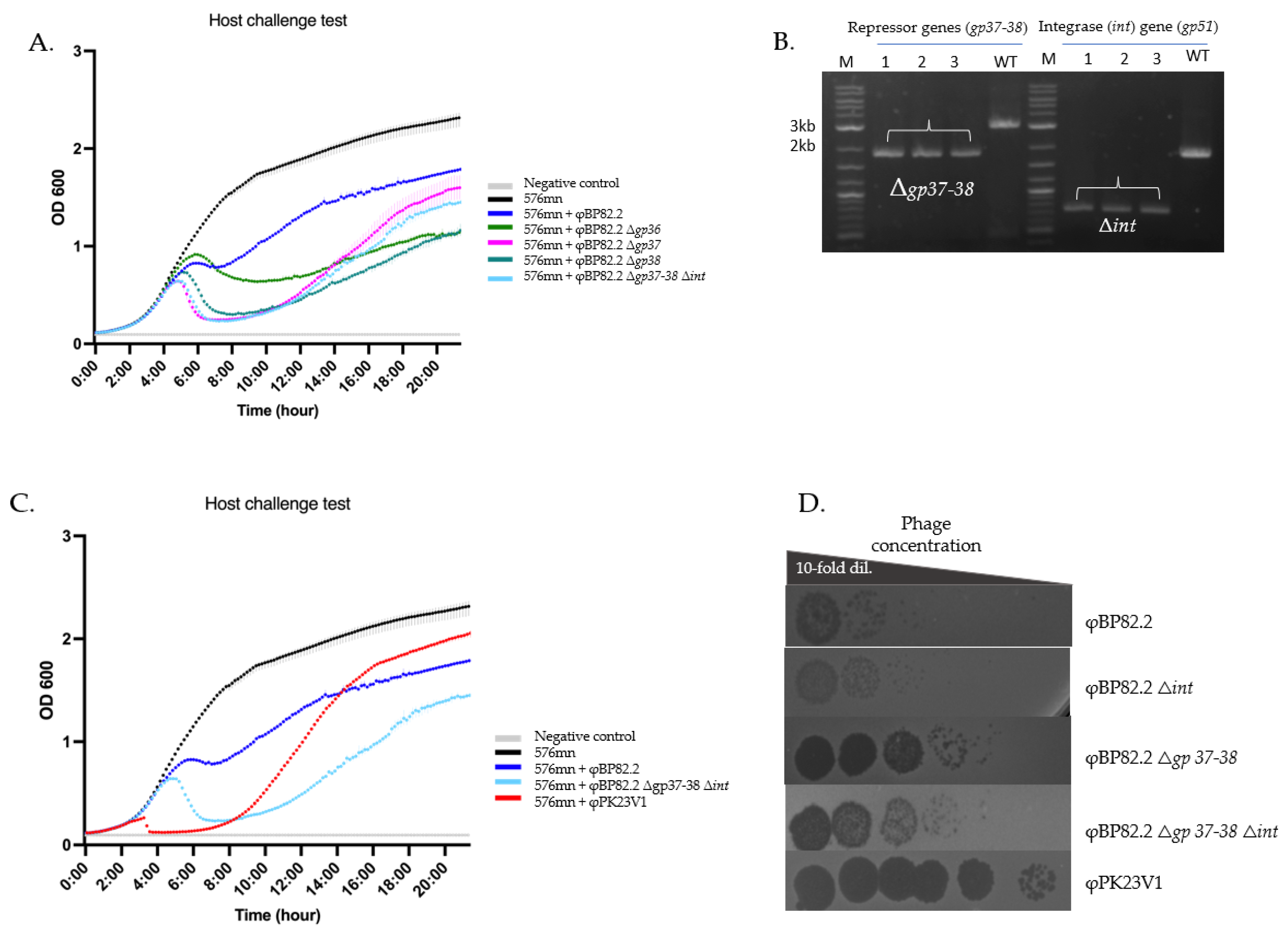
2.5. Repressor Gene Involvement in the Lysogenic-Lytic Switch
3. Discussion
3.1. Temperate Phages are Common in B. pseudomallei and May Contribute to Host Survival
3.2. The Integrase Gene is Required for Phage Excision but Does Not Enhance Lytic Activity
3.3. The Deletion of Repressor Genes Activates the Lysogenic - Lytic Switch
4. Materials and Methods
4.1. Bacterial Culture Condition
4.2. Prophage Genome Analysis
4.3. Phage Induction and Lysogen Isolation
4.4. RNA Sequencing
4.5. Phage Integrase Gene Mutagenesis
4.6. Construction of Replicative Plasmid for Integrase Supplementation
4.7. Phage Repressor Gene Mutagenesis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- White, N.J., Melioidosis. Lancet, 2003. 361(9370): p. 1715-22.
- Wiersinga, W.J., et al., Melioidosis: insights into the pathogenicity of Burkholderia pseudomallei. Nat Rev Microbiol, 2006. 4(4): p. 272-82. [CrossRef]
- Pongmala, K., et al., Distribution of Burkholderia pseudomallei within a 300-cm deep soil profile: implications for environmental sampling. Sci Rep, 2022. 12(1): p. 8674. [CrossRef]
- Limmathurotsakul, D., et al., Predicted global distribution of Burkholderia pseudomallei and burden of melioidosis. Nat Microbiol, 2016. 1(1). [CrossRef]
- Elschner, M.C., et al., Isolation of the highly pathogenic and zoonotic agent Burkholderia pseudomallei from a pet green Iguana in Prague, Czech Republic. BMC Vet Res, 2014. 10: p. 283. [CrossRef]
- Limmathurotsakul, D., et al., Melioidosis in animals, Thailand, 2006-2010. Emerg Infect Dis, 2012. 18(2): p. 325-7. [CrossRef]
- Wiersinga, W.J., et al., Melioidosis. Nat Rev Dis Primers, 2018. 4: p. 17107.
- Schweizer, H.P., Mechanisms of antibiotic resistance in Burkholderia pseudomallei: implications for treatment of melioidosis. Future Microbiol, 2012. 7(12): p. 1389-99. [CrossRef]
- Crowe, A., et al., Current antimicrobial susceptibility of first-episode melioidosis Burkholderia pseudomallei isolates from the Northern Territory, Australia. Int J Antimicrob Agents, 2014. 44(2): p. 160-2. [CrossRef]
- Dance, D., Treatment and prophylaxis of melioidosis. Int J Antimicrob Agents, 2014. 43(4): p. 310-8.
- Dance, D.A., et al., Trimethoprim/sulfamethoxazole resistance in Burkholderia pseudomallei. Int J Antimicrob Agents, 2014. 44(4): p. 368-9. [CrossRef]
- Sullivan, R.P., et al., 2020 Review and revision of the 2015 Darwin melioidosis treatment guideline; paradigm drift not shift. PLoS Negl Trop Dis, 2020. 14(9): p. e0008659. [CrossRef]
- Bugrysheva, J.V., et al., Antibiotic Resistance Markers in Burkholderia pseudomallei Strain Bp1651 Identified by Genome Sequence Analysis. Antimicrob Agents Chemother, 2017. 61(6). [CrossRef]
- Sarovich, D.S., et al., Development of ceftazidime resistance in an acute Burkholderia pseudomallei infection. Infect Drug Resist, 2012. 5: p. 129-32. [CrossRef]
- Sommanustweechai, A., et al., Environmental management procedures following fatal melioidosis in a captive chimpanzee (Pan troglodytes). J Zoo Wildl Med, 2013. 44(2): p. 475-9. [CrossRef]
- Lim, Y.M., et al., Effective Therapeutic Options for Melioidosis: Antibiotics versus Phage Therapy. Pathogens, 2022. 12(1). [CrossRef]
- Wang, Y., et al., A novel lytic phage potentially effective for phage therapy against Burkholderia pseudomallei in the tropics. Infect Dis Poverty, 2022. 11(1): p. 87. [CrossRef]
- Gatedee, J., et al., Isolation and characterization of a novel podovirus which infects burkholderia pseudomallei. Virol J, 2011. 8: p. 366. [CrossRef]
- Yordpratum, U., et al., Novel lytic bacteriopphages from soil that lyse Burkholderia pseudomallei. FEMS Microbiol Lett, 2011. 314(1): p. 81-88. [CrossRef]
- Guang-Han, O., et al., Experimental Phage Therapy for Burkholderia pseudomallei Infection. PLoS One, 2016. 11(7): p. e0158213. [CrossRef]
- Leclerc, H. and P. Sureau, [Research on bacteriophages of the Whitmore bacillus in stagnant waters of Hanoi]. Bull Soc Pathol Exot Filiales, 1956. 49(5): p. 874-82.
- Khrongsee, P., et al., A comprehensive study of prophage islands in Burkholderia pseudomallei complex. Frontiers in Bacteriology, 2024. 3: p. 1339809. [CrossRef]
- DeShazer, D., Genomic diversity of Burkholderia pseudomallei clinical isolates: subtractive hybridization reveals a Burkholderia mallei-specific prophage in B. pseudomallei 1026b. J Bacteriol, 2004. 186(12): p. 3938-50. [CrossRef]
- Woods, D.E., et al., Burkholderia thailandensis E125 harbors a temperate bacteriophage specific for Burkholderia mallei. J Bacteriol, 2002. 184(14): p. 4003-17. [CrossRef]
- Tuanyok, A., et al., Genomic islands from five strains of Burkholderia pseudomallei. BMC Genomics, 2008. 9: p. 566. [CrossRef]
- Khrongsee P, K.J., Alami-Rose M, Subramaniam K, Waltzek TB, Schweizer HP, Tuanyok A, Exploring Burkholderia pseudomallei-specific bacteriophages: overcoming O-antigen specificity and adaptive mutation in phage tail fiber. Front. Bacteriol, 2024. 3:1433593 .
- Lopez, C.M., et al., Versatile dual-technology system for markerless allele replacement in Burkholderia pseudomallei. Appl. Environ. Microbiol., 2009. 75: p. 6496-6503. [CrossRef]
- Dedrick, R.M., et al., Mycobacteriophage ZoeJ: A broad host-range close relative of mycobacteriophage TM4. Tuberculosis (Edinb), 2019. 115: p. 14-23. [CrossRef]
- Dedrick, R.M., et al., Engineered bacteriophages for treatment of a patient with a disseminated drug-resistant Mycobacterium abscessus. Nat Med, 2019. 25(5): p. 730-733. [CrossRef]
- Broussard, G.W., et al., Integration-dependent bacteriophage immunity provides insights into the evolution of genetic switches. Mol Cell, 2013. 49(2): p. 237-48. [CrossRef]
- Shitrit, D., et al., Genetic engineering of marine cyanophages reveals integration but not lysogeny in T7-like cyanophages. Isme j, 2022. 16(2): p. 488-499. [CrossRef]
- Christie, G.E. and R. Calendar, Bacteriophage P2. Bacteriophage, 2016. 6(1): p. e1145782.
- Guarente, L., et al., Mutant lambda phage repressor with a specific defect in its positive control function. Proc Natl Acad Sci U S A, 1982. 79(7): p. 2236-9. [CrossRef]
- Yao, G., et al., Phage Milagro: a platform for engineering a broad host range virulent phage for Burkholderia. J Virol, 2023. 97(11): p. e0085023. [CrossRef]
- Propst, K.L., et al., A Burkholderia pseudomallei delta-purM mutant is a avirulent in immune competent and immune deficient animals: candidate strain for exclusion from select agent lists. Infection and Immunity, 2010. 78: p. 3136-3143.
- Norris, M.H., et al., An avirulent Burkholderia pseudomallei ∆purM strain with atypical type B LPS: expansion of the toolkit for biosafe studies of melioidosis. BMC Microbiol, 2017. 17(1): p. 132. [CrossRef]
- Camacho, C., et al., BLAST+: architecture and applications. BMC Bioinformatics, 2009. 10(1): p. 421.
- Quinlan, A.R. and I.M. Hall, BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics, 2010. 26(6): p. 841-2. [CrossRef]
- Carver, T.J., et al., ACT: the Artemis Comparison Tool. Bioinformatics, 2005. 21(16): p. 3422-3. [CrossRef]
- Sullivan, M.J., N.K. Petty, and S.A. Beatson, Easyfig: a genome comparison visualizer. Bioinformatics, 2011. 27(7): p. 1009-10. [CrossRef]
- Bolger, A.M., M. Lohse, and B. Usadel, Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics, 2014. 30(15): p. 2114-20. [CrossRef]
- Li, H., Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv, 2013(preprint arXiv:1303.3997).
- Carver, T., et al., Artemis: an integrated platform for visualization and analysis of high-throughput sequence-based experimental data. Bioinformatics, 2012. 28(4): p. 464-9. [CrossRef]
- Choi, K.H., A. Kumar, and H.P. Schweizer, A 10-min method for preparation of highly electrocompetent Pseudomonas aeruginosa cells: application for DNA fragment transfer between chromosomes and plasmid transformation. J Microbiol Methods, 2006. 64(3): p. 391-7. [CrossRef]
- Lee, T.S., et al., BglBrick vectors and datasheets: A synthetic biology platform for gene expression. J Biol Eng, 2011. 5: p. 12. [CrossRef]
- Liljeruhm, J., et al., Engineering a palette of eukaryotic chromoproteins for bacterial synthetic biology. J Biol Eng, 2018. 12: p. 8. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



