Submitted:
23 February 2026
Posted:
25 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Evolutionary Triggers Behind Seed (Plant) Development
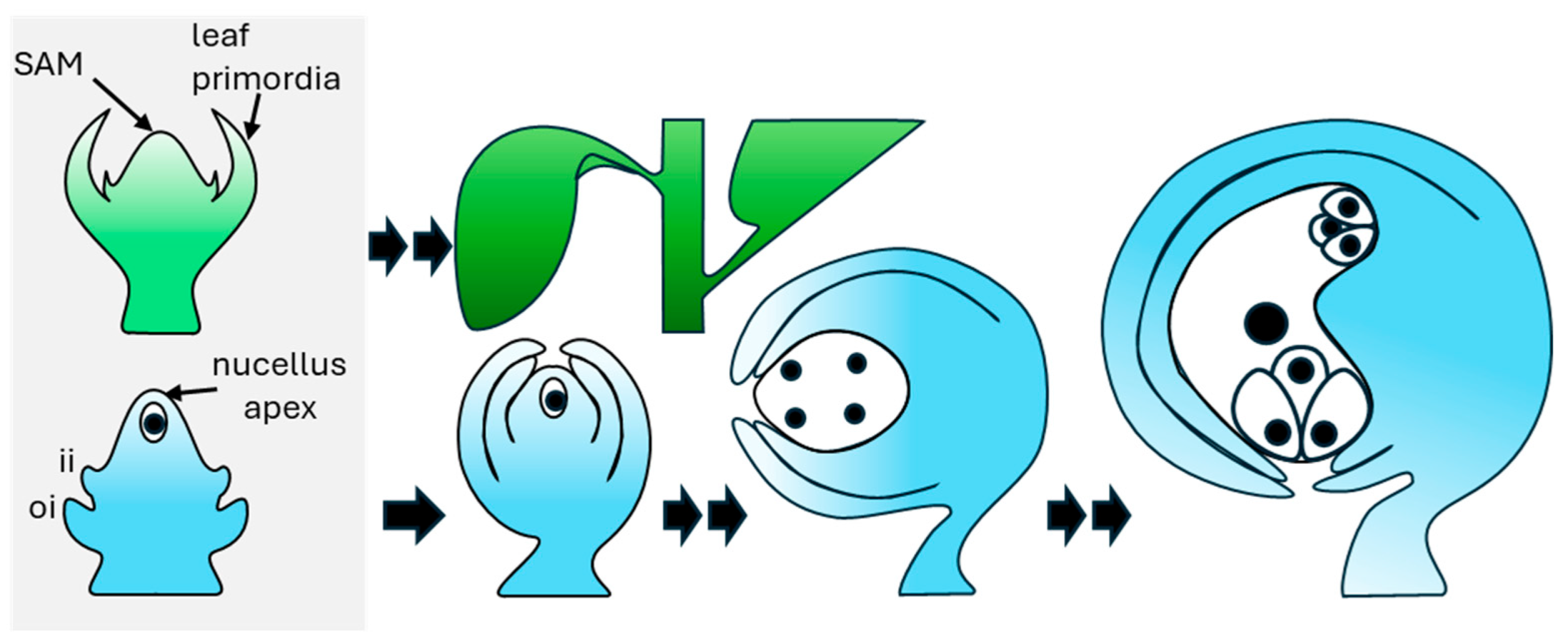
3. Ovule Development
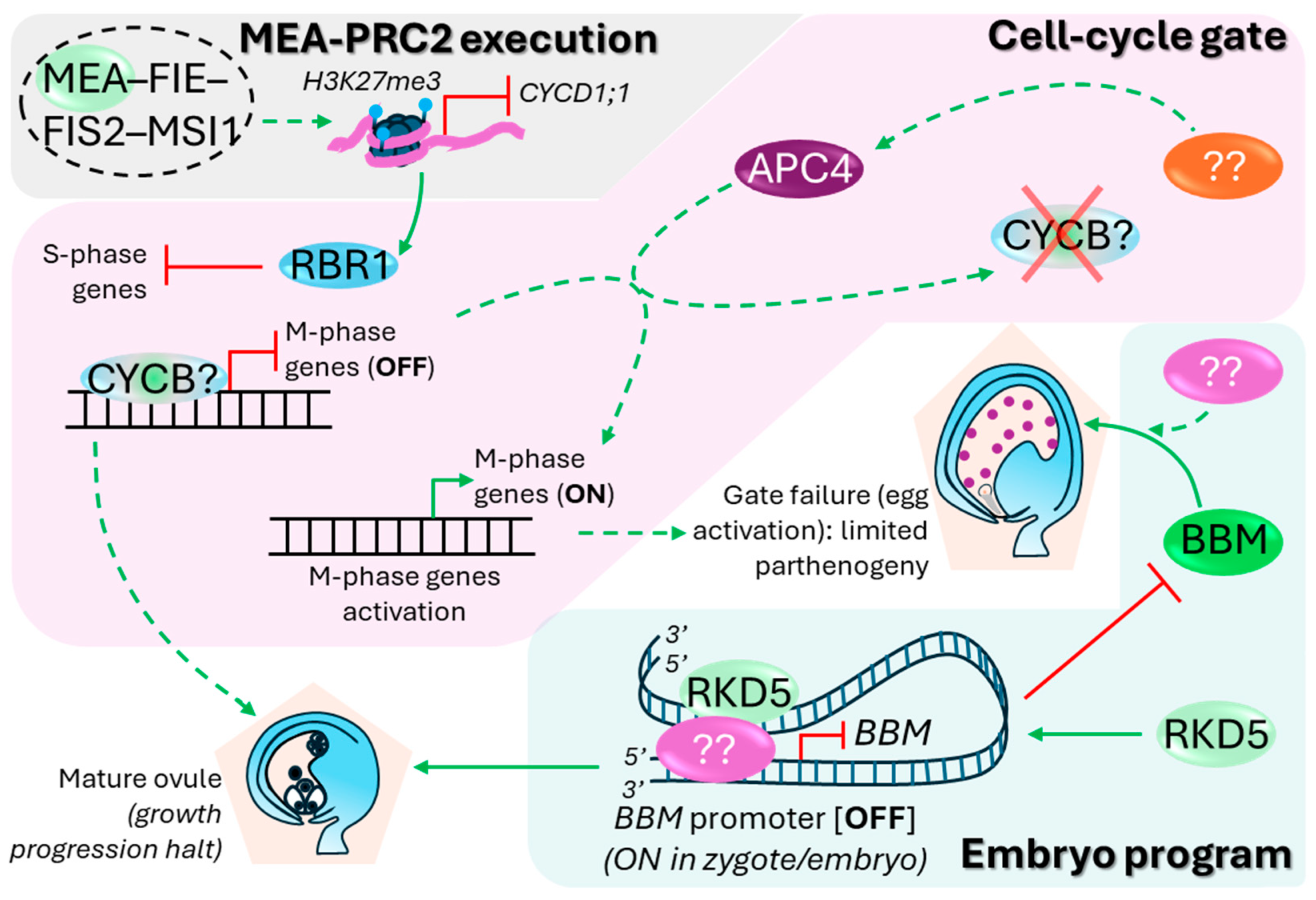
4. Ovule Maturity and Growth Progression Halt
5. Double Fertilization and Alleviation of the Molecular Block Against Seed Development
| Stage / window | Endosperm | Embryo | Maternal integument / testa |
|---|---|---|---|
| Pollen tube reception / pre-syngamy (POEM) | Central cell/endosperm-like responses can be triggered by pollen tube content release without gamete fusion (Kasahara et al., 2016, Honma et al., 2020). | No zygote formed. | PTC can initiate chalazal seed-coat wall formation; auxin can phenocopy aspects of initiation (Figueiredo et al., 2016, Liu et al., 2019). |
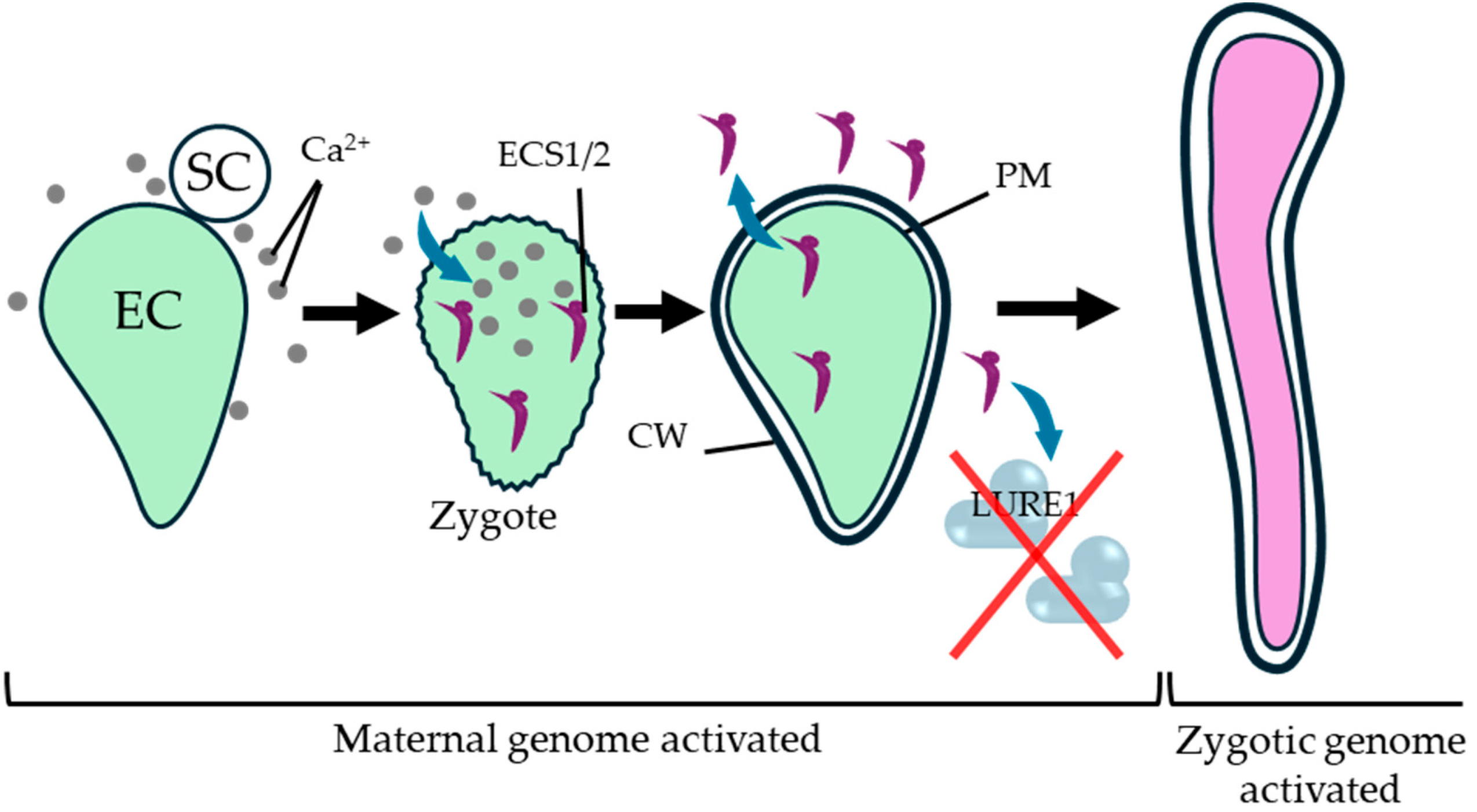
| Double fertilization | Paternal miR159 helps relieve maternal barriers by targeting GAMYB members (MYB33/MYB65) (Zhao et al., 2018b). | Fertilization initiates embryogenesis and relieves pre-fertilization arrest programs (Adhikari et al., 2020b). | Maternal tissues become responsive to fertilization-derived signals that promote seed coat initiation (Figueiredo et al., 2016). |
| Early coenocytic endosperm (pre-globular) | AGL62 supports/maintains the coenocytic program and prevents premature cellularization; an imprinted PHE1 network supports proliferation and delays cellularization (Hehenberger et al., 2012, Batista et al., 2019). | Early patterning is established as zygotic programs become active (Nodine and Bartel, 2012). | Endosperm expansion generates mechanical load on integuments (turgor) (Beauzamy et al., 2016). |
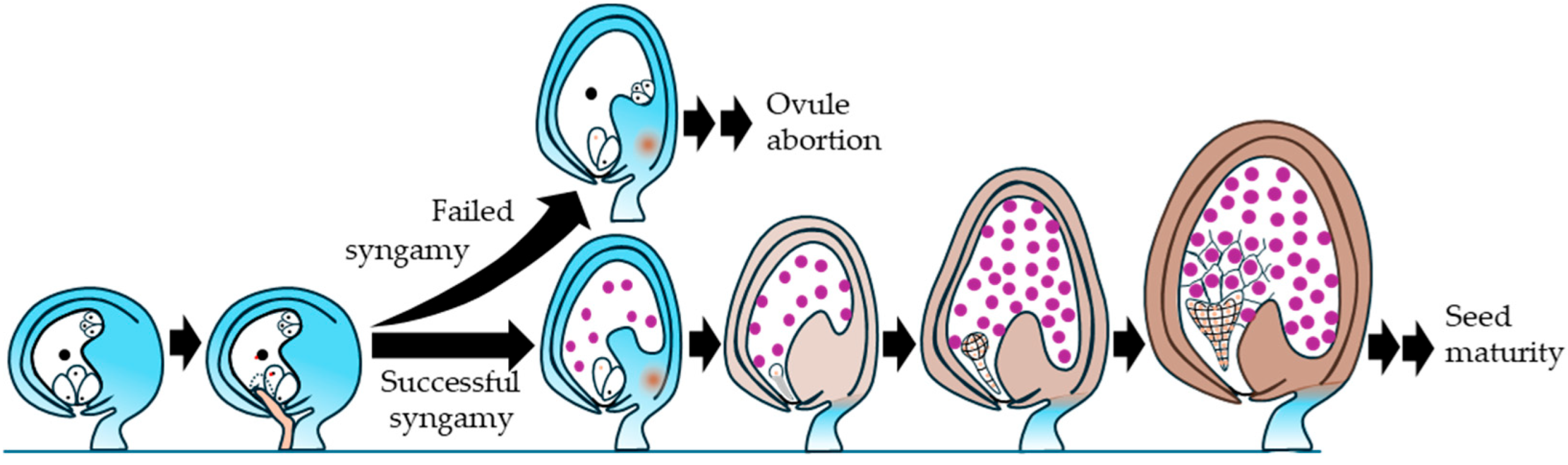
| Late coenocytic endosperm / late globular | Endosperm turgor approaches a maximum near the end of the coenocytic stage and then declines before cellularization (Beauzamy et al., 2016). | Embryo reaches globular stage prior to heart-stage organogenesis (Figure 10). | Chalazal seed coat differentiation includes activation of BAN and PA deposition (Debeaujon et al., 2003). |
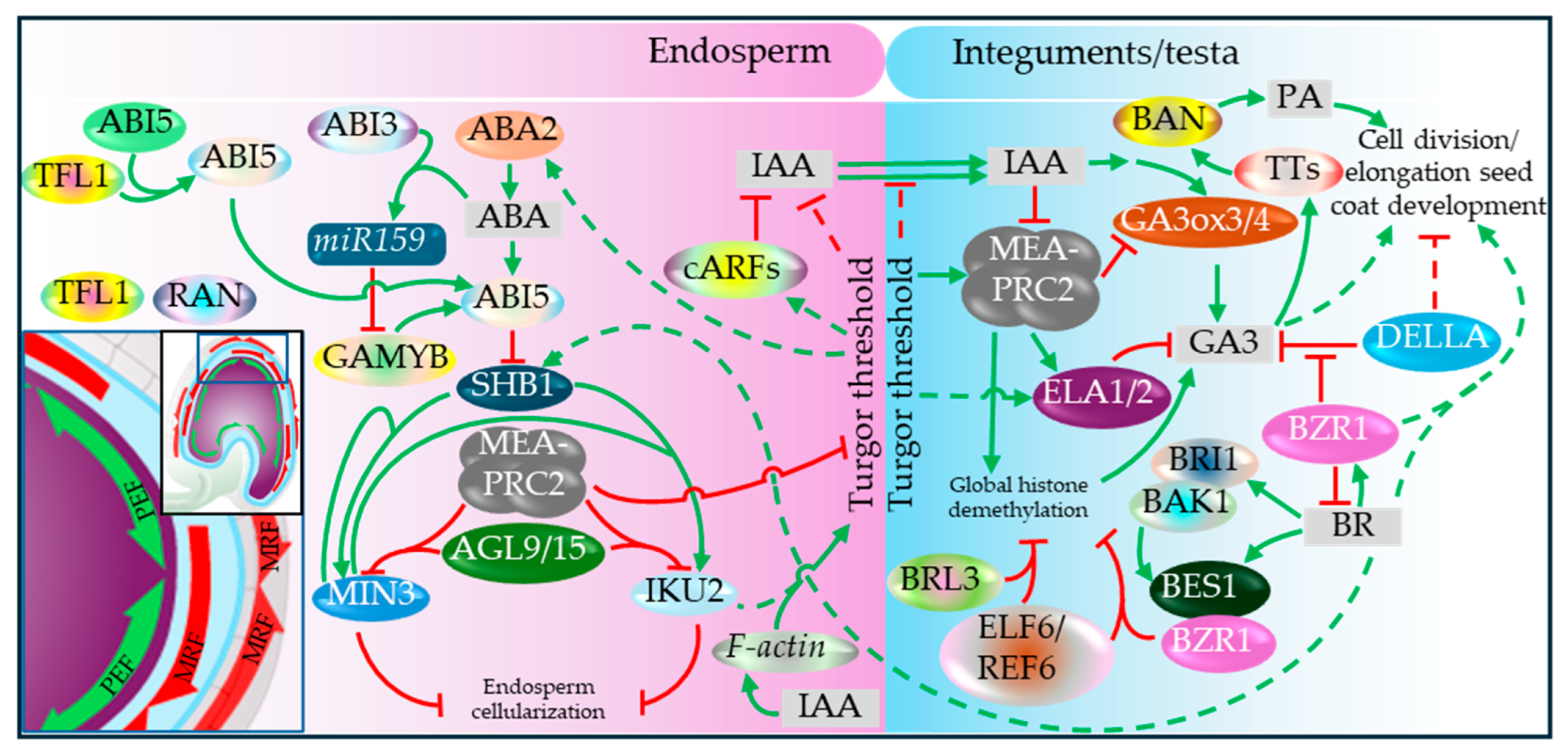
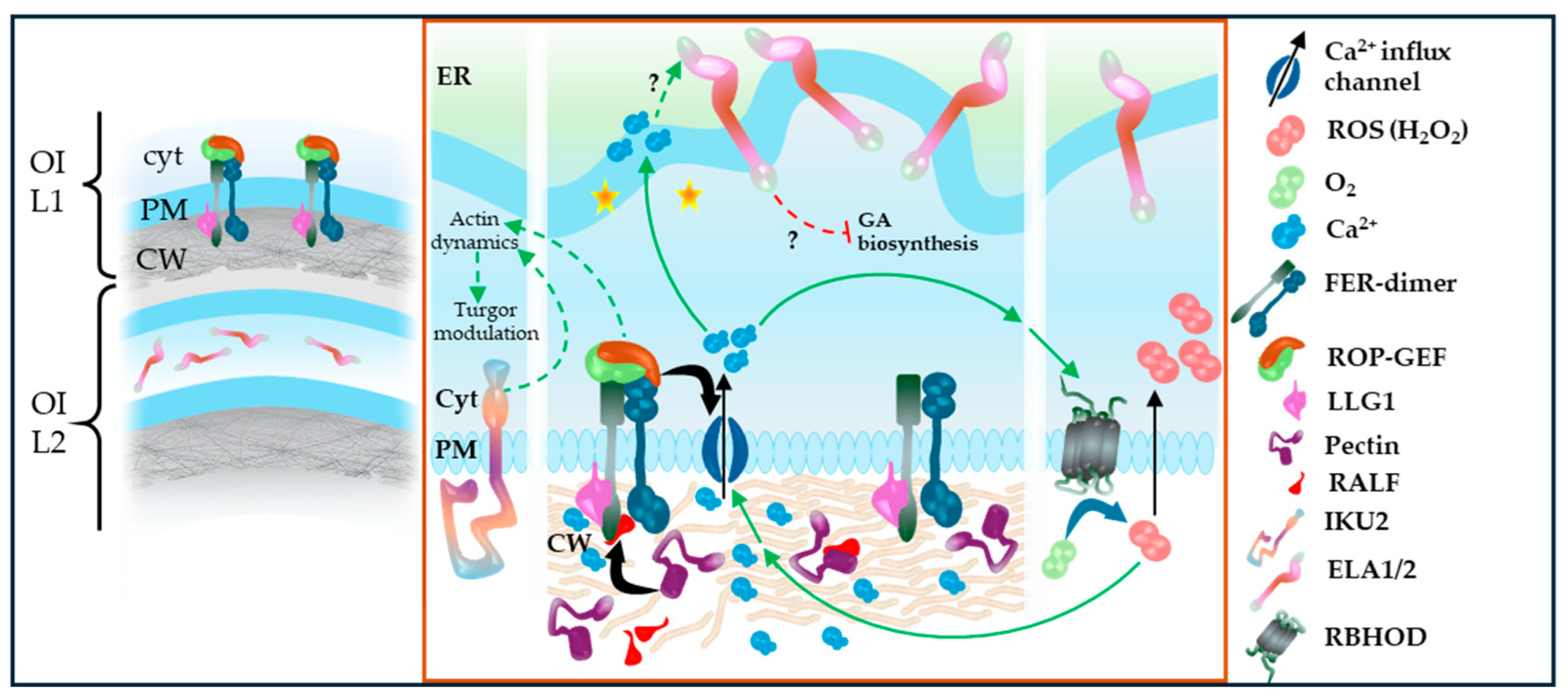
| Early heart stage (cellularization onset) | Cellularization initiates micropylar-to-chalazal; ABA/ABI5 represses the SHB1–MIN3–IKU2 proliferation module (Hehenberger et al., 2012, Cheng et al., 2014, Li et al., 2022). | Heart-stage morphogenesis proceeds alongside increasing zygotic genome activity (Zhao et al., 2019). | As cellularization proceeds, mechanosensitive testa responses are proposed to constrain growth via GA deactivation (ELA1/2) and wall stiffening (Creff et al., 2015, Creff et al., 2023). |
5.1. Pre-Syngamy Changes
5.2. The Coenocyte
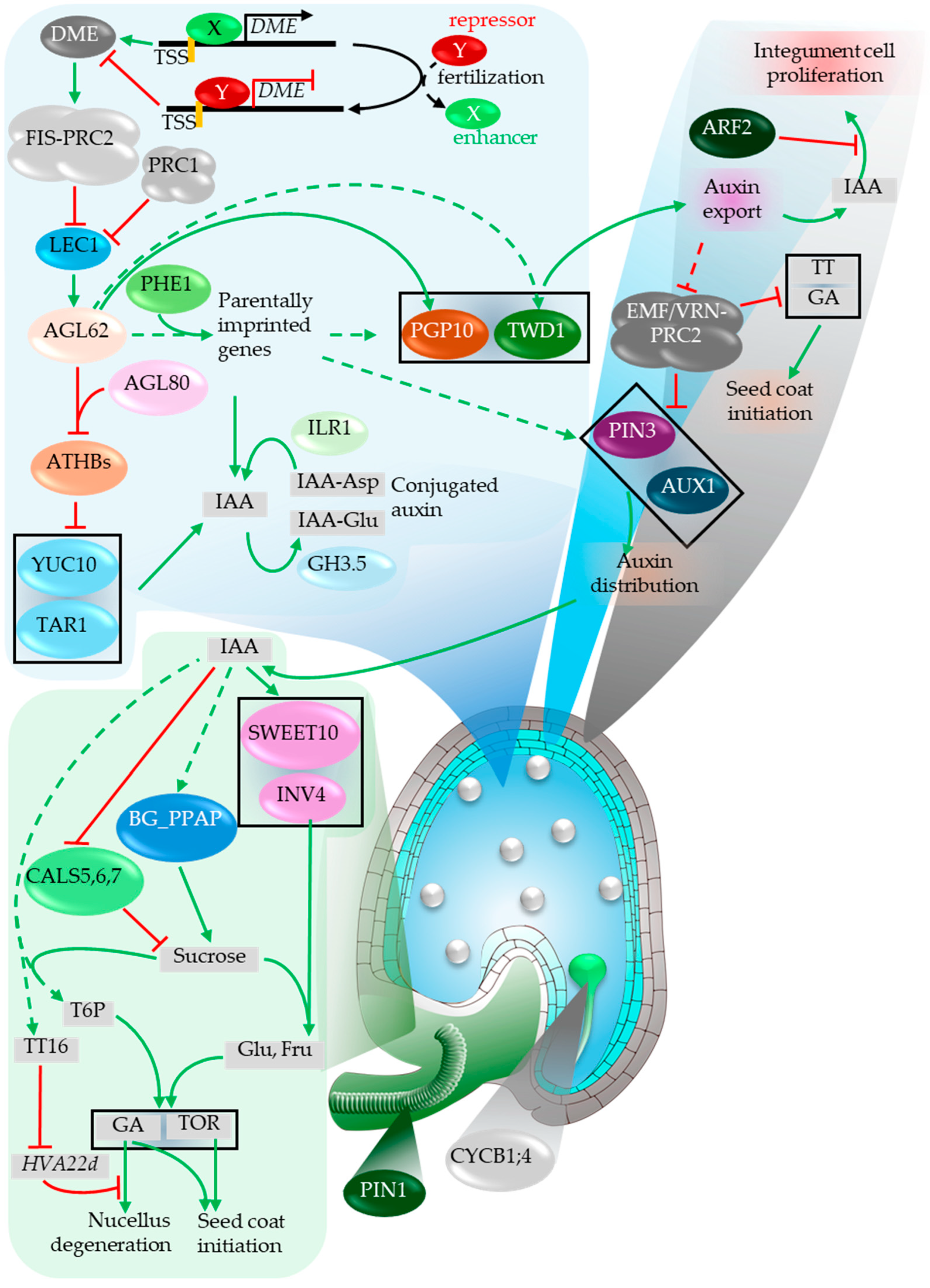
5.3. Seed Coat Development
5.4. Seed Size Regulation (Endosperm-Dependent)
5.5. Endosperm Cellularization
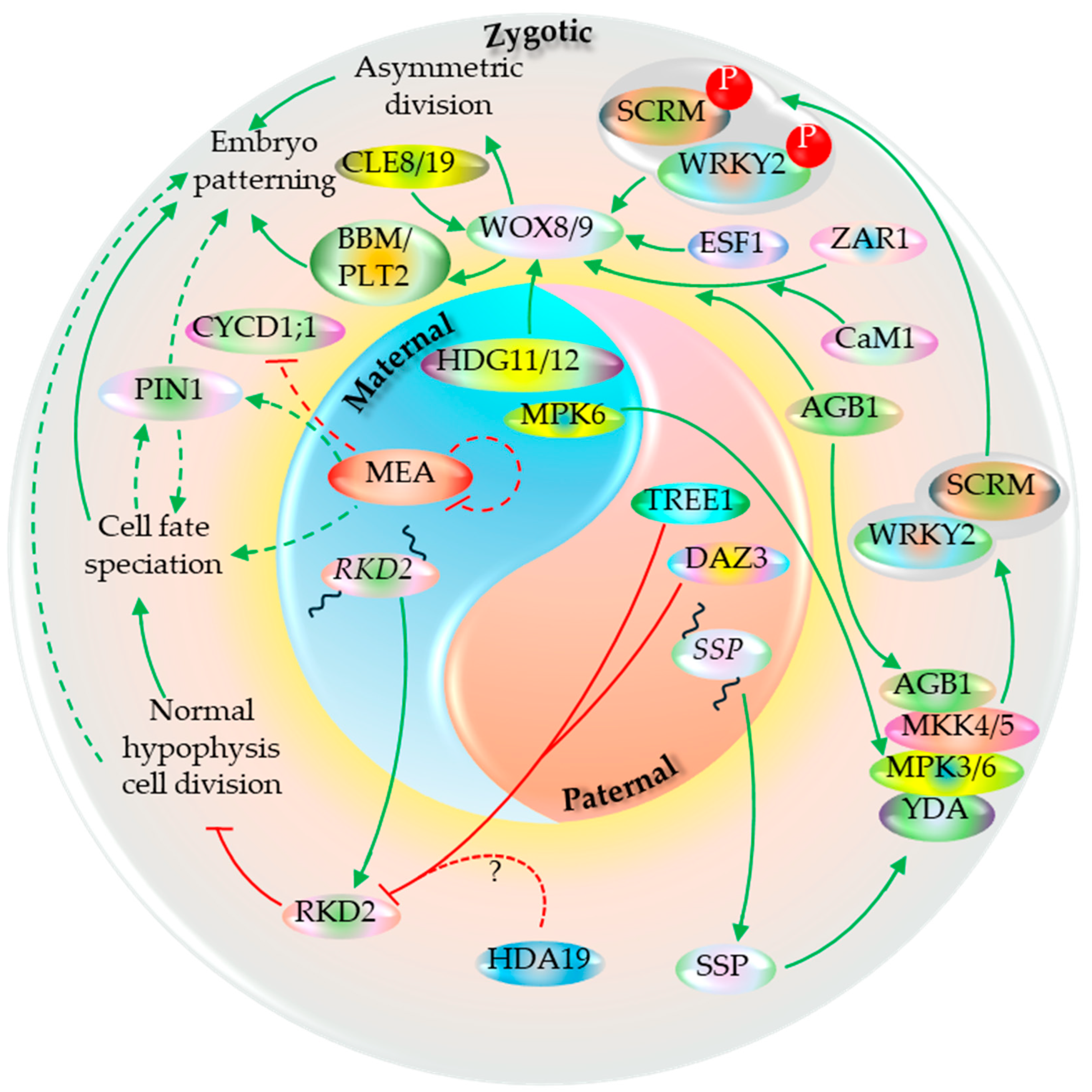
5.6. Embryo Development
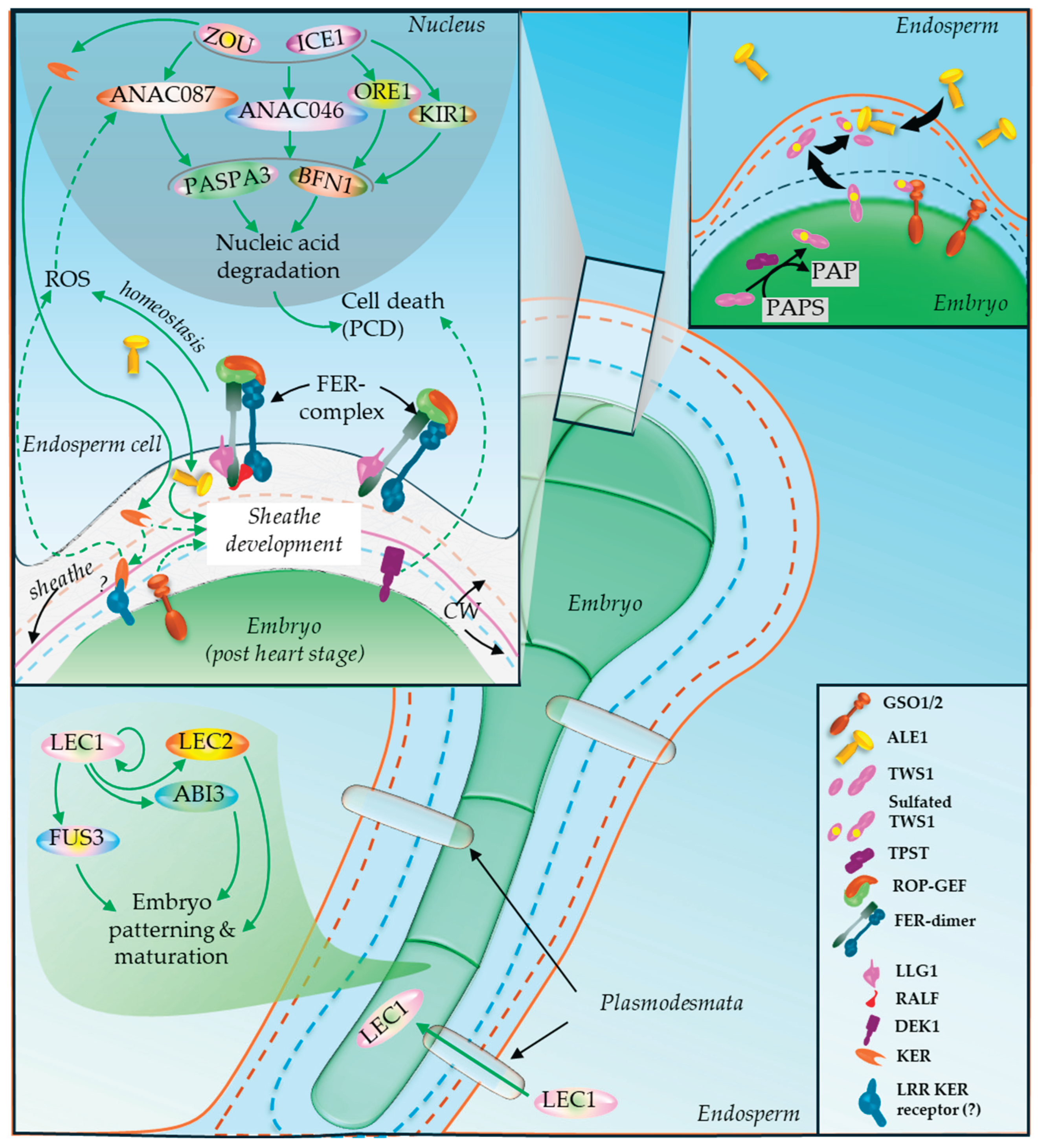
5.7. Embryo-Endosperm Growth Coordination
6. Bottlenecks and Future Perspectives
6.1. Correlative vs Causative Evidence Behind MMC Speciation
6.2. Sporophyte-Gametophyte Cross-Talk Conduits
6.3. Translational Perspective
7. Conclusion
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Abe, Y.; Mieda, K.; Ando, T.; Kono, I.; Yano, M.; Kitano, H.; Iwasaki, Y. The SMALL AND ROUND SEED1 (SRS1/DEP2) gene is involved in the regulation of seed size in rice. Genes & genetic systems 2010, 85, 327–339. [Google Scholar]
- Adamski, N.M.; Anastasiou, E.; Eriksson, S.; O'Neill, C.M.; Lenhard, M. Local maternal control of seed size by KLUH/ CYP78A5-dependent growth signaling. Proceedings of the National Academy of Sciences 2009, 106, 20115–20120. [Google Scholar] [CrossRef]
- Adhikari, P.B.; Liu, X.; Kasahara, R.D. Mechanics of pollen tube elongation: A perspective. Frontiers in Plant Science 2020a, 11, 589712. [Google Scholar] [CrossRef]
- Adhikari, P.B.; Liu, X.; Wu, X.; Zhu, S.; Kasahara, R.D. Fertilization in flowering plants: an odyssey of sperm cell delivery. Plant Molecular Biology 2020b, 103, 9–32. [Google Scholar] [CrossRef]
- Alexiadis, V.; Waldmann, T.; Andersen, J.; Mann, M.; Knippers, R.; Gruss, C. The protein encoded by the proto-oncogene DEK changes the topology of chromatin and reduces the efficiency of DNA replication in a chromatin-specific manner. Genes Dev 2000, 14, 1308–1312. [Google Scholar] [CrossRef]
- Ali, M.F.; Shin, J.M.; Fatema, U.; Kurihara, D.; Berger, F.; Yuan, L.; Kawashima, T. Cellular dynamics of coenocytic endosperm development in Arabidopsis thaliana. Nature Plants 2023, 9, 330–342. [Google Scholar] [CrossRef]
- Aloni, R.; Aloni, E.; Langhans, M.; Ullrich, C.I. Role of auxin in regulating Arabidopsis flower development. Planta 2006, 223, 315–328. [Google Scholar] [CrossRef]
- Antoine, A.F.; Faure, J.E.; Cordeiro, S.; Dumas, C.; Rougier, M.; Feijó, J.A. A calcium influx is triggered and propagates in the zygote as a wavefront during in vitro fertilization of flowering plants. Proceedings of the National Academy of Sciences 2000, 97, 10643–10648. [Google Scholar] [CrossRef]
- Arnault, G.; Vialette, A.C.M.; Andres-Robin, A.; Fogliani, B.; Gâteblé, G.; Scutt, C.P. Evidence for the extensive conservation of mechanisms of ovule integument development since the most recent common ancestor of living angiosperms. Frontiers in Plant Science 2018, 9–2018. [Google Scholar] [CrossRef]
- Aslam, M.; Fakher, B.; Jakada, B.H.; Cao, S.; Qin, Y. SWR1 chromatin remodeling complex: A key transcriptional regulator in plants. Cells 2019, 8. [Google Scholar] [CrossRef]
- Baroux, C.; Gagliardini, V.; Page, D.R.; Grossniklaus, U. Dynamic regulatory interactions of Polycomb group genes: MEDEA autoregulation is required for imprinted gene expression in Arabidopsis. Genes & Development 2006, 20, 1081–1086. [Google Scholar] [CrossRef]
- Baroux, C.; Grossniklaus, U. Seeds—An evolutionary innovation underlying reproductive success in flowering plants. In Current Topics in Developmental Biology; Grossniklaus, U., Ed.; Academic Press, 2019; pp. 605–642. [Google Scholar]
- Baroux, C.; Spillane, C.; Grossniklaus, U. Evolutionary origins of the endosperm in flowering plants. Genome Biology 2002, 3, reviews1026.1021. [Google Scholar] [CrossRef]
- Batista, R.A.; Moreno-Romero, J.; Qiu, Y.; van Boven, J.; Santos-González, J.; Figueiredo, D.D.; Köhler, C. The MADS-box transcription factor PHERES1 controls imprinting in the endosperm by binding to domesticated transposons. eLife 2019, 8, e50541. [Google Scholar] [CrossRef]
- Bauer, M.J.; Fischer, R.L. Genome demethylation and imprinting in the endosperm. Current Opinion in Plant Biology 2011, 14, 162–167. [Google Scholar] [CrossRef]
- Bayer, M.; Nawy, T.; Giglione, C.; Galli, M.; Meinnel, T.; Lukowitz, W. Paternal control of embryonic patterning in Arabidopsis thaliana. Science 2009, 323, 1485–1488. [Google Scholar] [CrossRef]
- Beauzamy, L.; Fourquin, C.; Dubrulle, N.; Boursiac, Y.; Boudaoud, A.; Ingram, G.C. Endosperm turgor pressure decreases during early Arabidopsis seed development. Development 2016, 143, 3295–3299. [Google Scholar] [CrossRef]
- Becker, A.; Chen, X.; Dresselhaus, T.; Gutsche, N.; Müller-Schüssele, S.J.; Sprunck, S.; Theißen, G.; de Vries, S.; Zachgo, S. Sexual reproduction in land plants: an evolutionary perspective. Plant Reproduction 2025, 38, 12. [Google Scholar] [CrossRef]
- Belmonte, M.F.; Kirkbride, R.C.; Stone, S.L.; Pelletier, J.M.; Bui, A.Q.; Yeung, E.C.; Hashimoto, M.; Fei, J.; Harada, C.M.; Munoz, M.D.; Le, B.H.; Drews, G.N.; Brady, S.M.; Goldberg, R.B.; Harada, J.J. Comprehensive developmental profiles of gene activity in regions and subregions of the Arabidopsis seed. Proceedings of the National Academy of Sciences 2013, 110, E435–E444. [Google Scholar] [CrossRef]
- Bencivenga, S.; Simonini, S.; Benková, E.; Colombo, L. The transcription factors BEL1 and SPL are required for cytokinin and auxin signaling during ovule development in Arabidopsis. The Plant Cell 2012, 24, 2886–2897. [Google Scholar] [CrossRef]
- Benton, M.J.; Wilf, P.; Sauquet, H. The angiosperm terrestrial revolution and the origins of modern biodiversity. New Phytologist 2022, 233, 2017–2035. [Google Scholar] [CrossRef]
- Berger, F. Meiosis as a mechanism for epigenetic reprogramming and cellular rejuvenation. Development 2024, 151. [Google Scholar] [CrossRef]
- Boyko, J.D.; Vasconcelos, T. Trait lability as a predictor of diversification dynamics in flowering plants. bioRxiv. 2025. 2024.2006.2003.597046.
- Brand, A.; Quimbaya, M.; Tohme, J.; Chavarriaga-Aguirre, P. Arabidopsis LEC1 and LEC2 orthologous genes are key regulators of somatic embryogenesis in cassava. Frontiers in Plant Science 2019, 10–2019. [Google Scholar] [CrossRef]
- Brantley, S.; Di Talia, S. The maternal-to-zygotic transition. Current Biology 2024, 34, R519–R523. [Google Scholar] [CrossRef]
- Butel, N.; Qiu, Y.; Xu, W.; Santos-González, J.; Köhler, C. Parental conflict driven regulation of endosperm cellularization by a family of Auxin Response Factors. Nature Plants 2024, 10, 1018–1026. [Google Scholar] [CrossRef]
- Cavalleri, A.; Astori, C.; Manrique, S.; Bruzzaniti, G.; Smaczniak, C.; Mizzotti, C.; Ruiu, A.; Spano, M.; Movilli, A.; Gregis, V.; Xu, X.; Kaufmann, K.; Colombo, L. (2025) SPOROCYTELESS/NOZZLE acts together with MADS-domain transcription factors to regulate an auxin-dependent network controlling the Megaspore Mother Cell development. bioRxiv 2025.2003, 2011.641985. [Google Scholar]
- Chen, B.; Maas, L.; Figueiredo, D.; Zhong, Y.; Reis, R.; Li, M.; Horstman, A.; Riksen, T.; Weemen, M.; Liu, H.; Siemons, C.; Chen, S.; Angenent, G.C.; Boutilier, K. BABY BOOM regulates early embryo and endosperm development. Proceedings of the National Academy of Sciences 2022, 119, e2201761119. [Google Scholar] [CrossRef]
- Chen, D.; Molitor, A.M.; Xu, L.; Shen, W.-H. Arabidopsis PRC1 core component AtRING1 regulates stem cell-determining carpel development mainly through repression of class I KNOX genes. BMC Biology 2016, 14, 112. [Google Scholar] [CrossRef]
- Cheng, T.; Liu, Z.; Li, H.; Huang, X.; Wang, W.; Shi, C.; Zhang, X.; Chen, H.; Yao, Z.; Zhao, P.; Peng, X.; Sun, M.-X. Sperm-origin paternal effects on root stem cell niche differentiation. Nature 2024, 634, 220–227. [Google Scholar] [CrossRef]
- Cheng, Z.J.; Zhao, X.Y.; Shao, X.X.; Wang, F.; Zhou, C.; Liu, Y.G.; Zhang, Y.; Zhang, X.S. Abscisic acid regulates early seed development in Arabidopsis by ABI5-mediated transcription of SHORT HYPOCOTYL UNDER BLUE1. The Plant Cell 2014, 26, 1053–1068. [Google Scholar] [CrossRef]
- Choi, Y.; Gehring, M.; Johnson, L.; Hannon, M.; Harada, J.J.; Goldberg, R.B.; Jacobsen, S.E.; Fischer, R.L. DEMETER, a DNA glycosylase domain protein, is required for endosperm gene imprinting and seed viability in Arabidopsis. Cell 2002, 110, 33–42. [Google Scholar] [CrossRef]
- Conner, J.A.; Mookkan, M.; Huo, H.; Chae, K.; Ozias-Akins, P. A parthenogenesis gene of apomict origin elicits embryo formation from unfertilized eggs in a sexual plant. Proceedings of the National Academy of Sciences 2015, 112, 11205–11210. [Google Scholar] [CrossRef]
- Costa, L.M.; Marshall, E.; Tesfaye, M.; Silverstein, K.A.T.; Mori, M.; Umetsu, Y.; Otterbach, S.L.; Papareddy, R.; Dickinson, H.G.; Boutiller, K.; VandenBosch, K.A.; Ohki, S.; Gutierrez-Marcos, J.F. Central cell–derived peptides regulate early embryo patterning in flowering plants. Science 2014, 344, 168–172. [Google Scholar] [CrossRef] [PubMed]
- Creff, A.; Ali, O.; Bied, C.; Bayle, V.; Ingram, G.; Landrein, B. Evidence that endosperm turgor pressure both promotes and restricts seed growth and size. Nature Communications 2023, 14, 67. [Google Scholar] [CrossRef]
- Creff, A.; Brocard, L.; Ingram, G. A mechanically sensitive cell layer regulates the physical properties of the Arabidopsis seed coat. Nature Communications 2015, 6, 6382. [Google Scholar] [CrossRef]
- Dai, D.; Mudunkothge, J.S.; Galli, M.; Char, S.N.; Davenport, R.; Zhou, X.; Gustin, J.L.; Spielbauer, G.; Zhang, J.; Barbazuk, W.B.; Yang, B.; Gallavotti, A.; Settles, A.M. Paternal imprinting of dosage-effect defective1 contributes to seed weight xenia in maize. Nature Communications 2022, 13, 5366. [Google Scholar] [CrossRef]
- De Giorgi, J.; Fuchs, C.; Iwasaki, M.; Kim, W.; Piskurewicz, U.; Gully, K.; Utz-Pugin, A.; Mene-Saffrane, L.; Waridel, P.; Nawrath, C. The Arabidopsis mature endosperm promotes seedling cuticle formation via release of sulfated peptides. Developmental cell 2021, 56, 3066–3081. e3065. [Google Scholar] [CrossRef]
- Debeaujon, I.; Nesi, N.; Perez, P.; Devic, M.; Grandjean, O.; Caboche, M.; Lepiniec, L. Proanthocyanidin-accumulating cells in Arabidopsis testa: regulation of differentiation and role in seed development. Plant Cell 2003, 15, 2514–2531. [Google Scholar] [CrossRef]
- Delesalle, C.; Montiel-Jorda, A.; Aiba, R.; Spielmann, J.; Neveu, J.; Fujita, S.; Vert, G. Arabidopsis microtubule-BRI1-associated proteins negatively regulate hypocotyl elongation by controlling brassinosteroid-dependent cortical microtubule reorientation. Plant Communications 2025, 101637. [Google Scholar] [CrossRef]
- Denay, G.; Creff, A.; Moussu, S.; Wagnon, P.; Thévenin, J.; Gérentes, M.-F.; Chambrier, P.; Dubreucq, B.; Ingram, G. Endosperm breakdown in Arabidopsis requires heterodimers of the basic helix-loop-helix proteins ZHOUPI and INDUCER OF CBP EXPRESSION 1. Development 2014, 141, 1222–1227. [Google Scholar] [CrossRef]
- Denninger, P.; Bleckmann, A.; Lausser, A.; Vogler, F.; Ott, T.; Ehrhardt, D.W.; Frommer, W.B.; Sprunck, S.; Dresselhaus, T.; Grossmann, G. Male-female communication triggers calcium signatures during fertilization in Arabidopsis. Nat Commun 2014, 5, 4645. [Google Scholar] [CrossRef]
- Deslauriers, S.D.; Larsen, P.B. FERONIA is a key modulator of brassinosteroid and ethylene responsiveness in Arabidopsis hypocotyls. Molecular Plant 2010, 3, 626–640. [Google Scholar] [CrossRef] [PubMed]
- Doll, N.M.; Bovio, S.; Gaiti, A.; Marsollier, A.-C.; Chamot, S.; Moussu, S.; Widiez, T.; Ingram, G. The endosperm-derived embryo sheath is an anti-adhesive structure that facilitates cotyledon emergence during germination in Arabidopsis. Current Biology 2020a, 30, 909–915.e904. [Google Scholar] [CrossRef]
- Doll, N.M.; Fierlej, Y.; Eekhout, T.; Elias, L.; Bellot, C.; Sun, G.; Grones, C.; Aesaert, S.; Coussens, G.; De Rycke, R.; Šimášková, M.; Montes, E.; Plagnard, C.; Rogowsky, P.M.; Abebaw, Y.M.; Bendahmane, M.; De Rybel, B.; Pauwels, L.; Widiez, T.; Nowack, M.K. KIL transcription factors facilitate embryo growth in maize by promoting endosperm elimination via lytic cell death. The Plant Cell 2025, 37. [Google Scholar] [CrossRef]
- Doll, N.M.; Royek, S.; Fujita, S.; Okuda, S.; Chamot, S.; Stintzi, A.; Widiez, T.; Hothorn, M.; Schaller, A.; Geldner, N.; Ingram, G. A two-way molecular dialogue between embryo and endosperm is required for seed development. Science 2020b, 367, 431–435. [Google Scholar] [CrossRef]
- Doll, N.M.; Van Hautegem, T.; Schilling, N.; De Rycke, R.; De Winter, F.; Fendrych, M.; Nowack, M.K. Endosperm cell death promoted by NAC transcription factors facilitates embryo invasion in Arabidopsis. Current Biology 2023, 33, 3785–3795.e3786. [Google Scholar] [CrossRef] [PubMed]
- Domínguez, F.; Cejudo, F.J. Programmed cell death (PCD): an essential process of cereal seed development and germination. Frontiers in Plant Science 2014, 5–2014. [Google Scholar] [CrossRef] [PubMed]
- Du, G.; Zhao, Y.; Xiao, C.; Ren, D.; Ding, Y.; Xu, J.; Jin, H.; Jiao, H. Mechanism analysis of calcium nitrate application to induce gibberellin biosynthesis and signal transduction promoting stem elongation of Dendrobium officinale. Industrial Crops and Products 2023, 195, 116495. [Google Scholar] [CrossRef]
- Duan, Q.; Kita, D.; Li, C.; Cheung, A.Y.; Wu, H.-M. FERONIA receptor-like kinase regulates RHO GTPase signaling of root hair development. Proceedings of the National Academy of Sciences 2010, 107, 17821–17826. [Google Scholar] [CrossRef]
- Ebel, C.; Mariconti, L.; Gruissem, W. Plant retinoblastoma homologues control nuclear proliferation in the female gametophyte. Nature 2004, 429, 776–780. [Google Scholar] [CrossRef]
- Faure, J.E.; Digonnet, C.; Dumas, C. An in vitro system for adhesion and fusion of maize gametes. Science 1994, 263, 1598–1600. [Google Scholar] [CrossRef]
- Feng, W.; Kita, D.; Peaucelle, A.; Cartwright, H.N.; Doan, V.; Duan, Q.; Liu, M.-C.; Maman, J.; Steinhorst, L.; Schmitz-Thom, I.; Yvon, R.; Kudla, J.; Wu, H.-M.; Cheung, A.Y.; Dinneny, J.R. The FERONIA receptor kinase maintains cell-wall integrity during salt stress through Ca2+ signaling. Current Biology 2018, 28, 666–675.e665. [Google Scholar] [CrossRef] [PubMed]
- Figueiredo, D.D.; Batista, R.A.; Roszak, P.J.; Hennig, L.; Köhler, C. Auxin production in the endosperm drives seed coat development in Arabidopsis. eLife 2016, 5, e20542. [Google Scholar] [CrossRef]
- Figueiredo, D.D.; Batista, R.A.; Roszak, P.J.; Köhler, C. Auxin production couples endosperm development to fertilization. Nature Plants 2015, 1, 15184. [Google Scholar] [CrossRef]
- Figueiredo, D.D.; Köhler, C. Auxin: a molecular trigger of seed development. Genes & Development 2018, 32, 479–490. [Google Scholar] [CrossRef]
- Fiume, E.; Coen, O.; Xu, W.; Lepiniec, L.; Magnani, E. Developmental patterning of sub-epidermal cells in the outer integument of Arabidopsis seeds. PLOS ONE 2017, 12, e0188148. [Google Scholar] [CrossRef]
- Fiume, E.; Fletcher, J.C. Regulation of Arabidopsis embryo and endosperm development by the polypeptide signaling molecule CLE8. Plant Cell 2012, 24, 1000–1012. [Google Scholar] [CrossRef]
- Floyd, Sandra K.; Friedman, William E. Evolution of endosperm developmental patterns among basal flowering plants. International Journal of Plant Sciences 2000, 161, S57–S81. [Google Scholar] [CrossRef]
- Fourquin, C.; Beauzamy, L.; Chamot, S.; Creff, A.; Goodrich, J.; Boudaoud, A.; Ingram, G. Mechanical stress mediated by both endosperm softening and embryo growth underlies endosperm elimination in Arabidopsis seeds. Development 2016, 143, 3300–3305. [Google Scholar] [CrossRef]
- Gai, W.; Liu, C.; Yang, M.; Li, F.; Xin, H.; Gai, S. Calcium signaling facilitates chilling- and GA- induced dormancy release in tree peony. Frontiers in Plant Science 2024, 15–2024. [Google Scholar] [CrossRef]
- Gallego-Bartolomé, J.; Minguet, E.G.; Grau-Enguix, F.; Abbas, M.; Locascio, A.; Thomas, S.G.; Alabadí, D.; Blázquez, M.A. Molecular mechanism for the interaction between gibberellin and brassinosteroid signaling pathways in Arabidopsis. Proceedings of the National Academy of Sciences 2012, 109, 13446–13451. [Google Scholar] [CrossRef] [PubMed]
- Gehring, M.; Bubb, K.L.; Henikoff, S. Extensive demethylation of repetitive elements during seed development underlies gene imprinting. Science 2009, 324, 1447–1451. [Google Scholar] [CrossRef] [PubMed]
- Gehring, M.; Huh, J.H.; Hsieh, T.-F.; Penterman, J.; Choi, Y.; Harada, J.J.; Goldberg, R.B.; Fischer, R.L. DEMETER DNA glycosylase establishes MEDEA polycomb gene self-imprinting by allele-specific demethylation. Cell 2006, 124, 495–506. [Google Scholar] [CrossRef] [PubMed]
- Goeckeritz, C.Z.; Zheng, X.; Harkess, A.; Dresselhaus, T. Widespread application of apomixis in agriculture requires further study of natural apomicts. iScience 2024, 27, 110720. [Google Scholar] [CrossRef] [PubMed]
- Gombos, M.; Raynaud, C.; Nomoto, Y.; Molnár, E.; Brik-Chaouche, R.; Takatsuka, H.; Zaki, A.; Bernula, D.; Latrasse, D.; Mineta, K.; Nagy, F.; He, X.; Iwakawa, H.; Őszi, E.; An, J.; Suzuki, T.; Papdi, C.; Bergis, C.; Benhamed, M.; Bögre, L.; Ito, M.; Magyar, Z. The canonical E2Fs together with RETINOBLASTOMA-RELATED are required to establish quiescence during plant development. Commun Biol 2023, 6, 903. [Google Scholar] [CrossRef]
- Guitton, A.-E.; Berger, F. Loss of function of MULTICOPY SUPPRESSOR OF IRA 1 produces nonviable parthenogenetic embryos in Arabidopsis. Current Biology 2005, 15, 750–754. [Google Scholar] [CrossRef]
- Guo, C.; Jiang, Y.; Shi, M.; Wu, X.; Wu, G. ABI5 acts downstream of miR159 to delay vegetative phase change in Arabidopsis. New Phytologist 2021, 231, 339–350. [Google Scholar] [CrossRef]
- Guo, L.; Jiang, L.; Zhang, Y.; Lu, X.-L.; Xie, Q.; Weijers, D.; Liu, C.-M. The anaphase-promoting complex initiates zygote division in Arabidopsis through degradation of cyclin B1. The Plant Journal 2016, 86, 161–174. [Google Scholar] [CrossRef]
- Guo, L.; Luo, X.; Li, M.; Joldersma, D.; Plunkert, M.; Liu, Z. Mechanism of fertilization-induced auxin synthesis in the endosperm for seed and fruit development. Nature Communications 2022, 13, 3985. [Google Scholar] [CrossRef]
- Hamamura, Y.; Nishimaki, M.; Takeuchi, H.; Geitmann, A.; Kurihara, D.; Higashiyama, T. Live imaging of calcium spikes during double fertilization in Arabidopsis. Nature Communications 2014, 5, 4722. [Google Scholar] [CrossRef]
- Hamann, T. Plant cell wall integrity maintenance as an essential component of biotic stress response mechanisms. Frontiers in plant science 2012, 3, 77. [Google Scholar] [CrossRef]
- He, W.; Li, W.; Luo, X.; Tang, Y.; Wang, L.; Yu, F.; Lin, Q. Rice FERONIA-LIKE RECEPTOR 3 and 14 affect grain quality by regulating redox homeostasis during endosperm development. Journal of Experimental Botany 2023, 74, 3003–3018. [Google Scholar] [CrossRef]
- Hehenberger, E.; Kradolfer, D.; Köhler, C. Endosperm cellularization defines an important developmental transition for embryo development. Development 2012, 139, 2031–2039. [Google Scholar] [CrossRef]
- Heidemann, B.; Primetis, E.; Zahn, I.E.; Underwood, C.J. To infinity and beyond: recent progress, bottlenecks, and potential of clonal seeds by apomixis. Plant J 2025, 121, e70054. [Google Scholar] [CrossRef]
- Honma, Y.; Adhikari, P.B.; Kuwata, K.; Kagenishi, T.; Yokawa, K.; Notaguchi, M.; Kurotani, K.; Toda, E.; Bessho-Uehara, K.; Liu, X.; Zhu, S.; Wu, X.; Kasahara, R.D. High-quality sugar production by osgcs1 rice. Communications Biology 2020, 3, 617. [Google Scholar] [CrossRef]
- Howell, S.H. Endoplasmic reticulum stress responses in plants. Annual Review of Plant Biology 2013, 64, 477–499. [Google Scholar] [CrossRef]
- Hu, Y.; Zhou, L.; Huang, M.; He, X.; Yang, Y.; Liu, X.; Li, Y.; Hou, X. Gibberellins play an essential role in late embryogenesis of Arabidopsis. Nature Plants 2018, 4, 289–298. [Google Scholar] [CrossRef]
- Huang, J.; Zhao, L.; Malik, S.; Gentile, B.R.; Xiong, V.; Arazi, T.; Owen, H.A.; Friml, J.; Zhao, D. Specification of female germline by microRNA orchestrated auxin signaling in Arabidopsis. Nature Communications 2022, 13, 6960. [Google Scholar] [CrossRef]
- Huang, W.J.; Liu, H.K.; McCormick, S.; Tang, W.H. Tomato pistil factor STIG1 promotes in vivo pollen tube growth by binding to phosphatidylinositol 3-phosphate and the extracellular domain of the pollen receptor kinase LePRK2. The Plant Cell 2014, 26, 2505–2523. [Google Scholar] [CrossRef]
- Ingouff, M.; Hamamura, Y.; Gourgues, M.; Higashiyama, T.; Berger, F. Distinct dynamics of HISTONE3 variants between the two fertilization products in plants. Curr Biol 2007, 17, 1032–1037. [Google Scholar] [CrossRef]
- Ingram, G.C. Family life at close quarters: communication and constraint in angiosperm seed development. Protoplasma 2010, 247, 195–214. [Google Scholar] [CrossRef]
- Jiang, T.; Li, L.; Hu, Q.; Kuang, X.; Zhang, L.; Zeng, W.; Miki, D.; Zheng, B. The DNA methylation–demethylation balance prevents development of multiple megaspore mother cells in Arabidopsis. The Plant Cell 2025, 37. [Google Scholar] [CrossRef]
- Jiang, W.-B.; Huang, H.-Y.; Hu, Y.-W.; Zhu, S.-W.; Wang, Z.-Y.; Lin, W.-H. Brassinosteroid regulates seed size and shape. Arabidopsis Plant Physiology 2013, 162, 1965–1977. [Google Scholar] [CrossRef]
- Johnson, K.L.; Degnan, K.A.; Ross Walker, J.; Ingram, G.C. AtDEK1 is essential for specification of embryonic epidermal cell fate. The Plant Journal 2005, 44, 114–127. [Google Scholar] [CrossRef]
- Johnston, A.J.; Matveeva, E.; Kirioukhova, O.; Grossniklaus, U.; Gruissem, W. A dynamic reciprocal RBR-PRC2 regulatory circuit controls Arabidopsis gametophyte development. Current Biology 2008, 18, 1680–1686. [Google Scholar] [CrossRef]
- Jullien, P.E.; Mosquna, A.; Ingouff, M.; Sakata, T.; Ohad, N.; Berger, F. Retinoblastoma and its binding partner MSI1 control imprinting in Arabidopsis. PLoS Biol 2008, 6, e194. [Google Scholar] [CrossRef]
- Jun, J.; Fiume, E.; Roeder, A.H.K.; Meng, L.; Sharma, V.K.; Osmont, K.S.; Baker, C.; Ha, C.M.; Meyerowitz, E.M.; Feldman, L.J.; Fletcher, J.C. Comprehensive analysis of CLE polypeptide signaling gene expression and overexpression activity in Arabidopsis. Plant Physiology 2010, 154, 1721–1736. [Google Scholar] [CrossRef]
- Kang, X.; Li, W.; Zhou, Y.; Ni, M. A WRKY Transcription Factor Recruits the SYG1-Like Protein SHB1 to Activate Gene Expression and Seed Cavity Enlargement. PLOS Genetics 2013, 9, e1003347. [Google Scholar] [CrossRef]
- Kao, P.; Nodine, M.D. Transcriptional activation of Arabidopsis zygotes is required for initial cell divisions. Scientific Reports 2019, 9, 17159. [Google Scholar] [CrossRef]
- Kasahara, R.D. Fertilization initiates seed nutrition via phloem end by a callose degradation enzyme. DNA and Cell Biology 2025, 44, 407–410. [Google Scholar] [CrossRef]
- Kasahara, R.D.; Notaguchi, M.; Nagahara, S.; Suzuki, T.; Susaki, D.; Honma, Y.; Maruyama, D.; Higashiyama, T. Pollen tube contents initiate ovule enlargement and enhance seed coat development without fertilization. Science Advances 2016, 2, e1600554. [Google Scholar] [CrossRef]
- Ke, Y.; Podio, M.; Conner, J.; Ozias-Akins, P. Single-cell transcriptome profiling of buffelgrass (Cenchrus ciliaris) eggs unveils apomictic parthenogenesis signatures. Sci Rep 2021, 11, 9880. [Google Scholar] [CrossRef]
- Khanday, I.; Skinner, D.; Yang, B.; Mercier, R.; Sundaresan, V. A male-expressed rice embryogenic trigger redirected for asexual propagation through seeds. Nature 2019, 565, 91–95. [Google Scholar] [CrossRef]
- Köhler, C.; Hennig, L.; Bouveret, R.; Gheyselinck, J.; Grossniklaus, U.; Gruissem, W. Arabidopsis MSI1 is a component of the MEA/FIE Polycomb group complex and required for seed development. The EMBO Journal 2003, 22, 4804–4814. [Google Scholar] [CrossRef]
- Kradolfer, D.; Hennig, L.; Kohler, C. Increased maternal genome dosage bypasses the requirement of the FIS Polycomb Repressive Complex 2 in Arabidopsis seed development. PLoS Genet 2013, 9, e1003163. [Google Scholar] [CrossRef]
- Kranz, E.; Lörz, H. In vitro fertilisation of maize by single egg and sperm cell protoplast fusion mediated by high calcium and high pH. Zygote 1994, 2, 125–128. [Google Scholar] [CrossRef]
- Kujirai, T.; Echigoya, K.; Kishi, Y.; Saeki, M.; Ito, T.; Kato, J.; Negishi, L.; Kimura, H.; Masumoto, H.; Takizawa, Y.; Gotoh, Y.; Kurumizaka, H. Structural insights into how DEK nucleosome binding facilitates H3K27 trimethylation in chromatin. Nature Structural & Molecular Biology 2025, 32, 1183–1192. [Google Scholar] [CrossRef]
- Lafon-Placette, C.; Köhler, C. Embryo and endosperm, partners in seed development. Current Opinion in Plant Biology 2014, 17, 64–69. [Google Scholar] [CrossRef]
- Larsson, E.; Vivian-Smith, A.; Offringa, R.; Sundberg, E. Auxin homeostasis in arabidopsis ovules is anther-dependent at maturation and changes dynamically upon fertilization. Frontiers in Plant Science 2017, 8. [Google Scholar] [CrossRef]
- Laux, T.; Mayer, K.F.X.; Berger, J.; Jürgens, G. The WUSCHEL gene is required for shoot and floral meristem integrity in Arabidopsis. Development 1996, 122, 87–96. [Google Scholar] [CrossRef]
- Lee, H.; Fischer, R.L.; Goldberg, R.B.; Harada, J.J. Arabidopsis LEAFY COTYLEDON1 represents a functionally specialized subunit of the CCAAT binding transcription factor. Proceedings of the National Academy of Sciences 2003, 100, 2152–2156. [Google Scholar] [CrossRef]
- Li, C.; Hu, F.; Chen, H.; Zhao, J. Transcriptome characteristics during cell wall formation of endosperm cellularization and embryo differentiation in Arabidopsis. Front Plant Sci 2022, 13, 998664. [Google Scholar] [CrossRef]
- Lid, S.E.; Gruis, D.; Jung, R.; Lorentzen, J.A.; Ananiev, E.; Chamberlin, M.; Niu, X.; Meeley, R.; Nichols, S.; Olsen, O.-A. The defective kernel 1 (dek1) gene required for aleurone cell development in the endosperm of maize grains encodes a membrane protein of the calpain gene superfamily. Proceedings of the National Academy of Sciences 2002, 99, 5460–5465. [Google Scholar] [CrossRef]
- Lima, R.B.; Pankaj, R.; Ehlert, S.T.; Finger, P.; Fröhlich, A.; Bayle, V.; Landrein, B.; Sampathkumar, A.; Figueiredo, D.D. Seed coat-derived brassinosteroid signaling regulates endosperm development. Nature Communications 2024, 15, 9352. [Google Scholar] [CrossRef]
- Liu, H.; Luo, Q.; Tan, C.; Song, J.; Zhang, T.; Men, S. Biosynthesis- and transport-mediated dynamic auxin distribution during seed development controls seed size in Arabidopsis. The Plant Journal 2023, 113, 1259–1277. [Google Scholar] [CrossRef]
- Liu, X.; Adhikari, P.; Kasahara, R. Pollen tube contents from failed fertilization contribute to seed coat initiation in Arabidopsis. F1000Research 2019, 8. [Google Scholar] [CrossRef]
- Liu, X.; Nakajima, K.P.; Adhikari, P.B.; Wu, X.; Zhu, S.; Okada, K.; Kagenishi, T.; Kurotani, K.-i.; Ishida, T.; Nakamura, M.; Sato, Y.; Kawakatsu, Y.; Xie, L.; Huang, C.; He, J.; Yokawa, K.; Sawa, S.; Higashiyama, T.; Bradford, K.J.; Notaguchi, M.; Kasahara, R.D. Fertilization-dependent phloem end gate regulates seed size. Current Biology 2025a, 35, 2049–2063.e2043. [Google Scholar] [CrossRef]
- Liu, X.Q.; Shi, J.J.; Fan, H.; Jiao, J.; Gao, L.; Tan, L.; Nagawa, S.; Wang, D.Y. Nuclear DNA replicates during zygote development in Arabidopsis and Torenia fournieri. Plant Physiology 2020, 185, 137–145. [Google Scholar] [CrossRef]
- Liu, Y.; Xiao, S.; Yang, M.; Guo, G.; Zhou, Y. The impact of Polycomb Group proteins on 3D chromatin structure and environmental stresses in plants. Plants 2025b, 14, 1038. [Google Scholar] [CrossRef]
- Lu, B.; Wang, S.; Feng, H.; Wang, J.; Zhang, K.; Li, Y.; Wu, P.; Zhang, M.; Xia, Y.; Peng, C.; Li, C. FERONIA-mediated TIR1/AFB2 oxidation stimulates auxin signaling in Arabidopsis. Molecular Plant 2024, 17, 772–787. [Google Scholar] [CrossRef]
- Lucas, W.J.; Bouché-Pillon, S.; Jackson, D.P.; Nguyen, L.; Baker, L.; Ding, B.; Hake, S. Selective trafficking of KNOTTED1 homeodomain protein and its mRNA through plasmodesmata. Science 1995, 270, 1980–1983. [Google Scholar] [CrossRef]
- Lukowitz, W.; Roeder, A.; Parmenter, D.; Somerville, C. A MAPKK kinase gene regulates extra-embryonic cell fate in Arabidopsis. Cell 2004, 116, 109–119. [Google Scholar] [CrossRef]
- Lv, Y.; Li, J.; Wang, Z.; Liu, Y.; Jiang, Y.; Li, Y.; Lv, Z.; Huang, X.; Peng, X.; Cao, Y.; Yang, H. Polycomb proteins RING1A/B promote H2A monoubiquitination to regulate female gametophyte development in Arabidopsis. Journal of Experimental Botany 2024, 75, 4822–4836. [Google Scholar] [CrossRef]
- Maruyama, D.; Völz, R.; Takeuchi, H.; Mori, T.; Igawa, T.; Kurihara, D.; Kawashima, T.; Ueda, M.; Ito, M.; Umeda, M.; Nishikawa, S.I.; Groß-Hardt, R.; Higashiyama, T. Rapid elimination of the persistent synergid through a cell fusion mechanism. Cell 2015, 161, 907–918. [Google Scholar] [CrossRef]
- Mathews, S.; Kramer, E.M. The evolution of reproductive structures in seed plants: a re-examination based on insights from developmental genetics. New Phytologist 2012, 194, 910–923. [Google Scholar] [CrossRef]
- Matzke, M.A.; Mosher, R.A. RNA-directed DNA methylation: an epigenetic pathway of increasing complexity. Nature Reviews Genetics 2014, 15, 394–408. [Google Scholar] [CrossRef]
- Meixner, A.; Boldt, K.; Van Troys, M.; Askenazi, M.; Gloeckner, C.J.; Bauer, M.; Marto, J.A.; Ampe, C.; Kinkl, N.; Ueffing, M. A QUICK screen for Lrrk2 interaction partners--leucine-rich repeat kinase 2 is involved in actin cytoskeleton dynamics. Mol Cell Proteomics 2011, 10, M110.001172. [Google Scholar] [CrossRef]
- Mendes, M.A.; Petrella, R.; Cucinotta, M.; Vignati, E.; Gatti, S.; Pinto, S.C.; Bird, D.C.; Gregis, V.; Dickinson, H.; Tucker, M.R.; Colombo, L. The RNA-dependent DNA methylation pathway is required to restrict SPOROCYTELESS/NOZZLE expression to specify a single female germ cell precursor in Arabidopsis. Development 2020, 147. [Google Scholar] [CrossRef]
- Migicovsky, Z.; Kovalchuk, I. Epigenetic modifications during angiosperm gametogenesis. Front Plant Sci 2012, 3, 20. [Google Scholar] [CrossRef]
- Möller, B.; Weijers, D. Auxin control of embryo patterning. Cold Spring Harb Perspect Biol 2009, 1, a001545. [Google Scholar] [CrossRef]
- Montgomery, S.A.; Berger, F. Epigenetic reprogramming of imprinting at meiosis. bioRxiv 2023, 2017.541143. [Google Scholar] [CrossRef]
- Moreno-Romero, J.; Jiang, H.; Santos-González, J.; Köhler, C. Parental epigenetic asymmetry of PRC2-mediated histone modifications in the Arabidopsis endosperm. The EMBO Journal 2016, 35, 1298–1311. [Google Scholar] [CrossRef]
- Moussu, S.; Doll, N.M.; Chamot, S.; Brocard, L.; Creff, A.; Fourquin, C.; Widiez, T.; Nimchuk, Z.L.; Ingram, G. ZHOUPI and KERBEROS mediate embryo/endosperm separation by promoting the formation of an extracuticular sheath at the embryo surface. Plant Cell 2017, 29, 1642–1656. [Google Scholar] [CrossRef]
- Muller, H.J. The relation of recombination to mutational advance. Mutation Research/Fundamental and Molecular Mechanisms of Mutagenesis 1964, 1, 2–9. [Google Scholar] [CrossRef]
- Murata, Y.; Pei, Z.M.; Mori, I.C.; Schroeder, J. Abscisic acid activation of plasma membrane Ca2+ channels in guard cells requires cytosolic NAD(P)H and is differentially disrupted upstream and downstream of reactive oxygen species production in abi1-1 and abi2-1 protein phosphatase 2C mutants. Plant Cell 2001, 13, 2513–2523. [Google Scholar] [CrossRef]
- Nakajima, K.P.; Kasahara, R.D. How do seeds get bigger? A newly identified phloem-terminal tissue controls seed enlargement through callose degradation. Academia Molecular Biology and Genomics 2025, 2. [Google Scholar] [CrossRef]
- Nakamura, M.; Bente, H.; Derkacheva, M.; Gentry, M.; Landberg, K.; Thelander, M.; Sundberg, E.; Hennig, L.; Köhler, C. Chromatin-associated DEK proteins maintain H3K27me3 balance and coordinate developmental transitions in plants. New Phytol 2026, 249, 930–944. [Google Scholar] [CrossRef]
- Nodine, M.D.; Bartel, D.P. Maternal and paternal genomes contribute equally to the transcriptome of early plant embryos. Nature 2012, 482, 94–97. [Google Scholar] [CrossRef]
- Nowack, M.K.; Grini, P.E.; Jakoby, M.J.; Lafos, M.; Koncz, C.; Schnittger, A. A positive signal from the fertilization of the egg cell sets off endosperm proliferation in angiosperm embryogenesis. Nature Genetics 2006, 38, 63–67. [Google Scholar] [CrossRef]
- O'Neill, J.P.; Colon, K.T.; Jenik, P.D. The onset of embryo maturation in Arabidopsis is determined by its developmental stage and does not depend on endosperm cellularization. Plant J 2019, 99, 286–301. [Google Scholar] [CrossRef]
- Ogawa, E.; Yamada, Y.; Sezaki, N.; Kosaka, S.; Kondo, H.; Kamata, N.; Abe, M.; Komeda, Y.; Takahashi, T. ATML1 and PDF2 play a redundant and essential role in Arabidopsis embryo development. Plant and Cell Physiology 2015, 56, 1183–1192. [Google Scholar] [CrossRef]
- Olmedo-Monfil, V.; Durán-Figueroa, N.; Arteaga-Vázquez, M.; Demesa-Arévalo, E.; Autran, D.; Grimanelli, D.; Slotkin, R.K.; Martienssen, R.A.; Vielle-Calzada, J.P. Control of female gamete formation by a small RNA pathway in Arabidopsis. Nature 2010, 464, 628–632. [Google Scholar] [CrossRef] [PubMed]
- Pankaj, R.; Lima, R.B.; Figueiredo, D.D. Hormonal regulation and crosstalk during early endosperm and seed coat development. Plant Reprod 2024a, 38, 5. [Google Scholar] [CrossRef]
- Pankaj, R.; Lima, R.B.; Luo, G.-Y.; Ehlert, S.; del Toro-de León, G.; Bente, H.; Finger, P.; Sato, H.; Figueiredo, D.D. BRI1-mediated removal of seed coat H3K27me3 marks is a brassinosteroid-independent process. bioRxiv 2024b, 2007.569203. [Google Scholar]
- Park, J.-S.; Frost, J.M.; Park, K.; Ohr, H.; Park, G.T.; Kim, S.; Eom, H.; Lee, I.; Brooks, J.S.; Fischer, R.L.; Choi, Y. Control of DEMETER DNA demethylase gene transcription in male and female gamete companion cells in Arabidopsis thaliana. Proceedings of the National Academy of Sciences 2017, 114, 2078–2083. [Google Scholar] [CrossRef]
- Pelletier, J.M.; Kwong, R.W.; Park, S.; Le, B.H.; Baden, R.; Cagliari, A.; Hashimoto, M.; Munoz, M.D.; Fischer, R.L.; Goldberg, R.B.; Harada, J.J. LEC1 sequentially regulates the transcription of genes involved in diverse developmental processes during seed development. Proceedings of the National Academy of Sciences 2017, 114, E6710–E6719. [Google Scholar] [CrossRef]
- Plong, A.; Rodriguez, K.; Alber, M.; Chen, W.; Reddy, G.V. CLAVATA3 mediated simultaneous control of transcriptional and post-translational processes provides robustness to the WUSCHEL gradient. Nature Communications 2021, 12, 6361. [Google Scholar] [CrossRef]
- Qin, Y.; Zhao, L.; Skaggs, M.I.; Andreuzza, S.; Tsukamoto, T.; Panoli, A.; Wallace, K.N.; Smith, S.; Siddiqi, I.; Yang, Z.; Yadegari, R.; Palanivelu, R. ACTIN-RELATED PROTEIN6 regulates female meiosis by modulating meiotic gene expression in Arabidopsis. The Plant Cell 2014, 26, 1612–1628. [Google Scholar] [CrossRef]
- Rampey, R.A.; LeClere, S.; Kowalczyk, M.; Ljung, K.; Sandberg, G.; Bartel, B. A family of auxin-conjugate hydrolases that contributes to free indole-3-acetic acid levels during Arabidopsis germination. Plant Physiol 2004, 135, 978–988. [Google Scholar] [CrossRef]
- Rani, S.; Roy, S.; Nandi, A.K. Pathogen induces MEDEA gene in a parent-of-origin nonspecific manner and causes embryo lethality in MEDEA overexpressing plants. Journal of Plant Biochemistry and Biotechnology 2025, 34, 933–940. [Google Scholar] [CrossRef]
- Reyes, J.L.; Chua, N.-H. ABA induction of miR159 controls transcript levels of two MYB factors during Arabidopsis seed germination. The Plant Journal 2007, 49, 592–606. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Leal, D.; León-Martínez, G.; Abad-Vivero, U.; Vielle-Calzada, J.-P. Natural variation in epigenetic pathways affects the specification of female gamete precursors in Arabidopsis. The Plant Cell 2015, 27, 1034–1045. [Google Scholar] [CrossRef]
- Ronceret, A.; Devic, M. Redrawing the borderline: Control of DNA replication at fertilization. Plant Signal Behav 2008, 3, 554–555. [Google Scholar] [CrossRef]
- Roszak, P.; Köhler, C. Polycomb group proteins are required to couple seed coat initiation to fertilization. Proceedings of the National Academy of Sciences 2011, 108, 20826–20831. [Google Scholar] [CrossRef] [PubMed]
- Rudall, P.J. Evolution and patterning of the ovule in seed plants. Biological Reviews 2021, 96, 943–960. [Google Scholar] [CrossRef] [PubMed]
- Rydin, C.; Khodabandeh, A.; Endress, P.K. The female reproductive unit of Ephedra (Gnetales): comparative morphology and evolutionary perspectives. Botanical Journal of the Linnean Society 2010, 163, 387–430. [Google Scholar] [CrossRef]
- Sabelli, P.A.; Liu, Y.; Dante, R.A.; Lizarraga, L.E.; Nguyen, H.N.; Brown, S.W.; Klingler, J.P.; Yu, J.; LaBrant, E.; Layton, T.M.; Feldman, M.; Larkins, B.A. Control of cell proliferation, endoreduplication, cell size, and cell death by the retinoblastoma-related pathway in maize endosperm. Proc Natl Acad Sci U S A 2013, 110, E1827-1836. [Google Scholar] [CrossRef]
- Schröder, J.A.; Bonnet, D.M.V.; Jullien, P.E. Non-cell-autonomous small RNA silencing in Arabidopsis female gametes. Current Biology 2023, 33, 183–188.e183. [Google Scholar] [CrossRef] [PubMed]
- Schruff, M.C.; Spielman, M.; Tiwari, S.; Adams, S.; Fenby, N.; Scott, R.J. The AUXIN RESPONSE FACTOR 2 gene of Arabidopsis links auxin signalling, cell division, and the size of seeds and other organs. Development 2006, 133, 251–261. [Google Scholar] [CrossRef]
- Simonini, S. Regulation of cell cycle in plant gametes: when is the right time to divide? Development 2025, 152. [Google Scholar] [CrossRef]
- Simonini, S.; Bemer, M.; Bencivenga, S.; Gagliardini, V.; Pires, N.D.; Desvoyes, B.; Van der Graaff, E.; Gutierrez, C.; Grossniklaus, U. The Polycomb group protein MEDEA controls cell proliferation and embryonic patterning in Arabidopsis. Developmental Cell 2021, 56, 1945–1960. e1947. [Google Scholar] [CrossRef]
- Simonini, S.; Bencivenga, S.; Grossniklaus, U. A paternal signal induces endosperm proliferation upon fertilization in Arabidopsis. Science 2024, 383, 646–653. [Google Scholar] [CrossRef] [PubMed]
- Solfanelli, C.; Ceron, F.; Paolicchi, F.; Giorgetti, L.; Geri, C.; Ceccarelli, N.; Kamiya, Y.; Picciarelli, P. Expression of two genes encoding gibberellin 2- and 3-oxidases in developing seeds of Phaseolus coccineus. Plant and Cell Physiology 2005, 46, 1116–1124. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Xie, X.; Chen, C.; Shu, J.; Thapa, R.K.; Nguyen, V.; Bian, S.; Kohalmi, S.E.; Marsolais, F.; Zou, J.; Cui, Y. LEAFY COTYLEDON1 expression in the endosperm enables embryo maturation in Arabidopsis. Nature Communications 2021, 12, 3963. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Xie, X.; Mavraganis, I.; Yu, B.; Shen, W.; Yang, H.; Xiang, D.; Wei, Y.; Cui, Y.; Zou, J. Tracking the genome-wide occupancy of Arabidopsis LEAFY COTYLEDON1 in endosperm development. Commun Biol 2025, 8, 771. [Google Scholar] [CrossRef] [PubMed]
- Sornay, E.; Forzani, C.; Forero-Vargas, M.; Dewitte, W.; Murray, J.A.H. Activation of CYCD7;1 in the central cell and early endosperm overcomes cell-cycle arrest in the Arabidopsis female gametophyte, and promotes early endosperm and embryo development. The Plant Journal 2015, 84, 41–55. [Google Scholar] [CrossRef]
- Szczuka, E.; Szczuka, A. Cuticle fluorescence during embryogenesis of Arabidopsis thaliana (L.) Heynh. Acta Biol. Cracov. Ser. Bot 2003, 45, 63–67. [Google Scholar]
- Takeuchi, N.; Kaneko, K.; Koonin, E.V. Horizontal gene transfer can rescue prokaryotes from muller’s ratchet: Benefit of DNA from dead cells and population subdivision. G3 GenesGenomesGenetics 2014, 4, 325–339. [Google Scholar] [CrossRef]
- Tanaka, H.; Onouchi, H.; Kondo, M.; Hara-Nishimura, I.; Nishimura, M.; Machida, C.; Machida, Y. A subtilisin-like serine protease is required for epidermal surface formation in Arabidopsis embryos and juvenile plants. Development 2001, 128, 4681–4689. [Google Scholar] [CrossRef]
- Tang, W.; Kelley, D.; Ezcurra, I.; Cotter, R.; McCormick, S. LeSTIG1, an extracellular binding partner for the pollen receptor kinases LePRK1 and LePRK2, promotes pollen tube growth in vitro. The Plant Journal 2004, 39, 343–353. [Google Scholar] [CrossRef] [PubMed]
- Tovar-Aguilar, A.; Grimanelli, D.; Acosta-García, G.; Vielle-Calzada, J.-P.; Badillo-Corona, J.A.; Durán-Figueroa, N. The miRNA822 loaded by ARGONAUTE9 modulates the monosporic female gametogenesis in Arabidopsis thaliana. Plant Reproduction 2024, 37, 243–258. [Google Scholar] [CrossRef]
- Tsering, T.; Di Donato, M.; Samakovli, D.; Milioni, D.; Iacobini, F.R.; Panagiotopoulos, K.; Plitsi, P.-K.; Azarello, E.; Mancuso, S.; Pukyšová, V.; Zwiewka, M.; Nodzynski, T.; Stumpe, M.; Ludwig-Müller, J.; Bailly, A.; Hatzopoulos, P.; Geisler, M.M. HSP90 differentially stabilizes plant ABCB-type auxin transporters on the plasma membrane. Nature Communications 2025, 16, 8643. [Google Scholar] [CrossRef]
- Tsuwamoto, R.; Fukuoka, H.; Takahata, Y. GASSHO1 and GASSHO2 encoding a putative leucine-rich repeat transmembrane-type receptor kinase are essential for the normal development of the epidermal surface in Arabidopsis embryos. The Plant Journal 2008, 54, 30–42. [Google Scholar] [CrossRef]
- Turck, F.; Roudier, F.; Farrona, S.; Martin-Magniette, M.-L.; Guillaume, E.; Buisine, N.; Gagnot, S.; Martienssen, R.A.; Coupland, G.; Colot, V. Arabidopsis TFL2/LHP1 specifically associates with genes marked by trimethylation of histone H3 lysine 27. PLOS Genetics 2007, 3, e86. [Google Scholar] [CrossRef]
- Ueda, M.; Aichinger, E.; Gong, W.; Groot, E.; Verstraeten, I.; Dai Vu, L.; De Smet, I.; Higashiyama, T.; Umeda, M.; Laux, T. Transcriptional integration of paternal and maternal factors in the Arabidopsis zygote. Genes & development 2017, 31, 617–627. [Google Scholar]
- Ueda, M.; Zhang, Z.; Laux, T. Transcriptional activation of Arabidopsis axis patterning genes WOX8/9 links zygote polarity to embryo development. Developmental Cell 2011, 20, 264–270. [Google Scholar] [CrossRef]
- Van Durme, M.; Olvera-Carrillo, Y.; Pfeiffer, M.L.; Doll, N.M.; De Winter, F.; Lin, Z.; Nowack, M.K. Fertility loss in senescing Arabidopsis ovules is controlled by the maternal sporophyte via a NAC transcription factor triad. Proceedings of the National Academy of Sciences 2023, 120, e2219868120. [Google Scholar] [CrossRef]
- Vanneste, S.; Pei, Y.; Friml, J. Mechanisms of auxin action in plant growth and development. Nature Reviews Molecular Cell Biology 2025, 26, 648–666. [Google Scholar] [CrossRef] [PubMed]
- Veluchamy, A.; Jégu, T.; Ariel, F.; Latrasse, D.; Mariappan, K.G.; Kim, S.K.; Crespi, M.; Hirt, H.; Bergounioux, C.; Raynaud, C.; Benhamed, M. LHP1 regulates H3K27me3 spreading and shapes the three-dimensional conformation of the Arabidopsis genome. PLoS One 2016, 11, e0158936. [Google Scholar] [CrossRef]
- Wang, H.; Ngwenyama, N.; Liu, Y.; Walker, J.C.; Zhang, S. Stomatal development and patterning are regulated by environmentally responsive mitogen-activated protein kinases in Arabidopsis. Plant Cell 2007, 19, 63–73. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Ko, E.E.; Tran, J.; Qiao, H. TREE1-EIN3–mediated transcriptional repression inhibits shoot growth in response to ethylene. Proceedings of the National Academy of Sciences 2020, 117, 29178–29189. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Wan, J.; Dang, K.; Meng, S.; Hu, D.; Lin, Y.; Qiu, X.; Guo, Z.; Fu, Z.; Ding, D.; Tang, J. zma-miR159 targets ZmMYB74 and ZmMYB138 transcription factors to regulate grain size and weight in maize. Plant Physiology 2023, 193, 2430–2441. [Google Scholar] [CrossRef]
- Wang, Y.; Hou, Y.; Gu, H.; Kang, D.; Chen, Z.; Liu, J.; Qu, L.-J. The Arabidopsis APC4 subunit of the anaphase-promoting complex/cyclosome (APC/C) is critical for both female gametogenesis and embryogenesis. The Plant Journal 2012, 69, 227–240. [Google Scholar] [CrossRef]
- Wei, B.; Zhang, J.; Pang, C.; Yu, H.; Guo, D.; Jiang, H.; Ding, M.; Chen, Z.; Tao, Q.; Gu, H.; Qu, L.-J.; Qin, G. The molecular mechanism of SPOROCYTELESS/NOZZLE in controlling Arabidopsis ovule development. Cell Research 2015, 25, 121–134. [Google Scholar] [CrossRef]
- Weimer, A.K.; Matos, J.L.; Sharma, N.; Patell, F.; Murray, J.A.H.; Dewitte, W.; Bergmann, D.C. Lineage- and stage-specific expressed CYCD7;1 coordinates the single symmetric division that creates stomatal guard cells. Development (Cambridge, England) 2018, pp. dev160671. [Google Scholar] [CrossRef]
- Weinhofer, I.; Hehenberger, E.; Roszak, P.; Hennig, L.; Köhler, C. H3K27me3 profiling of the endosperm implies exclusion of polycomb group protein targeting by DNA methylation. PLOS Genetics 2010, 6, e1001152. [Google Scholar] [CrossRef] [PubMed]
- Wu, G.; Otegui, M.S.; Spalding, E.P. The ER-Localized TWD1 immunophilin is necessary for localization of multidrug resistance-like proteins required for polar auxin transport in Arabidopsis roots. The Plant Cell 2010, 22, 3295–3304. [Google Scholar] [CrossRef]
- Wu, Y.; Zhao, X.; Chen, J.X.; Chaumier, T.; Tirichine, L. The RING1 subunit of Polycomb Repressive Complex 1 monoubiquitinates histone H2A and mediates repression independently of Polycomb Repressive Complex 2 in the model diatom Phaeodactylum tricornutum. New Phytologist 2025. [Google Scholar] [CrossRef] [PubMed]
- Wyrzykowska, A.; Bielewicz, D.; Plewka, P.; Sołtys-Kalina, D.; Wasilewicz-Flis, I.; Marczewski, W.; Jarmolowski, A.; Szweykowska-Kulinska, Z. The MYB33, MYB65, and MYB101 transcription factors affect Arabidopsis and potato responses to drought by regulating the ABA signaling pathway. Physiologia Plantarum 2022, 174, e13775. [Google Scholar] [CrossRef]
- Xiao, W.; Gehring, M.; Choi, Y.; Margossian, L.; Pu, H.; Harada, J.J.; Goldberg, R.B.; Pennell, R.I.; Fischer, R.L. Imprinting of the MEA polycomb gene is controlled by antagonism between MET1 methyltransferase and DME glycosylase. Developmental Cell 2003, 5, 891–901. [Google Scholar] [CrossRef] [PubMed]
- Xiong, H.; Wang, W.; Sun, M.-X. Endosperm development is an autonomously programmed process independent of embryogenesis. The Plant Cell 2021, 33, 1151–1160. [Google Scholar] [CrossRef]
- Xu, T.-T.; Ren, S.-C.; Song, X.-F.; Liu, C.-M. CLE19 expressed in the embryo regulates both cotyledon establishment and endosperm development in Arabidopsis. Journal of Experimental Botany 2015, 66, 5217–5227. [Google Scholar] [CrossRef]
- Xu, W.; Fiume, E.; Coen, O.; Pechoux, C.; Lepiniec, L.; Magnani, E. Endosperm and nucellus develop antagonistically in Arabidopsis seeds. Plant Cell 2016, 28, 1343–1360. [Google Scholar] [CrossRef]
- Xu, W.; Iannaccone, M.; Gomez-Paez, D.-M.; Choinard, S.; Lu, J.; Le Hir, R.; Dinant, S.; Kalmbach, L.; Feil, R.; Lunn, J.E.; Meyer, C.; Magnani, E. Too much sugar makes plants ‘pregnant’: maternal sucrose signals fertilization in Arabidopsis seeds. bioRxiv 2025, 2025.2005.2016.654273. [Google Scholar] [CrossRef]
- Yamada, T.; Yokota, S.y.; Hirayama, Y.; Imaichi, R.; Kato, M.; Gasser, C.S. Ancestral expression patterns and evolutionary diversification of YABBY genes in angiosperms. The Plant Journal 2011, 67, 26–36. [Google Scholar] [CrossRef]
- Yang, S.; Johnston, N.; Talideh, E.; Mitchell, S.; Jeffree, C.; Goodrich, J.; Ingram, G. The endosperm-specific ZHOUPI gene of Arabidopsis thaliana regulates endosperm breakdown and embryonic epidermal development. Development 2008, 135, 3501–3509. [Google Scholar] [CrossRef] [PubMed]
- Yoshikawa, M.; Peragine, A.; Park, M.Y.; Poethig, R.S. A pathway for the biogenesis of trans-acting siRNAs in Arabidopsis. Genes & Development 2005, 19, 2164–2175. [Google Scholar]
- Yoshimura, K.; Iida, K.; Iida, H. MCAs in Arabidopsis are Ca2+-permeable mechanosensitive channels inherently sensitive to membrane tension. Nature Communications 2021, 12, 6074. [Google Scholar] [CrossRef]
- Yu, F.; Li, J.; Huang, Y.; Liu, L.; Li, D.; Chen, L.; Luan, S. FERONIA receptor kinase controls seed size in Arabidopsis thaliana. Molecular Plant 2014, 7, 920–922. [Google Scholar] [CrossRef]
- Yu, T.-Y.; Wang, P.; Lv, Y.; Wang, B.; Zhao, M.-R.; Dong, X.-W. Auxin orchestrates germ cell specification in Arabidopsis. International Journal of Molecular Sciences 2025, 26, 3257. [Google Scholar] [CrossRef]
- Yu, T.Y.; Shi, D.Q.; Jia, P.F.; Tang, J.; Li, H.J.; Liu, J.; Yang, W.C. The Arabidopsis receptor kinase ZAR1 is required for zygote asymmetric division and its daughter cell fate. PLoS Genet 2016, 12, e1005933. [Google Scholar] [CrossRef]
- Yu, X.; Zhang, X.; Zhao, P.; Peng, X.; Chen, H.; Bleckmann, A.; Bazhenova, A.; Shi, C.; Dresselhaus, T.; Sun, M.X. Fertilized egg cells secrete endopeptidases to avoid polytubey. Nature 2021, 592, 433–437. [Google Scholar] [CrossRef]
- Zhang, B.; Li, C.; Li, Y.; Yu, H. Mobile TERMINAL FLOWER1 determines seed size in Arabidopsis. Nature Plants 2020, 6, 1146–1157. [Google Scholar] [CrossRef]
- Zhang, J.; Yao, J.; He, K.; Yu, C.; Du, J.; Du, J.; Fu, Q.; Yao, R.; Howe, G.A.; Han, X.; Hu, Y. Jasmonate signaling coordinates with the SOD7–KLU pathway to regulate seed size in Arabidopsis thaliana. The Plant Cell 2025, 37. [Google Scholar] [CrossRef]
- Zhang, M.; Wu, H.; Su, J.; Wang, H.; Zhu, Q.; Liu, Y.; Xu, J.; Lukowitz, W.; Zhang, S. Maternal control of embryogenesis by MPK6 and its upstream MKK4/MKK5 in Arabidopsis. The Plant Journal 2017, 92, 1005–1019. [Google Scholar] [CrossRef]
- Zhang, S.; Mohanty, D.; Muzaffar, A.; Ni, M. Two MADS-box proteins, AGL9 and AGL15, recruit the FIS-PRC2 complex to trigger the phase transition from endosperm proliferation to embryo development in Arabidopsis. Molecular Plant 2024, 17, 1110–1128. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, B.; Yan, D.; Dong, W.; Yang, W.; Li, Q.; Zeng, L.; Wang, J.; Wang, L.; Hicks, L.M.; He, Z. Two Arabidopsis cytochrome P450 monooxygenases, CYP714A1 and CYP714A2, function redundantly in plant development through gibberellin deactivation. The Plant Journal 2011, 67, 342–353. [Google Scholar] [CrossRef]
- Zhao, H.; Wu, D.; Kong, F.; Lin, K.; Zhang, H.; Li, G. The Arabidopsis thaliana nuclear factor Y transcription factors. Front Plant Sci 2016, 7, 2045. [Google Scholar] [CrossRef]
- Zhao, L.; Cai, H.; Su, Z.; Wang, L.; Huang, X.; Zhang, M.; Chen, P.; Dai, X.; Zhao, H.; Palanivelu, R.; Chen, X.; Qin, Y. KLU suppresses megasporocyte cell fate through SWR1-mediated activation of WRKY28 expression in Arabidopsis. Proceedings of the National Academy of Sciences 2018a, 115, E526–E535. [Google Scholar] [CrossRef] [PubMed]
- Zhao, P.; Zhou, X.; Shen, K.; Liu, Z.; Cheng, T.; Liu, D.; Cheng, Y.; Peng, X.; Sun, M.-x. Two-step maternal-to-zygotic transition with two-phase parental genome contributions. Developmental Cell 2019, 49, 882–893.e885. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Chen, A.; Gao, Z.; Hou, F.; Chen, Y.; Liu, Y. Overexpression of HmWOX8 promotes callus proliferation and shoot regeneration by regulating hormone signaling and shoot development-related genes. Ornamental Plant Research 2024, 4, e026. [Google Scholar] [CrossRef]
- Zhao, Y.; Wang, S.; Wu, W.; Li, L.; Jiang, T.; Zheng, B. Clearance of maternal barriers by paternal miR159 to initiate endosperm nuclear division in Arabidopsis. Nature Communications 2018b, 9, 5011. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Q.; Zheng, Y.; Ji, H.; Burnie, W.; Perry, S.E. Gene regulation by the AGL15 transcription factor reveals hormone interactions in somatic embryogenesis. Plant Physiology 2016, 172, 2374–2387. [Google Scholar] [CrossRef] [PubMed]
- Ziegler, D.J.; Khan, D.; Kalichuk, J.L.; Becker, M.G.; Belmonte, M.F. Transcriptome landscape of the early Brassica napus seed. Journal of Integrative Plant Biology 2019, 61, 639–650. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




