Submitted:
15 February 2026
Posted:
27 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Methods
2.1. Animals
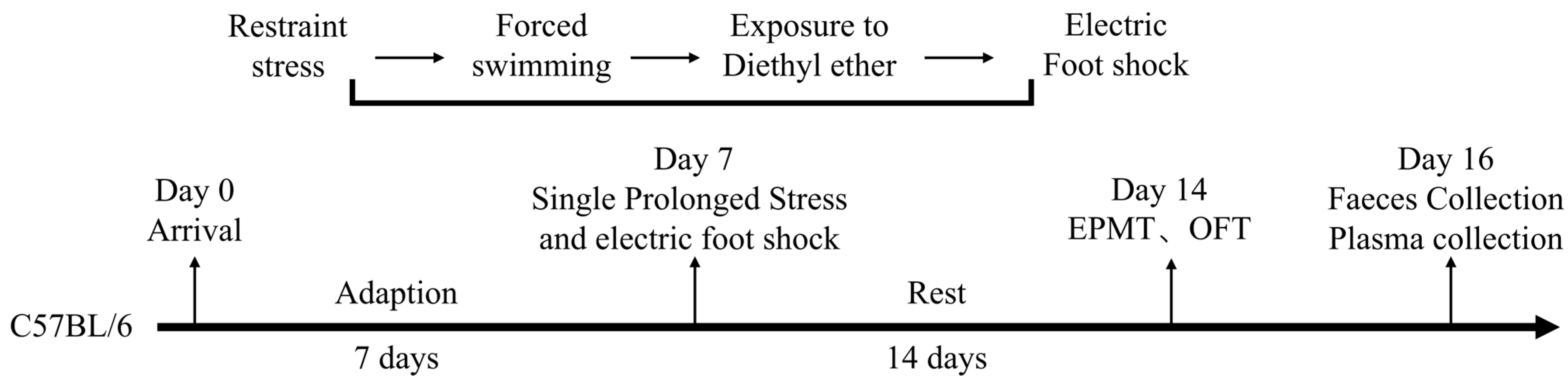
2.2. PTSD Model Establishment
2.3. Behavioral Test
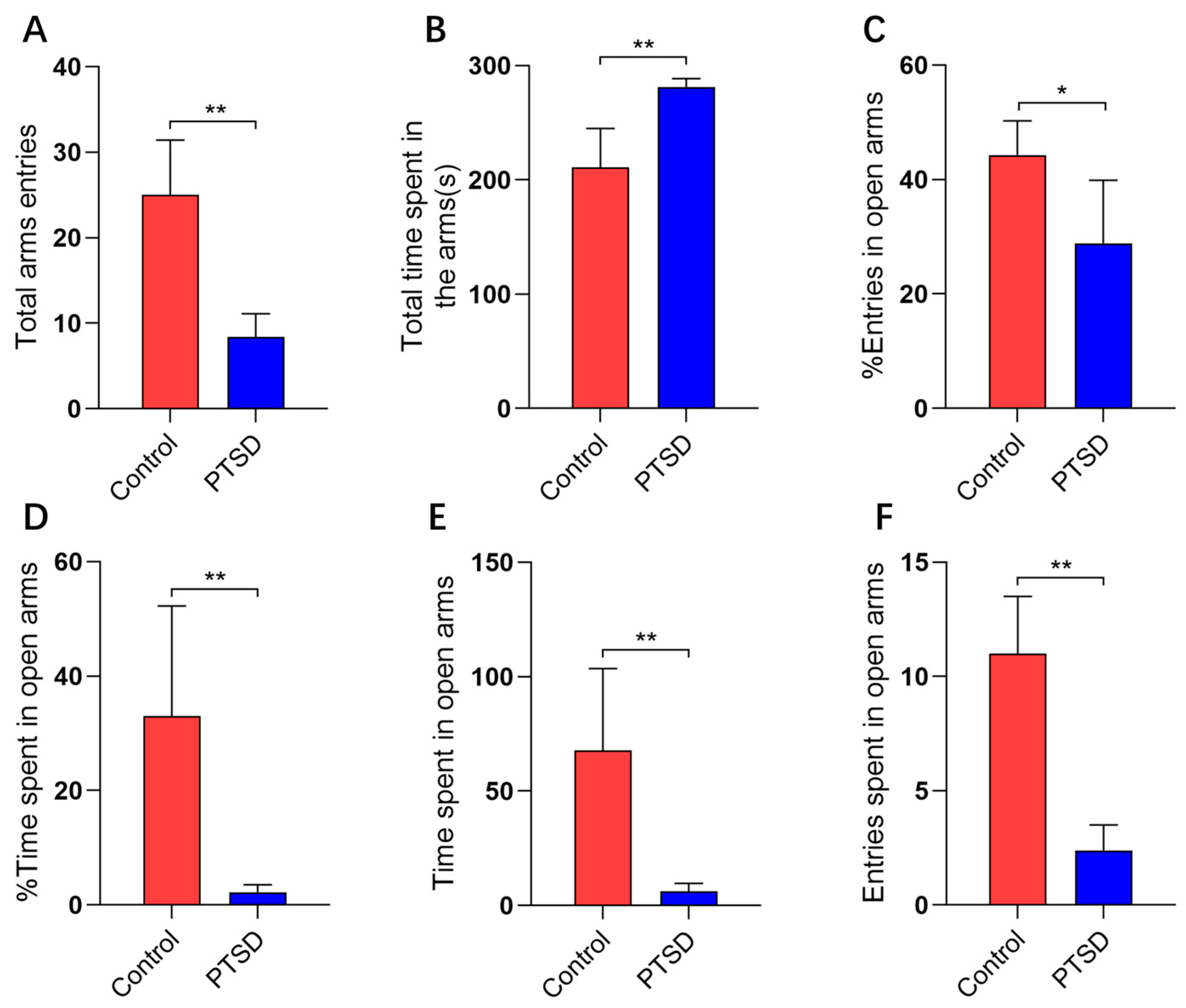
2.4. Elevated Plus Maze Test
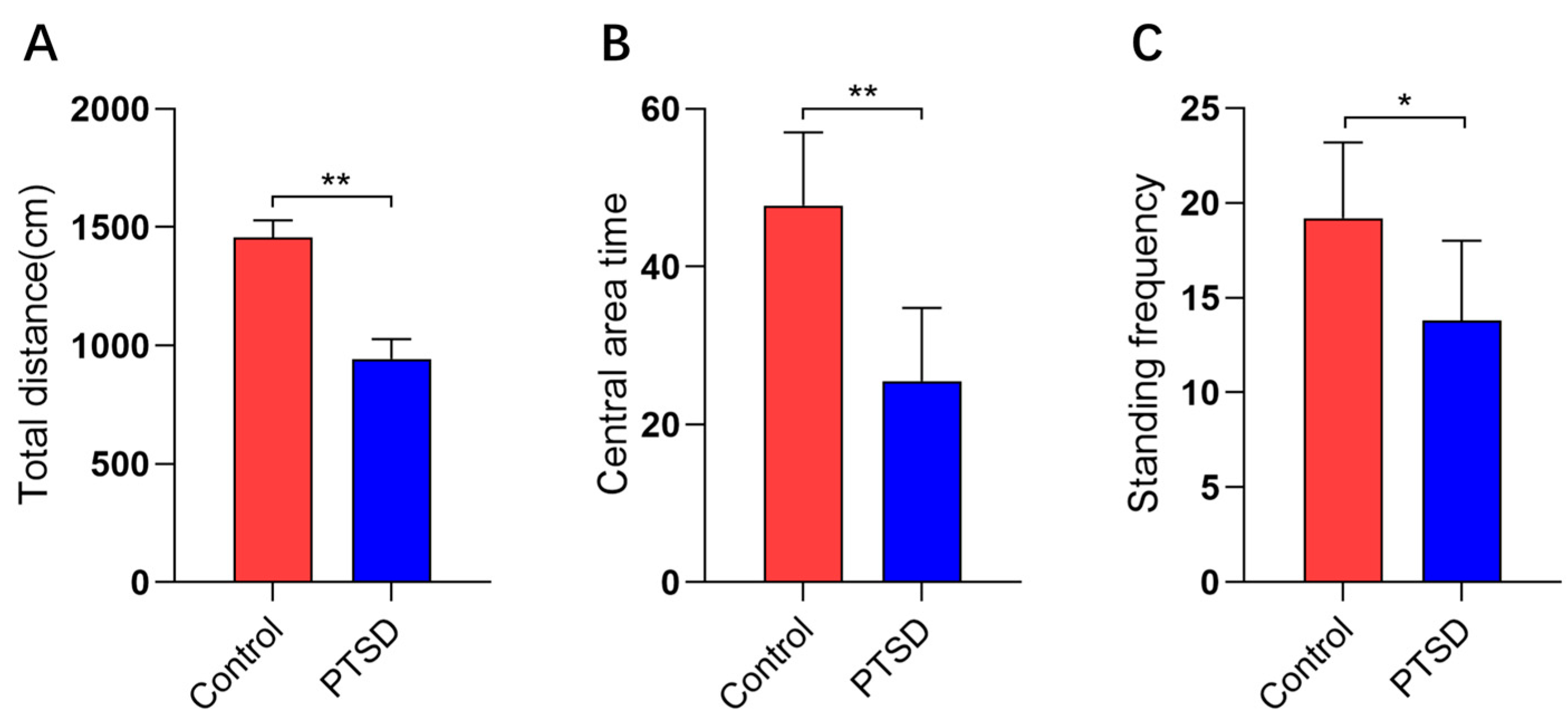
2.5. Open Field Test
2.6. Sample Collection and Preparation
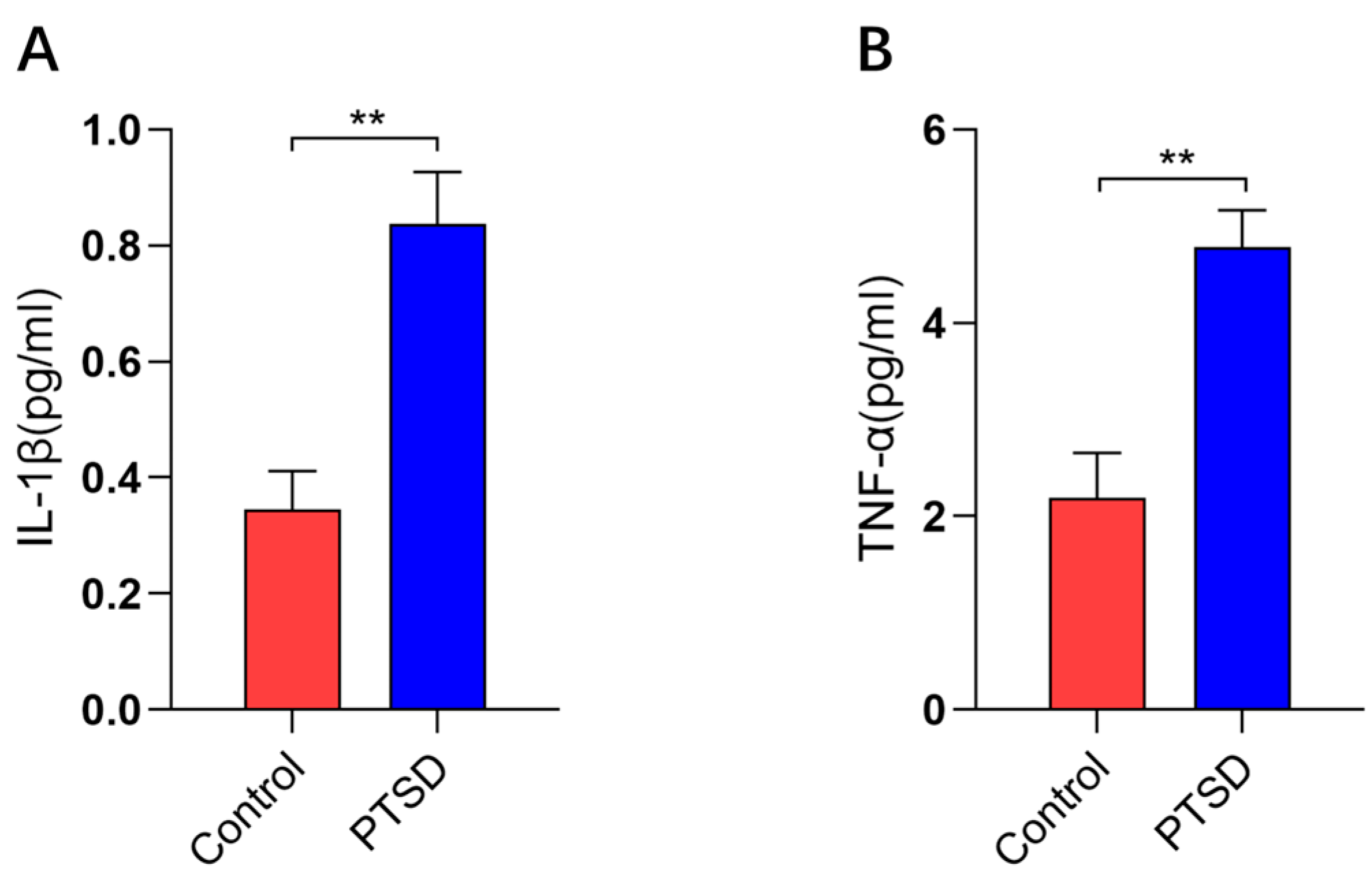
2.7. Cytokine Analysis
2.8. Metabolomics Analysis
2.9. Metabolomic Data Processing and Analysis
2.10. 16S rRNA Sequencing of Gut Microbiota
2.11. Bioinformatic and Statistical Analysis of 16S rRNA Data
2.12. Statistical Analysis
3. Results
3.1. Elevated Plus Maze Test
3.2. Open Field Test
3.3. Cytokines
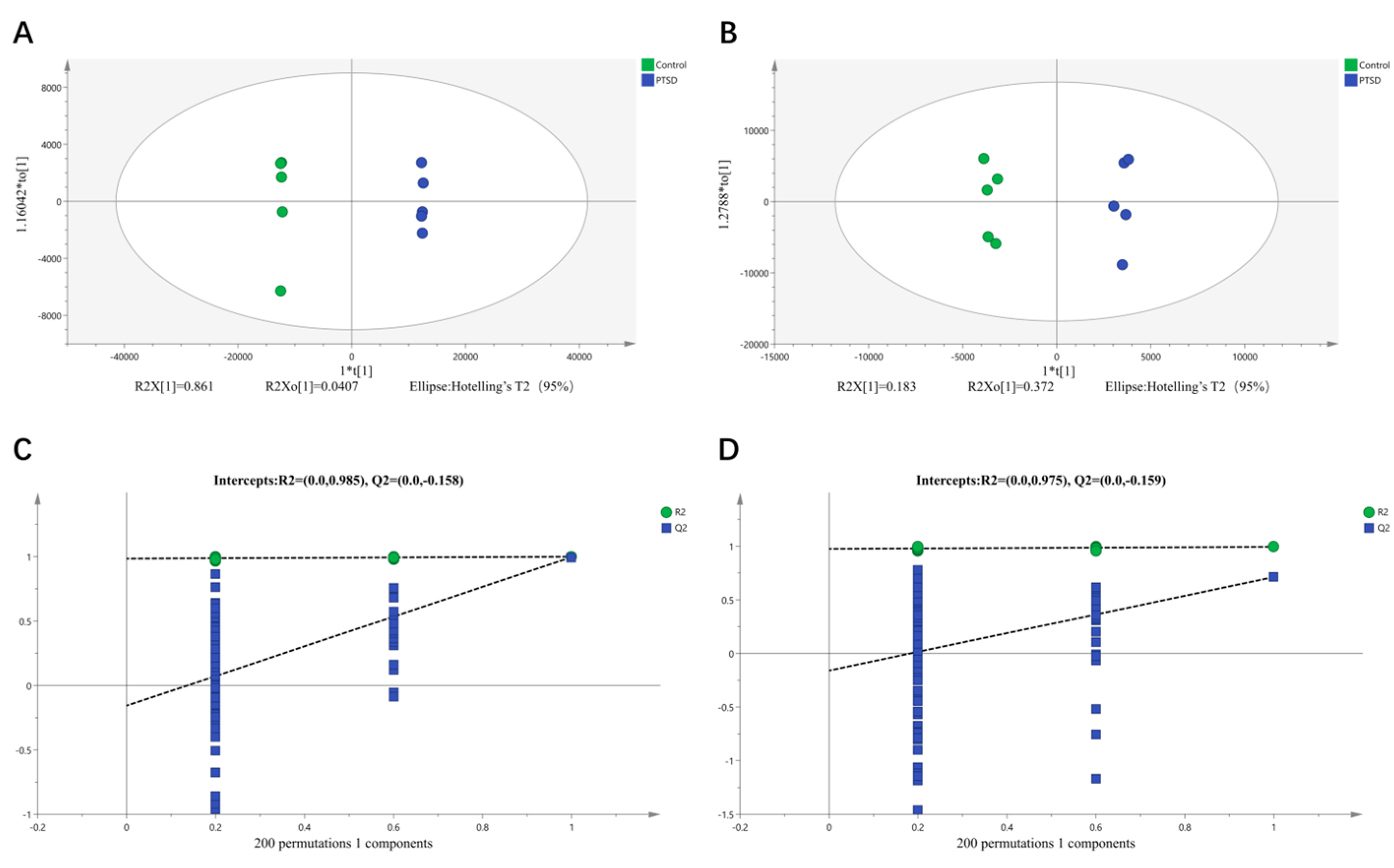
3.4. Metabolomics Analysis
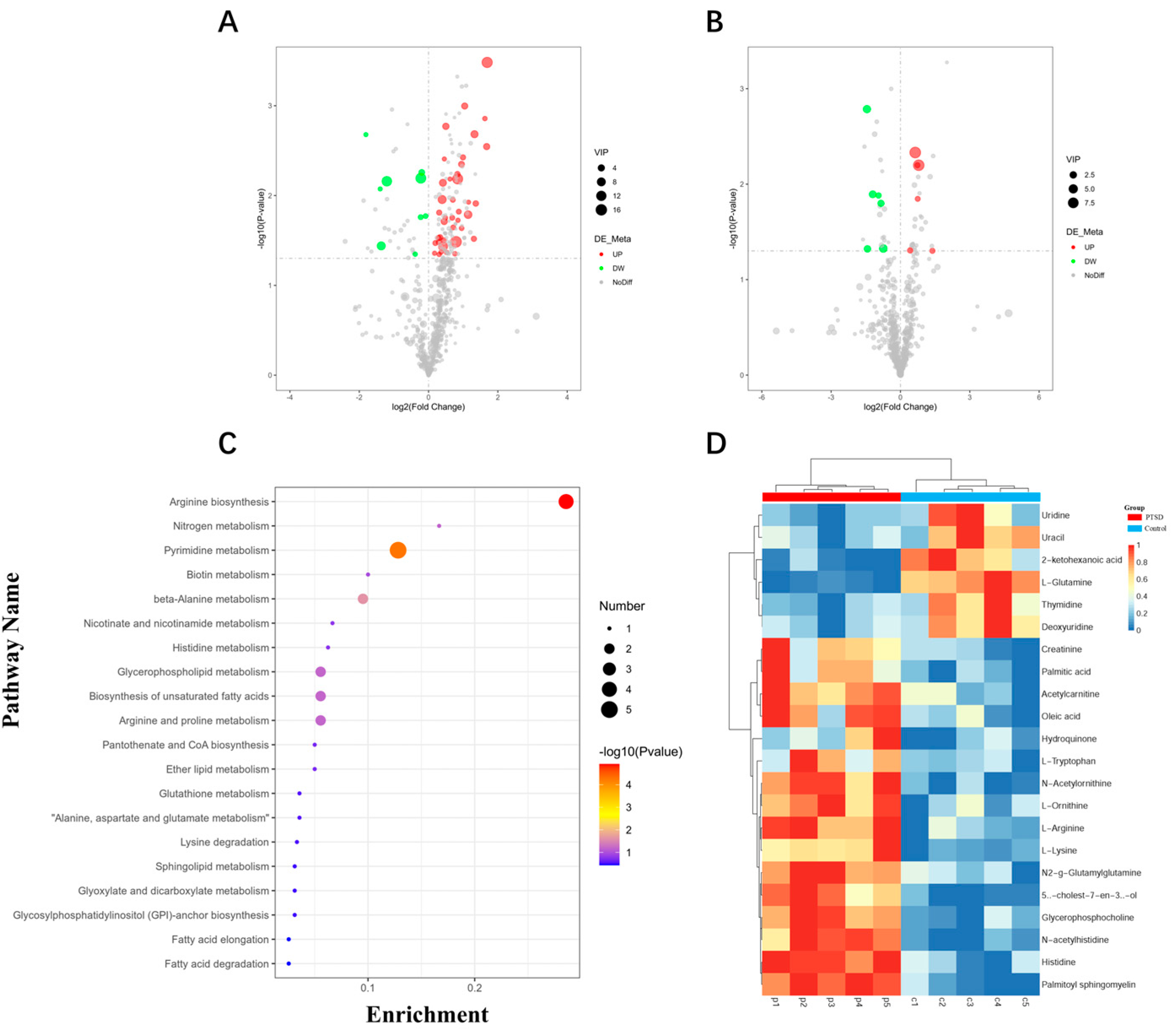
3.5. Differential Metabolites
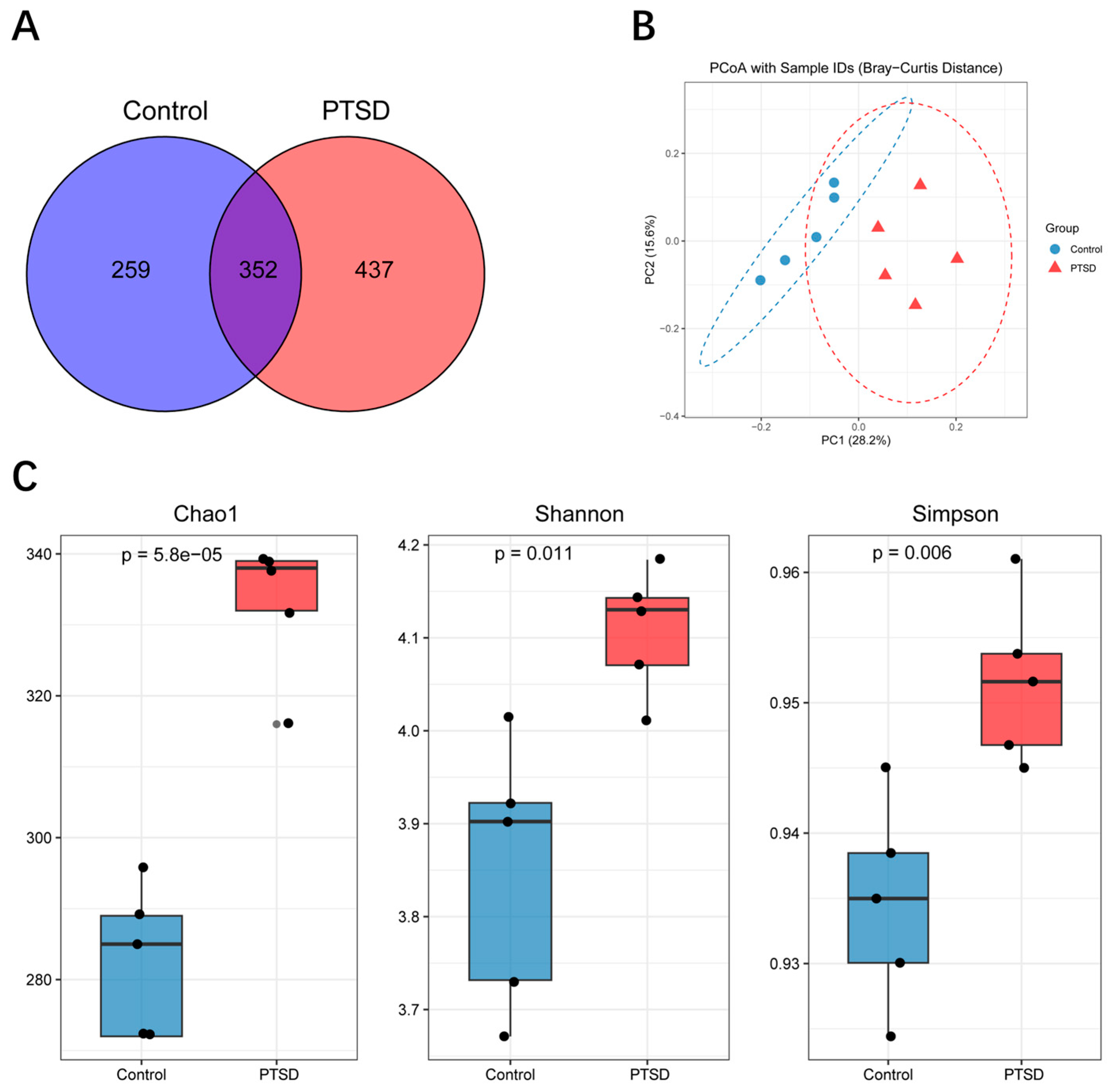
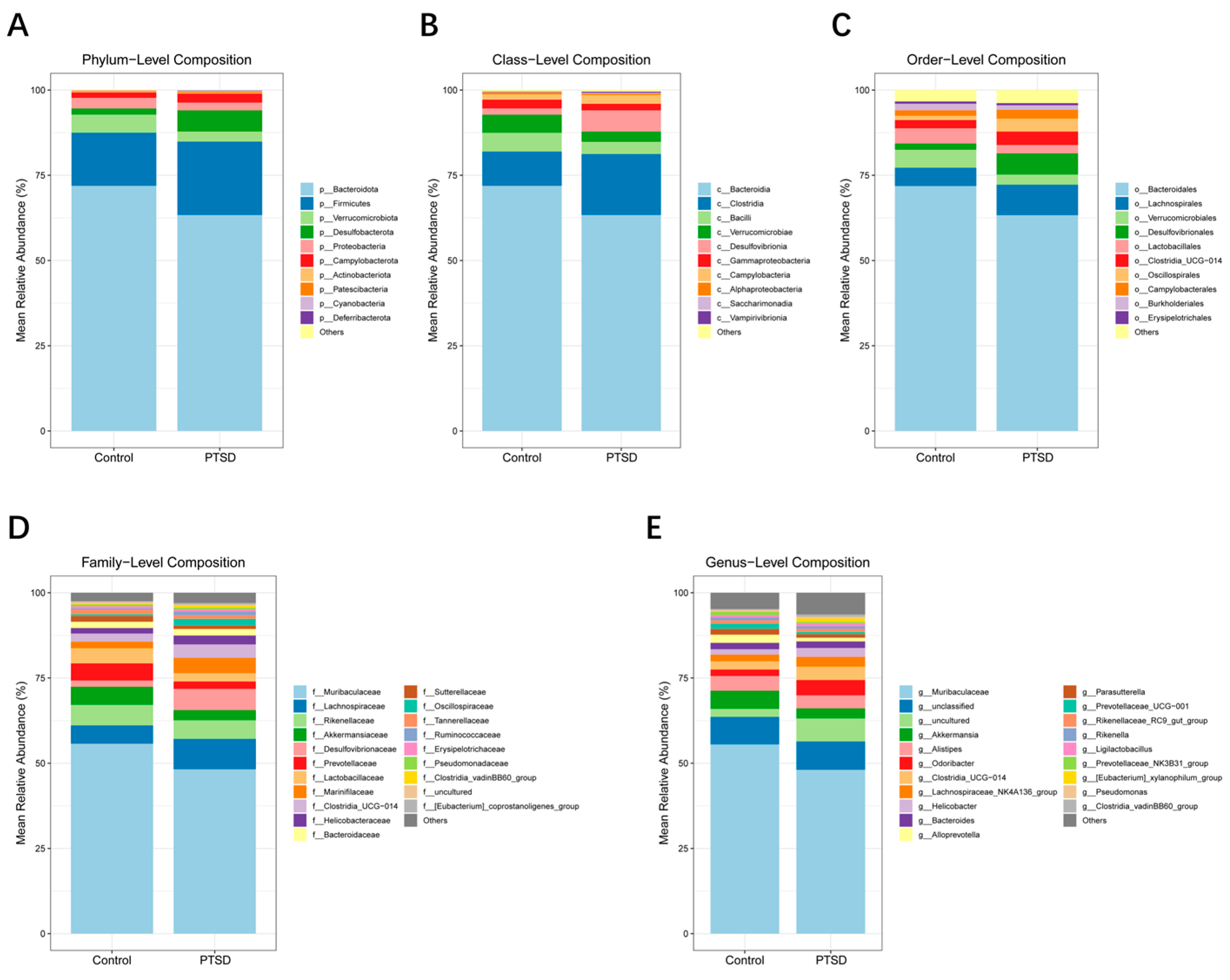
3.6. Gut Microbial Community Diversity Analysis
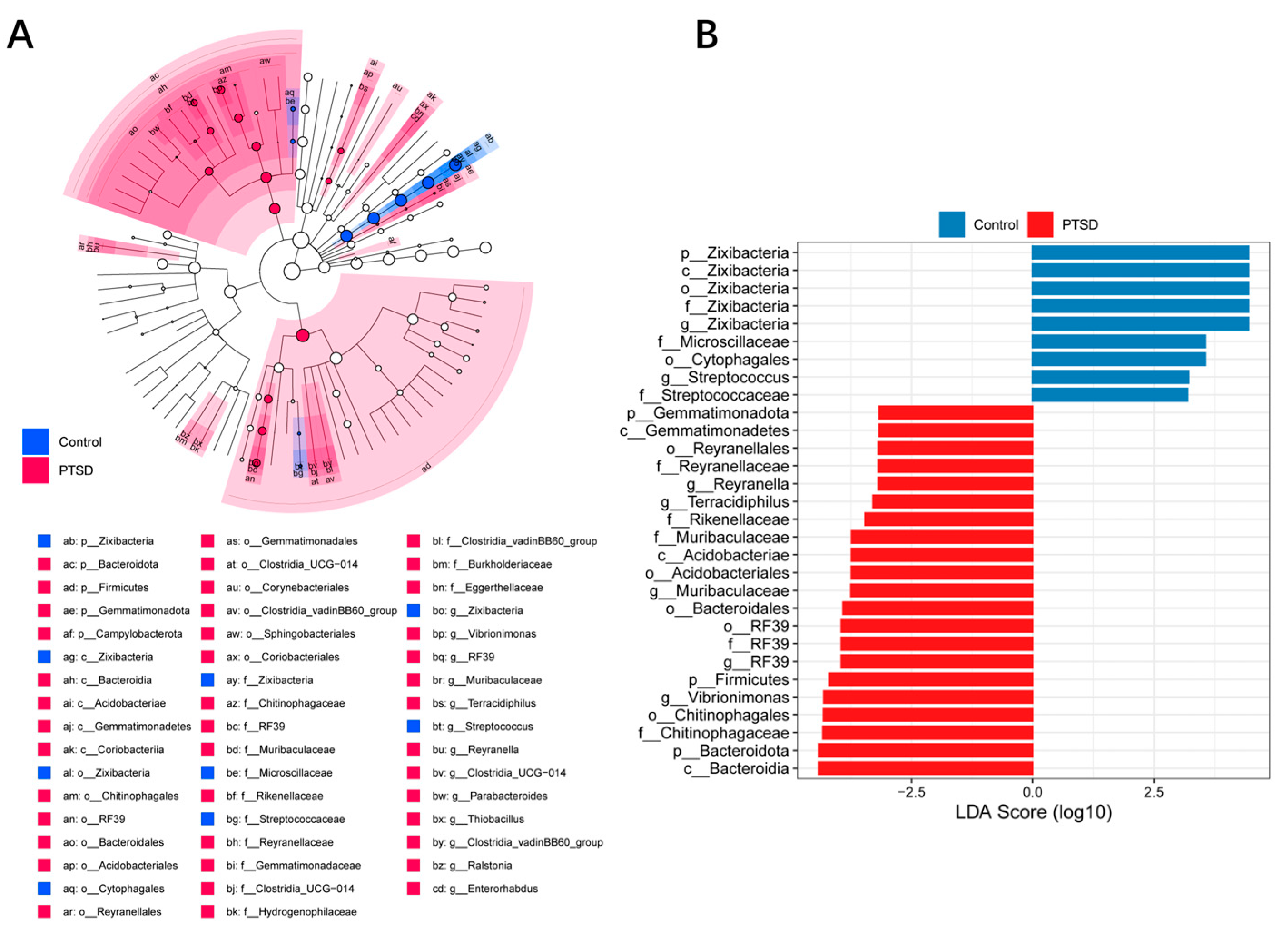
3.7. LEfSe Analysis
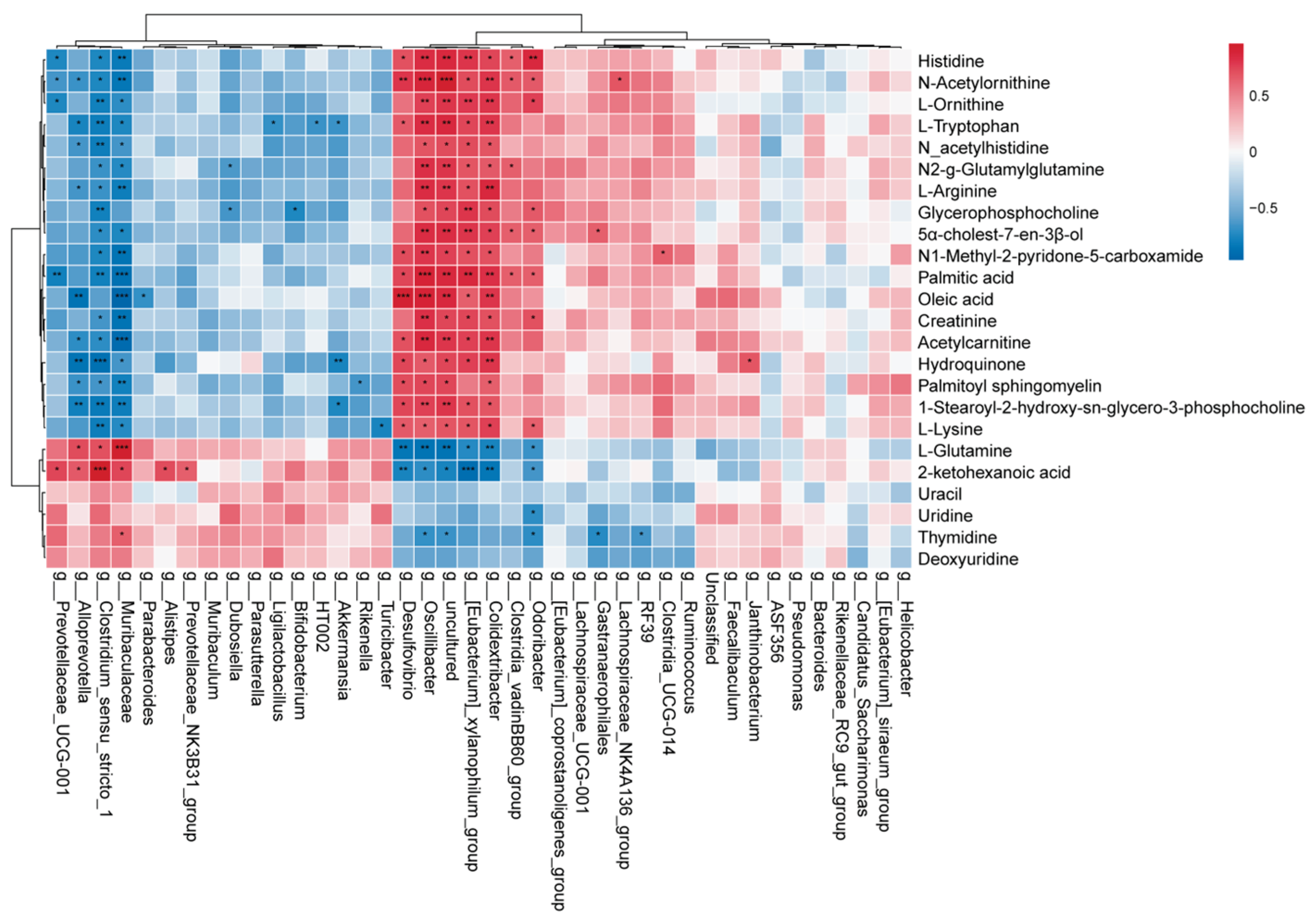
3.8. Correlation of Gut Microbiota with Plasma Metabolites
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| PTSD | Post-traumatic stress disorder |
| SPS&S | Single prolonged stress and electric foot shock |
| UHPLC-MS/MS | Ultra-high-performance liquid chromatography–tandem mass spectrometry |
| GABR | global arginine bioavailability ratio |
| MGBA | microbiota–gut–brain axis |
| FMT | fecal microbiota transplantation |
| EPMT | Elevated plus maze test |
| OFT | Open field test |
| PCA | Principal component analysis |
| PLS-DA | Partial least squares discriminant analysis |
| OPLS-DA | Orthogonal partial least squares discriminant analysis |
| HMDB | Human metabolome database |
| KEGG | Kyoto encyclopedia of genes and genomes |
| OUT | Operational taxonomic units |
| PCoA | Principal coordinates analysis |
| LEfSe | Linear discriminant analysis effect size |
| ANOVA | Analysis of variance |
| IL-1β | Interleukin-1β |
| TNF-α | Tumor necrosis factor-α |
| NOS | Nitric oxide synthase |
| NO | Nitric oxide |
| BD | Bipolar disorder |
| SFAs | Saturated fatty acids |
| PA | Palmitic acid |
| LPS | Lipopolysaccharides |
| NLRP3 | NOD-like receptor containing a pyrin domain 3 |
| 5-HIAA | 5-hydroxyindoleacetic acid |
| OA | Oleic acid |
| AMPK | AMP-activated protein kinase |
References
- Wilker, S.; Schneider, A.; Conrad, D.; Pfeiffer, A.; Boeck, C.; Lingenfelder, B.; Freytag, V.; Vukojevic, V.; Vogler, C.; Milnik, A.; et al. Genetic variation is associated with PTSD risk and aversive memory: Evidence from two trauma-Exposed African samples and one healthy European sample. Transl Psychiatry 2018, 8, 251. [Google Scholar] [CrossRef]
- Wingo, T.S.; Gerasimov, E.S.; Liu, Y.; Duong, D.M.; Vattathil, S.M.; Lori, A.; Gockley, J.; Breen, M.S.; Maihofer, A.X.; Nievergelt, C.M.; et al. Integrating human brain proteomes with genome-wide association data implicates novel proteins in post-traumatic stress disorder. Mol Psychiatry 2022, 27, 3075–3084. [Google Scholar] [CrossRef]
- Franzosa, E.A.; Sirota-Madi, A.; Avila-Pacheco, J.; Fornelos, N.; Haiser, H.J.; Reinker, S.; Vatanen, T.; Hall, A.B.; Mallick, H.; McIver, L.J.; et al. Gut microbiome structure and metabolic activity in inflammatory bowel disease. Nat Microbiol 2018, 4, 293–305. [Google Scholar] [CrossRef]
- Hendrix, J.; Ranginani, D.; Montero, A.M.; Lockett, C.; Xu, H.; James-Stevenson, T.; Shin, A. Early adverse life events and post-traumatic stress disorder in patients with constipation and suspected disordered defecation. Neurogastroenterol Motil 2021, 34, e14195. [Google Scholar] [CrossRef] [PubMed]
- Malhotra, D.; Boyle, S.H.; Gifford, E.J.; Sullivan, B.A.; Nguyen Wenker, T.H.; Abs, N.-D.; Ahmed, S.T.; Upchurch, J.; Vahey, J.; Stafford, C.; et al. Self-reported gastrointestinal disorders among veterans with gulf war illness with and without posttraumatic stress disorder. Neurogastroenterol Motil 2023, 35, e14548. [Google Scholar] [CrossRef] [PubMed]
- Mellon, S.H.; Gautam, A.; Hammamieh, R.; Jett, M.; Wolkowitz, O.M. Metabolism, Metabolomics, and Inflammation in Posttraumatic Stress Disorder. Biol Psychiatry 2018, 83, 866–875. [Google Scholar] [CrossRef] [PubMed]
- Wolf, E.J.; Bovin, M.J.; Green, J.D.; Mitchell, K.S.; Stoop, T.B.; Barretto, K.M.; Jackson, C.E.; Lee, L.O.; Fang, S.C.; Trachtenberg, F.; et al. Longitudinal associations between post-traumatic stress disorder and metabolic syndrome severity. Psychol Med 2016, 46, 2215–2226. [Google Scholar] [CrossRef]
- Yunitri, N.; Chu, H.; Kang, X.L.; Jen, H.-J.; Pien, L.-C.; Tsai, H.-T.; Kamil, A.R.; Chou, K.-R. Global prevalence and associated risk factors of posttraumatic stress disorder during COVID-19 pandemic: A meta-analysis. Int J Nurs Stud 2021, 126, 104136. [Google Scholar] [CrossRef]
- Mastrangelo, A.; Panadero, M.I.; Pérez, L.M.; Gálvez, B.G.; García, A.; Barbas, C.; Rupérez, F.J. New insight on obesity and adipose-derived stem cells using comprehensive metabolomics. Biochem J 2016, 473, 2187–2203. [Google Scholar] [CrossRef]
- Quinones, M.P.; Kaddurah-Daouk, R. Metabolomics tools for identifying biomarkers for neuropsychiatric diseases. Neurobiol Dis 2009, 35, 165–176. [Google Scholar] [CrossRef]
- Pontes, J.G.M.; Brasil, A.J.M.; Cruz, G.C.F.; de Souza, R.N.; Tasic, L. NMR-based metabolomics strategies: plants, animals and humans. ANALYTICAL METHODS 2017, 9, 1078–1096. [Google Scholar] [CrossRef]
- Nedic Erjavec, G.; Konjevod, M.; Nikolac Perkovic, M.; Svob Strac, D.; Tudor, L.; Barbas, C.; Grune, T.; Zarkovic, N.; Pivac, N. Short overview on metabolomic approach and redox changes in psychiatric disorders. Redox Biol 2017, 14, 178–186. [Google Scholar] [CrossRef]
- Johnson, C.H.; Ivanisevic, J.; Siuzdak, G. Metabolomics: beyond biomarkers and towards mechanisms. Nat Rev Mol Cell Biol 2016, 17, 451–459. [Google Scholar] [CrossRef] [PubMed]
- Karabatsiakis, A.; Hamuni, G.; Wilker, S.; Kolassa, S.; Renu, D.; Kadereit, S.; Schauer, M.; Hennessy, T.; Kolassa, I.-T. Metabolite profiling in posttraumatic stress disorder. J Mol Psychiatry 2015, 3, 2. [Google Scholar] [CrossRef]
- Shin, S.-Y.; Fauman, E.B.; Petersen, A.-K.; Krumsiek, J.; Santos, R.; Huang, J.; Arnold, M.; Erte, I.; Forgetta, V.; Yang, T.-P.; et al. An atlas of genetic influences on human blood metabolites. Nat Genet 2014, 46, 543–550. [Google Scholar] [CrossRef] [PubMed]
- Wei, Y.; Huang, L.; Sui, J.; Liu, C.; Qi, M. Human blood metabolites and risk of post-traumatic stress disorder: A Mendelian randomization study. J Affect Disord 2024, 372, 227–233. [Google Scholar] [CrossRef]
- Bersani, F.S.; Wolkowitz, O.M.; Lindqvist, D.; Yehuda, R.; Flory, J.; Bierer, L.M.; Makotine, I.; Abu-Amara, D.; Coy, M.; Reus, V.I.; et al. Global arginine bioavailability, a marker of nitric oxide synthetic capacity, is decreased in PTSD and correlated with symptom severity and markers of inflammation. Brain Behav Immun 2015, 52, 153–160. [Google Scholar] [CrossRef]
- Inoue, T.; Ogawa, S.; Narita, Z.; Sekiguchi, M.; Asari, Y.; Kataoka, Y.; Hattori, J.; Hori, H.; Kim, Y.; Inada, K. A Longitudinal Study of the Association of Blood Unsaturated Fatty Acids With Posttraumatic Stress Disorder (PTSD). Neuropsychopharmacol Rep 2025, 45, e12522. [Google Scholar] [CrossRef]
- Chen, C.; Liao, J.; Xia, Y.; Liu, X.; Jones, R.; Haran, J.; McCormick, B.; Sampson, T.R.; Alam, A.; Ye, K. Gut microbiota regulate Alzheimer's disease pathologies and cognitive disorders via PUFA-associated neuroinflammation. Gut 2022, 71, 2233–2252. [Google Scholar] [CrossRef]
- Tan, A.H.; Lim, S.Y.; Lang, A.E. The microbiome-gut-brain axis in Parkinson disease - from basic research to the clinic. Nat Rev Neurol 2022, 18, 476–495. [Google Scholar] [CrossRef]
- Zhao, Q.; Baranova, A.; Cao, H.; Zhang, F. Gut microbiome and major depressive disorder: insights from two-sample Mendelian randomization. BMC Psychiatry 2024, 24, 493. [Google Scholar] [CrossRef] [PubMed]
- Foster, J.A.; Baker, G.B.; Dursun, S.M. The Relationship Between the Gut Microbiome-Immune System-Brain Axis and Major Depressive Disorder. Front Neurol 2021, 12, 721126. [Google Scholar] [CrossRef] [PubMed]
- Ioannou, M.; Borkent, J.; Andreu-Sánchez, S.; Wu, J.; Fu, J.; Sommer, I.E.C.; Haarman, B.C.M. Reproducible gut microbial signatures in bipolar and schizophrenia spectrum disorders: A metagenome-wide study. Brain Behav Immun 2024, 121, 165–175. [Google Scholar] [CrossRef] [PubMed]
- He, Q.; Wang, W.; Xu, D.; Xiong, Y.; Tao, C.; You, C.; Ma, L.; Ma, J. Potential causal association between gut microbiome and posttraumatic stress disorder. Transl Psychiatry 2024, 14, 67. [Google Scholar] [CrossRef]
- Gorecki, A.M.; Preskey, L.; Bakeberg, M.C.; Kenna, J.E.; Gildenhuys, C.; MacDougall, G.; Dunlop, S.A.; Mastaglia, F.L.; Akkari, P.A.; Koengten, F.; et al. Altered Gut Microbiome in Parkinson's Disease and the Influence of Lipopolysaccharide in a Human α-Synuclein Over-Expressing Mouse Model. Front Neurosci 2019, 13, 839. [Google Scholar] [CrossRef]
- Qi, Z.; Lyu, M.; Yang, L.; Yuan, H.; Cao, Y.; Zhai, L.; Dang, W.; Liu, J.; Yang, F.; Li, Y. A Novel and Reliable Rat Model of Autism. Front Psychiatry 2021, 12, 549810. [Google Scholar] [CrossRef]
- Ait-Belgnaoui, A.; Payard, I.; Rolland, C.; Harkat, C.; Braniste, V.; Théodorou, V.; Tompkins, T.A. Bifidobacterium longum and Lactobacillus helveticus Synergistically Suppress Stress-related Visceral Hypersensitivity Through Hypothalamic-Pituitary-Adrenal Axis Modulation. J Neurogastroenterol Motil 2018, 24, 138–146. [Google Scholar] [CrossRef]
- Schretter, C.E.; Vielmetter, J.; Bartos, I.; Marka, Z.; Marka, S.; Argade, S.; Mazmanian, S.K. A gut microbial factor modulates locomotor behaviour in Drosophila. Nature 2018, 563, 402–406. [Google Scholar] [CrossRef] [PubMed]
- Cowan, C.S.M.; Hoban, A.E.; Ventura-Silva, A.P.; Dinan, T.G.; Clarke, G.; Cryan, J.F. Gutsy Moves: The Amygdala as a Critical Node in Microbiota to Brain Signaling. Bioessays 2017, 40. [Google Scholar] [CrossRef]
- Ruth, F.; Karen, Y.; Sondra, T.; Oshrit, S.; Orna, Z.-S.; Lelyan, M.; Yoram, L.; Omry, K. Microbiome Mediates Development of PTSD and Resilience. Research Square 2022, Preprint (version 1). [Google Scholar] [CrossRef]
- Zhou, Q.; Sun, T.; Wu, F.; Li, F.; Liu, Y.; Li, W.; Dai, N.; Tan, L.; Li, T.; Song, Y. Correlation of gut microbiota and neurotransmitters in a rat model of post-traumatic stress disorder. Journal of Traditional Chinese Medical Sciences 2020, 7, 375–385. [Google Scholar] [CrossRef]
- Xi, K.; Xiao, H.; Huang, X.; Yuan, Z.; Liu, M.; Mao, H.; Liu, H.; Ma, G.; Cheng, Z.; Xie, Y.; et al. Reversal of hyperactive higher-order thalamus attenuates defensiveness in a mouse model of PTSD. Sci Adv 2023, 9, eade5987. [Google Scholar] [CrossRef]
- Lindqvist, D.; Wolkowitz, O.M.; Mellon, S.; Yehuda, R.; Flory, J.D.; Henn-Haase, C.; Bierer, L.M.; Abu-Amara, D.; Coy, M.; Neylan, T.C.; et al. Proinflammatory milieu in combat-related PTSD is independent of depression and early life stress. Brain Behav Immun 2014, 42, 81–88. [Google Scholar] [CrossRef]
- Guo, M.; Liu, T.; Guo, J.-C.; Jiang, X.-L.; Chen, F.; Gao, Y.-S. Study on serum cytokine levels in posttraumatic stress disorder patients. Asian Pac J Trop Med 2012, 5, 323–325. [Google Scholar] [CrossRef]
- Spitzer, C.; Barnow, S.; Völzke, H.; Wallaschofski, H.; John, U.; Freyberger, H.J.; Löwe, B.; Grabe, H.J. Association of posttraumatic stress disorder with low-grade elevation of C-reactive protein: evidence from the general population. J Psychiatr Res 2009, 44, 15–21. [Google Scholar] [CrossRef]
- Bronte, V.; Zanovello, P. Regulation of immune responses by L-arginine metabolism. Nat Rev Immunol 2005, 5, 641–654. [Google Scholar] [CrossRef]
- Morris, S.M. Arginine metabolism: boundaries of our knowledge. J Nutr 2007, 137, 1602S–1609S. [Google Scholar] [CrossRef]
- Yoshimi, N.; Futamura, T.; Kakumoto, K.; Salehi, A.M.; Sellgren, C.M.; Holmén-Larsson, J.; Jakobsson, J.; Pålsson, E.; Landén, M.; Hashimoto, K. Blood metabolomics analysis identifies abnormalities in the citric acid cycle, urea cycle, and amino acid metabolism in bipolar disorder. BBA Clin 2016, 5, 151–158. [Google Scholar] [CrossRef]
- Yoshimi, N.; Futamura, T.; Bergen, S.E.; Iwayama, Y.; Ishima, T.; Sellgren, C.; Ekman, C.J.; Jakobsson, J.; Pålsson, E.; Kakumoto, K.; et al. Cerebrospinal fluid metabolomics identifies a key role of isocitrate dehydrogenase in bipolar disorder: evidence in support of mitochondrial dysfunction hypothesis. Mol Psychiatry 2016, 21, 1504–1510. [Google Scholar] [CrossRef]
- Ozden, A.; Angelos, H.; Feyza, A.; Elizabeth, W.; John, P. Altered plasma levels of arginine metabolites in depression. J Psychiatr Res 2019, 120, 21–28. [Google Scholar] [CrossRef]
- Watts, D.; Pfaffenseller, B.; Wollenhaupt-Aguiar, B.; Paul Géa, L.; Cardoso, T.D.A.; Kapczinski, F. Agmatine as a potential therapeutic intervention in bipolar depression: the preclinical landscape. Expert Opin Ther Targets 2019, 23, 327–339. [Google Scholar] [CrossRef]
- Ralston, J.C.; Metherel, A.H.; Stark, K.D.; Mutch, D.M. SCD1 mediates the influence of exogenous saturated and monounsaturated fatty acids in adipocytes: Effects on cellular stress, inflammatory markers and fatty acid elongation. J Nutr Biochem 2015, 27, 241–248. [Google Scholar] [CrossRef]
- Pifferi, F.; Laurent, B.; Plourde, M. Lipid Transport and Metabolism at the Blood-Brain Interface: Implications in Health and Disease. Front Physiol 2021, 12, 645646. [Google Scholar] [CrossRef]
- Ogawa, S.; Hori, H.; Niwa, M.; Itoh, M.; Lin, M.; Yoshida, F.; Ino, K.; Kawanishi, H.; Narita, M.; Nakano, W.; et al. Serum lipid and plasma fatty acid profiles in PTSD patients and healthy individuals: Associations with symptoms, cognitive function, and inflammatory markers. Prog Neuropsychopharmacol Biol Psychiatry 2025, 138, 111298. [Google Scholar] [CrossRef]
- Lu, Z.; Liu, S.; Lopes-Virella, M.F.; Wang, Z. LPS and palmitic acid Co-upregulate microglia activation and neuroinflammatory response. Compr Psychoneuroendocrinol 2021, 6, 100048. [Google Scholar] [CrossRef]
- Chaurasia, B.; Summers, S.A. Ceramides - Lipotoxic Inducers of Metabolic Disorders. Trends Endocrinol Metab 2015, 26, 538–550. [Google Scholar] [CrossRef]
- Moon, M.L.; Joesting, J.J.; Lawson, M.A.; Chiu, G.S.; Blevins, N.A.; Kwakwa, K.A.; Freund, G.G. The saturated fatty acid, palmitic acid, induces anxiety-like behavior in mice. Metabolism 2014, 63, 1131–1140. [Google Scholar] [CrossRef]
- Neufeld-Cohen, A.; Evans, A.K.; Getselter, D.; Spyroglou, A.; Hill, A.; Gil, S.; Tsoory, M.; Beuschlein, F.; Lowry, C.A.; Vale, W.; et al. Urocortin-1 and -2 double-deficient mice show robust anxiolytic phenotype and modified serotonergic activity in anxiety circuits. Mol Psychiatry 2009, 15. [Google Scholar] [CrossRef]
- Finucane, O.M.; Lyons, C.L.; Murphy, A.M.; Reynolds, C.M.; Klinger, R.; Healy, N.P.; Cooke, A.A.; Coll, R.C.; McAllan, L.; Nilaweera, K.N.; et al. Monounsaturated fatty acid-enriched high-fat diets impede adipose NLRP3 inflammasome-mediated IL-1β secretion and insulin resistance despite obesity. Diabetes 2015, 64, 2116–2128. [Google Scholar] [CrossRef]
- Holland, W.L.; Miller, R.A.; Wang, Z.V.; Sun, K.; Barth, B.M.; Bui, H.H.; Davis, K.E.; Bikman, B.T.; Halberg, N.; Rutkowski, J.M.; et al. Receptor-mediated activation of ceramidase activity initiates the pleiotropic actions of adiponectin. Nat Med 2010, 17, 55–63. [Google Scholar] [CrossRef]
- Mironova, G.D.; Khrenov, M.O.; Talanov, E.Y.; Glushkova, O.V.; Parfenyuk, S.B.; Novoselova, T.V.; Lunin, S.M.; Belosludtseva, N.V.; Novoselova, E.G.; Lemasters, J.J. The role of mitochondrial KATP channel in anti-inflammatory effects of uridine in endotoxemic mice. Arch Biochem Biophys 2018, 654, 70–76. [Google Scholar] [CrossRef]
- Zhou, D.; Long, C.; Shao, Y.; Li, F.; Sun, W.; Zheng, Z.; Wang, X.; Huang, Y.; Pan, F.; Chen, G.; et al. Integrated Metabolomics and Proteomics Analysis of Urine in a Mouse Model of Posttraumatic Stress Disorder. Front Neurosci 2022, 16, 828382. [Google Scholar] [CrossRef]
- Traut, T.W. Physiological concentrations of purines and pyrimidines. Mol Cell Biochem 1994, 140. [Google Scholar] [CrossRef]
- Connolly, G.P.; Duley, J.A. Uridine and its nucleotides: biological actions, therapeutic potentials. Trends Pharmacol Sci 1999, 20, 218–225. [Google Scholar] [CrossRef]
- Ley, R.E.; Peterson, D.A.; Gordon, J.I. Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell 2006, 124, 837–848. [Google Scholar] [CrossRef]
- Smith, B.J.; Miller, R.A.; Ericsson, A.C.; Harrison, D.C.; Strong, R.; Schmidt, T.M. Changes in the gut microbiome and fermentation products concurrent with enhanced longevity in acarbose-treated mice. BMC Microbiol 2019, 19, 130. [Google Scholar] [CrossRef]
- Ormerod, K.L.; Wood, D.L.A.; Lachner, N.; Gellatly, S.L.; Daly, J.N.; Parsons, J.D.; Dal'Molin, C.G.O.; Palfreyman, R.W.; Nielsen, L.K.; Cooper, M.A.; et al. Genomic characterization of the uncultured Bacteroidales family S24-7 inhabiting the guts of homeothermic animals. Microbiome 2016, 4, 36. [Google Scholar] [CrossRef]
- Zhang, F.; Zhou, Y.; Chen, H.; Jiang, H.; Zhou, F.; Lv, B.; Xu, M. Curcumin Alleviates DSS-Induced Anxiety-Like Behaviors via the Microbial-Brain-Gut Axis. Oxid Med Cell Longev 2022, 2022, 6244757. [Google Scholar] [CrossRef]
- Cani, P.D.; Depommier, C.; Derrien, M.; Everard, A.; de Vos, W.M. Akkermansia muciniphila: paradigm for next-generation beneficial microorganisms. Nat Rev Gastroenterol Hepatol 2022, 19, 625–637. [Google Scholar] [CrossRef]
- Cani, P.D.; de Vos, W.M. Next-Generation Beneficial Microbes: The Case of Akkermansia muciniphila. Front Microbiol 2017, 8, 1765. [Google Scholar] [CrossRef]
- Xia, T.; Duan, W.; Zhang, Z.; Li, S.; Zhao, Y.; Geng, B.; Zheng, Y.; Yu, J.; Wang, M. Polyphenol-rich vinegar extract regulates intestinal microbiota and immunity and prevents alcohol-induced inflammation in mice. Food Res Int 2021, 140, 110064. [Google Scholar] [CrossRef] [PubMed]
- Shen, C.-L.; Wang, R.; Ji, G.; Elmassry, M.M.; Zabet-Moghaddam, M.; Vellers, H.; Hamood, A.N.; Gong, X.; Mirzaei, P.; Sang, S.; et al. Dietary supplementation of gingerols- and shogaols-enriched ginger root extract attenuate pain-associated behaviors while modulating gut microbiota and metabolites in rats with spinal nerve ligation. J Nutr Biochem 2021, 100, 108904. [Google Scholar] [CrossRef]
- Wu, H.-J.; Wu, E. The role of gut microbiota in immune homeostasis and autoimmunity. Gut Microbes 2012, 3. [Google Scholar] [CrossRef] [PubMed]
- Doney, E.; Cadoret, A.; Dion-Albert, L.; Lebel, M.; Menard, C. Inflammation-driven brain and gut barrier dysfunction in stress and mood disorders. Eur J Neurosci 2021, 55, 2851–2894. [Google Scholar] [CrossRef] [PubMed]
| No | Metabolites | FC | P-value | VIP | Trend |
|---|---|---|---|---|---|
| 1 | 1-Stearoyl-2-hydroxy-sn-glycero-3-phosphocholine | 5.18128 | 1.31926E-08 | 7.04406 | ↑ |
| 2 | L-Glutamine | 0.14185 | 6.57121E-07 | 2.32125 | ↓ |
| 3 | Palmitoyl sphingomyelin | 1.86521 | 1.88291E-06 | 5.016 | ↑ |
| 4 | N-acetylhistidine | 2.45764 | 1.29928E-05 | 1.22055 | ↑ |
| 5 | Histidine | 4.01037 | 1.35614E-05 | 2.5638 | ↑ |
| 6 | N-Acetylornithine | 2.49663 | 2.26593E-05 | 1.90844 | ↑ |
| 7 | Glycerophosphocholine | 2.09156 | 2.48749E-05 | 7.36942 | ↑ |
| 8 | 5α-cholest-7-en-3β-ol | 4.03121 | 6.21691E-05 | 1.71005 | ↑ |
| 9 | N2-g-Glutamylglutamine | 3.52854 | 7.68233E-05 | 1.83453 | ↑ |
| 10 | L-Lysine | 2.14282 | 9.18155E-05 | 2.14269 | ↑ |
| 11 | L-Arginine | 1.82076 | 0.00011 | 2.41993 | ↑ |
| 12 | L-Ornithine | 1.93682 | 0.00041 | 1.0943 | ↑ |
| 13 | N1-Methyl-2-pyridone-5-carboxamide | 2.29938 | 0.00053 | 1.45207 | ↑ |
| 14 | Acetylcarnitine | 2.04835 | 0.00078 | 2.17828 | ↑ |
| 15 | 2-ketohexanoic acid | 0.36609 | 0.00164 | 3.56613 | ↓ |
| 16 | Palmitic acid | 1.55761 | 0.00466 | 9.30956 | ↑ |
| 17 | Creatinine | 1.56381 | 0.00618 | 1.09553 | ↑ |
| 18 | Oleic acid | 1.73006 | 0.00634 | 9.0672 | ↑ |
| 19 | Thymidine | 0.43574 | 0.01278 | 2.98816 | ↓ |
| 20 | Deoxyuridine | 0.51843 | 0.01316 | 1.45515 | ↓ |
| 21 | L-Tryptophan | 2.15465 | 0.01503 | 1.18498 | ↑ |
| 22 | Uracil | 0.53818 | 0.02080 | 0.98912 | ↓ |
| 23 | Uridine | 0.37175 | 0.04768 | 2.57269 | ↓ |
| 24 | Hydroquinone | 2.60759 | 0.04993 | 1.2828 | ↑ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




