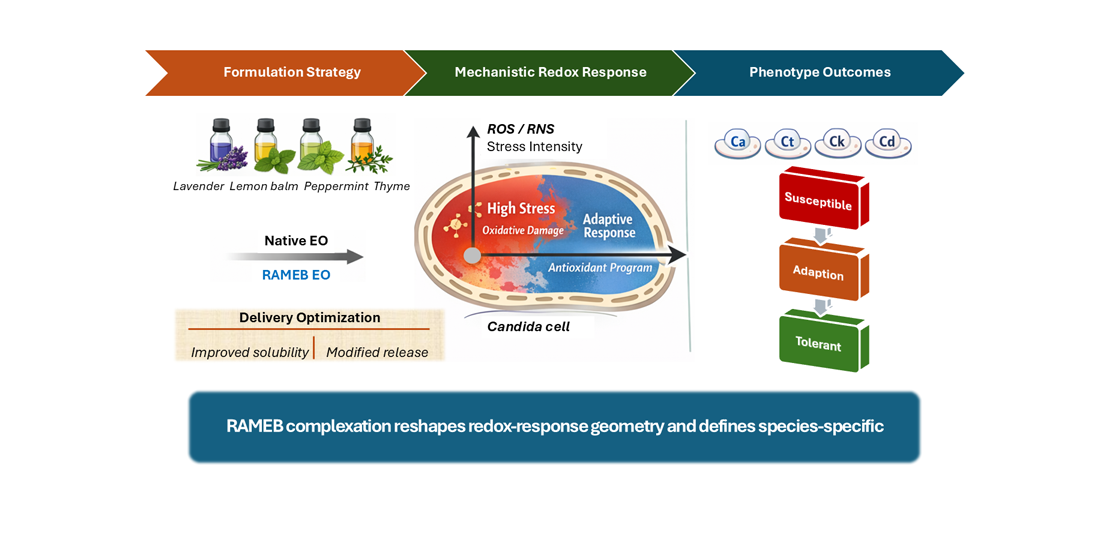
3. Results
3.1. Species-Level Susceptibility, Phase Plasticity, and Survival Effects
To define baseline interspecies susceptibility architecture, we quantified growth-inhibitory (MIC
90) and sub-inhibitory (EC
10) concentration thresholds for native essential oils and their corresponding RAMEB inclusion complexes across
C. albicans,
C. tropicalis,
C. krusei, and
C. dubliniensis. The MIC
90 values reflect near-complete inhibition following prolonged exposure of early-phase cells, whereas EC
10 values capture early functional suppression under short-term exposure in mid-log phase populations. These parameters therefore interrogate distinct physiological states and exposure kinetics. Comparative evaluation across species enables discrimination between formulation-dependent potency shifts and intrinsic taxon-specific resistance characteristics. The distributional structure of MIC
90 and EC
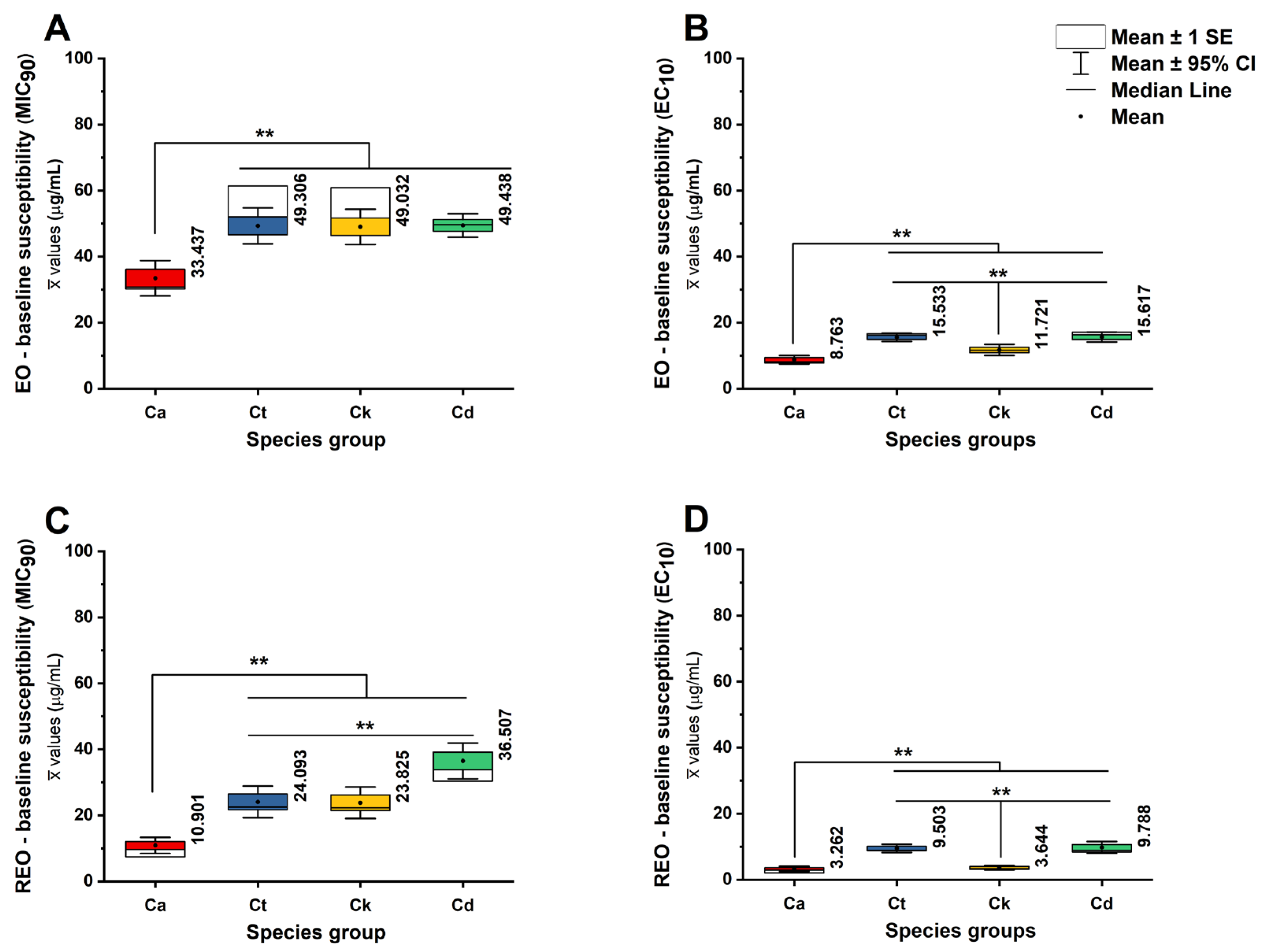
10 values is summarized in
Figure 1 and supplementary
Table S3-S4 (MIC
90 and EC
10 raw data, respectively).
To characterize species-dependent susceptibility patterns and quantify the separation between sublethal and inhibitory concentration ranges, raw MIC90 and EC10 values were compared across the four Candida species using nonparametric methods, as expected for concentration-response endpoints with non-normal distributions. Analyses were performed on the experimental essential oil/RAMEB treatment set (controls excluded), with N representing experimental repeat measurements contributing to each species distribution (total N = 162 repeats; C. albicans n = 54, C. tropicalis n = 36, C. krusei n = 36, C. dubliniensis n = 36)
Species differed significantly in MIC90, indicating that the growth-inhibitory threshold is strongly species-structured (Kruskal-Wallis H(3) = 193.329, p < .001). Mean MIC90 values (weighted average) were lowest for C. albicans (22.186 ± 1.005 µg/mL) and increased markedly for non-albicans species, including C. tropicalis (36.557 ± 1.273 µg/mL), C. krusei (36.371 ± 1.257 µg/mL), and most prominently C. dubliniensis (42.789 ± 1.003 µg/mL) for both EO and REO groups. Consistent with these medians, mean rank ordering placed C. dubliniensis highest and C. albicans lowest, supporting a shift toward reduced inhibitory susceptibility in the non-albicans species (p < .001).
The MIC
90 values demonstrated clear species-specific structuring and varied significantly across the eight experimental treatments (L, B, P, T, RL, RB, RP, RT) (see
Table S2 for strain vs treatment-wise comparisons). Treatment potency of RT (
C. albicans: 2.785 ± 0.179 µg/mL), (
C. krusei: 9.168 ± 0.88 µg/mL), (
C. tropicalis: 9.902 ± 1.02 µg/mL), (
C. dubliniensis: 16.44 ± 0.343 µg/mL) and RB (
C. albicans: 5.812 ± 0.376 µg/mL), (
C. tropicalis: 18.719 ± 1.811 µg/mL), (
C. krusei: 19.25 ± 1.977 µg/mL), and (
C. dubliniensis: 27.357 ± 0.571 µg/mL) emerged as the most potent inhibitory conditions across all taxa. The
C. albicans exhibited the highest susceptibility in these groups (p < .001). Conversely, the highest inhibitory thresholds were observed in treatments L, P, and RP. Notably,
C. dubliniensis showed a distinct resistance profile under treatment RP (65.559 ± 0.454 µg/mL), while
C. tropicalis and
C. krusei displayed their maximum MIC
90 values under treatment L (64.16 ± 0.483 µg/mL, and 62.656 ± 0.32 µg/mL, respectively). Across nearly all tested conditions,
C. albicans remained the most sensitive species (p < .001). Non-albicans species (NACs) required consistently higher concentrations for inhibition, with
C. tropicalis and
C. krusei showing nearly identical resistance patterns in several treatments, such as P (62.447 ± 0.325 µg/mL and 62.076 ± 0.623 µg/mL) and RL (22.395 ± 1.356 µg/mL, and 23.134 ± 1.33 µg/mL), respectively. Treatments T, B, and RL resulted in intermediate MIC
90 values, further emphasizing a gradient of susceptibility that is both treatment-dependent and species-specific.
A similarly strong species effect was observed for EC10 (Kruskal-Wallis H(3) = 103.63, p < .001), demonstrating that sensitivity within the sublethal concentration range also varies by species. The EC10 was lowest for C. albicans (overall distribution of 6.026 ± 0.254 µg/mL) and increased in C. krusei (7.679 ± 0.35 µg/mL) but was substantially higher in C. tropicalis (12.524 ± 0.307 µg/mL) and C. dubliniensis (12.772 ± 0.375 µg/mL). The rank pattern for EC10 placed C. tropicalis and C. dubliniensis at the highest mean ranks, suggesting that these species require higher concentrations to elicit early measurable inhibition, consistent with diminished low-dose sensitivity or enhanced tolerance-like behavior under subinhibitory exposure (p < .001).
The analysis of EC
10 values revealed a distinct species-specific structuring of sublethal thresholds across the eight treatment conditions (
Table S3 for strain vs treatment-wise comparisons). The
C. albicans consistently emerged as the most sensitive species, particularly under treatments RT (0.731 ± 0.507 µg/mL) and RB (1.652 ± 0.103 µg/mL), where the onset of inhibitory effects occurred at the lowest concentrations. In contrast, non-albicans species required significantly higher concentrations to reach this 10% effective threshold. While RT remained the most potent treatment across all taxa, maintaining mean values under ~10 µg/mL for both
C. tropicalis and
C. krusei treatments, L (17.905 ± 0.484 µg/mL, and 16.655 ± 0.409 µg/mL) and P (16.87 ± 0.093 µg/mL, and 17.139 ± 0.324 µg/mL) demonstrated the highest sublethal thresholds. Specifically,
C. tropicalis and
C. krusei exhibited peak EC
10 values under treatment L, while
C. dubliniensis reached its maximum threshold under treatment RP (19.276 ± 0.264 µg/mL). Overall, these data clearly demonstrate that species-specific differences are most evident at the sublethal level, with non-albicans species displaying a broader range of resistance before the initiation of inhibitory activity, at their mid-log phase when compared to early growth phase.
Because absolute inhibitory thresholds do not fully describe the dynamic transition between sublethal and inhibitory exposure regimes, the geometric relationship between EC
10 and MIC
90 values for each strain–treatment combination were furthermore analyzed. The resulting Δs (EC
10-MIC
90) and EC
10/MIC
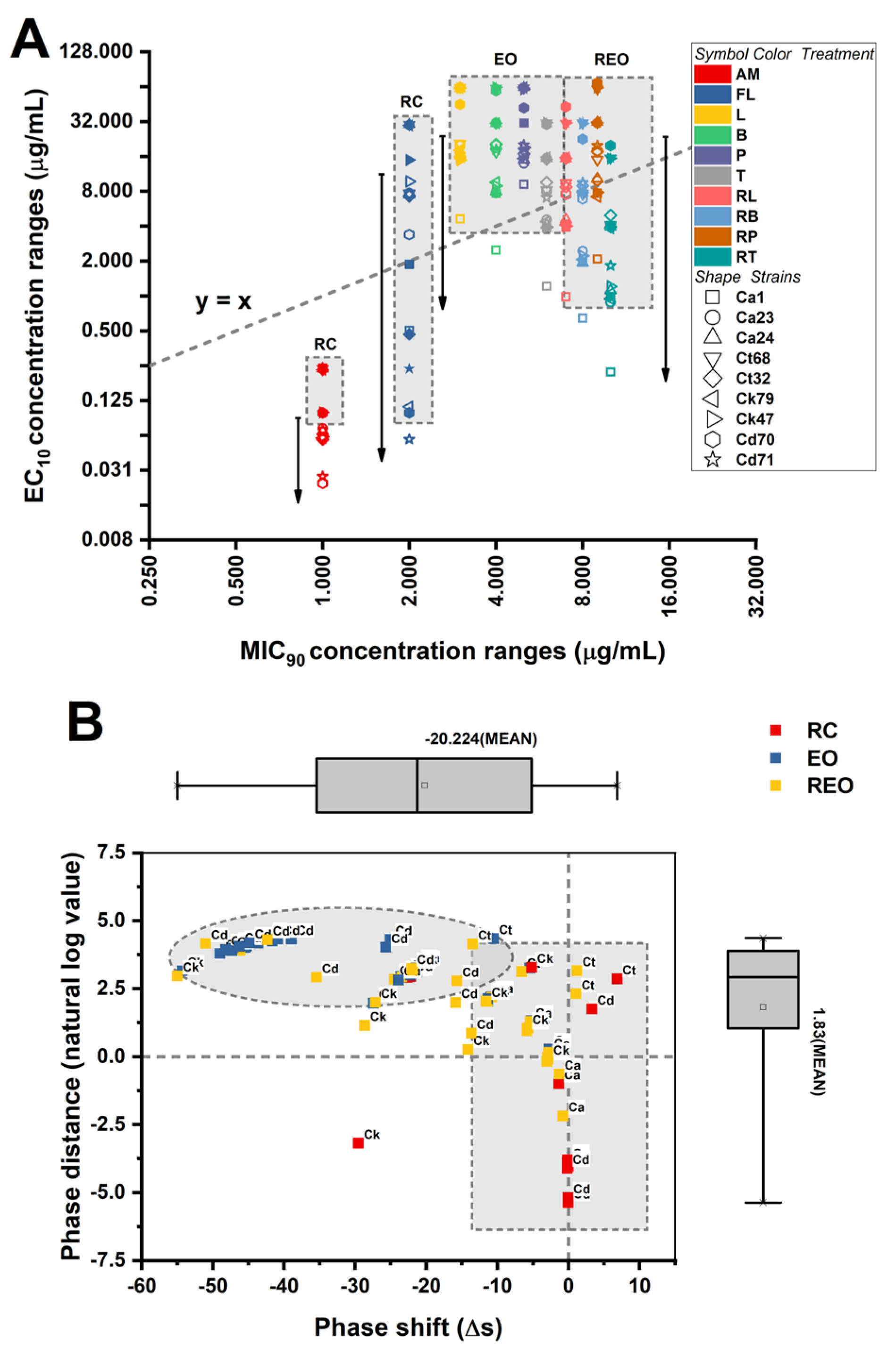
90 metrics quantify the separation between early measurable growth perturbation and near-complete growth inhibition, thereby operationalizing the concept of “phase plasticity.” This framework allows assessment of whether species differ in the breadth of their sub-inhibitory tolerance window and whether RAMEB complexation alters the positioning of treatments within the MIC-EC susceptibility landscape. The multidimensional plasticity architecture is depicted in
Figure 2.
To capture how the sublethal response region relates to the inhibitory threshold, two plasticity metrics were derived: phase shift (Δs = EC10 - MIC90 (absolute separation)) and phase distance = EC10/MIC90 (relative separation). Both metrics differed significantly across species (phase shift: Kruskal-Wallis H(3) = 189.555, p < .001; phase distance: H(3) = 232.148, p < .001), indicating that EC10 is positioned differently relative to MIC90 depending on the species. For phase shift, means were negative across all species, consistent with EC10 occurring at concentrations below the MIC90 threshold. However, the magnitude of this separation differed: C. albicans showed the smallest absolute separation (-16.159 ± 0.757), whereas C. tropicalis (-24.033 ± 1.061) and especially C. krusei (-28.691 ± 0.982) and C. dubliniensis (-30.066 ± 0.693) exhibited more negative shifts. Interpreted biologically, a more negative phase shifts indicates a larger gap between the concentration where early inhibition begins and the concentration required for near-complete growth inhibition, suggesting species-dependent breadth of the sublethal “adaptive window.” For phase distance, species differences were also evident. The C. krusei exhibited the lowest EC10/MIC90 ratio (0.209 ± 0.004), consistent with the largest proportional separation between sublethal and inhibitory thresholds. In contrast, ratios were higher for C. albicans (0.281 ± 0.001), C. dubliniensis (0.285 ± 0.004), and C. tropicalis (0.473 ± 0.018). Taken together, these plasticity analyses indicate that non-albicans Candida species, particularly C. krusei and C. dubliniensis exhibit a broader separation between EC10 and MIC90, supporting the interpretation that adaptive/tolerance-linked concentration ranges may differ by species (p < .001).
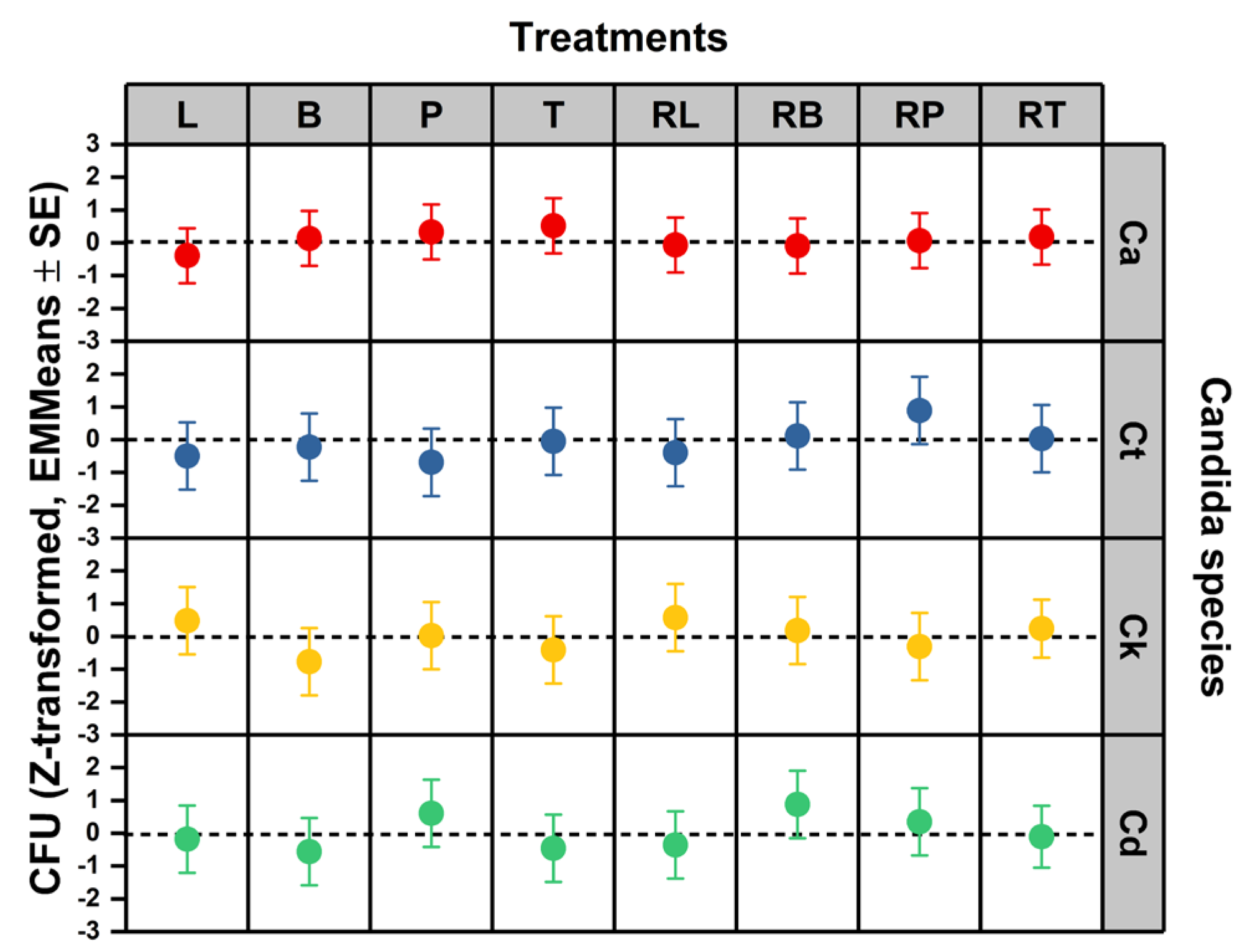
To determine whether species- and formulation-dependent differences in concentration thresholds translate into functional survival outcomes under working exposure conditions, colony-forming unit (CFU) dynamics were analyzed using species-wise standardized linear mixed models. Z-transformation within species isolates treatment-driven deviations from baseline survival variability and permits direct comparison of EO and RAMEB–EO effects across taxa. This analysis tests whether susceptibility and plasticity differences observed at the concentration-response level manifest as measurable shifts in viable population burden. Species-stratified estimated marginal means of standardized CFU responses are presented in
Figure 3.
Survival was quantified by calculating the area under the curve (AUC) from CFU counts across all time points. To account for inherent growth variations between strains, these AUC values were Z-transformed. This standardization ensures that values represent within-strain deviations from the baseline, focusing the analysis on relative survival rather than absolute differences in growth capacity. In linear mixed models accounting for experimental repeats, there was no evidence that standardized CFU differed by treatment, species, or their interaction within the experimental EO/RAMEB set (treatment: F(7, 40.538) = 0.744, p = .637; species class: F(3, 41.579) = 0.223, p = 0.880; treatment x species class: F(21, 41.918) = 0.891, p = 0.602). The EMMeans therefore indicated only directional trends (e.g., more negative values suggesting stronger CFU suppression), but these differences were not statistically robust under the model. Importantly, this lack of effect in standardized CFU suggests that, under the tested working conditions and within-strain normalization, survival responses were comparatively uniform across experimental treatments. Susceptibility and plasticity analyses demonstrated clear species-dependent variations at the concentration-response level. However, under the specific working concentrations utilized, survival measured via CFU remained consistent (non-significant) across all tested Candida strains (
Figure 3).
3.2. Treatment-Induced Oxidative and Nitrosative Stress Responses to Stratify Exposure Signatures
To determine whether essential oil and RAMEB-EO formulations elicit distinct oxidative and nitrosative stress programs across Candida species, standardized mechanistic markers encompassing intracellular reactive oxygen species (ROS), reactive nitrogen species (RNS), and key antioxidant response genes (CAT1, GPX1, SOD1) were analyzed. Because these variables were Z-transformed within species, the resulting profiles reflect treatment-induced deviations from each species’ baseline stress architecture rather than absolute interspecies differences. This approach was used to enable direct comparison of mechanistic exposure signatures across taxa while controlling for inherent species-level variability in redox homeostasis.
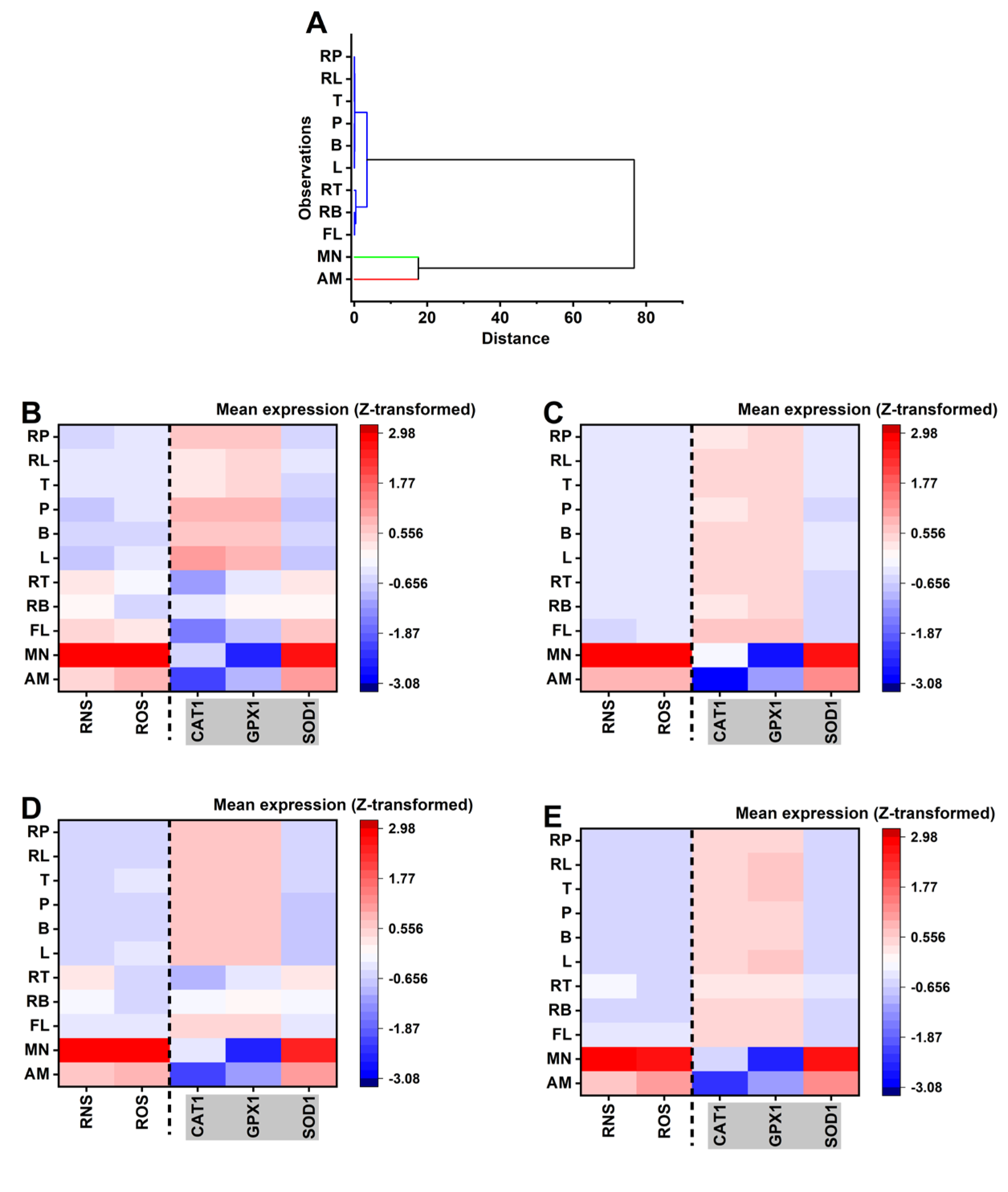
Hierarchical clustering was first applied to the aggregated treatment signatures to define global mechanistic structure. Species-stratified heatmaps were then constructed to visualize how each treatment perturbs the balance between stress generation (ROS/RNS) and antioxidant compensation (CAT1/GPX1/SOD1). Together, these analyses interrogate whether formulation effects primarily alter oxidative burden, antioxidant engagement, or the coordination between these systems. The resulting mechanistic landscape and oxidative-nitrosative response cluster classification is presented in
Figure 4 and
Table 1 respectively.
Visual inspection of the species-specific heatmaps (
Figure 4: panel B-E) for
C. albicans,
C. tropicalis,
C. krusei, and
C. dubliniensis demonstrates that these oxidative-nitrosative exposure signatures are broadly conserved across the tested taxa. Despite the significant differences observed in absolute concentration thresholds (EC
10 values), the relative mechanistic deviations-most notably the high-stress profile of MN and the suppressed response of AM remain consistent across all species. This conservation suggests that while species differ in their individual sensitivity thresholds, the underlying cellular “mode of action” for these treatments is largely species independent. Hierarchical clustering (
Figure 4A) confirms a clear binary split between high-stress anchors and the experimental treatment set. The experimental treatments (B, L, P, T, RL, RB, RP) form a tightly knit hierarchical group characterized by low ROS/RNS generation and a coordinated CAT1/GPX1-driven adaptive response. The proximity of RT and RB to the reference antifungal FL in the dendrogram further suggests that these specific combinations represent a transition state toward higher inhibitory activity while maintaining the general mechanistic signature of the experimental group (see supplementary
Tables S4 and S7 for raw percentage reactive oxygen-nitrogen species generation compared to untreated control and pairwise dendrogram distance matrix for hierarchical clustering, respectively).
To define treatment-specific oxidative-nitrosative stress exposure signatures and the associated antioxidant response programs, standardized mechanistic endpoints (ROS, RNS, CAT1, GPX1, SOD1) across
C. albicans,
C. tropicalis,
C. krusei, and
C. dubliniensis we analyzed by k-clustering (
Table 1). Because all mechanistic variables were Z-transformed within species (split by species standardization), this analysis emphasizes relative mechanistic deviations (mode of action patterns) rather than absolute potency or concentration effects. Accordingly, the experimental treatments were interpreted together with reference anchors: AM and FL as antifungal controls and MN as an oxidative stress positive control. Mechanistic profiles were summarized at the treatment × species level (mean Z-values), yielding N = 44 exposure signatures corresponding to 11 treatments measured in each of 4 species (11 signatures per species). Treatments included A, F, L, B, P, T, RL, RB, RP, RT, MN. The k-means clustering (k = 3) was applied to the five-dimensional signature space (ROS, RNS, CAT1, GPX1, SOD1) and resolved three mechanistic classes (cluster 1 n = 4, cluster 2 n = 7, cluster 3 n = 33). Cluster centers (Z units) indicated that class separation was driven by both the magnitude of ROS/RNS generation and the balance of antioxidant response gene shifts.
In
Table 1, cluster 1 exhibited the strongest stress phenotype, with markedly elevated RNS (+2.904) and ROS (+2.839). This was accompanied by a strong SOD1 upshift (+2.616) and pronounced GPX1 suppression (-2.6), while CAT1 remained near baseline (-0.372). Importantly, Cluster 1 consisted exclusively of MN (4/4 signatures), indicating that MN produced a conserved high-stress signature across all four species and served as a robust “high oxidative-nitrosative burden” anchor in the mechanistic space.
Cluster 2 showed modest elevations in stress markers (RNS +0.484; ROS +0.443) but a disproportionately reduced antioxidant response, particularly CAT1 (-1.88) and GPX1 (-0.832), with only a modest positive shift in SOD1 (+0.839). In terms of composition, this mechanistic class was dominated by AM (4/7 signatures; one per species), with additional membership from RT (2 signatures) and FL (1 signature). Thus, AM displayed a reproducible cross-species signature consistent with stress present but CAT1/GPX1 response suppressed, whereas FL and RT showed occasional drift into this phenotype. Cluster 3 formed the dominant mechanistic class and was characterized by below baseline ROS/RNS (RNS -0.454; ROS -0.449) together with relative increases in CAT1 (+0.469) and GPX1 (+0.507) and a decrease in SOD1 (-0.51). This profile is consistent with an adaptive/low stress signature in which oxidative/nitrosative stress markers are not elevated and antioxidant response patterns preferentially involve CAT1/GPX1 rather than SOD1. Cluster 3 included all experimental treatments across all species (B, L, P, T, RL, RB, RP each contributed 4/4 species signatures) and contained most signatures for F (3/4) and RT (2/4). Distances between cluster centers supported clear separation among the mechanistic classes, particularly between the MN-driven high-stress class and the dominant low-stress/adaptive class (cluster 1 vs 3 distance = 6.498; cluster 1 vs 2 = 4.490; cluster 2 vs 3 = 3.287). Despite distinct mechanistic classes, cluster membership was not associated with species under the species-wise standardized framework. Pearson’s chi-square test showed no enrichment (χ²(6) = 1.905, p = 0.928), indicating that the clustering structure is primarily treatment-signature driven rather than species-driven in this mechanistic space. This suggests that the oxidative-nitrosative exposure signatures are broadly conserved across C. albicans, C. tropicalis, C. krusei, and C. dubliniensis, with species effects more likely to emerge in downstream phenotype/efficacy endpoints than in the top-level mechanistic stratification. Within the mechanistic framework (species-wise Z-standardized ROS/RNS and antioxidant gene responses), the most favorable exposure signature was the dominant cluster 3 pattern-low ROS/RNS with CAT1/GPX1 upshift-capturing all experimental treatments (B, L, P, T, RL, RB, RP) consistently across species and most signatures for FL and RT. In contrast, A reproducibly mapped to cluster 2, characterized by moderate ROS/RNS elevation coupled to suppressed CAT1/GPX1 response, representing a mechanistically less desirable pattern in this signature space. As expected, MN formed cluster 1 exclusively, producing the strongest oxidative-nitrosative burden and serving as the high-stress positive control anchor. Overall, these results indicate that the experimental treatment set predominantly expresses a mechanistic profile consistent with low oxidative/nitrosative burden and adaptive antioxidant engagement, whereas AM is associated with a stress-plus-suppressed-response signature and MN represents maximal stress induction.
3.3. Convergence of Planktonic Efficacy and Mature Biofilm Eradication Across Candida Species
To determine whether early planktonic inhibitory effects translate into suppression of structured biofilm-associated phenotypes, we next evaluated treatment performance across complementary functional endpoints spanning planktonic metabolism (PMT), planktonic viability (PVA), biofilm-associated metabolic activity (BMT), biofilm-associated viability (BVA), and biofilm biomass (BB). Linear mixed models were applied to species-wise Z-transformed responses to enable comparison of relative within-species shifts across treatments. This integrative framework allowed direct assessment of whether treatments that perform strongly under planktonic conditions retain inhibitory hierarchy in the more structured and stress-resilient biofilm context, and whether such convergence is conserved across experimented Candida species. The resulting estimated marginal means profiles are shown in
Figure 5 (see supplementary
Table S5-S6 for reduced percentage PMT, PVA, BB, BMT and BVA raw data).
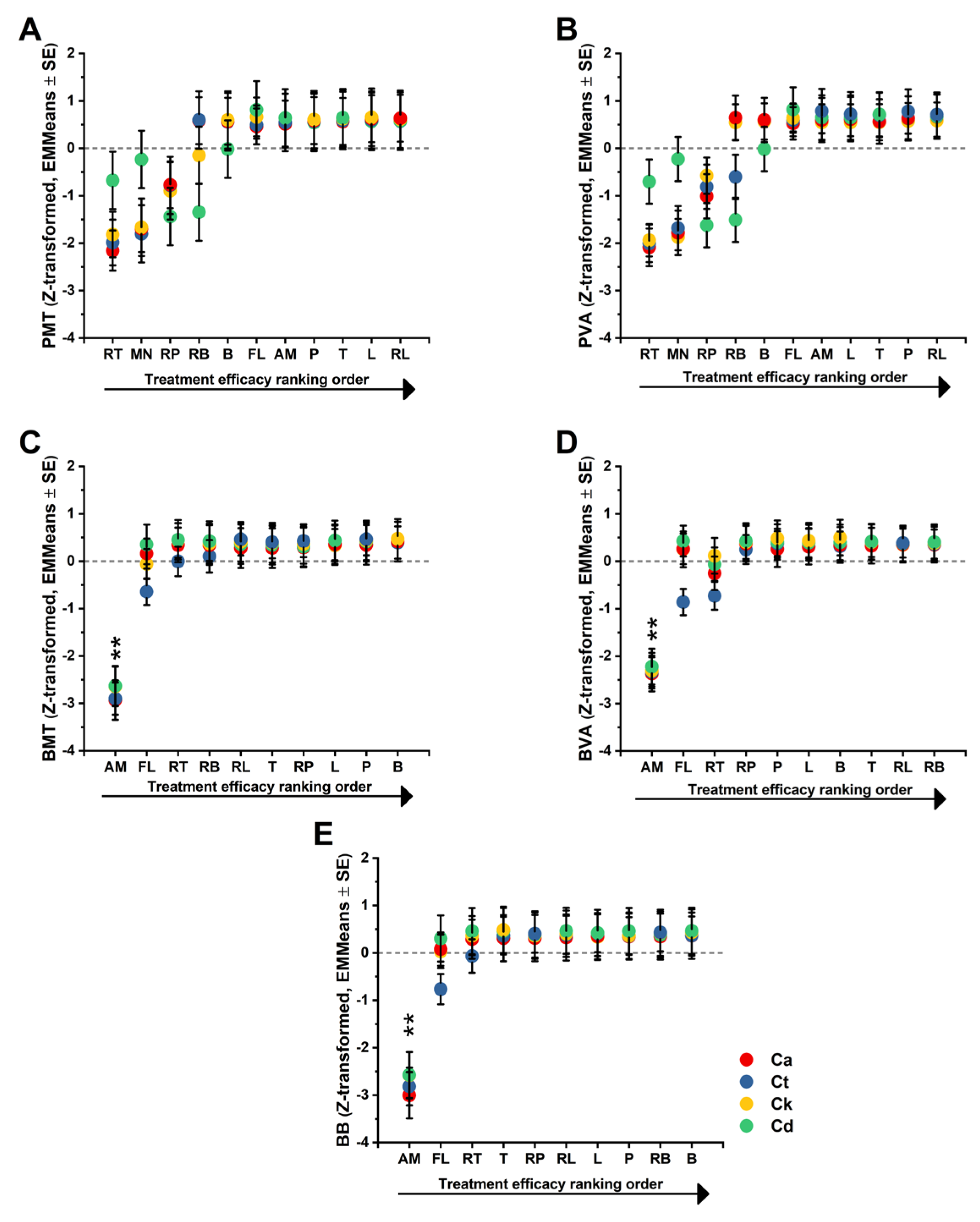
The results summarized in
Figure 5 are supported by two primary visual analyses. First, the estimated marginal means plot (EMMeans) illustrates the overall ranking of treatments, highlighting the significant suppression of metabolism and viability by RT and RP relative to the neutral baseline of the remaining panel. Second, the treatment × species interaction plot visualizes the biological basis of the statistical interaction; it specifically highlights the crossover effects where RB and RP achieve maximal efficacy in
C. dubliniensis compared to their more moderate profiles in
C. albicans and
C. tropicalis. These visualizations confirm the sensitivity of the Z-standardized scale in detecting species-specific physiological shifts.
Strong treatment effects were observed for both planktonic endpoints, demonstrating that Candida planktonic physiology was substantially modulated by the experimental treatment panel. For metabolism, a highly significant overall treatment effect was found (PMT: F(7, 41.506) = 42.829, p < .001), and this effect was even more pronounced for viability (PVA: F(7, 39.861) = 88.926, p < .001), indicating that viability provided high sensitivity for treatment discrimination. The species’ main effect was not found to be significant for PMT (p = 0.3) and was borderline for PVA (p = 0.051). Given the species-wise standardization, these results were expected and indicated that the dominant signal was treatment-driven rather than a reflection of baseline between-species offsets. A significant treatment × species interaction was exhibited by both endpoints, demonstrating that treatment efficacy was not uniform across the four species. This was statistically confirmed for both PMT (F(21, 43.84) = 2.765, p = 0.00224) and PVA (F(21, 41.521) = 6.42, p < .001), while simple-effect analyses confirmed that treatment differences remained significant within each species for both endpoints (all p < .001), supporting a species-stratified interpretation. A concise ranking of treatments on the standardized scale was provided by estimated marginal means (EMMeans), showing consistency between metabolic and viability suppression as efficacy indicators. The RT was identified as the most inhibitory treatment, showing the strongest suppression of metabolism (EMMeans: -1.659 ± 0.133) and viability (-1.679 ± 0.096), while RP (metabolic activity: -0.971 ± 0.134, viability: -1.004 ± 0.096) was ranked second and remained significantly more inhibitory than most of the panels. Whereas RB (0.078 ± 0.134) was found to be near neutral for metabolism but showed a modest, significant reduction in viability (-0.229 ± 0.096), while positive EMMeans were displayed by L, B, P, T, and RL, reflecting weaker inhibition compared to RT/RP. The dominance of RT and RP was reinforced by Bonferroni-adjusted pairwise comparisons, which identified RT as significantly more inhibitory than all other treatments. The biological basis of the interaction was revealed by inspection of EMMs within each species, where RT was characterized as a broad-spectrum inhibitor with strongly negative EMMs produced in all four species for both endpoints, although the magnitude of suppression was attenuated in C. dubliniensis (p < .001). A particularly pronounced inhibitory profile was shown by RP in C. dubliniensis (PMT: -1.438 ± 0.279; PVA: -1.621 ± 0.201), suggesting specific effectiveness against this species, while a striking species-dependent pattern was shown by RB, which explains a major portion of the interaction by showing pronounced inhibition in C. dubliniensis (PVA: -1.509 ± 0.203) despite not being consistently inhibitory in C. albicans and C. tropicalis.
This physiological modulation extended to the inhibition of premature biofilms, where efficacy was similarly quantified through biofilm biomass (BB), metabolic activity (BMT), and viability (BVA). For biofilm metabolism, no significant main effect was detected for treatment (F(7, 40.226) = 0.572, p = 0.774) or species (F(3, 41.071) = 0.893, p = 0.453), indicating that no single treatment performed as the “best” across all species. Global EMM trends for metabolism showed RT at the lowest level (0.306; 95% CI: 0.183-0.429) and RL at the highest (0.377; 95% CI: 0.254-0.501). Analyzing the reduced metabolic activity (compared to UC), shift from essential oil to RAMEB-complexed form, T → RT showed the non-significant metabolic shift (0.049; 95% CI -0.24-0.338). Overall, there was no globally significant difference observed among the treatments, as the collective mean for metabolic inhibition (BMT) for all tested essential oils and their RAMEB-complexed counterparts remained consistently within the 50–60% range relative to the untreated control (UC) (p < .001).
Biofilm viability was identified as the most decisive and treatment-dependent readout in the premature biofilm model (F(7, 39.997) = 10.258, p < .001). A non-significant treatment × species interaction was present (F(21, 40.674) = 1.037, p = 0.446), and species effect remained borderline non-significant (F(3, 40.625) = 2.664, p = 0.446). The RT produced the lowest overall viability (EMMeans: -0.232; 95% CI: -0.369 to -0.096), while B showed the highest (0.392; 95% CI: 0.255 to 0.529). The T → RT transition provided a very large improvement in suppression (shift: 0.62; 95% CI: 0.3 to 0.939, p < .001), marking the strongest “RAMEB-benefit” signal. In contrast to the physiological markers, premature biofilm biomass appeared robust and was not detectably altered by treatments or formulation status on the Z-standardized scale. No significant effects were detected for treatment (F(7, 40.334) = 0.356, p = 0.922), species (F(3, 41.474) = 1.054, p = 0.379), or the interaction (F(21, 41.564) = 0.516, p = 0.947). Overall, EMM trends showed RT with the lowest biomass (0.268; 95% CI: 0.114 to 0.423) and RB with the highest (0.398; 95% CI: 0.243 to 0.552), though these differences were not statistically significant. This highlights a critical decoupling in early biofilm inhibition: while biological markers like viability and metabolism are highly responsive to specialized treatments, the physical biofilm biomass remains insensitive at this stage. In conclusion, treatment efficacy is readily detected at multiple levels and is characterized by a clear species-dependence. The identification of broad-spectrum (RT) versus species-selective (RB, RP) patterns, where RAMEB complexation is not uniformly beneficial, successfully informs subsequent biofilm maturation and mechanistic integration analyses.
3.4. Integrated Treatment Ranking Identifies the Most Effective Regimens Across Endpoints
To move beyond endpoint-specific comparisons and establish a unified hierarchy of treatment performance, a composite efficacy index integrating planktonic and biofilm readouts into a single standardized score were constructed. This integrated metric captures the net biological impact of each formulation across metabolic activity, viability, and biomass endpoints, thereby reducing endpoint-specific bias and highlighting consistent treatment-level effects. Linear mixed modeling was applied to the species-wise Z-transformed composite score to preserve within-species normalization while enabling cross-treatment comparison. This approach highlights the identification of globally superior regimens and, critically, permits direct evaluation of whether RAMEB complexation confers a systematic efficacy advantage within each essential oil family. The integrated ranking and formulation-shift analysis are presented in
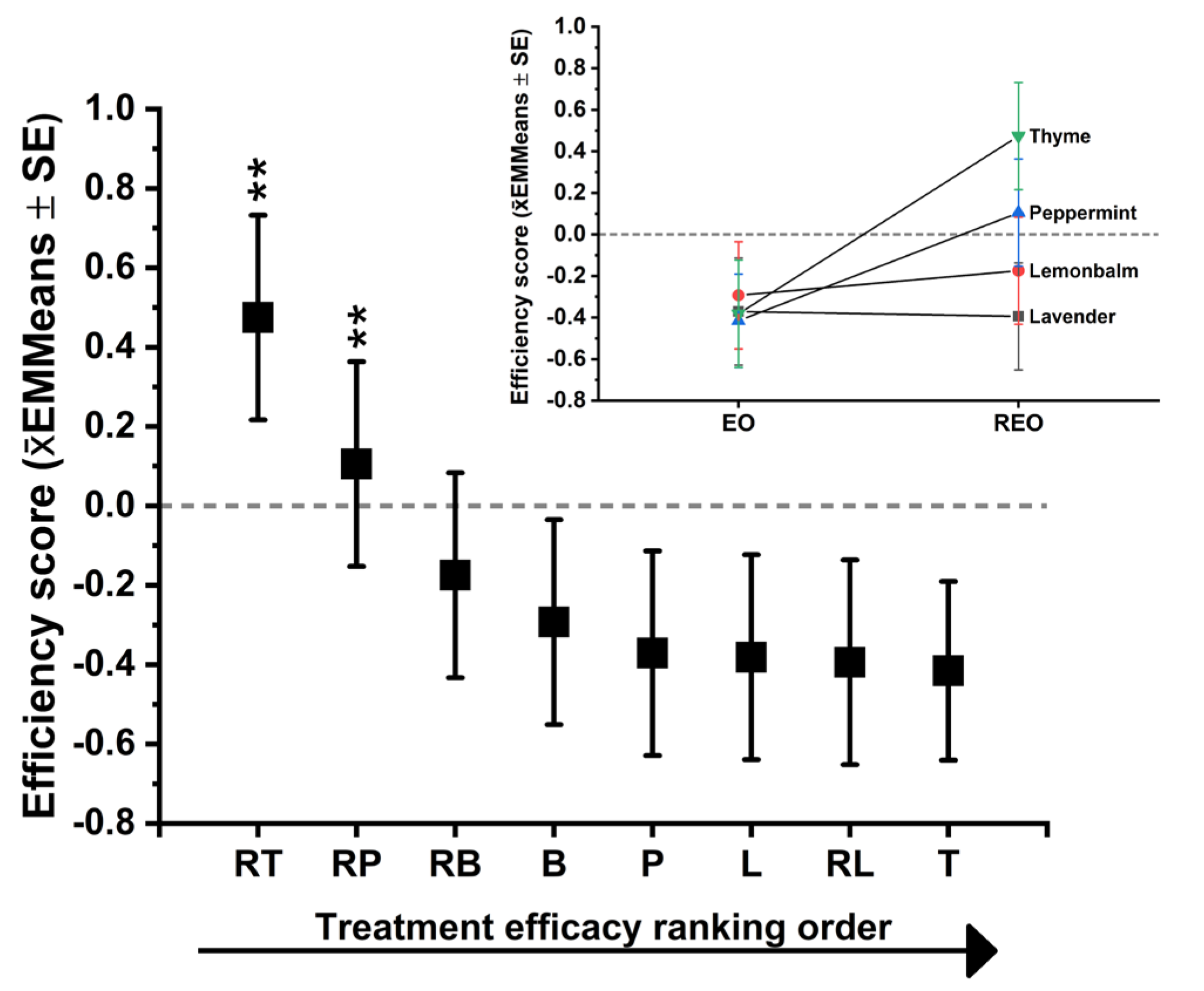
Figure 6.
Inset: Slope plot summarizing the direction and magnitude of formulation effects within each EO family, comparing the parent EO to its corresponding RAMEB inclusion complex (R-EO) using the same integrated score. Upward slopes indicate improved efficacy after complexation, while downward slopes indicate reduced efficacy, illustrating that RAMEB reformulation produces family-specific shifts rather than a uniform advantage across all EOs.
The mixed model demonstrated a pronounced overall regimen effect on the composite outcome (treatment: F(7, 40.752) = 24.552, p < .001), confirming that the experimental treatments differ substantially in their global efficacy across endpoints when assessed in an integrated framework. In contrast, the species main effect was negligible (F(3, 42.523) = 0.106, p = 0.956), which is consistent with the analytic design: because endpoints were Z-standardized within species, the composite score reflects within-species deviations rather than between-species baseline differences. Importantly, the model detected significant species dependence of integrated efficacy, evidenced by a treatment × species interaction (F(21, 43.016) = 2.26, p = 0.012). This indicates that while some regimens are broadly effective, others show species-selective performance, and the “best regimen” cannot be assumed to be identical for all Candida species under the integrated scoring scheme. Estimated marginal means (EMMeans) for treatment, averaged across species, established a clear hierarchy of integrated efficacy (higher EMMeans = better overall inhibition). The RT ranked as the strongest overall regimen (EMMeans = 0.475, 95% CI 0.346-0.603). Its confidence interval was entirely above zero, indicating consistently high performance relative to the grand mean across endpoints and species. Whereas RP ranked second (EMMeans = 0.106, 95% CI -0.024 to 0.235), representing a modest integrated benefit; its CI slightly crossed zero, suggesting greater variability across species/endpoints compared with RT. The RB, however, followed with a negative mean (EMMeans = -0.175, 95% CI -0.304 to -0.045), indicating that, on average across species, RB did not match the integrated efficacy of RT and was not uniformly suppressive across all included endpoints. The remaining treatments formed a lower-performing group with negative EMMs: B (-0.293), L (-0.371), T (-0.381), RL (-0.394), and P (-0.415) (lowest). These negative composite scores indicate comparatively weaker global suppression across the endpoint set, consistent with limited broad-spectrum efficacy under the integrated scoring framework. Multiplicity-controlled pairwise evidence supports RT as the top regimen among all treatments. The RT was significantly superior to every other regimen, including the second-ranked RP (RT > RP: mean difference = 0.369, p = 0.005, Bonferroni-adjusted pairwise comparison). The RT also exceeded RB and the lower-performing group by larger margins (e.g., RT > RB: difference = 0.649, p < .001), confirming that RT’s leading rank is statistically robust under conservative correction. For RP, pairwise comparisons showed that RP was clearly above the low-performing treatments overall, but the difference between RP and RB did not remain significant after Bonferroni adjustment (RP vs RB: p = 0.100). This result supports that RP is generally a strong regimen, but its advantage over RB is not consistently large enough across the dataset to survive the most stringent correction, reflecting species-structured effects. In C. albicans, RT was decisively the top regimen, with a markedly elevated integrated score (RT = 0.862, 95% CI 0.671 -1.054), indicating strong and consistent inhibition across the endpoint set. The RP was near neutral (RP = 0.028, CI crossing zero), and the weakest integrated performance in this species was observed for T (-0.466). This pattern supports RAMEB encapsulated thyme oil as a broad and high-confidence regimen for C. albicans.
In C. tropicalis, RT again ranked highest (RT = 0.529, 95% CI 0.310-0.749). The RP was close to neutral (-0.015), suggesting limited overall integrated benefit in this species. Notably, RB showed the lowest integrated mean (-0.405), indicating that RB’s performance is not uniformly favorable and may be comparatively weak against C. tropicalis when outcomes are integrated. This species therefore contributes to the interaction by separating RT from RB more strongly.
In case of C. krusei, RT remained at the top regimen (RT = 0.453, 95% CI 0.233-0.672), and RP performed relatively well (0.143), consistent with RP having stronger efficacy in some species contexts than others. The weakest regimen in this species was RL (-0.517). Overall, C. krusei preserves the “RT-best” pattern but differs in the ordering and spacing of the remaining treatments, contributing to the interaction signal.
In contrast to the other three species, C. dubliniensis displayed a shifted integrated ranking, where RP emerged as the top regimen (RP = 0.266, 95% CI -0.004 to 0.536) and RB ranked second (0.141), while RT was comparatively weaker (RT = 0.054). Although RP’s CI narrowly crossed zero, the ordering indicates that C. dubliniensis responds preferentially to RP/RB relative to RT in the integrated framework. The poorest composite performance in this species was P (-0.529).
3.5. Response Classes Reveal Susceptible, Tolerant/resistant, and Hormetic/adaptive Phenotypes with Species Enrichment
To translate multidimensional efficacy patterns into biologically interpretable outcome states, treatment–species signatures into discrete response phenotypes based on their integrated endpoint profiles were classified. Using unsupervised clustering of inhibition-coded functional readouts, each signature was assigned to one of three classes representing strong inhibition (susceptible), weak inhibition (tolerant/resistant), or mixed/hormetic-adaptive behavior. This phenotype-level abstraction enables visualization of how each formulation distributes species across inhibitory states, rather than focusing solely on mean shifts. Importantly, this approach further revealed whether treatments drive uniform suppression across tested Candida species or generate heterogeneous responses suggestive of adaptation or tolerance emergence. The distribution of response classes across treatments is summarized in
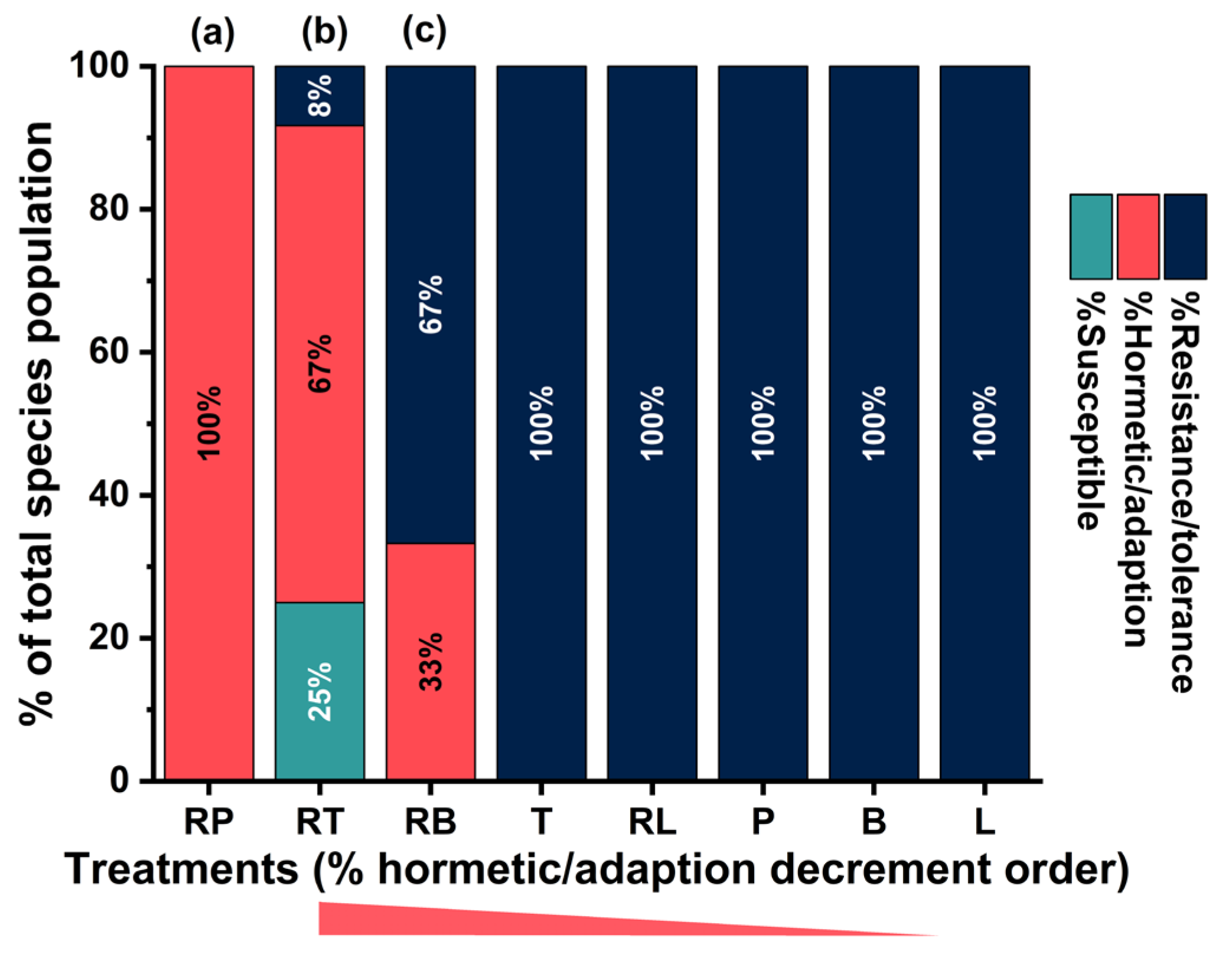
Figure 7.
To translate multi-endpoint responses into biologically interpretable phenotypes, response classes were derived using unsupervised clustering of aggregated strain × treatment signatures. This approach integrated planktonic metabolism and viability with biofilm biomass, metabolic activity, and viability into a single multivariate response fingerprint, thereby enabling each strain–treatment pair to be assigned to one of three phenotypes: susceptible, hormetic/adaptive, or tolerant/resistant. Importantly, all endpoints contributing to these signatures had been Z-standardized within species, such that class membership reflected within-species deviations in response magnitude (i.e., the extent to which a given strain shifted relative to typical behavior of its own species), rather than baseline differences between species. Aggregated response fingerprints were constructed for four Candida species, including C. albicans, C. tropicalis, C. krusei, and C. dubliniensis. For each strain × treatment combination, mean inhibition-coded responses were computed across endpoints representing planktonic inhibition (In-PMT, reflecting planktonic metabolism; In-PVA, reflecting planktonic viability) and biofilm eradication (Er-BB, reflecting biofilm biomass; In-BMT, reflecting biofilm metabolic activity; In-BVA, reflecting biofilm attached cellular viability). Because inhibition-coded metrics were analyzed (defined as the negative of the original species-wise Z values), larger positive values indicated stronger inhibition for the corresponding endpoint, whereas negative values indicated relative maintenance or enhancement of function under exposure in the standardized response space. Response-class analysis was performed across the experimental regimen set encoded as L, B, P, and T, representing base essential oil exposures in non-encapsulated form, and RL, RB, RP, and RT, representing the corresponding RAMEB-encapsulated formulations of the same treatment types. Consistent with earlier integrated ranking analyses in which RT and RP were placed at the top overall, this phenotype-based framework was used to provide an orthogonal interpretation focused on the qualitative response patterns induced across biological compartments.
Unsupervised learning was then applied to the aggregated inhibition-coded fingerprints using k-means clustering with k = 3, yielding three reproducible response classes. Biological meaning was inferred directly from the final cluster centers, which summarized the multivariate inhibition patterns associated with each phenotype. The first class was interpreted as a susceptible phenotype characterized by global, high-magnitude inhibition and therefore representing “true responders” under the standardized framework. In this class, strong planktonic suppression was observed at the cluster center (In-PMT = 1.57; In-PVA = 2.27), accompanied by marked reduction in biofilm viability (In-BVA = 3.36). By contrast, the center values for biofilm biomass and biofilm metabolism were more modest and variable (Er-BB = -0.37; In-BMT = -0.58). This pattern supported an interpretation in which viability was compromised across compartments particularly within biofilms even when biomass and metabolic readouts did not show immediate or uniform inhibition, consistent with mechanisms that disrupt viability or core physiology without necessarily precipitating rapid biomass collapse.
The second class was interpreted as a hormetic/adaptive phenotype, capturing mixed or compartment-shifted inhibition patterns in which planktonic suppression was apparent while biofilm inhibition remained limited. At the class center, planktonic inhibition remained clearly positive (In-PMT = 1.40; In-PVA = 1.39), whereas biofilm endpoints clustered near neutral or negative values, most notably for biofilm viability (In-BVA = -0.07). Under this phenotype, strain treatment pairs were inferred to exhibit strong inhibition in planktonic assays while failing to translate that inhibition into comparably strong suppression of biofilm traits, a pattern consistent with adaptive compensation, biofilm-associated tolerance emergence, or hormetic-like responses in which stress exposure preferentially triggers protective pathways rather than producing uniform inhibition across compartments.
The third class was interpreted as a tolerant/resistant phenotype, representing weak inhibition across endpoints and consistent with minimal global efficacy under exposure. Negative inhibition-coded values were observed across the included endpoints at the class center (In-PMT = -0.57; In-PVA = -0.57; biofilm measures approximately -0.38 to -0.40), indicating that strains assigned to this class-maintained function across both planktonic and biofilm compartments relative to species-typical behavior in the standardized space. Under this interpretation, treatment regimens mapping to this phenotype were inferred to impose insufficient inhibitory pressure and/or to be countered by strain-level tolerance mechanisms that preserved viability and activity under the tested conditions. Across N = 75 aggregated strain × treatment fingerprints, the tolerant/resistant phenotype dominated the response landscape. Specifically, the susceptible phenotype (Class 1) was rare (n = 3; 4.0%), the hormetic/adaptive phenotype (Class 2) was observed at moderate frequency (n = 20; 26.7%), and the tolerant/resistant phenotype (Class 3) comprised the majority of signatures (n = 52; 69.3%). This distribution indicated that globally susceptible response profiles were uncommon, whereas partial or ineffective inhibition patterns predominated, thereby supporting the broader inference that many strain–treatment combinations failed to produce coordinated suppression across endpoints and compartments in this dataset.
Species representation in the clustered dataset was balanced, with totals of n = 25 for C. albicans, n = 17 for C. tropicalis, n = 17 for C. krusei, and n = 16 for C. dubliniensis. To improve statistical stability for enrichment testing, classes were collapsed into a binary phenotype in which “Responder” combined the susceptible and hormetic/adaptive classes, and “tolerant/resistant” corresponded to the tolerant/resistant class alone. Under this collapsed definition, no evidence of association between responder status and species was detected (Pearson χ²(3) = 0.270, p = 0.966; N = 75), and expected counts were adequate (minimum expected count 4.91). These results indicated that responder versus tolerant/resistant phenotypes were not enriched within any species under the species-wise standardized response framework, suggesting that interspecies baseline differences were effectively controlled and that response heterogeneity was primarily driven by strain-level and regimen-level factors rather than species identity.
In contrast to the null species association, response classes were strongly structured by regimen type, with a pronounced contrast between non-encapsulated exposures and RAMEB-encapsulated formulations. The susceptible phenotype was observed exclusively under RT exposure, indicating that RT uniquely generated globally strong inhibitory signatures across planktonic and biofilm endpoints in this dataset. This exclusivity provided phenotype-level corroboration of earlier integrated ranking findings in which RT consistently emerged as the top-performing regimen, and it further suggested that RT was the only regimen capable of reliably driving strain–treatment signatures into the “true responder” state defined by coordinated inhibition. The hormetic/adaptive phenotype was enriched among the encapsulated regimens, being dominated by RP and RT and additionally represented by RB. Under this pattern, strong planktonic inhibition was frequently induced while biofilm inhibition remained incomplete, consistent with a regimen profile that imposed substantial stress yet permitted a compartmental shift toward biofilm-associated tolerance or adaptive compensation in a subset of strains. By contrast, the tolerant/resistant phenotype was heavily populated by the non-encapsulated regimens (L, B, P, and T) and by RL, indicating comparatively weak global inhibition across endpoints for those exposures. In practical terms, these regimens were most often associated with response fingerprints that remained outside the responder phenotypes, thereby providing a mechanistic and phenotypic explanation for their comparatively lower integrated efficacy. Taken together, the response-class framework complemented the integrated ranking results by specifying not only which regimen performed best, but also the phenotypes they most induced. RT was identified as the most reliable driver of the susceptible phenotype, reflecting global inhibition across compartments and aligning with its consistently high overall performance in prior analyses. RP, RT, and RB were frequently associated with hormetic/adaptive profiles, characterized by strong planktonic inhibition coupled to incomplete suppression of biofilm traits, a pattern consistent with adaptive transitions toward biofilm-associated tolerant states. Meanwhile, the predominance of non-encapsulated regimens within the tolerant/resistant phenotype provided a clear phenotypic rationale for their weaker integrated performance, as these exposures most commonly failed to generate coordinated inhibitory pressure across the multi-endpoint response space.
3.6. Mechanistic Inference Stress and Antioxidant Gene Programs Predict Inhibitory Phenotypes
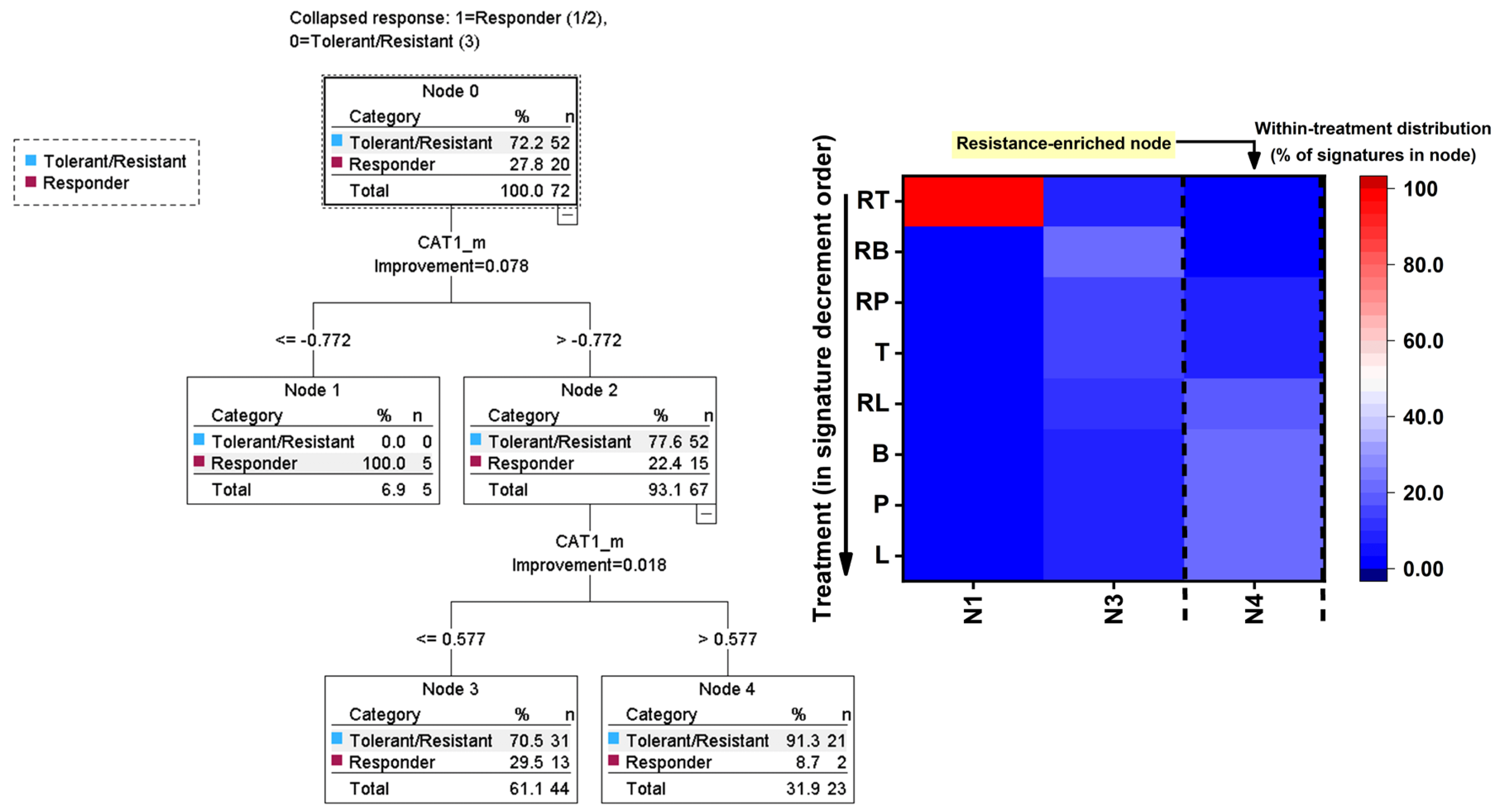
To determine whether mechanistic stress signatures could predict functional inhibitory outcomes, the relationship between oxidative–nitrosative markers (ROS/RNS) and antioxidant response genes (CAT1, GPX1, SOD1) and the derived inhibitory phenotype classes were modeled. Using aggregated treatment × species mechanistic signatures, a classification and regression tree (CRT) approach with Responder (responder vs tolerant/resistant) as the dependent variable was applied. This analysis allowed the identification of hierarchical decision rules linking specific stress-response programs to phenotypic outcomes, thereby moving beyond descriptive clustering toward predictive mechanistic inference. The resulting tree structure and treatment-wise enrichment across terminal nodes are shown in
Figure 8.
To connect the CRT-defined mechanistic programs to regimen identity, terminal node membership was cross-tabulated against treatment, and regimen enrichment was interpreted in the context of the node-wise class composition defined by the Responder outcome. In the full analysis (supplementary
Figure S2) that retained the antifungal and stress controls (AM, FL, and MN), treatment signatures were distributed non-randomly across the terminal nodes of the GPX1/CAT1-structured tree, indicating that specific regimens preferentially occupied distinct regions of the stress/antioxidant marker space rather than mapping uniformly across mechanistic programs. In this full model, the controls anchored opposite mechanistic extremes, with AM mapping exclusively to the high-confidence responder terminal program (Node 3; 9/9, 100%) and MN mapping exclusively to the tolerant/resistant-enriched terminal program (Node 5; 9/9, 100%), thereby validating that the CRT terminal states captured coherent and biologically separable mechanistic configurations. The second antifungal control, FL, was distributed across responder-enriched terminal programs, with strongest representation in Node 7 (7/12, 58.3%) and additional mapping to Node 2 (3/12, 25.0%) and Node 3 (2/12, 16.7%), and no mapping to the tolerant/resistant node, consistent with an overall responder-aligned mechanistic profile. Within the experimental regimens, responder-enriched programs were not confined to a single formulation, as the GPX1-high responder program (Node 2) was preferentially occupied by L (6/9, 66.7%), B (5/9, 55.6%), T (5/9, 55.6%), and RL (5/9, 55.6%), with substantial contribution from P (4/9, 44.4%), whereas a second responder-enriched program (Node 7) was preferentially occupied by RB (6/9, 66.7%), with additional contributions from RL (3/9, 33.3%) and P (3/9, 33.3%). In contrast, RT exhibited pronounced heterogeneity in mechanistic mapping, with a substantial fraction aligning with the tolerant/resistant program (Node 5; 6/12, 50.0%) and the remainder distributed across responder-enriched and mixed programs, while RP preferentially occupied the mixed/ambiguous terminal program (Node 8; 4/9, 44.4%), consistent with context-dependent mechanistic engagement under the CRT rule set.
Because control-driven anchoring can inflate apparent separability and obscure finer regimen structure among experimental exposures, a second CRT was evaluated after exclusion of A, F, and MN (
Figure 8). In this experimental-only dataset (N = 72), node membership differed significantly by treatment identity (Pearson χ²(14) = 51.162, p < .001), although the contingency table remained sparse and enrichment was therefore interpreted descriptively. In this restricted analysis, terminal node membership segregated strongly by regimen, and a single terminal node was composed exclusively of RT signatures (Node 1; 5/5, 100%), indicating that RT occupied a distinct CAT1-defined mechanistic region when controls were removed. This RT-only node represented 55.6% of all RT observations in the experimental-only subset (5/9), while the remaining RT signatures mapped to the dominant mixed node (Node 3; 4/9, 44.4%) and did not appear in Node 4. The largest terminal node (Node 3; n = 44) captured the majority of experimental signatures and contained all RB observations (9/9, 100%), along with high within-treatment representation for B (7/9, 77.8%), T (7/9, 77.8%), and RP (7/9, 77.8%), and moderate representation for P (5/9, 55.6%), RL (5/9, 55.6%), and L (4/9, 44.4%). The remaining terminal node (Node 4; n = 23) concentrated L signatures (5/9, 55.6%) and included substantial contributions from P (4/9, 44.4%) and RL (4/9, 44.4%), with smaller contributions from B (2/9, 22.2%), T (2/9, 22.2%), and RP (2/9, 22.2%), while remaining entirely unoccupied by RB (0/9, 0%) and RT (0/9, 0%). Thus, after exclusion of controls, regimen stratification persisted but was expressed primarily as differential occupation of CAT1-defined terminal programs, with RB consistently mapping to the dominant mixed terminal node, RT partially isolating into a treatment-specific terminal node, and L/P/RL showing relatively greater representation in the alternative terminal node.
3.7. Multivariate Mode-Of-Action Landscape Defines Treatment Signatures and Mechanistic Axes
To integrate oxidative-nitrosative stress markers and antioxidant gene responses into a unified mechanistic framework, principal component analysis (PCA) on the aggregated treatment × species signatures were performed . Because all variables were standardized within species, this multivariate approach captures relative mechanistic positioning rather than potency differences, enabling comparison of mode-of-action geometry across treatments. The PCA has reduced the five-dimensional stress-response space (RNS, ROS, CAT1, GPX1, SOD1) into orthogonal axes that summarize dominant patterns of covariation. The resulting biplot (
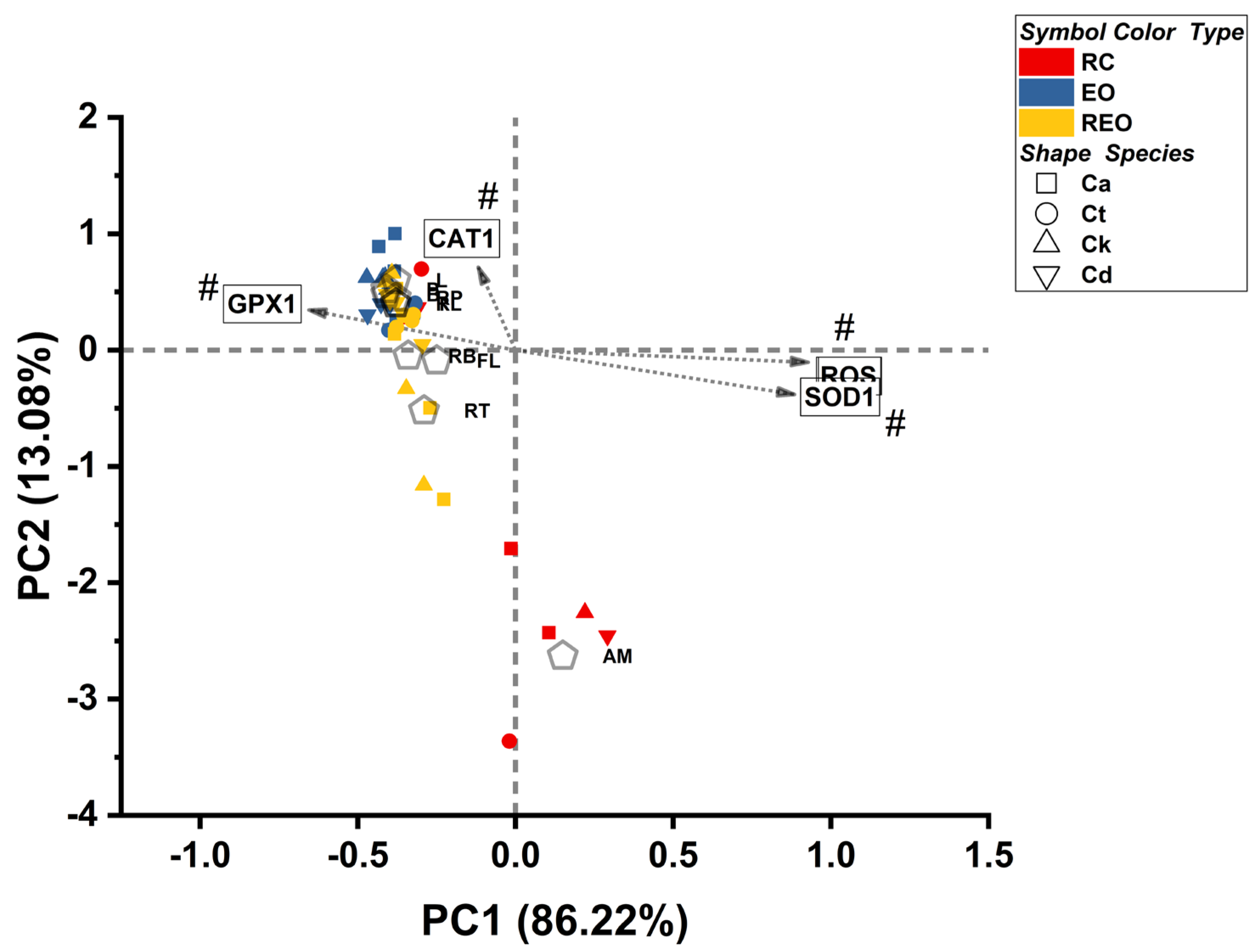
Figure 9) visualizes how treatments and species distribute along a primary stress-intensity axis and a secondary antioxidant-program axis, thereby defining the global mechanistic landscape of the EO and RAMEB-EO formulations.
Principal component analysis (PCA) was performed on aggregated treatment × species mechanistic signatures (RNS, ROS, CAT1, GPX1, SOD1; Z-transformed within species). The biplot shows PC1 (86.22%) versus PC2 (13.08%) for each treatment–species signature. Points are colored by treatment type (RC = antifungal reference controls, EO = essential oils, REO = RAMEB-EO complexes) and shaped by Candida species (Ca, Ct, Ck, Cd). Dashed crosshairs indicate the origin (0,0). Grey arrows depict rotated loading vectors: PC1 captures a dominant stress/redox intensity axis (positive direction aligned with ROS and SOD1, opposed to GPX1), whereas PC2 reflects a CAT1-centered antioxidant-response axis (positive direction aligned with CAT1). Centroid labels (treatment codes) indicate the mean position of each treatment across species. Hash marks (#) denote variables with the strongest contributions to the displayed axes (highest absolute loadings). Note: MN is excluded from this panel to prevent stress-control anchor from compressing the experimental space; full PCA including MN is provided in supplementary materials (
Figure S3).
Component interpretation was guided by variable vectors projected onto PC space. PC1 was aligned with a strong oxidative-stress direction, as ROS projected prominently in the positive PC1 direction and co-varied with the SOD1 vector, while GPX1 projected strongly in the opposing (negative PC1) direction. The PC2 was dominated by the catalase axis, with CAT1 projecting primarily along the positive PC2 direction. Thus, the MoA plane was interpreted as contrasting ROS/SOD1-associated stress programs against GPX1-associated antioxidant configurations along PC1, with CAT1-associated variation represented orthogonally along PC2. Because treatment centroids and strain-level points were overlaid in the same space, these axes were interpreted as integrated MoA dimensions reflecting coordinated shifts in mechanistic marker configuration across strain–treatment signatures rather than isolated single-marker effects. Two visualizations were required to preserve interpretability across the full mechanistic range of the dataset. In the full MoA map (supplementary
Figure S3), inclusion of the oxidative stress control MN generated an extreme displacement along positive PC1 in the direction of ROS (and toward the SOD1-aligned quadrant), thereby anchoring the oxidative-stress end of the landscape but simultaneously compressing the remaining treatments near the origin when a single global scale was applied. For clarity of the experimental regimen structure, a zoomed view (the first image) was therefore examined in which MN was excluded from the plotted space, allowing separation among the remaining treatments to be resolved without altering the underlying PCA solution or its mechanistic interpretation.
Treatment positioning in PC space was non-random and mechanistically interpretable in both views. In the full map, MN was separated far to the right on PC1, consistent with a distinct ROS-dominated stress signature that was not approached by any experimental regimen, thereby providing an internal validation that the PCA captured a coherent oxidative-stress program. The antifungal control AM separated strongly along negative PC2, placing it opposite the CAT1 vector and indicating a mechanistically distinct program characterized by displacement away from the CAT1-aligned direction relative to the experimental cluster. The remaining regimens formed a compact treatment-defined constellation near the negative PC1 region, indicating broad alignment toward the GPX1-associated direction and away from the MN-defined ROS extreme. Within this constellation, the base (non-encapsulated) experimental regimens (L, B, P, and T) clustered tightly with the corresponding encapsulated formulations (RL, RB, and RP), indicating that most experimental exposures shared a broadly similar multivariate stress/antioxidant configuration in this MoA space, with relatively subtle between-regimen differences compared with the control anchors. Notably, RT showed the most consistent displacement among the experimental regimens by shifting downward relative to the main cluster (negative PC2), placing it farther from the CAT1-aligned direction than most other experimental signatures and indicating a distinctive catalase-opposed mechanistic positioning within regimen space. The FL mapped within the central experimental constellation and overlapped most strongly with the region occupied by RB and the densely clustered L/B/P/T/RL/RP signatures, consistent with a responder-aligned but not extreme mechanistic configuration relative to the two control anchors. Formal multivariate testing of PCA scores (listwise N = 96) supported the visual inference that the MoA landscape was treatment-defined. Treatment identity significantly structured the multivariate PC space (Pillai’s Trace = 1.667; F(18,172) = 47.82; p < .001), with significant treatment effects detected on both axes (PC1: F(9,86) = 46.27, p < .001, R² = 0.829; PC2: F(9,86) = 49.46, p < .001, R² = 0.838). In contrast, species did not structure the global PC1-PC2 MoA space (Pillai’s Trace = 0.0147; F(6,184) = 0.226; p = 0.968), indicating that the dominant multivariate organization reflected regimen-associated signature shifts rather than baseline species separation.
Overall, the MoA landscape was defined primarily by treatment identity, with MN and AM providing mechanistically interpretable anchors at opposite extremes of the space. MN uniquely occupied the ROS/SOD1-aligned stress extreme on PC1, whereas AM separated most strongly along PC2 in the direction opposing CAT1. Most experimental regimens (L, B, P, T, RL, RB, and RP) occupied a compact region toward negative PC1 consistent with relative alignment to the GPX1-associated direction and away from ROS-driven stress, while RT exhibited the clearest within-experimental deviation by shifting toward negative PC2, indicating a distinctive catalase-opposed positioning within the treatment-defined MoA space. The complete data on the eigenvalues, rotated component matrix of the mechanistic markers and principle component centroid values can be found in supplementary
Tables S8-S10.