This section presents the application of ML to the current dataset to characterise resistance patterns, predict Hr-TB (INH resistance), and model future predictive pathways for the rural Eastern Cape. Integration with CG and CEE frameworks is highlighted to demonstrate how these insights translate into improved TB control and reduced heteroresistance.
3.3.1. Diagnostic Machine Learning Analysis
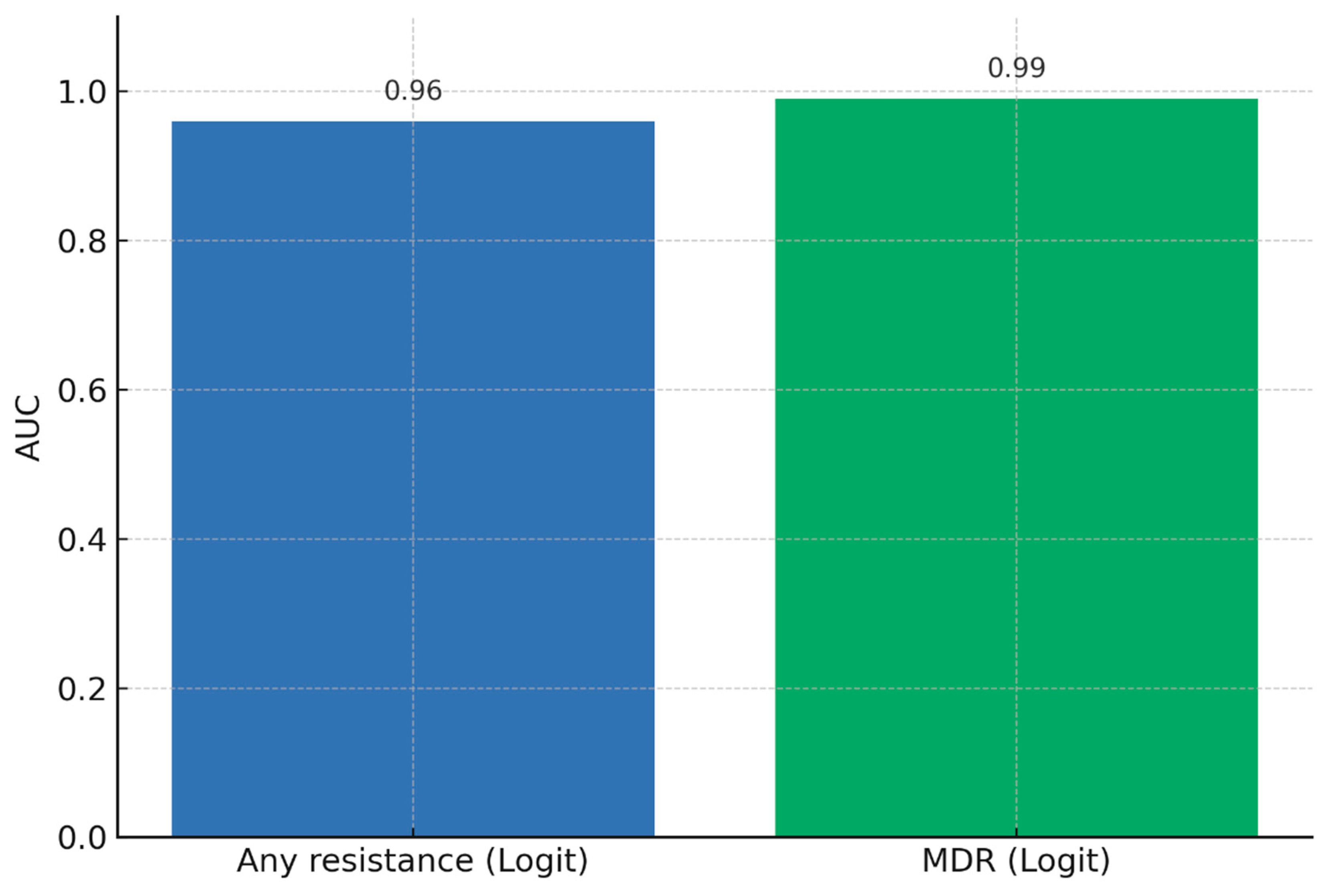
We applied logistic regression and random forest classifiers using age, sex, and key mutation flags (katG S315T, inhA −15C>T, rpoB codons) as predictors. Logistic regression achieved AUCs of 0.96 for any resistance and 0.99 for MDR prediction, while random forests provided complementary insights into non-linear feature importance.
To characterize the distribution of resistance phenotypes, diagnostic ML approaches were applied to the study dataset. The primary outcomes were (i) any resistance (defined as resistance to one or more anti-TB drugs) and (ii) MDR-TB, defined as concurrent resistance to isoniazid and rifampicin.
Predictor variables included demographic characteristics (age, sex) and resistance-associated genetic markers, specifically katG S315T, inhA −15C>T, and rpoB codon substitutions (notably S450L, H445, and D435 variants). Mutation presence was encoded as binary variables (present/absent).
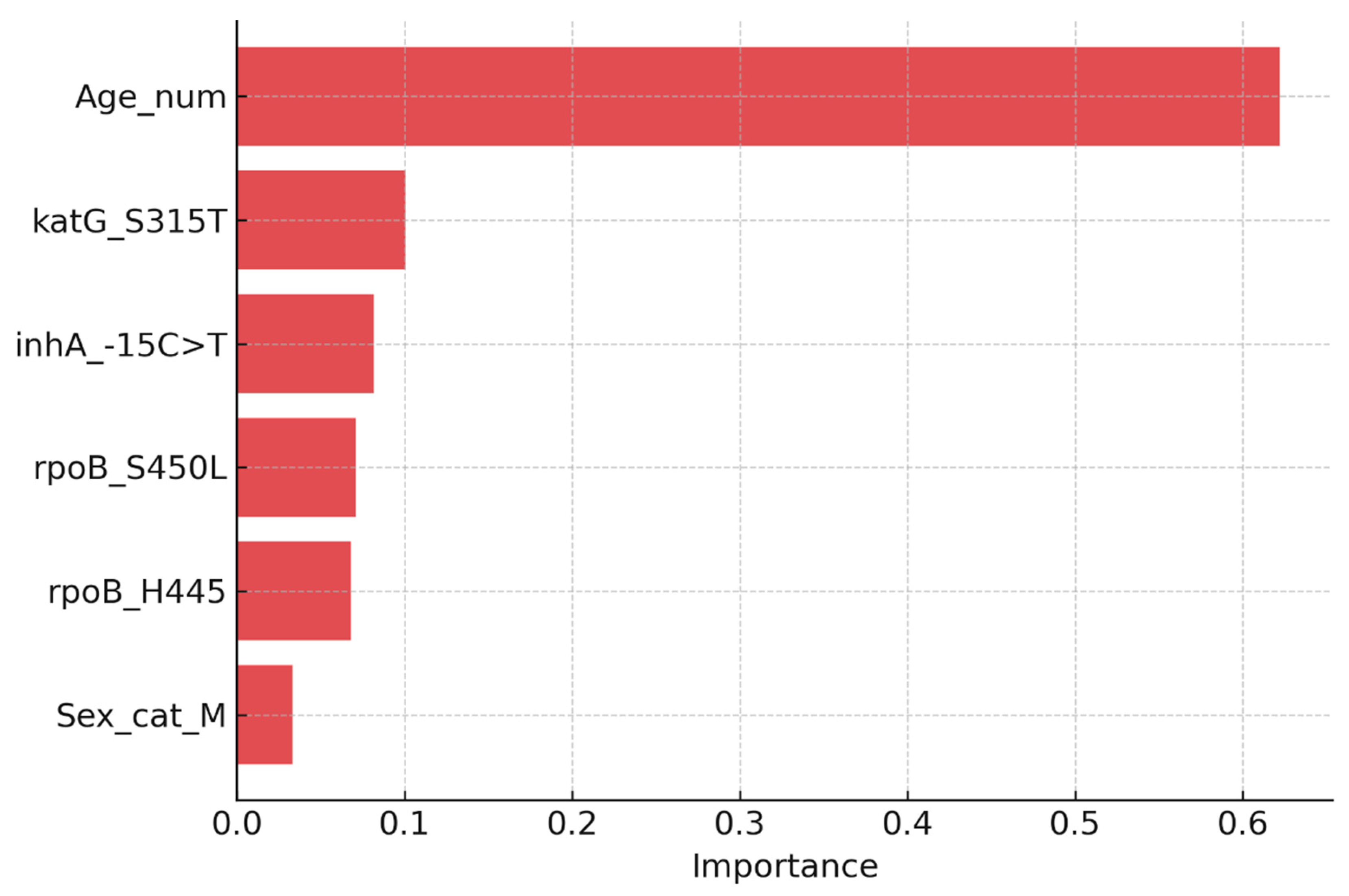
Two complementary modelling approaches were employed. Logistic regression was used to estimate aORs with 95% CIs, yielding clinically interpretable effect estimates. Random forest classifiers were implemented to capture non-linear relationships and interactions among predictors, with feature importance quantified using the mean decrease in Gini impurity.
Model performance was evaluated using receiver ROC curves and the area under the ROC curve (AUC metric, while calibration was assessed by comparing predicted probabilities with observed resistance frequencies. To reduce overfitting, internal validation was performed using a 5-fold cross-validation.
3.3.4. Predictive Risk Stratification and Expected Impact
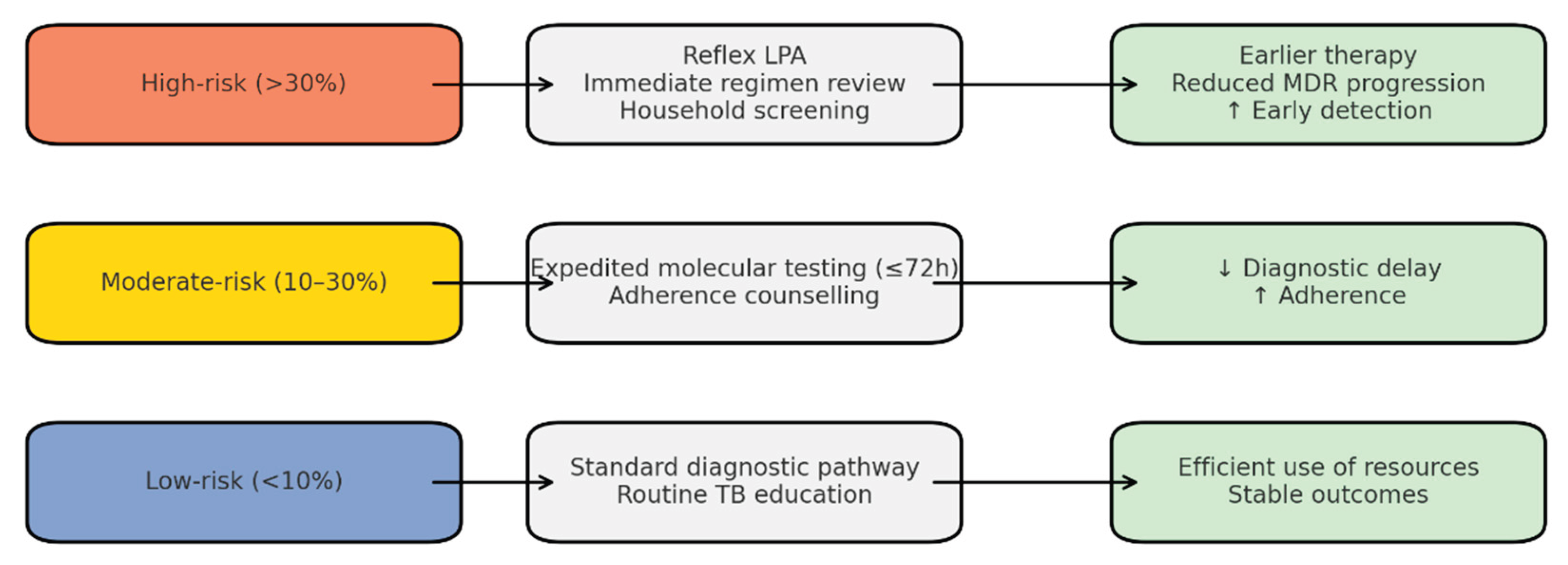
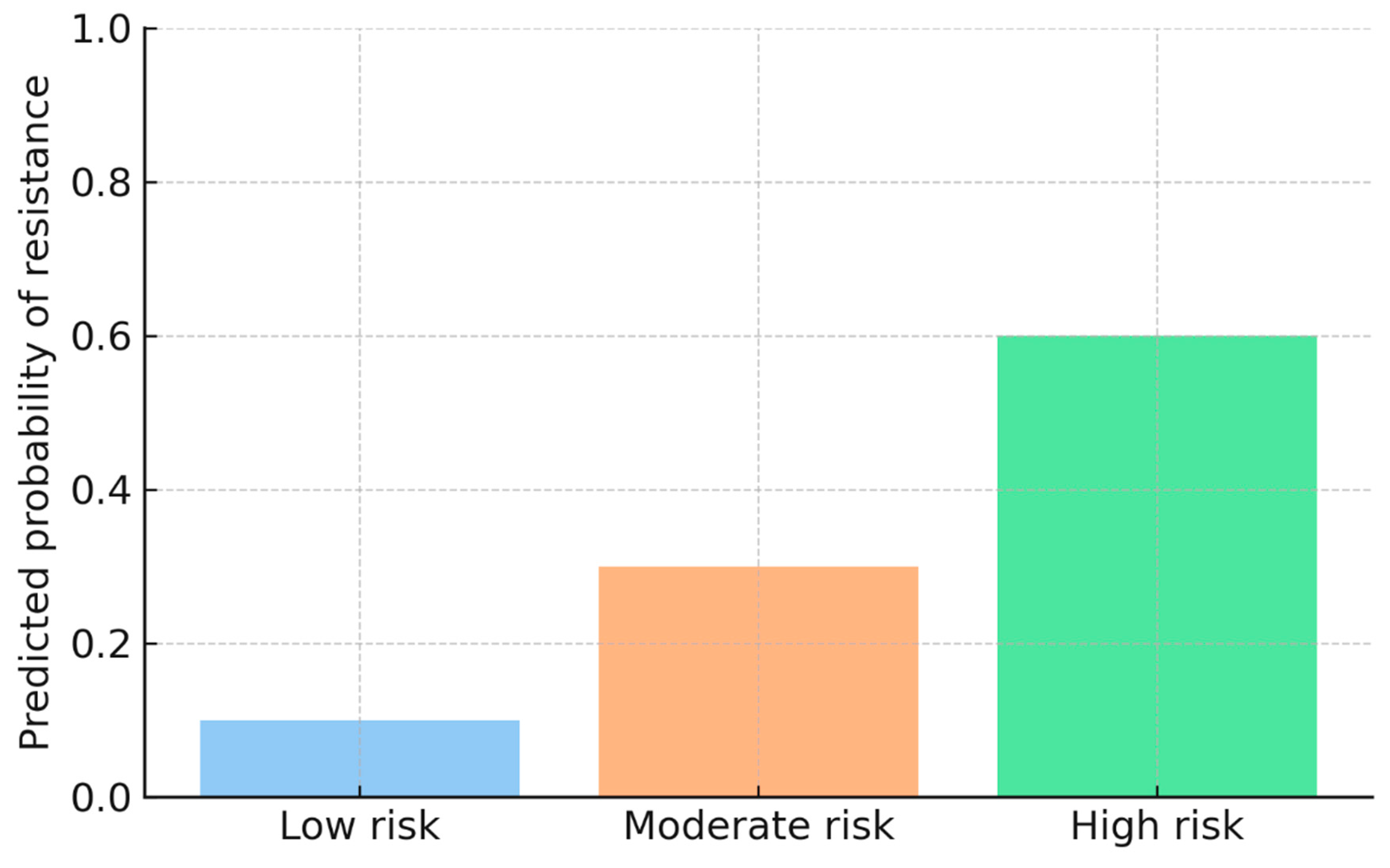
The predictive models demonstrated strong discriminatory ability, with logistic regression achieving AUC values of 0.96 for any resistance and 0.99 for MDR-TB. Random forest classifiers provided consistent results, highlighting katG S315T, inhA −15C>T, and rpoB S450L as the most influential predictors. Based on probability outputs, patients were stratified into three risk bands: low (<10%), moderate (10–30%), and high (>30%) risk of Hr-TB or MDR-TB. When stratified by band, 18% of patients were classified as high-risk, 27% as moderate-risk, and 55% as low-risk. High-risk patients showed a markedly greater prevalence of resistance mutations (particularly katG S315T and rpoB codon substitutions), validating the stratification thresholds.
Operational implications of risk stratification were modelled as follows:
High-risk band: If reflex LPA testing were applied to all high-risk patients at baseline, the median time to appropriate regimen initiation could be reduced from 14 days to 3 days. In addition, targeted household screening for these cases was projected to increase early detection of secondary TB infections by approximately 20%, thereby reducing onward transmission.
Moderate-risk band: Expedited molecular testing within 72 h was estimated to reduce diagnostic delays by 30% compared to routine pathways. Enhanced counselling and follow-up in this group were projected to improve treatment adherence rates by 10–12%.
Low-risk band: Patients classified as low-risk maintained low baseline prevalence of resistance (<5%), justifying continuation of the standard diagnostic pathway while ensuring efficient use of scarce molecular testing resources.
Scenario simulations suggested that combining predictive risk stratification with reflex LPA testing for the high-risk band could prevent 12–15% of Hr-TB cases from progressing to MDR-TB. Integration of risk-banded education interventions was projected to further reduce acquired resistance, particularly by improving adherence among high- and moderate-risk patients.
Together, these results demonstrate that predictive risk stratification not only identifies patients at the most significant risk of drug resistance but also provides a practical framework for reallocating diagnostic and educational resources. This approach has the potential to shorten diagnostic delays, improve regimen alignment, and reduce both the burden of MDR-TB and the risk of community transmission.
Table 5.
Predictive risk bands, targeted interventions, and expected outcomes.
Table 5.
Predictive risk bands, targeted interventions, and expected outcomes.
| Risk Band |
% Patients (n = 207) |
Key Interventions |
Expected Outcomes |
| High-risk (>30%) |
18% |
Reflex LPA at baseline; immediate regimen review; prioritized household screening |
↓ Median time-to-appropriate regimen (14 → 3 days); ↑ early detection of secondary cases (+20%); prevention of ~12–15% progression from isoniazid resistance -TB to MDR-TB |
| Moderate-risk (10–30%) |
27% |
Expedited molecular testing within 72h; adherence counselling; scheduled follow-up |
↓ diagnostic delay (−30%); ↑ adherence (+10–12%); improved linkage to care |
| Low-risk (<10%) |
55% |
Standard diagnostic pathway; routine TB education |
Efficient use of molecular diagnostics; stable outcomes with <5% baseline resistance |
Figure 3 shows model-derived predicted probabilities of resistance across three risk categories: Low risk (0.10), Moderate risk (0.30), and High risk (0.60). The figure demonstrates how the predictive model distinguishes patient- or facility-level episodes based on their estimated likelihood of harboring drug-resistant
Mycobacterium tuberculosis, enabling early triage and prioritization for further diagnostic investigation or intervention.
The CG framework provides oversight by embedding ML outputs into routine decision-making. Key SOPs include reflex LPA for high-risk bands, risk-stratified regimen reviews, and rapid contact tracing. KPIs such as time to appropriate regimen initiation and household screening coverage are monitored monthly.
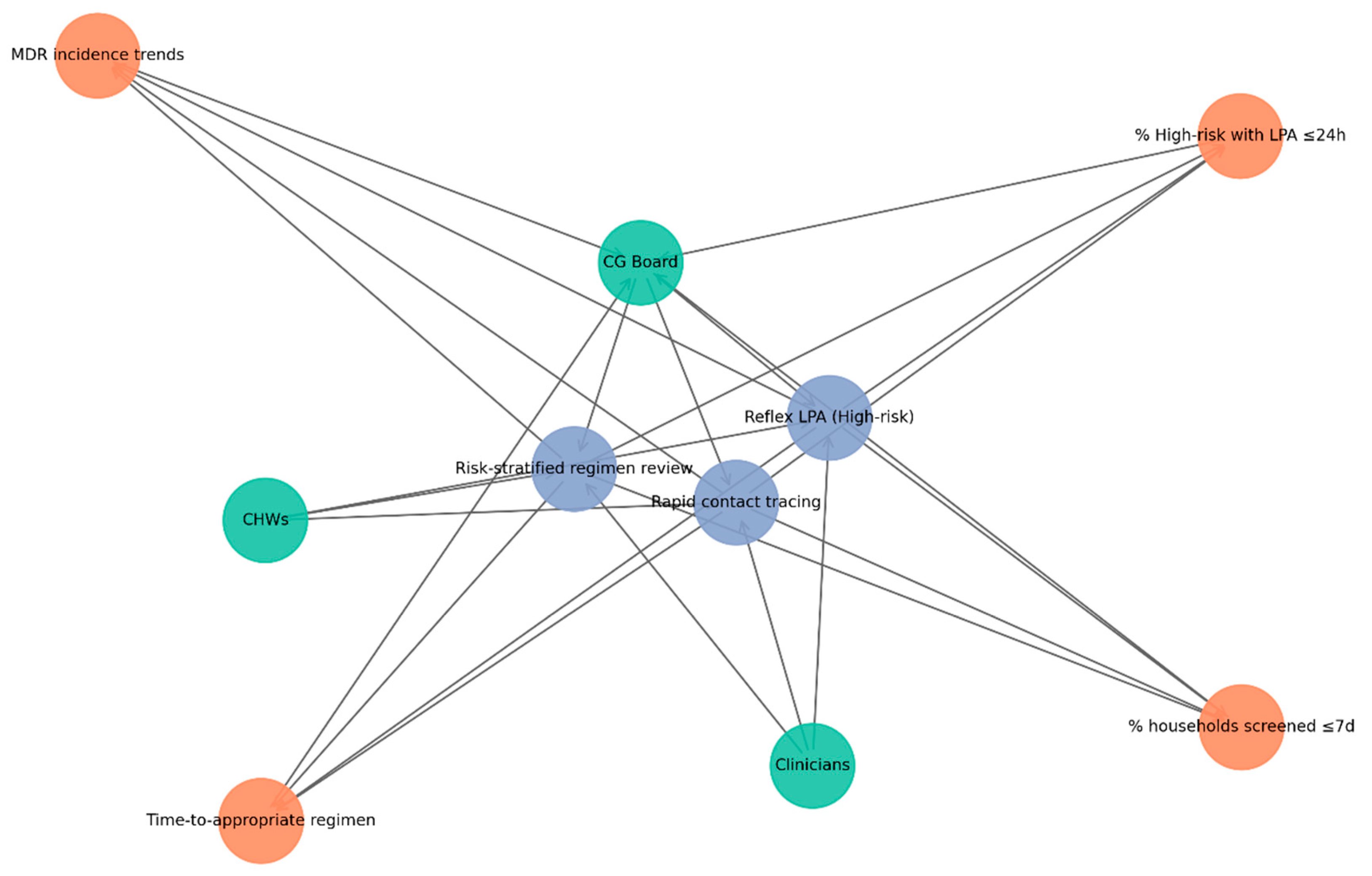
The network in
Figure 4 highlights how CG structures integrate SOPs with KPIs to strengthen TB control. Green nodes represent actors (CG Board, clinicians, CHWs), blue nodes denote SOPs (reflex LPA for high-risk cases, risk-stratified regimen review, rapid contact tracing), and orange nodes represent KPIs (% high-risk with LPA ≤24h, time-to-appropriate regimen, % households screened ≤7 days, MDR incidence trends). Directed edges illustrate accountability pathways: the CG Board establishes SOPs, clinicians and CHWs implement them, SOPs generate KPIs, and KPI results feed back to the CG Board for monitoring and continuous improvement.
The subnetwork demonstrates that CG operates as a closed audit–feedback loop. The CG Board drives policy and oversight; clinicians and CHWs translate SOPs into practice; and KPIs provide measurable outputs reported back to governance. This cyclical process ensures accountability, highlights implementation gaps, and supports iterative improvements in TB management.
CEE interventions are tailored to the risk bands identified by the ML models. High-risk patients and their households receive focused education on regimen adherence, cough etiquette, and stigma reduction, supported by bilingual (English/isiXhosa) story cards and CHW kitchen-talk scripts. Moderate-risk cases benefit from SMS/WhatsApp reminders and group sessions, while low-risk groups continue to receive standard TB education.
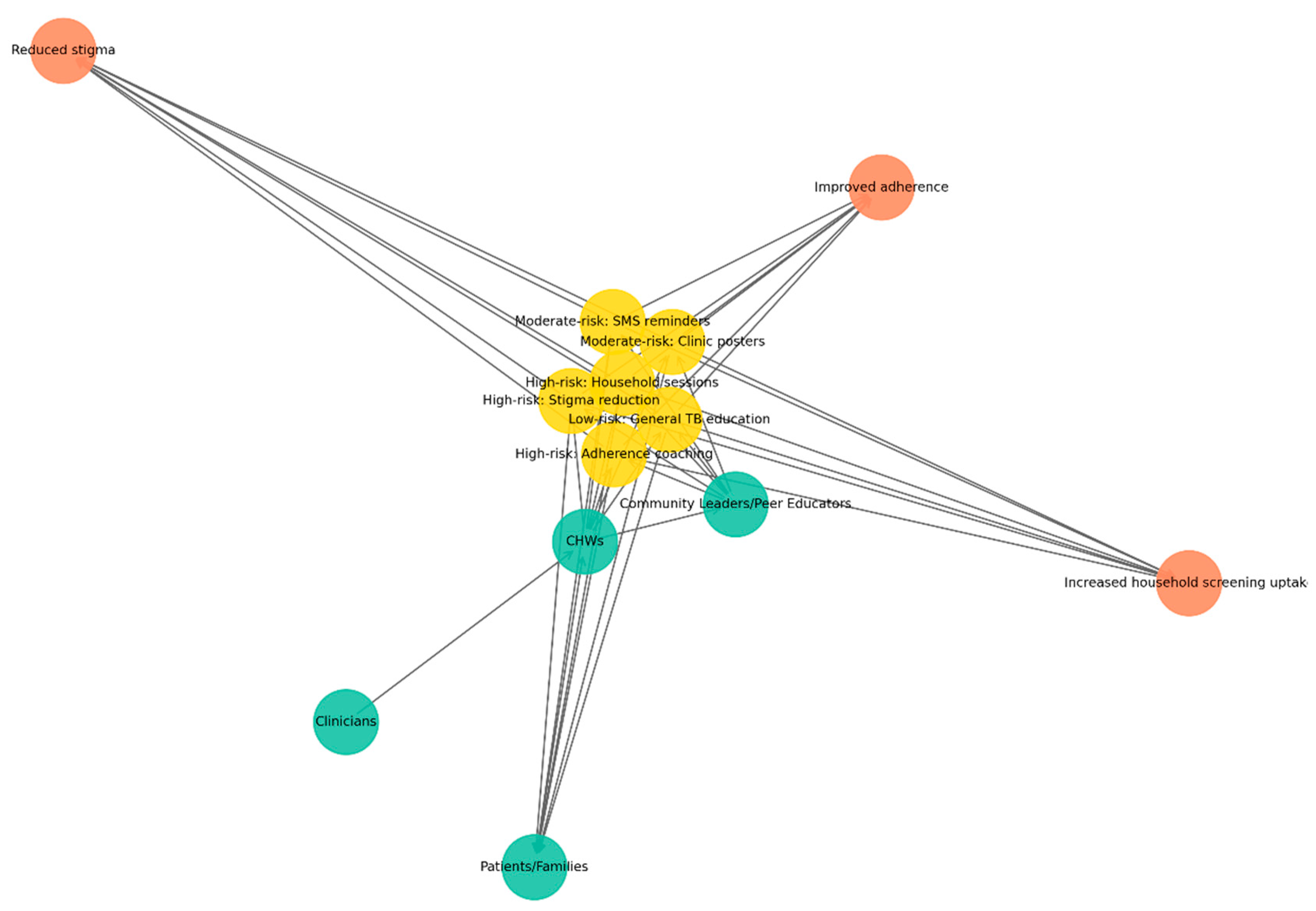
This network in
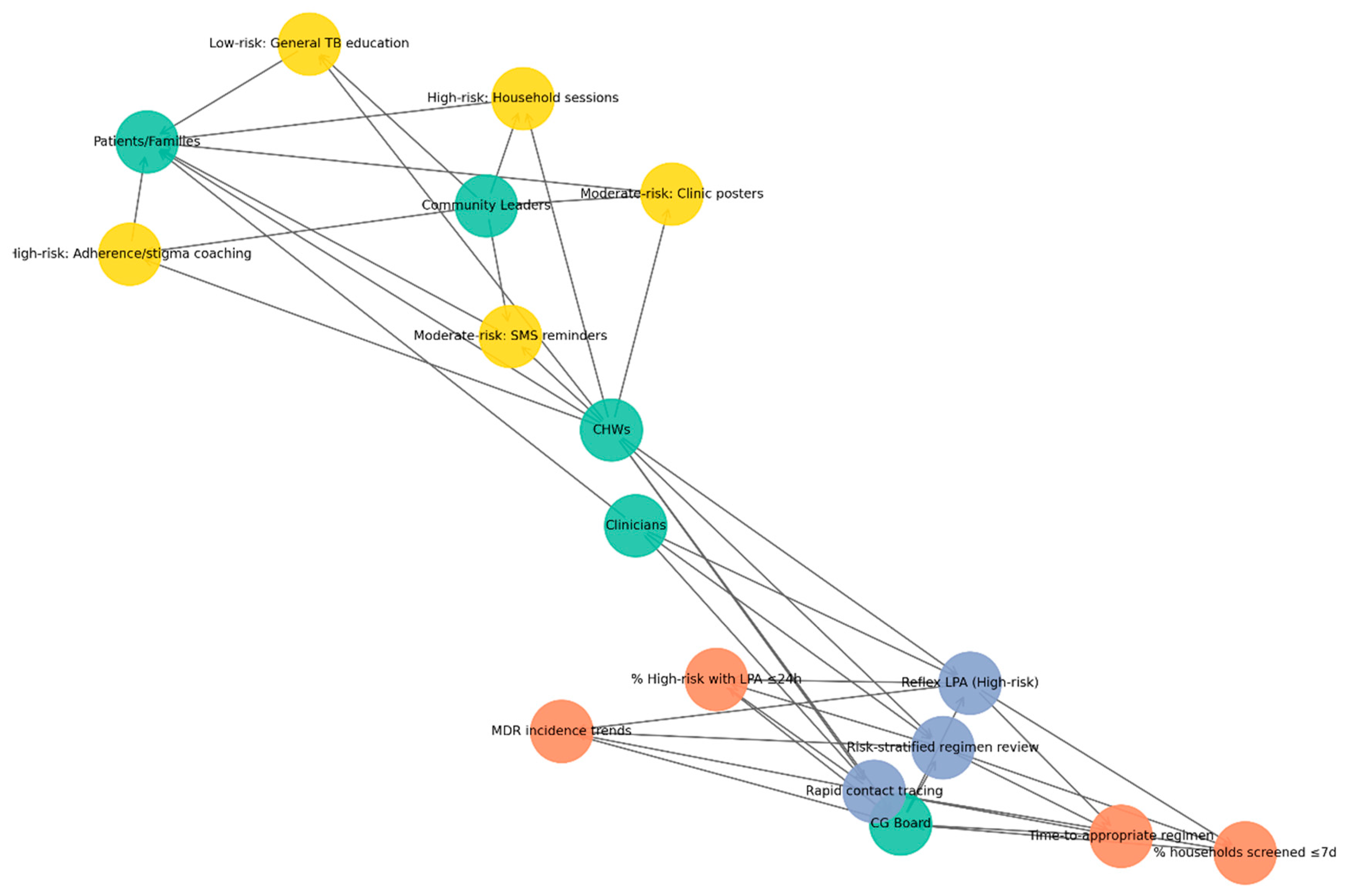
Figure 5 illustrates how CHWs serve as central agents in delivering risk-banded TB education. Green nodes represent actors (CHWs, clinicians, patients/families, and community leaders or peer educators), yellow nodes denote interventions stratified by risk: high-risk households (household sessions, stigma reduction, adherence coaching), moderate-risk groups (SMS/WhatsApp reminders, clinic posters), and low-risk groups (general TB education). In contrast, orange nodes represent outcomes (improved adherence, reduced stigma, increased household screening uptake). Directed edges depict flows of delivery, support, and reinforcement: CHWs deliver interventions to patients and families, clinicians provide technical back-up, and community leaders reinforce education messages. Feedback loops are visible when patients report challenges to CHWs, who then engage leaders to mobilize responses.
The CEE subnetwork highlights the crucial role of CHWs in delivering targeted educational interventions across diverse risk levels. By linking clinicians, households, and community leaders, CHWs ensure that health messages are not only delivered but also contextualized and reinforced. This configuration strengthens patient adherence, reduces stigma, and promotes timely household screening, demonstrating how community engagement complements clinical governance in sustaining TB control.
The figure illustrates a network model linking risk-tiered TB interventions (yellow nodes: low-, moderate-, and high-risk strategies) with community and health-system actors (green nodes: community health workers, clinicians, community leaders, patients/families) and expected outcomes (orange nodes: reduced stigma, improved adherence, increased household screening uptake). Interventions such as SMS reminders, clinic posters, household education sessions, stigma-reduction activities, general TB education, and adherence coaching are positioned centrally, demonstrating how different risk-stratified actions mobilize various stakeholders to achieve targeted public health effects. The network highlights the multi-sectoral nature of TB control and the central role of community-based approaches in strengthening prevention, adherence, and screening uptake.
Green nodes represent key actors, including clinicians, CHWs, patients and families, and community leaders. Blue nodes denote CG SOPs, such as reflex LPAs for high-risk cases, risk-stratified regimen reviews, and rapid contact tracing. Orange nodes indicate KPIs, including the proportion of high-risk patients receiving LPAs within 24 h, time to initiation of the appropriate regimen, household screening coverage within seven days, and monitoring of MDR incidence trends. Yellow nodes highlight CEE interventions tailored by risk-band, with high-risk households receiving targeted sessions and adherence/stigma coaching, moderate-risk groups supported through SMS reminders and clinic posters, and low-risk individuals provided with general TB education. Directed edges capture accountability, implementation, and feedback loops between governance structures, health workers, and communities, with CHWs emerging as pivotal bridging agents linking governance oversight with community-based interventions.
The systems map (
Figure 6,
Table 6) illustrates the interconnected roles of CG and CEE in strengthening TB control. Nodes were grouped into four categories: actors (clinicians, CHWs, the CG Board/M&M committee, patients and families, and community leaders or peer educators); processes (SOPs) on the CG side (reflex LPAs for high-risk cases, risk-stratified regimen reviews, and rapid contact tracing); KPIs (proportion of high risk patients with LPAs within 24 h, time to appropriate regimen initiation, household screening coverage within seven days, and monitoring of MDR incidence trends); and CEE interventions stratified by risk-band (high risk households receiving targeted sessions, stigma reduction and adherence coaching; moderate-risk groups supported through SMS/WhatsApp reminders and clinic posters; and low-risk individuals provided with general TB education).
Directed edges represent accountability, operational, and feedback relationships. Accountability edges link the CG Board to SOPs and KPIs, with feedback loops returning to clinicians and CHWs. Operational edges connect clinicians and CHWs through bi-directional risk communication, while community edges capture CHWs delivering education to households and peer educators engaging with community leaders. Feedback edges further demonstrate CHWs’ reporting barriers and adherence challenges, which are ultimately returned to the CG Board. Overall, the network underscores CHWs as pivotal bridging agents, integrating governance oversight with community-based action and sustaining accountability loops between policy, clinical practice, and patient education.
Figure 6.
Integration of CG and CEE in TB control.
Figure 6.
Integration of CG and CEE in TB control.
This system map (
Figure 6) illustrates the dynamic relationships between key actors, CG structures, and CEE interventions within the TB care continuum. The framework shows how CG processes such as oversight, quality assurance, adherence monitoring, and coordination of care interact with community-centred educational strategies, including outreach, peer-led learning, and household-level engagement. Together, these components create a feedback-driven system designed to strengthen early detection, treatment adherence, patient support, and public health responsiveness in high-burden, resource-limited settings.
Table 6.
Summary of nodes included in the network analysis, organized by category.
Table 6.
Summary of nodes included in the network analysis, organized by category.
| Category |
Nodes |
| Actor |
Clinicians |
| Actor |
CHWs |
| Actor |
CG Board |
| Actor |
Patients/Families |
| Actor |
Community Leaders |
| CG SOP |
Reflex LPA (High risk) |
| CG SOP |
Risk-stratified regimen review |
| CG SOP |
Rapid contact tracing |
| KPI |
% High risk with LPA ≤24h |
| KPI |
Time-to-appropriate regimen |
| KPI |
% households screened ≤7d |
| KPI |
MDR incidence trends |
| CEE intervention |
High risk: Household sessions |
| CEE intervention |
High risk: Adherence/stigma coaching |
| CEE intervention |
Moderate risk: SMS reminders |
| CEE intervention |
Moderate risk: Clinic posters |
| CEE intervention |
Low risk: General TB education |