Submitted:
14 February 2026
Posted:
26 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Cell Culture
2.3. Preparation of Cell Lysates and Western Blot Analysis
2.4. RNA Extraction and RT‒PCR
2.5. Cell Cycle Analysis
2.6. Immunofluorescence
2.7. Immunophenotyping
2.8. Clonogenic Assay
2.9. Statistical Analysis
3. Results and Discussion
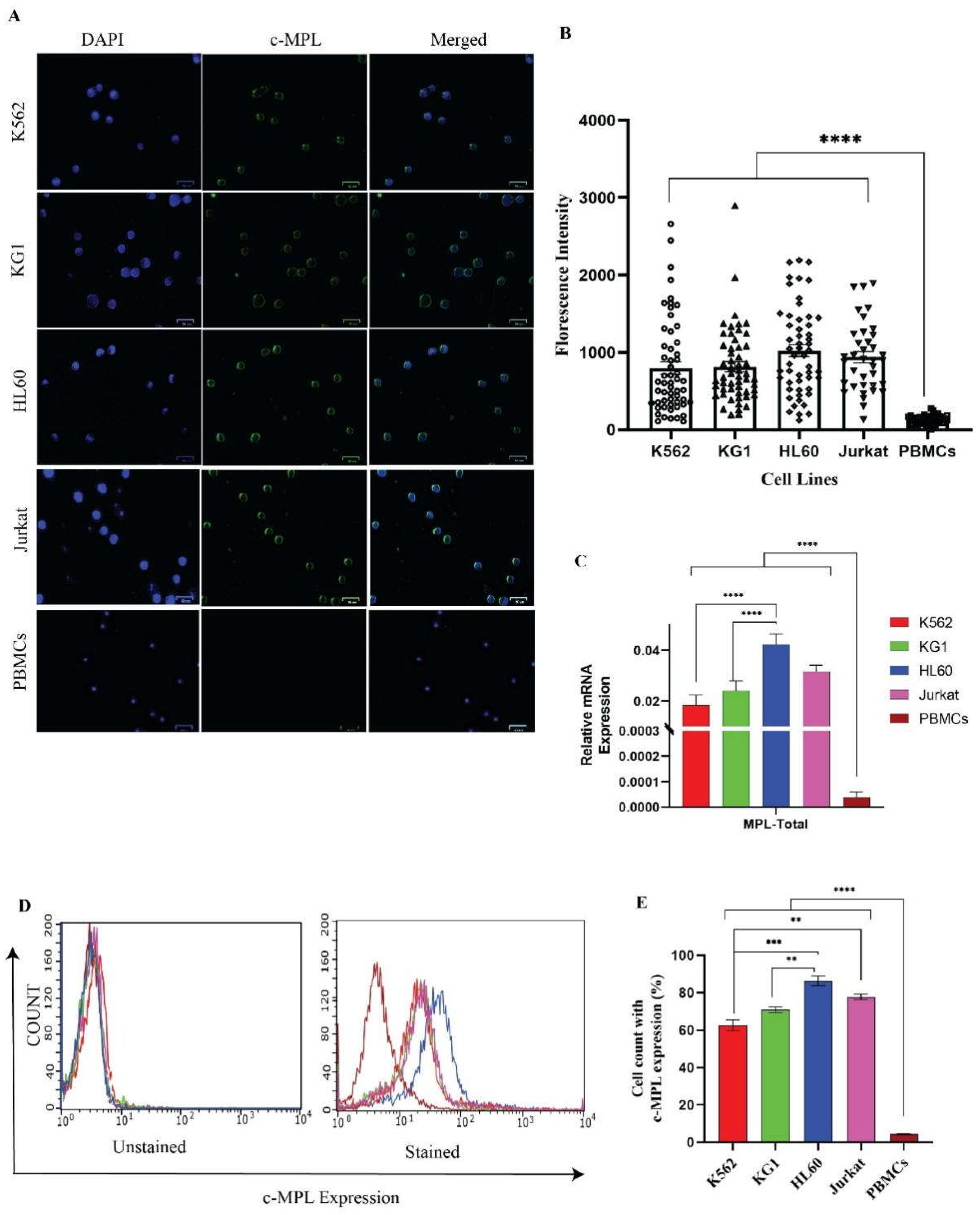
3.1. Over-Expression of Total c-MPL in Different Leukaemic Cell Lines Compared to WT Cells
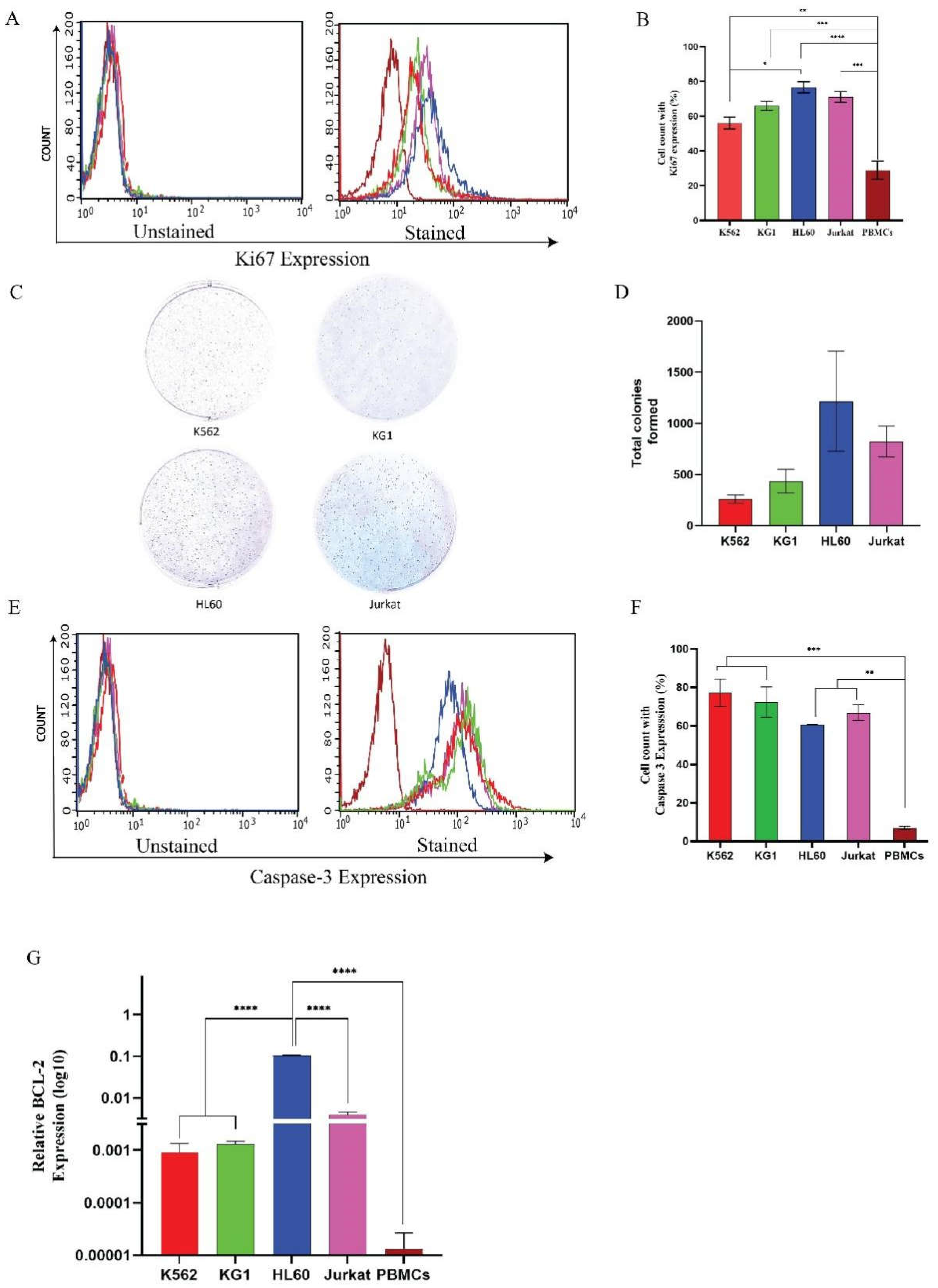
3.2. Different Leukaemia Cell Lines Exhibit Different Levels of Severity Even with Overexpression of c-MPL
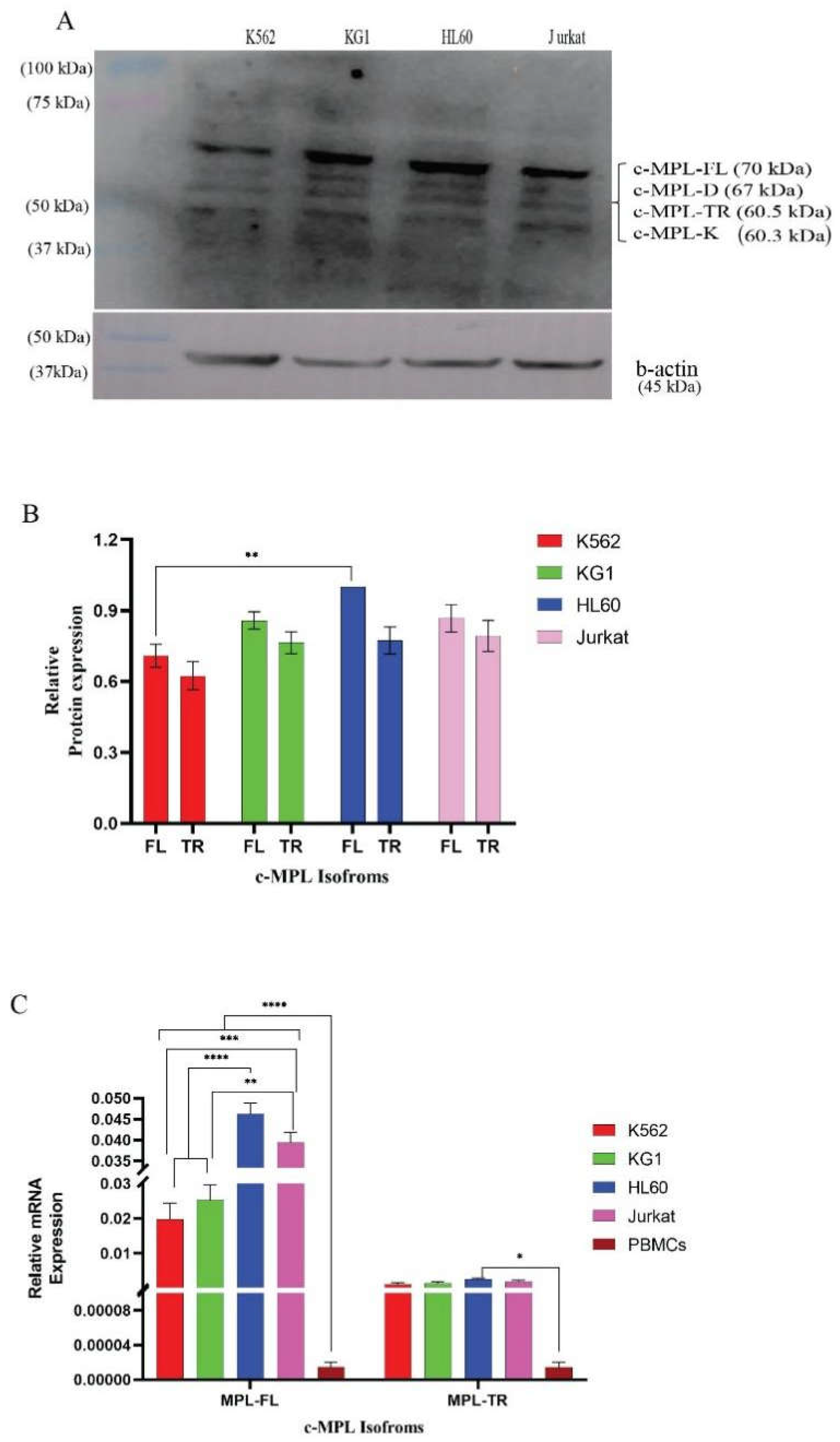
3.3. The Severe Leukaemia Conditions Show an Increase in MPL-FL Isoforms
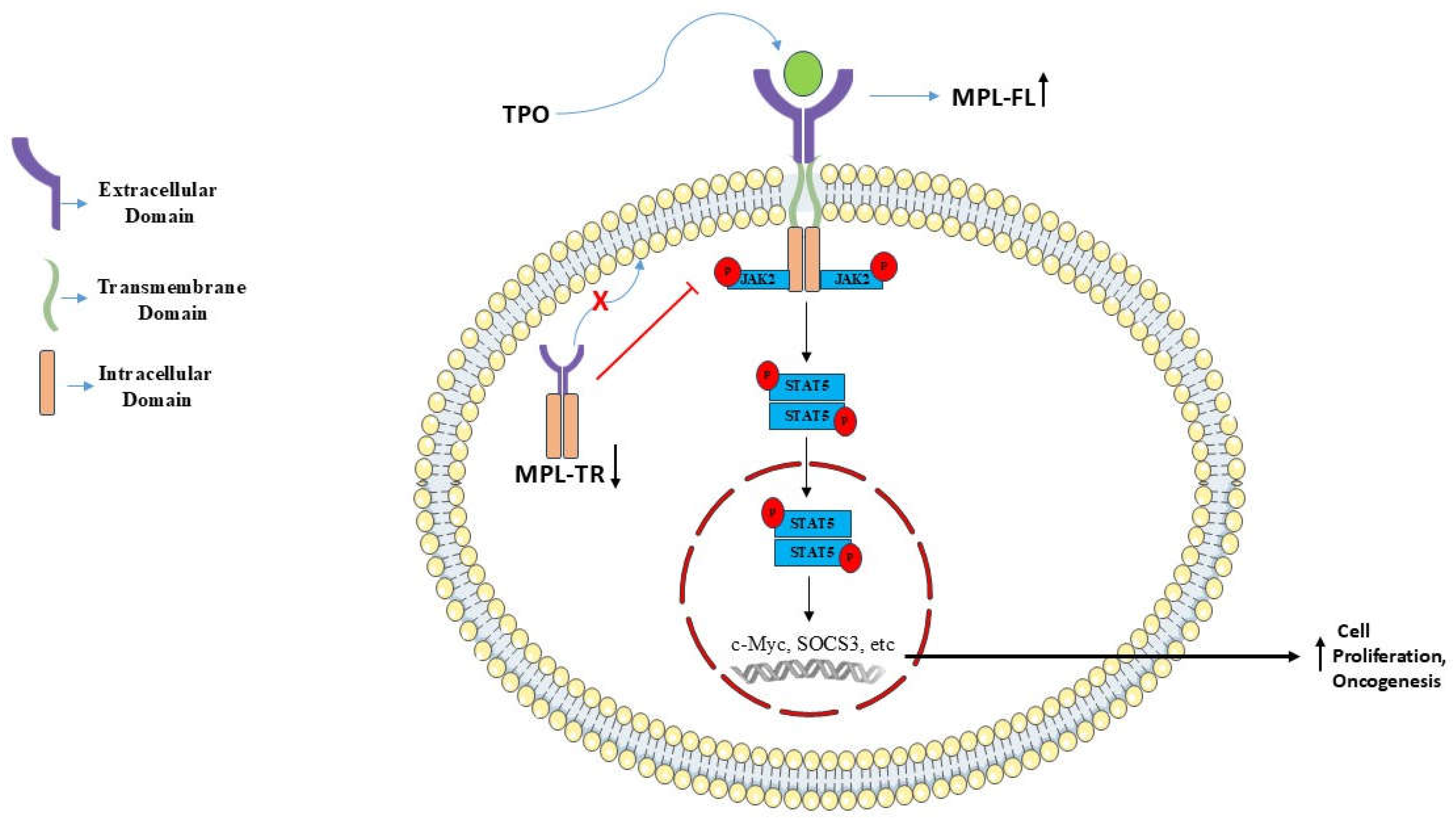
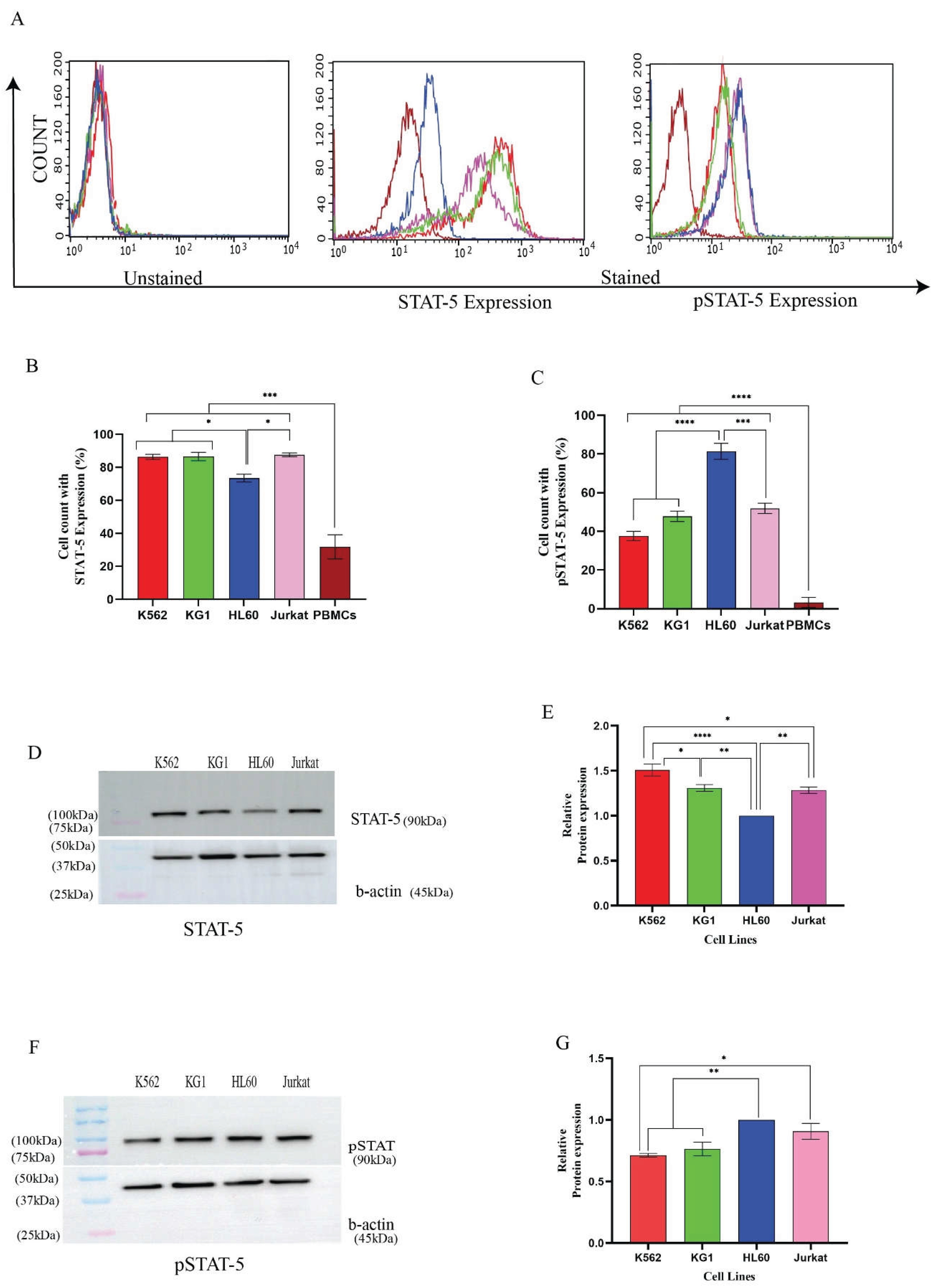
3.4. Overexpression of c-MPL-FL Exacerbates Leukemic Conditions Through Activation of the STAT Pathway
4. Discussion and Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Nemkov, T; D’Alessandro, A; Reisz, JA. Metabolic underpinnings of leukemia pathology and treatment. Cancer Reports 2019, 2(2), e1139. [Google Scholar] [CrossRef] [PubMed]
- Kondo, M. Lymphoid and myeloid lineage commitment in multipotent hematopoietic progenitors. Immunological reviews 2010, 238(1), 37–46. [Google Scholar] [CrossRef] [PubMed]
- Li, H; Zhao, N; Li, Y; Xing, H; Chen, S; Xu, Y; Tang, K; Tian, Z; Wang, M; Rao, Q; Wang, J. c-MPL is a candidate surface marker and confers self-renewal, quiescence, chemotherapy resistance, and leukemia initiation potential in leukemia stem cells. Stem Cells 2018, 36(11), 1685–96. [Google Scholar] [CrossRef]
- Ng, A.P.; Kauppi, M.; Metcalf, D.; Hyland, C.D.; Josefsson, E.C.; Lebois, M.; Zhang, J.G.; Baldwin, T.M.; Di Rago, L.; Hilton, D.J.; Alexander, W.S. Mpl expression on megakaryocytes and platelets is dispensable for thrombopoiesis but essential to prevent myeloproliferation. Proceedings of the National Academy of Sciences 2014, 111(16), 5884–5889. [Google Scholar] [CrossRef]
- Hitchcock, I.S.; Hafer, M.; Sangkhae, V.; Tucker, J.A. The thrombopoietin receptor: revisiting the master regulator of platelet production. Platelets 2021, 32(6), 770–778. [Google Scholar] [CrossRef] [PubMed]
- Hussain, MA; Das, SP; Kulkarni, M; Laha, S. A review on the functional characteristics of the c-Myeloproliferative Leukaemia (c-MPL) gene and its isoforms. Cellular Oncology 2024, 47(5), 1607–26. [Google Scholar] [CrossRef]
- Pikman, Y.; Lee, B.H.; Mercher, T.; McDowell, E.; Ebert, B.L.; Gozo, M.; Cuker, A.; Wernig, G.; Moore, S.; Galinsky, I.; DeAngelo, D.J. MPLW515L is a novel somatic activating mutation in myelofibrosis with myeloid metaplasia. PLoS medicine 2006, 3(7), e270. [Google Scholar] [CrossRef]
- Ding, J.; Komatsu, H.; Wakita, A.; Kato-Uranishi, M.; Ito, M.; Satoh, A.; Tsuboi, K.; Nitta, M.; Miyazaki, H.; Iida, S.; Ueda, R. Familial essential thrombocythemia associated with a dominant-positive activating mutation of the c-MPL gene, which encodes for the receptor for thrombopoietin. Blood 2004, 103(11), 4198–4200. [Google Scholar] [CrossRef]
- Wang, T; Wang, Z; Yang, R. Thrombopoietic growth factors in the treatment of immune thrombocytopenic purpura. Critical reviews in oncology/hematology 2011, 77(3), 172–83. [Google Scholar] [CrossRef]
- Duensing, S.; Duensing, A.; Meran, J.G.; Kreft, A.; Büsche, G.; Ganser, A.; Georgii, A. Molecular detection of c-MPL thrombopoietin receptor gene expression in chronic myeloproliferative disorders. Molecular Pathology 1999, 52(3), 146. [Google Scholar] [CrossRef]
- Bridgford, J.L.; Lee, S.M.; Lee, C.M.; Guglielmelli, P.; Rumi, E.; Pietra, D.; Wilcox, S.; Chhabra, Y.; Rubin, A.F.; Cazzola, M.; Vannucchi, A.M. Novel drivers and modifiers of MPL-dependent oncogenic transformation identified by deep mutational scanning. Blood, The Journal of the American Society of Hematology 2020, 135(4), 287–292. [Google Scholar] [CrossRef]
- Chen, Y.; Xie, J.; Shen, Z.; Shi, J.; Chen, S.; Wang, G. Clinical and molecular characteristics of acute myeloid leukemia with MPL mutation. Hematology 2022, 27(1), 530–534. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Chen, Z.; Jiang, Y.; Qiu, X.; Zhao, X. Different mutations of the human c-MPL gene indicate distinct haematopoietic diseases. Journal of Hematology & Oncology 2013, 6, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Yu, P.; Qiu, S.W.; Rao, Q.; Lin, D.; Xing, H.Y.; Tang, K.J.; Tian, Z.; Wang, M.; Wang, J.X. Expression of c-MPL in leukemic stem cells from acute myeloid patients. Zhongguo shi yan xue ye xue za zhi 2012, 20(5), 1052–1055. [Google Scholar]
- Albitar, M.; Manshouri, T.; Kantarjian, H.; Keating, M.; Estrov, Z.; Faber, J.; Freireich, E.J.; Pierce, S.; Estey, E. Correlation between lower c-MPL protein expression and favorable cytogenetic groups in acute myeloid leukemia. Leukemia research 1999, 23(1), 63–69. [Google Scholar] [CrossRef]
- Wang, J.Y.; Yu, P.; Chen, S.; Xing, H.; Chen, Y.; Wang, M.; Tang, K.; Tian, Z.; Rao, Q.; Wang, J. Activation of Rac1 GTPase promotes leukemia cell chemotherapy resistance, quiescence and niche interaction. Molecular oncology 2013, 7(5), 907–916. [Google Scholar] [CrossRef] [PubMed]
- Klairmont, MM; Cheng, J; Schwartzberg, L; Ho, HH; Gradowski, JF. Chronic myeloid leukemia, BCR-ABL1-positive with CALR and MPL mutations. International Journal of Laboratory Hematology 2018, 40(3). [Google Scholar] [CrossRef]
- Zhang, B; Li, L; Ho, Y; Li, M; Marcucci, G; Tong, W; Bhatia, R. Heterogeneity of leukemia-initiating capacity of chronic myelogenous leukemia stem cells. The Journal of clinical investigation 2016, 126(3), 975–91. [Google Scholar] [CrossRef]
- Iijima, K; Sugita, K; Inukai, T; Goi, K; Tezuka, T; Uno, K; Sato, H; Kagami, K; Nakazawa, S. Expression of thrombopoietin receptor and its functional role in human B-precursor leukemia cells with 11q23 translocation or Philadelphia chromosome. Leukemia 2000, 14(9), 1598–605. [Google Scholar] [CrossRef]
- Yamazaki, H; Nishida, H; Iwata, S; Dang, NH; Morimoto, C. CD90 and CD110 correlate with cancer stem cell potentials in human T-acute lymphoblastic leukemia cells. Biochemical and biophysical research communications 2009, 383(2), 172–7. [Google Scholar] [CrossRef]
- Katheeja, M.N.; Das, S.P.; Das, R.; Laha, S. BRCA1 interactors, RAD50 and BRIP1, as prognostic markers for triple-negative breast cancer severity. Frontiers in Genetics 2023, 14, 1035052. [Google Scholar] [CrossRef]
- Adel, A.; Djida, A.A.; Hamid, N.; Lila, B.H.; Souhil, T.; Abderrezak, G.; Idir, B. TRIzol-Based RNA extraction for detection protocol for SARS-CoV-2 of the coronavirus Disease 2019. In New Microbes and New Infections.
- Laboratory Techniques in Cellular and Molecular Medicine; Hajdúch, M., Drábek, J., Eds.; Palacky University Olomouc, 2021. [Google Scholar]
- Yoshihara, H.; Arai, F.; Hosokawa, K.; Hagiwara, T.; Takubo, K.; Nakamura, Y.; Gomei, Y.; Iwasaki, H.; Matsuoka, S.; Miyamoto, K.; Miyazaki, H. Thrombopoietin/MPL signaling regulates hematopoietic stem cell quiescence and interaction with the osteoblastic niche. Cell stem cell 2007, 1(6), 685–697. [Google Scholar] [CrossRef]
- Maktoof, S.A.; Al-Ammar, N.S.; Al-Kashwan, T.A.; Jumaa, A.K.; Bader, P.; Kreyenberg, H. The expressions of myeloid differentiation and non-lineage specific differentiation antigens among FAB subtypes of acute myeloid leukemia in Iraqi patients. Romanian medical JouRnal 2024, 71(4). [Google Scholar] [CrossRef]
- Jensen, H.A.; Yourish, H.B.; Bunaciu, R.P.; Varner, J.D.; Yen, A. Induced myelomonocytic differentiation in leukemia cells is accompanied by noncanonical transcription factor expression. FEBS open bio 2015, 5, 789–800. [Google Scholar] [CrossRef] [PubMed]
- Skopek, R.; Palusińska, M.; Kaczor-Keller, K.; Pingwara, R.; Papierniak-Wyglądała, A.; Schenk, T.; Lewicki, S.; Zelent, A.; Szymański, Ł. Choosing the right cell line for acute myeloid leukemia (AML) research. International journal of molecular sciences 2023, 24(6), 5377. [Google Scholar] [CrossRef]
- Borowicz, S; Van Scoyk, M; Avasarala, S; Rathinam, MK; Tauler, J; Bikkavilli, RK; Winn, RA. The soft agar colony formation assay. Journal of visualized experiments: JoVE 2014, 27(92), 51998. [Google Scholar]
- Wu, Y.; Zhao, B.; Chen, X.; Geng, X.; Zhang, Z. Circ_0009910 sponges miR-491-5p to promote acute myeloid leukemia progression through modulating B4GALT5 expression and PI3K/AKT signaling pathway. International journal of laboratory hematology 2022, 44(2), 320–332. [Google Scholar] [CrossRef]
- Salavatipour, MS; Kouhbananinejad, SM; Lashkari, M; Bardsiri, MS; Moghadari, M; Kashani, B; Farsinejad, A; Vahidi, R. Kermanian propolis induces apoptosis through upregulation of Bax/Bcl-2 ratio in acute myeloblastic leukemia cell line (NB4). Journal of Cancer Research and Therapeutics 2023, 19(2), 327–34. [Google Scholar] [CrossRef] [PubMed]
- Mehta, SV; Shukla, SN; Vora, HH. Overexpression of Bcl-2 protein predicts chemoresistance in acute myeloid leukemia: its correlation with FLT3. Neoplasma 2013, 60(6), 666–75. [Google Scholar] [CrossRef]
- Coers, J; Ranft, C; Skoda, RC. A truncated isoform of c-MPL with an essential C-terminal peptide targets the full-length receptor for degradation. Journal of Biological Chemistry 2004, 279(35), 36397–404. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Bunting, K.D. STAT5 in hematopoietic stem cell biology and transplantation. Jak-stat 2013, 2(4), e27159. [Google Scholar] [CrossRef] [PubMed]
- Kaushansky, K. The molecular mechanisms that control thrombopoiesis. The Journal of clinical investigation 2005, 115(12), 3339–3347. [Google Scholar] [CrossRef]
- Xiao, N.; Laha, S.; Das, S.P.; Morlock, K.; Jesneck, J.L.; Raffel, G.D. Ott1 (Rbm15) regulates thrombopoietin response in hematopoietic stem cells through alternative splicing of c-MPL. Blood, The Journal of the American Society of Hematology 2015, 125(6), 941–948. [Google Scholar] [CrossRef]
- Spivak, J.L.; Moliterno, A.R. The thrombopoietin receptor, MPL, is a therapeutic target of opportunity in the MPN. Frontiers in Oncology 2021, 11, 641613. [Google Scholar] [CrossRef] [PubMed]
- Skoda, R.C.; Seldin, D.C.; Chiang, M.K.; Peichel, C.L.; Vogt, T.F.; Leder, P. Murine c-mpl: a member of the hematopoietic growth factor receptor superfamily that transduces a proliferative signal. The EMBO journal 1993, 12(7), 2645–2653. [Google Scholar] [CrossRef] [PubMed]
| S. No | Antibody | Purpose | Concentration | Company | Secondary Antibody concentration |
| 1 | c-MPL | Western Blotting | 1:1000 | Invitrogen | 1:2000 |
| 2 | STAT5 | Western Blotting | 1:1000 | CST | 1:2000 |
| 3 | p-STAT5 | Western Blotting | 1:1000 | CST | 1:2000 |
| 4 | c-MPL | Immunofluorescence | 1:200 | Invitrogen | 1:250 |
| 5 | c-MPL | Immunophenotyping | 1:200 | Invitrogen | 1:250 |
| 6 | Caspase | Immunophenotyping | 1:200 | CST | 1:250 |
| 7 | STAT | Immunophenotyping | 1:200 | CST | 1:250 |
| 8 | p-STAT | Immunophenotyping | 1:200 | CST | 1:250 |
| 9 | CD44 | Immunophenotyping | 1:100 | Abcam | Conjugated |
| 10 | Ki67 | Immunophenotyping | 1:200 | Invitrogen | 1:250 |
| Primers | Position | Sequence |
| c-MPL-FL (MPL-P) | Forward Primer | 5′ GCGATCTCGCTACCGTTTAC 3′ |
| Reverse Primer | 5′ AGGAAACTGCCACCTCAGC 3′ | |
| c-MPL -TR (MPL-S) | Forward Primer | 5′ AGGACTGGAAGGAGAC 3′ |
| Reverse Primer | 5′ TCAGGCTGCAGTGTCCTAAG 3′ | |
| c-MPL -Total | Forward Primer | 5′ GAGAAGCTTCAGCTCTGAC 3′ |
| Reverse Primer | 5′ CAAGTGCCACTGCATCTCCA 3′ | |
| Beta-actin | Forward Primer | 5′ACTGGAACGGTGAAGGTGAC 3′ |
| Reverse Primer | 5′AGAGAAGTGGGGTGGCTTTT3′ | |
| BCL-2 | Forward Primer | 5′GACTGAGTACCTGAACCGGC 3′ |
| Reverse Primer | 5′GTTCCACAAAGGCATCCCAGC 3′ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





