Submitted:
14 February 2026
Posted:
26 February 2026
You are already at the latest version
Abstract
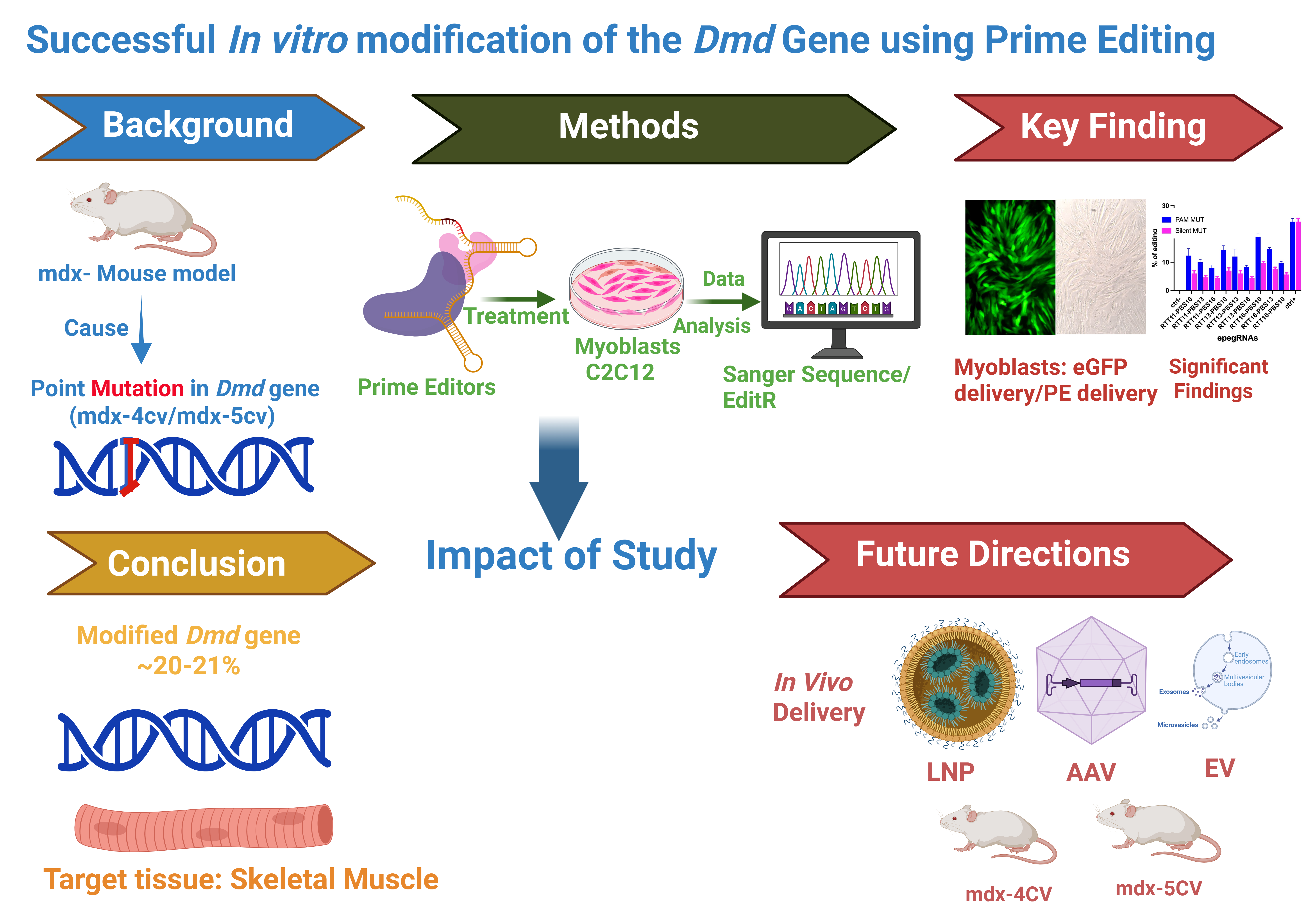
Keywords:
1. Introduction
2. Materials and Methods
2.1. Cell Lines
2.2. Plasmids
2.3. Cell Culture Conditions
2.4. Plasmid Electroporation in Mouse C2C12 Myoblasts
2.5. Genomic DNA Extraction and PCR Amplification
2.6. Sanger Sequencing and Editing Analysis
2.7. Statistical Analysis
3. Results
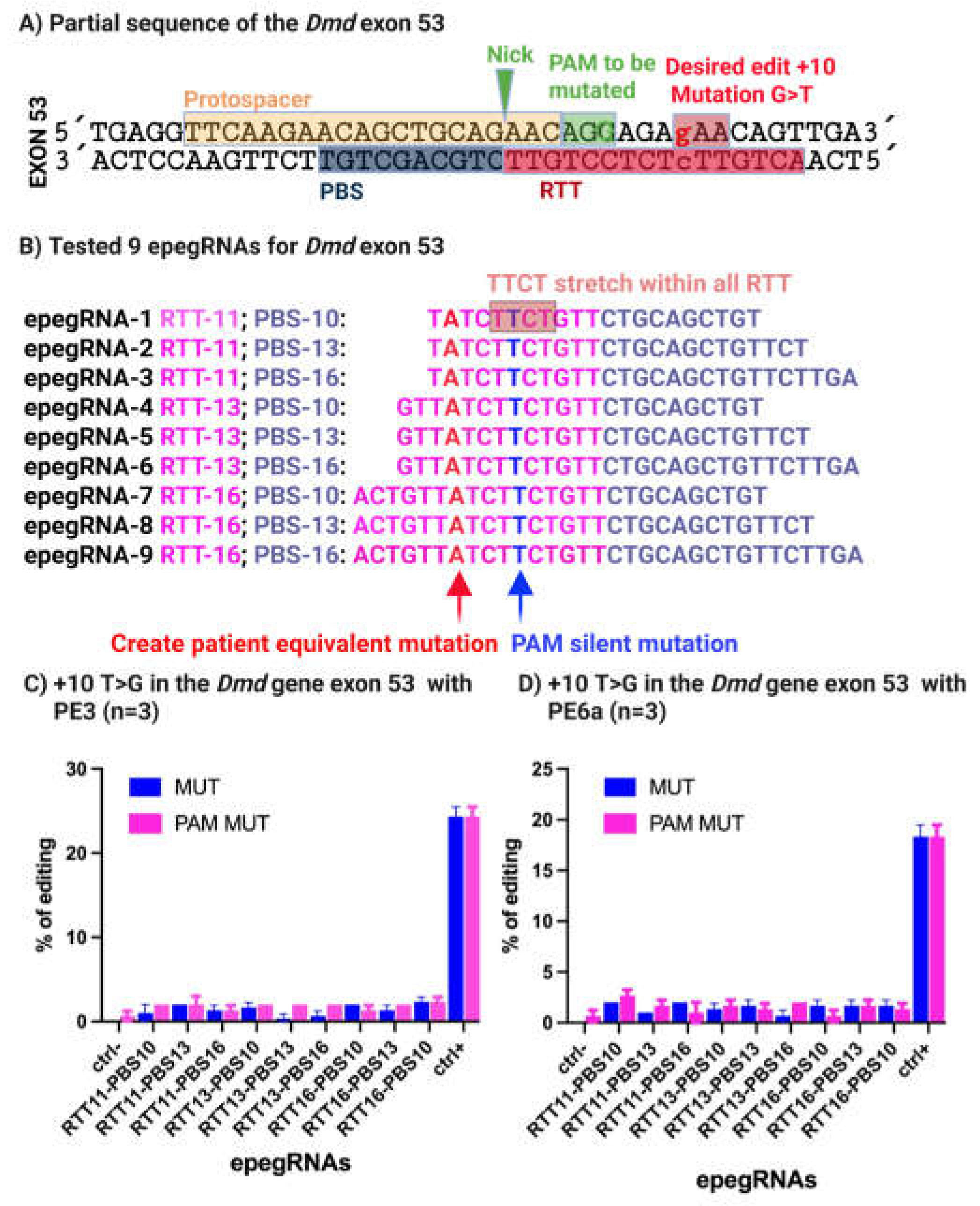
3.1. Installation of the 4cv Nonsense Mutation in Exon 53 of the Dmd Gene in C2C12 Myoblasts Using Prime Editing
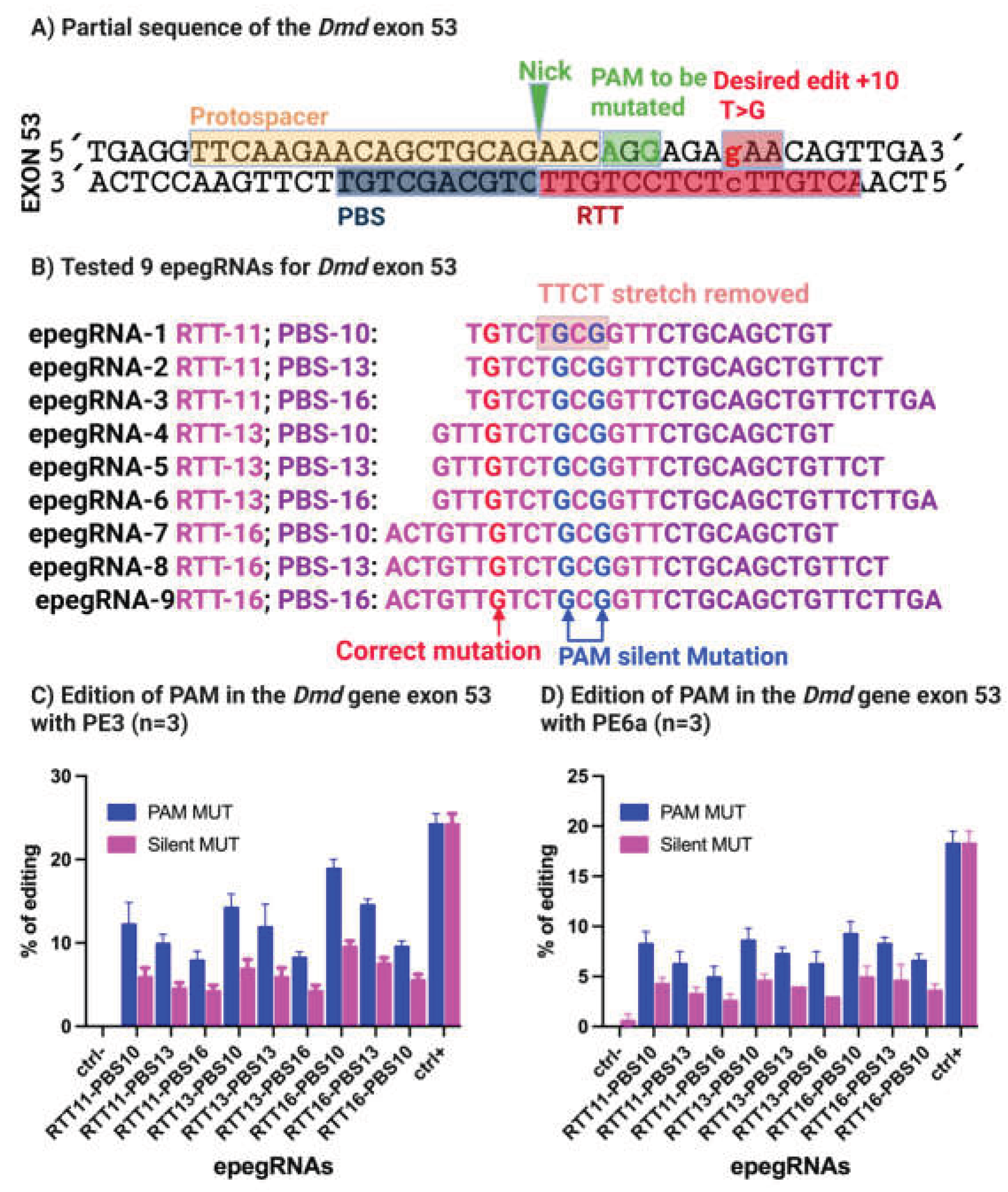
3.2. Optimization of epegRNAs by Removal of the TTCT Stretch Enables Efficient Correction of the 4cv Mutation In Vitro
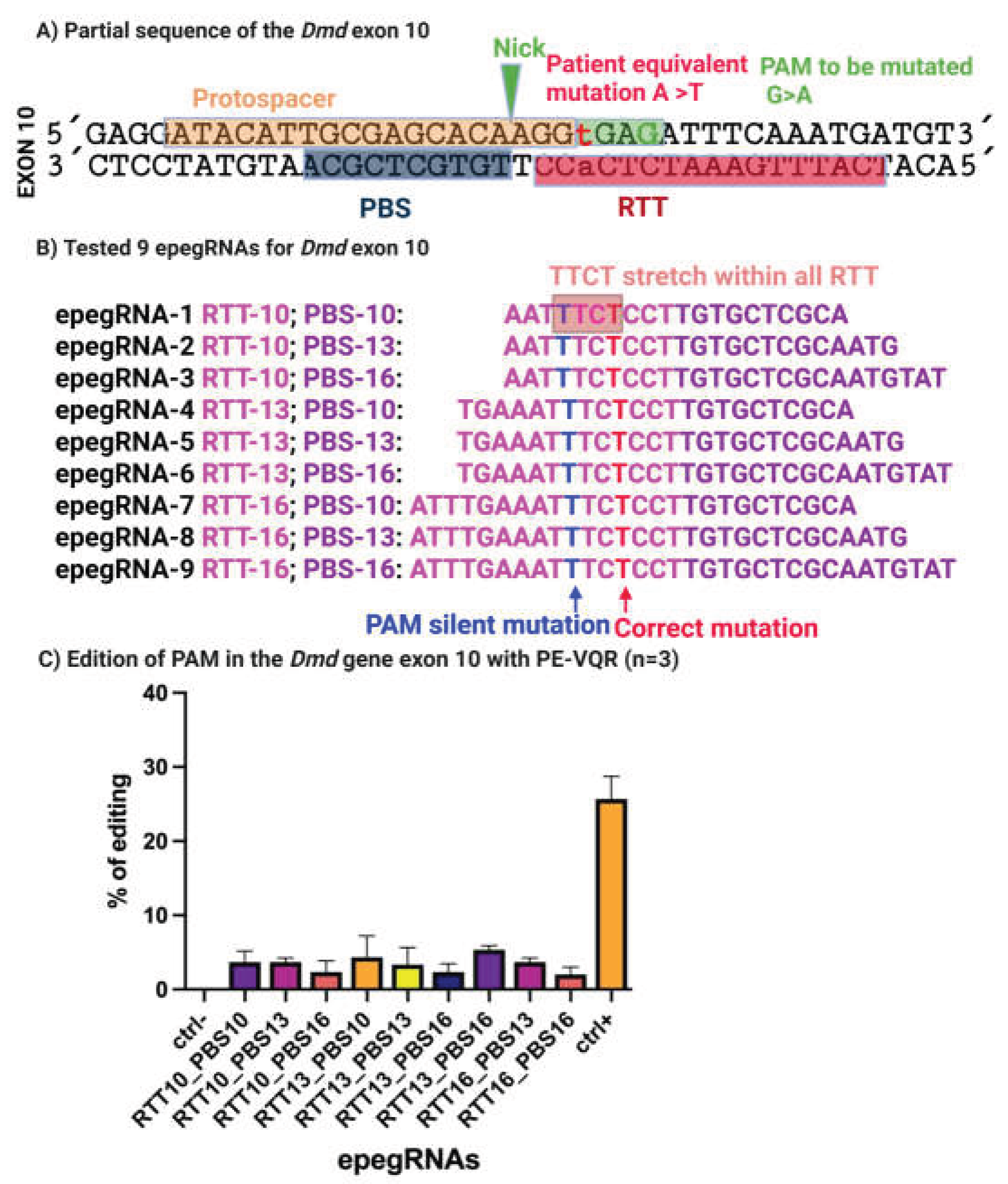
3.3. In Vitro Prime Editing of the 5cv Mutation Using an NGAG PAM
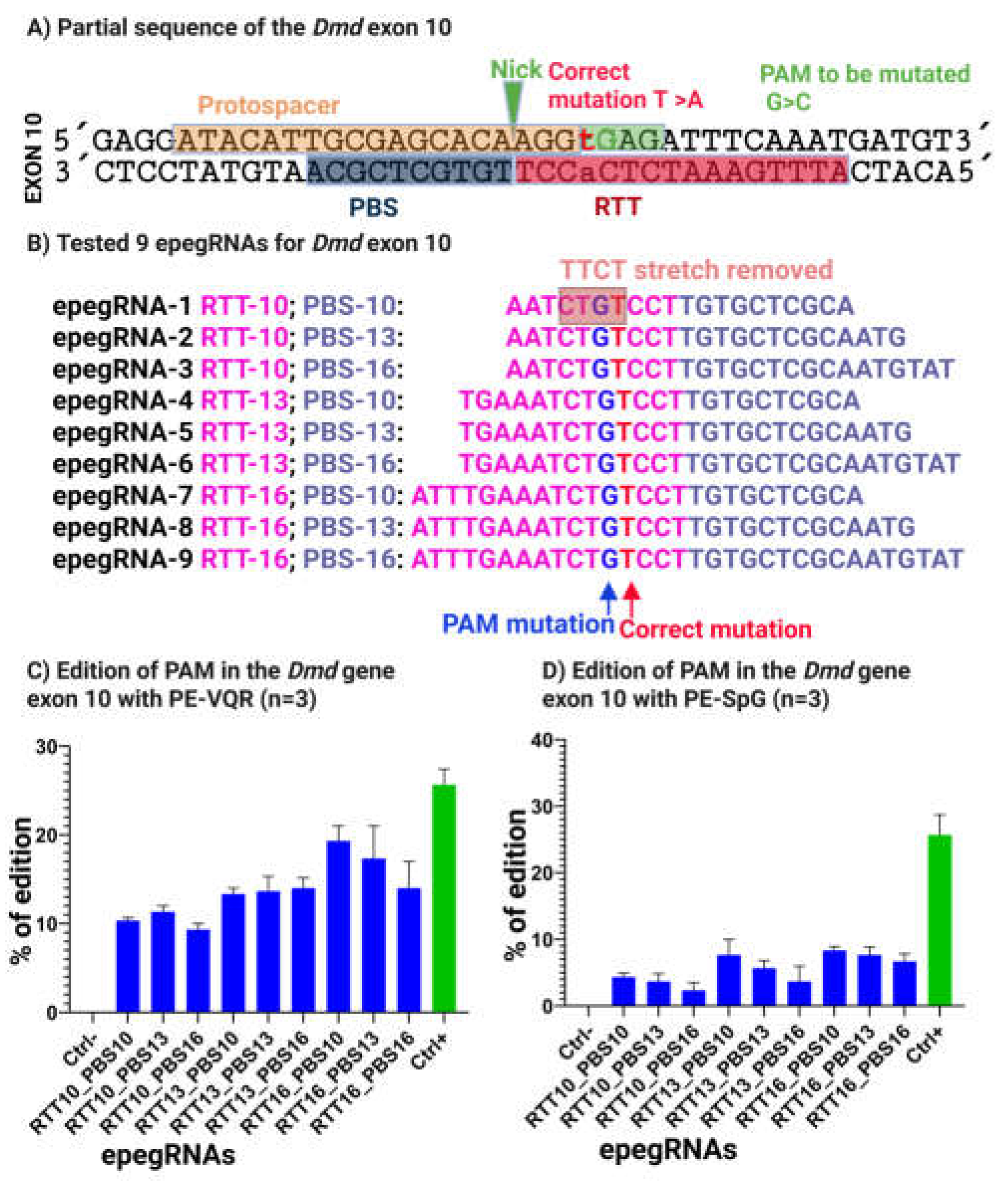
3.4. Removal of the TTCT Stretch Enhances Prime Editing Efficiency for Correction of the 5cv Mutation In Vitro
4. Discussion
Prime Editing Efficiency Is Highly Sensitive to RTT–PBS Configuration
Identification of a Cryptic Transcription Termination Constraint in epegRNAs
Prime Editing of the mdx-5cv Mutation Highlights the Importance of PAM Expansion and Guide Optimization
5. Implications for Therapeutic Prime Editing of DMD
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Ginn, S.L., et al., Gene therapy clinical trials worldwide to 2017: An update. The journal of gene medicine, 2018. 20(5): p. e3015.
- Doudna, J.A. and E. Charpentier, The new frontier of genome engineering with CRISPR-Cas9. Science, 2014. 346(6213): p. 1258096. [CrossRef]
- Porteus, M.H., A new class of medicines through DNA editing. New England Journal of Medicine, 2019. 380(10): p. 947-959. [CrossRef]
- Villiger, L., et al., CRISPR technologies for genome, epigenome and transcriptome editing. Nature Reviews Molecular Cell Biology, 2024. 25(6): p. 464-487. [CrossRef]
- Pacesa, M., O. Pelea, and M. Jinek, Past, present, and future of CRISPR genome editing technologies. Cell, 2024. 187(5): p. 1076-1100. [CrossRef]
- Urnov, F.D., et al., Genome editing with engineered zinc finger nucleases. Nature Reviews Genetics, 2010. 11(9): p. 636-646. [CrossRef]
- Sun, N. and H. Zhao, Transcription activator-like effector nucleases (TALENs): a highly efficient and versatile tool for genome editing. Biotechnology and bioengineering, 2013. 110(7): p. 1811-1821. [CrossRef]
- Sander, J.D. and J.K. Joung, CRISPR-Cas systems for editing, regulating and targeting genomes. Nature biotechnology, 2014. 32(4): p. 347-355. [CrossRef]
- Komor, A.C., et al., Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature, 2016. 533(7603): p. 420-424. [CrossRef]
- Gaudelli, N.M., et al., Programmable base editing of A• T to G• C in genomic DNA without DNA cleavage. Nature, 2017. 551(7681): p. 464-471. [CrossRef]
- Kurt, I.C., et al., CRISPR C-to-G base editors for inducing targeted DNA transversions in human cells. Nature biotechnology, 2021. 39(1): p. 41-46. [CrossRef]
- Anzalone, A.V., et al., Search-and-replace genome editing without double-strand breaks or donor DNA. Nature, 2019. 576(7785): p. 149-157. [CrossRef]
- Anzalone, A.V., L.W. Koblan, and D.R. Liu, Genome editing with CRISPR–Cas nucleases, base editors, transposases and prime editors. Nature biotechnology, 2020. 38(7): p. 824-844. [CrossRef]
- Nelson, J.W., et al., Engineered pegRNAs improve prime editing efficiency. Nature biotechnology, 2022. 40(3): p. 402-410. [CrossRef]
- EP, H., Dystrophin: the protein product of the Duchenne muscular dystrophy locus. Cell, 1987. 51: p. 919-928.
- Ryder, S., et al., The burden, epidemiology, costs and treatment for Duchenne muscular dystrophy: an evidence review. Orphanet journal of rare diseases, 2017. 12(1): p. 79. [CrossRef]
- Okubo, M., et al., Genetic diagnosis of Duchenne/Becker muscular dystrophy using next-generation sequencing: validation analysis of DMD mutations. Journal of human genetics, 2016. 61(6): p. 483-489.
- Ricotti, V., et al., NorthStar Clinical Network. Long-term benefits and adverse effects of intermittent versus daily glucocorticoids in boys with Duchenne muscular dystrophy. J Neurol Neurosurg Psychiatry, 2013. 84(6): p. 698-705. [CrossRef]
- Kourakis, S., et al., Standard of care versus new-wave corticosteroids in the treatment of Duchenne muscular dystrophy: Can we do better? Orphanet Journal of Rare Diseases, 2021. 16(1): p. 117. [CrossRef]
- Gloss, D., et al., Practice guideline update summary: Corticosteroid treatment of Duchenne muscular dystrophy [RETIRED] Report of the Guideline Development Subcommittee of the American Academy of Neurology. Neurology, 2016. 86(5): p. 465-472.
- Birnkrant, D.J., et al., Diagnosis and management of Duchenne muscular dystrophy, part 2: respiratory, cardiac, bone health, and orthopaedic management. The Lancet Neurology, 2018. 17(4): p. 347-361.
- Hoffman, E.P., et al., Vamorolone trial in Duchenne muscular dystrophy shows dose-related improvement of muscle function. Neurology, 2019. 93(13): p. e1312-e1323. [CrossRef]
- Wilton-Clark, H. and T. Yokota, Antisense and gene therapy options for Duchenne muscular dystrophy arising from mutations in the N-terminal hotspot. Genes, 2022. 13(2): p. 257. [CrossRef]
- Komaki, H., et al., Systemic administration of the antisense oligonucleotide NS-065/NCNP-01 for skipping of exon 53 in patients with Duchenne muscular dystrophy. Science Translational Medicine, 2018. 10(437): p. eaan0713.
- Aartsma-Rus, A., et al., Theoretic applicability of antisense-mediated exon skipping for Duchenne muscular dystrophy mutations. Human mutation, 2009. 30(3): p. 293-299. [CrossRef]
- Yucel, N., et al., Humanizing the mdx mouse model of DMD: the long and the short of it. NPJ Regenerative medicine, 2018. 3(1): p. 4.
- Im, W.B., et al., Differential expression of dystrophin isoforms in strains of mdx mice with different mutations. Human molecular genetics, 1996. 5(8): p. 1149-1153. [CrossRef]
- Beastrom, N., et al., mdx5cv mice manifest more severe muscle dysfunction and diaphragm force deficits than do mdx mice. The American journal of pathology, 2011. 179(5): p. 2464-2474.
- Murray, J.B., P.T. Harrison, and J. Scholefield, Prime editing: therapeutic advances and mechanistic insights. Gene Therapy, 2025. 32(2): p. 83-92. [CrossRef]
- Fu, Y., et al., Prime editing: current advances and therapeutic opportunities in human diseases. Science bulletin, 2023. 68(24): p. 3278-3291.
- Kluesner, M.G., et al., EditR: a method to quantify base editing from Sanger sequencing. The CRISPR journal, 2018. 1(3): p. 239-250. [CrossRef]
- Ui-Tei, K., S. Maruyama, and Y. Nakano, Enhancement of single guide RNA transcription for efficient CRISPR/Cas-based genomic engineering. Genome, 2017. 60(6): p. 537-545. [CrossRef]
- Doman, J.L., et al., Designing and executing prime editing experiments in mammalian cells. Nature protocols, 2022. 17(11): p. 2431-2468.
- Anders, C., K. Bargsten, and M. Jinek, Structural plasticity of PAM recognition by engineered variants of the RNA-guided endonuclease Cas9. Molecular cell, 2016. 61(6): p. 895-902.
- Walton, R.T., et al., Unconstrained genome targeting with near-PAMless engineered CRISPR-Cas9 variants. Science, 2020. 368(6488): p. 290-296.
- Arechavala-Gomeza, V., et al., Revertant fibres and dystrophin traces in Duchenne muscular dystrophy: implication for clinical trials. Neuromuscular Disorders, 2010. 20(5): p. 295-301. [CrossRef]
- Happi Mbakam, C., et al., Prime editing strategies to mediate exon skipping in DMD gene. Frontiers in Medicine, 2023. 10: p. 1128557.
- Wang, Q., et al., Selection-free precise gene repair using high-capacity adenovector delivery of advanced prime editing systems rescues dystrophin synthesis in DMD muscle cells. Nucleic Acids Research, 2024. 52(5): p. 2740-2757.
- Long, C., et al., Postnatal genome editing partially restores dystrophin expression in a mouse model of muscular dystrophy. Science, 2016. 351(6271): p. 400-403.
- Nelson, C.E., et al., In vivo genome editing improves muscle function in a mouse model of Duchenne muscular dystrophy. Science, 2016. 351(6271): p. 403-407.
- Briner, A.E., et al., Guide RNA functional modules direct Cas9 activity and orthogonality. Molecular cell, 2014. 56(2): p. 333-339.
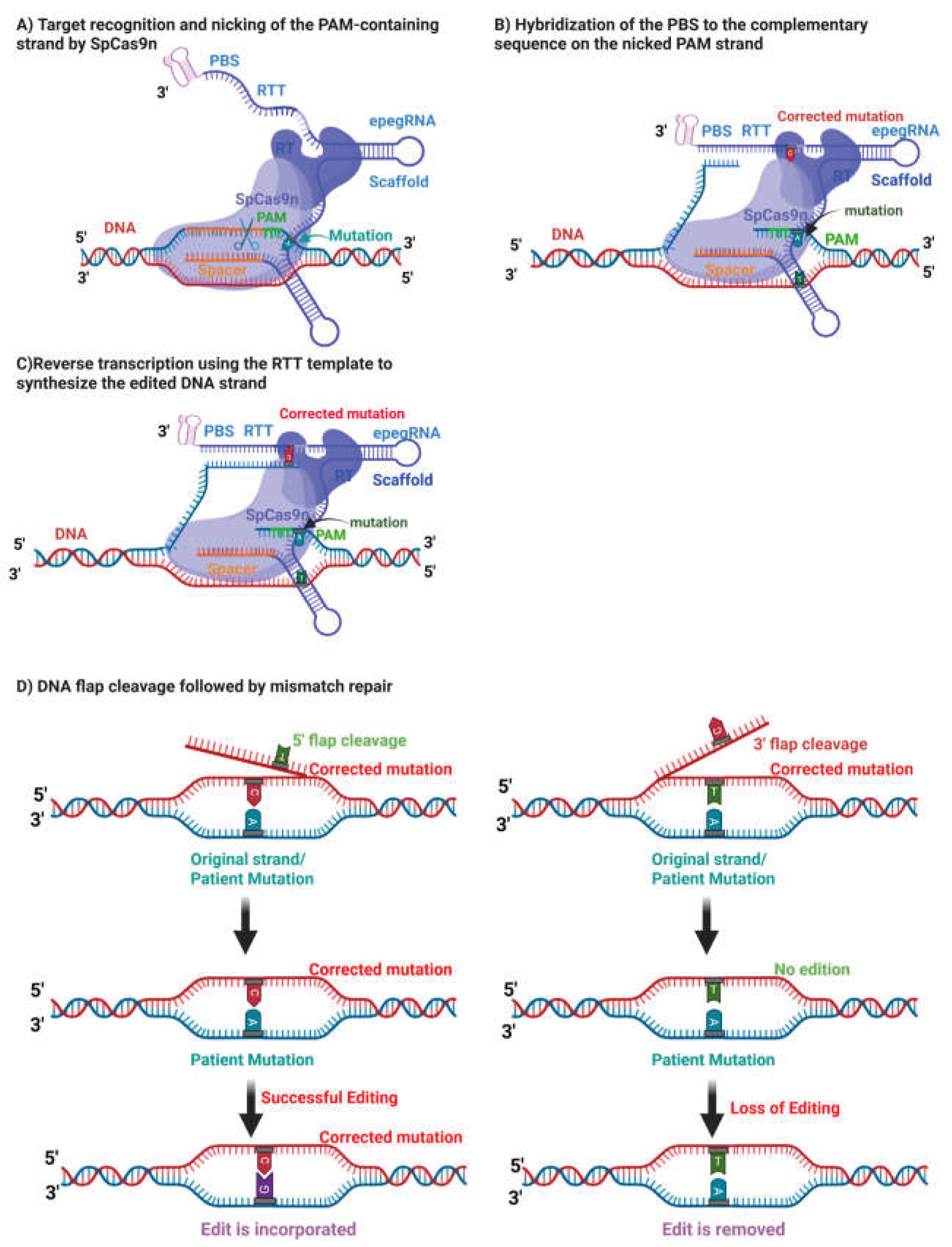
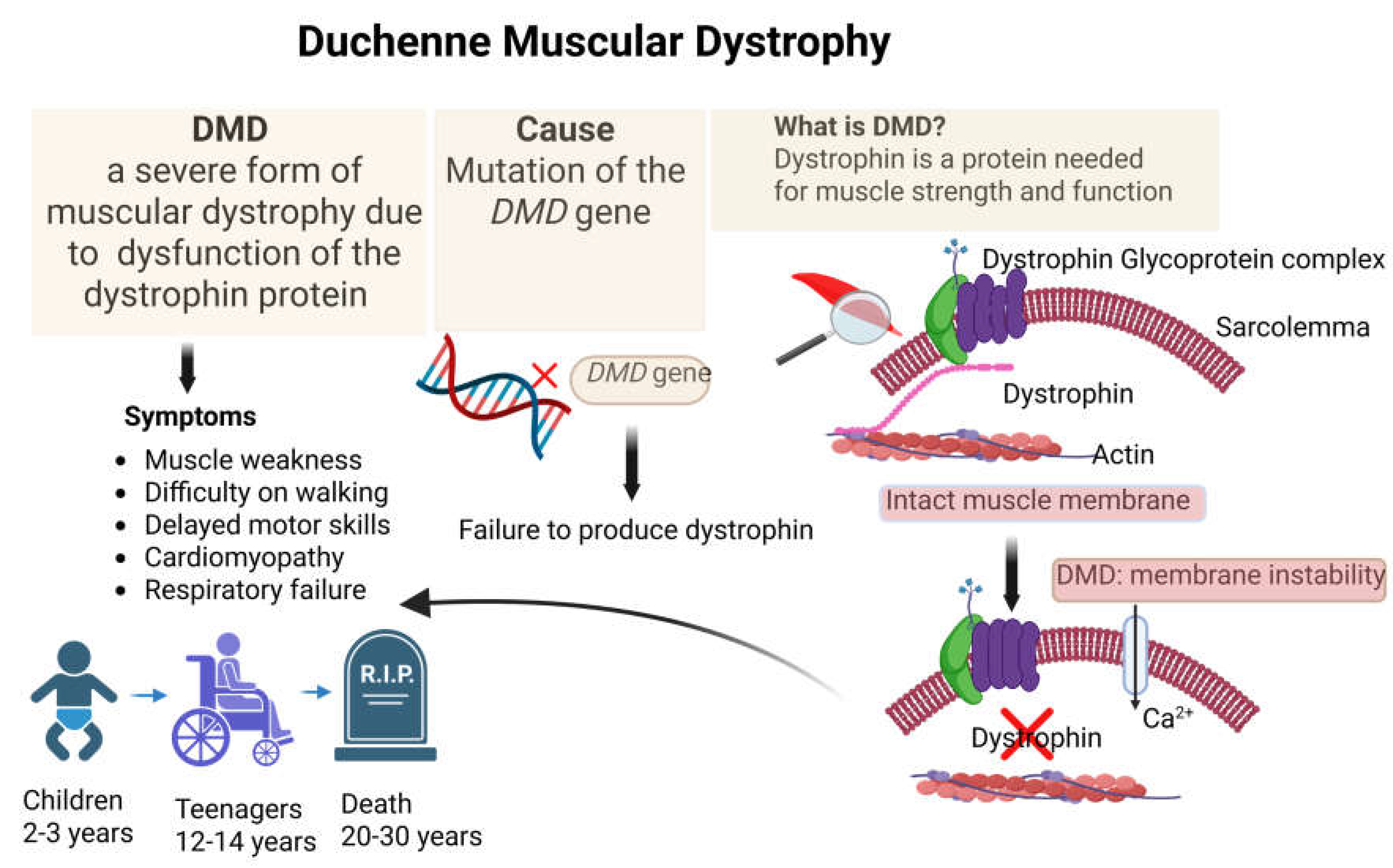
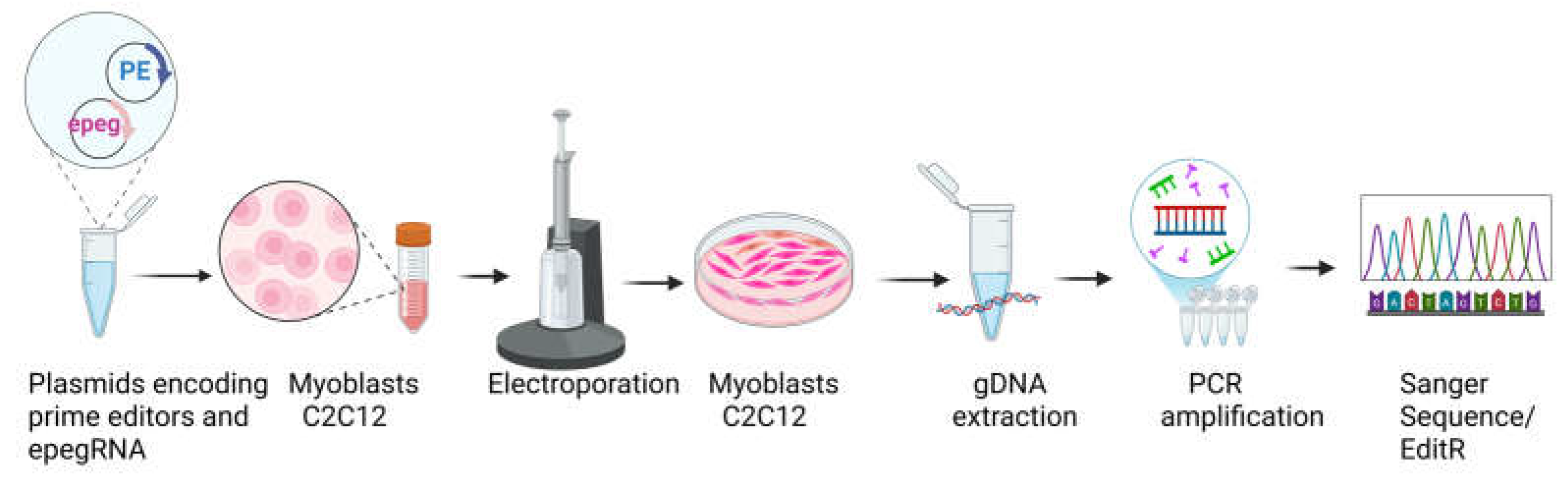
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



