Submitted:
15 February 2026
Posted:
16 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Method Development
2.1.1. Preparation of Peptide Analyte and Inhibitors
2.1.2. Standards and Working Solutions
2.1.3. Development of Chromatographic Conditions
2.1.4. Instrument
2.1.5. Development of Enzyme Assay Conditions
2.1.6. Chromatographic Conditions
2.2. Method Validation
2.2.1. Specificity
2.2.2. Linearity
2.2.3. Accuracy
2.2.4. Precision
2.2.5. Robustness
2.3. Commercial Inhibitor Assay
3. Results
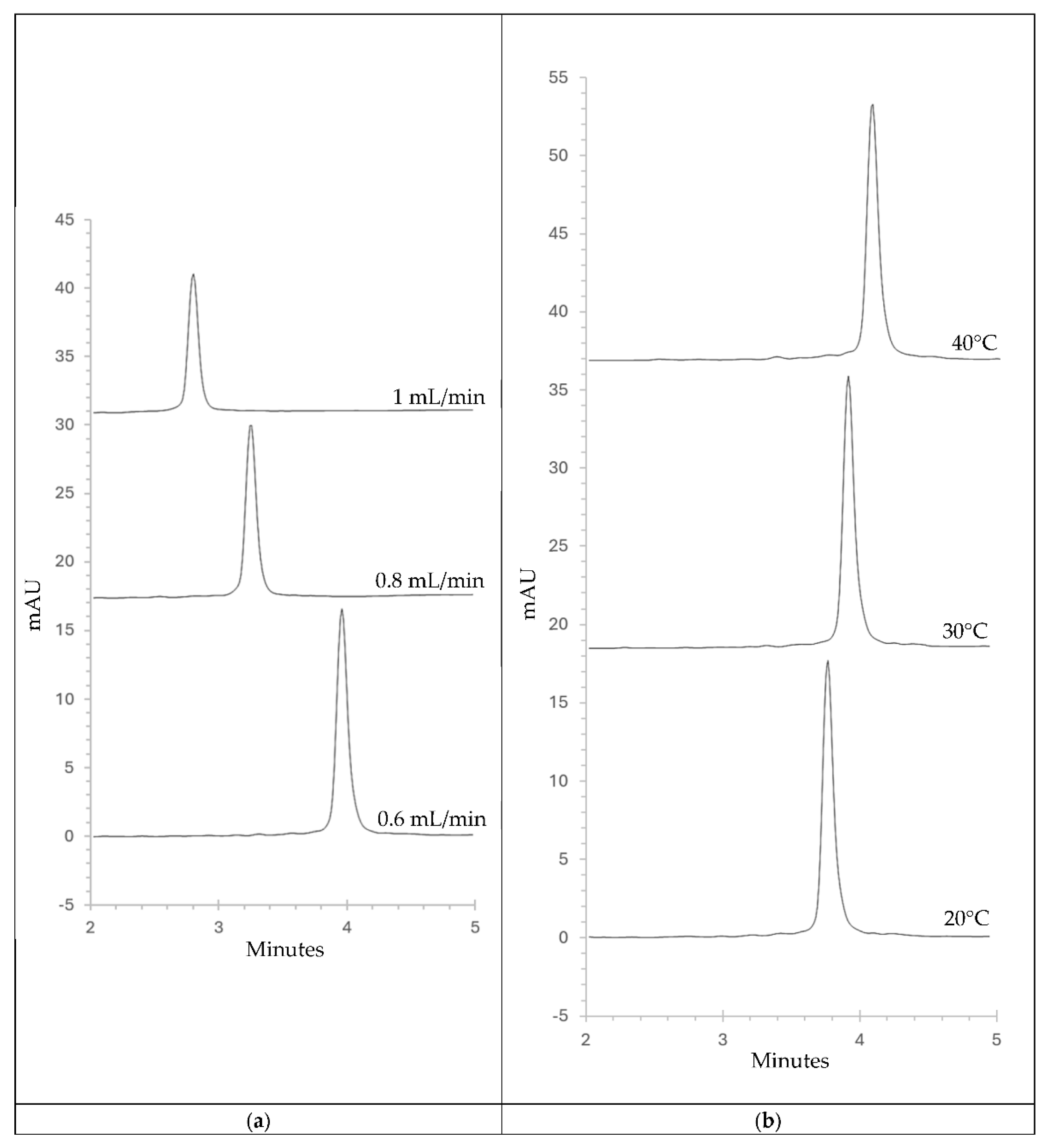
3.1. Method Development and Optimisation
3.2. Method Validation
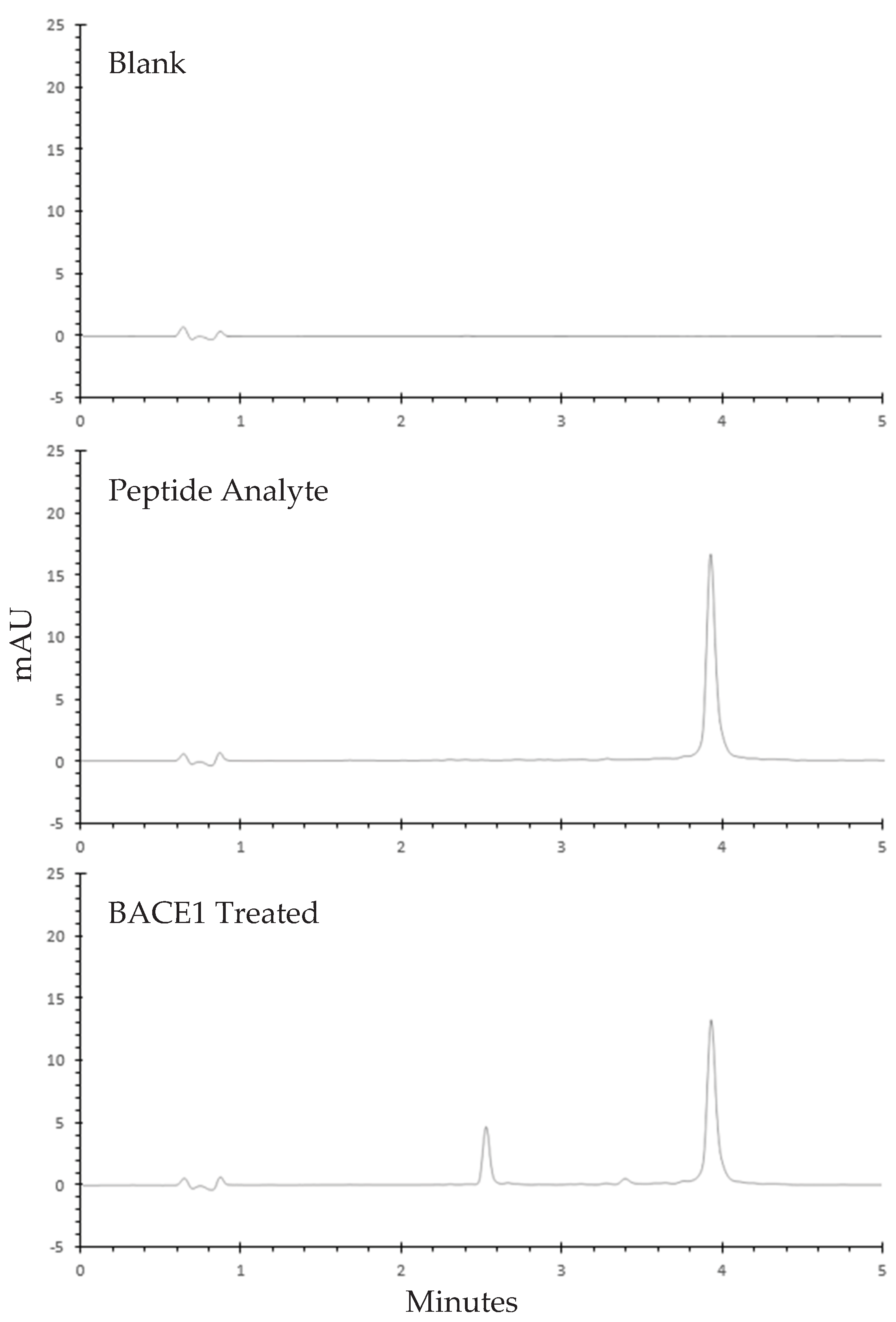
3.2.1. Specificity
3.2.2. Linearity
3.2.3. Accuracy
3.2.4. Precision
3.2.5. Robustness
3.3. Commercial Inhibitor Assay
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| HPLC | High-performance liquid chromatography |
| BACE1 | Beta-site amyloid precursor protein cleaving enzyme 1 |
| ICH | International council for harmonisation |
| TFA | Trifluoroacetic acid |
| RSD | Relative standard deviation |
| APP | Amyloid precursor protein |
| AD | Alzheimer’s disease |
| Aβ | β-amyloid |
| ELISA | Enzyme-linked immunosorbent assay |
| FIA | Fluorescence immunoassay |
| FRET | Fluorescence resonance energy transfer |
| SPR | Surface plasmon resonance |
| RP-HPLC | Reverse phase high-performance liquid chromatography |
| LC | Liquid chromatography |
| FA | Formic acid |
| PA | Phosphoric acid |
| LoD | Limit of detection |
| LoQ | Limit of quantification |
| SD | Standard deviation |
| QC | Quality control |
| DMSO | Dimethyl sulfoxide |
| tR | Retention time |
References
- Cho, Y; Bae, HG; Okun, E; Arumugam, TV; Jo, DG. Physiology and pharmacology of amyloid precursor protein. Pharmacol Ther. 2022, 235, 108122. [Google Scholar] [CrossRef] [PubMed]
- Yuksel, M; Tacal, O. Trafficking and proteolytic processing of amyloid precursor protein and secretases in Alzheimer’s disease development: An up-to-date review. European Journal of Pharmacology 2019, 856, 172415. [Google Scholar] [CrossRef] [PubMed]
- Rahman, MM; Lendel, C. Extracellular protein components of amyloid plaques and their roles in Alzheimer’s disease pathology. Molecular Neurodegeneration 2021, 16(1), 59. [Google Scholar] [CrossRef] [PubMed]
- Nandi, A; Counts, N; Chen, S; Seligman, B; Tortorice, D; Vigo, D; et al. Global and regional projections of the economic burden of Alzheimer’s disease and related dementias from 2019 to 2050: A value of statistical life approach. EClinicalMedicine 2022, 51, 101580. [Google Scholar] [CrossRef] [PubMed]
- Slack, MA; Gordon, SM. Protease Activity in Vascular Disease. Arterioscler Thromb Vasc Biol. 2019, 39(10), e210–e8. [Google Scholar] [CrossRef] [PubMed]
- Famutimi, OG; Adebiyi, VG; Akinmolu, BG; Dada, OV; Adewale, IO. Trypsin, chymotrypsin and elastase in health and disease. Future Journal of Pharmaceutical Sciences 2024, 10(1), 126. [Google Scholar] [CrossRef]
- Alhajj, M; Zubair, M; Farhana, A. Enzyme Linked Immunosorbent Assay. In StatPearls; StatPearls Publishing: Treasure Island (FL).
- Terzapulo, X; Kassenova, A; Bukasov, R. Immunoassays: Analytical and Clinical Performance, Challenges, and Perspectives of SERS Detection in Comparison with Fluorescent Spectroscopic Detection. Int J Mol Sci. 2024, 25(4). [Google Scholar] [CrossRef] [PubMed]
- Lim, J; Petersen, M; Bunz, M; Simon, C; Schindler, M. Flow cytometry based-FRET: basics, novel developments and future perspectives. Cell Mol Life Sci. 2022, 79(4), 217. [Google Scholar] [CrossRef] [PubMed]
- Fan, J; Wei, X; Dong, H; Zhang, Y; Zhou, Y; Xu, M; et al. Advancement in Analytical Techniques for Determining the Activity of β-Site Amyloid Precursor Protein Cleaving Enzyme 1. Crit Rev Anal Chem. 2024, 54(6), 1797–809. [Google Scholar] [CrossRef] [PubMed]
- Yi, X; Chen, H; He, Y; Wang, J. Assay of Biomarkers for Alzheimer’s Disease by Surface Plasmon Resonance. Journal of Analysis and Testing 2024, 8(3), 251–61. [Google Scholar] [CrossRef]
- Wiesner, R; Scheller, C; Krebs, F; Wätzig, H; Oltmann-Norden, I. A comparative study of CE-SDS, SDS-PAGE, and Simple Western: Influences of sample preparation on molecular weight determination of proteins. Electrophoresis 2021, 42(3), 206–18. [Google Scholar] [CrossRef] [PubMed]
- Halde, S; Mungantiwar, A; Chintamaneni, M. Simple, precise and accurate HPLC method of analysis for nevirapine suspension from human plasma. Indian J Pharm Sci. 2011, 73(4), 416–21. [Google Scholar] [CrossRef] [PubMed]
- Yi, X; Hao, Y; Xia, N; Wang, J; Quintero, M; Li, D; et al. Sensitive and continuous screening of inhibitors of β-site amyloid precursor protein cleaving enzyme 1 (BACE1) at single SPR chips. Anal Chem. 2013, 85(7), 3660–6. [Google Scholar] [CrossRef] [PubMed]
- Q2(R1); Validation of Analytical Procedures: Text and Methodology. 2005.
- Penwell, A; Sharp, K; Mansour, M; Sammatur, L. Development and validation of an HPLC/UV assay for separation and quantification of peptide antigens from a liposomal vaccine delivery platform. J Pharm Biomed Anal. 2012, 66, 176–82. [Google Scholar] [CrossRef] [PubMed]
- Pinto, I; Cerqueira, C; Freitas, Z; Santos, E; do Carmo, F; Junior, E. Development and validation of an analytical method using High Performance Liquid Chromatography (HPLC) to determine ethyl butylacetylaminopropionate in topical repellent formulations. Brazilian Journal of Pharmaceutical Sciences 2017, 53. [Google Scholar] [CrossRef]
- Thompson, M; Ellison, SLR; Wood, R. Harmonized guidelines for single-laboratory validation of methods of analysis (IUPAC Technical Report). Pure and Applied Chemistry 2002, 74(5), 835–55. [Google Scholar] [CrossRef]
- Salazar, J; Samhan-Arias, AK; Gutierrez-Merino, C. Hexa-Histidine, a Peptide with Versatile Applications in the Study of Amyloid-β(1–42) Molecular Mechanisms of Action. Molecules 2023, 28(20). [Google Scholar] [CrossRef] [PubMed]
- Moldoveanu, SC; David, V. Chapter 2 - Parameters that Characterize HPLC Analysis; Moldoveanu, SC, David, V, Eds.; Elsevier: Essentials in Modern HPLC Separations, 2013; pp. 53–83. [Google Scholar] [CrossRef]
- Ghosh, AK; Shin, D; Downs, D; Koelsch, G; Lin, X; Ermolieff, J; et al. Design of Potent Inhibitors for Human Brain Memapsin 2 (β-Secretase). J Am Chem Soc. 2000, 122(14), 3522–3. [Google Scholar] [CrossRef] [PubMed]
- Jeppsson, F; Eketjäll, S; Janson, J; Karlström, S; Gustavsson, S; Olsson, LL; et al. Discovery of AZD3839, a potent and selective BACE1 inhibitor clinical candidate for the treatment of Alzheimer disease. J Biol Chem. 2012, 287(49), 41245–57. [Google Scholar] [CrossRef]
| Parameter | Value |
|---|---|
| Slope | 1.5265 |
| Intercept | -5.213 |
| Coefficient of determination (r2) | 0.9997 |
| Coefficient of correlation (r) | 0.9998 |
| Linear range (µM) | 14.92 – 72 |
| Limit of detection (µM) | 2.40 |
| Limit of quantification (µM) | 7.26 |
| Curve no. or parameter | Slope | Intercept | Correlation Coefficient |
|---|---|---|---|
| 1 | 1.5512 | -4.9216 | 0.9998 |
| 2 | 1.4860 | -5.2224 | 0.9956 |
| 3 | 1.5446 | -6.2621 | 0.9998 |
| 4 | 1.5330 | -5.2946 | 0.9997 |
| 5 | 1.4711 | -3.2737 | 0.9997 |
| 6 | 1.5732 | -6.3036 | 0.9991 |
| Mean (n = 6) | 1.5265 | -5.2130 | 0.9990 |
| SD | 0.0397 | 0.0016 | |
| RSD (%) | 2.5979 | 0.1644 |
| QC Level (µM) | Recovery (%) | Mean Recovery (%) | RSD (%) |
|---|---|---|---|
| Low QC (25.21 µM) | 97.74 | 96.62 | 2.86 |
| 97.22 | |||
| 94.62 | |||
| 100.60 | |||
| 96.96 | |||
| 92.54 | |||
| Medium QC (42.60 µM) | 99.36 | 98.38 | 1.37 |
| 98.44 | |||
| 96.90 | |||
| 99.20 | |||
| 99.81 | |||
| 96.58 | |||
| High QC (72 µM) | 103.28 | 96.97 | 3.99 |
| 99.27 | |||
| 97.27 | |||
| 95.00 | |||
| 94.45 | |||
| 92.54 |
| Sample | Day 1 Recovery (µM)* | Day 2 Recovery (µM)** |
|---|---|---|
| 1 | 63.88 | 60.21 |
| 2 | 61.06 | 59.03 |
| 3 | 61.19 | 61.06 |
| 4 | 61.78 | 59.62 |
| 5 | 61.78 | 61.32 |
| 6 | 62.24 | 60.80 |
| Repeatability | ||
| Mean Recovery | 61.99 | 60.34 |
| SD | 1.02 | 0.89 |
| RSD (%) | 1.65 | 1.47 |
| Intermediate Precision | ||
| Mean Recovery | 61.17 | |
| SD | 1.25 | |
| RSD (%) | 2.05 |
| Parameter | Mean Recovery (µM) | % Change in Recovery | Mean Retention Time (min) | % Change in Retention Time |
|---|---|---|---|---|
| Temperature (30 °C) | ||||
| -10% | 59.93 | -0.12 | 3.97 | 1.60 |
| +10% | 59.84 | -0.27 | 3.90 | -0.15 |
| Flow Rate (0.6 mL/min) |
||||
| -10% | 67.77 | 12.95 | 4.07 | 4.25 |
| +10% | 54.62 | -8.96 | 3.73 | -4.48 |
| Sample | Analyte Remaining (µM) | Change in Activity (%) | t-Test (BACE1) |
|---|---|---|---|
| OM99-2 | 39.77 | -50.74 | 5.24E-07 |
| AZD3839 | 40.86 | -53.36 | 2.89E-06 |
| BACE1 | 22.40 | ||
| APP | 58.83 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




