Submitted:
12 February 2026
Posted:
14 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. Phenotypic Variation of Mixograph Parameters
2.2. QTL Mapping Results
2.3. Effect Analysis of Stable Loci
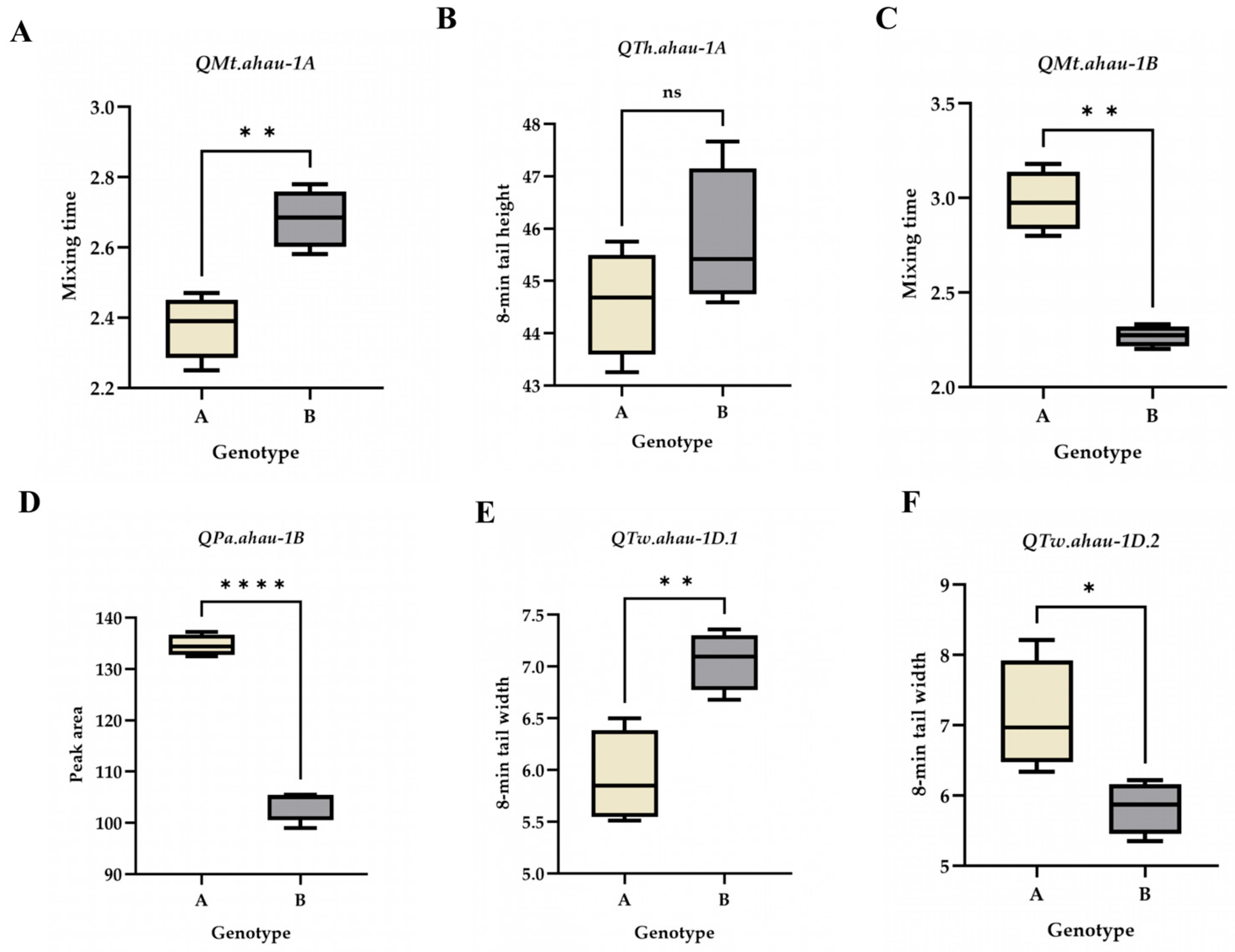
2.4. Development and Validation of KASP Markers
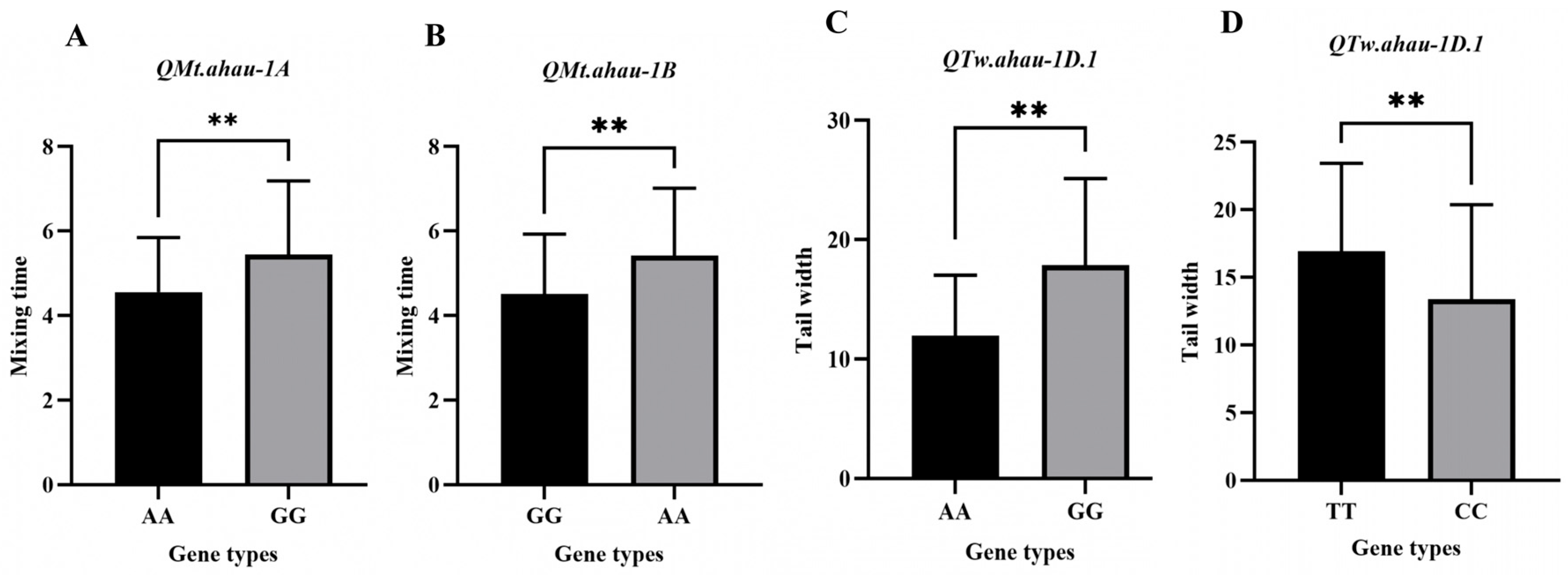
2.5. Prediction of Candidate Genes
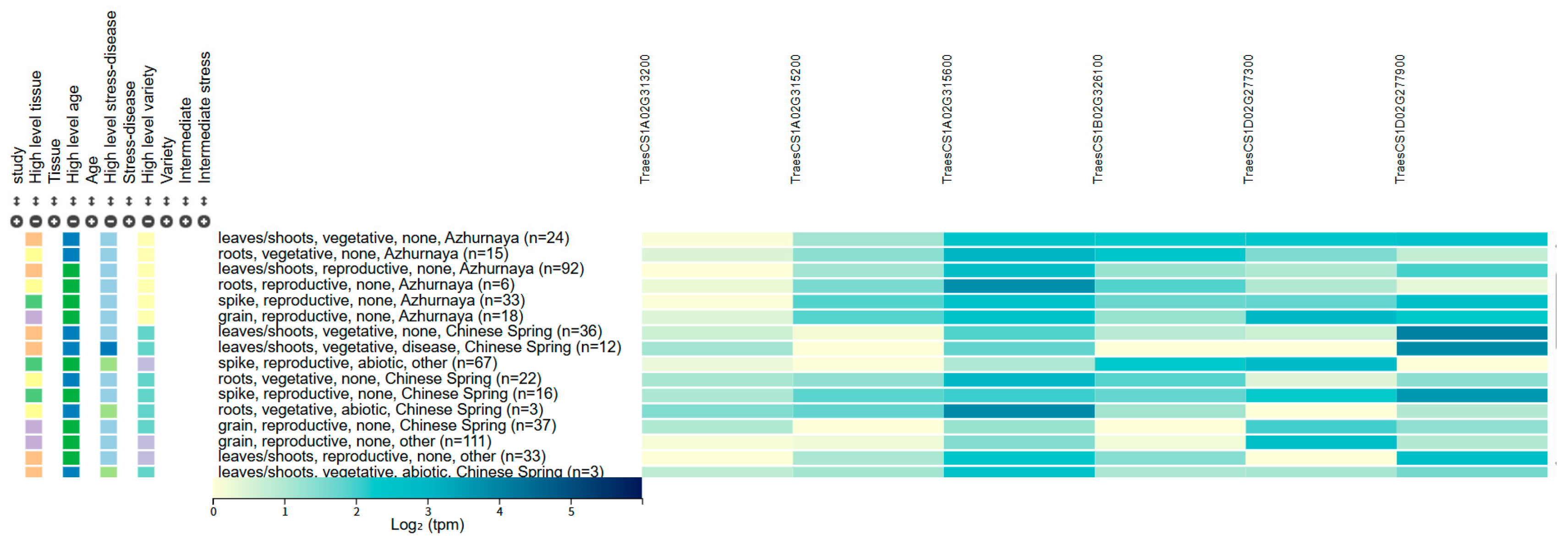
3. Discussion
3.1. Comparison with Known QTL Loci
3.2. Development of Molecular Markers for Major Loci QMt.ahau-1A, QMt.ahau-1B, and QTw.ahau-1D.1
3.3. Candidate Genes
4. Materials and Methods
4.1. Plant Materials and Field Experimental Design
4.2. Phenotypic Measurement
4.3. Statistical Analysis of Phenotypic Data
4.4. Construction of Genetic Linkage Map and QTL Mapping
4.5. Development and Validation of KASP Markers
4.6. Screening of Candidate Genes
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Wang, F.C.; Zhang, W.; Chen, W.Y. Mixograph and its application in the determination of wheat flour quality. Grain and Feed Industry. 2004, 12, 10-12.
- Sun, C.L.; Tian, J.C.; Zhang, Y.X. Application of texture analyzer in the evaluation of the wheat paste quality. Experiment Science and Technology. 2007, 5, 46-49.
- Mohamed,B.;,Abdullah, A.; Khaled,M.;Mohamed,M.;Mohamed,A.; Mohamed,M.;Mohamed,S.; Ibrahim, A. QTL analysis of farinograph and Mixograph related traits in spring wheat under heat stress conditions. Molecular Biology Reports.2020,47,5477-5486.
- Jiang, X.L.; Li, X.J.; Feng,S.V.;Hu,T.Z.;Ru,Z.G. Genetic variation of dough Mixograph characters and their relationships with other quality traits in wheat. Journal of Triticeae Crops.2013, 33, 806-811.
- Shen, X.Y.; Yan, J.; Chen, X.M.; Zhang,Y.;Li,H.L.;Wang,D.S.,He,Z.H.;Zhang,Y. Relationship of Mixograph parameters with farinograph and extensograph parameters,and bread-making quality traits. Acta Agronomica Sinica. 2010, 36, 1037-1043.
- James, C.N.; Cristina, A.; Flavio, B.; Patrick,L.F.Daisy,G. G.;Christine,J. B.;Roberto,J. P.;Marie,R.P.;Philippe,L.;Calvin,O. Q.; Mark,E. S.Quantitative trait locus analysis of wheat quality traits. Euphytica,2006, 149, 145-159.
- Morgan, E.S.; Ajay, K.;Shahryar,K.;Senay,S.;Mohammed,S.;Eder,E.M.;Phillip,E.M.;Edward,L.D.;Elias,E.;Blaine,S.;Steven,S.X.;Mohamed,M.A et al. New QTL alleles for quality-related traits in spring wheat revealed by RIL population derived from supernumerary×non-supernumerary spikelet genotupes. Theor. Appl. Genet. 2015, 128, 893-912.
- Huang,X.;Cloutier,S.;Lycar,L.;Radovanovic,N.;Humphreys,D.; Noll,J.;Somers,D.;Brown,P.Molecular detection of qtls for agronomic and quality traits in a doubled haploid population derived from two canadian wheats (Triticum aestivum L.).Theor. Appl. Genet.2006, 113, 753-766.
- Nelson,J. C.;Andreescu,C.; Breseghello,F.; Finney,P.L.;Gualberto,D.G.; Bergman,C.J.;Pena,R.J.;Perretant,M.R.;Leroy,P.;Qualset,C.O. Quantitative trait locus analysis of wheat quality traits. Euphytica, 2006, 149,145-159.
- Barakat, M.; Al-Doss, A.; Moustafa, K.; Motawei, M.; Alamri, M.; Mergoum, M.; Sallam, M.; Al-Ashkar, I. QTL analysis of farinograph and Mixograph related traits in spring wheat under heat stress conditions. Mol. Biol.2020, 47, 5477–5486.
- Chen, J.S. Construction of high denstiy genetic map of wheat (Triticum aestivum L.) and QTL analysis for quality and molecular marker exploration.Shandong Agricultural University, China,2015.
- Zheng, F.F.; Deng, Z.Y.; Shi, C.L.; Zhang, X.Y.; Tian, J.C. QTL mapping for dough mixing characteristics in a recombinant inbred population derived from a waxy × strong gluten wheat (Triticum aestivum L.). J. Integr. Agr. 2013, 12, 951–961.
- Deng,Z.; Tian,J.;Chen,F.;Li,W.;Zheng,F.; Chen,J.; Shi,C.; Sun,C.;Wang,S.; Zhang,Y. Genetic dissection on wheat flour quality traits in two related populations. Euphytica, 2015,203: 221-235.
- Xu, K.L. Identification of novel loci Qphs.ahau-5A and Qphs.ahau-6D for pre-harvest sprouting resistance in common wheat.Anhui Agricultural University, China, 2021.
- Jin, H.; Tian, Y.;Zhang,Y.;Zhang,R.;Zhao,H.B.;YangX.;Song,X.Z.;Dimitrov,Y.;Wu,Y.;Gao,Q.;Liu,J.D.;Zhang,J.M.;He,Z.H.Genome-wide association mapping of processing quality traits in common wheat (Triticum aestivum L.). Genes.2023, 14, 1816.
- Kong, X.S. Discovery of genetic loci controling wheat gluten quality and Mixograph parameters and identification of high quality germplasm.Shandong Agricultural University, China, 2022.
- James, C.N.;Cristina, A.;Flavio, B.;Patrick, L.F.;Daisy,G.G.;Christine,J.B.;Roberto,J.P.;Marie,R.P.;Philippe, L.;Calvin,O.Q.;Mark,E.S. Quantitative trait locus analysis of wheat quality traits. Euphytica.2006, 149, 145-159.
- Guo, H.; Fu, X.Y.; Ji, M.Q. Association analysis of dough mixing parameters using a natural population. Shandong Agricultural Sciences.2020, 52, 1-7.
- Yu,H.X.;An,Y.L.; Wang,A.P.;Guan,X.; Tian,J.C.; Ning,T.Y.; Fan.K.X.;Li,H.; Liu,Q.Q.;Wang, D.X.;Chen,J.S.Genetic dissection of the mixing properties of wheat flour (Triticum aestivum L.) using unconditional and conditional QTL mapping.J.Geonmics.2022,10:8–15.
- Sun,C.W.; Jing,Z.H.;Chen,X.Q.;Chen. J.H.; Shang,Q.Q.;Jin.H.;Jia.J,Z.;Ren.Y.; Zhao,L.; Gao,L.F.;He, Z.H.;Chen,F.Reconciliation of wheat 660K and 90K SNP arrays and their utilization in dough rheological properties of bread wheat.Journal of Advanced Research.2025,77: 1-13.
- Rasheed, A.; Hao, Y.F.; Xia, X.C.; Khan, A.; Xu, Y.B.; Varshney, R.K.; He, Z. H.Crop breeding chips and genotyping platforms: progress, challenges, and perspectives. Mol. Plant. 2017, 10, 1047–1064.
- Wu, H.; Graaf , B.; Mariani, C.; Cheung, A.Y. Hydroxyproline-rich glycoproteins in plant reproductive tissues: structure, functions and regulation. Cell. Mol. Life Sci. 2001, 58, 1418–1429.
- Showalter, A.M.; Keppler, B.D.; Liu, X.; et al. Bioinformatic identification and analysis of Hydroxyproline-rich glycoproteins in populus trichocarpa. BMC Plant Biol. 2016, 16, 229.
- Sun, A.L.; Li, Y.L.; He, Y.; Zou,X.X.;Chen,F.L.;Ji,R.Z.;You,C.Q.;Yu,K.Y.;Li,Y.;Xiao,W.J.;Guo,X.H.Comprehensive genome-wide identification, characterization, and expression analysis of CCHC-Type zinc ginger gene family in wheat (Triticum aestivum L.). Front. Plant Sci. 2022, 13, 892105.
- Wu, Z.M.; Shen, S.H.; Wang, Y.D.;Tao,W.Q.;Zhao,Z.Q.;Hu,X.L.;Yu,P. Genome-wide identification and expression analysis of the zinc finger protein gene subfamilies under drought stress in Triticum aestivum L. Plants,2022, 11, 2511.
- Ai, Q.; Pan, W.Q.; Zeng, Y.; Li,Y.H.;Cui,L.C.CCCH zinc finger genes in barley: genome-wide identification, evolution, expression and haplotype analysis. BMC Plant Biol. 2022, 22, 117.
- Chen, J.J.; Ouyang, Y.D.; Wang, L.;Xie,W.B.; Zhang,Q.F. Aspartic proteases gene family in rice: gene structure and expression, predicted protein features and phylogenetic relation. Gene. 2009, 442, 108-118.
- Wei, K.F.; Li, Y.X. Functional genomics of the protein kinase superfamily from wheat. Mol. Breed. 2019, 39, 141.
- Ral, J.P.;Bowerman, A.F.;Li, Z.Y.; Sirault,X.;Furbank,R.;Bloemsma,M.;Cavanagh,C.R.;Howitt,C.A.;Morell,M.K.Down-regulation of glucan, water-dikinase activity in wheat endosperm increases vegetative biomass and yield. Plant Biotech. J. 2012, 10, 871-882.
- Wei, K.F.; Han, P. Comparative Functional Genomics of the TPR Gene Family in Arabidopsis, Rice and Maize. Mol. Breed. 2017,37,152.
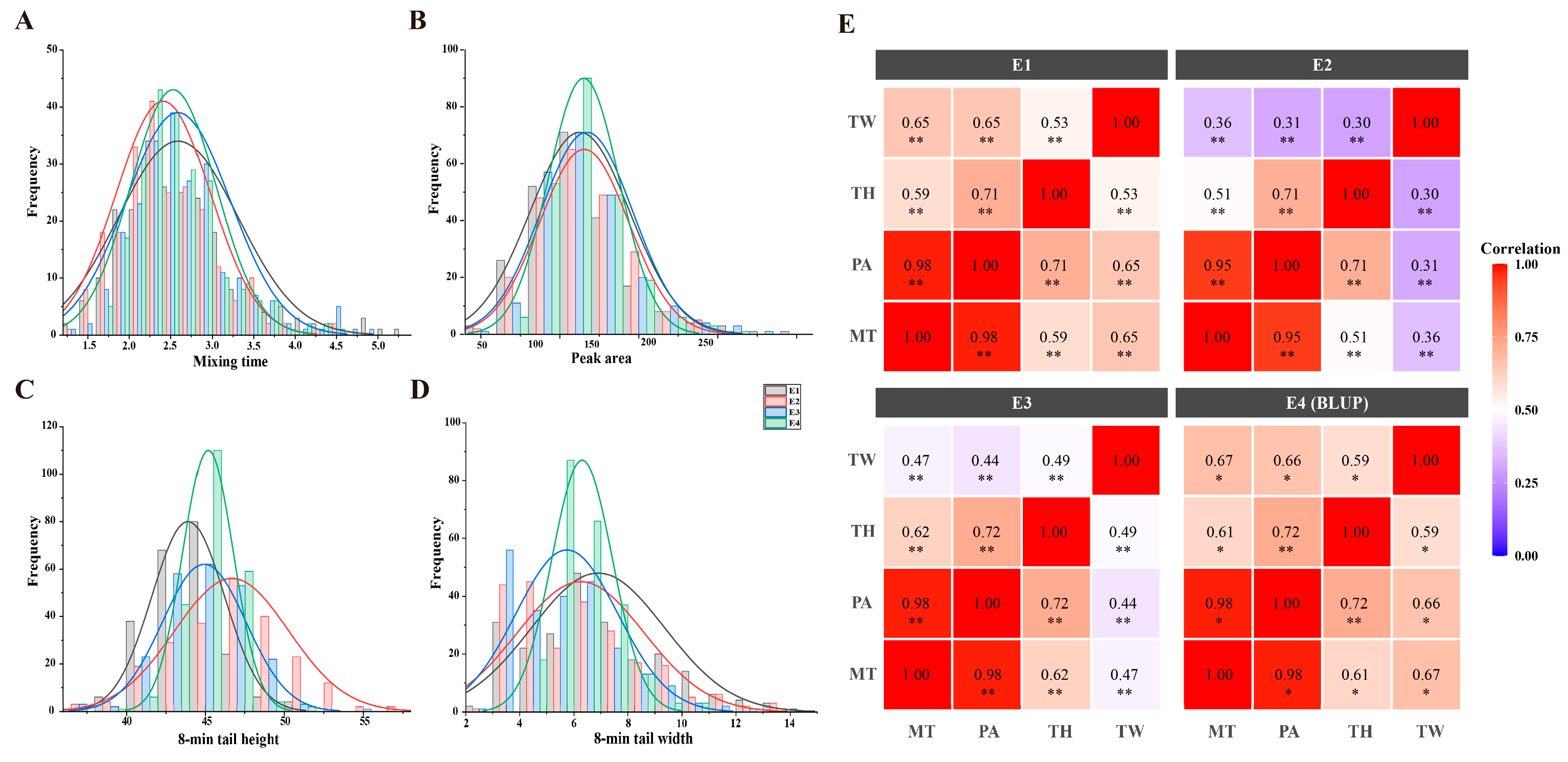
| Trait | Environment | Parents | RILs | ||||
| Zhongyou 9507 | Yangxiaomai | Minimum | Maximum | Mean | Cofficient of Variation | ||
| MT | 21SX (E1) | 4.4 | 2.1 | 1.3 | 5.2 | 2.6 | 0.28 |
| 22SX (E2) | 3.9 | 1.6 | 1.3 | 4.9 | 2.6 | 0.24 | |
| 22HF (E3) | 3.1 | 1.8 | 1.3 | 4.4 | 2.4 | 0.24 | |
| E4 (BLUP) | 3.8 | 1.8 | 1.6 | 4.4 | 2.5 | 0.2 | |
| PA | 21SX (E1) | 177.04 | 84.04 | 53.05 | 247.81 | 112.68 | 0.28 |
| 22SX (E2) | 180.49 | 75.11 | 58.96 | 239.54 | 117.03 | 0.25 | |
| 22HF (E3) | 139.03 | 79.58 | 56.12 | 205.98 | 115.45 | 0.24 | |
| E4 (BLUP) | 165.52 | 79.58 | 66.09 | 201.12 | 115.05 | 0.19 | |
| TH | 21SX (E1) | 43.63 | 41.12 | 37.93 | 51.81 | 43.91 | 0.05 |
| 22SX (E2) | 49.4 | 44.23 | 37.32 | 55.57 | 44.92 | 0.06 | |
| 22HF (E3) | 48.09 | 42.67 | 36.22 | 57.04 | 46.66 | 0.08 | |
| E4 (BLUP) | 47.04 | 42.67 | 39.56 | 50.12 | 45.16 | 0.04 | |
| TW | 21SX (E1) | 10.34 | 3.91 | 2.77 | 14.47 | 6.9 | 0.37 |
| 22SX (E2) | 5.89 | 5.67 | 2.59 | 12.53 | 5.74 | 0.33 | |
| 22HF (E3) | 6.71 | 4.79 | 2.59 | 17.07 | 6.28 | 0.39 | |
| E4 (BLUP) | 7.65 | 4.79 | 4.17 | 10.32 | 6.3 | 0.18 | |
| Trait | Mean square | F-value | H² | ||||
| Genotype | Environment | G×E | Genotype | Environment | G×E | ||
| MT | 3.02 | 7.18 | 0.34 | 174.58** | 414.23** | 19.83** | 0.90 |
| PA | 6169.98 | 3304.69 | 885.31 | 24089.16** | 12902.33** | 3456.49** | 0.87 |
| TH | 46.98 | 1320.2 | 14.11 | 411.48** | 11562.92** | 123.59** | 0.77 |
| TW | 26.37 | 234.67 | 9.47 | 1916.38** | 17051.77** | 687.97** | 0.74 |
| Trait | QTL | Flanking markers | Physical interval (Mb, IWGSC v1.0) | LOD value | Phenotypic variation explanation (PVE, %) | Additive effect | Environment |
| MT | QMt.ahau-1A | AX-110911310-AX-108914408 | 513.62-514.56 | 4.59 | 15.34 | 0.23 | E2/E3 |
| QMt.ahau-1B | AX-111503025-AX-110519906 | 551.72-552.48 | 21.23 | 23.62 | 0.39 | E1/E2/E4 | |
| TW | QTw.ahau-1D.1 | AX-109827648-AX-110069135 | 374.05-375.94 | 5.05 | 9.89 | -0.54 | E1/E4 |
| QTw.ahau-1D.2 | AX-109079356-AX-111804778 | 410.07-413.66 | 6.28 | 10.32 | 0.84 | E1/E3/E4 | |
| PA | QPa.ahau-1B | AX-111503025-AX-110519906 | 551.72-552.48 | 23.2 | 21.72 | 14.18 | E1/E4 |
| TH | QTh.ahau-1A | AX-109055961-AX-111496677 | 504.82-508.05 | 13.08 | 20.38 | -1.11 | E2/E4 |
| KASP marker | SNP marker | FAM-compatible primers (5'-3') | HEX-compatible primers (5'-3') | Common primers (5'-3') |
| QMt-1A-KASP | AX-110911310-AX-108914408 | gcatgtcatcagcactgtgA | gcatgtcatcagcactgtgG | cctttctcacataTggcgcC |
| QMt-1B-KASP | AX-111503025-AX-110519906 | ccacaactttctgcctagctA | ccacaactttctgcctagctG | ttatgcggtCgaggatccG |
| QTw-1D.1-KASP-1 | AX-109827648-AX-110069135 | tcaacacTaaatgctccaaaaacaG | tcaacacTaaatgctccaaaaacaA | gcttgctaacctggaagatccG |
| QTw-1D.1-KASP-2 | AX-109827648-AX-110069135 | cttCgtccacgtccAttagT | cttCgtccacgtccAttagC | acacaagagcatgtaataccgG |
| Candidate Gene | Chromosome | Position | Annotation |
| (Mb) | |||
| TraesCS1A02G313200 | 1A | 505.27 | Hydroxyproline-rich glycoprotein family protein |
| TraesCS1A02G315200 | 1A | 506.55 | zinc finger CCCH domain protein |
| TraesCS1A02G315600 | 1A | 507.08 | Protease-related family protein |
| TraesCS1B02G326100 | 1B | 551.72 | Kinase family protein |
| TraesCS1D02G277300 | 1D | 374.40 | Phosphoglucan, water dikinase, chloroplastic |
| TraesCS1D02G277900 | 1D | 375.20 | Tetratricopeptide repeat (TPR)-like superfamily protein |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




