2. Results
2.1. The Oncogenic Core Network
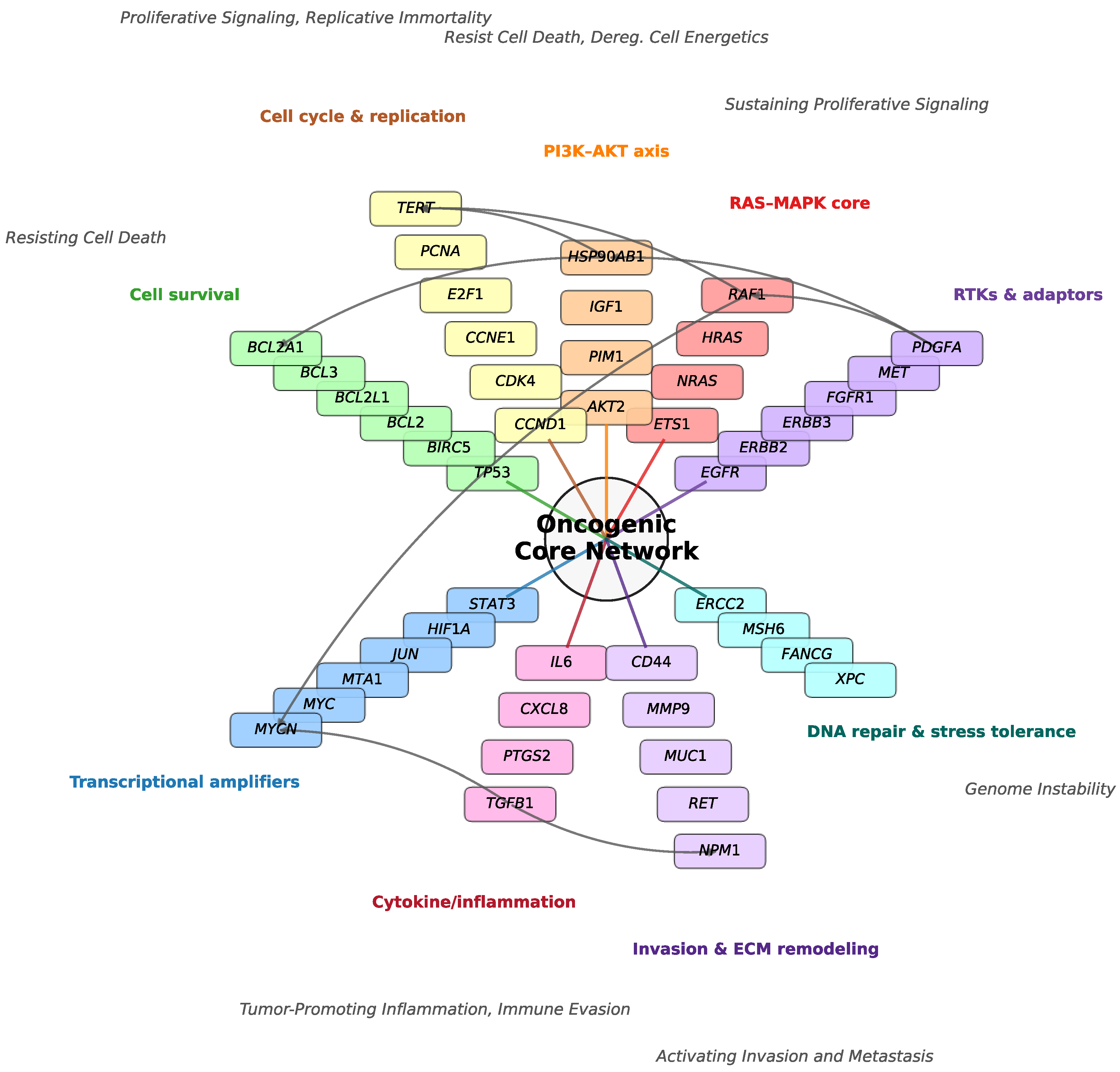
To define the baseline architecture of the study, we first mapped the 102 selected consensus cancer genes onto a canonical oncogenic core network (
Figure 1). This network illustrates how the selected genes organize into interconnected functional modules that collectively sustain malignant phenotypes. RTKs & Adaptors (e.g.,
EGFR,
ERBB2,
MET) act as upstream hubs, transmitting extracellular growth signals into the RAS–MAPK core and PI3K–AKT axis. The RAS–MAPK module—constitutively locked in an active state by the gain-of-function
KRAS G12S mutation—links directly to Cell Cycle & Replication (e.g.,
CCND1,
CDK4) and Transcriptional Amplifiers include canonical oncogenes (e.g.
MYC) and contextual promoters (e.g.,
STAT3,
JUN), collectively enforcing the Hallmarks of Sustaining Proliferative Signaling and Enabling Replicative Immortality.
In parallel, the PI3K–AKT axis functions as a critical survival node. In the specific context of A549 cells, this axis connects to Cell Survival to override the apoptotic potential of wild-type TP53 while engaging anti-apoptotic effectors like BCL3. This active suppression ensures Resisting Cell Death and Deregulating Cellular Energetics, maintaining viability despite oncogenic stress. Cytokine/Inflammatory Signaling (e.g., IL6, TGFB1— contextual tumor promoters) feeds directly into transcriptional amplification and Invasion & ECM Remodeling (e.g., CD44, MMP9), linking Tumor-Promoting Inflammation to Activating Invasion & Metastasis—a connection that is particularly amplified by the STK11 deficiency inherent to this cell line.
Finally, DNA Repair & Stress Tolerance (e.g., ERCC2, MSH6) anchors the hallmark of Genome Instability. Critically, this module is shaped by the loss-of-function KEAP1 G333C mutation found in A549 cells. By disabling the "brake" on oxidative stress responses, this mutation creates a constitutive antioxidant shield that protects the tumor against therapeutic insults, maintaining a permissive state for oncogenic evolution.
2.2. Classification of Genes by Pathway and Global Centroid Dynamics
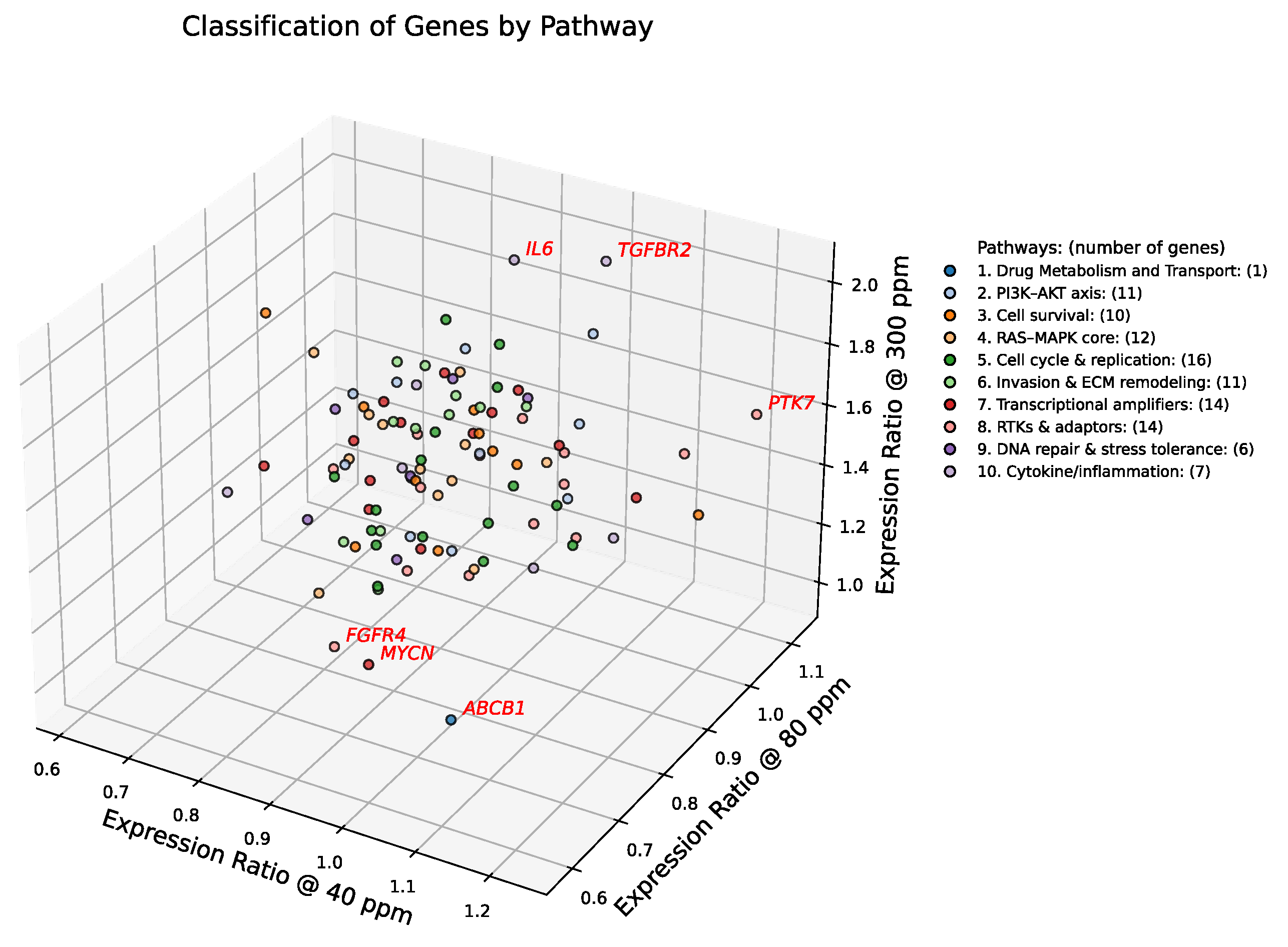
To map the transcriptional landscape of A549 cells under deuterium stress, we plotted the 102 consensus genes in a three-dimensional expression space defined by their ratios at 40, 80, and 300 ppm relative to the 150 ppm control (
Figure 2). Genes were color-coded by pathway, and their collective behavior was quantified by calculating pathway-specific centroids (
Table 1). To identify transcriptional outliers, we ranked all genes by their Euclidean distance from the global center. The top six genes—
TGFBR2,
IL6,
FGFR4,
ABCB1,
MYCN, and
PTK7—were selected based on their pronounced divergence and a clear inflection point in the distance distribution (highlighted in red in
Figure 2).
The quantitative analysis of pathway centroids (
Table 1) reveals a monotonic increase in transcriptional activity correlating with deuterium concentration. The
Global Center coordinates (0.884 → 0.940 → 1.436) serve as the system-wide baseline:
At 40 ppm, the global centroid (0.884) is well below unity, indicating that extreme deuterium depletion exerts a broad suppressive effect on the oncogenic transcriptome.
At 80 ppm, expression remains dampened (0.940), reinforcing the concept of depletion as a "metabolic brake."
At 300 ppm, the global centroid surges to 1.436, reflecting a robust, system-wide activation (~44% increase).
Using this global 1.436 value as a benchmark reveals distinct pathway sensitivities. Pathways exceeding this threshold at 300 ppm—including Cytokine/Inflammation (1.567), Cell Survival (1.476), and Invasion & ECM Remodeling (1.469)—are hyper-responsive to deuterium enrichment. In contrast, RTKs & Adaptors (1.377) and Cell Cycle & Replication (1.371) respond more modestly, falling below the global average. Notably, Drug Metabolism and Transport (ABCB1) exhibits a unique trajectory with extreme suppression at 80 ppm (0.585), diverging sharply from the global trend. Together, these results indicate that A549 cells adapt to deuterium stress by suppressing energetically costly programs (like drug efflux) under depletion, while enrichment drives a coordinated reprogramming that favors survival, invasion, and inflammation.
Centroid values represent the mean relative expression ratios at 40, 80, and 300 ppm compared to the 150 ppm control. The Global center corresponds to the overall average across all pathways and serves as a reference baseline. Pathways are annotated with representative genes.
2.3. Density-Based Spatial Clustering
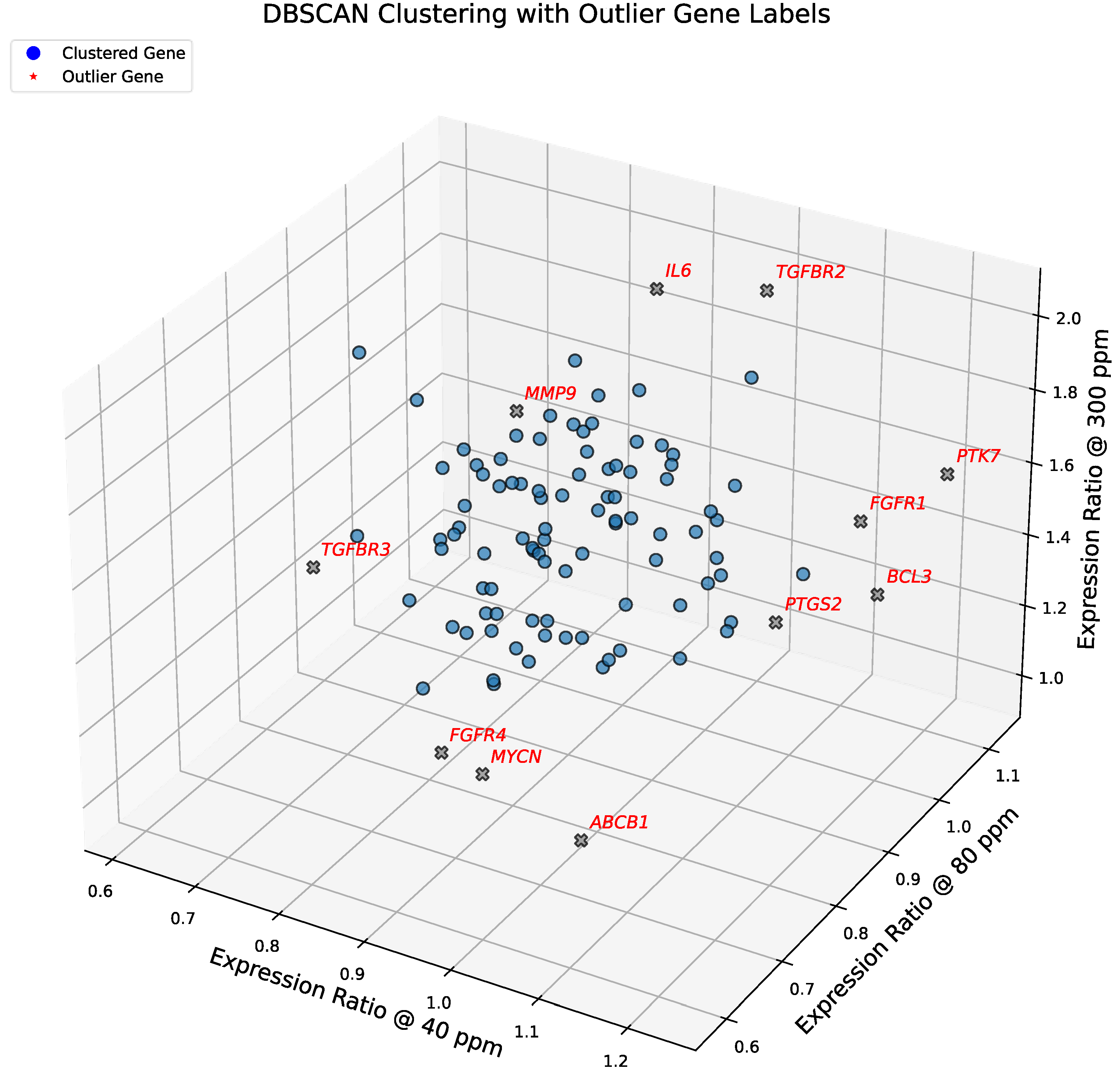
To complement centroid-based identification of transcriptionally sensitive genes, we applied DBSCAN to the 3D expression space defined by relative expression ratios at 40 ppm, 80 ppm, and 300 ppm deuterium as shown in
Figure 3. A 5-distance plot (
Figure S1) was used to select the ε parameter, consistent with the standard practice of setting
min_samples = 5. This choice balances sensitivity to small transcriptional modules with robustness against noise, enabling detection of biologically meaningful outliers in the 102-gene dataset. Unlike centroid-based methods that emphasize spherical pathway-level coherence, DBSCAN identifies density-based outliers with unique expression ratios, independent of canonical pathway clustering. These outliers may represent sentinel responders or context-specific regulators whose behavior diverges from pathway averages, offering additional insight into deuterium-dependent transcriptional heterogeneity.
To visually summarize the directional behavior of gene expression ratios across three deuterium concentrations (40, 80, 300 ppm), we assigned each outlier a pattern symbol triplet (e.g., → ↓ ↗) based on its relative expression values. These symbols reflect the magnitude and direction of change using a biologically interpretable scale: ↑ for strong upregulation (>1.37), ↗ for moderate upregulation (1.17–1.37), → for stable expression (0.82–1.17), ↙ for moderate downregulation (0.70–0.82), and ↓ for strong downregulation (<0.70). The thresholds were rigorously selected based on statistical distributions and centroid benchmarks (e.g., the 1.37 upper threshold aligns with the lowest centroid of upregulated clusters at 300 ppm).
2.4. Pathway-Specific Analysis of DBSCAN Outlier Genes
To elucidate the biological relevance of the 11 outlier genes identified by DBSCAN clustering (ε = 0.13), we grouped them by functional pathway and analyzed their expression dynamics.
2.5. Drug Metabolism and Transport
ABCB1 (→ ↓ ↗) encodes P-glycoprotein, a member of the ATP-Binding Cassette (ABC) transporter superfamily. It functions as an ATP-powered efflux pump that actively exports a wide range of hydrophobic molecules, including chemotherapeutic agents, making it a central mediator of multidrug resistance (MDR). In our dataset, ABCB1 exhibited a non-linear response: mild induction at 40 ppm (+12%), suppression at 80 ppm (−42%) (with small 7% error margin), and reactivation at 300 ppm (+27%). This distinct, significant suppression at 80 ppm is particularly significant and identifies ABCB1 as a sentinel gene highly sensitive to moderate deuterium depletion.
2.6. Cell Survival
BCL3 (↗ → ↑) is an IκB-like protein that modulates NF-κB signaling, promoting transcription of anti-apoptotic genes and enabling tumor survival under stress. It is frequently upregulated in hematologic and solid malignancies. In this dataset, BCL3 exhibited progressive upregulation across all deuterium concentrations, consistent with a robust survival response. Its low total error reinforces the reliability of this measurement and supports its role as a transcriptional amplifier of cell survival under deuterium concentration change.
2.7. RTKs and Adaptors
This functional group highlights a striking divergence in receptor sensitivity. FGFR1 (→ → ↑) exhibited stable expression at physiological levels but surged at 300 ppm (+57%), indicating a threshold-dependent response to deuterium enrichment. In sharp contrast, FGFR4 (↙ ↙ →) was consistently suppressed at 40 and 80 ppm (~−19%), suggesting that deuterium depletion dampens specific growth factor receptivity. PTK7 (↗ → ↑), an orphan receptor involved in Wnt/Planar Cell Polarity signaling, displayed a progressive upward trend across all concentrations. Together, these profiles demonstrate that while enrichment fuels broad receptor activation (FGFR1, PTK7), depletion selectively compromises specific signaling nodes (FGFR4).
2.8. Cytokine and Inflammation
IL6 (→ → ↑), PTGS2 (↗ → ↑), TGFBR2 (→ → ↑), and TGFBR3 (↓ → ↗) are key regulators of inflammation and immune modulation. IL6 and PTGS2 promote tumor growth, angiogenesis, and immune suppression, with IL6 showing strong induction at 300 ppm (+99%). TGFBR2 and TGFBR3 mediate TGF-β signaling, which plays context-dependent roles in cancer progression. TGFBR2 was strongly induced at 300 ppm (+105%), while TGFBR3 showed distinct suppression at 40 ppm (−36%) followed by recovery. These patterns suggest that deuterium enrichment amplifies inflammatory signaling, contributing to Tumor-Promoting Inflammation.
2.9. Invasion and ECM Remodeling
MMP9 (↓ → ↑) encodes a matrix metalloproteinase that degrades extracellular matrix components, facilitating tumor invasion. It was suppressed at 40 ppm (−31%) and progressively induced at higher concentrations. As a driver of Activating Invasion & Metastasis, its suppression under depletion reflects a dose-dependent inhibition of invasive behavior.
2.10. Transcriptional Amplifiers
MYCN (→ ↙ →) is a member of the MYC family, regulating cell cycle and metabolism. It showed mild suppression at 80 ppm (−24%) followed by stability at 300 ppm (+1%). As a master regulator, MYCN’s modulation under deuterium stress may have cascading effects on downstream oncogenic programs.
2.11. Core Pathway Stability
In the current dataset of 102 genes, four pathways—Cell cycle & replication, DNA repair & stress tolerance, PI3K–AKT axis, and RAS–MAPK core—contained no outlier genes. This consistency shows that all genes in these core oncogenic programs responded in a moderate, coordinated manner to deuterium concentration changes, indicating that these pathways act as stable, coherent modules. In contrast, heterogeneous pathways like Cytokine and Inflammation showed larger variations, producing multiple outliers.
2.12. Symbolic Clustering of Gene Expression Patterns
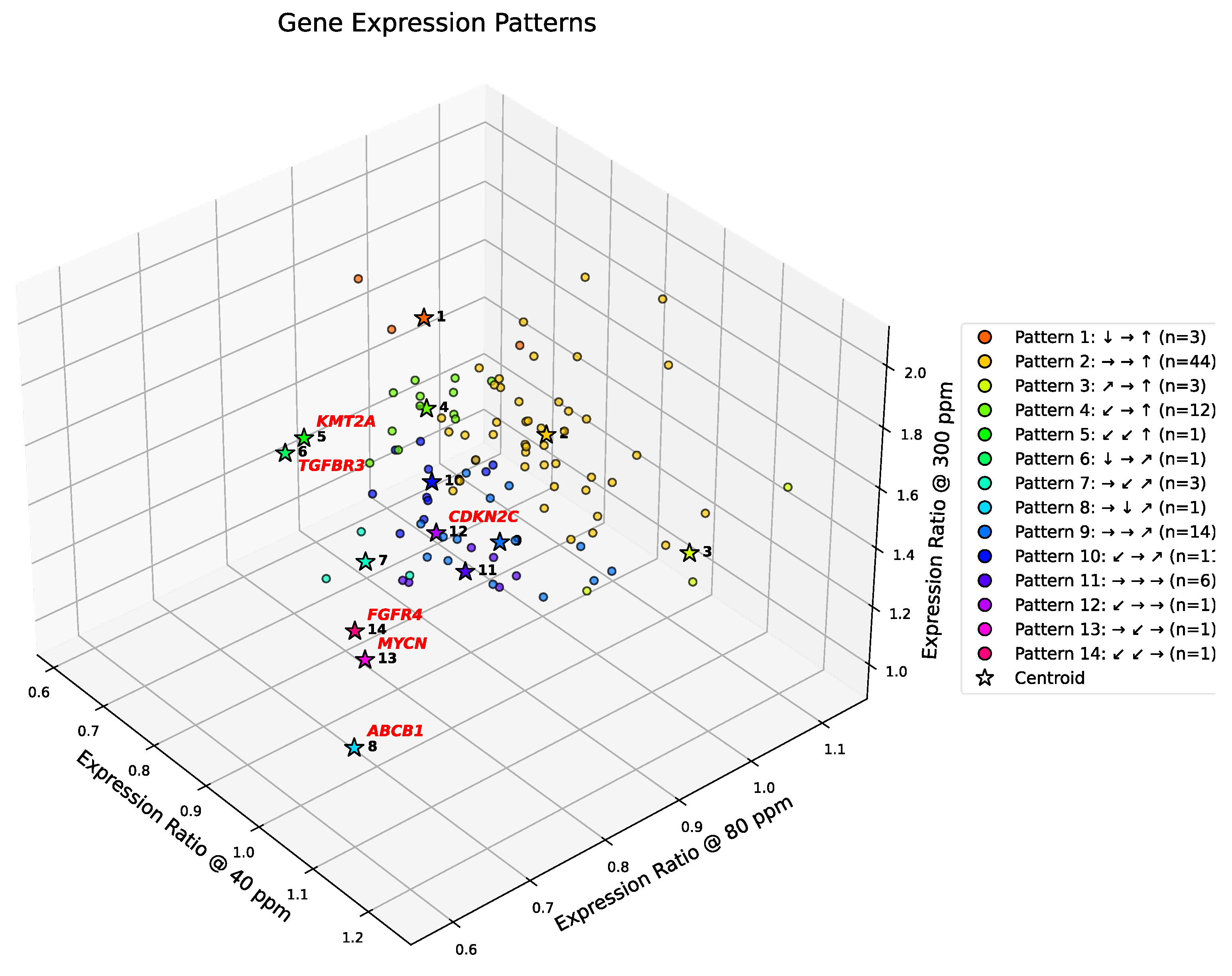
In this section, we analyze gene expression patterns in the 102-gene dataset using the discretized thresholds introduced above. Symbolic clustering across deuterium concentrations reveals a striking sparsity in realized transcriptional patterns. Although the symbolic encoding scheme allows for 125 theoretical combinations, only 14 distinct patterns were observed (cf.
Figure 4). This >90% reduction in theoretical complexity reflects strong biological constraints, suggesting that A549 cells converge onto a limited set of coordinated regulatory programs to manage deuterium stress.
2.13. Dominant Transcriptional Trajectories
The distribution of genes across these 14 patterns is highly skewed. Most genes converge into just three dominant clusters, with Pattern 2 emerging as the overarching transcriptional program.
Pattern 2 (→ → ↑): This dominant group (n=44) represents genes that remain stable under depletion but surge under enrichment. It encapsulates the "Enrichment as Accelerator" phenotype, comprising nearly half the dataset and driving the Hallmarks of Resisting Cell Death, Sustaining Proliferative Signaling, and Tumor-Promoting Inflammation.
Minor Clusters: Two other patterns (Patterns 4 and 9) contain between 12–14 genes, representing alternative regulatory modes. Patterns 1, 3, 7, and 11 contain between 3–6 genes.
2.14. Singleton Outliers and Methodological Consensus
In contrast to the dominant groups, six genes—
ABCB1, CDKN2C, FGFR4, KMT2A, MYCN, and
TGFBR3—exhibit unique symbolic patterns not shared by any other gene. Importantly, four of these six singleton genes align with the outliers identified by DBSCAN, reinforcing their status as biologically distinct transcriptional entities (
Figure 4). For instance,
ABCB1 and
FGFR4 consistently diverge from the population mean, spanning critical hallmarks of Multidrug Resistance and Growth Factor Signaling.
Comparative Outlier Detection and Dataset Refinement
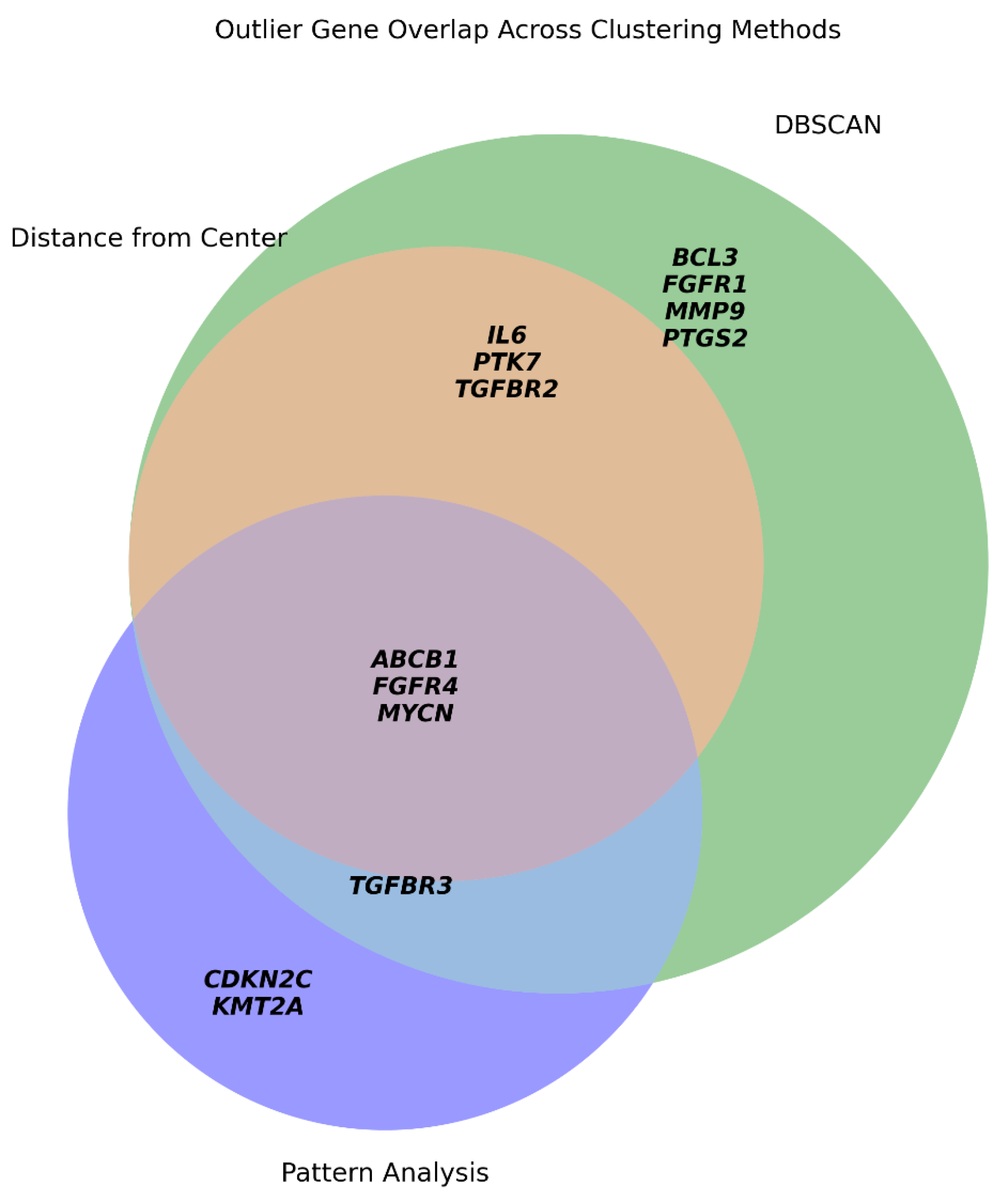
We evaluated transcriptional outliers using three complementary approaches: global centroid distance, DBSCAN clustering, and symbolic pattern analysis (
Figure 5).
Centroid-based: Flagged six genes, all of which were re-confirmed by DBSCAN.
Symbolic Analysis: Overlapped with four DBSCAN-defined genes and identified unique trajectories for CDKN2C and KMT2A.
DBSCAN: Proved the most sensitive method, identifying the "core" outliers captured by other methods plus five additional context-specific regulators.
This comparison demonstrates that while distance-based metrics provide baseline selectivity, DBSCAN offers superior sensitivity in detecting transcriptional divergence. Accordingly, for the subsequent GMM clustering, we excluded the full set of DBSCAN-defined outliers to improve cluster resolution and biological interpretability, reducing the dataset to 91 genes.
In summary, the symbolic clustering framework delineates a biologically constrained transcriptional landscape, characterized by a limited set of dominant high-expression patterns and punctuated by distinct outliers. The sparsity of realized patterns, the centrality of Pattern 2, and the functional relevance of singleton genes highlight the importance of integrated clustering and pathway-level interpretation in future analyses. Nonetheless, propagated measurement error in the current dataset may reach 10–15%, potentially causing genes near classification boundaries to overlap with adjacent patterns. This ambiguity (see Figure S2) highlights the need for complementary unsupervised clustering approaches. Moreover, the five-category classification scheme may be overly rigid for capturing extreme expression levels—for example, it assigns the same symbol (↑) to 40% and 100% overexpression. To complement the symbolic classification, we next applied unsupervised clustering using Gaussian Mixture Models (GMM), which provide a probabilistic framework for identifying continuous transcriptional variation and resolving ambiguous boundary assignments. Gaussian Mixture Models are particularly suited to this dataset because they treat expression ratios as overlapping probability distributions, allowing genes with propagated error near symbolic boundaries to be assigned probabilistically rather than rigidly. This flexibility captures subtle transcriptional shifts and resolves the ambiguity inherent in discrete symbolic categories.
2.15. GMM Clustering 91 Genes with Tied Covariance Ellipsoids
We evaluated Gaussian Mixture Models (GMMs) using silhouette analysis across seven candidate cluster counts (
k = 2 to 9), where
k denotes the number of components the algorithm attempts to identify (cf.
Figure S3). Each cluster ideally captures a distinct gene expression pattern under varying deuterium concentrations.
2.16. Statistical Model Selection
A critical observation from the silhouette analysis is that all scores remained below 0.5. In clustering topology, a coefficient < 0.5 indicates that the data structure is not characterized by distinct, widely separated "islands," but rather represents a continuous biological gradient with overlapping boundaries. Consequently, the silhouette score alone was insufficient to determine the optimal partition, requiring a balance between statistical separation and biological resolution.
Rejection ofk= 2: Although the tied covariance model with k = 2 yielded the highest numerical silhouette score, inspection revealed this solution was uninformative, providing only a trivial binary partition (high vs. low expression at 300 ppm) that masked pathway-specific dynamics.
Selection ofk= 4 andk= 6: The tied covariance model with k = 4 yielded the second-highest score, closely followed by k = 6. This configuration achieved a favorable Bayesian Information Criterion (BIC = −438) and log-likelihood (LogLik = 267), suggesting an optimal balance between model fit and complexity.
2.17. Model Robustness and Sensitivity Analysis
While full covariance models produced higher LogLik values, their substantially worse BIC scores pointed to overfitting, justifying the use of tied covariance. Overall, GMM clustering (silhouette = 0.320) outperformed k-means (silhouette = 0.255), confirming that the probabilistic, ellipsoid-based boundaries of GMM better capture the complex covariance structure of oncogenic gene expression than the spherical assumptions of k-means.
To assess clustering robustness, we performed a leave-one-out sensitivity analysis comparing GMM models with 4 and 6 clusters (GMM-4 and GMM-6, cf.
Table S2). For each gene, we computed the Adjusted Rand Index (ARI) after its removal. The average ARI difference was −0.043, indicating that GMM-6 is marginally more stable overall. However, gene-specific impacts varied:
Stabilizers: Genes such as
BIRC5 and
MUC1 stabilized the GMM-4 solution, reflecting their roles in core cell cycle/survival modules, cf.
Table S2.
Destabilizers: Genes like
AKT2 and
BCL2A1 destabilized GMM-4, suggesting they sit at the boundaries of broader clusters and require the higher resolution of GMM-6 to be accurately partitioned, cf.
Table S2.
Instability ofk= 9: Notably, the 9-cluster solution proved unstable; removal of a single gene triggered complete cluster rearrangement, confirming that k = 9 over-fragmented the data into artificial groups.
2.18. Cluster Visualization and Boundary Drivers
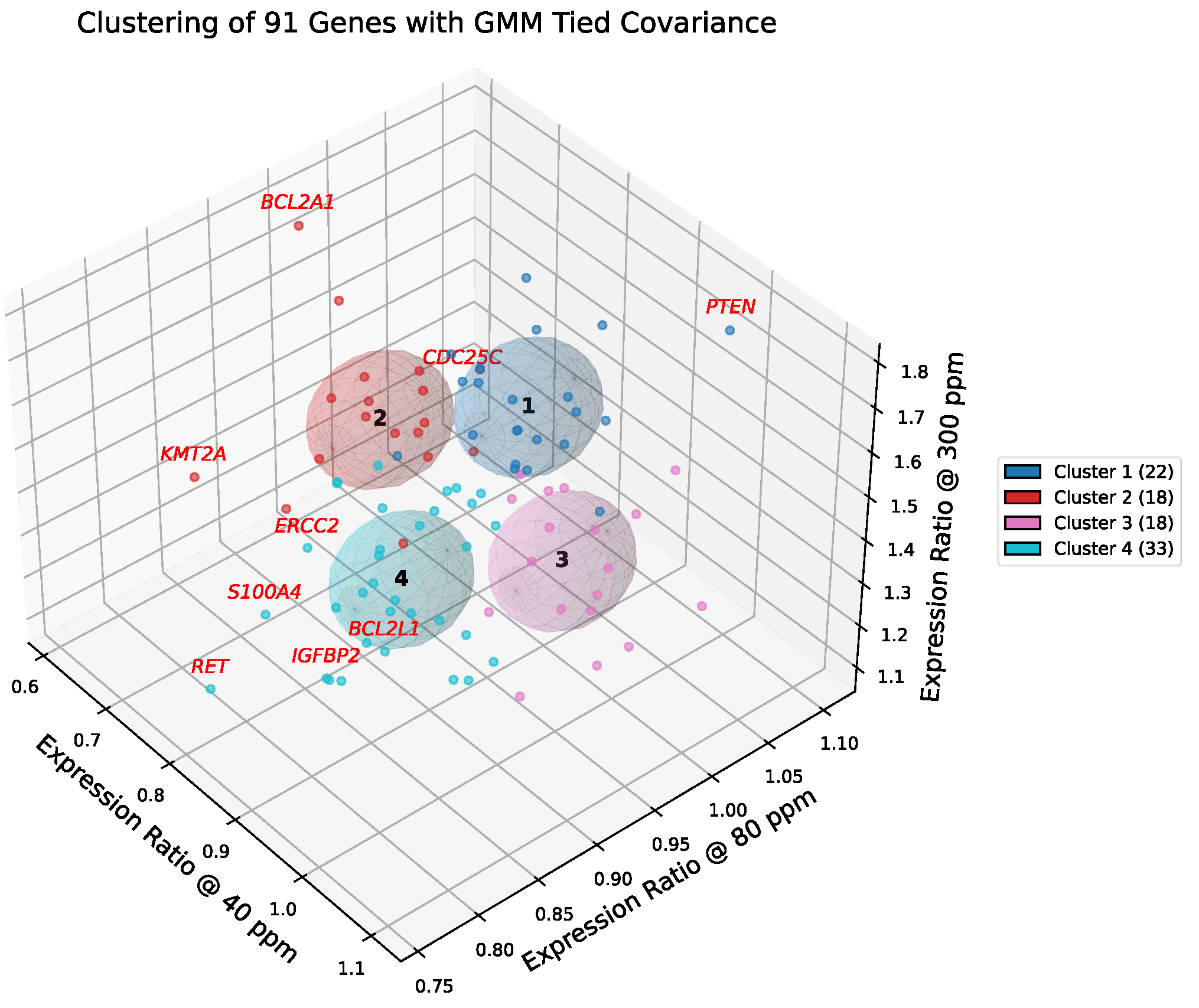
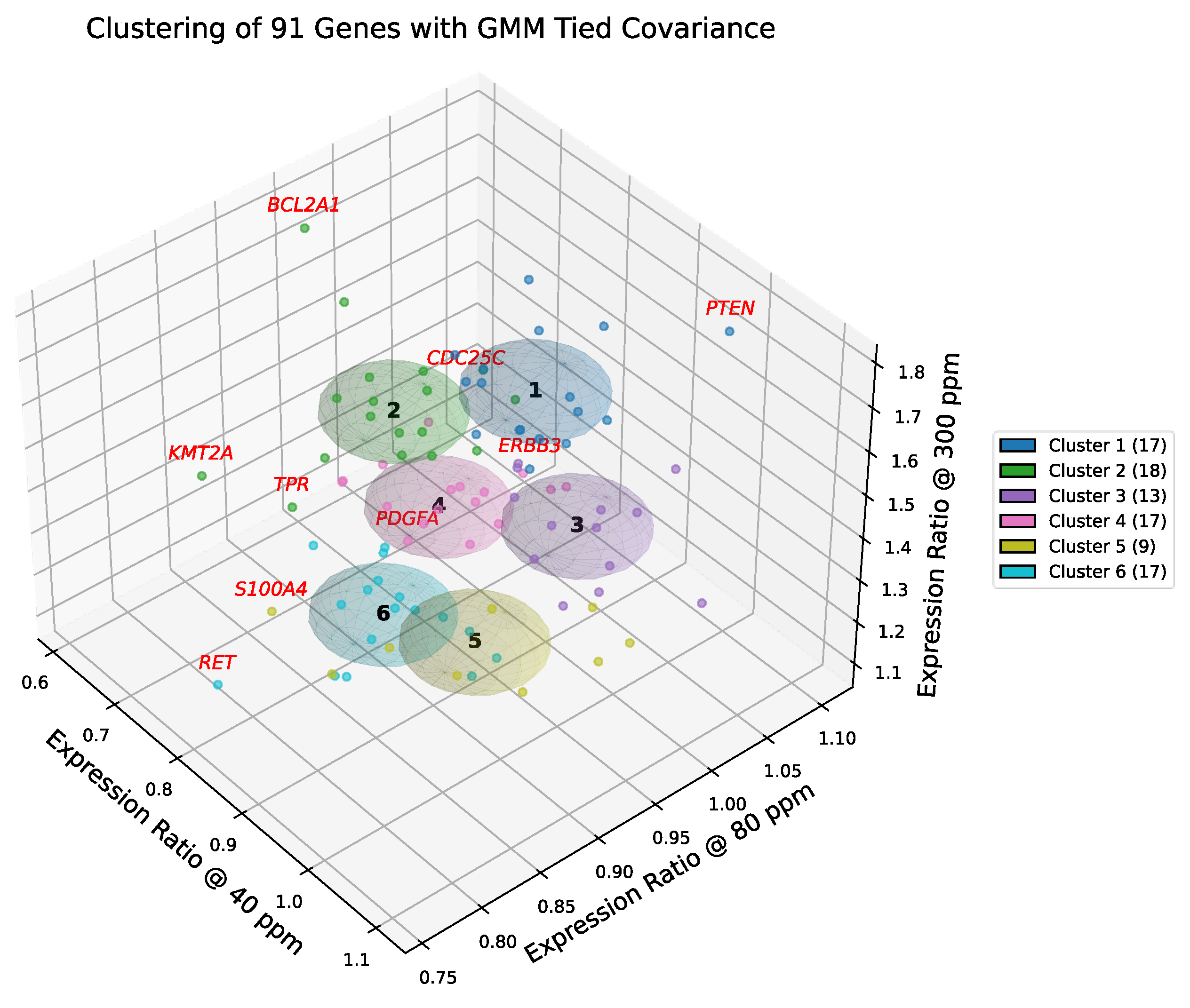
Figure 6 and
Figure 7 present the results of GMM-4 and GMM-6 applied to the curated set of 91 cancer-related genes. In the selected model, each cluster is represented by a shared covariance ellipsoid centered at its coordinate mean. To assess the confidence of assignments, we computed the Mahalanobis distance of each gene from its cluster centroid, cf.
Table S2. Unlike Euclidean distance, Mahalanobis distance accounts for the covariance structure, penalizing directions of low variance. Genes with high Mahalanobis distances (e.g.,
PTEN, CDC25C, ERBB3, BCL2A1) were identified as boundary drivers—genes that are statistically peripheral to their cluster yet biologically critical for defining the cluster's functional edge (e.g., balancing tumor suppression with amplification).
2.19. Cluster Transitions and Heterogeneity (GMM-4 to GMM-6)
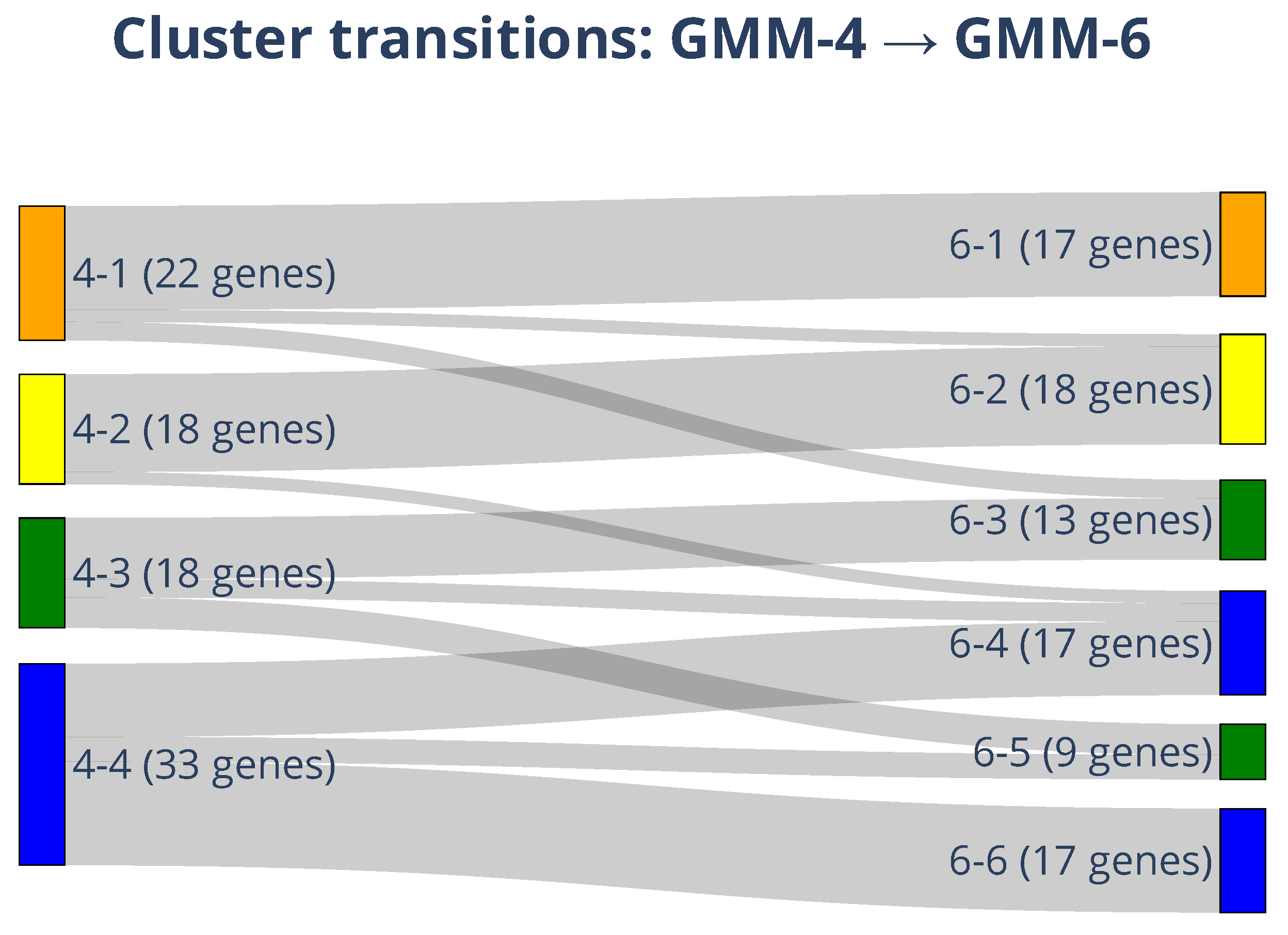
To visualize how increasing model granularity redistributes genes, we mapped the transitions from GMM-4 to GMM-6 using a Sankey diagram (cf.
Figure 8). The analysis revealed clear patterns of stability and fragmentation:
Stable Modules: Genes assigned to GMM-4 clusters 4-1 and 4-2 (22 and 18 genes, respectively) largely remained together, mapping almost exclusively into GMM-6 clusters 6-1 and 6-2. This indicates that these transcriptional programs (associated with Cellular Regulation and KRAS Effector signaling) are robust across model resolutions.
Fragmented Heterogeneity: In contrast, the largest GMM-4 cluster (4-4, n = 33) was partitioned into multiple GMM-6 clusters (6-4, 6-5, and 6-6). This dissociation unmasked critical heterogeneity: it separated the Invasive/Plasticity drivers (TGFB1, now Cluster 5) and Growth Factors (PDGFA, now Cluster 4) from the Basal Stability genes (BIRC5, now Cluster 6). The proportional widths of the flows emphasize that while some transcriptional states are robust, others fragment into finer substructures, proving that the six-cluster resolution is necessary to distinguish Invasion programs from general survival mechanisms.
2.20. Composition and Biological Identity of GMM-6 Clusters
Gaussian mixture modeling with tied covariance ellipsoids (k = 6), validated by silhouette analysis, delineated six distinct gene groups whose centroid trajectories across deuterium concentrations define the transcriptional landscape of A549 cells.
Clusters 1 and 2 showed the strongest induction at 300 ppm (cf. Figure 7), representing stress-responsive proliferative and survival modules:
Cluster 1 integrated cell cycle regulators and DNA repair factors, with boundary outliers (PTEN, CDC25C, ERBB3) marking the tension between tumor suppression and receptor-driven proliferation. Canonical oncogenes (MYC, STAT3, TERT) reinforced its proliferative bias within the context of a wild-type TP53 axis.
Cluster 2 encompassed transcriptional amplifiers and apoptosis regulators, with outliers (BCL2A1, KMT2A, TPR) highlighting survival stability. Importantly, this cluster contained the core RAS effector RAF1 alongside EGFR, underscoring its role as the primary signaling engine driven by the G12S mutation.
Cluster 3 was cohesive, balancing oncogenic drivers (ERBB2, JUN, IGF1) with the tumor suppressor TP53. This module reflects a signaling core where proliferative drive is counterweighted by suppressive control. While basal transcriptional load is driven by oncogenic RAS, the co-clustering of TP53 with homeostatic regulators confirms that A549 cells retain a competent, stress-responsive p53 axis. This module preserves canonical regulatory architecture, enabling a coordinated—though ultimately overwhelmed—response to deuterium modulation.
Cluster 4 contained accessory growth factor and stress response genes, with PDGFA defining its proliferative edge. Oncogenes (NRAS, MET, LYN) were present alongside apoptosis regulators (BCL2, FAS), highlighting a dual role in growth signaling and stress adaptation distinct from the primary RAS driver in Cluster 2.
Cluster 5 was enriched for ECM remodeling and invasion, with S100A4 marking the invasive boundary. Key members included the oncogene HRAS alongside the tumor promoter TGFB1. The enrichment of these plasticity drivers is consistent with the phenotype of STK11 (LKB1) deficiency inherent to A549 cells, identifying this cluster as the primary engine of microenvironmental adaptation and metastatic potential.
Cluster 6 grouped cell cycle and repair genes that remained relatively stable across conditions, with RET delineating its invasion-linked outlier. Oncogenes (AKT2, CDK4, FGFR3) were central, together with survival factors such as BIRC5 (Survivin). This defines a “Basal Maintenance” cluster that sustains viability even when more plastic modules (Cluster 5) are suppressed.
Together, these clusters reveal a structured landscape in which proliferative and survival modules dominate the stress response, while invasion and repair pathways provide adaptive support. Outlier genes stretch cluster boundaries, identifying the most stress-responsive drivers of instability. Notably, several highlighted genes—including TP53, EGFR, and RET—are clinically relevant in A549 LUAD cell line, underscoring the translational importance of these clustering results.