Submitted:
09 February 2026
Posted:
10 February 2026
You are already at the latest version
Abstract
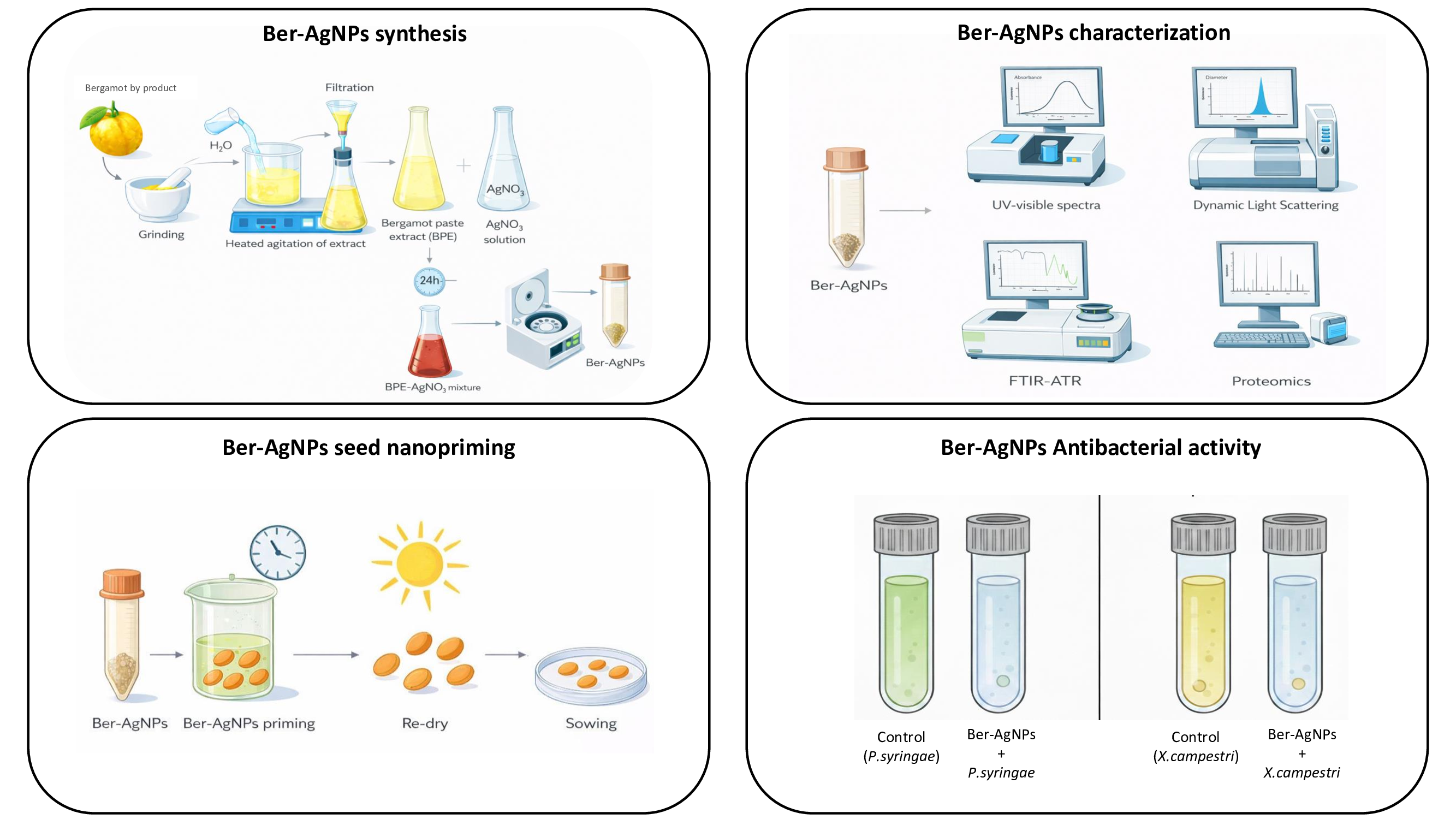
Keywords:
1. Introduction
2. Results and Discussion
2.1. Optimization of Reaction Conditions
2.1.1. Effect of Bergamot Pomace Extract (BPE) Concentration
2.1.2. Effect of pH
2.1.3. Effect of extract-to-metal ratio
2.1.4. Effect of Incubation Time
2.1.5. Effect of Incubation Temperature
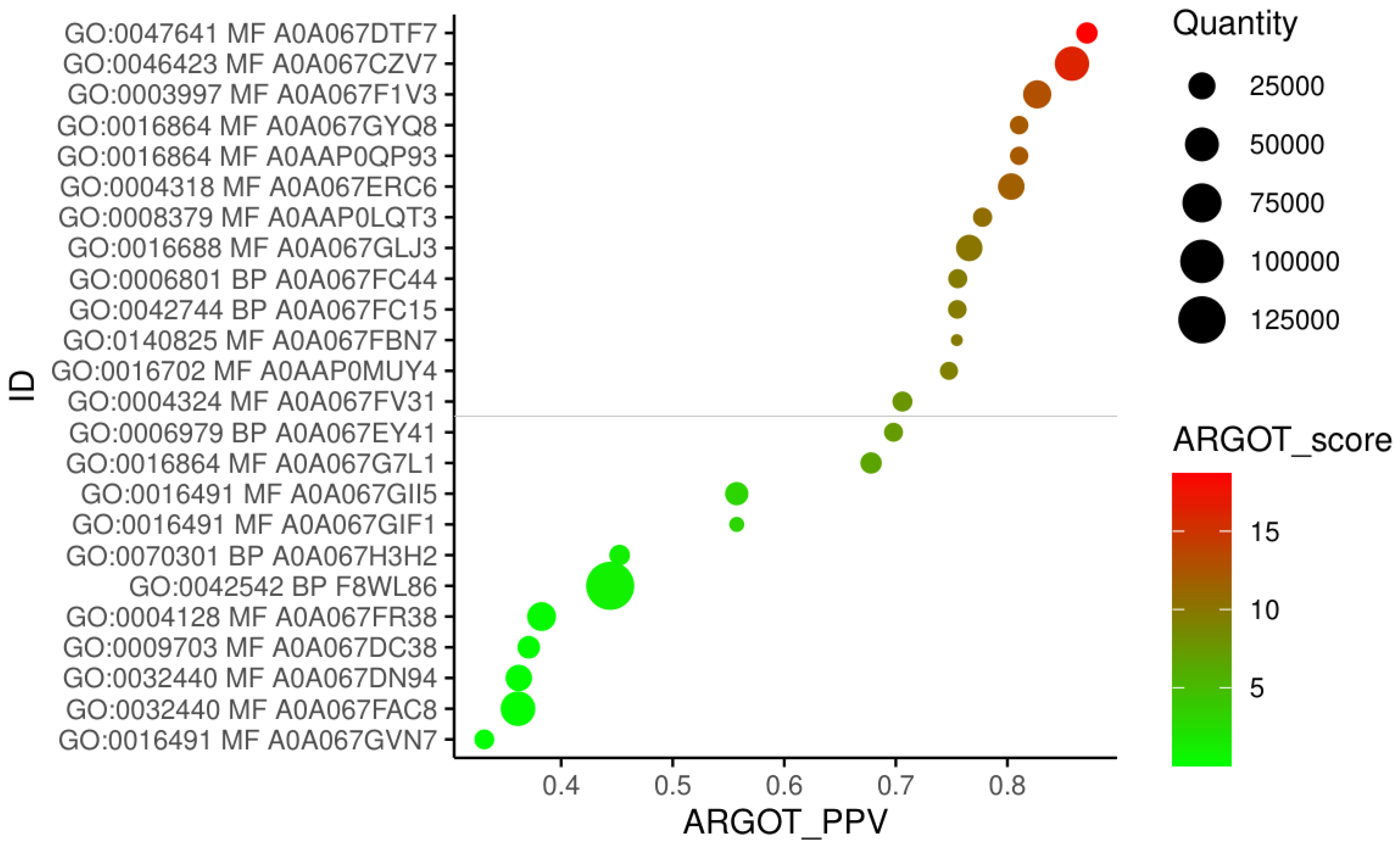
2.3. Proteomic Analysis
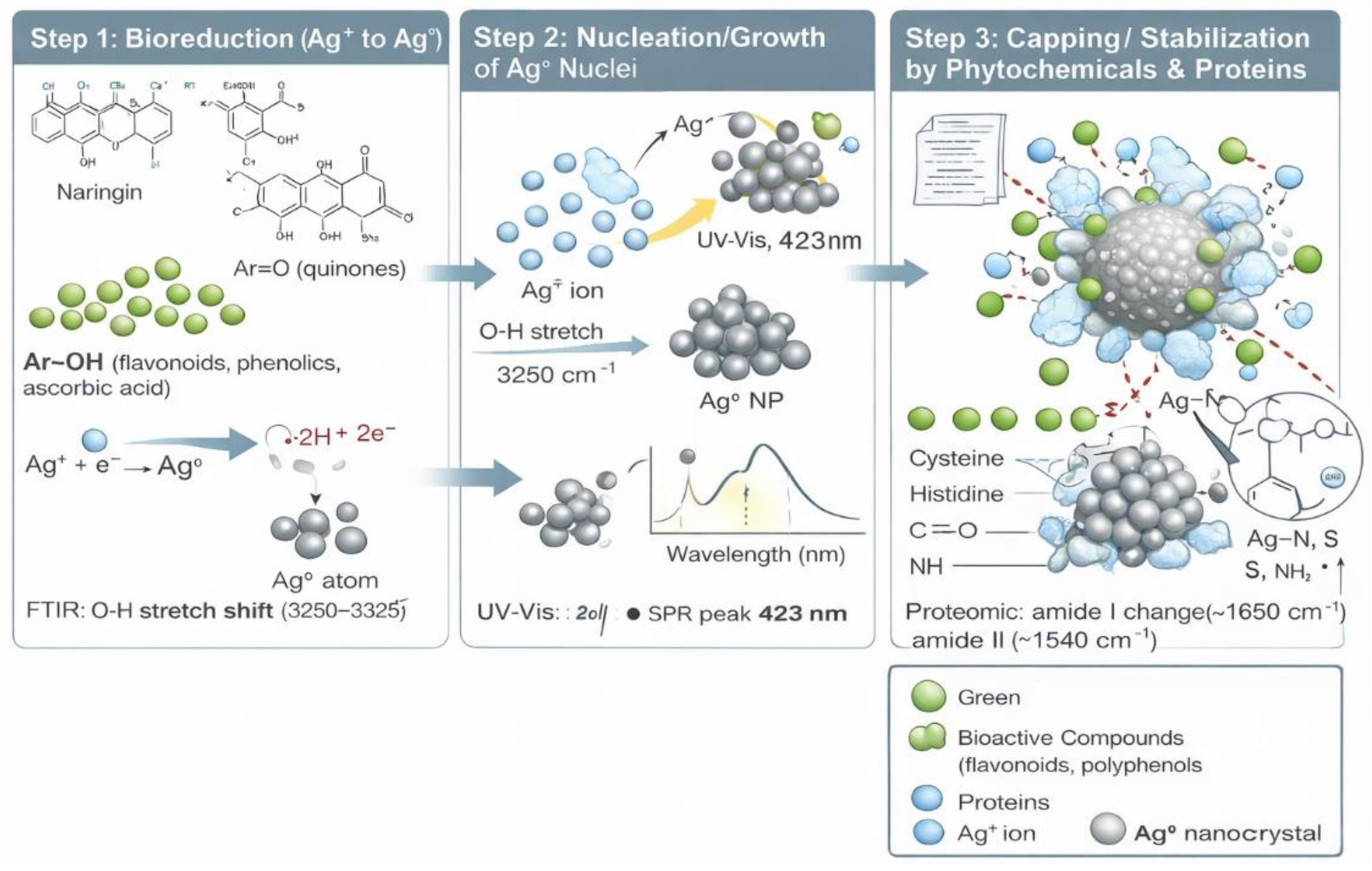
2.4. Putative Mechanism of Ber-AgNPs Synthesis
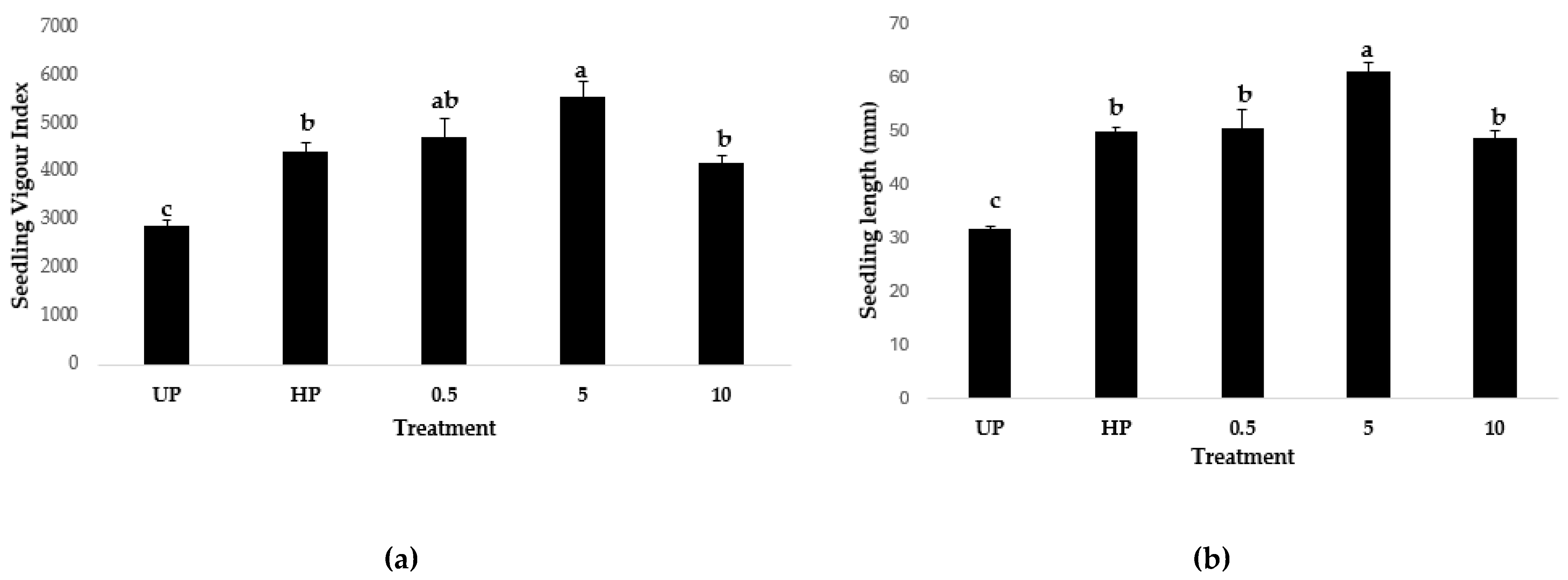
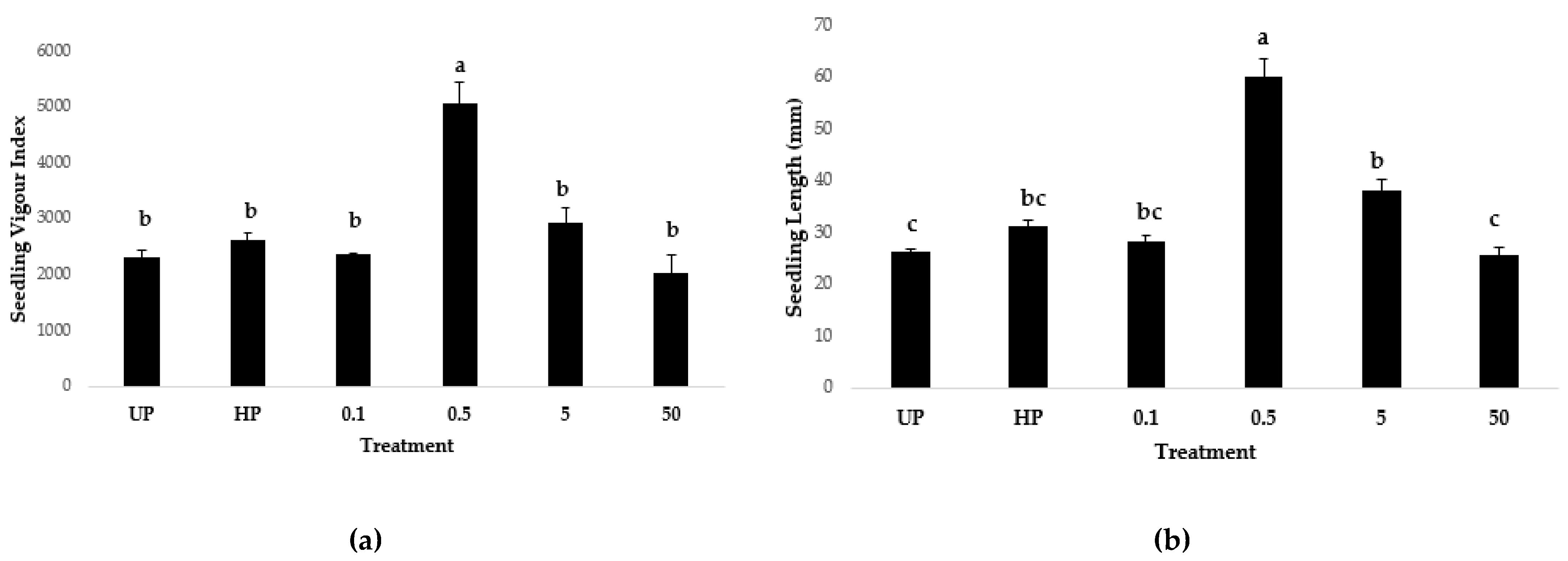
2.5. Seed Nanopriming
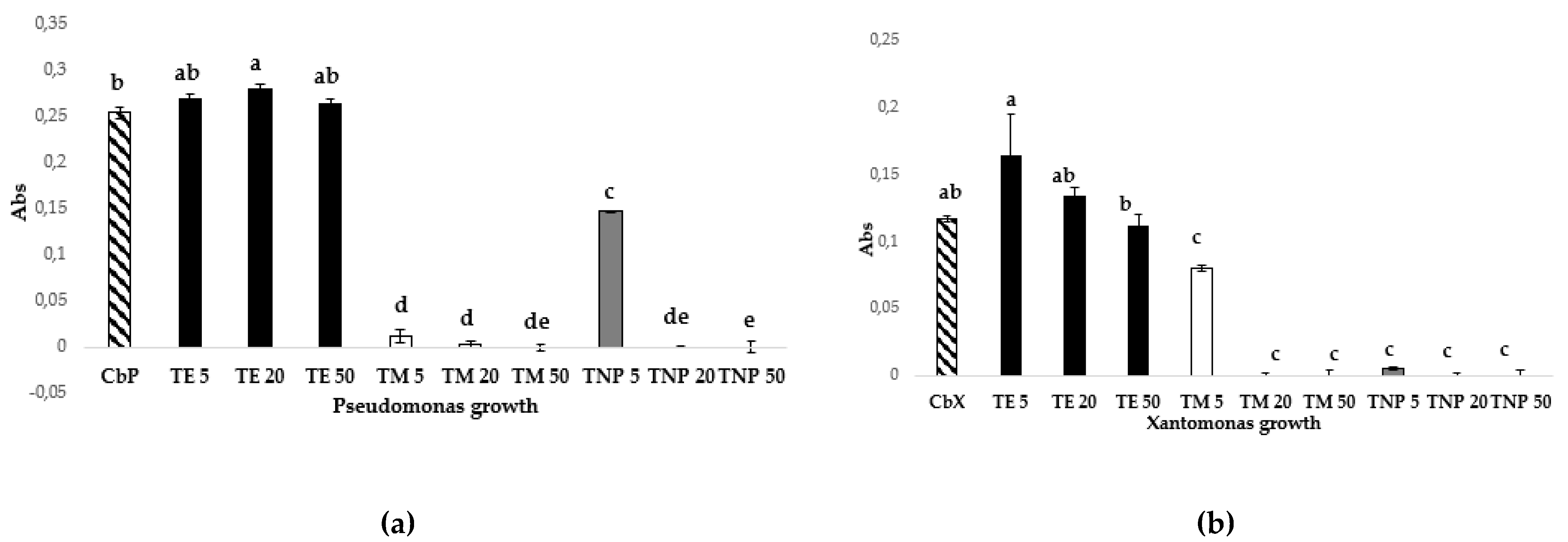
2.6. In Vitro Antibacterial Activity
3. Materials and Methods
3.1. Waste Biomaterial and Chemicals
3.2. Optimization of Reaction Conditions
3.2.1. BPE Concentration
3.2.2. pH
3.2.3. Extract:Metal Ratio
3.2.4. Incubation Time
3.2.5. Incubation Temperature
3.3. Ber-AgNPs Chemical-Physical Characterization
3.4. Proteomic analysis of the Ber-AgNPs
3.4.1. Sample preparation
3.4.2. Protein Aggregation Capture (PAC)
3.4.3. Protein Amount Estimation
3.4.4. LC-MS/MS Analysis
3.4.5. Data-Dependent Acquisition (DDA)
3.4.6. Data-Indipendent Acquisition (DIA)
3.4.7. Functional Annotation and Gene Ontology Analysis of Protein
3.5. Seed Nanopriming
3.5.1. Triphasic Germination Process of the Tomato and Lettuce Seeds
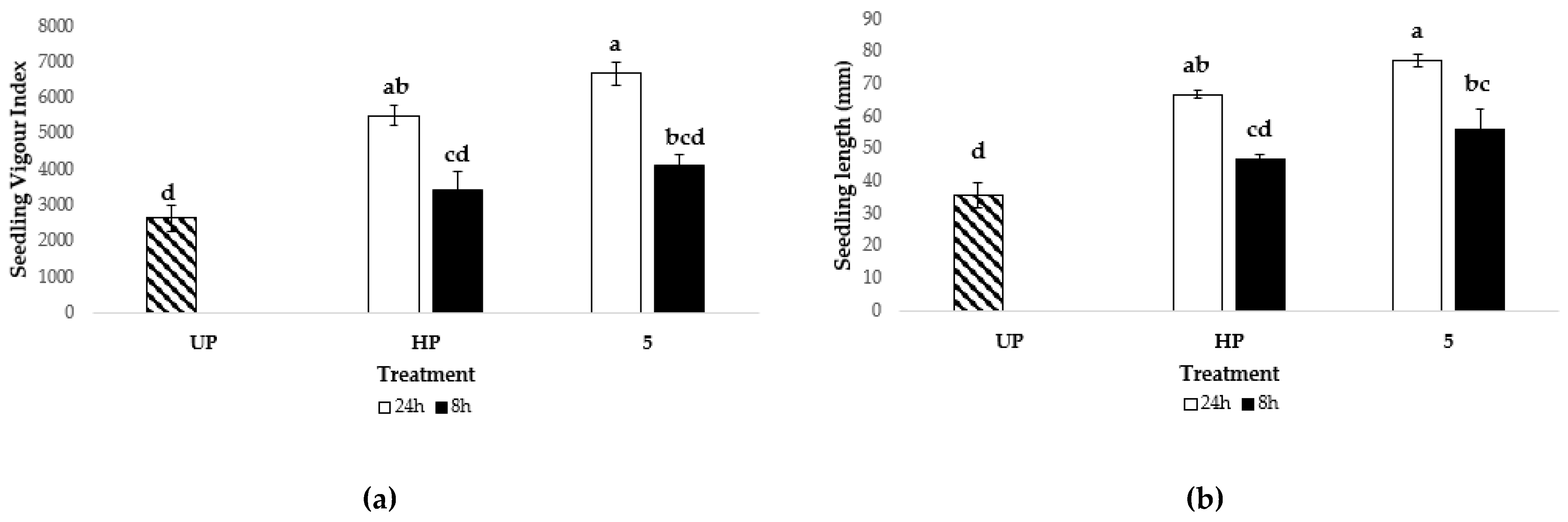
3.5.2. Tomato seed Nanopriming
3.5.2.1. Optimization of Ber-AgNPs Concentration
3.5.2.2. Optimization of Seed Nanopriming Time.
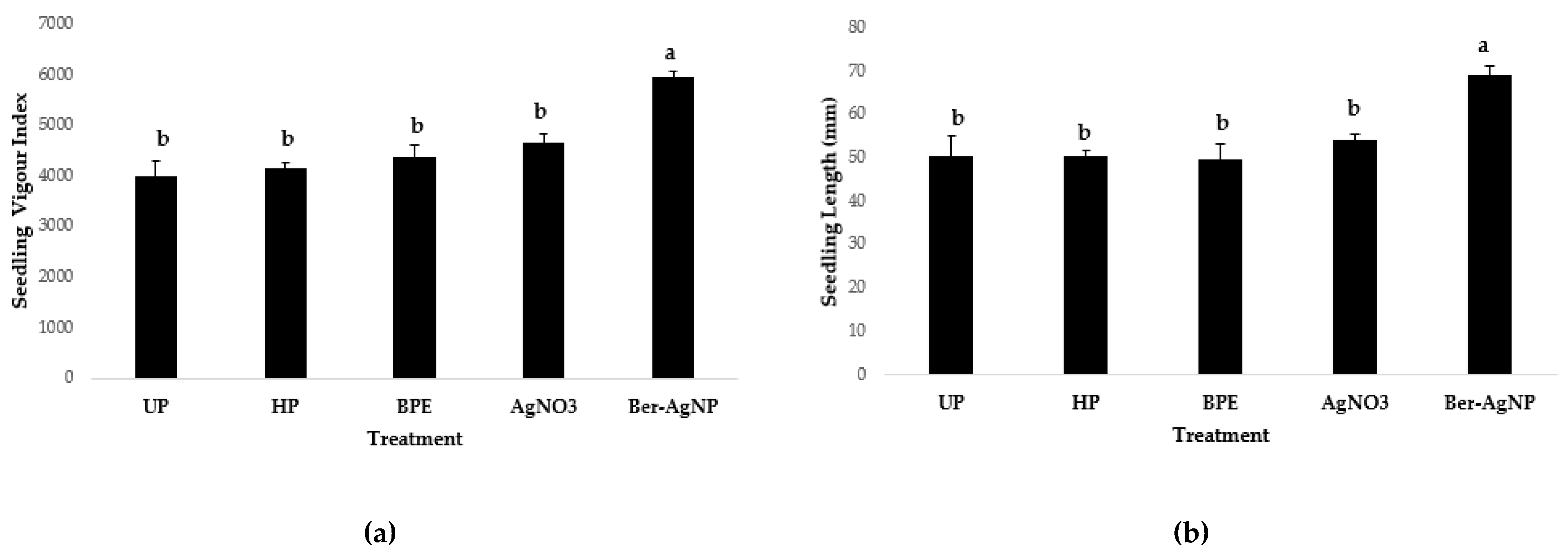
3.5.2.3. Comparison with Metal and BPE Controls
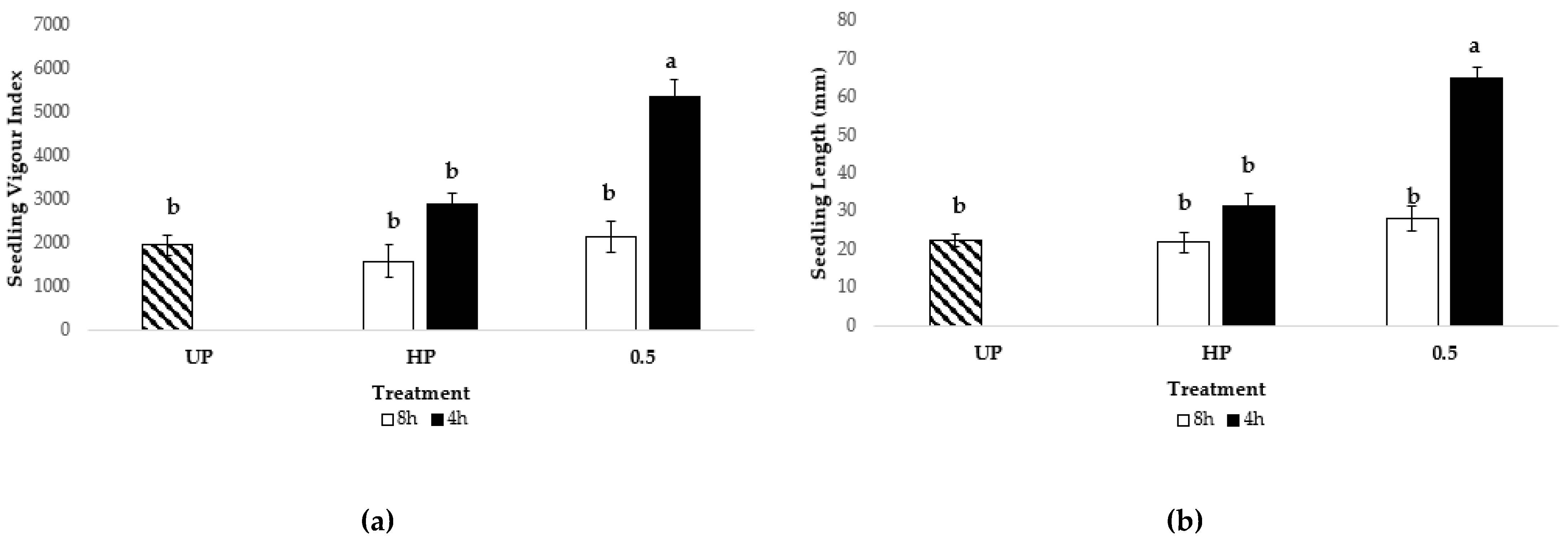
3.5.3. Lettuce Seed Nanopriming
3.5.3.1. Optimization of Ber-AgNPs Concentration
3.5.3.2. Optimization of Seed Nanopriming Time
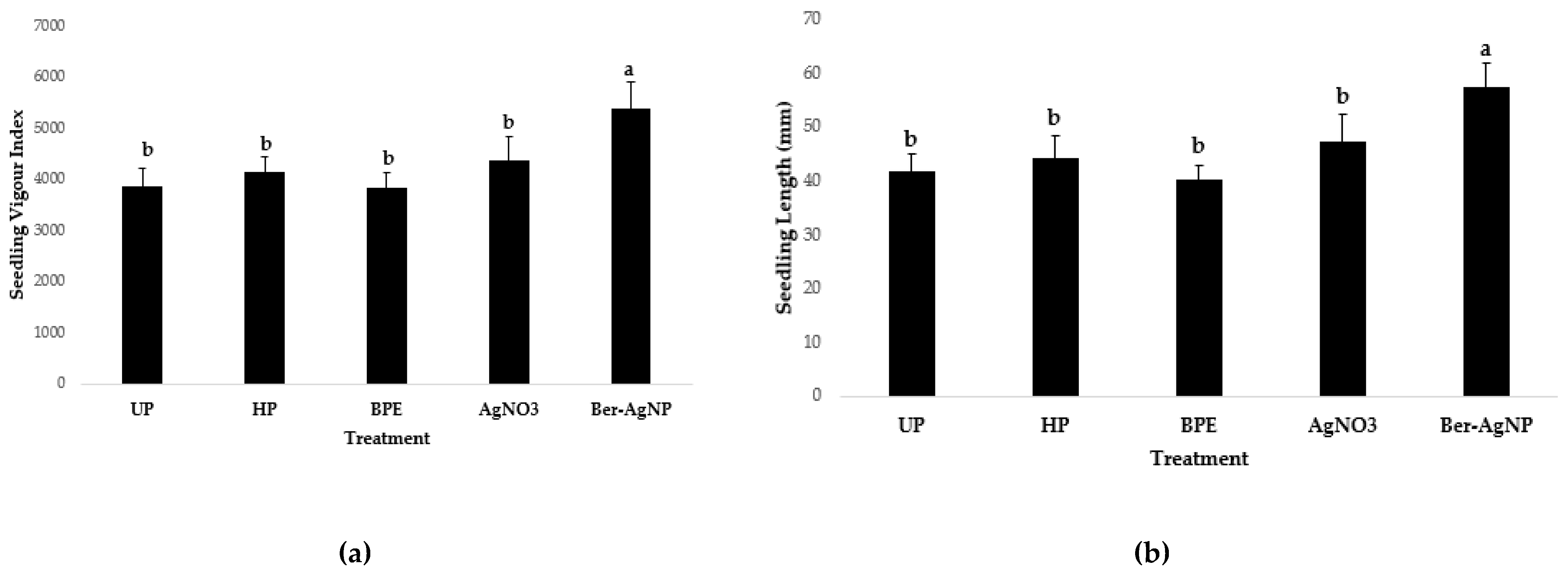
3.5.3.3 Comparison with Metal and Bergamot Pomace Extract Controls
3.6. Statistical Analysis
3.7. Antibacterial Activity of Ber-AgNPs
5. Conclusions
Supplementary Materials
Author Contributions
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
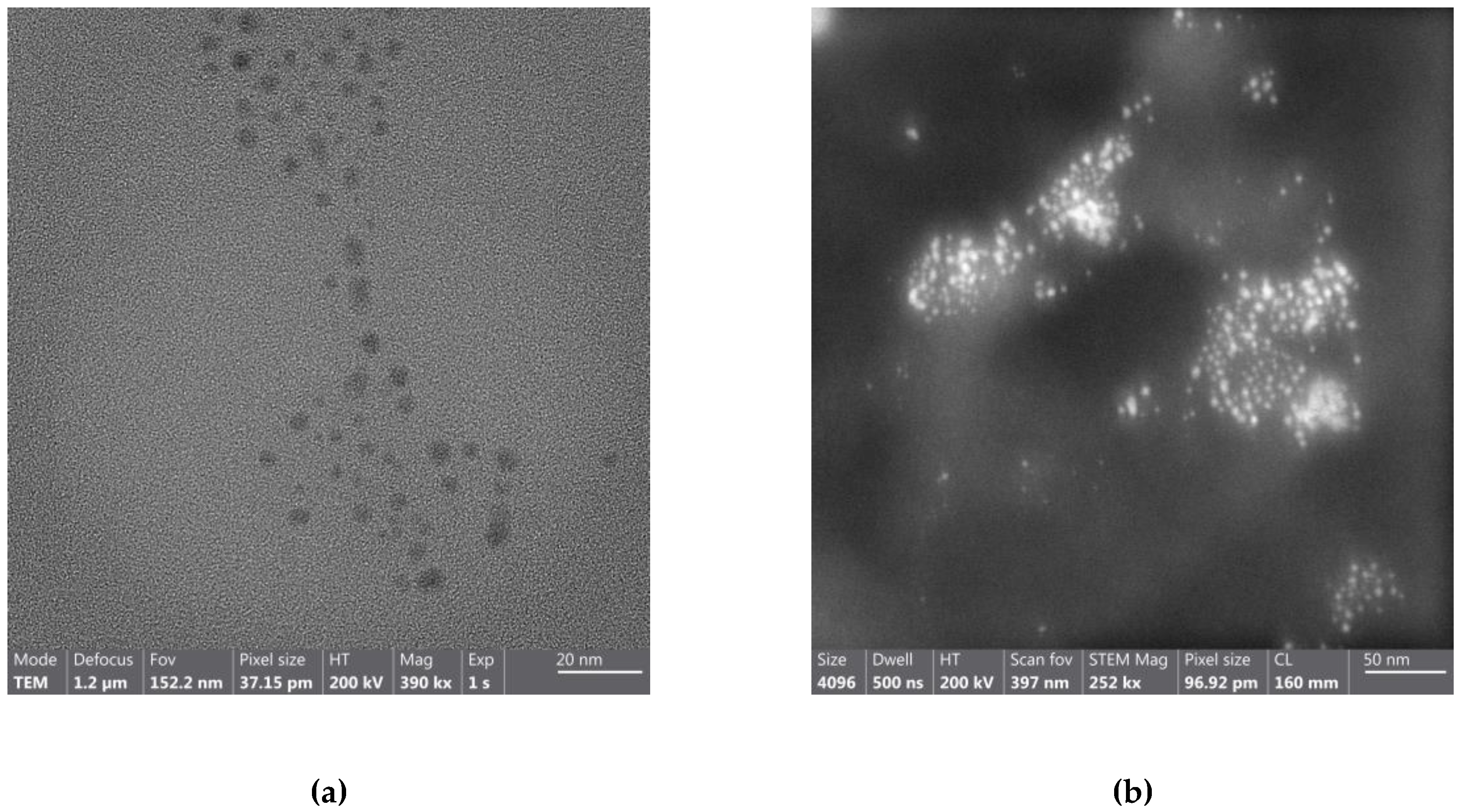
| TEM | Transmission Electron Microscopy |
| STEM | Scanning Transmission Electron Microscopy |
| ATR-FTIR | attenuated total reflectance – Fourier transform infrared spectroscopy |
| NP | Nanoparticle |
| M-NPs | Metal/Metalloids nanoparticles |
| GS-M-NPs | Green synthesis of Metal/Metalloids nanoparticles |
| Ber-AgNPs | bergamot pomace-based silver nanoparticles |
| BPE | bergamot pomace extract |
| SPR | surface plasmon resonance |
| PdI | polydispersity index |
| DLS | Dynamic Light Scattering |
References
- Khan, Y.; Sadia, H.; Ali Shah, S.Z.; Khan, M.N.; Shah, A.A.; Ullah, N.; Ullah, M.F.; Bibi, H.; Bafakeeh, O.T.; Ben Khedher, N.; et al. Classification, Synthetic, and Characterization Approaches to Nanoparticles, and Their Applications in Various Fields of Nanotechnology: A Review. Catalysts 2022, 12. [Google Scholar] [CrossRef]
- Rhaman, M.S.; Tania, S.S.; Imran, S.; Rauf, F.; Kibria, M.G.; Ye, W.; Hasanuzzaman, M.; Murata, Y. Seed Priming with Nanoparticles: An Emerging Technique for Improving Plant Growth, Development, and Abiotic Stress Tolerance. J. Soil Sci. Plant Nutr. 2022, 22, 4047–4062. [Google Scholar] [CrossRef]
- Moulick, R.G.; Das, S.; Debnath, N.; Bandyopadhyay, K. Potential Use of Nanotechnology in Sustainable and ‘Smart’ Agriculture: Advancements Made in the Last Decade. Plant Biotechnol. Rep. 2020, 14, 505–513. [Google Scholar] [CrossRef]
- Miu, B.A.; Dinischiotu, A. New Green Approaches in Nanoparticles Synthesis: An Overview. Molecules 2022, 27. [Google Scholar] [CrossRef] [PubMed]
- Saha, J.; Begum, A.; Mukherjee, A.; Kumar, S. A Novel Green Synthesis of Silver Nanoparticles and Their Catalytic Action in Reduction of Methylene Blue Dye. Sustainable Environment Research 2017, 27, 245–250. [Google Scholar] [CrossRef]
- Rodríguez-Félix, F.; López-Cota, A.G.; Moreno-Vásquez, M.J.; Graciano-Verdugo, A.Z.; Quintero-Reyes, I.E.; Del-Toro-Sánchez, C.L.; Tapia-Hernández, J.A. Sustainable-Green Synthesis of Silver Nanoparticles Using Safflower (Carthamus Tinctorius L.) Waste Extract and Its Antibacterial Activity. Heliyon 2021, 7. [Google Scholar] [CrossRef] [PubMed]
- Flores-Contreras, E.A.; González-González, R.B.; Pablo Pizaña-Aranda, J.J.; Parra-Arroyo, L.; Rodríguez-Aguayo, A.A.; Iñiguez-Moreno, M.; González-Meza, G.M.; Araújo, R.G.; Ramírez-Gamboa, D.; Parra-Saldívar, R.; et al. Agricultural Waste as a Sustainable Source for Nanoparticle Synthesis and Their Antimicrobial Properties for Food Preservation. Frontiers in Nanotechnology 2024, 6. [Google Scholar] [CrossRef]
- Xu, C.; Nasrollahzadeh, M.; Selva, M.; Issaabadi, Z.; Luque, R. Waste-to-Wealth: Biowaste Valorization into Valuable Bio(Nano)Materials.
- Bagheri, S.; Moradi, P.; Nematollahi, F.; Zarinnia, V.; Abdossi, V. Investigating Green Synthesis of Silver Nanoparticles from Orange Peel (Citrus Sinensis) and the Effects of Chitosan, Sylamol, Nanosilver on Rhizopus Stolanifer in Tomato (Solanum Lycopersicum). Indian Journal of Ecology 2020, 47, 82–86. [Google Scholar]
- Pratikno, H.; Anggya, P.B.; Fadhila, F.; Chafidz, A.; Wara, D.P.R.W. Biosynthesis of Silver Nanoparticles Using Banana Raja (Musa Paradisiaca Var. Raja) Peel Extract: Effect of Different Concentrations of the AgNO3 Solution. Key Eng. Mater. 2021, 61–66. [Google Scholar] [CrossRef]
- Chen, Q.; Liu, G.; Chen, G.; Chen, T.; Mi, T. Preparation of Antibacterial Paper 2017, Vol. 12.
- Saratale, R.G.; Saratale, G.D.; Ghodake, G.; Cho, S.K.; Kadam, A.; Kumar, G.; Jeon, B.H.; Pant, D.; Bhatnagar, A.; Shin, H.S. Wheat Straw Extracted Lignin in Silver Nanoparticles Synthesis: Expanding Its Prophecy towards Antineoplastic Potency and Hydrogen Peroxide Sensing Ability. Int. J. Biol. Macromol. 2019, 128, 391–400. [Google Scholar] [CrossRef]
- Anadebe, V.C.; Chukwuike, V.I.; Ebenso, E.E.; Barik, R.C. Chapter 14 - Trends and Perspectives in Waste-Derived Nanoparticles and Circular Economy. Waste-Derived Nanoparticles 2024, 367–379. [Google Scholar]
- Guimarães, R.; Barros, L.; Barreira, J.C.M.; Sousa, M.J.; Carvalho, A.M.; Ferreira, I.C.F.R. Targeting Excessive Free Radicals with Peels and Juices of Citrus Fruits: Grapefruit, Lemon, Lime and Orange. Food and Chemical Toxicology 2010, 48, 99–106. [Google Scholar] [CrossRef] [PubMed]
- Shofinita, D.; Feng, S.; Langrish, T.A.G. Comparing Yields from the Extraction of Different Citrus Peels and Spray Drying of the Extracts. Advanced Powder Technology 2015, 26, 1633–1638. [Google Scholar] [CrossRef]
- Alkhulaifi, M.M.; Alshehri, J.H.; Alwehaibi, M.A.; Awad, M.A.; Al-Enazi, N.M.; Aldosari, N.S.; Hatamleh, A.A.; Abdel- Raouf, N. Green Synthesis of Silver Nanoparticles Using Citrus Limon Peels and Evaluation of Their Antibacterial and Cytotoxic Properties. Saudi J. Biol. Sci. 2020, 27, 3434–3441. [Google Scholar] [CrossRef]
- Gupta, S.; Choudhary, D.K.; Sundaram, S. Green Synthesis and Characterization of Silver Nanoparticles Using Citrus Sinensis (Orange Peel) Extract and Their Antidiabetic, Antioxidant, Antimicrobial and Anticancer Activity. Waste Biomass Valorization 2025, 16, 1101–1114. [Google Scholar] [CrossRef]
- Hussain, M.; Raja, N.I.; Mashwani, Z.U.R.; Naz, F.; Iqbal, M.; Aslam, S. Green Synthesis and Characterisation of Silver Nanoparticles and Their Effects on Antimicrobial Efficacy and Biochemical Profiling in Citrus Reticulata. IET Nanobiotechnol. 2018, 12, 514–519. [Google Scholar] [CrossRef]
- Khane, Y.; Benouis, K.; Albukhaty, S.; Sulaiman, G.M.; Abomughaid, M.M.; Al Ali, A.; Aouf, D.; Fenniche, F.; Khane, S.; Chaibi, W.; et al. Green Synthesis of Silver Nanoparticles Using Aqueous Citrus Limon Zest Extract: Characterization and Evaluation of Their Antioxidant and Antimicrobial Properties. Nanomaterials 2022, 12. [Google Scholar] [CrossRef]
- Mickky, B.; Elsaka, H.; Abbas, M.; Gebreil, A.; Shams Eldeen, R. Orange Peel-Mediated Synthesis of Silver Nanoparticles with Antioxidant and Antitumor Activities. BMC Biotechnol. 2024, 24. [Google Scholar] [CrossRef]
- Saratale, R.G.; Shin, H.S.; Kumar, G.; Benelli, G.; Ghodake, G.S.; Jiang, Y.Y.; Kim, D.S.; Saratale, G.D. Exploiting Fruit Byproducts for Eco-Friendly Nanosynthesis: Citrus × Clementina Peel Extract Mediated Fabrication of Silver Nanoparticles with High Efficacy against Microbial Pathogens and Rat Glial Tumor C6 Cells. Environmental Science and Pollution Research 2018, 25, 10250–10263. [Google Scholar] [CrossRef]
- Saravanan, M.; Vahidi, H.; Medina Cruz, D.; Vernet-Crua, A.; Mostafavi, E.; Stelmach, R.; Webster, T.J.; Ali Mahjoub, M.; Rashedi, M.; Barabadi, H. Emerging Antineoplastic Biogenic Gold Nanomaterials for Breast Cancer Therapeutics: A Systematic Review. Int. J. Nanomedicine 2020, 15, 3577–3595. [Google Scholar] [CrossRef]
- Usman, O.; Mohsin Baig, M.M.; Ikram, M.; Iqbal, T.; Islam, S.; Syed, W.; Al-Rawi, M.B.A.; Naseem, M. Green Synthesis of Metal Nanoparticles and Study Their Anti-Pathogenic Properties against Pathogens Effect on Plants and Animals. Sci. Rep. 2024, 14. [Google Scholar] [CrossRef]
- Mishra, P.; Ahmed, T.; Singh, L. A Comparative Study of Biosynthesized Silver-Nanoparticles from Citrus Maxima Peel, Pulp and Seed: A Special Retrospect for Antimicrobial Activity. J. Pharm. Res. Int. 2021, 454–463. [Google Scholar] [CrossRef]
- Sujitha, M. V.; Kannan, S. Green Synthesis of Gold Nanoparticles Using Citrus Fruits (Citrus Limon, Citrus Reticulata and Citrus Sinensis) Aqueous Extract and Its Characterization. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2013, 102, 15–23. [Google Scholar] [CrossRef]
- Hashem, A.M.; Abuzeid, H.; Kaus, M.; Indris, S.; Ehrenberg, H.; Mauger, A.; Julien, C.M. Green Synthesis of Nanosized Manganese Dioxide as Positive Electrode for Lithium-Ion Batteries Using Lemon Juice and Citrus Peel. Electrochim. Acta 2018, 262, 74–81. [Google Scholar] [CrossRef]
- Vankar, P.S.; Shukla, D. Biosynthesis of Silver Nanoparticles Using Lemon Leaves Extract and Its Application for Antimicrobial Finish on Fabric. Applied Nanoscience (Switzerland) 2012, 2, 163–168. [Google Scholar] [CrossRef]
- Akhter, N.; Batool, M.; Yaqoob, A.; Shahid, M.; Muhammad, F.; Khan, J.; Mudassir, M.A.; Rasheed, M.; Javed, S.; Al Farraj, D.A.; et al. Potential Biological Application of Silver Nanoparticles Synthesized from Citrus Paradisi Leaves. Sci. Rep. 2024, 14. [Google Scholar] [CrossRef]
- Dikshit, P.K.; Kumar, J.; Das, A.K.; Sadhu, S.; Sharma, S.; Singh, S.; Gupta, P.K.; Kim, B.S. Green Synthesis of Metallic Nanoparticles: Applications and Limitations. Catalysts 2021, 11. [Google Scholar] [CrossRef]
- Kolbert, Z.; Szollosi, R.; Rónavári, A.; Molnár, Á. Nanoforms of Essential Metals: From Hormetic Phytoeffects to Agricultural Potential. J. Exp. Bot. 2022, 73, 1825–1840. [Google Scholar] [CrossRef]
- Nile, S.H.; Thiruvengadam, M.; Wang, Y.; Samynathan, R.; Shariati, M.A.; Rebezov, M.; Nile, A.; Sun, M.; Venkidasamy, B.; Xiao, J.; et al. Nano-Priming as Emerging Seed Priming Technology for Sustainable Agriculture—Recent Developments and Future Perspectives. J. Nanobiotechnology 2022, 20. [Google Scholar] [CrossRef] [PubMed]
- Song, K.; He, X. How to Improve Seed Germination with Green Nanopriming. Seed Science and Technology 2021, 49, 81–92. [Google Scholar] [CrossRef]
- Ntolia, A.; Chatzigiannakou, T.; Michailidis, N.; Aggeli, A. A Comprehensive Physicochemical Characterization of Silver Nanoparticles as a Prerequisite for Their Successful Biomedical Applications. Inorganics (Basel). 2025, 13. [Google Scholar] [CrossRef]
- Ahangar, E. The PH Role in Nanotechnology, Electrochemistry, and Nano-Drug Delivery Vol. 41.
- Masarudin, M.J.; Cutts, S.M.; Evison, B.J.; Phillips, D.R.; Pigram, P.J. Factors Determining the Stability, Size Distribution, and Cellular Accumulation of Small, Monodisperse Chitosan Nanoparticles as Candidate Vectors for Anticancer Drug Delivery: Application to the Passive Encapsulation of [14C]-Doxorubicin. Nanotechnol. Sci. Appl. 2015, 8, 67–80. [Google Scholar] [CrossRef]
- Saratale, R.G.; Shin, H.S.; Kumar, G.; Benelli, G.; Ghodake, G.S.; Jiang, Y.Y.; Kim, D.S.; Saratale, G.D. Exploiting Fruit Byproducts for Eco-Friendly Nanosynthesis: Citrus × Clementina Peel Extract Mediated Fabrication of Silver Nanoparticles with High Efficacy against Microbial Pathogens and Rat Glial Tumor C6 Cells. Environmental Science and Pollution Research 2018, 25, 10250–10263. [Google Scholar] [CrossRef]
- Pasieczna-Patkowska, S.; Cichy, M.; Flieger, J. Application of Fourier Transform Infrared (FTIR) Spectroscopy in Characterization of Green Synthesized Nanoparticles. Molecules 2025, 30. [Google Scholar] [CrossRef] [PubMed]
- Nikam, S.A.; Chaudhari, S.P. Biosynthesis of Silver Nanoparticles from Polyphenolic Extract of Baliospermun Solanifolium Using Central Composite Design. Pharmacognosy Res. 2022, 14, 405–411. [Google Scholar] [CrossRef]
- Rautela, A.; Rani, J.; Debnath (Das), M. Green Synthesis of Silver Nanoparticles from Tectona Grandis Seeds Extract: Characterization and Mechanism of Antimicrobial Action on Different Microorganisms. J. Anal. Sci. Technol. 2019, 10. [Google Scholar] [CrossRef]
- Kayed, K.; Issa, M.; Al-ourabi, H. The FTIR Spectra of Ag/Ag2O Composites Doped with Silver Nanoparticles. J. Exp. Nanosci. 2024, 19. [Google Scholar] [CrossRef]
- Malik, M.A.; Batterjee, M.G.; Kamli, M.R.; Alzahrani, K.A.; Danish, E.Y.; Nabi, A. Polyphenol-Capped Biogenic Synthesis of Noble Metallic Silver Nanoparticles for Antifungal Activity against Candida Auris. Journal of Fungi 2022, 8. [Google Scholar] [CrossRef]
- Edo, G.I.; Mafe, A.N.; Ali, A.B.M.; Akpoghelie, P.O.; Yousif, E.; Isoje, E.F.; Igbuku, U.A.; Zainulabdeen, K.; Owheruo, J.O.; Essaghah, A.E.A.; et al. Eco-Friendly Nanoparticle Phytosynthesis via Plant Extracts: Mechanistic Insights, Recent Advances, and Multifaceted Uses. Nano TransMed 2025, 4. [Google Scholar] [CrossRef]
- Hasan, S.M.; Hussain, S.; Yousuf, M.; Tapia-Hernández, J.A.; Raja, D.A. Protein-Based Silver Nanoparticles: Synthesis, Characterization, Administration, and Nanomedicine Applications. Int. J. Biomater. 2025, 2025. [Google Scholar] [CrossRef] [PubMed]
- Chugh, G.; Singh, B.R.; Adholeya, A.; Barrow, C.J. Role of Proteins in the Biosynthesis and Functioning of Metallic Nanoparticles. Crit. Rev. Biotechnol. 2022, 42, 1045–1060. [Google Scholar] [CrossRef] [PubMed]
- Hietzschold, S.; Walter, A.; Davis, C.; Taylor, A.A.; Sepunaru, L. Does Nitrate Reductase Play a Role in Silver Nanoparticle Synthesis? Evidence for NADPH as the Sole Reducing Agent. ACS Sustain. Chem. Eng. 2019, 7, 8070–8076. [Google Scholar] [CrossRef]
- Ahmed, K.B.A.; Senthilnathan, R.; Megarajan, S.; Anbazhagan, V. Sunlight Mediated Synthesis of Silver Nanoparticles Using Redox Phytoprotein and Their Application in Catalysis and Colorimetric Mercury Sensing. J. Photochem. Photobiol. B 2015, 151, 39–45. [Google Scholar] [CrossRef]
- Černý, M.; Habánová, H.; Berka, M.; Luklová, M.; Brzobohatý, B. Hydrogen Peroxide: Its Role in Plant Biology and Crosstalk with Signalling Networks. Int. J. Mol. Sci. 2018, 19. [Google Scholar] [CrossRef]
- Yang, J.; Yue, L.; Yang, Z.; Miao, Y.; Ouyang, R.; Hu, Y. Metal-Based Nanomaterials: Work as Drugs and Carriers against Viral Infections. Nanomaterials 2021, 11. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, Y.M.A.; Elshahawy, I.E. Antifungal Activity of Photo-Biosynthesized Silver Nanoparticles (AgNPs) from Organic Constituents in Orange Peel Extract against Phytopathogenic Macrophomina Phaseolina. Eur. J. Plant Pathol. 2022, 162, 725–738. [Google Scholar] [CrossRef]
- Kodak, M.; Pratim Sarma, M.; Shamim, M.Z.; Bordoloi, N. Green Synthesis Of Silver Nanoparticles Using Citrus Sinensis Peel Extract Vol. 11.
- Trouillas, P.; Marsal, P.; Siri, D.; Lazzaroni, R.; Duroux, J.L. A DFT Study of the Reactivity of OH Groups in Quercetin and Taxifolin Antioxidants: The Specificity of the 3-OH Site. Food Chem. 2006, 97, 679–688. [Google Scholar] [CrossRef]
- Bose, D.; Chatterjee, S. Biogenic Synthesis of Silver Nanoparticles Using Guava (Psidium Guajava) Leaf Extract and Its Antibacterial Activity against Pseudomonas Aeruginosa. Applied Nanoscience (Switzerland) 2016, 6, 895–901. [Google Scholar] [CrossRef]
- Chiorcea-Paquim, A.M. Electrochemistry of Flavonoids: A Comprehensive Review. Int. J. Mol. Sci. 2023, 24. [Google Scholar] [CrossRef] [PubMed]
- Shahidi, F.; Costa De Camargo, A.; Fuentes, J.; Speisky, H.; Shahidi, F.; Costa De Camargo, A.; Fuentes, J. Citation: Speisky, H Revisiting the Oxidation of Flavonoids: Loss, Conservation or Enhancement of Their Antioxidant Properties. 2022. [Google Scholar]
- Karthik, R.; Govindasamy, M.; Chen, S.-M.; Cheng, Y.-H.; Muthukrishnan, P.; Padmavathy, S.; Elangovan, A. Biosynthesis of Silver Nanoparticles by Using Camellia Japonica Leaf Extract for the Electrocatalytic Reduction of Nitrobenzene and Photocatalytic Degradation of Eosin-Y. J. Photochem. Photobiol. B 2017, 170, 164–172. [Google Scholar] [CrossRef]
- Paparella, S.; Araújo, S.S.; Rossi, G.; Wijayasinghe, M.; Carbonera, D.; Balestrazzi, A. Seed Priming: State of the Art and New Perspectives. Plant Cell Rep. 2015, 34, 1281–1293. [Google Scholar] [CrossRef]
- Lutts, S.; Benincasa, P.; Wojtyla, L.; Kubala, S.; Pace, R.; Lechowska, K.; Quinet, M.; Garnczarska, M. Seed Priming: New Comprehensive Approaches for an Old Empirical Technique. In New Challenges in Seed Biology - Basic and Translational Research Driving Seed Technology; InTech, 2016. [Google Scholar]
- Charachimwe, R.R.; Chandiposha, M.; Manjeru, P. The Effect of Seed Priming to Improve Germination Parameters and Early Growth of Chickpea (Cicer Arietnum L). International Journal of Agronomy 2023. [Google Scholar] [CrossRef]
- Ansari, M.; Ahmed, S.; Abbasi, A.; Khan, M.T.; Subhan, M.; Bukhari, N.A.; Hatamleh, A.A.; Abdelsalam, N.R. Plant Mediated Fabrication of Silver Nanoparticles, Process Optimization, and Impact on Tomato Plant. Sci. Rep. 2023, 13. [Google Scholar] [CrossRef] [PubMed]
- Rutkowski, M.; Krzemińska-Fiedorowicz, L.; Khachatryan, G.; Bulski, K.; Kołton, A.; Khachatryan, K. Biodegradable Silver Nanoparticles Gel and Its Impact on Tomato Seed Germination Rate in In Vitro Cultures. Applied Sciences (Switzerland) 2022, 12. [Google Scholar] [CrossRef]
- Sonawane, H.; Arya, S.; Math, S.; Shelke, D. Myco-Synthesized Silver and Titanium Oxide Nanoparticles as Seed Priming Agents to Promote Seed Germination and Seedling Growth of Solanum Lycopersicum: A Comparative Study. Int. Nano Lett. 2021, 11, 371–379. [Google Scholar] [CrossRef]
- Antunes, D.R.; Forini, M.M.L.H.; Coqueiro, Y.A.; Pontes, M.S.; Lima, P.H.C.; Cavalcante, L.A.F.; Sanches, A.O.; Caires, A.R.L.; Santiago, E.F.; Grillo, R. Effect of Hyaluronic Acid-Stabilized Silver Nanoparticles on Lettuce (Lactuca Sativa L.) Seed Germination. Chemosphere 2024, 364. [Google Scholar] [CrossRef]
- Méndez-Andrade, R.; Ruiz-Torres, N.A.; Wang, Y.; García-Cerda, L.A.; Vera-Reyes, I. Enhanced Tomato Seed Germination and Seedling Growth Through Oxidative Stress Using Biogenic Silver Nanoparticles. Terra Latinoamericana 2024, 42, 1–10. [Google Scholar] [CrossRef]
- Yang, L.; Zhang, L.; Zhang, Q.; Wei, J.; Zhao, X.; Zheng, Z.; Chen, B.; Xu, Z. Nanopriming Boost Seed Vigor: Deeper Insights into the Effect Mechanism. Plant Physiology and Biochemistry 2024, 214. [Google Scholar] [CrossRef]
- Karami Mehrian, S.; Heidari, R.; Rahmani, F.; Najafi, S. Effect of Chemical Synthesis Silver Nanoparticles on Germination Indices and Seedlings Growth in Seven Varieties of Lycopersicon Esculentum Mill (Tomato) Plants. J. Clust. Sci. 2016, 27, 327–340. [Google Scholar] [CrossRef]
- Bano, N.; Khan, S.; Hamid, Y.; Ullah, M.A.; Khan, A.G.; Bano, F.; Luo, J.; Li, T. Effect of Foliar Application of Nanoparticles on Growth, Physiology, and Antioxidant Enzyme Activities of Lettuce (Lactuca Sativa L.) Plants under Cadmium Toxicity. Environmental Science and Pollution Research 2023, 30, 99310–99325. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.; Xiang, S.; Lv, X.; Wang, X.; Li, F.; Liu, W.; Liu, C.; Ran, M.; Huang, J.; Xu, X.; et al. Biosynthesized Silver Nanoparticles Inhibit Pseudomonas Syringae Pv. Tabaci by Directly Destroying Bacteria and Inducing Plant Resistance in Nicotiana Benthamiana. Phytopathology Research 2022, 4. [Google Scholar] [CrossRef]
- Li, M.; Lei, T.; Li, J.; Li, X.; Wei, Z. Antibacterial Activity of Silver Nanoparticles against Xanthomonas Compestris Var. Compestris. ScienceAsia 2025, 51. [Google Scholar] [CrossRef]
- Khalifa, H.O.; Oreiby, A.; Mohammed, T.; Abdelhamid, M.A.A.; Sholkamy, E.N.; Hashem, H.; Fereig, R.M. Silver Nanoparticles as Next-Generation Antimicrobial Agents: Mechanisms, Challenges, and Innovations against Multidrug-Resistant Bacteria. Front. Cell. Infect. Microbiol. 2025, 15. [Google Scholar] [CrossRef]
- Loo, Y.Y.; Rukayadi, Y.; Nor-Khaizura, M.A.R.; Kuan, C.H.; Chieng, B.W.; Nishibuchi, M.; Radu, S. In Vitro Antimicrobial Activity of Green Synthesized Silver Nanoparticles against Selected Gram-Negative Foodborne Pathogens. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef]
- Zhang, H.; Zheng, T.; Wang, Y.; Li, T.; Chi, Q. Multifaceted Impacts of Nanoparticles on Plant Nutrient Absorption and Soil Microbial Communities. Front. Plant Sci. 2024, 15. [Google Scholar] [CrossRef] [PubMed]
- Mikhailova, E.O. Silver Nanoparticles: Mechanism of Action and Probable Bio-Application. J. Funct. Biomater. 2020, 11. [Google Scholar] [CrossRef]
- Zheng, S.; Zhou, Q.; Chen, C.; Yang, F.; Cai, Z.; Li, D.; Geng, Q.; Feng, Y.; Wang, H. Role of Extracellular Polymeric Substances on the Behavior and Toxicity of Silver Nanoparticles and Ions to Green Algae Chlorella Vulgaris. Science of the Total Environment 2019, 660, 1182–1190. [Google Scholar] [CrossRef]
- Elfadadny, A.; Ragab, R.F.; AlHarbi, M.; Badshah, F.; Ibáñez-Arancibia, E.; Farag, A.; Hendawy, A.O.; De los Ríos-Escalante, P.R.; Aboubakr, M.; Zakai, S.A.; et al. Antimicrobial Resistance of Pseudomonas Aeruginosa: Navigating Clinical Impacts, Current Resistance Trends, and Innovations in Breaking Therapies. Front. Microbiol. 2024, 15. [Google Scholar] [CrossRef] [PubMed]
- Kamer, A.M.A.; El Maghraby, G.M.; Shafik, M.M.; Al-Madboly, L.A. Silver Nanoparticle with Potential Antimicrobial and Antibiofilm Efficiency against Multiple Drug Resistant, Extensive Drug Resistant Pseudomonas Aeruginosa Clinical Isolates. BMC Microbiol. 2024, 24. [Google Scholar] [CrossRef]
- Trzcińska-Wencel, J.; Wypij, M.; Terzyk, A.P.; Rai, M.; Golińska, P. Biofabrication of Novel Silver and Zinc Oxide Nanoparticles from Fusarium Solani IOR 825 and Their Potential Application in Agriculture as Biocontrol Agents of Phytopathogens, and Seed Germination and Seedling Growth Promoters. Front. Chem. 2023, 11. [Google Scholar] [CrossRef] [PubMed]
- More, P.R.; Pandit, S.; De Filippis, A.; Franci, G.; Mijakovic, I.; Galdiero, M. Silver Nanoparticles: Bactericidal and Mechanistic Approach against Drug Resistant Pathogens. Microorganisms 2023, 11. [Google Scholar] [CrossRef] [PubMed]
- Shahid, A.; Faizan, M.; Raza, M.A. POTENTIAL ROLE OF SILVER NANOPARTICLES (AgNPs) AND ZINC NANOPARTICLES (ZnNPs) FOR PLANT DISEASE MANAGEMENT. Agrobiological Records 2023, 14, 59–69. [Google Scholar] [CrossRef]
- Rodrigues, A.S.; Batista, J.G.S.; Rodrigues, M.Á.V.; Thipe, V.C.; Minarini, L.A.R.; Lopes, P.S.; Lugão, A.B. Advances in Silver Nanoparticles: A Comprehensive Review on Their Potential as Antimicrobial Agents and Their Mechanisms of Action Elucidated by Proteomics. Front. Microbiol. 2024, 15. [Google Scholar] [CrossRef]
- Prathna, T.C.; Chandrasekaran, N.; Raichur, A.M.; Mukherjee, A. Biomimetic Synthesis of Silver Nanoparticles by Citrus Limon (Lemon) Aqueous Extract and Theoretical Prediction of Particle Size. Colloids Surf. B Biointerfaces 2011, 82, 152–159. [Google Scholar] [CrossRef]
- Kaviya, S.; Santhanalakshmi, J.; Viswanathan, B.; Muthumary, J.; Srinivasan, K. Biosynthesis of Silver Nanoparticles Using Citrus Sinensis Peel Extract and Its Antibacterial Activity. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2011, 79, 594–598. [Google Scholar] [CrossRef]
- Dell’Annunziata, F.; Mosidze, E.; Folliero, V.; Lamparelli, E.P.; Lopardo, V.; Pagliano, P.; Porta, G. Della; Galdiero, M.; Bakuridze, A.D.; Franci, G. Eco-Friendly Synthesis of Silver Nanoparticles from Peel and Juice C. Limon and Their Antiviral Efficacy against HSV-1 and SARS-CoV-2. Virus Res. 2024, 349. [Google Scholar] [CrossRef]
- Niluxsshun, M.C.D.; Masilamani, K.; Mathiventhan, U. Green Synthesis of Silver Nanoparticles from the Extracts of Fruit Peel of Citrus Tangerina, Citrus Sinensis, and Citrus Limon for Antibacterial Activities. Bioinorg. Chem. Appl. 2021, 2021. [Google Scholar] [CrossRef]
- Vi Thien Thao, N.; Thi Mai, N.; Do Thanh Thang, T.; Huy Vu, P.; Thi Luong, N.; Thi Hong Thuy, L. OPTIMISATION OF THE GREEN SYNTHESIS PROCESS OF COPPER NANOPARTICLES USING LIME (Citrus Aurantifolia) EXTRACT USING RESPONSE SURFACE METHODOLOGY WITH THE BOX-BEHNKEN MODEL; 2025; Vol. 25. [Google Scholar]
- Chand, K.; Cao, D.; Eldin Fouad, D.; Hussain Shah, A.; Qadeer Dayo, A.; Zhu, K.; Nazim Lakhan, M.; Mehdi, G.; Dong, S. Green Synthesis, Characterization and Photocatalytic Application of Silver Nanoparticles Synthesized by Various Plant Extracts. Arabian Journal of Chemistry 2020, 13, 8248–8261. [Google Scholar] [CrossRef]
- Batth, T.S.; Tollenaere, M.A.X.; Rüther, P.; Gonzalez-Franquesa, A.; Prabhakar, B.S.; Bekker-Jensen, S.; Deshmukh, A.S.; Olsen, J. V. Protein Aggregation Capture on Microparticles Enables Multipurpose Proteomics Sample Preparation. Molecular and Cellular Proteomics 2019, 18, 1027–1035. [Google Scholar] [CrossRef] [PubMed]
- Törönen, P.; Medlar, A.; Holm, L. PANNZER2: A Rapid Functional Annotation Web Server. Nucleic Acids Res. 2018, 46, W84–W88. [Google Scholar] [CrossRef] [PubMed]
- Supek, F.; Bošnjak, M.; Škunca, N.; Šmuc, T. Revigo Summarizes and Visualizes Long Lists of Gene Ontology Terms. PLoS One 2011, 6. [Google Scholar] [CrossRef]
- Abenavoli, M.R.; Cacco, G.; Sorgonà, A.; Marabottini, R.; Paolacci, A.R.; Ciaffi, M.; Badiani, M. The Inhibitory Effects of Coumarin on the Germination of Durum Wheat (Triticum Turgidum Ssp. Durum, CV. Simeto) Seeds. J. Chem. Ecol. 2006, 32, 489–506. [Google Scholar] [CrossRef] [PubMed]
- Bewley, J.D.; Bradford, K.J.; Hilhorst, H.W.M.; Nonogaki, H. Seeds: Physiology of Development, Germination and Dormancy; 2013. [Google Scholar]
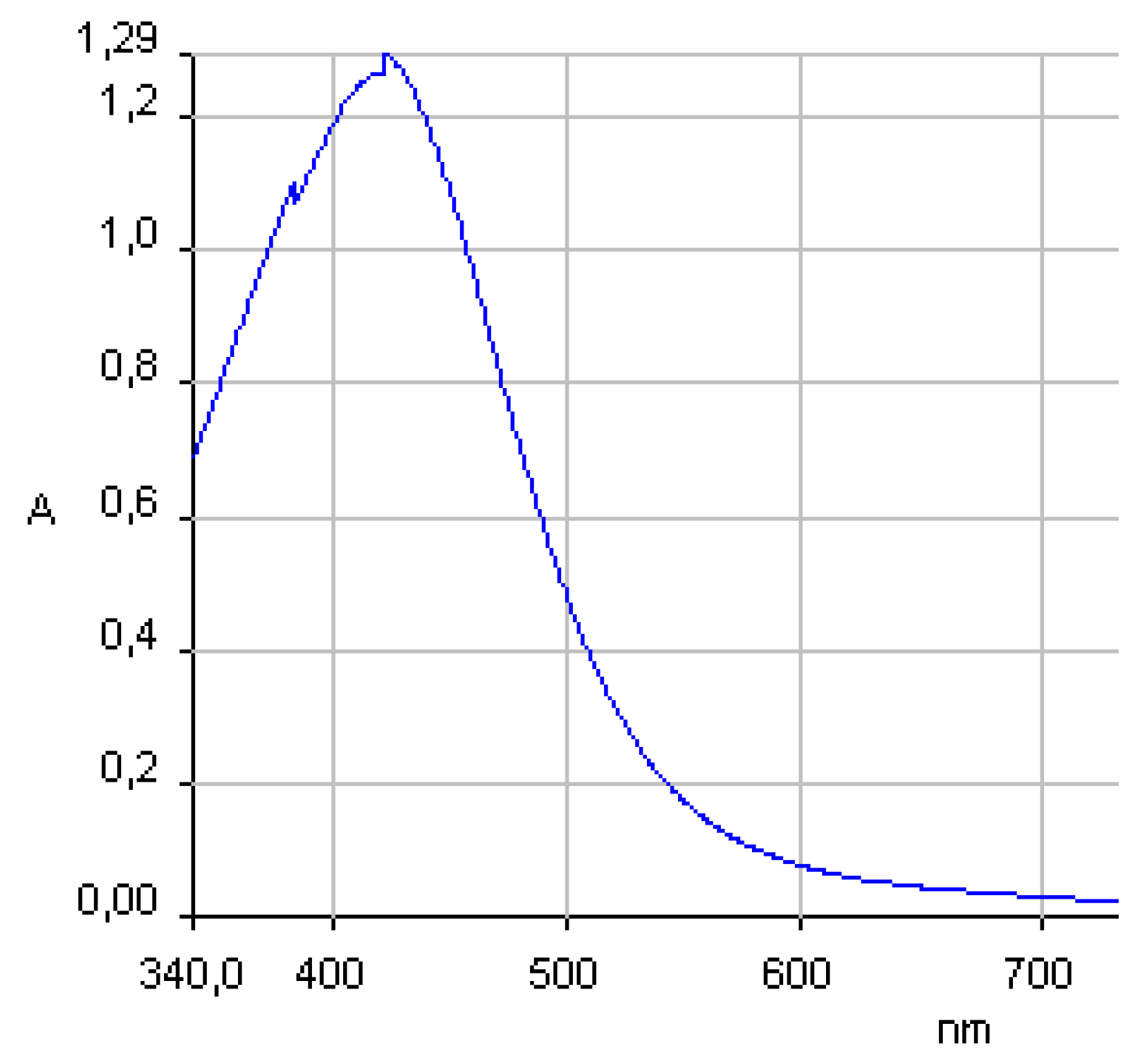

| Sample | Z-Average (d.nm) | PdI | Z-potential (mV) |
| BPE1 | 91.06 | 0.296 | -22.3 |
| BPE0.1 | 89.44 | 0.187 | -29.7 |
| Sample | Z-Average (d.nm) | PdI | Z-potential (mV) |
| pH7 | 115.8 | 0.237 | -31.3 |
| pH8.5 | 86.81 | 0.289 | -41.4 |
| pH9 | 86.08 | 0.262 | -34.1 |
| Sample | Z-Average (d.nm) | PdI | Z-potential (mV) |
| 2:1 ratio | 86.35 | 0.238 | -30.9 |
| 1:1 ratio | 89.44 | 0.187 | -29.7 |
| Sample | Z-Average (d.nm) | PdI | Z-potential (mV) |
| t1 | 53.35 | 0.285 | -32.3 |
| t3 | 46.57 | 0.371 | -33 |
| t24 | 58.81 | 0.268 | -32.5 |
| Sample | Z-Average (d.nm) | PdI | Z-potential |
| T15 | 30.56 | 0.518 | -22.8 |
| T20 | 41 | 0.507 | -32.6 |
| T25 | 50 | 0.313 | -30.4 |
| Sample | Z-Average (d.nm) | PdI | Z-potential |
| Ber-AgNPs | 74.9 | 0.248 | -32.74 |
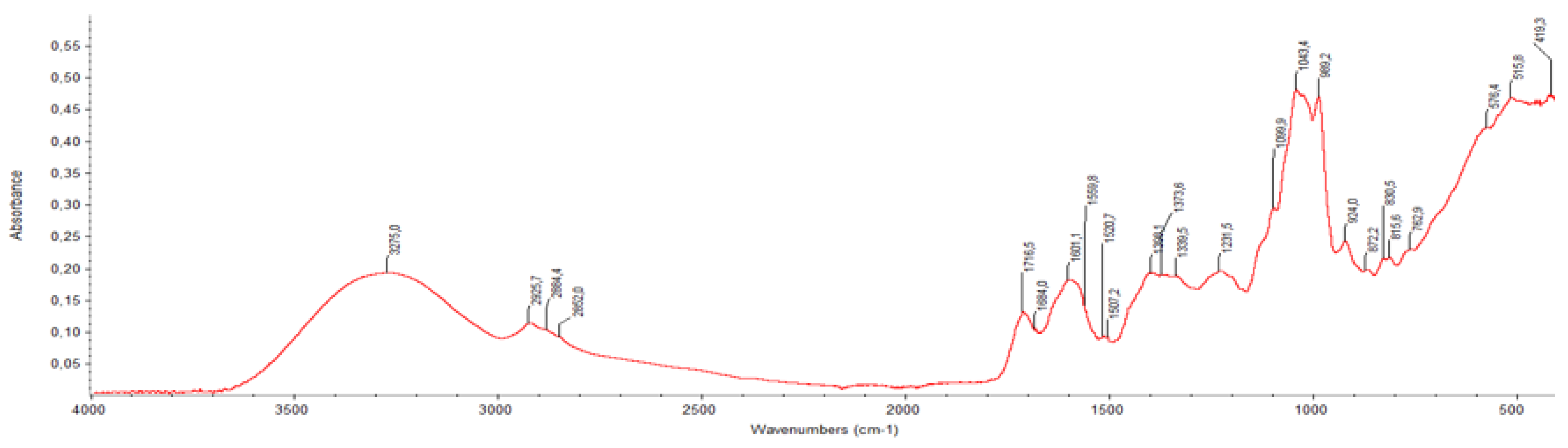
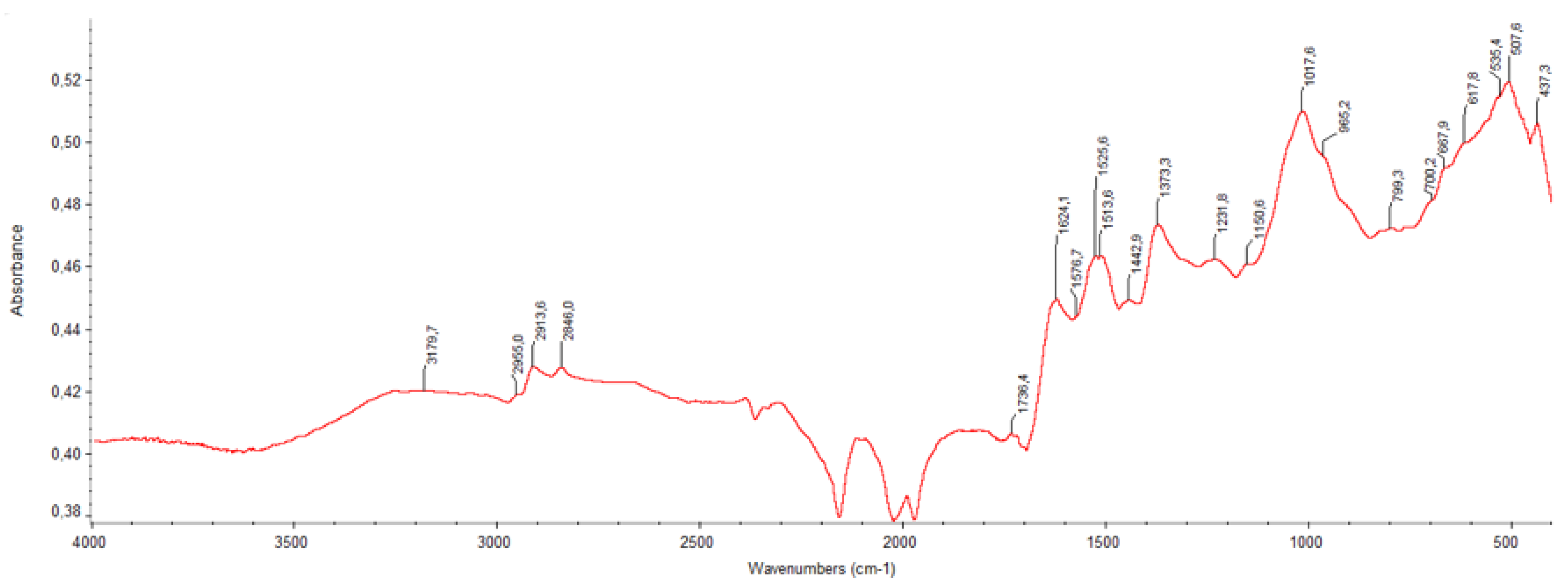
| Wavenumber (cm⁻¹) | BPE | Ber-Ag-NPs | Compound Assignment / Notes |
| ~3250 | Broad, intense | Slightly shifted/sharper | O–H stretching (phenols, flavonoids, alcohols) |
| 2923, 2854 | Present | Decreased intensity | C–H stretching (aliphatic –CH₂/–CH₃) |
| 1710–1690 | Strong | Shifted (~1680) | C=O stretching (carboxylic acids, ketones, esters) |
| 1610–1580 | Medium-strong | Slightly reduced | C=C aromatic ring stretching; possibly conjugated C=O |
| 1454–1430 | Visible | Less intense | CH₂ bending, asymmetric COO⁻ stretching |
| 1384 | Distinct band | Intensity drop | Phenolic O–H bending; COO⁻ symmetric stretch |
| 1323–1245 | Multiple peaks | Slight shift/loss | C–O stretching (phenols, ethers); C–N in proteins |
| 1170–1030 | Complex region | Peak displacement | C–O–C ether linkages; C–O phenolic and alcoholic groups |
| ~698–617 | Weak/moderate | New/shifted peaks | Possible Ag–O or Ag–N interactions (Ag coordination signals) |
| ~516 | Weak shoulder | Slightly intensified | Fingerprint region; may include metal–oxygen stretching (Ag–O) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




