Submitted:
08 February 2026
Posted:
10 February 2026
You are already at the latest version
Abstract
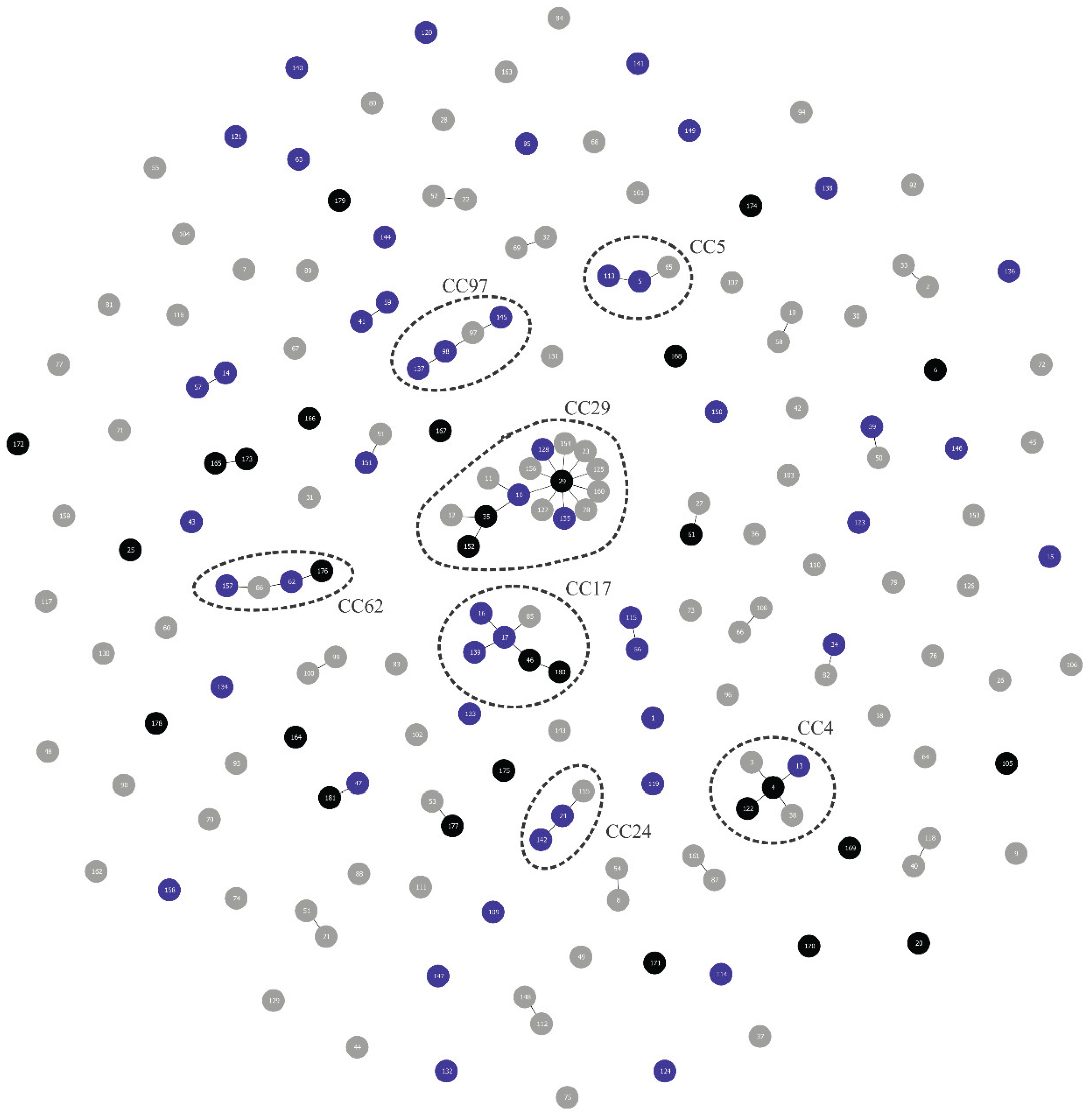
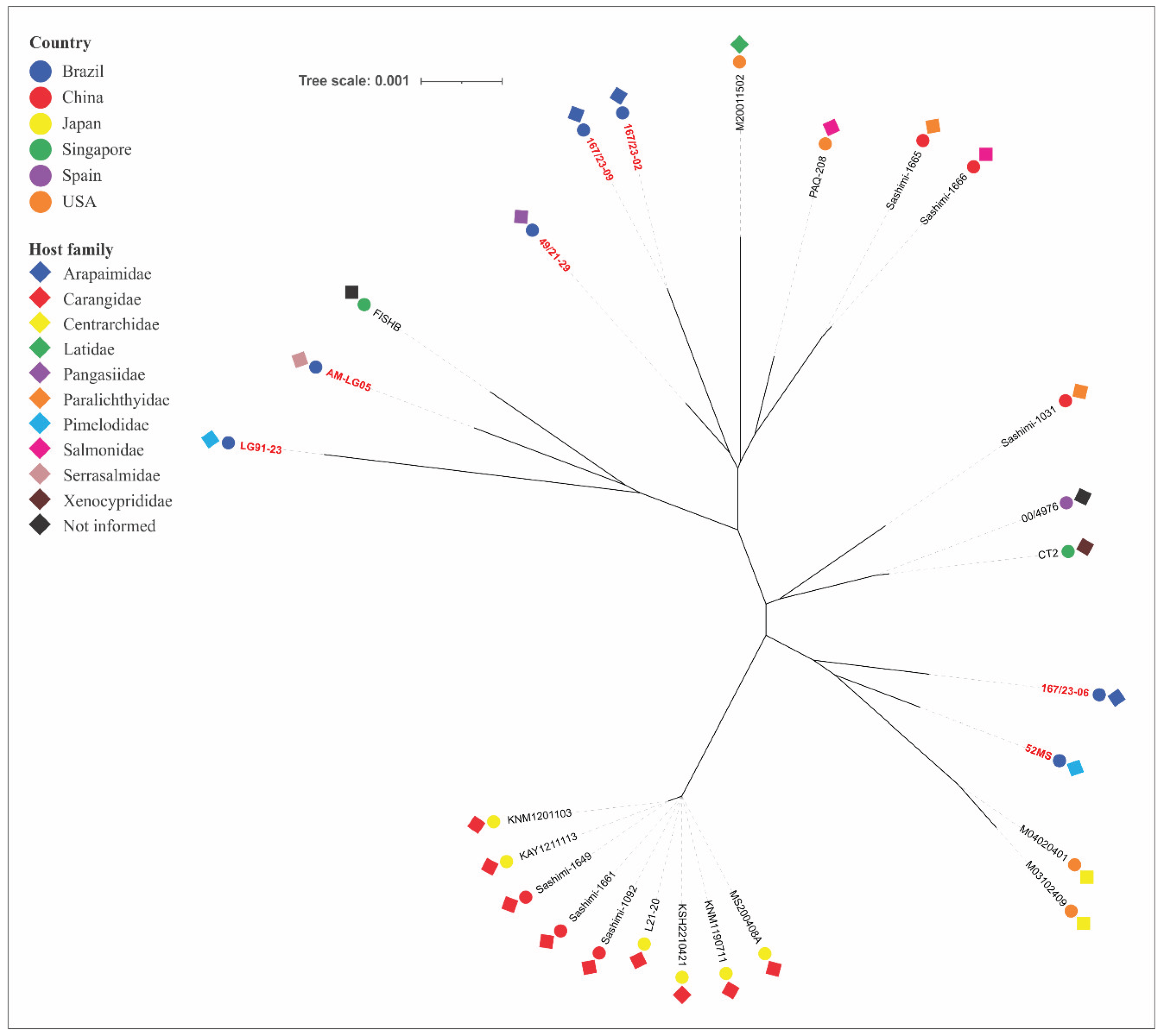
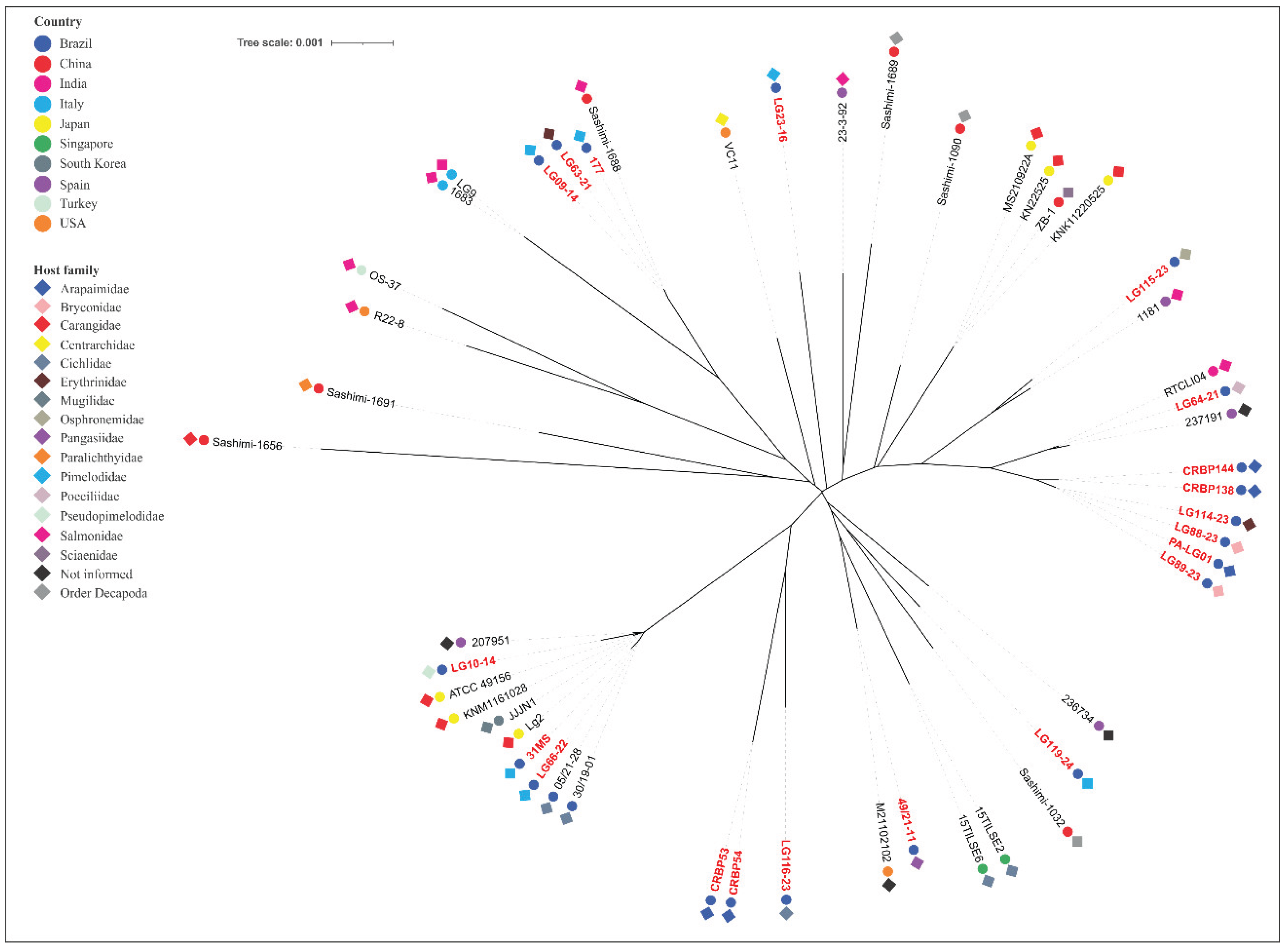
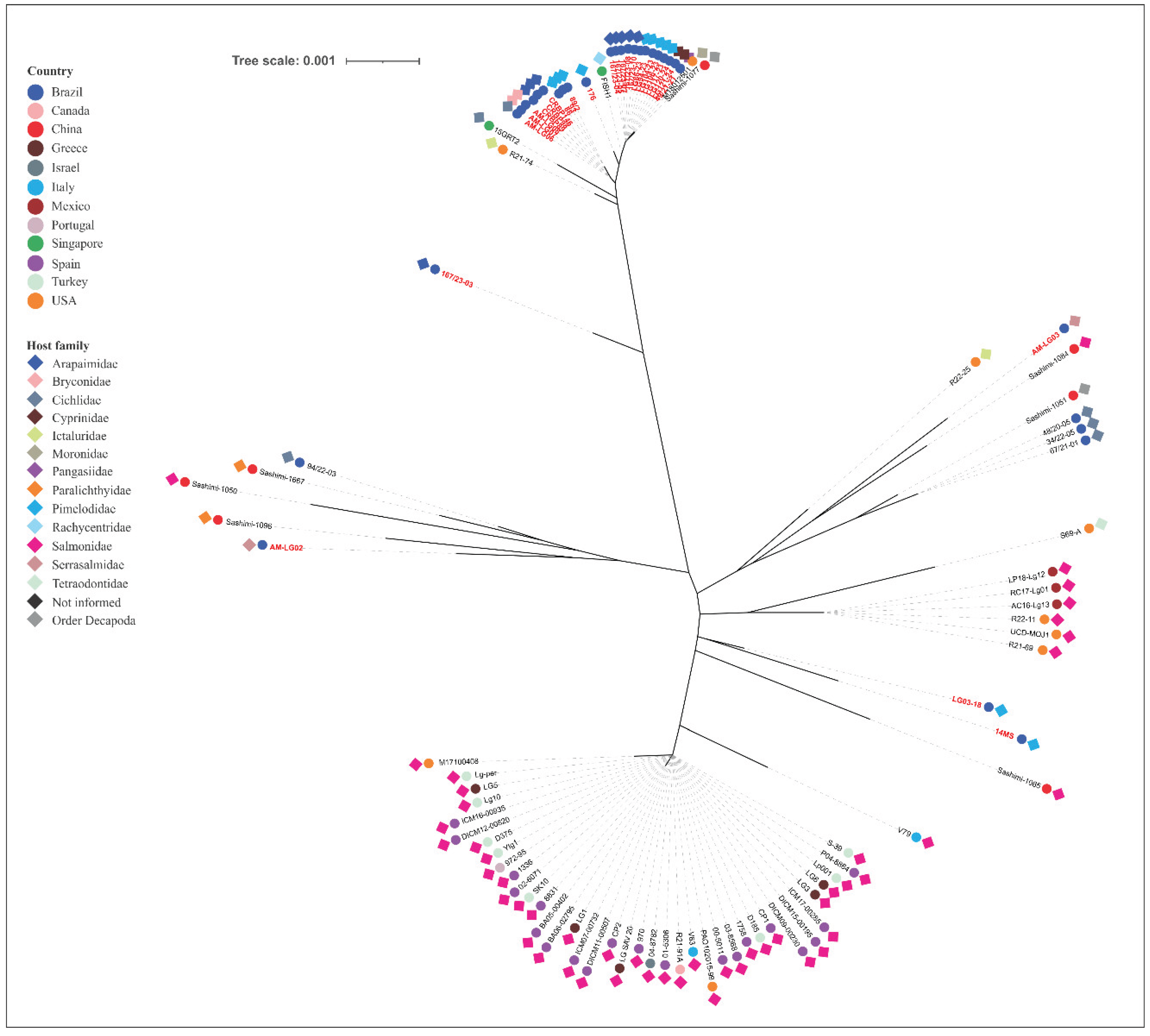
Lactococcosis has emerged as an economically and ecologically significant disease in aquatic animals worldwide. This study employed multilocus sequence typing (MLST) to investigate the genetic diversity of Lactococcus spp. strains from Brazilian fish species and evaluate their phylogenetic relationships with global isolates to elucidate potential epidemiological connections involving multiple host species and distinct geographic regions. A total of 55 isolates had their DNA extracted, followed by the amplification and sequencing of the internal fragments of seven housekeeping genes (als, atpA, tuf, gapC, gyrB, rpoC and galP). Sequence types (STs) and clonal complexes (CCs) were defined. An unrooted neighbor-joining phylogenetic tree was generated using allele profiles from this study and those previously reported from other aquatic animal species. The isolates comprised 29 STs (11 previously reported, 18 novel ones), which were grouped into species-specific CCs: CC5 (L. formosensis); CC4, CC17, CC62 (L. garvieae); CC24, CC29, CC97 (L. petauri). Considerable genetic divergence was observed, with L. formosensis and L. garvieae forming heterogeneous populations, while L. petauri was more homogeneous. Phylogenetics confirmed groupings within the CCs and revealed significant genetic arrangements. In conclusion, the Brazilian Lactococcus spp. strains analyzed in this study constitute a genetically diverse population based on their STs. MLST and phylogenetic analysis demonstrated genetic relatedness between the L. garvieae and L. formosensis isolates from this study and those from other aquatic animal species. In contrast, all the STs identified for L. petauri in this study were unrelated to the MLST lineages responsible for outbreaks in Brazilian Nile tilapia (Oreochromis niloticus) and North American rainbow trout (Oncorhynchus mykiss). This suggests that piscine L. petauri populations in the Americas evolved from distinct ancestral origins.
Keywords:
Introduction
Methods
Bacterial Strains and Identification
DNA Extraction
Multilocus Sequence Typing
Data Analysis
Results
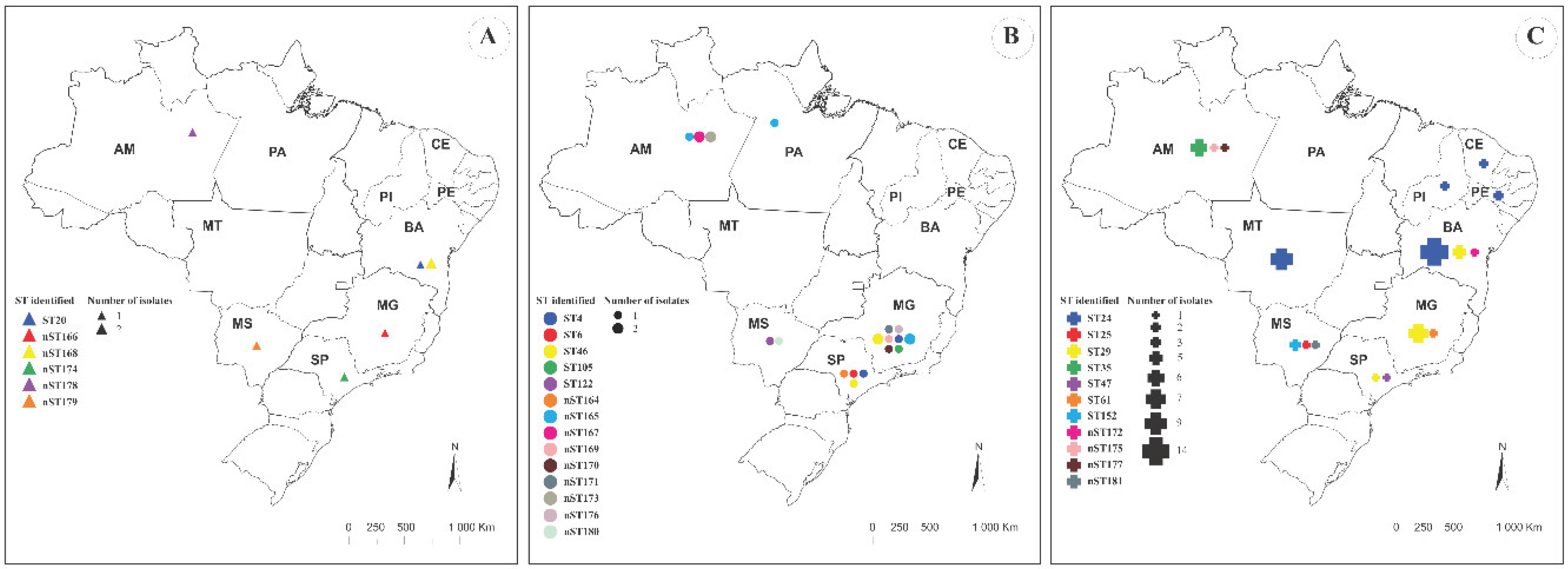
MLST Analysis
Discussion
Supplementary Materials
Author Contributions
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Shahin, K; Abdel-Glil, M; Saticıoğlu, IB; Duman, M; Altun, S; Colussi, S; et al. Diving into the depths: unveiling the main etiologies of piscine lactococcosis with a novel multiplex qPCR assay. J Fish Dis. 2025, e14147. [Google Scholar] [CrossRef]
- Rao, S; Pham, TH; Poudyal, S; Cheng, L-W; Nazareth, SC; Wang, P-C; et al. First report on genetic characterization, cell-surface properties and pathogenicity of Lactococcus garvieae, emerging pathogen isolated from cage-cultured cobia (Rachycentron canadum). Transbound Emerg Dis. 2022, 69, 1197–211. [Google Scholar] [CrossRef]
- Egger, RC; Rosa, JCC; Resende, LFL; de Pádua, SB; de Oliveira Barbosa, F; Zerbini, MT; et al. Emerging fish pathogens Lactococcus petauri and L. garvieae in Nile tilapia (Oreochromis niloticus) farmed in Brazil. Aquaculture 2023, 565, 739093. [Google Scholar] [CrossRef]
- Abraham, T; Yazdi, Z; Littman, E; Shahin, K; Heckman, TI; Quijano Cardé, EM; et al. Detection and virulence of Lactococcus garvieae and L. petauri from four lakes in southern California. J Aquat Anim Health 2023, 35, 187–98. [Google Scholar] [CrossRef] [PubMed]
- Salogni, C; Bertasio, C; Accini, A; Gibelli, LR; Pigoli, C; Susini, F; et al. The Characterisation of Lactococcus garvieae isolated in an outbreak of septicaemic disease in farmed sea bass (Dicentrarchus labrax, Linnaues 1758) in Italy. Pathogens 2024, 13. [Google Scholar] [CrossRef]
- Salif, M; Ogawa, R; Mikami, A; Daibata, M; Imajoh, M. Complete genome sequence of Lactococcus garvieae isolated from a greater amberjack (Seriola dumerili) farmed in Japan in 2022. Microbiol Resour Announc. 2024, 13, e00436-24. [Google Scholar] [CrossRef]
- Balan, R; Pandey, S; Wang, P-C; Byadgi, OV; Chen, S-C. Insights on the virulence and genomic features of Lactococcus garvieae isolated from giant freshwater prawn Macrobrachium rosenbergii (de Man 1879). J Fish Dis. 2024, 47, e14011. [Google Scholar] [CrossRef]
- Wongkaew, J; Chatchaiphan, S; Taengphu, S; Dong, HT; Senapin, S; Piyapattanakorn, S. Identification and pathogenicity of Lactococcus species in Nile tilapia (Oreochromis niloticus) and Asian sea bass (Lates calcarifer). J Fish Dis. 2025, 48, e14113. [Google Scholar] [CrossRef] [PubMed]
- Shahin, K; Veek, T; Heckman, TI; Littman, E; Mukkatira, K; Adkison, M; et al. Isolation and characterization of Lactococcus garvieae from rainbow trout, Onchorhyncus mykiss, from California, USA. Transbound Emerg Dis. 2022, 69, 2326–43. [Google Scholar] [CrossRef] [PubMed]
- Chang, PH; Lin, CW; Lee, YC. Lactococcus garvieae infection of cultured rainbow trout, Oncorhynchus mykiss, in Taiwan and associated biophysical characteristics and histopathology. Bull Eur Assoc Fish Pathol. 2002, 22, 319–27. [Google Scholar]
- Kang, SH; Shin, GW; Shin, YS; Palaksha, K. J; Rim, Kim Y; Hee, Yang H; et al. Experimental evaluation of pathogenicity of Lactococcus garvieae in black rockfish (Sebastes schlegeli). J Vet Sci. 2004, 5, 387–90. [Google Scholar] [CrossRef]
- Fukuda, Y; Tue, Y; Oinaka, D; Wada, Y; Yamashita, A; Urasaki, S; et al. Pathogenicity and immunogenicity of non-agglutinating Lactococcus garvieae with anti-KG- phenotype rabbit serum in Seriola spp. Fish Pathol 2015, 50, 200–6. [Google Scholar] [CrossRef]
- Meyburgh, CM; Bragg, RR; Boucher, CE. Lactococcus garvieae: an emerging bacterial pathogen of fish. Dis Aquat Organ. 2017, 123, 67–79. [Google Scholar] [CrossRef]
- Fukushima, HCS; Leal, CAG; Cavalcante, RB; Figueiredo, HCP; Arijo, S; Moriñigo, MA; et al. Lactococcus garvieae outbreaks in Brazilian farms: lactococcosis in Pseudoplatystoma sp. – development of an autogenous vaccine as a control strategy. J Fish Dis. 2017, 40, 263–72. [Google Scholar] [CrossRef]
- Pastorino, P; Vela Alonso, AI; Colussi, S; Cavazza, G; Menconi, V; Mugetti, D; et al. A summer mortality outbreak of lactococcosis by Lactococcus garvieae in a raceway system affecting farmed rainbow trout (Oncorhynchus mykiss) and brook trout (Salvelinus fontinalis). Animals 2019, 9. [Google Scholar] [CrossRef] [PubMed]
- Choi, HJ; Hur, JW; Cho, JB; Park, KH; Jung, HJ; Kang, YJ. Introduction of bacterial and viral pathogens from imported ornamental finfish in South Korea. Fish Aquat Sci. 2019, 22, 5. [Google Scholar] [CrossRef]
- Cardoso, PHM; Moreno, LZ; de Oliveira, CH; Gomes, VTM; Silva, APS; Barbosa, MRF; et al. Main bacterial species causing clinical disease in ornamental freshwater fish in Brazil. Folia Microbiol (Praha) 2021, 66, 231–9. [Google Scholar] [CrossRef] [PubMed]
- Neupane, S; Rao, S; Yan, W-X; Wang, P-C; Chen, S-C. First identification, molecular characterization, and pathogenicity assessment of Lactococcus garvieae isolated from cultured pompano in Taiwan. J Fish Dis. 2023, 46, 1295–309. [Google Scholar] [CrossRef] [PubMed]
- Barbanti, ACC; do Rosário, AEC; da Silva Maia, CRM; Rocha, VP; Costa, HL; Trindade, JM; et al. Genetic characterization of lactococcosis-causing bacteria isolated from Brazilian native fish species. Aquaculture 2024, 593, 741305. [Google Scholar] [CrossRef]
- Bondavalli, F; Colussi, S; Pastorino, P; Zanoli, A; Bezzo Llufrio, T; Fernández-Garayzábal, JF; et al. First report of Lactococcus petauri in the pumpkinseed (Lepomis gibbosus) from Candia lake (Northwestern Italy). Fishes 2024, 9. [Google Scholar] [CrossRef]
- Esposito, G; Bignami, G; Colussi, S; Pastorino, P; Bondavalli, F; Fioravanti, M; et al. Expanding horizons: the first reported outbreak of piscine lactococcosis in farmed gilthead seabream Sparus aurata in the Northern Tyrrhenian sea. J Fish Dis. 2025, 48, e14121. [Google Scholar] [CrossRef] [PubMed]
- Ferrario, C; Ricci, G; Milani, C; Lugli, GA; Ventura, M; Eraclio, G; et al. Lactococcus garvieae: where is it from? A first approach to explore the evolutionary history of this emerging pathogen. PLoS One 2013, 8, e84796. [Google Scholar] [CrossRef]
- Reguera-Brito, M; Galán-Sánchez, F; Blanco, MM; Rodríguez-Iglesias, M; Domínguez, L; Fernández-Garayzábal, JF; et al. Genetic analysis of human clinical isolates of Lactococcus garvieae: relatedness with isolates from foods. Infect Genet Evol. 2016, 37, 185–91. [Google Scholar] [CrossRef]
- Thiry, D; Billen, F; Boyen, F; Duprez, J-N; Quenault, H; Touzain, F; et al. Genomic relatedness of a canine Lactococcus garvieae to human, animal and environmental isolates. Res Vet Sci. 2021, 137, 170–3. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y; Han, J; Barkema, HW; Wang, Y; Gao, J; Kastelic, JP; et al. Comparative genomic analyses of Lactococcus garvieae isolated from bovine mastitis in China. Microbiol Spectr. 2023, 11, e02995-22. [Google Scholar] [CrossRef]
- Chan, Y-X; Cao, H; Jiang, S; Li, X; Fung, K-K; Lee, C-H; et al. Genomic investigation of Lactococcus formosensis, Lactococcus garvieae, and Lactococcus petauri reveals differences in species distribution by human and animal sources. Microbiol Spectr. 2024, 12, e00541-24. [Google Scholar] [CrossRef] [PubMed]
- Eldar, A; Goria, M; Ghittino, C; Zlotkin, A; Bercovier, Herve. Biodiversity of Lactococcus garvieae strains isolated from fish in Europe, Asia, and Australia. Appl Environ Microbiol. 1999, 65, 1005–8. [Google Scholar] [CrossRef]
- Ferrario, C; Ricci, G; Borgo, F; Rollando, A; Fortina, MG. Genetic investigation within Lactococcus garvieae revealed two genomic lineages. FEMS Microbiol Lett. 2012, 332, 153–61. [Google Scholar] [CrossRef]
- Rao, S; Chen, M-Y; Sudpraseart, C; Lin, P; Yoshida, T; Wang, P-C; et al. Genotyping and phenotyping of Lactococcus garvieae isolates from fish by pulse-field gel electrophoresis (PFGE) and electron microscopy indicate geographical and capsular variations. J Fish Dis. 2022, 45, 771–81. [Google Scholar] [CrossRef]
- Evans, JJ; Bohnsack, JF; Klesius, PH; Whiting, A a; Garcia, JC; Shoemaker, C a; et al. Phylogenetic relationships among Streptococcus agalactiae isolated from piscine, dolphin, bovine and human sources: a dolphin and piscine lineage associated with a fish epidemic in Kuwait is also associated with human neonatal infections in Japan. J Med Microbiol. 2008, 57 Pt 11, 1369–76. [Google Scholar] [CrossRef]
- Barony, GM; Tavares, GC; Pereira, FL; Carvalho, AF; Dorella, FA; Leal, CAG; et al. Large-scale genomic analyses reveal the population structure and evolutionary trends of Streptococcus agalactiae strains in Brazilian fish farms. Sci Rep. 2017, 7, 13538. [Google Scholar] [CrossRef]
- Maiden, MCJ; van Rensburg, MJJ; Bray, JE; Earle, SG; Ford, SA; Jolley, KA; et al. MLST revisited: the gene-by-gene approach to bacterial genomics. Nat Rev Microbiol. 2013, 11, 728–36. [Google Scholar] [CrossRef] [PubMed]
- Jolley, KA; Bray, JE; Maiden, MCJ. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 2018, 3, 1–20. [Google Scholar] [CrossRef]
- Kozakai, M; Matsumoto, C; Matsumoto, M; Takakura, A; Matsubayashi, K; Satake, M. Different growth kinetics in blood components and genetic analysis of Lactococcus garvieae isolated from platelet concentrates. Transfusion 2020, 60, 1492–9. [Google Scholar] [CrossRef] [PubMed]
- Lin, YS; Kweh, KH; Koh, TH; Lau, QC; Abdul Rahman, NB. Genomic analysis of Lactococcus garvieae isolates. Pathology 2020, 52, 700–7. [Google Scholar] [CrossRef] [PubMed]
- Mahmoud, MM; Abdelsalam, M; Kawato, S; Harakawa, S; Kawakami, H; Hirono, I; et al. Comparative genome analyses of three serotypes of Lactococcus bacteria isolated from diseased cultured striped jack (Pseudocaranx dentex). J Fish Dis. 2023, 46, 829–39. [Google Scholar] [CrossRef]
- Heckman, TI; Yazdi, Z; Older, CE; Griffin, MJ; Waldbieser, GC; Chow, AM; et al. Redefining piscine lactococcosis. Appl Environ Microbiol 2024, 0, e02349-23. [Google Scholar] [CrossRef]
- Feil, EJ; Li, BC; Aanensen, DM; Hanage, WP; Spratt, Brian G. eBURST: Inferring patterns of evolutionary descent among clusters of related bacterial genotypes from multilocus sequence typing data. J Bacteriol. 2004, 186, 1518–30. [Google Scholar] [CrossRef]
- Francisco, AP; Bugalho, M; Ramirez, M; Carriço, JA. Global optimal eBURST analysis of multilocus typing data using a graphic matroid approach. BMC Bioinformatics 2009, 10, 152. [Google Scholar] [CrossRef]
- Nascimento, M; Sousa, A; Ramirez, M; Francisco, AP; Carriço, JA; Vaz, C. PHYLOViZ 2.0: providing scalable data integration and visualization for multiple phylogenetic inference methods. Bioinformatics 2017, 33, 128–9. [Google Scholar] [CrossRef]
- Hunter, PR; Gaston, MA. Numerical index of the discriminatory ability of typing systems: an application of Simpson’s index of diversity. J Clin Microbiol. 1988, 26, 2465–6. [Google Scholar] [CrossRef]
- Pinto, FR; Melo-Cristino, J; Ramirez, M. A confidence interval for the Wallace coefficient of concordance and its application to microbial typing methods. PLoS One 2008, 3, e3696. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S; Stecher, G; Suleski, M; Sanderford, M; Sharma, S; Tamura, K. MEGA12: Molecular evolutionary genetic analysis version 12 for adaptive and green computing. Mol Biol Evol. 2024, 41, msae263. [Google Scholar] [CrossRef]
- Tamura, K; Stecher, G; Kumar, S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol Biol Evol. 2021, 38, 3022–7. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I; Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 2024, 52, W78–82. [Google Scholar] [CrossRef]
- Goodman, LB; Lawton, MR; Franklin-Guild, RJ; Anderson, RR; Schaan, L; Thachil, AJ; et al. Lactococcus petauri sp. nov., isolated from an abscess of a sugar glider. Int J Syst Evol Microbiol. 2017, 67, 4397–404. [Google Scholar] [CrossRef] [PubMed]
- Vela, AI; del Mar Blanco, M; Colussi, S; Kotzamanidis, C; Prearo, M; Altinok, I; et al. The association of Lactococcus petauri with lactococcosis is older than expected. Aquaculture 2024, 578, 740057. [Google Scholar] [CrossRef]
- Rodrigues, RA; do Nascimento Silva, AL; Siqueira, MS; Pilarski, F; Leal, CRB; Kuibida, KV; et al. Hematological, biochemical, and histopathological responses in sorubim Pseudoplatystoma spp. experimentally infected with Lactococcus garvieae. Aquac Int. 2020, 28, 1907–23. [Google Scholar] [CrossRef]
- Ortega, C; Irgang, R; Valladares-Carranza, B; Collarte, C; Avendaño-Herrera, R. First identification and characterization of Lactococcus garvieae isolated from rainbow trout (Oncorhynchus mykiss) cultured in Mexico. Animals 2020, 10. [Google Scholar] [CrossRef]
- Rosário, AEC; Reis, FYT; Barbanti, ACC; da Costa, ÉJC; da Silva Maia, CRM; Kotzent, S; et al. Susceptibility and clinicopathological findings of three Amazonian fishes experimentally infected with. Lactococcus spp. Preprints 2025. [Google Scholar] [CrossRef]

| Isolate | Species | Origin | Year | Host | MLST | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Allele | ST | CC | |||||||||||
| als | atpA | tuf | gapC | gyrB | rpoC | galP | |||||||
| 167/23-02 | L. formosensis | BA | 2023 | Arapaima gigas | 22 | 62 | 18 | 3 | 20 | 4 | 78 | n168 | Singleton |
| 167/23-06 | L. formosensis | BA | 2023 | Arapaima gigas | 15 | 10 | 14 | 9 | 13 | 15 | 15 | 20 | Singleton |
| 167/23-09 | L. formosensis | BA | 2023 | Arapaima gigas | 22 | 62 | 18 | 3 | 20 | 4 | 78 | n168 | Singleton |
| 49/21-29 | L. formosensis | SP | 2021 | Pangasianodon hypophthalmus | 100 | 4 | 18 | 3 | 20 | 4 | 79 | n174 | Singleton |
| 52MS | L. formosensis | MS | 2012 | Pseudoplatystoma fasciatum | 91 | 60 | 14 | 9 | 20 | 33 | 81 | n179 | Singleton |
| AM-LG05 | L. formosensis | AM | 2022 | Colossoma macropomum | 90 | 35 | 14 | 9 | 20 | 38 | 81 | n178 | Singleton |
| LG91-23 | L. formosensis | MG | 2023 | Pseudoplatystoma sp. | 92 | 4 | 50 | 3 | 20 | 4 | 73 | n166 | Singleton |
| 177 | L. garvieae | MS | 2012 | Pseudoplatystoma fasciatum | 3 | 3 | 4 | 2 | 59 | 3 | 3 | 122 | CC4 |
| 31MS | L. garvieae | MS | 2012 | Pseudoplatystoma fasciatum | 12 | 8 | 54 | 7 | 27 | 13 | 12 | n180 | CC17 |
| 49/21-11 | L. garvieae | SP | 2021 | Pangasianodon hypophthalmus | 5 | 5 | 6 | 2 | 5 | 5 | 5 | 6 | Singleton |
| CRBP53 | L. garvieae | AM | 2023 | Arapaima gigas | 93 | 61 | 51 | 15 | 72 | 61 | 74 | n167 | Singleton |
| CRBP54 | L. garvieae | AM | 2023 | Arapaima gigas | 93 | 61 | 51 | 15 | 72 | 61 | 74 | n167 | Singleton |
| CRBP138 | L. garvieae | AM | 2023 | Arapaima gigas | 34 | 59 | 27 | 15 | 28 | 19 | 29 | n173 | - |
| CRBP144 | L. garvieae | AM | 2023 | Arapaima gigas | 34 | 59 | 27 | 15 | 28 | 19 | 29 | n173 | - |
| LG09-14 | L. garvieae | SP | 2014 | Pseudoplatystoma corruscans | 3 | 3 | 4 | 2 | 3 | 3 | 3 | 4 | CC4 |
| LG10-14 | L. garvieae | MG | 2014 | Lophiosilurus alexandri | 60 | 8 | 6 | 7 | 10 | 45 | 48 | 105 | Singleton |
| LG23-16 | L. garvieae | SP | 2016 | Pseudoplatystoma corruscans | 88 | 22 | 46 | 25 | 71 | 20 | 71 | n164 | Singleton |
| LG63-21 | L. garvieae | MG | 2021 | Hoplias macrophtalmus | 3 | 3 | 4 | 2 | 3 | 3 | 3 | 4 | CC4 |
| LG64-21 | L. garvieae | MG | 2021 | Xiphophorus maculatus | 87 | 24 | 27 | 15 | 28 | 27 | 29 | n176 | nCC62 |
| LG66-22 | L. garvieae | MG | 2022 | Phractocephalus hemioliopterus | 12 | 8 | 6 | 7 | 27 | 13 | 12 | 46 | CC17 |
| LG88-23 | L. garvieae | MG | 2023 | Brycon amazonicus | 34 | 59 | 27 | 15 | 28 | 19 | 72 | n165 | - |
| LG89-23 | L. garvieae | MG | 2023 | Brycon amazonicus | 34 | 59 | 27 | 15 | 28 | 19 | 72 | n165 | - |
| LG114-23 | L. garvieae | AM | 2023 | Hoplias malabaricus | 34 | 59 | 27 | 15 | 28 | 19 | 72 | n165 | - |
| LG115-23 | L. garvieae | MG | 2023 | Trichogaster lalius | 21 | 13 | 1 | 2 | 73 | 16 | 75 | n169 | Singleton |
| LG116-23 | L. garvieae | MG | 2023 | Cichla sp. | 94 | 3 | 52 | 2 | 73 | 62 | 76 | n170 | Singleton |
| LG119-24 | L. garvieae | MG | 2024 | Pseudoplatystoma sp. | 95 | 69 | 6 | 2 | 5 | 63 | 77 | n171 | Singleton |
| PA-LG01 | L. garvieae | PA | 2018 | Arapaima gigas | 34 | 59 | 27 | 15 | 28 | 19 | 72 | n165 | - |
| 86 | L. petauri | MS | 2012 | Pseudoplatystoma sp. | 9 | 7 | 3 | 2 | 37 | 9 | 9 | 152 | nCC29 |
| 93 | L. petauri | MS | 2012 | Pseudoplatystoma sp. | 9 | 7 | 3 | 2 | 37 | 9 | 9 | 152 | nCC29 |
| 176 | L. petauri | MS | 2012 | Pseudoplatystoma fasciatum | 9 | 7 | 3 | 4 | 16 | 9 | 17 | 25 | Singleton |
| 14MS | L. petauri | MS | 2012 | Pseudoplatystoma fasciatum | 32 | 21 | 7 | 2 | 7 | 11 | 6 | n181 | - |
| 167/23-03 | L. petauri | BA | 2023 | Arapaima gigas | 94 | 6 | 7 | 2 | 7 | 11 | 82 | n172 | Singleton |
| 167/23-04 | L. petauri | BA | 2023 | Arapaima gigas | 9 | 7 | 3 | 4 | 18 | 9 | 9 | 29 | nCC29 |
| 167/23-05 | L. petauri | BA | 2023 | Arapaima gigas | 9 | 7 | 3 | 4 | 18 | 9 | 9 | 29 | nCC29 |
| 167/23-07 | L. petauri | BA | 2023 | Arapaima gigas | 9 | 7 | 3 | 4 | 18 | 9 | 9 | 29 | nCC29 |
| 167/23-08 | L. petauri | BA | 2023 | Arapaima gigas | 9 | 7 | 3 | 4 | 18 | 9 | 9 | 29 | nCC29 |
| 167/23-10 | L. petauri | BA | 2023 | Arapaima gigas | 9 | 7 | 3 | 4 | 18 | 9 | 9 | 29 | nCC29 |
| 49/21-21 | L. petauri | SP | 2021 | Pangasianodon hypophthalmus | 9 | 7 | 3 | 4 | 18 | 9 | 9 | 29 | nCC29 |
| 89/2 | L. petauri | MS | 2012 | Pseudoplatystoma sp. | 9 | 7 | 3 | 2 | 37 | 9 | 9 | 152 | nCC29 |
| AM-LG02 | L. petauri | AM | 2020 | Colossoma macropomum | 61 | 6 | 7 | 35 | 7 | 11 | 8 | n175 | Singleton |
| AM-LG03 | L. petauri | AM | 2022 | Colossoma macropomum | 89 | 20 | 26 | 2 | 24 | 25 | 6 | n177 | - |
| AM-LG06 | L. petauri | AM | 2022 | Pterophyllum scalare | 9 | 7 | 3 | 2 | 7 | 9 | 9 | 35 | nCC29 |
| AM-LG07 | L. petauri | AM | 2022 | Brycon amazonicus | 9 | 7 | 3 | 2 | 7 | 9 | 9 | 35 | nCC29 |
| AM-LG08 | L. petauri | AM | 2022 | Brycon amazonicus | 9 | 7 | 3 | 2 | 7 | 9 | 9 | 35 | nCC29 |
| CRBP89 | L. petauri | AM | 2023 | Arapaima gigas | 9 | 7 | 3 | 2 | 7 | 9 | 9 | 35 | nCC29 |
| CRBP98 | L. petauri | AM | 2023 | Arapaima gigas | 9 | 7 | 3 | 2 | 7 | 9 | 9 | 35 | nCC29 |
| CRBP146 | L. petauri | AM | 2023 | Arapaima gigas | 9 | 7 | 3 | 2 | 7 | 9 | 9 | 35 | nCC29 |
| LG03-18 | L. petauri | MG | 2018 | Pseudoplatystoma corruscans | 33 | 6 | 10 | 2 | 7 | 11 | 8 | 61 | - |
| LG86-23 | L. petauri | MG | 2023 | Pseudoplatystoma sp. | 9 | 7 | 3 | 4 | 18 | 9 | 9 | 29 | nCC29 |
| LG94-23 | L. petauri | MG | 2023 | Pseudoplatystoma sp. | 9 | 7 | 3 | 4 | 18 | 9 | 9 | 29 | nCC29 |
| LG104-23 | L. petauri | MG | 2023 | Pseudoplatystoma sp. | 9 | 7 | 3 | 4 | 18 | 9 | 9 | 29 | nCC29 |
| LG106-23 | L. petauri | MG | 2023 | Pseudoplatystoma sp. | 9 | 7 | 3 | 4 | 18 | 9 | 9 | 29 | nCC29 |
| LG117-23 | L. petauri | MG | 2023 | Pseudoplatystoma sp. | 9 | 7 | 3 | 4 | 18 | 9 | 9 | 29 | nCC29 |
| LG120-24 | L. petauri | MG | 2024 | Carassius auratus | 9 | 7 | 3 | 4 | 18 | 9 | 9 | 29 | nCC29 |
| LG121-24 | L. petauri | MG | 2024 | Carassius auratus | 9 | 7 | 3 | 4 | 18 | 9 | 9 | 29 | nCC29 |
| Gene | Primer pairs (5’-3’) | Annealing temperature (ºC) | Size (bp) | Nº of alleles | Nº of polymorphic sites |
|---|---|---|---|---|---|
| als | F: ATTCGGCTCAGACTTAGTTG R: TTCAGCTGCTTCAACATCAA |
58 | 811 | 22 | 85 |
| atpA | F: TAYRTYGGKGAYGGDATYGC R: CCRCGRTTHARYTTHGCYTG |
56 | 803 | 18 | 51 |
| tuf | F: ATATGCGGCCGCCATYGGHCACGTBGACCA R: AAAATATGCGGCCGCTCNCCNGGCATNACCAT |
56 | 809 | 15 | 30 |
| gapC | F: AAGTTGGTATTAACGGTTTCG R: AAGTGTACGAACGAGGTTAG |
56 | 821 | 8 | 9 |
| gyrB | F: CATGCTGGTGGTAAATTTGG R: GTCATCCATTTCTCCTAAACC |
58 | 827 | 16 | 111 |
| rpoC | F: TTGGTCCACAAAAGGACTGG R: TCACGTCCTTTTGCTTCCAT |
58 | 830 | 18 | 58 |
| galP | F: TGGGGAAAATTTAAACCTTGG R: ATCATCAGAACGGCTGGAAG |
58 | 812 | 21 | 107 |
| Bacterial species | ST in aquatic animals/ST totala | STs identified in this study | STs identified in other studies |
|---|---|---|---|
| L. formosensis | 18/39 | ST20, nST166, nST168, nST174. nST178, nST179 | ST5, ST41, ST43, ST56, ST59, ST113, ST114, ST115, nST140, nST141, nST150, nST151 |
| L. garvieae | 33/55 | ST4, ST6, ST46, ST105, ST122, nST164, nST165, nST167, nST169, nST170, nST171, nST173, nST176, nST180 | ST1, ST13, ST16, ST17, ST39, ST62, ST63, ST95, ST109, ST119, ST120, ST121, ST123, ST124, ST139, nST144, nST147, nST157, nST158 |
| L. petauri | 29/85 | ST25, ST29, ST35, ST61, ST152, nST172, nST175, nST177, nST181 | ST10, ST14, ST15, ST24, ST34, ST47, ST57, ST98, ST128, ST132, ST133, ST134, ST135, ST136, ST137, ST138, nST142, nST145, nST146, nST149 |
| Lactococcus ssp.b | 0/2 | - | - |
| Total | 80/181 | 29/181 | 51/181 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




