Submitted:
09 February 2026
Posted:
11 February 2026
You are already at the latest version
Abstract
Keywords:
Introduction
Methods
Study Population
Patient Cohorts
Genotyping and Quality Control
- Call rate filter (> 0.95): Removed 31,671,940 variants
- Minor allele frequency filter (> 0.05): Removed 76,514,080 variants
- Hardy-Weinberg equilibrium filter (p > 1×10⁻⁶): Removed 485,562 variants
- Monomorphic variant check: Removed 0 variants
Statistical Analysis
- Statistical significance was defined as p < 5×10⁻⁸ for genome-wide significance and p < 1×10⁻⁵ for suggestive associations.
- Genomic inflation factor (λ) was calculated to assess potential population stratification or systematic biases.
- Results were visualized using Manhattan plots and quantile-quantile (QQ) plots.
Bioinformatic Analysis
Variant Annotation and Functional Prediction
Expression and Regulatory Analysis
Statistical Refinement Analyses
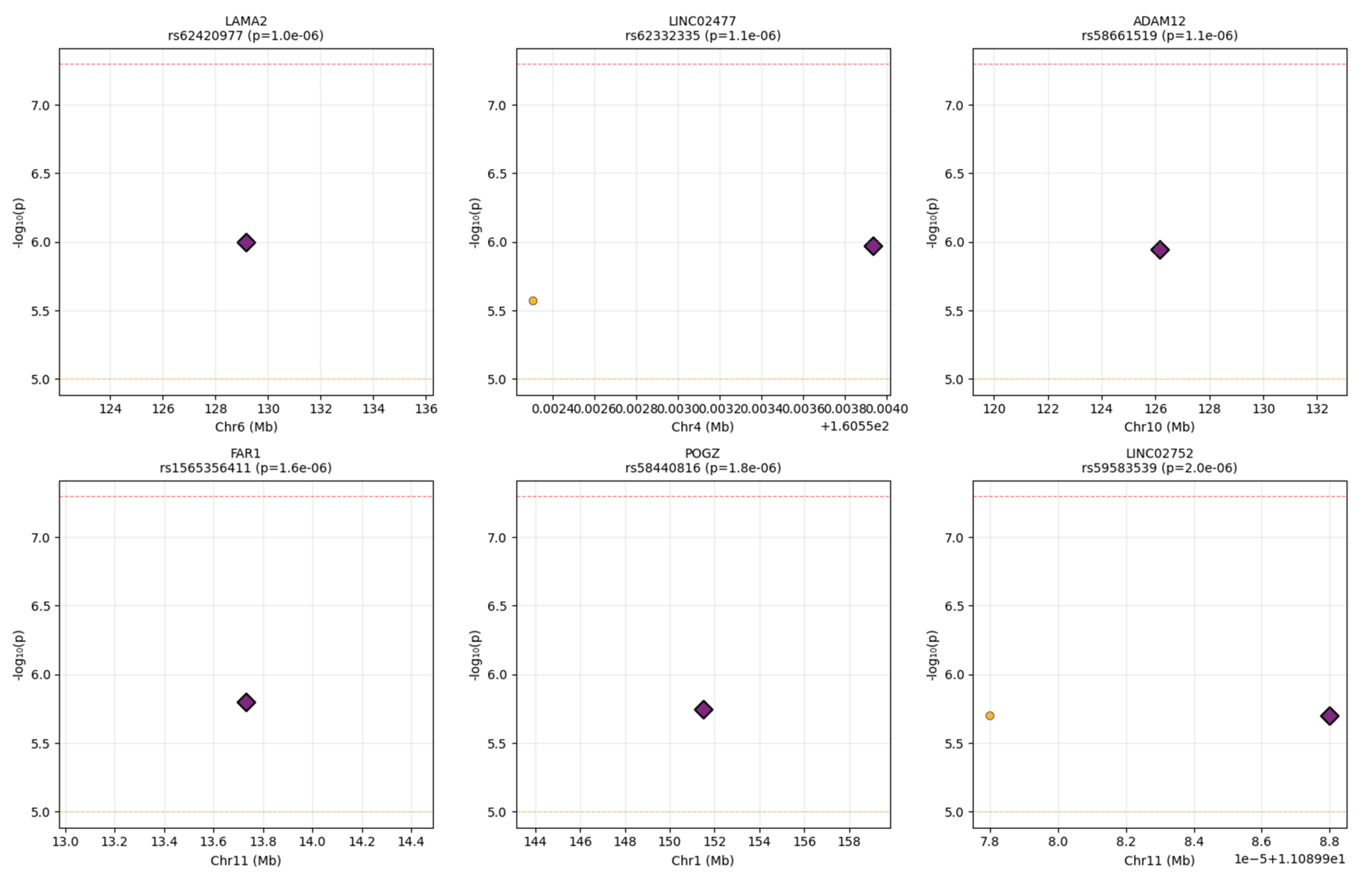
Regional Visualization
Results
ClinVar Analysis Results
ACAF Analysis Results
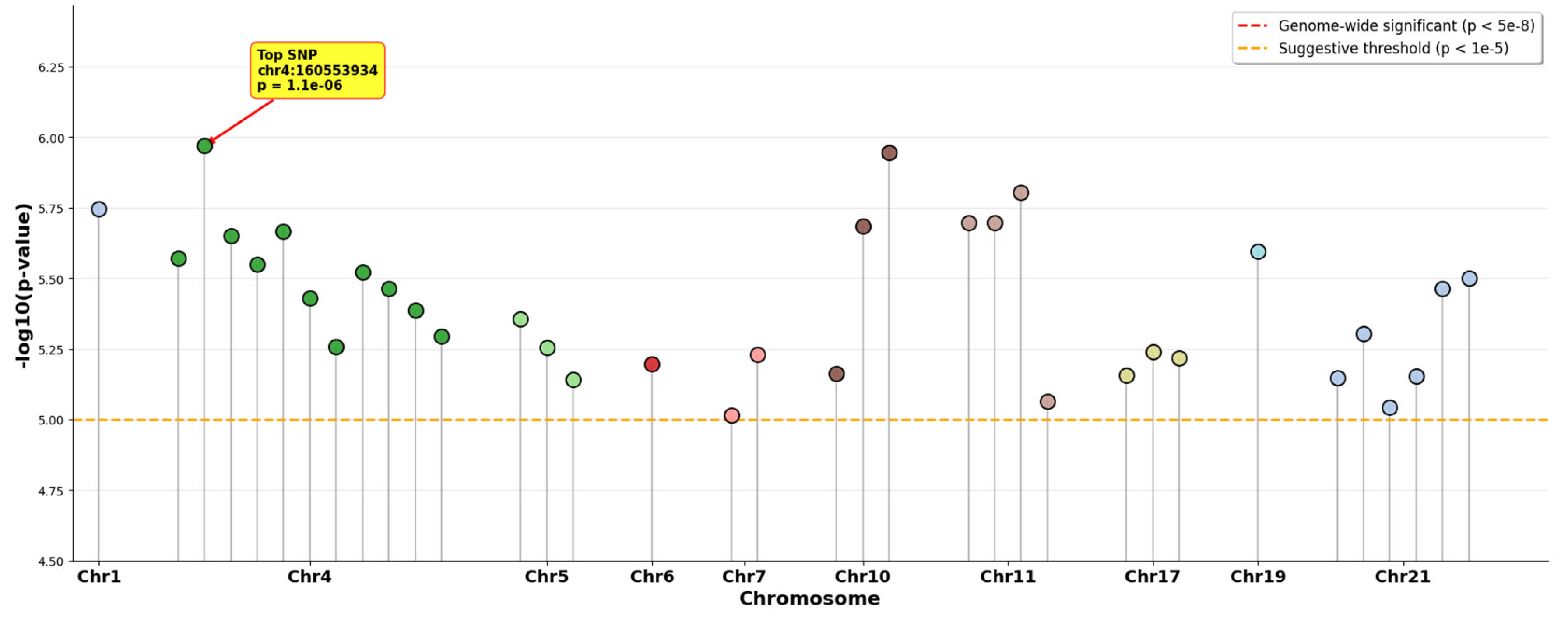
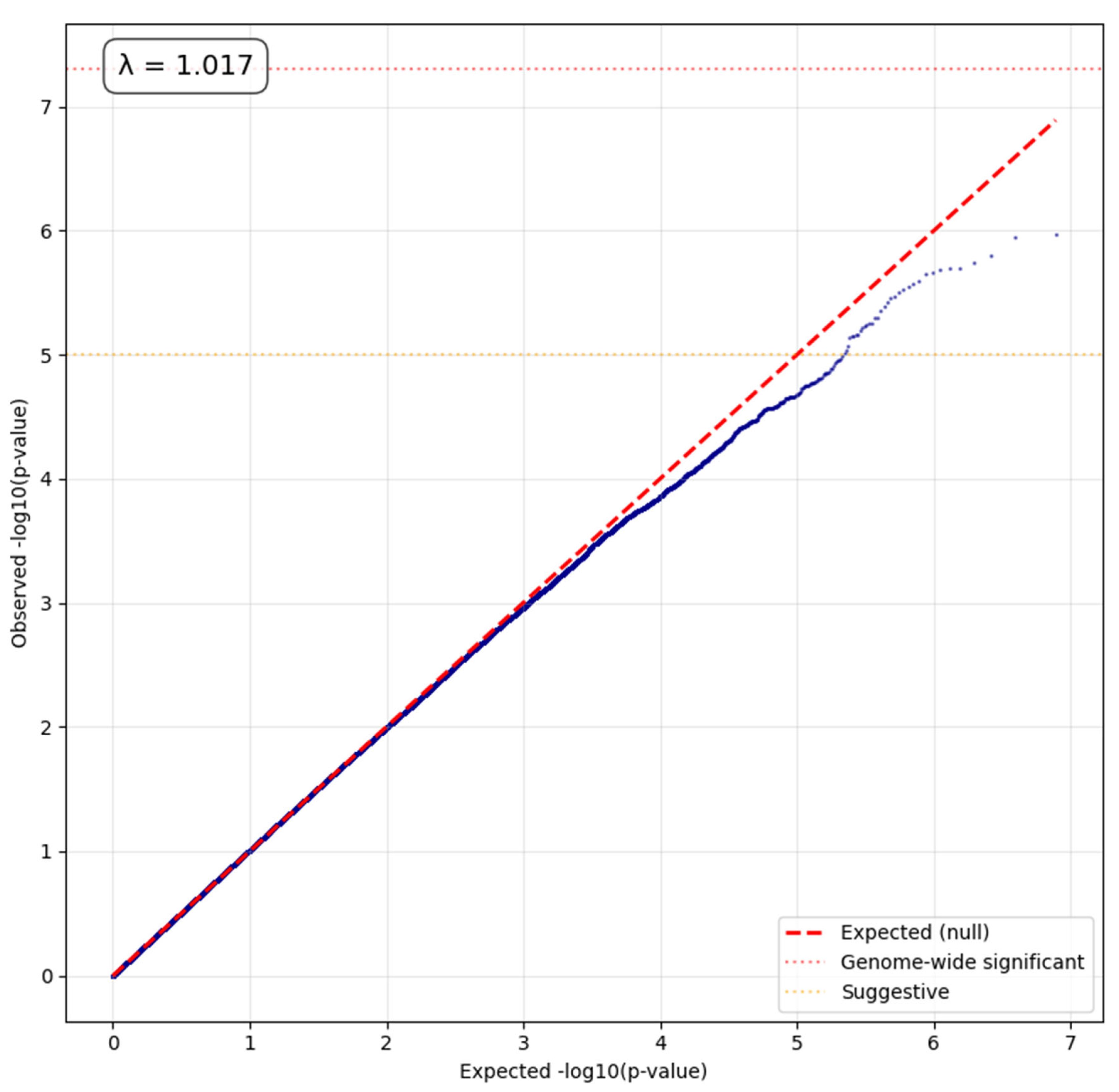
Functional and Regulatory Annotation of Suggestive Variants
Variant Characterization and Genomic Context
Population Genetics and Evolutionary Context
Pathogenicity Prediction
Splicing Analysis
Expression Quantitative Trait Loci Analysis
- rs6010165 (MLC1): Significant eQTL in whole blood (p = 2.9×10⁻¹³, NES = -0.18), affecting expression of MLC1, MOV10L1, and PANX2. Also showed splicing QTL activity.
- rs62420977 (LAMA2): Splicing QTL in thyroid tissue (p = 1.8×10⁻⁵, NES = -0.56).
- rs2460694 (B3GAT2): eQTL in tibial nerve (p = 3.5×10⁻⁴, NES = 0.13).
Regulatory Element Analysis
Chromatin Accessibility Analysis
Comparison of Results Across Analyses
Unified Genetic Architecture
Critical Impact of Study Design
Robustness to Demographic Confounding
Functional Annotation Integration
Regional Architecture Patterns
Discussion
Interpretation of Key Findings
Methodological Considerations
Biological and Clinical Implications
Comparison with Previous Literature
Strengths and Limitations
Future Directions
Conclusions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Conte, M. S.; Bradbury, A. W.; Kolh, P.; White, J. V.; Dick, F.; Fitridge, R.; Mills, J. L.; Ricco, J.; Suresh, K. R.; Murad, M. H.; Aboyans, V.; Aksoy, M.; Alexandrescu, V.; Armstrong, D.; Azuma, N.; Belch, J.; Bergoeing, M.; Bjorck, M.; Chakfé, N.; Wang, S. Global Vascular Guidelines on the Management of Chronic Limb-Threatening Ischemia. European Journal of Vascular and Endovascular Surgery 2019, 58(1), S1–S109.e33. [Google Scholar] [CrossRef]
- Firth, D. Bias reduction of maximum likelihood estimates. Biometrika 1993, 80(1), 27–38. [Google Scholar] [CrossRef]
- Fowkes, F. G. R.; Rudan, D.; Rudan, I.; Aboyans, V.; Denenberg, J. O.; McDermott, M. M.; Norman, P. E.; Sampson, U. K.; Williams, L. J.; Mensah, G. A.; Criqui, M. H. Comparison of global estimates of prevalence and risk factors for peripheral artery disease in 2000 and 2010: a systematic review and analysis. The Lancet 2013, 382(9901), 1329–1340. [Google Scholar] [CrossRef]
- Qvist, P.; Christensen, J. H.; Vardya, I.; Rajkumar, A. P.; Mørk, A.; Paternoster, V.; Füchtbauer, E.; Pallesen, J.; Fryland, T.; Dyrvig, M.; Hauberg, M. E.; Lundsberg, B.; Fejgin, K.; Nyegaard, M.; Jensen, K.; Nyengaard, J. R.; Mors, O.; Didriksen, M.; Børglum, A. D. The Schizophrenia-Associated BRD1 gene regulates behavior, neurotransmission, and expression of schizophrenia risk enriched gene sets in mice. Biological Psychiatry 2016, 82(1), 62–76. [Google Scholar] [CrossRef]
- Goodney, P. P.; Tarulli, M.; Faerber, A. E.; Schanzer, A.; Zwolak, R. M. Fifteen-Year trends in lower limb amputation, revascularization, and preventive measures among Medicare patients. JAMA Surgery 2015, 150(1), 84. [Google Scholar] [CrossRef]
- Klarin, D.; Lynch, J.; Aragam, K.; Chaffin, M.; Assimes, T. L.; Huang, J.; Lee, K. M.; Shao, Q.; Huffman, J. E.; Natarajan, P.; Arya, S.; Small, A.; Sun, Y. V.; Vujkovic, M.; Freiberg, M. S.; Wang, L.; Chen, J.; Saleheen, D.; Lee, J. S.; Damrauer, S. M. Genome-wide association study of peripheral artery disease in the Million Veteran Program. Nature Medicine 2019, 25(8), 1274–1279. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.; Blackwell, T.; Boehnke, M.; Scott, L. J. Recommended joint and Meta-Analysis strategies for Case-Control Association testing of Single Low-Count variants. Genetic Epidemiology 2013, 37(6), 539–550. [Google Scholar] [CrossRef]
- Mackness, M.; Mackness, B. Human paraoxonase-1 (PON1): Gene structure and expression, promiscuous activities and multiple physiological roles. Gene 2015, 567(1), 12–21. [Google Scholar] [CrossRef]
- Malik, R.; Chauhan, G.; Traylor, M.; Sargurupremraj, M.; Okada, Y.; Mishra, A.; Rutten-Jacobs, L.; Giese, A.; Van Der Laan, S. W.; Gretarsdottir, S.; Anderson, C. D.; Chong, M.; Adams, H. H. H.; Ago, T.; Almgren, P.; Amouyel, P.; Ay, H.; Bartz, T. M.; Benavente, O. R.; Dichgans, M. Multiancestry genome-wide association study of 520,000 subjects identifies 32 loci associated with stroke and stroke subtypes. Nature Genetics 2018, 50(4), 524–537. [Google Scholar] [CrossRef] [PubMed]
- Matsukura, M.; Ozaki, K.; Takahashi, A.; Onouchi, Y.; Morizono, T.; Komai, H.; Shigematsu, H.; Kudo, T.; Inoue, Y.; Kimura, H.; Hosaka, A.; Shigematsu, K.; Miyata, T.; Watanabe, T.; Tsunoda, T.; Kubo, M.; Tanaka, T. Genome-Wide Association study of peripheral arterial disease in a Japanese population. PLoS ONE 2015, 10(10), e0139262. [Google Scholar] [CrossRef] [PubMed]
- Mustapha, J. A.; Katzen, B. T.; Neville, R. F.; Lookstein, R. A.; Zeller, T.; Miller, L. E.; Jaff, M. R. Determinants of Long-Term Outcomes and Costs in the Management of Critical Limb Ischemia: A Population-Based Cohort Study. Journal of the American Heart Association 2018, 7(16). [Google Scholar] [CrossRef]
- A comprehensive 1000 Genomes–based genome-wide association meta-analysis of coronary artery disease. (2015). Nature Genetics 47(10), 1121–1130. [CrossRef]
- Norgren, L.; Hiatt, W.; Dormandy, J.; Nehler; Harris, K.; Fowkes, F. Inter-Society Consensus for the Management of Peripheral Arterial Disease (TASC II). Journal of Vascular Surgery 2007, 45(1), S5–S67. [Google Scholar] [CrossRef]
- Jones, S.; Swaminathan, A.; Patel, M.; Vemulapalli, S. Lower extremity amputation in peripheral artery disease: improving patient outcomes. Vascular Health and Risk Management 2014, 417. [Google Scholar] [CrossRef]
- Tam, V.; Patel, N.; Turcotte, M.; Bossé, Y.; Paré, G.; Meyre, D. Benefits and limitations of genome-wide association studies. Nature Reviews Genetics 2019, 20(8), 467–484. [Google Scholar] [CrossRef]
- Ma, Q.; Jacobi, P. M.; Emmer, B. T.; Kretz, C. A.; Ozel, A. B.; McGee, B.; Kimchi-Sarfaty, C.; Ginsburg, D.; Li, J. Z.; Desch, K. C. Genetic variants in ADAMTS13 as well as smoking are major determinants of plasma ADAMTS13 levels. Blood Advances 2017, 1(15), 1037–1046. [Google Scholar] [CrossRef] [PubMed]
- Wassel, C. L.; Lamina, C.; Nambi, V.; Coassin, S.; Mukamal, K. J.; Ganesh, S. K.; Jacobs, D. R.; Franceschini, N.; Papanicolaou, G. J.; Gibson, Q.; Yanek, L. R.; Van Der Harst, P.; Ferguson, J. F.; Crawford, D. C.; Waite, L. L.; Allison, M. A.; Criqui, M. H.; McDermott, M. M.; Mehra, R.; Murabito, J. M. Genetic determinants of the ankle-brachial index: A meta-analysis of a cardiovascular candidate gene 50K SNP panel in the candidate gene association resource (CARe) consortium. Atherosclerosis 2012, 222(1), 138–147. [Google Scholar] [CrossRef]
- Yang, J.; Weedon, M. N.; Purcell, S.; Lettre, G.; Estrada, K.; Willer, C. J.; Smith, A. V.; Ingelsson, E.; O’Connell, J. R.; Mangino, M.; Mägi, R.; Madden, P. A.; Heath, A. C.; Nyholt, D. R.; Martin, N. G.; Montgomery, G. W.; Frayling, T. M.; Hirschhorn, J. N.; McCarthy, M. I.; Visscher, P. M. Genomic inflation factors under polygenic inheritance. European Journal of Human Genetics 2011, 19(7), 807–812. [Google Scholar] [CrossRef]
- Yao, Y. Laminin: loss-of-function studies. Cellular and Molecular Life Sciences 2016, 74(6), 1095–1115. [Google Scholar] [CrossRef] [PubMed]
- Fox, C. S.; Golden, S. H.; Anderson, C.; Bray, G. A.; Burke, L. E.; De Boer, I. H.; Deedwania, P.; Eckel, R. H.; Ershow, A. G.; Fradkin, J.; Inzucchi, S. E.; Kosiborod, M.; Nelson, R. G.; Patel, M. J.; Pignone, M.; Quinn, L.; Schauer, P. R.; Selvin, E.; Vafiadis, D. K. Update on prevention of cardiovascular disease in adults with Type 2 diabetes mellitus in light of recent evidence: A scientific statement from the American Heart Association and the American Diabetes Association. Diabetes Care 2015, 38(9), 1777–1803. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, J. A.; Mulder, H.; Jones, W. S.; Rockhold, F. W.; Baumgartner, I.; Berger, J. S.; Blomster, J. I.; Fowkes, F. G. R.; Held, P.; Katona, B. G.; Mahaffey, K. W.; Norgren, L.; Hiatt, W. R.; Patel, M. R. Polyvascular disease and risk of major adverse cardiovascular events in peripheral artery disease. JAMA Network Open 2018, 1(7), e185239. [Google Scholar] [CrossRef]
- McDermott, M. M.; Kibbe, M.; Guralnik, J. M.; Pearce, W. H.; Tian, L.; Liao, Y.; Zhao, L.; Criqui, M. H. Comparative effectiveness study of self-directed walking exercise, lower extremity revascularization, and functional decline in peripheral artery disease. Journal of Vascular Surgery 2013, 57(4), 990–996.e1. [Google Scholar] [CrossRef]
- Ricco, J.; Bartelink, M.; Björck, M.; Brodmann, M.; Cohnert, T.; Czerny, M.; De Carlo, M.; Debus, S.; Espinola-Klein, C.; Kahan, T.; Kownator, S.; Mazzolai, L.; Naylor, A. R.; Röther, J.; Sprynger, M.; Tendera, M.; Tepe, G.; Venermo, M.; Vlachopoulos, C.; Obiekezie, A. 2017 ESC Guidelines on the Diagnosis and Treatment of Peripheral Arterial Diseases, in collaboration with the European Society for Vascular Surgery (ESVS). European Heart Journal 2017, 39(9), 763–816. [Google Scholar]
- Gerhard-Herman, M. D.; Gornik, H. L.; Barrett, C.; Barshes, N. R.; Corriere, M. A.; Drachman, D. E.; Fleisher, L. A.; Fowkes, F. G. R.; Hamburg, N. M.; Kinlay, S.; Lookstein, R.; Misra, S.; Mureebe, L.; Olin, J. W.; Patel, R. A.; Regensteiner, J. G.; Schanzer, A.; Shishehbor, M. H.; Stewart, K. J.; Walsh, M. E. 2016 AHA/ACC Guideline on the Management of Patients with Lower Extremity Peripheral Artery Disease: Executive summary: A report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines. Circulation 2016, 135(12). [Google Scholar] [CrossRef] [PubMed]
- Kullo, I. J.; Leeper, N. J. The genetic basis of peripheral arterial disease: current knowledge, challenges, and future directions. Circulation research 2015, 116(9), 1551–1560. [Google Scholar] [CrossRef]
- Leeper, N. J.; Kullo, I. J.; Cooke, J. P. Genetics of peripheral artery Disease. Circulation 2012, 125(25), 3220–3228. [Google Scholar] [CrossRef] [PubMed]
- Morley, R. L.; Sharma, A.; Horsch, A. D.; Hinchliffe, R. J. Peripheral artery disease. BMJ 2018, j5842. [Google Scholar] [CrossRef] [PubMed]
- Korutla, R.; Tedder, D.; Brogan, K.; Milosevic, M.; Wilczek, M. P.; Shehadeh, N.; Shara, N.; Ross, E. G.; Amal, S. Analysis and Visualization of Confounders and Treatment Pathways Leading to Amputation and Non-Amputation in Peripheral Artery Disease Patients Using Sankey Diagrams: Enhancing Explainability. Biomedicines 2025, 13(2), 258. [Google Scholar] [CrossRef]
- Barraza-Flores, P.; Bates, C. R.; Oliveira-Santos, A.; Burkin, D. J. Laminin and integrin in LAMA2-Related Congenital Muscular Dystrophy: From Disease to Therapeutics. Frontiers in Molecular Neuroscience 2020, 13. [Google Scholar] [CrossRef]
- Brignone, M. S.; Lanciotti, A.; Camerini, S.; De Nuccio, C.; Petrucci, T. C.; Visentin, S.; Ambrosini, E. MLC1 protein: a likely link between leukodystrophies and brain channelopathies. Frontiers in Cellular Neuroscience 2015, 09. [Google Scholar] [CrossRef]
- Zhang, M.; Bai, J.; Yuan, H.; Duan, X.; Yu, L.; Li, Y.; Li, K.; Rile, S.; Wang, X.; Wang, H.; Liu, P.; Yan, J.; Wang, C. BRD1 deficiency affects SREBF1-related lipid metabolism through regulating H3K9ac/H3K9me3 transition to inhibit HCC progression. Cell Death and Disease 2025, 16(1). [Google Scholar] [CrossRef] [PubMed]
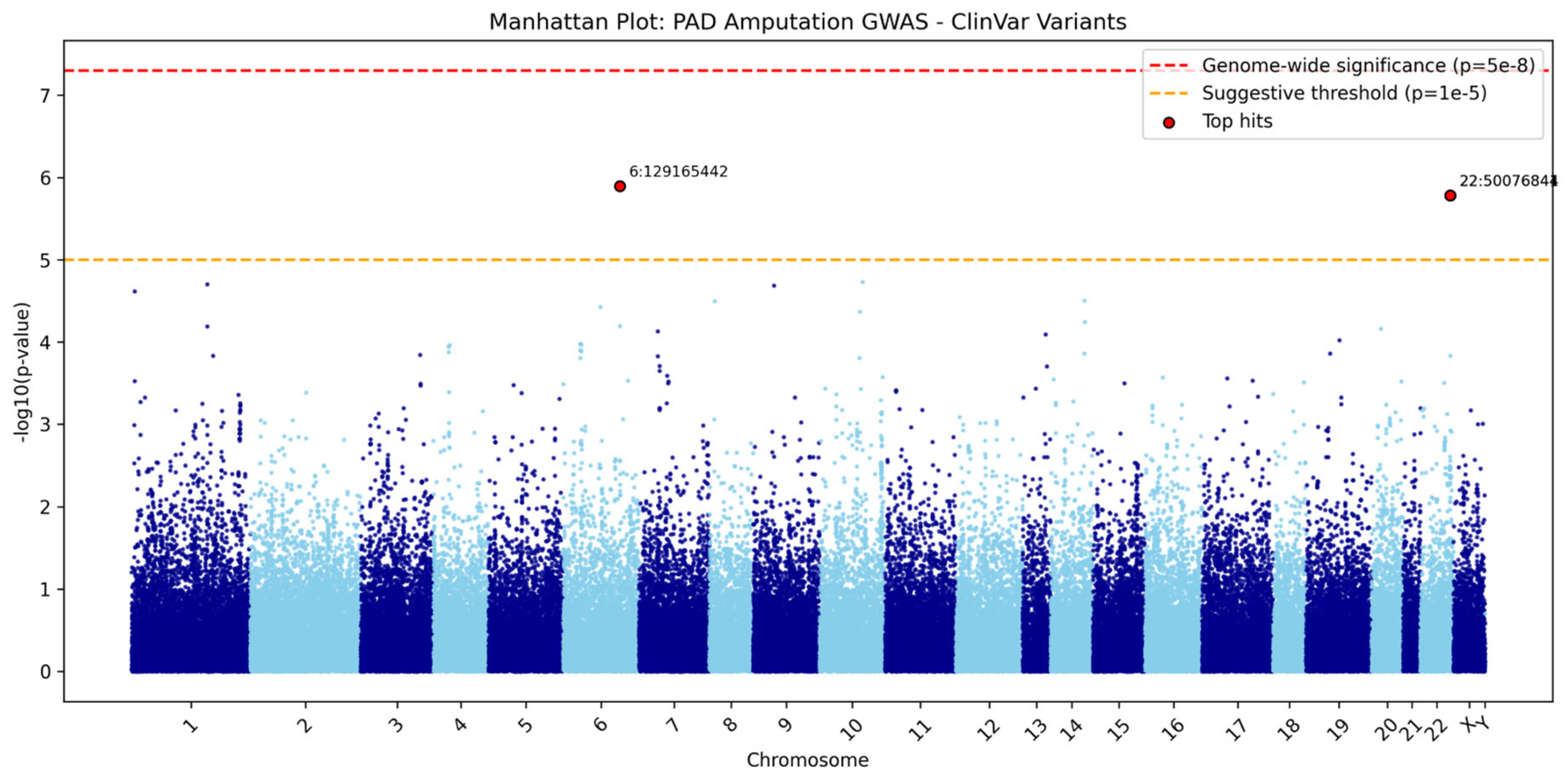
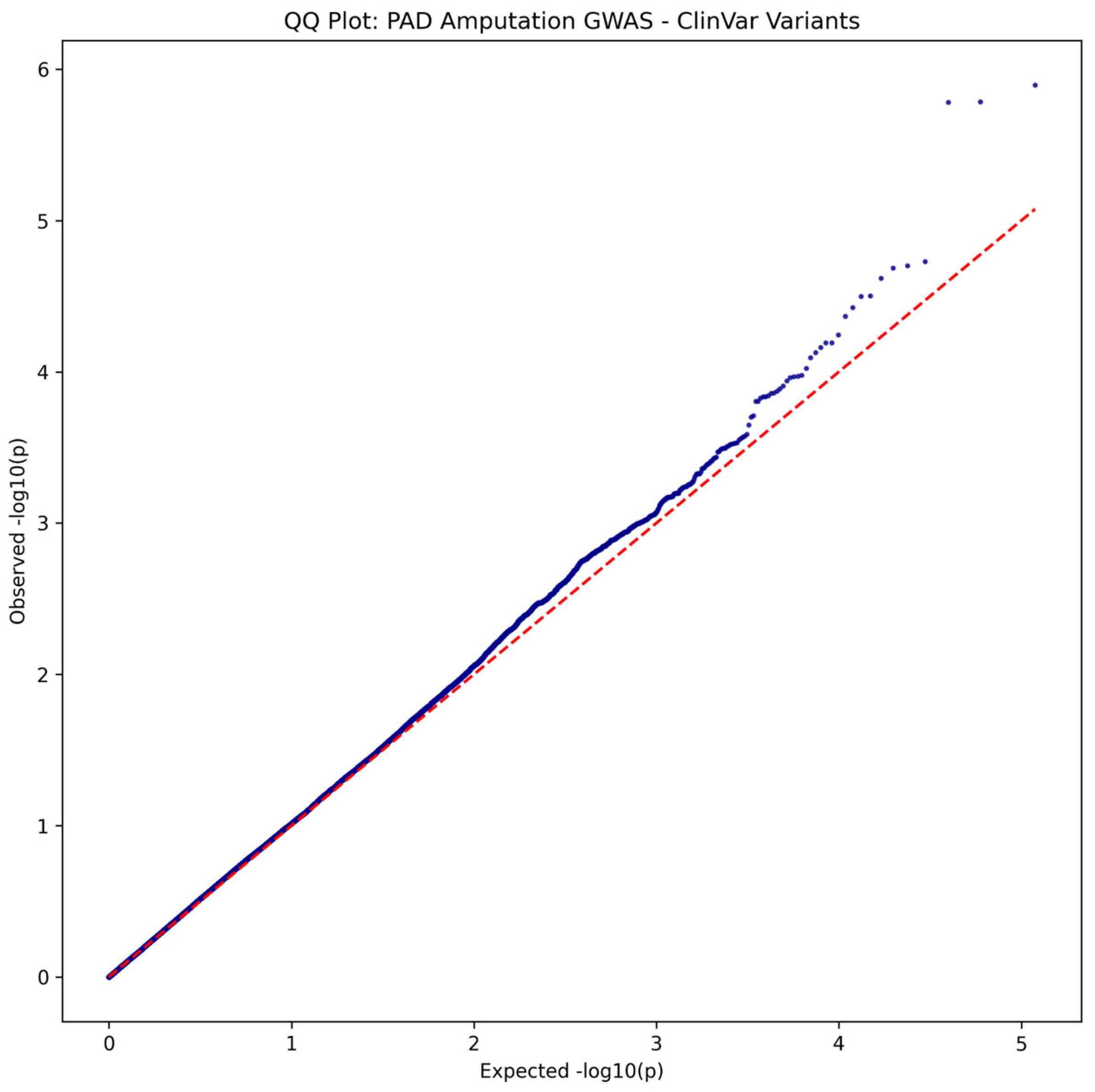
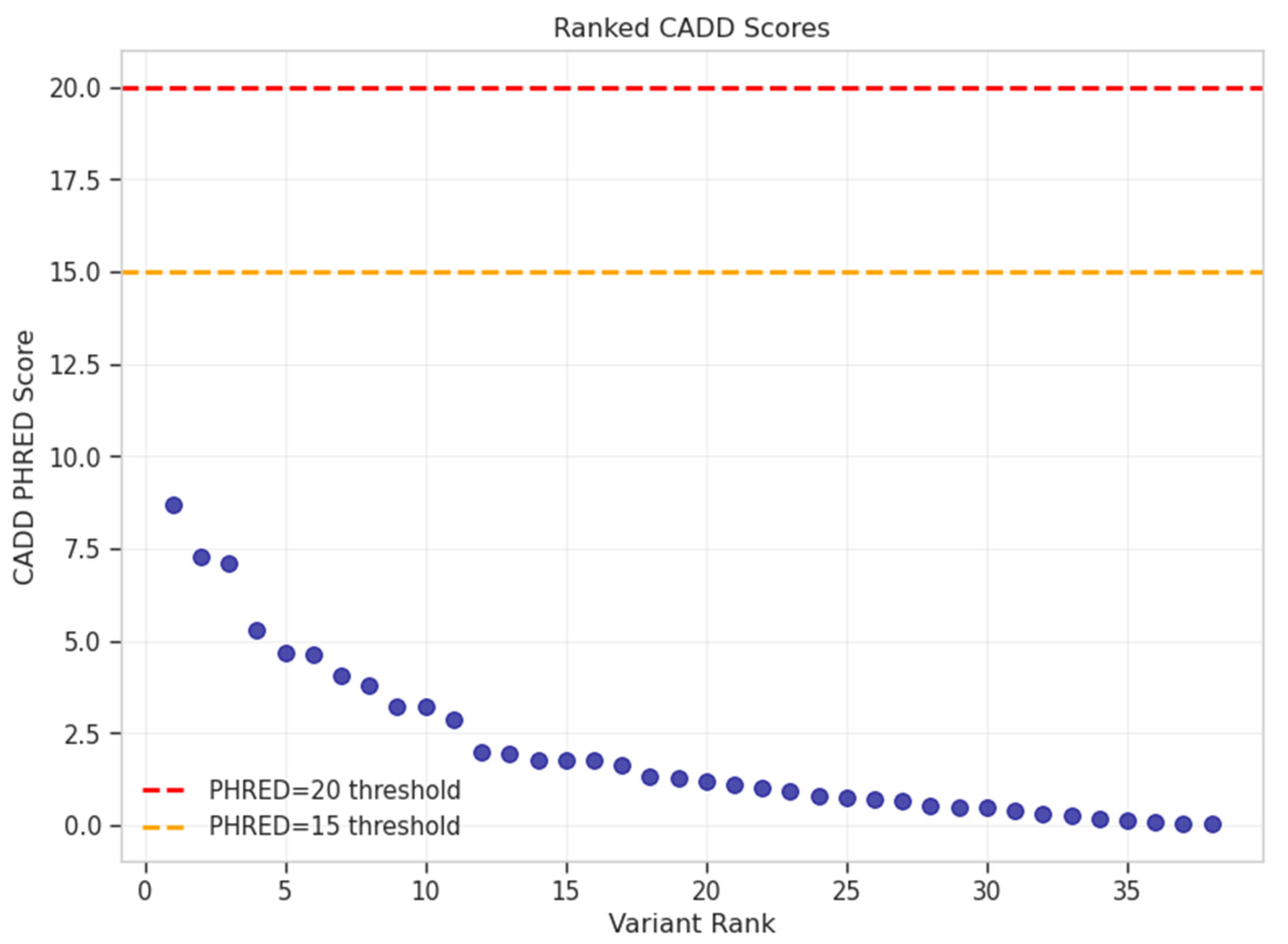
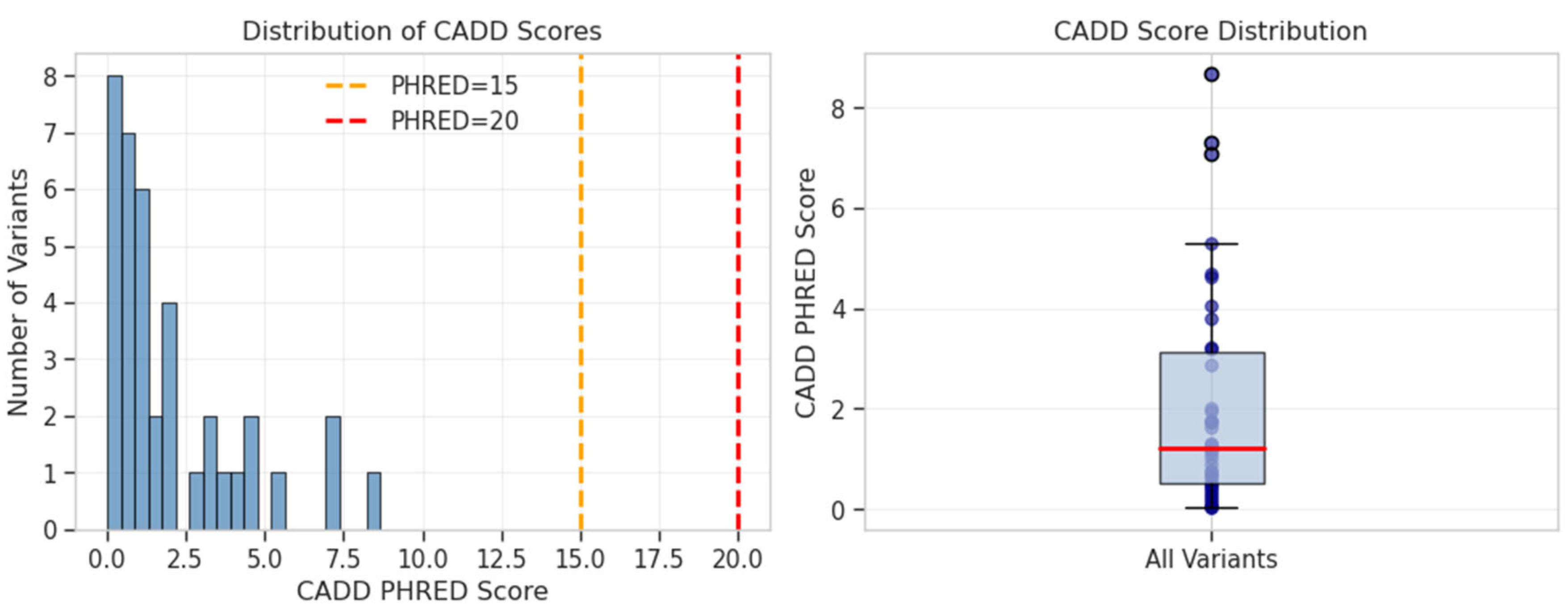
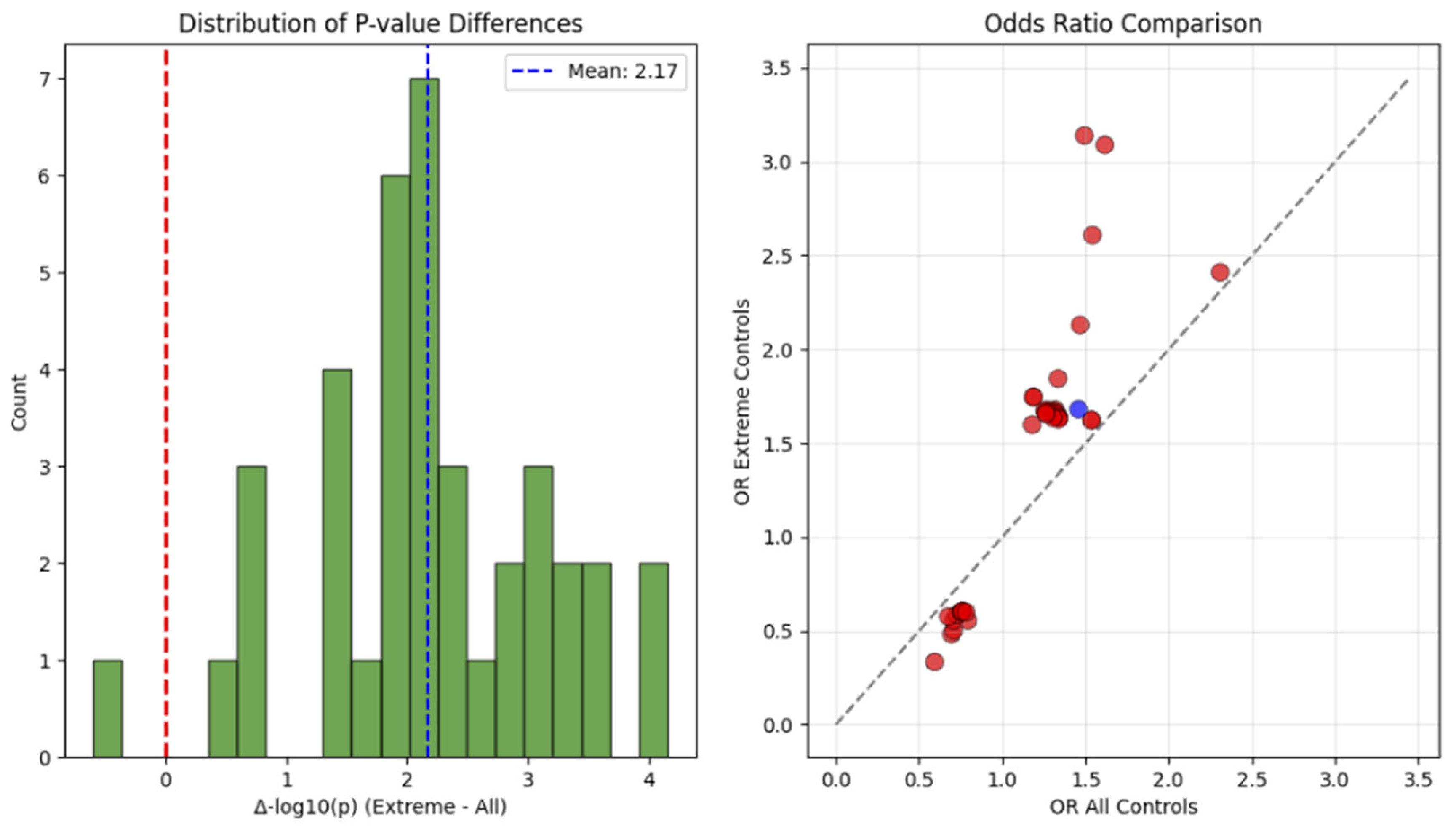
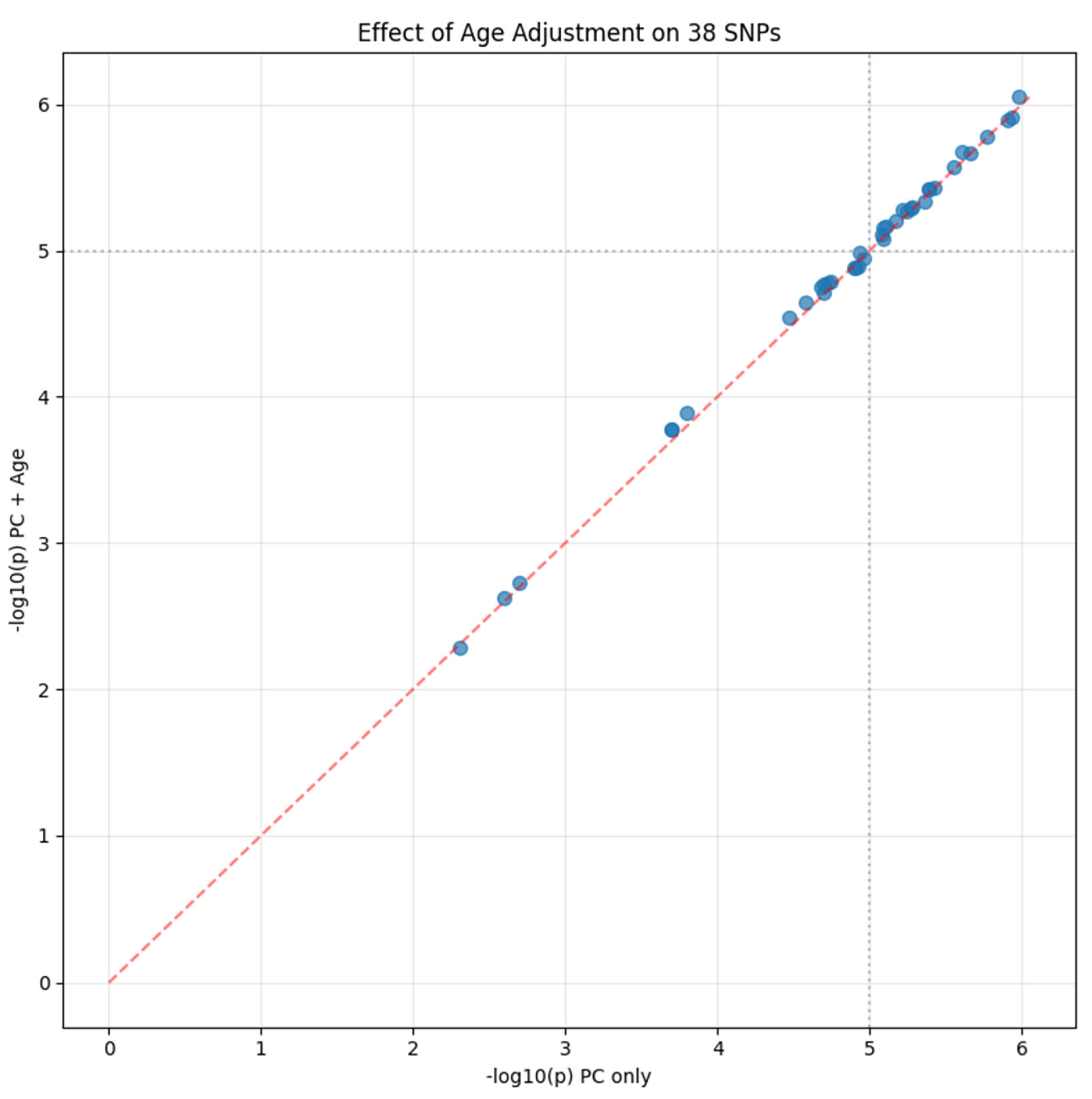
| Chr:Position | rsID | Ref>Alt | Beta | P-value | OR | Gene/Nearest Gene | Consequence | Distance (bp) | MAF (%) | eQTL | CADD PHRED |
| chr6:129165442 | rs62420977 | G>A | 0.88 | 1.00×10⁻⁶ | 2.41 | LAMA2 | intron_variant | 0 | 2.15 | Yes | 0.4 |
| chr22:50076844 | rs6010165 | G>A | 0.484 | 2.00×10⁻⁶ | 1.62 | MLC1 | synonymous_variant | 0 | 11.4 | Yes | 0.54 |
| chr22:50076841 | rs267607236 | T>C | 0.484 | 2.00×10⁻⁶ | 1.62 | MLC1 | splice_region_variant | 0 | 10.8 | No | 0.02 |
| Chr:Position | rsID | Ref>Alt | Beta | P-value | OR | Gene/Nearest Gene | Consequence | Distance (bp) | MAF (%) |
eQTL (presence) |
CADD PHRED |
| chr4:160553934 | rs62332335 | G>C | -0.729 | 1.07×10⁻⁶ | 0.48 | LINC02477 | intron_variant | 0 | 29.8 | No | 7.09 |
| chr10:126157589 | rs58661519 | C>T | 0.756 | 1.13×10⁻⁶ | 2.13 | ADAM12 | intron_variant | 0 | 15.5 | No | 5.3 |
| chr11:13732125 | rs1565356411 | TAA>T | 1.128 | 1.58×10⁻⁶ | 3.09 | FAR1 | 3_prime_UTR_variant | 0 | 6.4 | No | 0.27 |
| chr1:151481028 | rs58440816 | G>C | -0.588 | 1.80×10⁻⁶ | 0.56 | POGZ | intergenic_variant | 21,534 | 46.2 | No | 1.3 |
| chr11:11089988 | rs59583539 | T>C | 0.557 | 2.00×10⁻⁶ | 1.75 | LINC02752 | intron_variant | 0 | 12.4 | No | 7.29 |
| chr11:11089978 | rs59299975 | T>C | 0.557 | 2.00×10⁻⁶ | 1.75 | LINC02752 | intron_variant | 0 | 12.4 | No | 0.02 |
| chr10:16831081 | rs57775857 | C>T | 0.612 | 2.06×10⁻⁶ | 1.84 | CUBN | intron_variant | 0 | 33.9 | No | 2 |
| chr4:167358505 | rs10390166 | A>T | 0.517 | 2.17×10⁻⁶ | 1.68 | RN7SL776P | intergenic_variant | 41,990 | 30.6 | No | 4.06 |
| chr4:167355663 | rs58299204 | G>A | 0.515 | 2.23×10⁻⁶ | 1.67 | RN7SL776P | intergenic_variant | 44,832 | 29 | No | 1.03 |
| chr19:7375984 | rs60516505 | A>G | 0.519 | 2.54×10⁻⁶ | 1.68 | ARHGEF18-AS1, ARHGEF18 | intron_variant | 0 | 27.4 | No | 0.08 |
| chr4:160552306 | rs377194319 | T>TCTAA | -0.691 | 2.69×10⁻⁶ | 0.5 | LINC02477 | intron_variant | 0 | 29.8 | No | 1.74 |
| chr4:167358166 | rs12108386 | C>A | 0.51 | 2.82×10⁻⁶ | 1.67 | RN7SL776P | intergenic_variant | 42,329 | 32.5 | No | 0.68 |
| chr4:167372657 | rs17601234 | T>A | 0.5 | 3.02×10⁻⁶ | 1.65 | RN7SL776P | intergenic_variant | 27,838 | 29 | No | 1.11 |
| chr21:44596298 | rs17004490 | T>C | 1.144 | 3.15×10⁻⁶ | 3.14 | TSPEAR, KRTAP10-7, KRTAP10-6 | intron_variant | 0 | 14.9 | No | 4.69 |
| chr4:167373408 | rs111484533 | A>ACAGT | 0.494 | 3.43×10⁻⁶ | 1.64 | RN7SL776P | intergenic_variant | 27,087 | 31.1 | No | 2.88 |
| chr21:19075049 | rs143785792 | G>A | -0.587 | 3.45×10⁻⁶ | 0.56 | ENSG00000235965 | intergenic_variant | 27,365 | 37.2 | No | 3.79 |
| chr4:167362902 | rs60353699 | G>T | 0.504 | 3.73×10⁻⁶ | 1.66 | RN7SL776P | intergenic_variant | 37,593 | 29 | No | 1.62 |
| chr4:167373717 | rs59233454 | G>A | 0.491 | 4.13×10⁻⁶ | 1.63 | RN7SL776P | intergenic_variant | 26,778 | 29.1 | No | 1.77 |
| chr5:133657373 | rs59870923 | T>C | 0.516 | 4.40×10⁻⁶ | 1.68 | FSTL4 | intergenic_variant | 44,832 | 51.8 | No | 3.21 |
| chr21:19051267 | rs1985175965 | G>T | -0.534 | 5.00×10⁻⁶ | 0.59 | ENSG00000235965 | upstream_gene_variant | 3,583 | 33.8 | No | 0.32 |
| chr4:167374290 | rs58934032 | C>T | 0.488 | 5.09×10⁻⁶ | 1.63 | RN7SL776P | intergenic_variant | 26,205 | 29 | No | 1.77 |
| chr4:167367142 | rs59244366 | T>C | 0.492 | 5.52×10⁻⁶ | 1.64 | RN7SL776P | intergenic_variant | 33,353 | 29 | No | 8.66 |
| chr5:133661151 | rs1182945568 | T>C | 0.515 | 5.56×10⁻⁶ | 1.67 | FSTL4 | intergenic_variant | 48,610 | 52.8 | No | 0.75 |
| chr17:21269327 | rs2508018174 | G>A | -0.504 | 5.76×10⁻⁶ | 0.6 | ENSG00000289453 | intergenic_variant | 6,570 | 26.6 | No | 0.46 |
| chr7:73141070 | rs1801996601 | C>CCACA | 0.959 | 5.92×10⁻⁶ | 2.61 | SPDYE10 | intron_variant | 0 | 15.9 | No | 0.12 |
| chr17:21269385 | rs1045164134 | A>AT | -0.502 | 6.01×10⁻⁶ | 0.61 | ENSG00000289453 | intergenic_variant | 6,512 | 24.4 | No | 1.17 |
| chr6:70891807 | rs2460694 | T>C | -0.551 | 6.35×10⁻⁶ | 0.58 | B3GAT2 | intron_variant | 0 | 30.1 | Yes | 0.18 |
| chr10:49442 | rs1834436109 | AGCTCAGGTGTCCTT>A | -1.096 | 6.87×10⁻⁶ | 0.33 | TUBB8 | 5_prime_UTR_variant | 0 | 13.2 | No | 3.22 |
| chr17:21240761 | rs57745152 | G>A | 0.469 | 7.00×10⁻⁶ | 1.6 | NATD1, TMEM11-DT | 3_prime_UTR_variant | 0 | 10.7 | No | 0.66 |
| chr21:19056133 | rs28806580 | A>G | -0.509 | 7.05×10⁻⁶ | 0.6 | ENSG00000235965 | intergenic_variant | 8,449 | 26.4 | No | 0.78 |
| chr21:19051108 | rs9653673 | G>A | -0.515 | 7.26×10⁻⁶ | 0.6 | ENSG00000235965 | upstream_gene_variant | 3,424 | 26.3 | No | 4.62 |
| chr5:133664291 | rs56555387 | G>A | 0.508 | 7.28×10⁻⁶ | 1.66 | FSTL4 | intergenic_variant | 51,750 | 53.5 | No | 1.95 |
| chr11:20368197 | rs1256321231 | T>A | 0.505 | 8.68×10⁻⁶ | 1.66 | HTATIP2 | intron_variant | 0 | 12.8 | No | 0.49 |
| chr21:19051807 | rs1568913903 | T>TTA | -0.503 | 9.12×10⁻⁶ | 0.6 | ENSG00000235965 | upstream_gene_variant | 4,123 | 26.4 | No | 1.28 |
| chr7:37714274 | rs1215147035 | T>C | -0.513 | 9.65×10⁻⁶ | 0.6 | GPR141 | intron_variant | 0 | 29.9 | No | 0.9 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




