Submitted:
05 February 2026
Posted:
10 February 2026
You are already at the latest version
Abstract
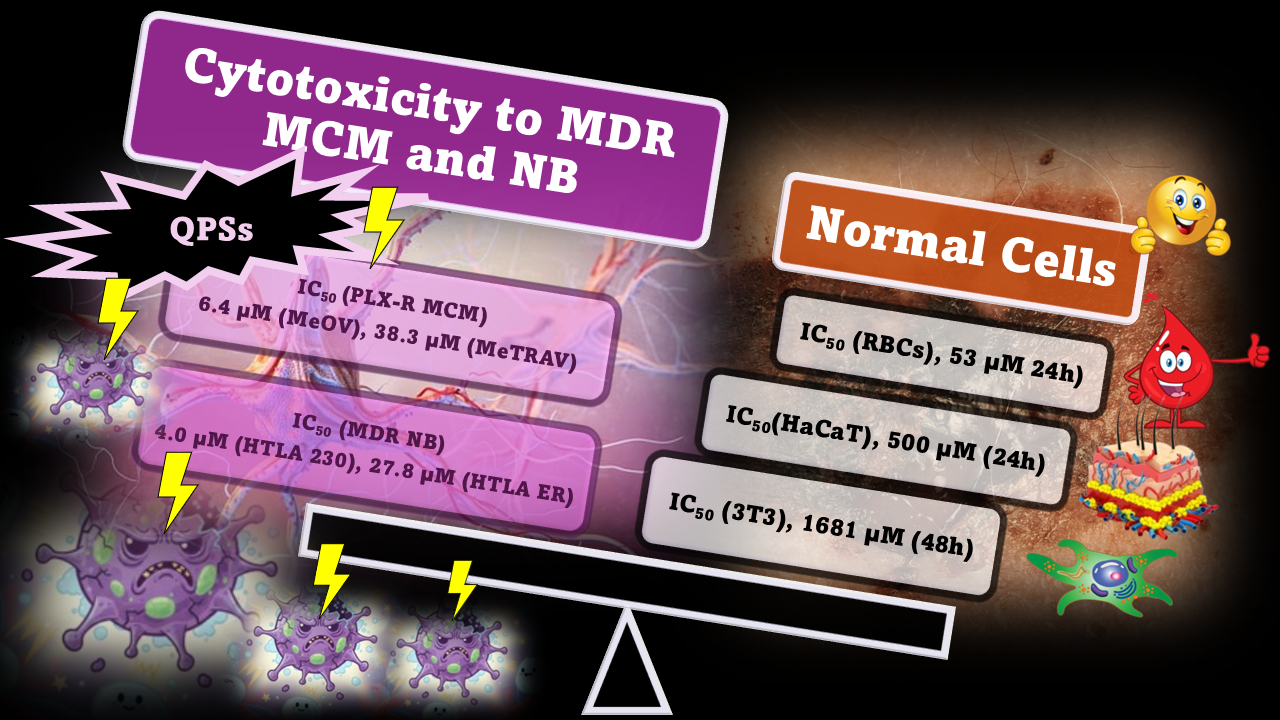

Keywords:
1. Introduction
1.1. The Rational of The Study
2. Results and Discussion
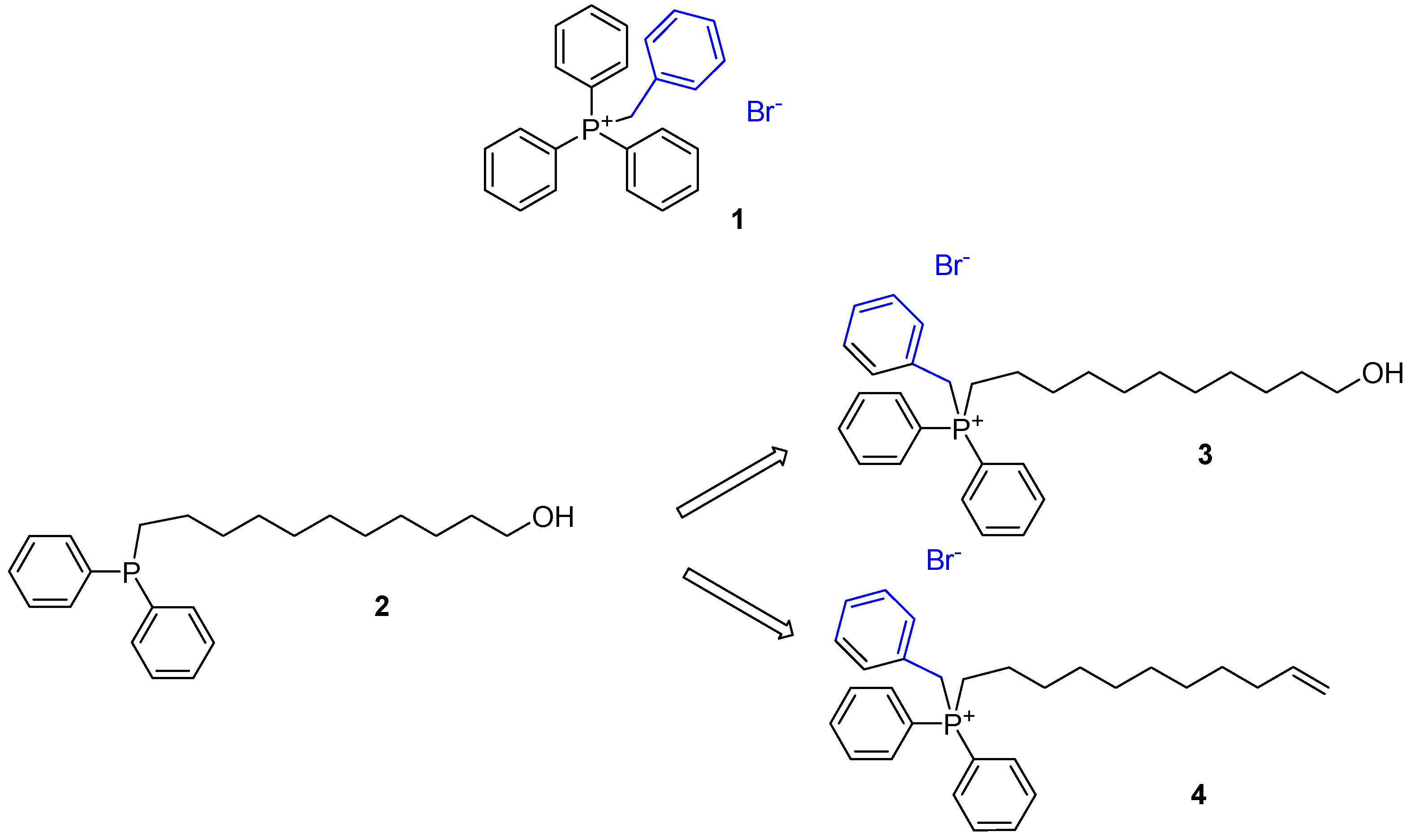
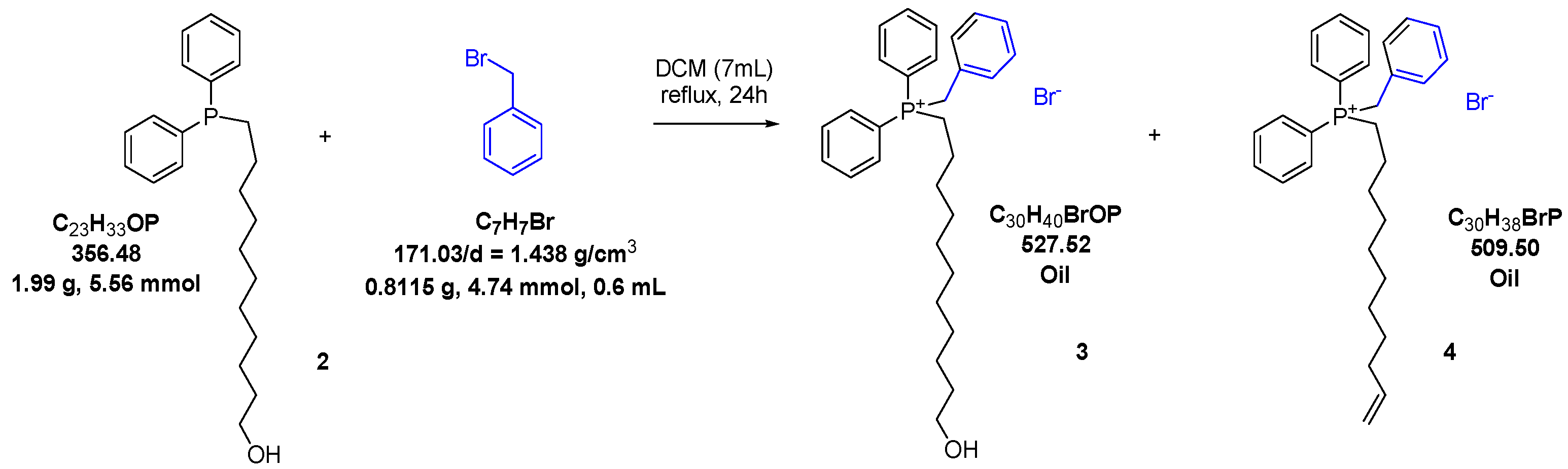
2.1. Synthesis of Phosphine Compound 2 and Benzyl Phosphonium Salts 1, 3 and 4
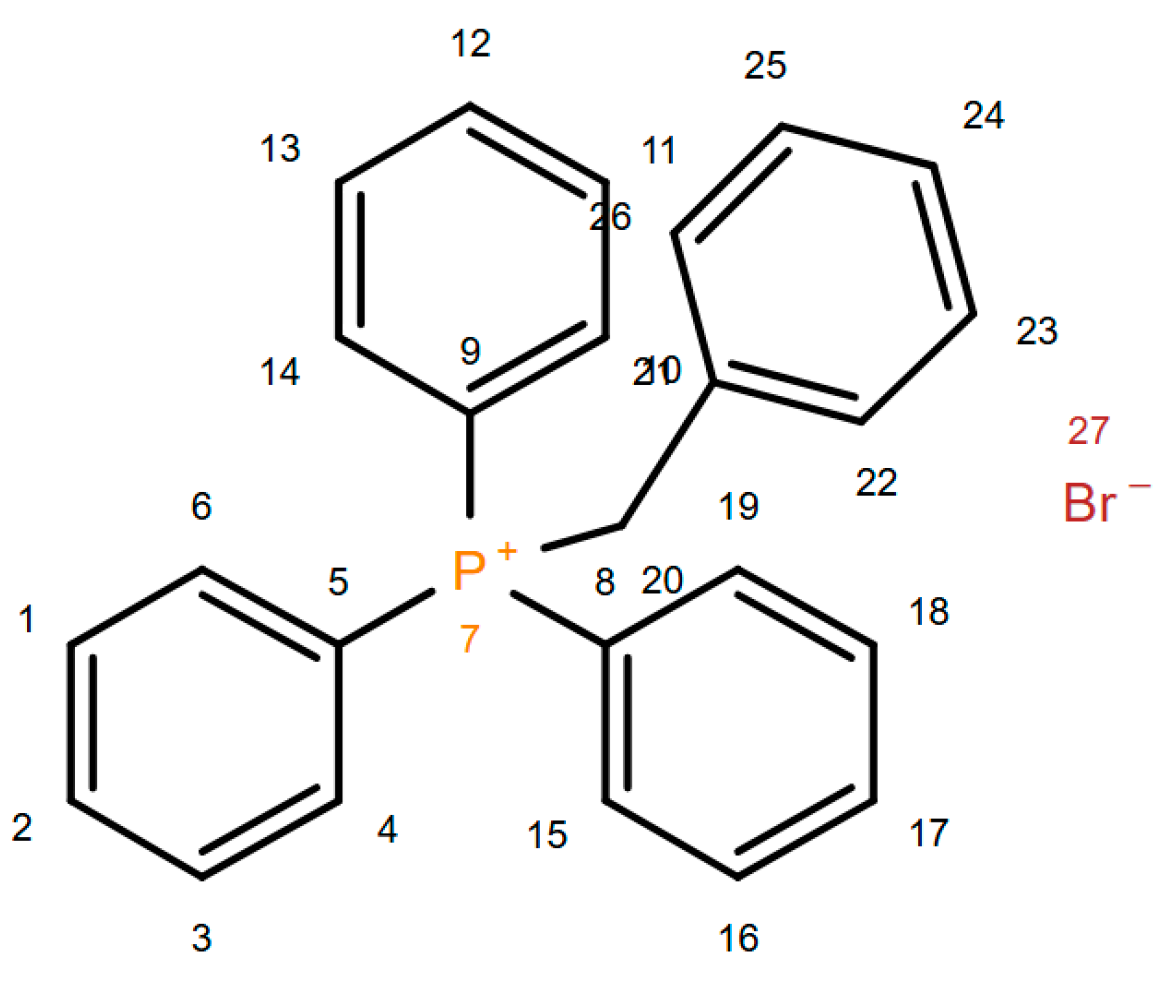
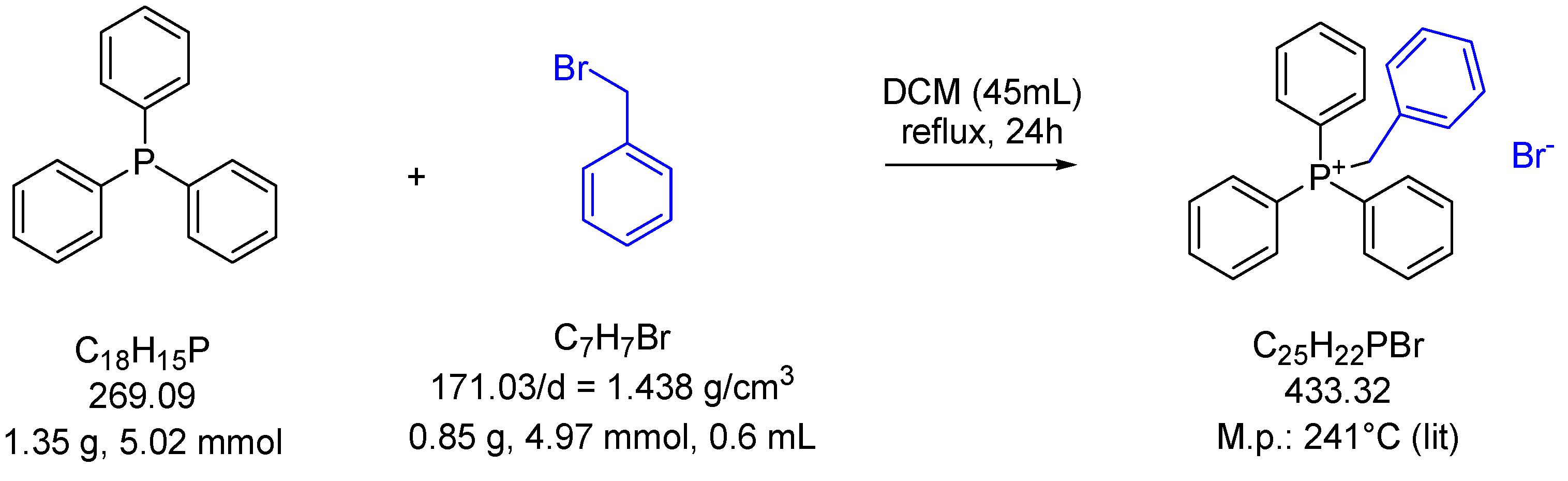
2.1.1. Benzyl Triphenyl Phosphonium Bromide 1
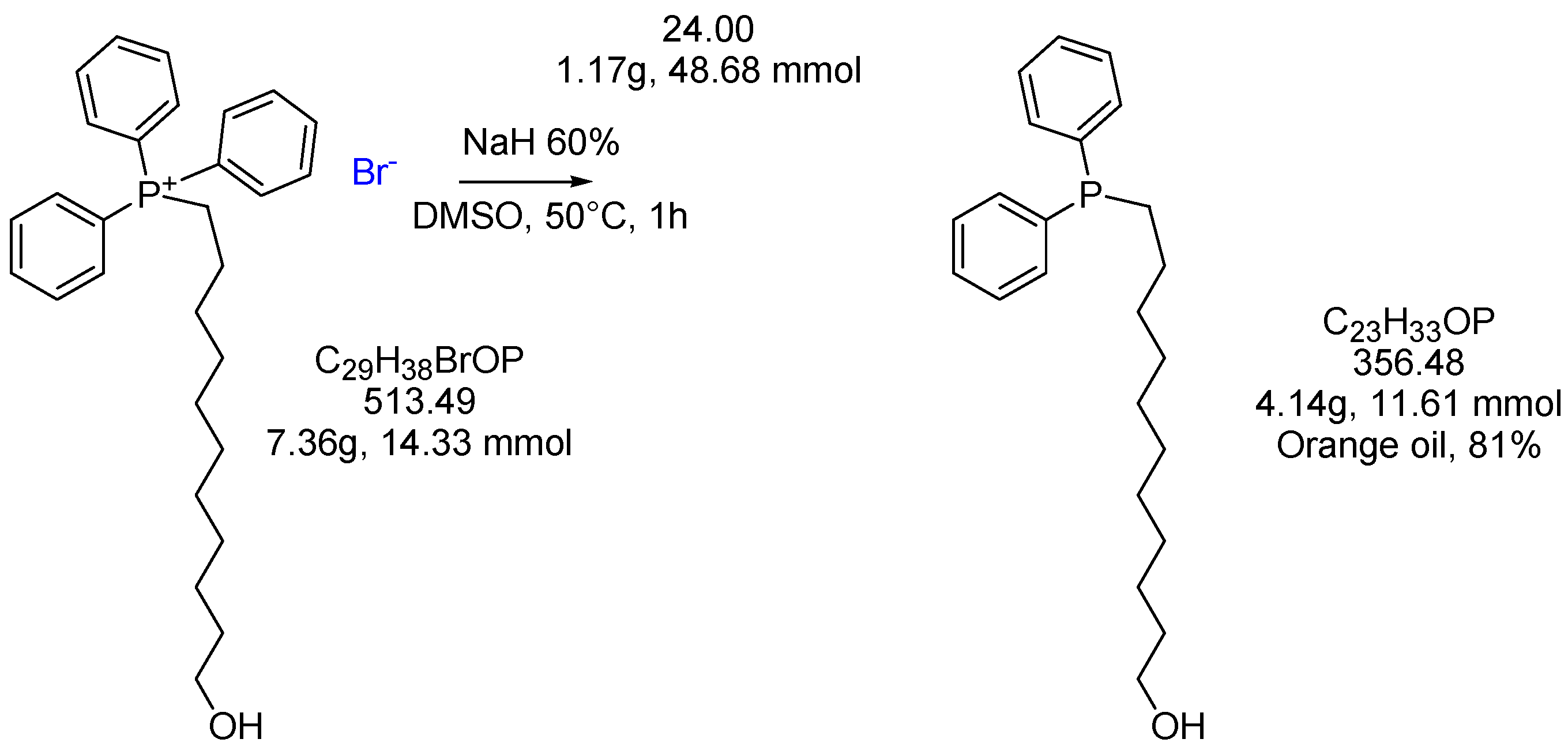
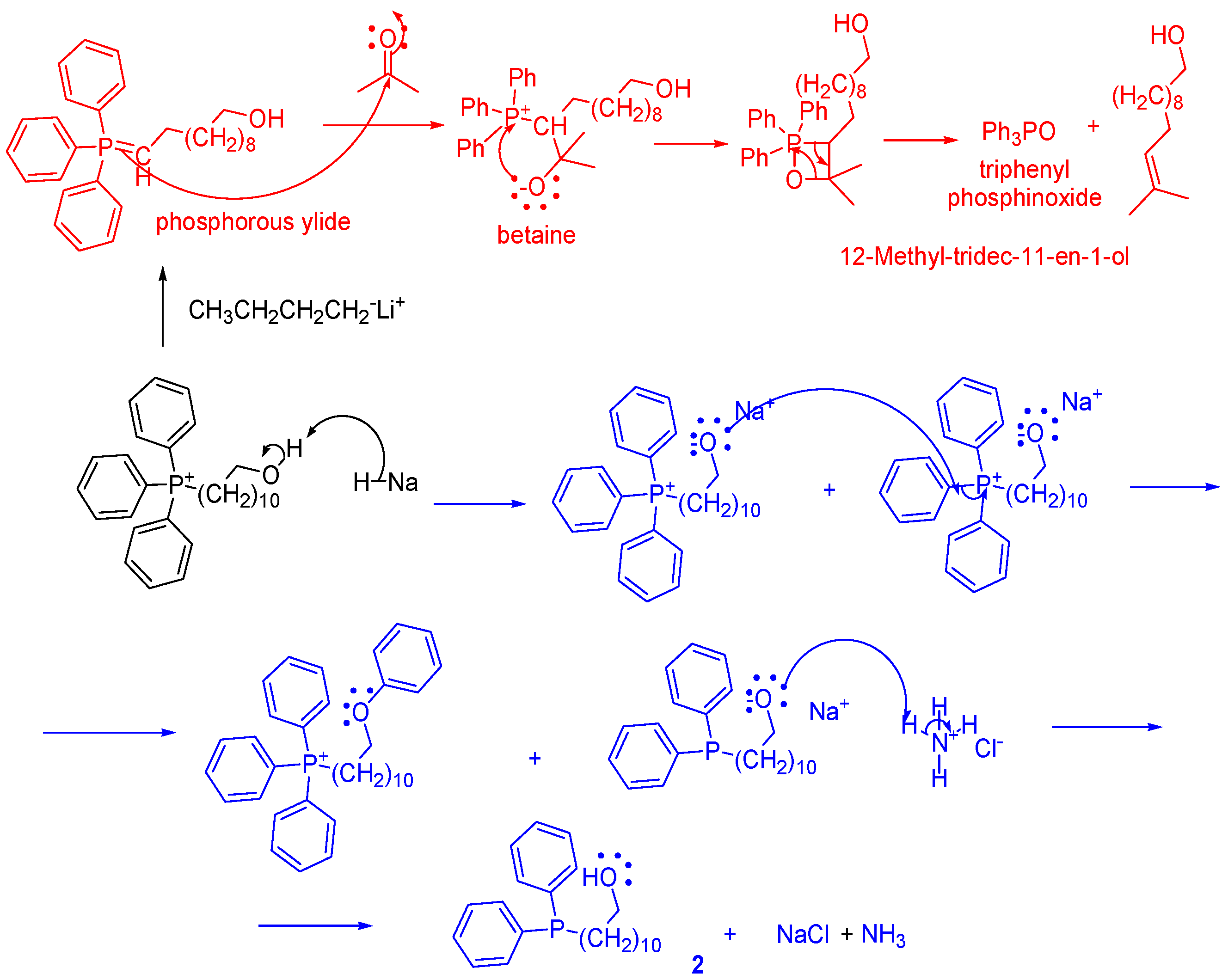
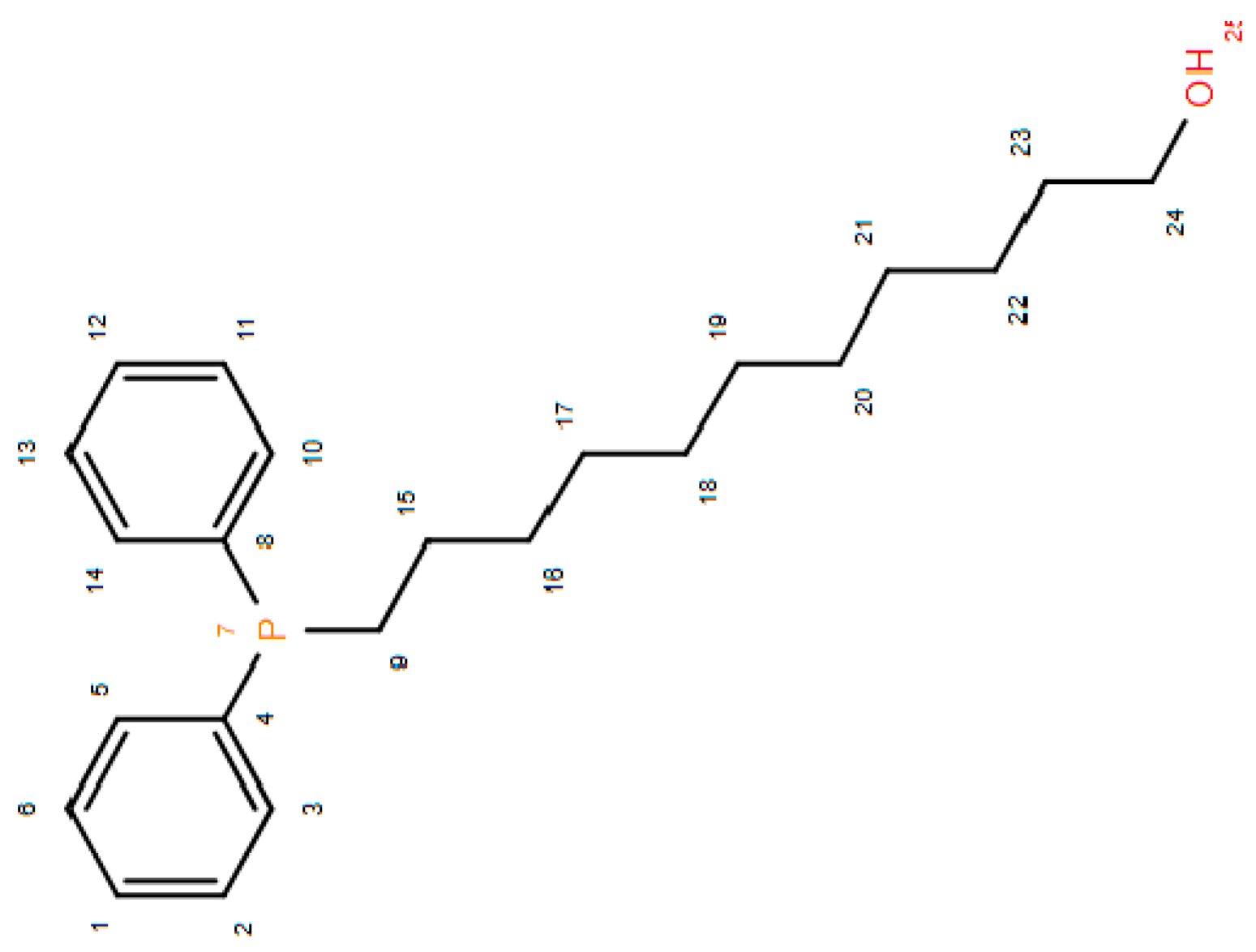
2.1.2. 11-(Di-Phenyl-Phosphine)-1-Undecanol (2) [45]
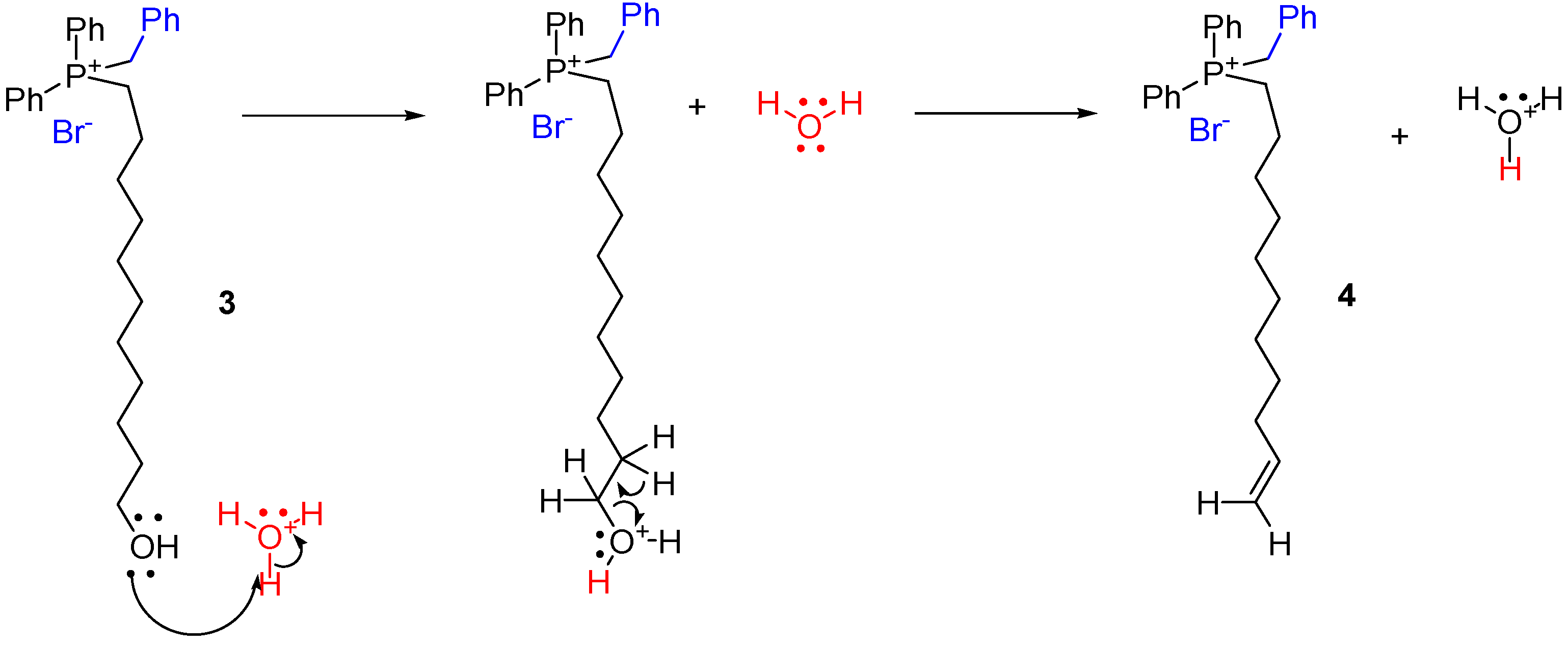
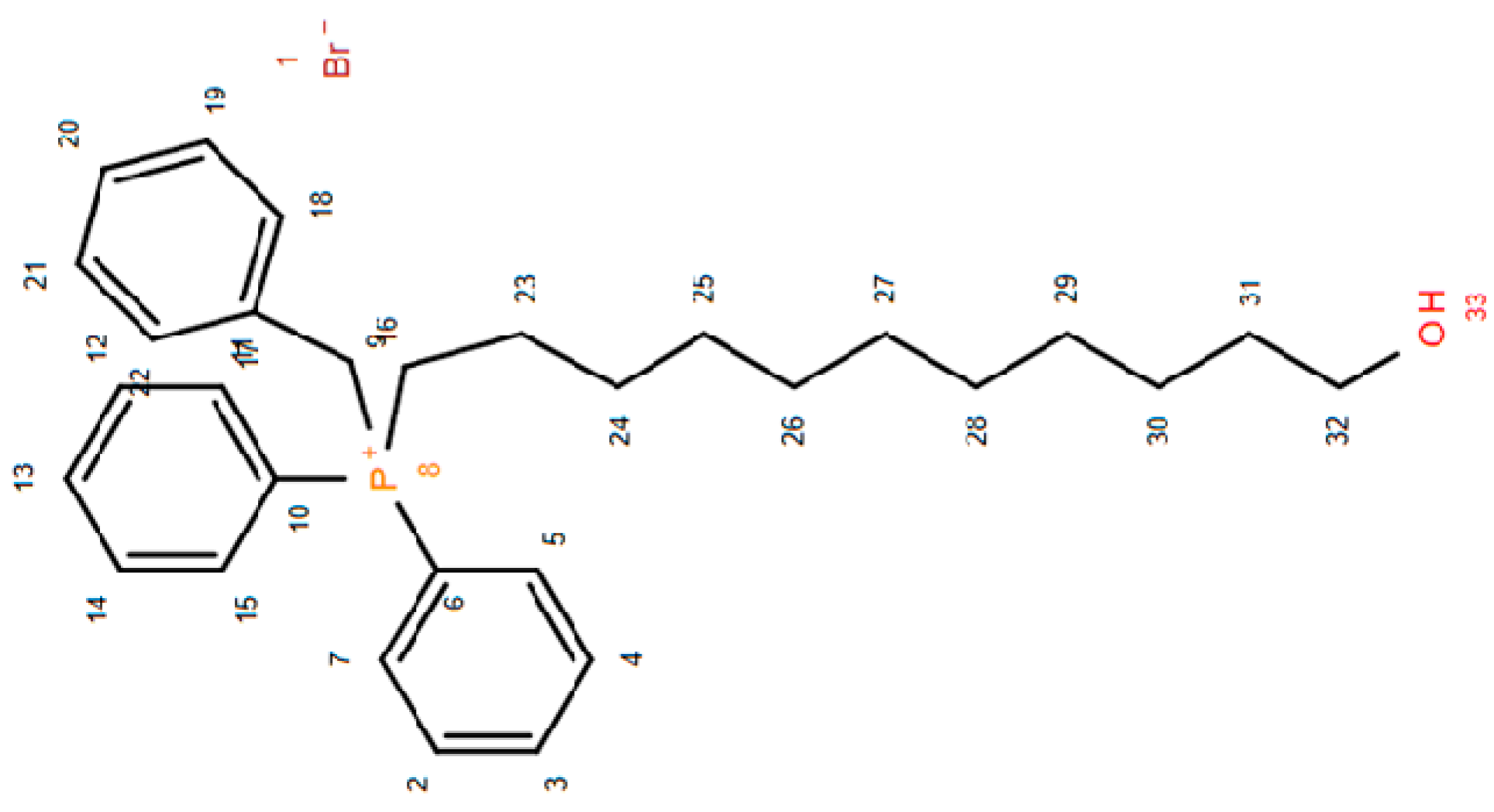
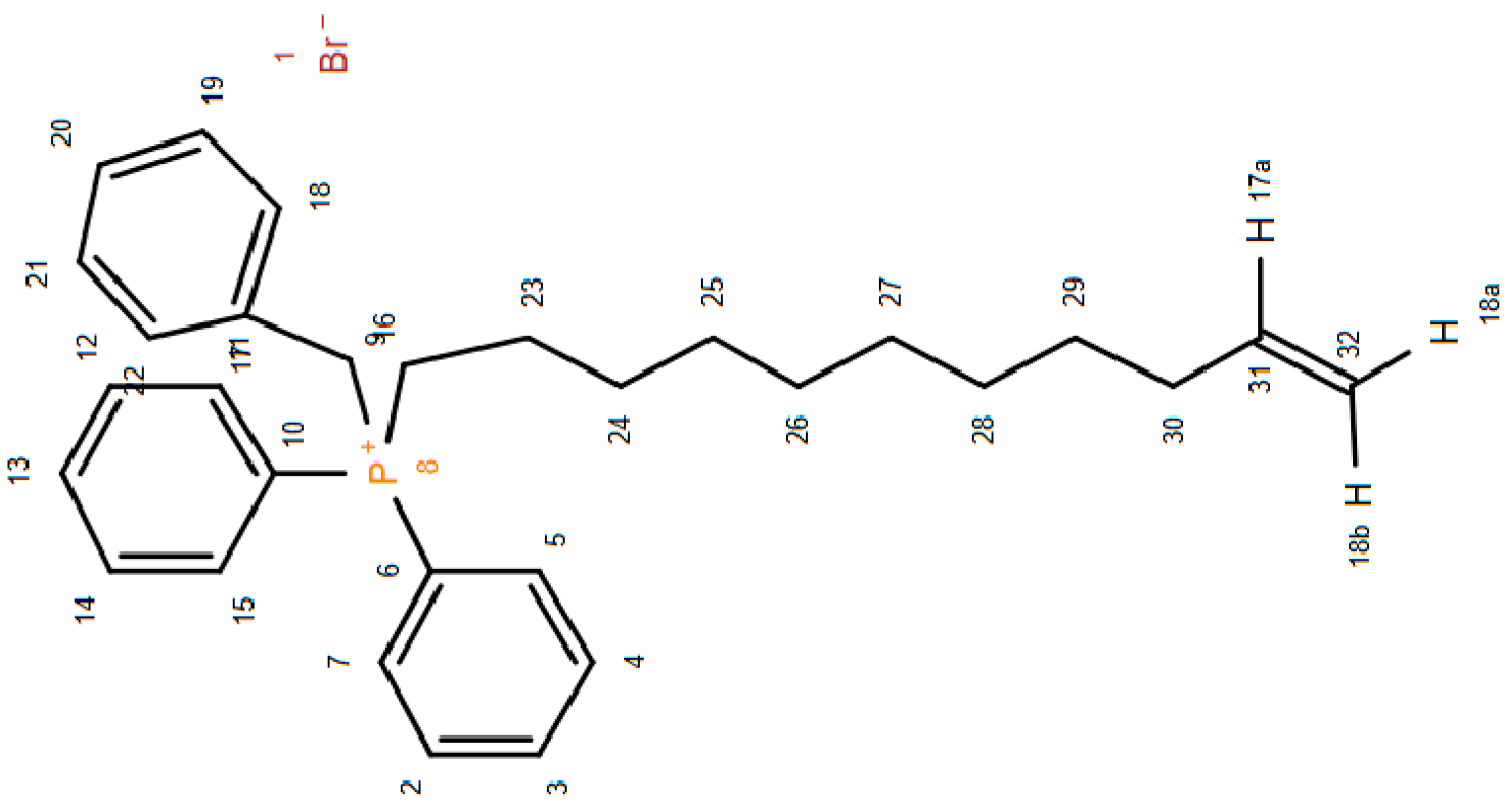
2.1.3. Benzyl-(11-Hydroxy-Undecyl)-Diphenyl-Phosphonium Bromide 3 and Benzyl-Diphenyl-Undec-10-Enyl-Phosphonium Bromide 4.
2.2. ATR-FTIR Spectra of Compounds 1-4
2.3. UV-Vis Spectra
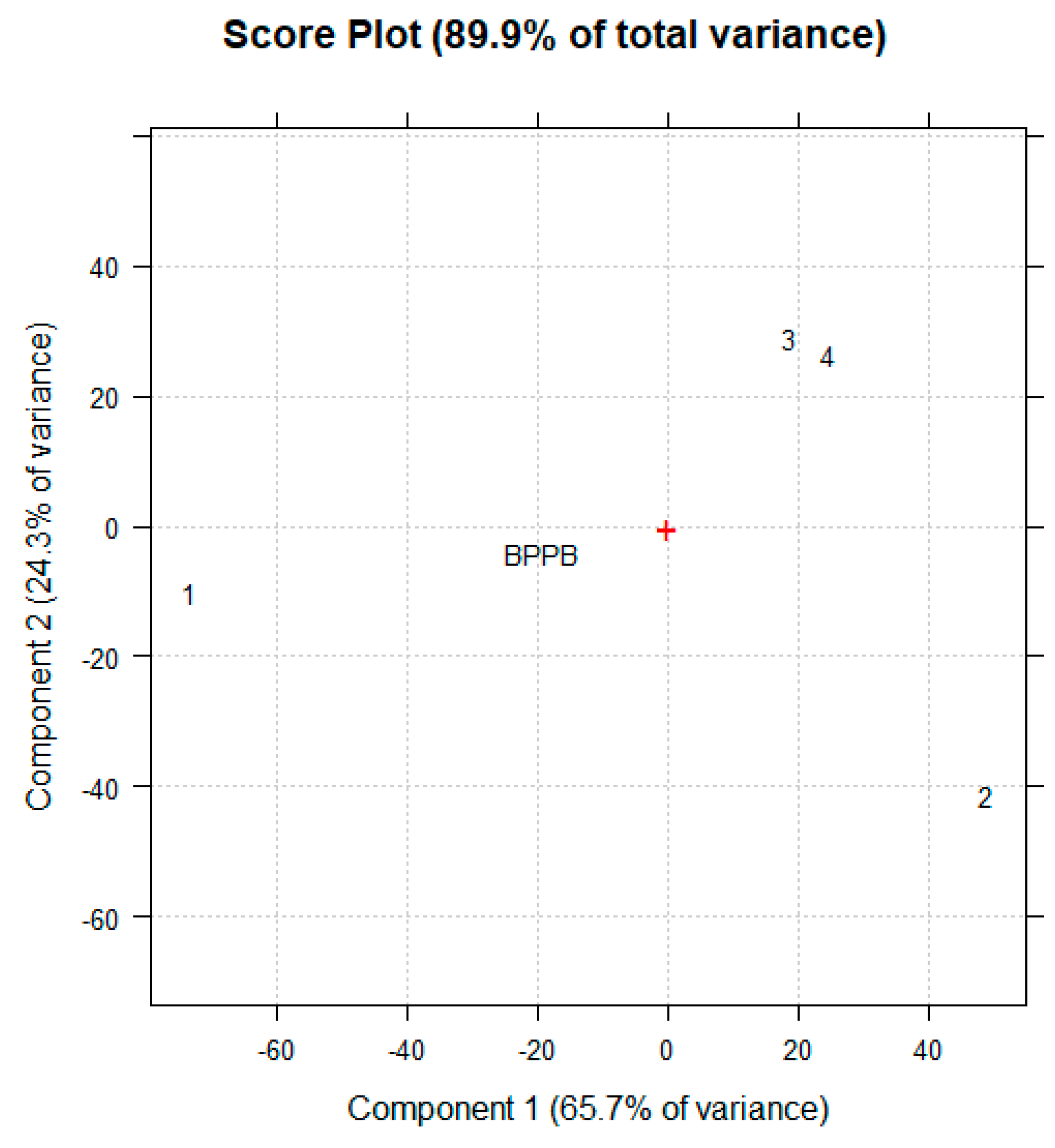
2.4. Principal Component Analysis (PCA) of ATR-FTIR UV-Vis and NMR Spectral Data
2.4.1. PCA of ATR-FTIR and UV-Vis Spectral Data
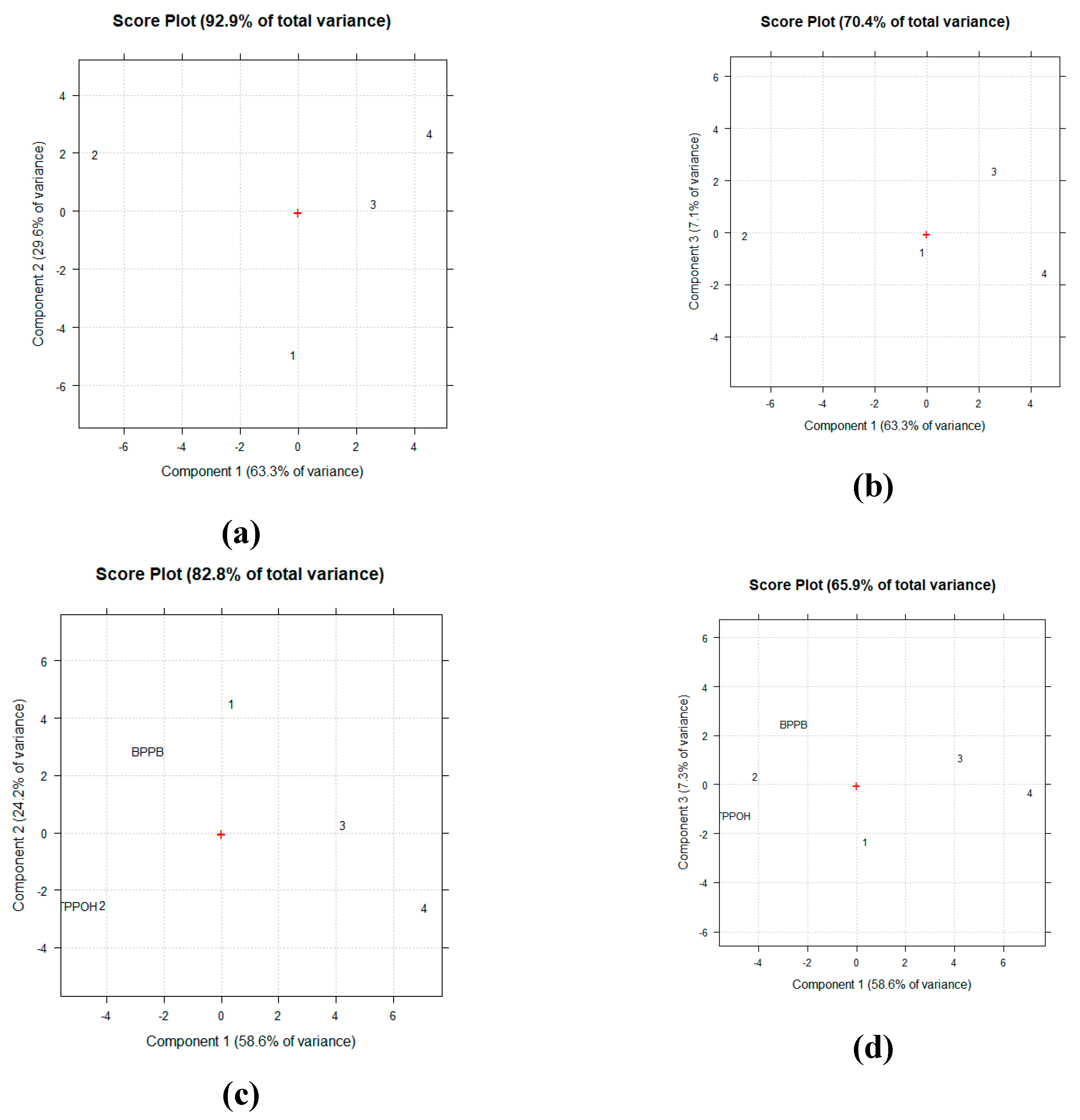
2.4.2. PCA of NMR Spectral Data
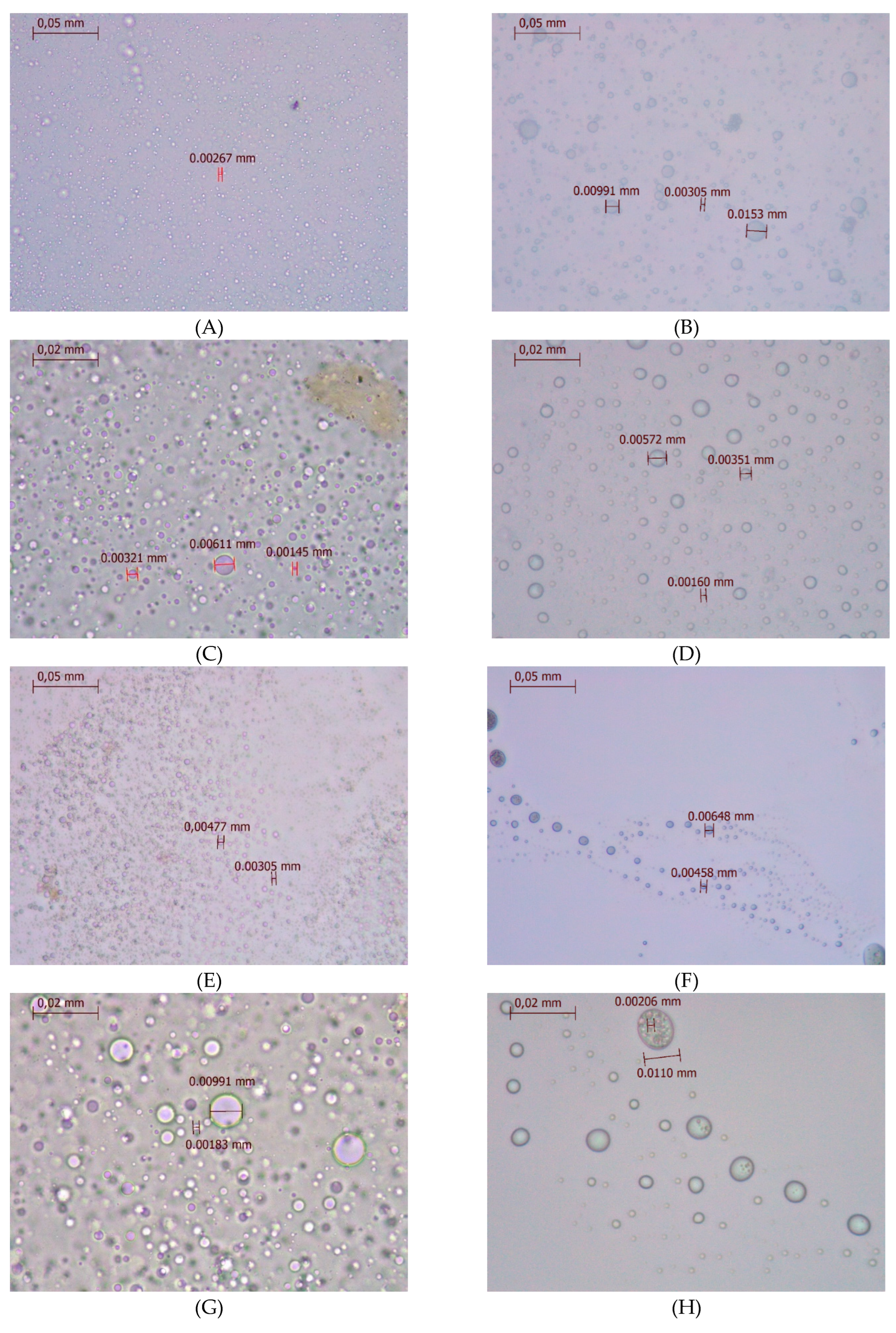
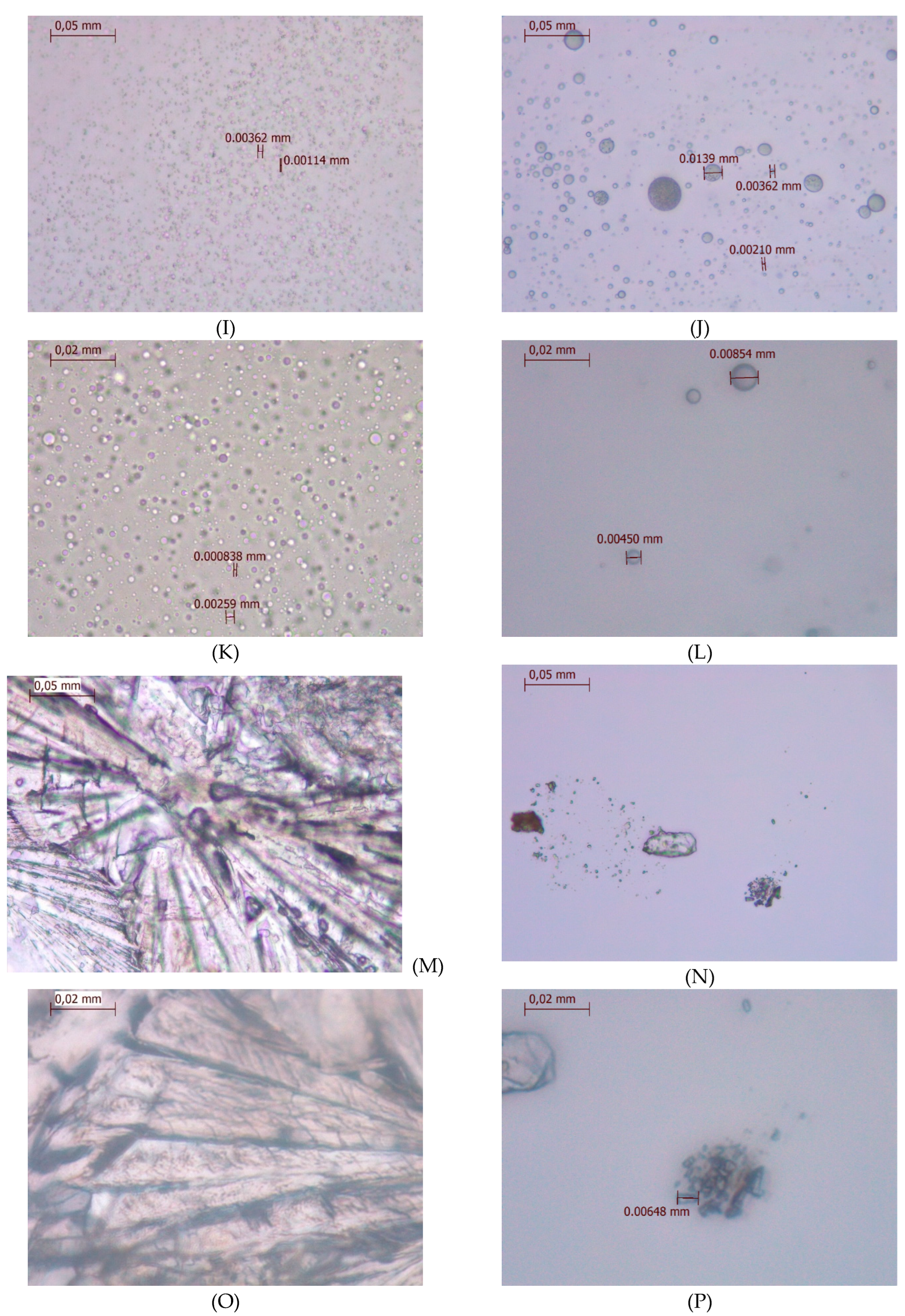
2.5. Optical Microscopy Analyses
2.6. Biological Effects of Compounds 1, 3 and 4 on Tumoral and Not Tumoral Human Cells Models
2.6.1. Concentration- and Time-Dependent Anticancer Effects of 1, 3 and 4 on MeOV PLX-R and MeTRAV PLX-R Cell Viability
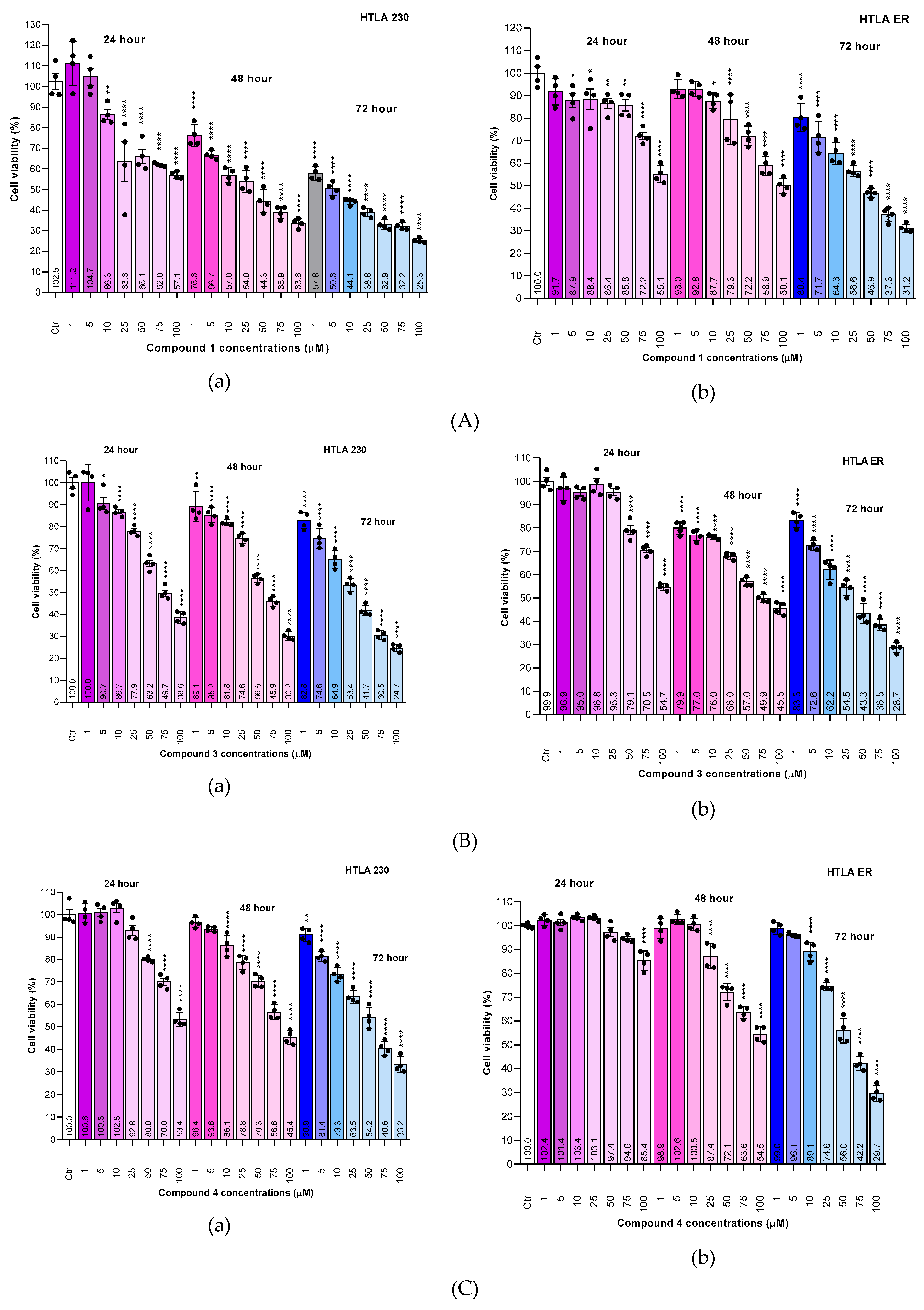
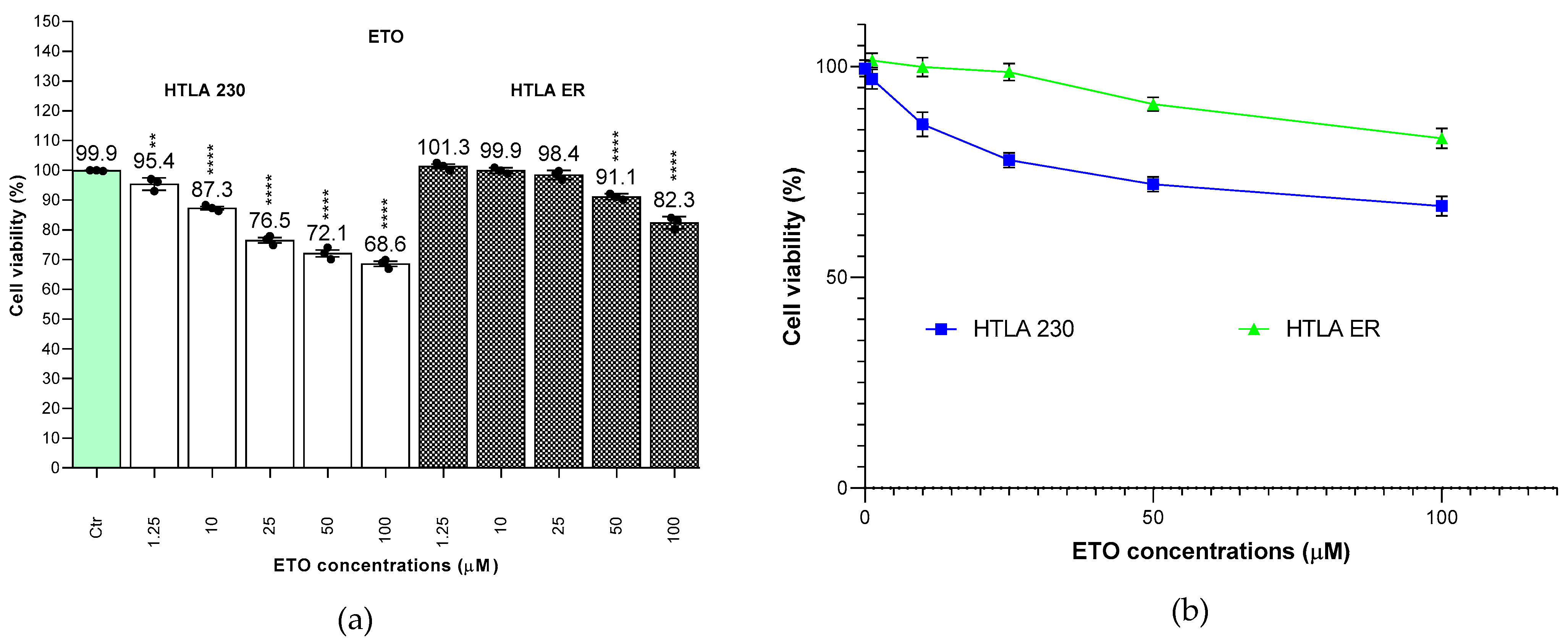
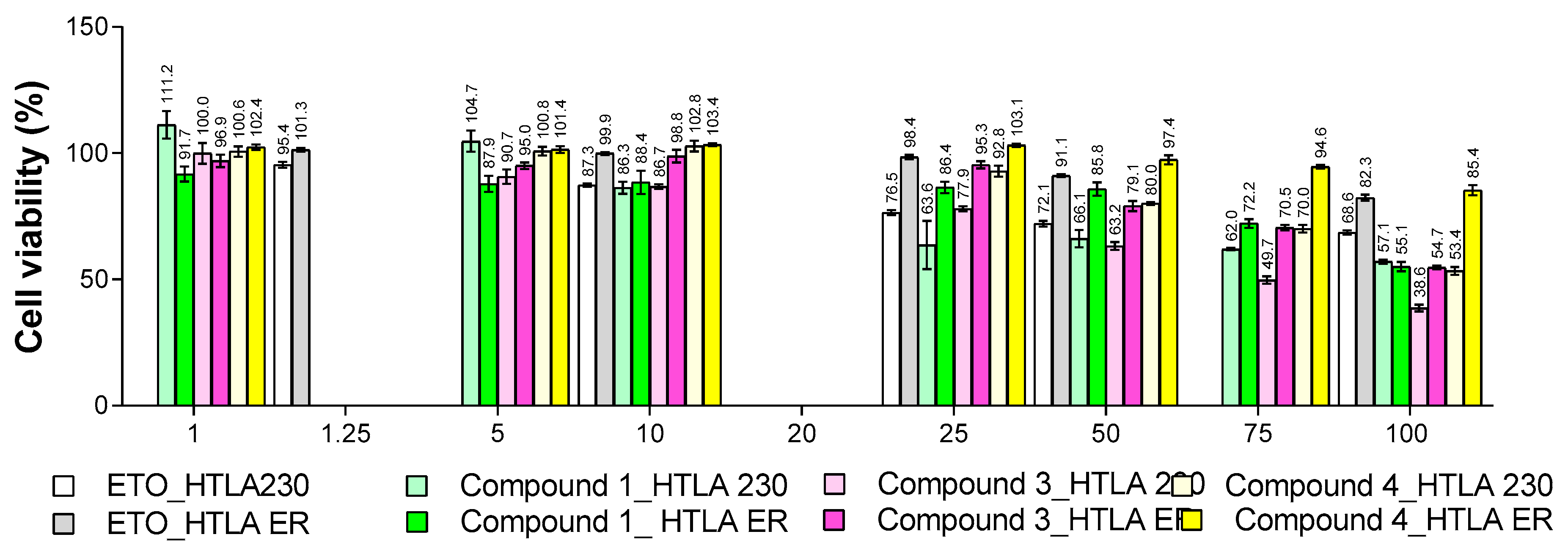
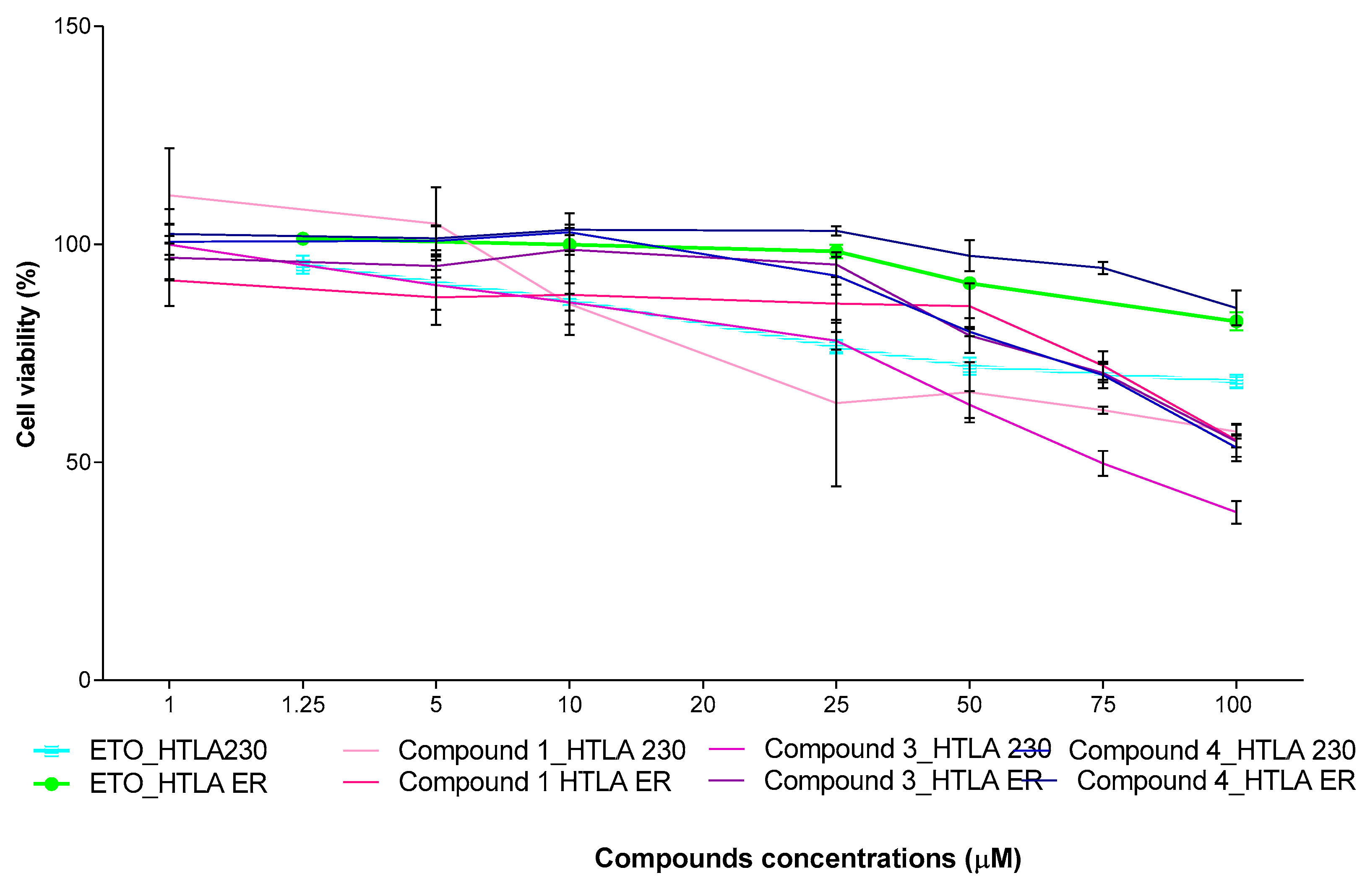
2.6.2. Concentration- and Time-Dependent Effects of 1, 3 and 4 on HTLA 230 and MDR HTLA ER NB Cell Viability
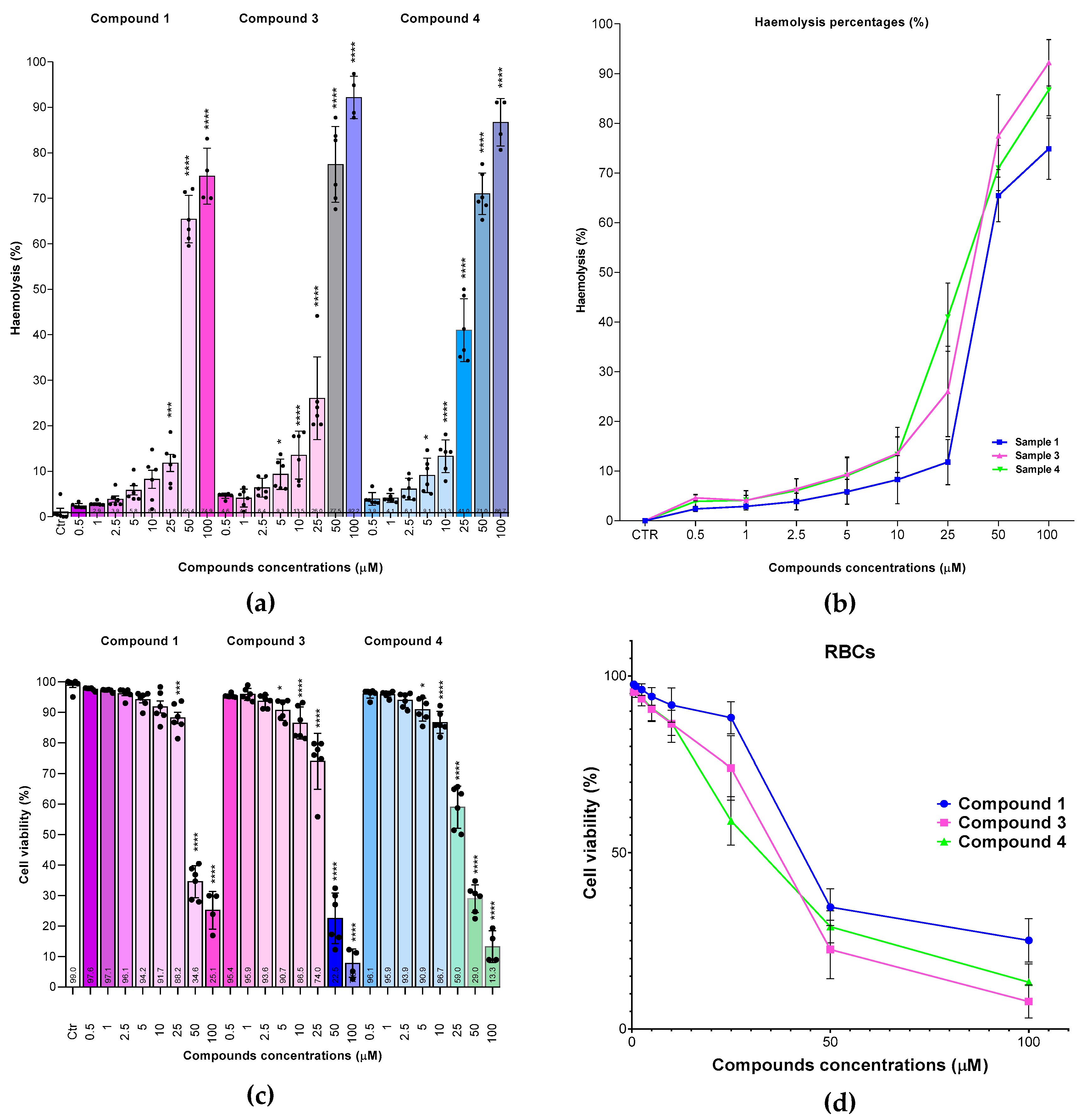
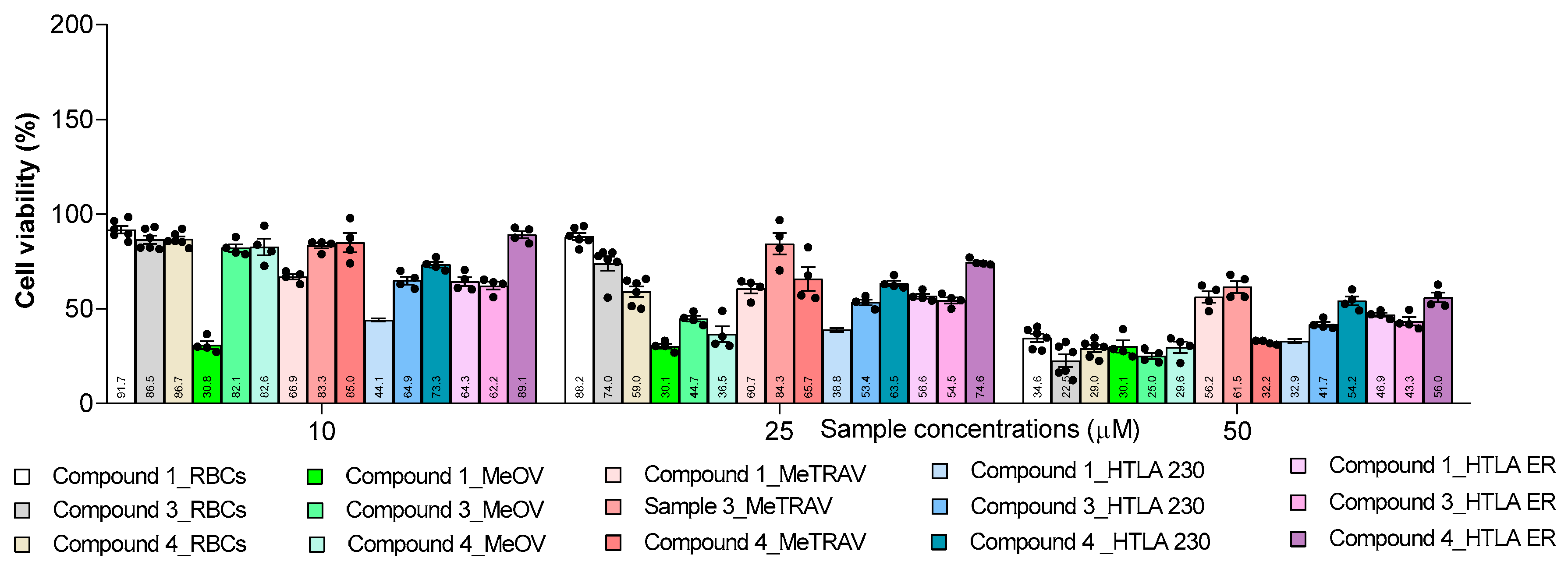
2.6.3. In vitro Haemolytic Toxicity of Compounds 1, 3 and 4 on Red Blood Cells (RBCs)
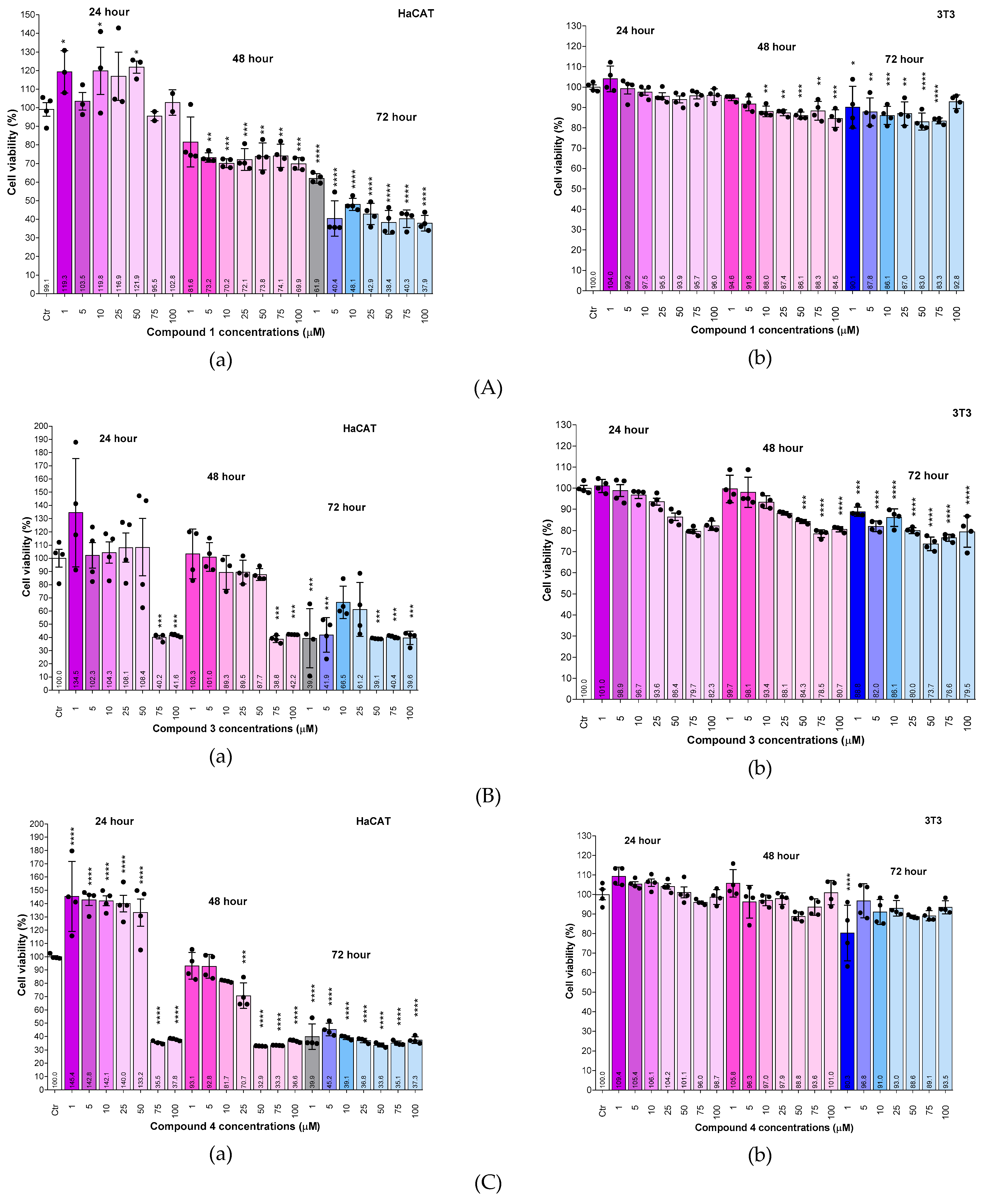
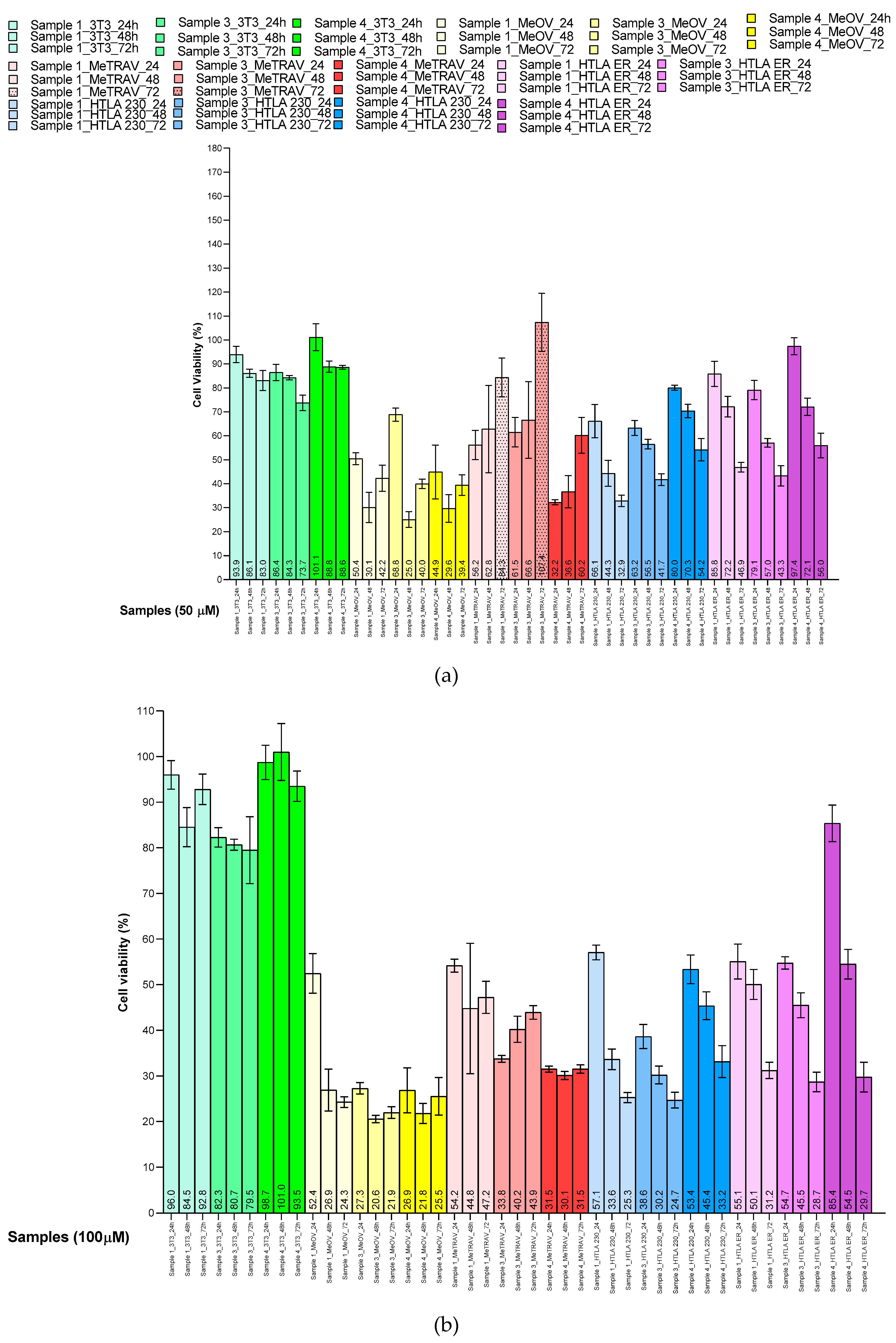
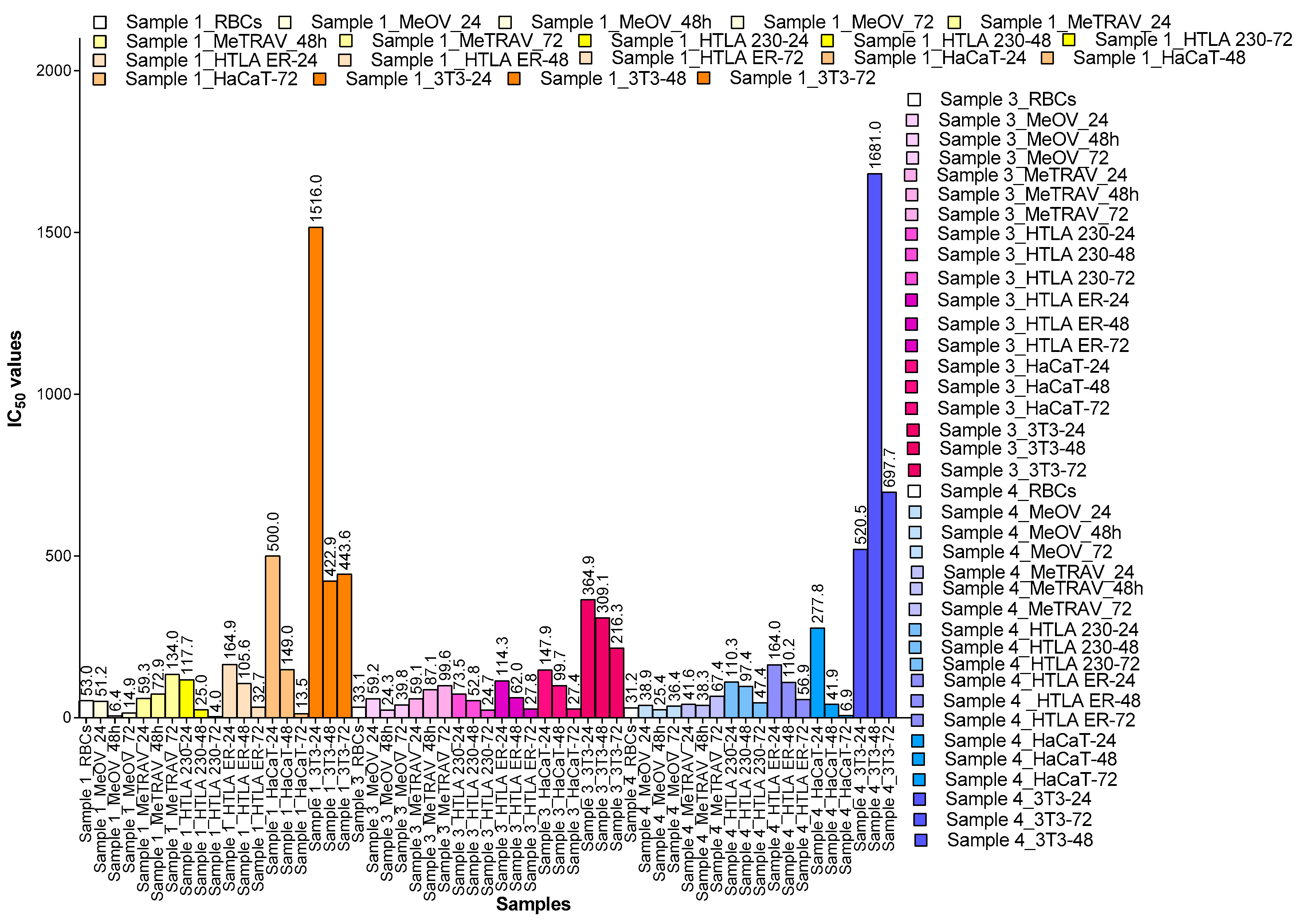
2.6.4. Concentration- and Time-Dependent Effects of samples 1, 3 and 4 on HaCaT and 3T3 Cell Viability
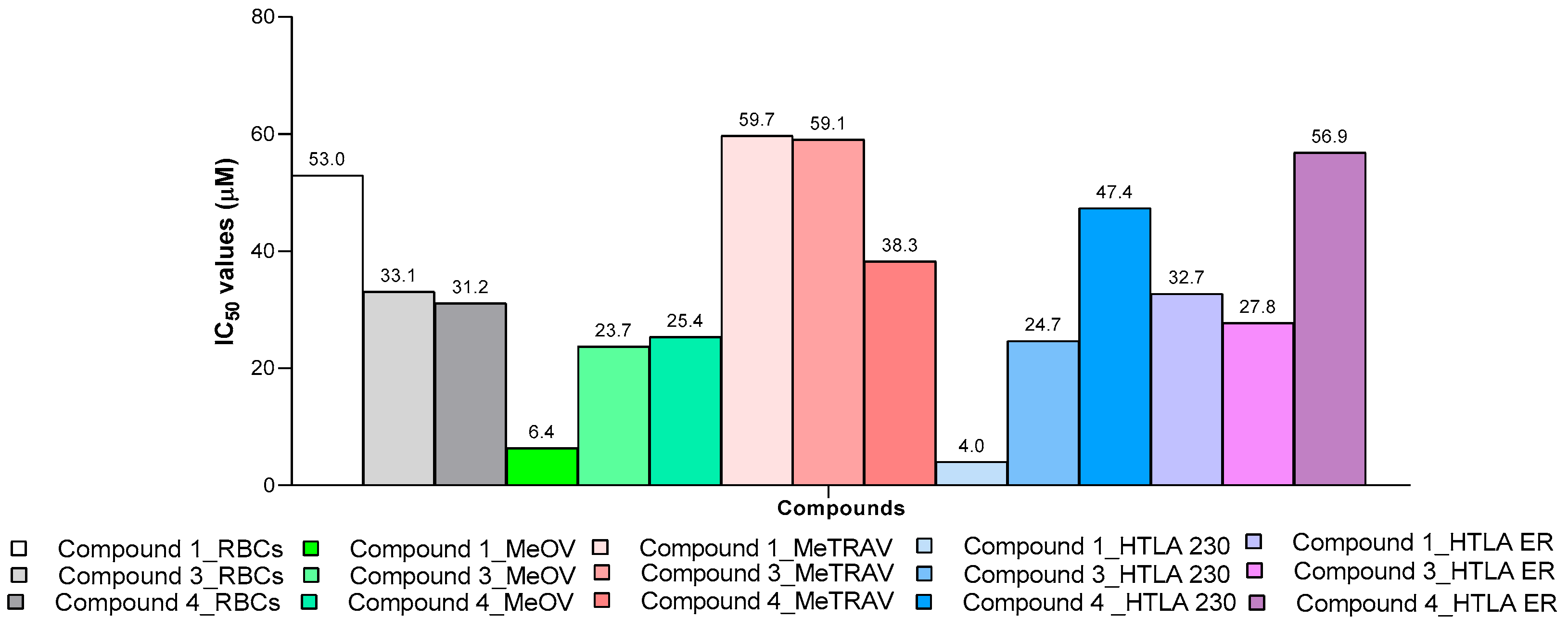
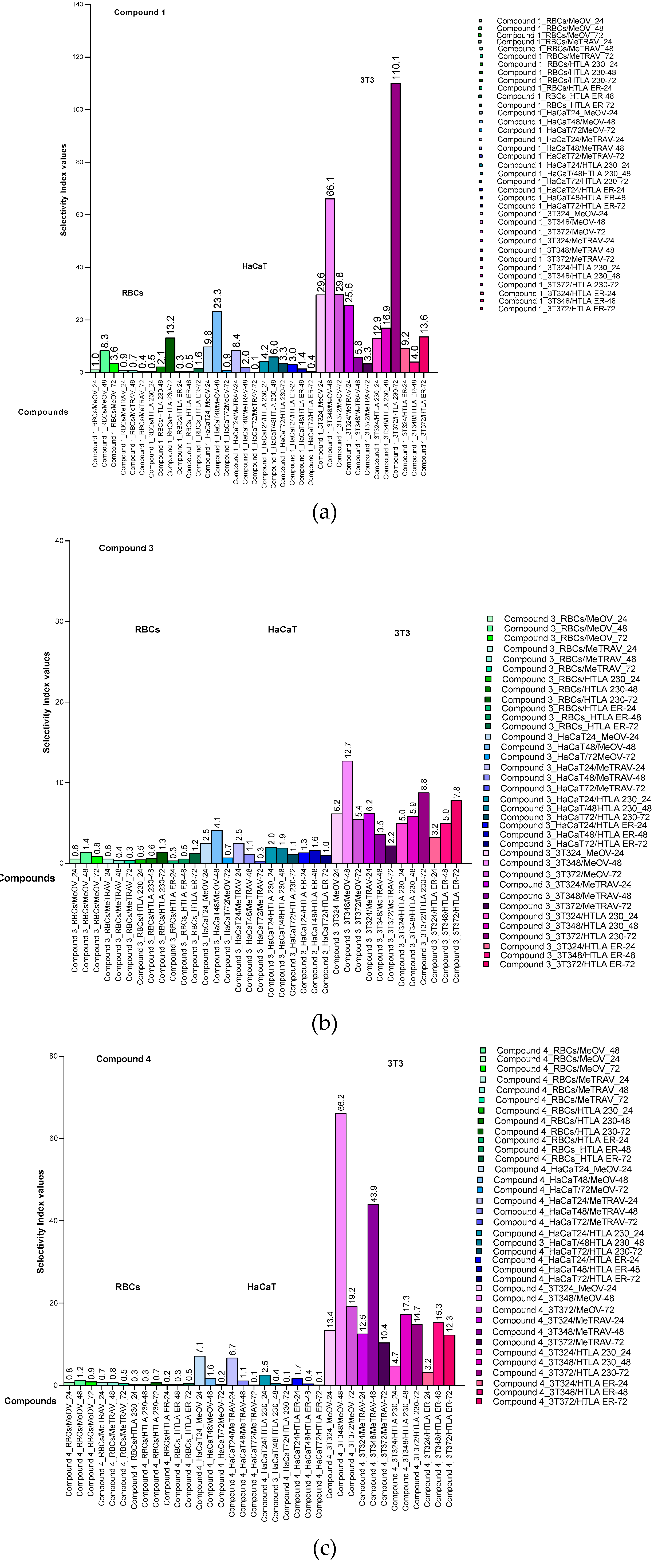
2.6.5. Selectivity Index
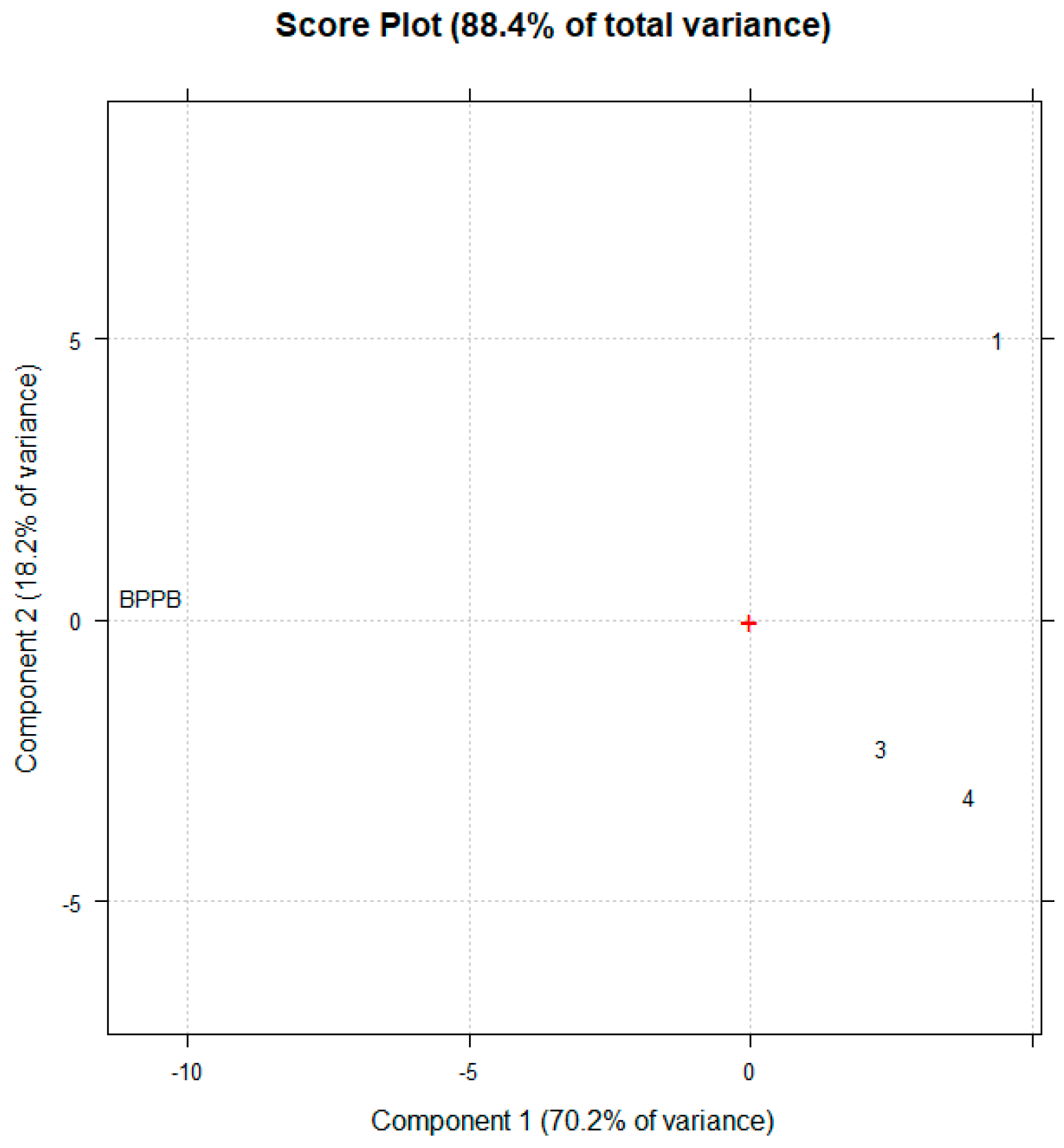
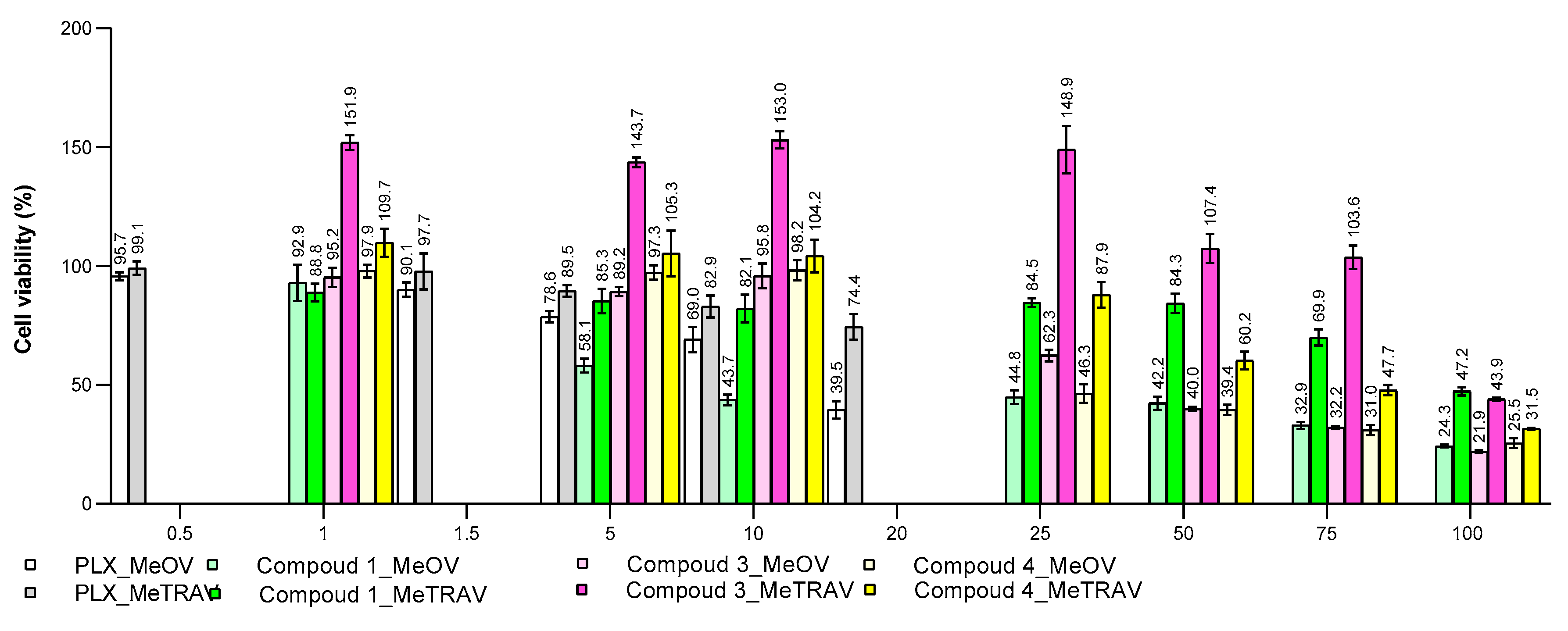
2.6.6. Comparison of 1, 3 and 4 Biological Effects with Those of BPPB Using PCA
3. Materials and Methods
3.1. Chemicals and Instruments
3.2. Synthesis of Phosphor-Containing Compounds 1-4
3.2.1. Benzyl triphenyl phosphonium bromide (1) [42]
3.2.2. 11-(Di-Phenyl-Phosphine)-1-Undecanol 2 [45]
3.2.3. Benzyl-(11-Hydroxyundecyl)-Diphenyl-Phosphonium Bromide (3) and Benzyl-diphenyl-undec-10-enyl-phosphonium bromide (4)
3.3. ATR-FTIR Spectroscopy of Compounds 1-4
3.4. UV-Vis Analyses
3.5. Multivariate Analysis of ATR-FTIR, UV-Vis and NMR Spectral Data
3.5.1. ATR-FTIR and UV-Vis Spectral Data
3.5.2. NMR Spectral Data
3.5.3. Biological Findings
3.6. Optical Microscopy Analyses
3.7. Cytotoxicity Experiments Using Compounds 1, 3 and 4 on PLX-resistant MCM (MeOV and MeTRAV) Cells, as Well as on ETO-sensitive (HTLA 230) and MDR (HTLA ER) NB Cells
3.7.1. Cell Lines and Culture Conditions
3.7.2. Treatments
3.7.3. Cell Viability Assay
3.8. In vitro Haemolytic Toxicity of Samples 1, 3 and 4, Using Red Blood Cells (RBCs)
3.9. Evaluation of Cytotoxicity of Samples 1, 3 and 4 on Human Keratinocytes (HaCaT) and Murine Fibroblasts (3T3)
3.9.1. Cell Culture
3.9.2. Treatments
3.9.3. Viability Assay
3.10. Comparison Between the Biological Effects of 1, 3 and 4 and Those of BPPB by PCA
3.11. Statistical Analyses
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Long, G. V; Swetter, S.M.; Menzies, A.M.; Gershenwald, J.E.; Scolyer, R.A. Cutaneous Melanoma. The Lancet 2023, 402, 485–502. [CrossRef]
- Drugs in Clinical Development for Melanoma. Pharmaceut. Med. 2012, 26, 171–183. [CrossRef]
- J Ferlay; M Ervik; FC Lam, et al Global Cancer Observatory: Cancer Today International Agency for Research on Cancer. Available online: https://gco.iarc.fr/today/en (accessed on 21 January 2025).
- Gandini, S.; Sera, F.; Cattaruzza, M.S.; Pasquini, P.; Abeni, D.; Boyle, P.; Melchi, C.F. Meta-Analysis of Risk Factors for Cutaneous Melanoma: I. Common and Atypical Naevi. Eur. J. Cancer 2005, 41, 28–44. [CrossRef]
- Bliss, J.M.; Ford, D.; Swerdlow, A.J.; Armstrong, B.K.; Cristofolini, M.; Elwood, J.M.; Green, A.; Holly, E.A.; Mack, T.; Mackie, R.M.; et al. Risk of Cutaneous Melanoma Associated with Pigmentation Characteristics and Freckling: Systematic Overview of 10 Case-control Studies. Int. J. Cancer 1995, 62, 367–376. [CrossRef]
- Davies, H.; Bignell, G.R.; Cox, C.; Stephens, P.; Edkins, S.; Clegg, S.; Teague, J.; Woffendin, H.; Garnett, M.J.; Bottomley, W.; et al. Mutations of the BRAF Gene in Human Cancer. Nature 2002, 417, 949–954. [CrossRef]
- Holderfield, M.; Deuker, M.M.; McCormick, F.; McMahon, M. Targeting RAF Kinases for Cancer Therapy: BRAF-Mutated Melanoma and Beyond. Nat. Rev. Cancer 2014, 14, 455–467. [CrossRef]
- Haq, R.; Fisher, D.E.; Widlund, H.R. Molecular Pathways: BRAF Induces Bioenergetic Adaptation by Attenuating Oxidative Phosphorylation. Clinical Cancer Research 2014, 20, 2257–2263. [CrossRef]
- Patel, B.; Patel, M.; Gami, B.; Patel, A. Cultivation of Bioenergy Crops in Gujarat State: A Consultative Survey Process to Understand the Current Practices of Landowners. Environ. Dev. Sustain. 2021, 23, 8991–9013. [CrossRef]
- Boutros, A.; Croce, E.; Ferrari, M.; Gili, R.; Massaro, G.; Marconcini, R.; Arecco, L.; Tanda, E.T.; Spagnolo, F. The Treatment of Advanced Melanoma: Current Approaches and New Challenges. Crit. Rev. Oncol. Hematol. 2024, 196, 104276. [CrossRef]
- Berk-Krauss, J.; Stein, J.A.; Weber, J.; Polsky, D.; Geller, A.C. New Systematic Therapies and Trends in Cutaneous Melanoma Deaths Among US Whites, 1986–2016. Am. J. Public Health 2020, 110, 731–733. [CrossRef]
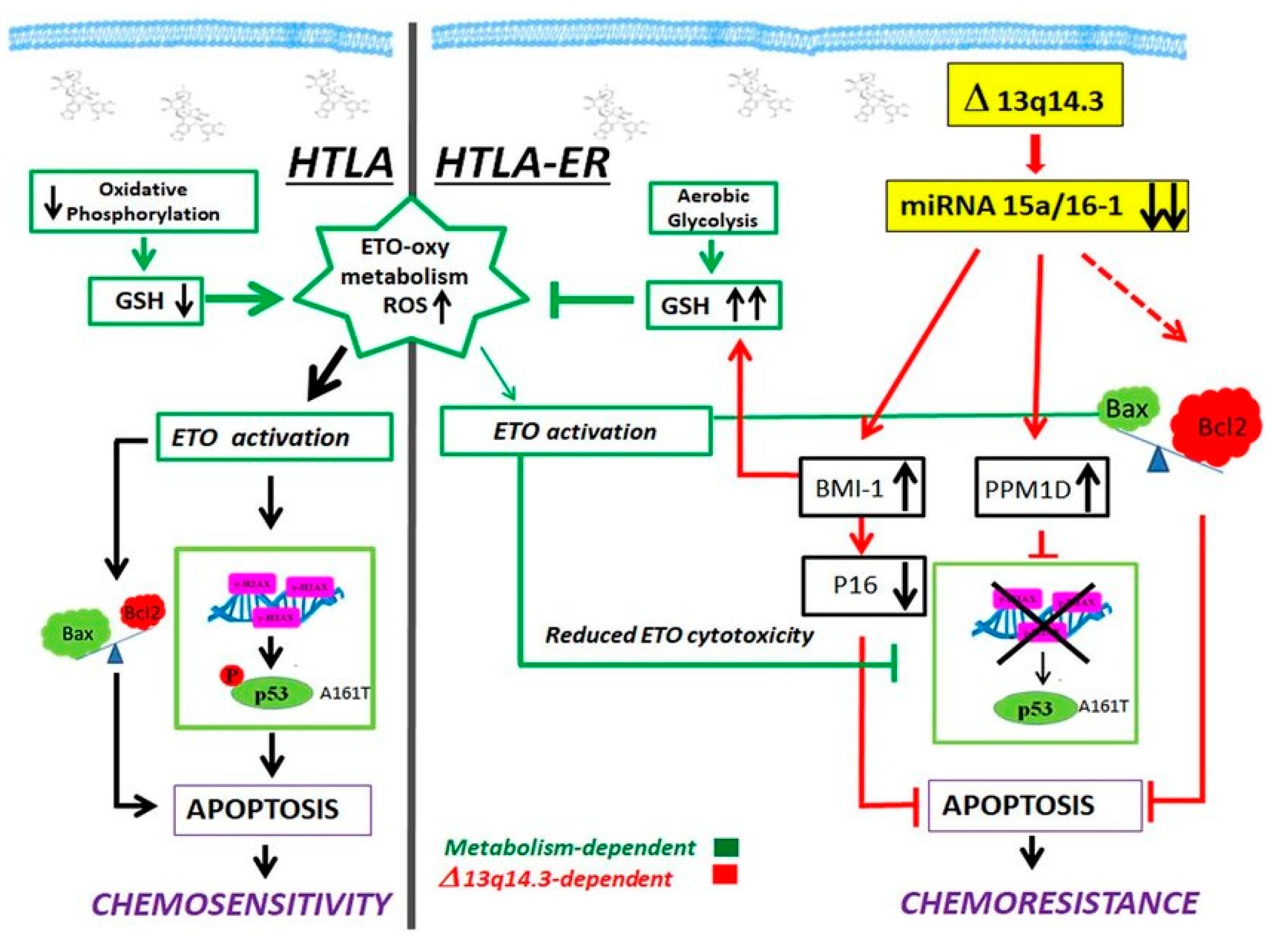
- Marengo, B.; Monti, P.; Miele, M.; Menichini, P.; Ottaggio, L.; Foggetti, G.; Pulliero, A.; Izzotti, A.; Speciale, A.; Garbarino, O.; et al. Etoposide-Resistance in a Neuroblastoma Model Cell Line Is Associated with 13q14.3 Mono-Allelic Deletion and MiRNA-15a/16-1 down-Regulation. Sci. Rep. 2018, 8. [CrossRef]
- Schwab, M.; Westermann, F.; Hero, B.; Berthold, F. Neuroblastoma: Biology and Molecular and Chromosomal Pathology. Lancet Oncol. 2003, 4, 472–480. [CrossRef]
- Cheung, N.-K. V.; Dyer, M.A. Neuroblastoma: Developmental Biology, Cancer Genomics and Immunotherapy. Nat. Rev. Cancer 2013, 13, 397–411. [CrossRef]
- Morandi, F.; Airoldi, I.; Marimpietri, D.; Bracci, C.; Faini, A.C.; Gramignoli, R. CD38, a Receptor with Multifunctional Activities: From Modulatory Functions on Regulatory Cell Subsets and Extracellular Vesicles, to a Target for Therapeutic Strategies. Cells 2019, 8, 1527. [CrossRef]
- Colla, R.; Izzotti, A.; De Ciucis, C.; Fenoglio, D.; Ravera, S.; Speciale, A.; Ricciarelli, R.; Furfaro, A.L.; Pulliero, A.; Passalacqua, M.; et al. Glutathione-Mediated Antioxidant Response and Aerobic Metabolism: Two Crucial Factors Involved in Determining the Multi-Drug Resistance of High-Risk Neuroblastoma. Oncotarget 2016, 7, 70715–70737. [CrossRef]
- Seeger, R.C.; Brodeur, G.M.; Sather, H.; Dalton, A.; Siegel, S.E.; Wong, K.Y.; Hammond, D. Association of Multiple Copies of the N- Myc Oncogene with Rapid Progression of Neuroblastomas. New England Journal of Medicine 1985, 313, 1111–1116. [CrossRef]
- Garbarino, O.; Valenti, G.E.; Monteleone, L.; Pietra, G.; Mingari, M.C.; Benzi, A.; Bruzzone, S.; Ravera, S.; Leardi, R.; Farinini, E.; et al. PLX4032 Resistance of Patient-Derived Melanoma Cells: Crucial Role of Oxidative Metabolism. Front. Oncol. 2023, 13. [CrossRef]
- Alfei, S.; Torazza, C.; Bacchetti, F.; Signorello, M.G.; Passalacqua, M.; Domenicotti, C.; Marengo, B. Tri-Phenyl-Phosphonium-Based Nano Vesicles: A New In Vitro Nanomolar-Active Weapon to Eradicate PLX-Resistant Melanoma Cells. Int. J. Mol. Sci. 2025, 26, 3227. [CrossRef]
- Alfei, S.; Torazza, C.; Bacchetti, F.; Milanese, M.; Passalaqua, M.; Khaledizadeh, E.; Vernazza, S.; Domenicotti, C.; Marengo, B. TPP-Based Nanovesicles Kill MDR Neuroblastoma Cells and Induce Moderate ROS Increase, While Exert Low Toxicity To-Wards Primary Cell Cultures: An in Vitro Study . IJMS 2025, 26, 4991. [CrossRef]
- Alfei, S.; Zuccari, G.; Athanassopoulos, C.M.; Domenicotti, C.; Marengo, B. Strongly ROS-Correlated, Time-Dependent, and Selective Antiproliferative Effects of Synthesized Nano Vesicles on BRAF Mutant Melanoma Cells and Their Hyaluronic Acid-Based Hydrogel Formulation. Int. J. Mol. Sci. 2024, 25, 10071. [CrossRef]
- Alfei, S.; Giannoni, P.; Signorello, M.G.; Torazza, C.; Zuccari, G.; Athanassopoulos, C.M.; Domenicotti, C.; Marengo, B. The Remarkable and Selective In Vitro Cytotoxicity of Synthesized Bola-Amphiphilic Nanovesicles on Etoposide-Sensitive and -Resistant Neuroblastoma Cells. Nanomaterials 2024, 14, 1505. [CrossRef]
- Alfei, S.; Giannoni, P.; Signorello, M.G.; Torazza, C.; Zuccari, G.; Athanassopoulos, C.M.; Domenicotti, C.; Marengo, B. The Remarkable and Selective In Vitro Cytotoxicity of Synthesized Bola-Amphiphilic Nanovesicles on Etoposide-Sensitive and -Resistant Neuroblastoma Cells. Nanomaterials 2024, 14, 1505. [CrossRef]
- Nuraje, N.; Bai, H.; Su, K. Bolaamphiphilic Molecules: Assembly and Applications. Prog. Polym. Sci. 2013, 38, 302–343. [CrossRef]
- Dhanya, D.; Giuseppe, P.; Rita, C.A.; Annaluisa, M.; Stefania, S.M.; Francesca, G.; Vitale, D. V.; Anna, R.; Claudio, A.; Pasquale, L.; et al. Phosphonium Salt Displays Cytotoxic Effects Against Human Cancer Cell Lines. Anticancer Agents Med. Chem. 2018, 17. [CrossRef]
- Lukáč, M.; Pisárčik, M.; Garajová, M.; Mrva, M.; Dušeková, A.; Vrták, A.; Horáková, R.; Horváth, B.; Devínsky, F. Synthesis, Surface Activity, and Biological Activities of Phosphonium and Metronidazole Salts. J. Surfactants Deterg. 2020, 23, 1025–1032. [CrossRef]
- Bachowska, B.; Kazmierczak-Baranska, J.; Cieslak, M.; Nawrot, B.; Szczęsna, D.; Skalik, J.; Bałczewski, P. High Cytotoxic Activity of Phosphonium Salts and Their Complementary Selectivity towards HeLa and K562 Cancer Cells: Identification of Tri- n -butyl- n -hexadecylphosphonium Bromide as a Highly Potent Anti-HeLa Phosphonium Salt. ChemistryOpen 2012, 1, 33–38. [CrossRef]
- Terekhova, N. V.; Tatarinov, D.A.; Shaihutdinova, Z.M.; Pashirova, T.N.; Lyubina, A.P.; Voloshina, A.D.; Sapunova, A.S.; Zakharova, L.Ya.; Mironov, V.F. Design and Synthesis of Amphiphilic 2-Hydroxybenzylphosphonium Salts with Antimicrobial and Antitumor Dual Action. Bioorg. Med. Chem. Lett. 2020, 30, 127234. [CrossRef]
- Bacchetti, F.; Schito, A.M.; Milanese, M.; Castellaro, S.; Alfei, S. Anti Gram-Positive Bacteria Activity of Synthetic Quaternary Ammonium Lipid and Its Precursor Phosphonium Salt. Int. J. Mol. Sci. 2024, 25, 2761. [CrossRef]
- Alfei, S.; Zuccari, G.; Bacchetti, F.; Torazza, C.; Milanese, M.; Siciliano, C.; Athanassopoulos, C.M.; Piatti, G.; Schito, A.M. Synthesized Bis-Triphenyl Phosphonium-Based Nano Vesicles Have Potent and Selective Antibacterial Effects on Several Clinically Relevant Superbugs. Nanomaterials 2024, 14, 1351. [CrossRef]
- Jędrzejczyk, M.; Sulik, M.; Mielczarek-Puta, M.; Lim, G.Y.; Podsiad, M.; Hoser, J.; Bednarczyk, P.; Struga, M.; Huczyński, A. Anticancer Activity of Salinomycin Quaternary Phosphonium Salts. Eur. J. Med. Chem. 2025, 282, 117055. [CrossRef]
- Jeena, M.T.; Kim, S.; Jin, S.; Ryu, J.-H. Recent Progress in Mitochondria-Targeted Drug and Drug-Free Agents for Cancer Therapy. Cancers (Basel). 2019, 12, 4. [CrossRef]
- Liu, Y.; Sun, Y.; Guo, Y.; Shi, X.; Chen, X.; Feng, W.; Wu, L.-L.; Zhang, J.; Yu, S.; Wang, Y.; et al. An Overview: The Diversified Role of Mitochondria in Cancer Metabolism. Int. J. Biol. Sci. 2023, 19, 897–915. [CrossRef]
- Borges, D.G.F.; Carvalho, D.S.; Bomfim, G.C.; Ramos, P.I.P.; Brzozowski, J.; Góes-Neto, A.; F. S. Andrade, R.; El-Hani, C. On the Origin of Mitochondria: A Multilayer Network Approach. PeerJ 2023, 11, e14571. [CrossRef]
- Ermolaev, V. V.; Arkhipova, D.M.; Miluykov, V.A.; Lyubina, A.P.; Amerhanova, S.K.; Kulik, N. V.; Voloshina, A.D.; Ananikov, V.P. Sterically Hindered Quaternary Phosphonium Salts (QPSs): Antimicrobial Activity and Hemolytic and Cytotoxic Properties. Int. J. Mol. Sci. 2021, 23, 86. [CrossRef]
- Nunes, B.; Cagide, F.; Borges, F.; Simões, M. Antimicrobial Activity and Cytotoxicity of Novel Quaternary Ammonium and Phosphonium Salts. J. Mol. Liq. 2024, 401, 124616. [CrossRef]
- Kodjo Amengor, C.D.; Amaning Danquah, C.; Adusei, E.B.A.; Kekessie, F.K.; Ofosu-Koranteng, F.; Peprah, P.; Harley, B.K.; Orman, E.; Adu, J.; Saaka, Y. Synthesized Phosphonium Compounds Demonstrate Resistant Modulatory and Antibiofilm Formation Activities against Some Pathogenic Bacteria. Heteroatom Chemistry 2022, 2022, 1–9. [CrossRef]
- Terekhova, N. V.; Tatarinov, D.A.; Shaihutdinova, Z.M.; Pashirova, T.N.; Lyubina, A.P.; Voloshina, A.D.; Sapunova, A.S.; Zakharova, L.Ya.; Mironov, V.F. Design and Synthesis of Amphiphilic 2-Hydroxybenzylphosphonium Salts with Antimicrobial and Antitumor Dual Action. Bioorg. Med. Chem. Lett. 2020, 30, 127234. [CrossRef]
- Galkina, I.; Bakhtiyarova, Y.; Andriyashin, V.; Galkin, V.; Cherkasov, R. Synthesis and Antimicrobial Activities of Phosphonium Salts on Basis of Triphenylphosphine and 3,5-Di-Tert-Butyl-4-Hydroxybenzyl Bromide. Phosphorus Sulfur Silicon Relat. Elem. 2013, 188, 15–18. [CrossRef]
- Milenković, M.R.; Živković-Radovanović, V.; Andjelković, L. Synthesis and Antimicrobial Activity of (3-Formyl-4-Hydroxybenzyl)Triphenylphosphonium Chloride Acylhydrazones. Russ. J. Gen. Chem. 2020, 90, 1716–1720. [CrossRef]
- Valenti, G.E.; Alfei, S.; Caviglia, D.; Domenicotti, C.; Marengo, B. Antimicrobial Peptides and Cationic Nanoparticles: A Broad-Spectrum Weapon to Fight Multi-Drug Resistance Not Only in Bacteria. Int. J. Mol. Sci. 2022, 23, 6108. [CrossRef]
- Cui, M.; Li, Z.; Tang, R.; Jia, H.; Liu, B. Novel (E)-5-Styryl-2,2′-Bithiophene Derivatives as Ligands for β-Amyloid Plaques. Eur. J. Med. Chem. 2011, 46, 2908–2916. [CrossRef]
- Cabiddu, S.; Maccioni, A.; Secci, M. Application of the Wittig Syntheses. III. Electronic Effects of Substituents and Stereochemistry of the Reaction. Annali di Chimica (Rome, Italy) 1964, 54, 1153–1164.
- Improvements in the Production of Quaternary Phosphonium Halides 1959.
- Hands, A.R.; Mercer, A.J.H. Reactions of Sodium Hydride with ω-Hydroxyalkyl-Phosphonium, -Arsonium and -Ammonium Salts. J. Chem. Soc. C 1968, 0, 1331–1337. [CrossRef]
- Petrucci, M.G.L.; Kakkar, A.K. Heterogenizing Homogeneous Catalysis Using Molecular Self-Assembly of Long Alkane Chain Phosphines Bound to Rh(I) Complexes. Chemistry of Materials 1999, 11, 269–276. [CrossRef]
- Ammer, J.; Nolte, C.; Karaghiosoff, K.; Thallmair, S.; Mayer, P.; Devivie-Riedle, R.; Mayr, H. Ion-Pairing of Phosphonium Salts in Solution: C-H…halogen and C-H…π Hydrogen Bonds. Chemistry - A European Journal 2013, 19. [CrossRef]
- Arunan, E.; Desiraju, G.R.; Klein, R.A.; Sadlej, J.; Scheiner, S.; Alkorta, I.; Clary, D.C.; Crabtree, R.H.; Dannenberg, J.J.; Hobza, P.; et al. Definition of the Hydrogen Bond (IUPAC Recommendations 2011). Pure and Applied Chemistry 2011, 83, 1637–1641. [CrossRef]
- Daasch, L.; Smith, D. Infrared Spectra of Phosphorus Compounds. Anal. Chem. 1951, 23, 853–868. [CrossRef]
- Bergel’son, L.D.; Solodovik, V.D.; Shemyakin, M.M. Stereoregulated Synthesis of Unsaturated Compounds Communication 9. Stereochemistry of the Reactions of Aldehydes with ?,?-Unsaturated Triphenylphosphonium Ylides [Alkylidenetriphenylphosphoranes]. Bulletin of the Academy of Sciences, USSR Division of Chemical Science 1966, 15, 468–473. [CrossRef]
- Ceccacci, F.; Sennato, S.; Rossi, E.; Proroga, R.; Sarti, S.; Diociaiuti, M.; Casciardi, S.; Mussi, V.; Ciogli, A.; Bordi, F.; et al. Aggregation Behaviour of Triphenylphosphonium Bolaamphiphiles. J. Colloid Interface Sci. 2018, 531, 451–462. [CrossRef]
- Alfei, S.; Brullo, C.; Caviglia, D.; Zuccari, G. Preparation and Physicochemical Characterization of Water-Soluble Pyrazole-Based Nanoparticles by Dendrimer Encapsulation of an Insoluble Bioactive Pyrazole Derivative. Nanomaterials 2021, 11, 2662. [CrossRef]
- Alfei, S.; Schito, A.M.; Zuccari, G. Considerable Improvement of Ursolic Acid Water Solubility by Its Encapsulation in Dendrimer Nanoparticles: Design, Synthesis and Physicochemical Characterization. Nanomaterials 2021, 11, 2196. [CrossRef]
- Graikioti, D.; Athanassopoulos, C.M.; Schito, A.M.; Alfei, S. Synthesis and Characterization of Triphenyl Phosphonium-Modified Triterpenoids with Never Reported Antibacterial Effects Against Clinically Relevant Gram-Positive Superbugs 2025.
- Alfei, S.; Oliveri, P.; Malegori, C. Assessment of the Efficiency of a Nanospherical Gallic Acid Dendrimer for Long-Term Preservation of Essential Oils: An Integrated Chemometric-Assisted FTIR Study. ChemistrySelect 2019, 4, 8891–8901. [CrossRef]
- Erdei, E.; Torres, S.M. A New Understanding in the Epidemiology of Melanoma. Expert Rev. Anticancer Ther. 2010, 10, 1811–1823. [CrossRef]
- Leary, M.; Heerboth, S.; Lapinska, K.; Sarkar, S. Sensitization of Drug Resistant Cancer Cells: A Matter of Combination Therapy. Cancers (Basel). 2018, 10, 483. [CrossRef]
- Bollag, G.; Hirth, P.; Tsai, J.; Zhang, J.; Ibrahim, P.N.; Cho, H.; Spevak, W.; Zhang, C.; Zhang, Y.; Habets, G.; et al. Clinical Efficacy of a RAF Inhibitor Needs Broad Target Blockade in BRAF-Mutant Melanoma. Nature 2010, 467, 596–599. [CrossRef]
- Hallmeyer, S.; Gonzalez, R.; Lawson, D.H.; Cranmer, L.D.; Linette, G.P.; Puzanov, I.; Taback, B.; Cowey, C.L.; Ribas, A.; Daniels, G.A.; et al. Vemurafenib Treatment for Patients with Locally Advanced, Unresectable Stage IIIC or Metastatic Melanoma and Activating Exon 15 BRAF Mutations Other than V600E. Melanoma Res. 2017, 27, 585–590. [CrossRef]
- Joseph, R.; Swaika, A.; Crozier, J.A. Vemurafenib: An Evidence-Based Review of Its Clinical Utility in the Treatment of Metastatic Melanoma. Drug Des. Devel. Ther. 2014, 775. [CrossRef]
- Tanda, E.T.; Vanni, I.; Boutros, A.; Andreotti, V.; Bruno, W.; Ghiorzo, P.; Spagnolo, F. Current State of Target Treatment in BRAF Mutated Melanoma. Front. Mol. Biosci. 2020, 7. [CrossRef]
- Li, K.; Xiao, G.; Richardson, J.J.; Tardy, B.L.; Ejima, H.; Huang, W.; Guo, J.; Liao, X.; Shi, B. Targeted Therapy against Metastatic Melanoma Based on Self-Assembled Metal-Phenolic Nanocomplexes Comprised of Green Tea Catechin. Advanced Science 2019, 6. [CrossRef]
- Chapman, P.B.; Robert, C.; Larkin, J.; Haanen, J.B.; Ribas, A.; Hogg, D.; Hamid, O.; Ascierto, P.A.; Testori, A.; Lorigan, P.C.; et al. Vemurafenib in Patients with BRAFV600 Mutation-Positive Metastatic Melanoma: Final Overall Survival Results of the Randomized BRIM-3 Study. Annals of Oncology 2017, 28, 2581–2587. [CrossRef]
- Chandrasekhar, B.; Gor, R.; Ramalingam, S.; Thiagarajan, A.; Sohn, H.; Madhavan, T. Repurposing FDA-Approved Compounds to Target JAK2 for Colon Cancer Treatment. Discover Oncology 2024, 15, 226. [CrossRef]
- Di Raimondo, C.; Lozzi, F.; Di Domenico, P.P.; Campione, E.; Bianchi, L. The Diagnosis and Management of Cutaneous Metastases from Melanoma. Int. J. Mol. Sci. 2023, 24, 14535. [CrossRef]
- Boukamp, P.; Petrussevska, R.T.; Breitkreutz, D.; Hornung, J.; Markham, A.; Fusenig, N.E. Normal Keratinization in a Spontaneously Immortalized Aneuploid Human Keratinocyte Cell Line. J. Cell Biol. 1988, 106, 761–771. [CrossRef]
- Ölschläger, V.; Schrader, A.; Hockertz, S. Comparison of Primary Human Fibroblasts and Keratinocytes with Immortalized Cell Lines Regarding Their Sensitivity to Sodium Dodecyl Sulfate in a Neutral Red Uptake Cytotoxicity Assay. Arzneimittelforschung 2011, 59, 146–152. [CrossRef]
- Schoop, V.M.; Fusenig, N.E.; Mirancea, N. Epidermal Organization and Differentiation of HaCaT Keratinocytes in Organotypic Coculture with Human Dermal Fibroblasts. Journal of Investigative Dermatology 1999, 112, 343–353. [CrossRef]
- Mukherjee, S.P.; Davoren, M.; Byrne, H.J. In Vitro Mammalian Cytotoxicological Study of PAMAM Dendrimers – Towards Quantitative Structure Activity Relationships. Toxicology in Vitro 2010, 24, 169–177. [CrossRef]
- Zhang, K.; Na, T.; Wang, L.; Gao, Q.; Yin, W.; Wang, J.; Yuan, B.-Z. Human Diploid MRC-5 Cells Exhibit Several Critical Properties of Human Umbilical Cord-Derived Mesenchymal Stem Cells. Vaccine 2014, 32, 6820–6827. [CrossRef]
- Alfei, S.; Caviglia, D.; Piatti, G.; Zuccari, G.; Schito, A.M. Synthesis, Characterization and Broad-Spectrum Bactericidal Effects of Ammonium Methyl and Ammonium Ethyl Styrene-Based Nanoparticles. Nanomaterials 2022, 12, 2743. [CrossRef]
- Alfei, S.; Marengo, B.; Domenicotti, C. Polyester-Based Dendrimer Nanoparticles Combined with Etoposide Have an Improved Cytotoxic and Pro-Oxidant Effect on Human Neuroblastoma Cells. Antioxidants 2020, 9, 50. [CrossRef]
- Alfei, S.; Marengo, B.; Zuccari, G.; Turrini, F.; Domenicotti, C. Dendrimer Nanodevices and Gallic Acid as Novel Strategies to Fight Chemoresistance in Neuroblastoma Cells. Nanomaterials 2020, 10, 1243. [CrossRef]
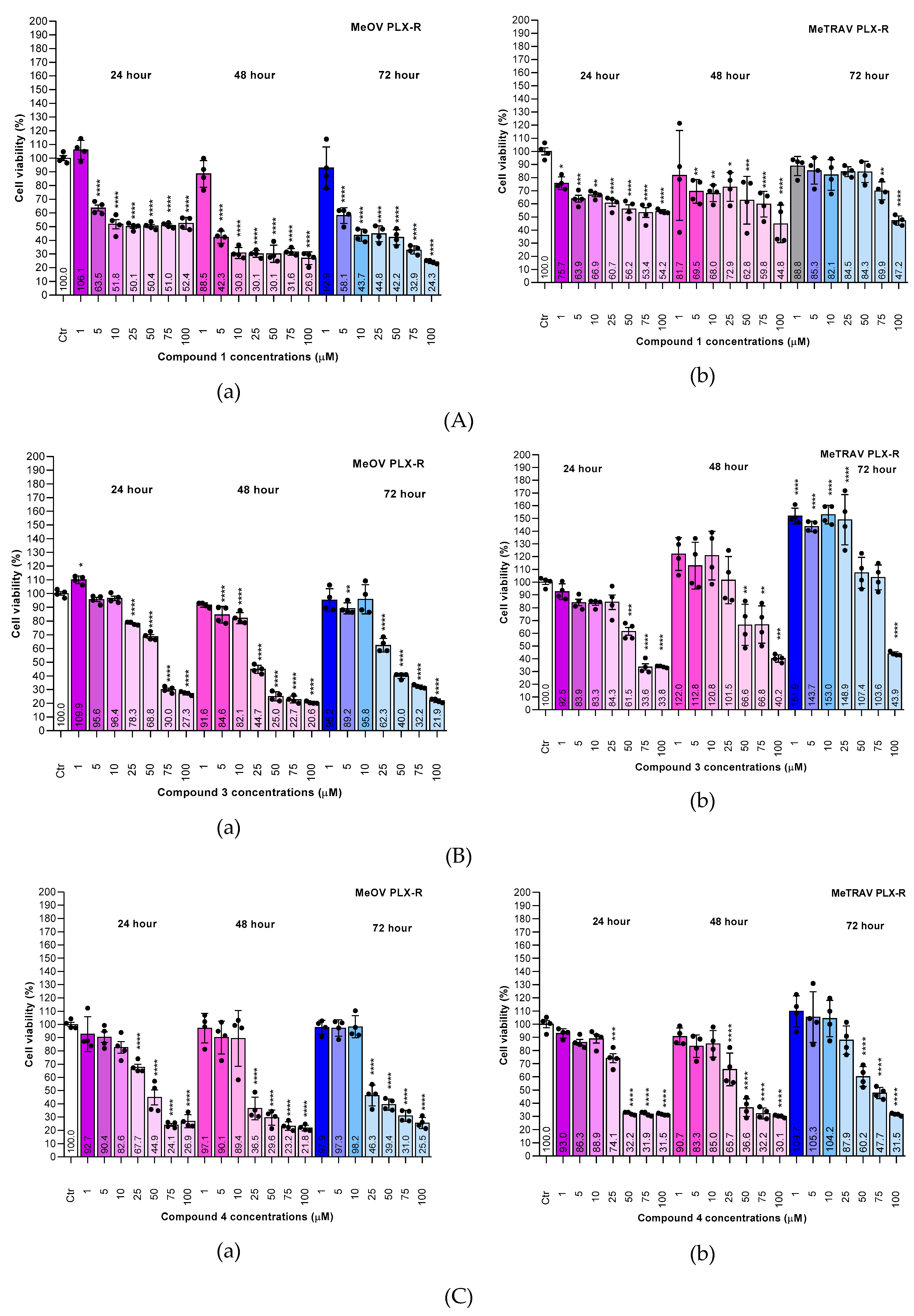
| Compound | Exposure time (hours) | IC50 MeOV PLX-R (µM) | IC50 MeTRAV PLX-R (µM) |
| 1 | 24 | 51.23 ± 42.33 * | 59.72 ± 27.23 * |
| 48 | 6.37 ± 3.02 | 72.93 ± 37.22 * | |
| 72 | 14.89 ± 5.25 | 134.0 ± 40.25 * | |
| 3 | 24 | 62.05 ± 14.00 * | 59.06 ± 11.03 * |
| 48 | 23.73 ± 3.22 | 87.11 ± 27.62 * | |
| 72 | 39.45 ± 6.71 | 99.64 ± V.W. * | |
| 4 | 24 | 38.86 ± 7.13 | 41.62 ± 6.24 |
| 48 | 25.40 ± 6.90 | 38.26 ± 7.00 | |
| 72 | 36.35 ± 8.13 | 67.40 ± 10.16 * | |
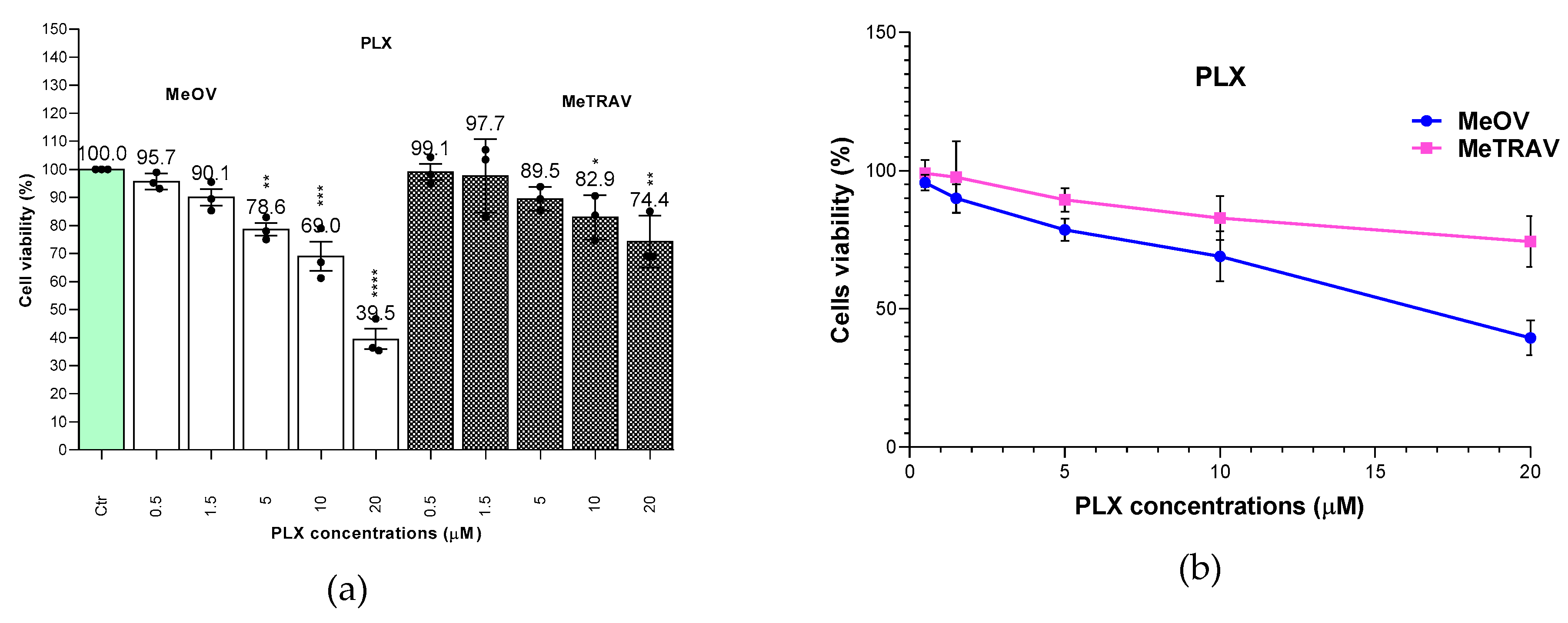
| Vemurafenib (PLX) | 72 | 16.86 ± 3.48 | 53.43 ± 20.91 * |
| Compound | Exposure time (hours) | IC50 HTLA 230 (µM) | IC50 MDR HATLA ER (µM) |
| 1 | 24 | 117.70 ± 71.64 * | 164.90 ± 38.30 * |
| 48 | 25.02 ± 4.73 | 105.60 ± 14.58 * | |
| 72 | 4.03 ± 0.91 | 32.71 ± 8.79 | |
| 3 | 24 | 73.47 ± 7.89 * | 114.30 ± 12.50 * |
| 48 | 57.92 ± 8.95 * | 62.03 ± 15.24 * | |
| 72 | 24.67 ± 3.29 | 27.76 ± 3.56 | |
| 4 | 24 | 110.30 ± 9.95 * | 164.00 ± 56.45 * |
| 48 | 97.40 ± 12.72 * | 110.20 ± 12.20 * | |
| 72 | 47.36 ± 5.81 | 56.88 ± 3.09 * | |
| ETO | 24 | 145.60 ± 35.40 * | 531.20 ± 147.35 * |
| Cells | Sample 1 | Sample 3 | Sample 4 |
| RBCs ° | 52.97±10.93 * | 33.12±6.91 | 31.15±4.13 |
| MeOV 24 h | 51.23 ± 42.33 * | 62.05 ± 14.00 * | 38.86 ± 7.13 |
| MeTRAV 24h | 59.72 ± 27.23 * | 59.06 ± 11.03 * | 41.62 ± 6.24 |
| HTLA 230 24h | 117.70±71.64 * | 73.47±7.98 * | 110.30±9.55 * |
| HTLA ER 24 h | 164.90±38.30 * | 114.30±12.50 * | 164.00±56.45 * |
| MeOV 48 h | 6.37 ± 3.02 | 23.73 ± 3.22 | 25.40 ± 6.90 |
| MeTRAV 48h | 72.93 ± 37.22 * | 87.11 ± 27.62 * | 38.26 ± 7.00 |
| HTLA 230 48h | 25.02±4.73 | 57.92±8.95 * | 97.40±12.72 * |
| HTLA ER 48h | 105.60±15.58 * | 62.03±15.24 * | 110.20±12.10 * |
| MeOV 72h | 14.89 ± 5.25 | 39.45 ± 6.71 | 36.35 ± 8.13 |
| MeTRAV 72h | 134.0 ± 40.25 * | 99.64 ± V.W. * | 67.40 ± 10.16 * |
| HTLA 230 72h | 4.03. ± 0.91 | 24.67±3.29 | 46.36±5.81 |
| HTLA ER 72h | 32.71±8.79 | 27.26±3.56 | 56.88±3.09 * |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.





