Submitted:
09 February 2026
Posted:
10 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Phenotypic Evaluation
2.2. DNA Preparation and Sequencing
2.3. SNP Calling and Annotation
2.4. Genetic structure Analysis
2.5. GWAS Analysis
2.6. Candidate Gene Screening
3. Results
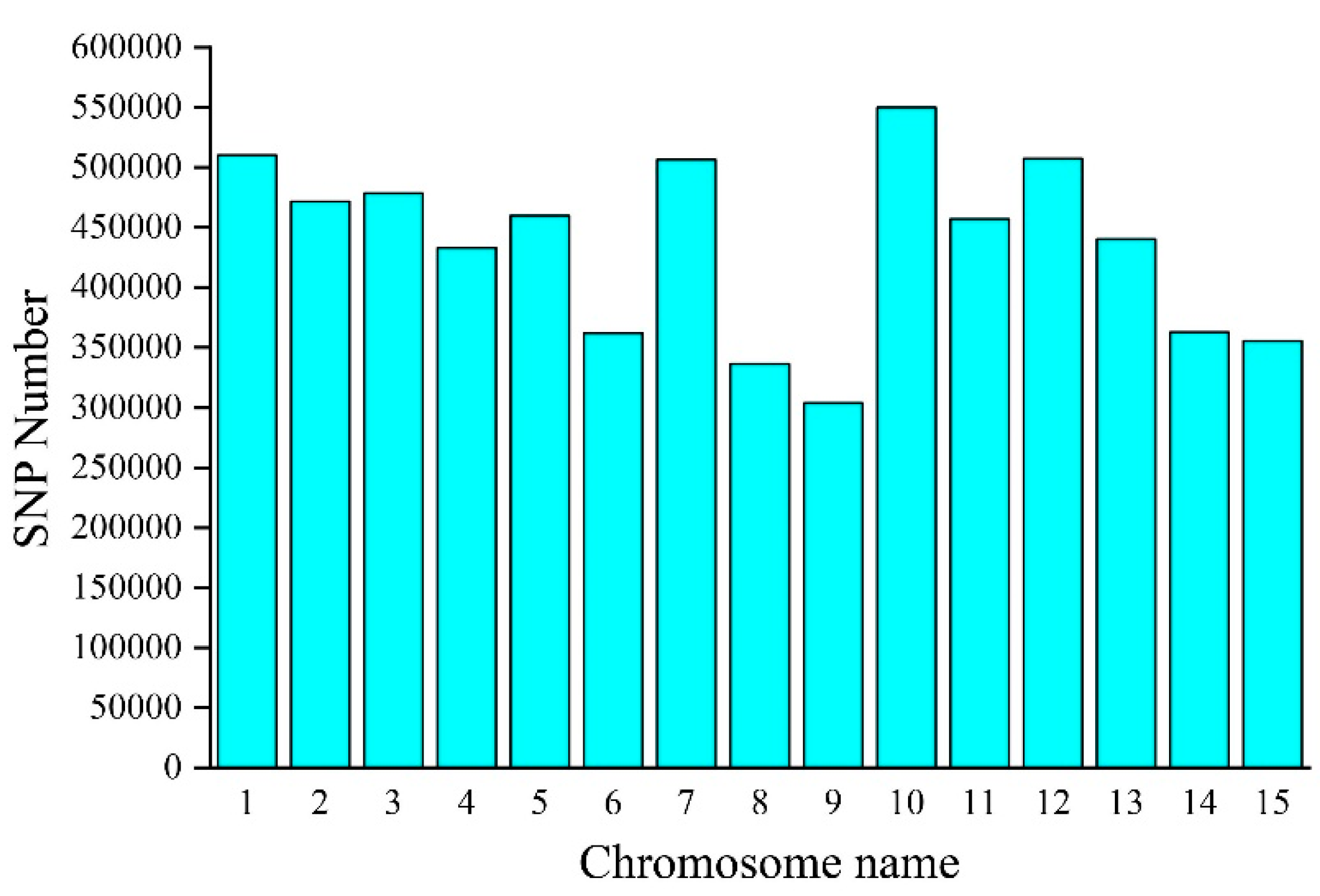
3.1. SNP Variation Detection
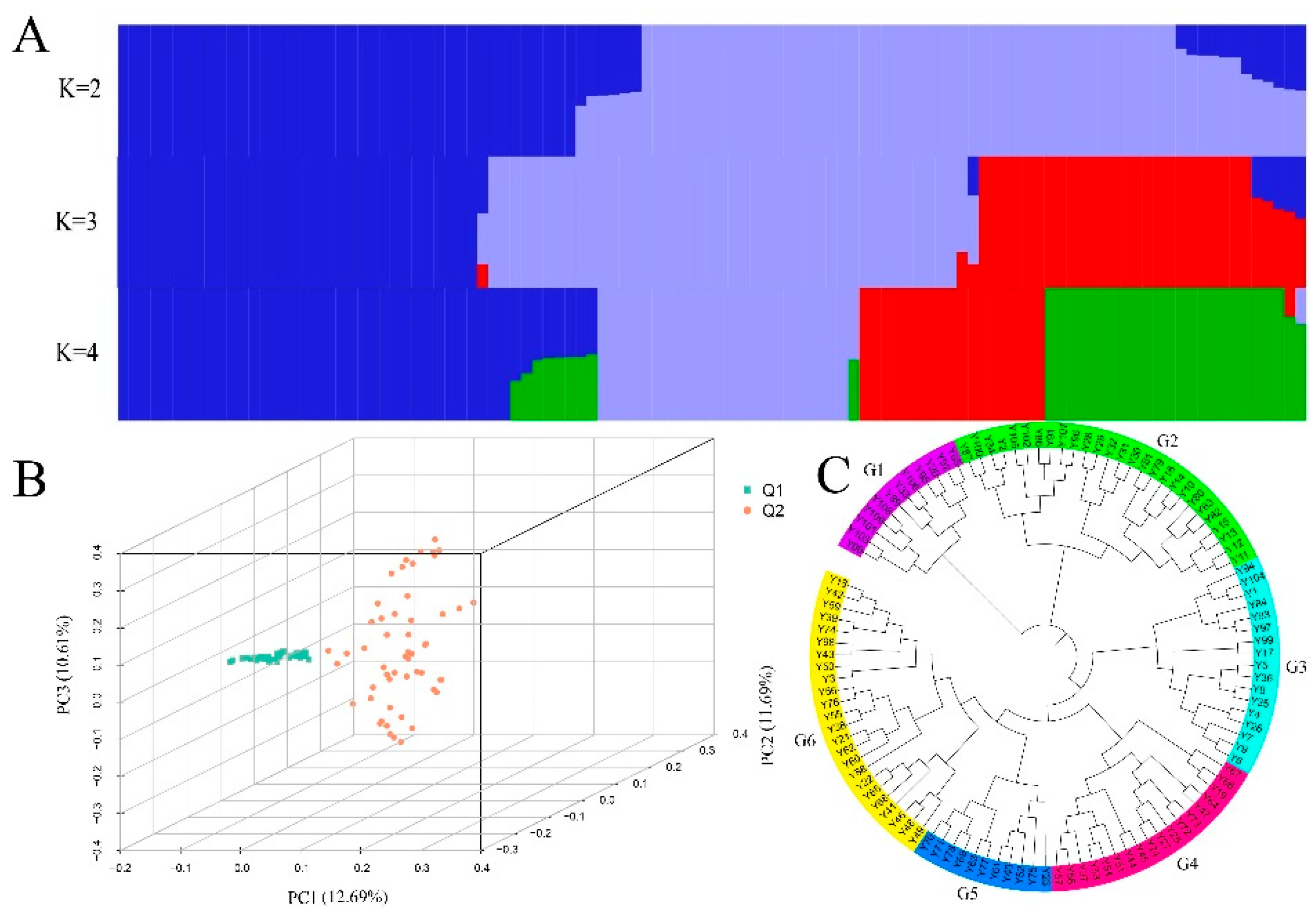
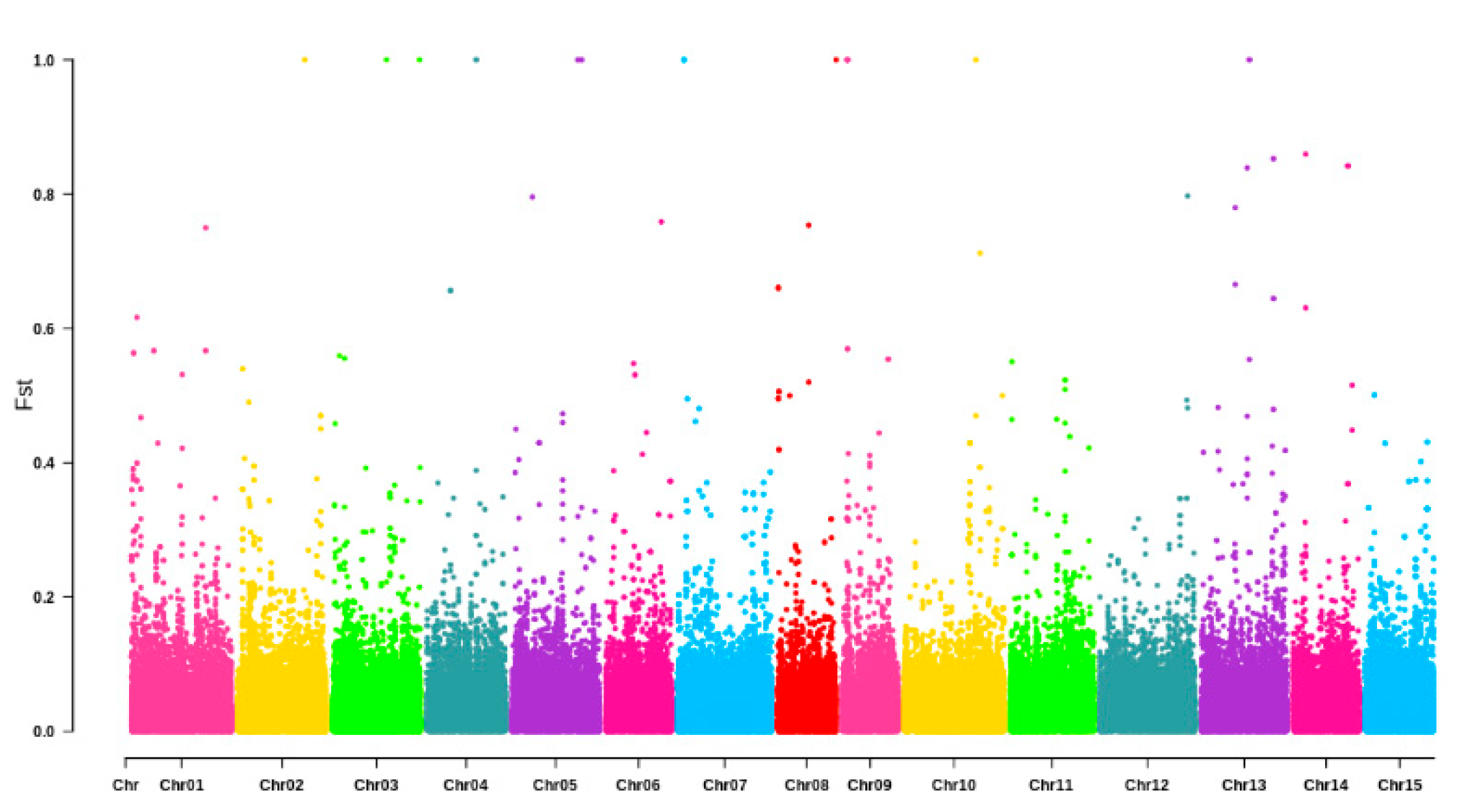
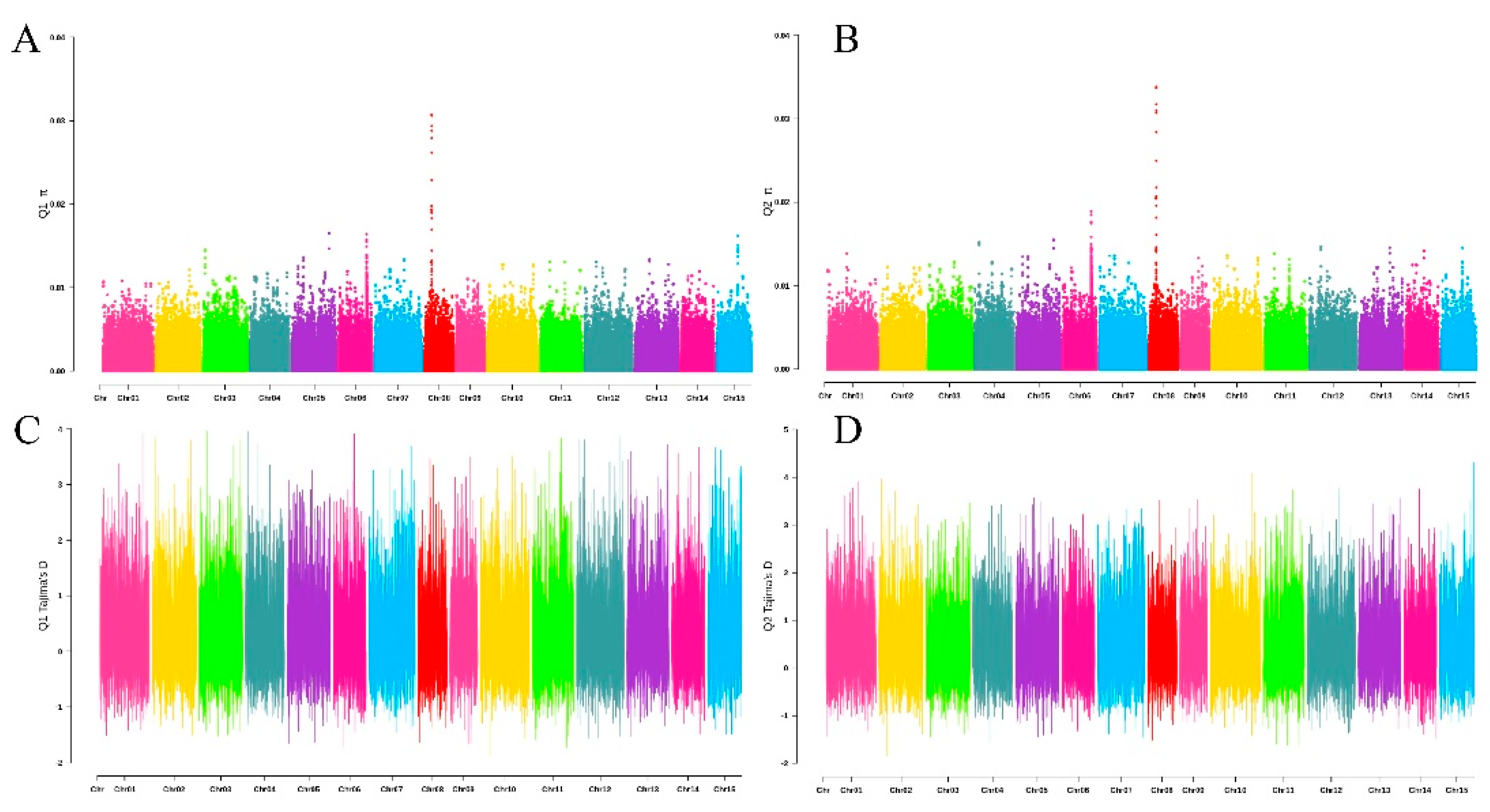
3.2. Genetic Structure
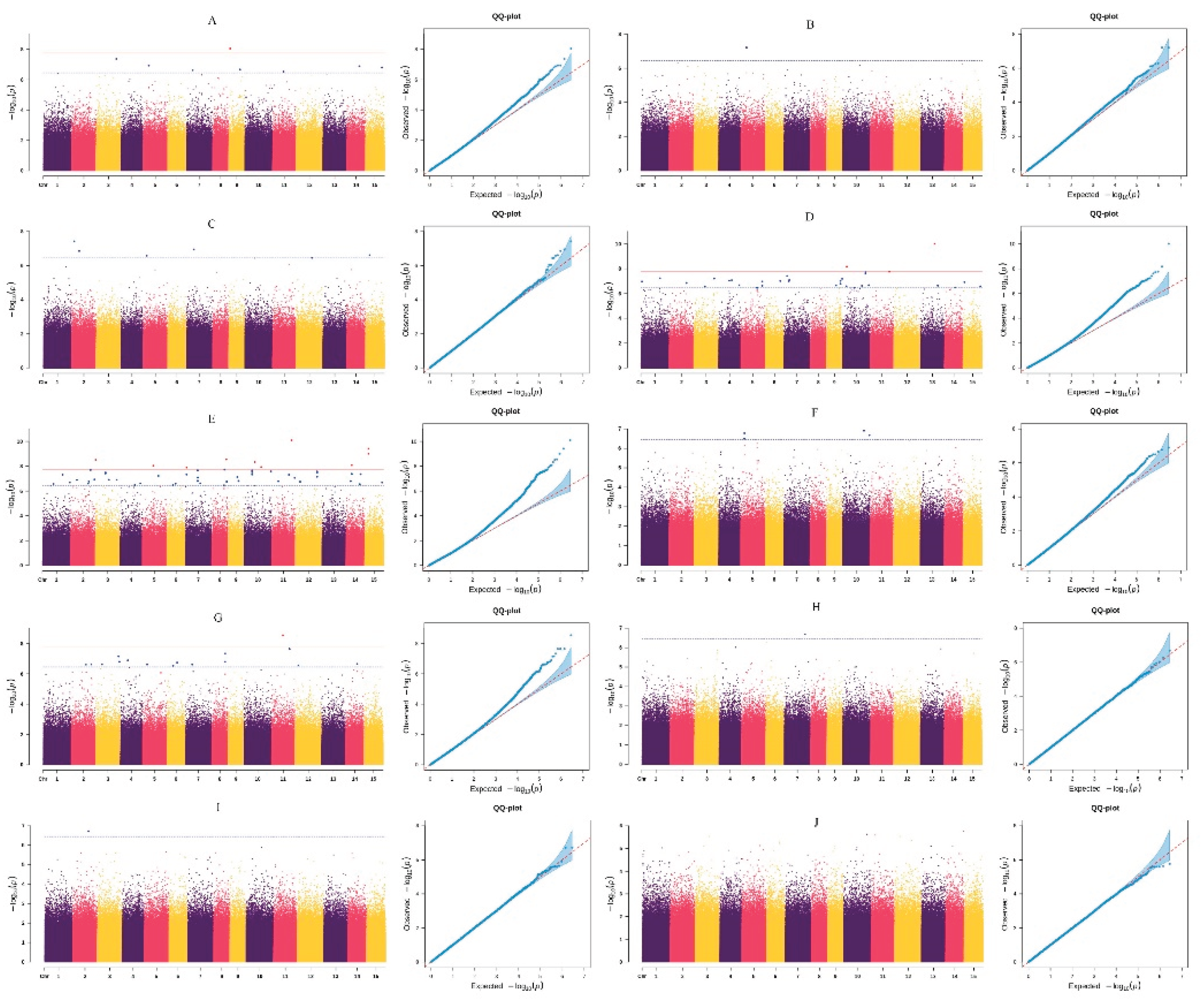
3.3. GWAS of Fruit Traits
3.4. Candidate Genes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Data Availability Statement
Conflicts of Interest
References
- Yin, D.D.; Li, S.S.; Wu, Q.; Feng, C.Y.; Li, B.; Wang, Q.Y.; Wang, L.S.; Xu, W.Z. Advances in Research of Six Woody Oil Crops in China. Chin. Bull. Bot. 2018, 53, 110–125. [Google Scholar] [CrossRef]
- Zhang, P.P.; Chu, Z.Q.; Jiao, Q.; Zhou, B.Q.; Zhang, D.Q. Research Progress in Processing and Utilization of Woody Oil By-Products. China For. Prod. Lnd. 2022, 59, 63–68. [Google Scholar] [CrossRef]
- Quan, W.X.; Wang, A.P.; Gao, C.; Li, C.C. Applications of Chinese Camellia Oleifera and Its By-Products: A Review. Front. Chem. 2022, 10, 921246. [Google Scholar] [CrossRef]
- Liu, J.B.; Liao, Z.P.; Sun, T.Y.; Feng, N.; Long, Q.Z.; Zhong, H.Y.; Zhou, B. Fatty Acids, Triacylglycerol and Sn-2 Fatty Acids Distributions Variations in Seed Oil from Camellia Cultivars. J. Food Nutr. Sci. 2018, 6, 143–153. [Google Scholar] [CrossRef]
- Li, H.; Zhou, G.Y.; Zhang, H.Y.; Liu, J.A. Research Progress on the Health Function of Tea Oil. J. Plant Med. 2011, 5, 485–489. [Google Scholar] [CrossRef]
- Yu, J.; Yan, H.Q.; Wu, Y.G.; Wang, Y.; Xia, P.G. Quality Evaluation of the Oil of Camellia Spp. Foods 2022, 11, 2221. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Liu, L.; Zhou, Y.L.; Zheng, Q.; Luo, S.Y.; Xiang, T.T.; Zhou, L.J.; Feng, S.L.; Yang, H.Y.; Ding, C.B. Characterization and Comprehensive Evaluation of Phenotypic Characters in Wild Camellia Oleifera Germplasm for Conservation and Breeding. Front. Plant Sci. 2023, 14, 1052890. [Google Scholar] [CrossRef]
- Wan, X.Q.; Sun, D.C.; Nie, Y.M.; Wang, Q.M.; Zhang, T.F.; Wang, R.; Li, F.F.; Zhao, X.D.; Gao, C. Analysis and Evaluation of Camellia Oleifera Abel. Germplasm Fruit Traits from the High-Altitude Areas of East Guizhou Province, China. Sci. Rep. 2024, 14, 18440. Available online: https://hub.uu2025.xyz/10.1038/s41598-024-69454-9. [CrossRef]
- Peng, S.F. Genome-Wide Association Studies of Important Economic Traits in Camellia oleifera Abel. Ph.D. Dissertation, Beijing Forestry University, Beijing, China, 2021. [Google Scholar] [CrossRef]
- Lv, H.H.; Wu, Y.F.; Li, Y.; Liang, T.; Zhou, L.X.; Chen, H. Variation and selection of excellent fruit traits in Camellia oleifera clones. J. For. Environ. 2021, 41, 519–526. [Google Scholar] [CrossRef]
- Chai, J.Y.; Wang, K.L.; Yao, X.H.; Teng, J.H.; Lin, P. Genetic Analysis of the Fruit and Oil Related Traits on Hybrid Offspring of Nested Mating of Camellia Oleifera. For. Res. 2023, 36, 1–10. [Google Scholar] [CrossRef]
- Zhou, W.C.; Wang, Z.W.; Dong, L.; Wen, Q.; Huang, W.Y.; Li, T.; Ye, J.S.; Xu, L.A. Analysis on the character diversity of fruit and seed of Camellia chekiangoleosa. J. Nanjing For. Univ. 2021, 45, 51–59. [Google Scholar] [CrossRef]
- Liu, L.X.; Yao, X.H.; Zeng, H.T.; Teng, J.H.; Xu, H.; Chen, J.J.; Chang, J. Effects of Flowering Biology and Pollination Methods on Fruit Development of Camellia oleifera. J. Southwest Univ. 2025, 47, 93–103. [Google Scholar] [CrossRef]
- Liang, T.; Li, X.S.; Li, Y.; Lv, H.H.; Zheng, G.H.; Chen, H. Variations in Fruit Traits in Cross-bred Camellia oleifera Clones. J. For. Environ. 2022, 42, 166–173. [Google Scholar] [CrossRef]
- Li, J.N.; Wu, M.Z.; Li, Y.; Ye, L.; Chen, S.P.; Chen, H. Comprehensive Analysis of the Phenotypic Variation among Seedling Populations of Camellia chekiangoleosa. J. For. Environ. 2024, 44, 274–282. [Google Scholar] [CrossRef]
- Ye, L.; Li, Y.; Liu, Y.X.; Zhou, L.X.; Li, J.N.; Liang, T.; Xie, W.W.; Xie, Y.Q.; Li, Z.Z.; Lv, H.H.; et al. Genetic Linkage Map Construction and QTL Mapping for Juvenile Leaf and Growth Traits in Camellia Oleifera. Agronomy 2024, 14, 3022. [Google Scholar] [CrossRef]
- Wang, Z.H.; Yong, C.W.; Chen, G.; Ding, Y.D.; Zhao, Z.G.; Lyu, A.Q.; Han, C.Y. Genetic Diversity Analysis of Different Cultivars of Camellia Oleifera Based on CDDP and ITS Molecular Markers. Chin. J. Oil Crop Sci. 2022, 44, 94–102. [Google Scholar] [CrossRef]
- Li, Y.M.; Yin, L.J.; He, X.Y.; Hu, C.L.; Wu, R.H.; Long, Q.; Yuan, D.Y.; Xiao, S.X. Ploidy and Fruit Trait Variation of Oil-Tea Camellia: Implication for Ploidy Breeding. J. Integr. Agric. 2024, 23, 2662–2673. [Google Scholar] [CrossRef]
- Ye, T.W.; Li, S.F.; Li, Y.M.; Xiao, S.X.; Yuan, D.Y. Impact of Polyploidization on Genome Evolution and Phenotypic Diversity in Oil-Tea Camellia. Ind. Crops Prod. 2024, 218, 118928. [Google Scholar] [CrossRef]
- Li, J.Q.; Liu, Y.C.; Wei, X.; Yang, Y.M.; Yang, Y.C.; Wang, S.; Gao, S.Q.; Lin, J.Q.; Xu, Y.G.; Sun, B.; et al. Research progress on genome-wide association analysis (GWAS) for fruit quality and resistance traits. Jiangsu Agric. Sci. 2024, 52, 10–19. [Google Scholar] [CrossRef]
- Jia, G.Q.; Huang, X.H.; Zhi, H.; Zhao, Y.; Zhao, Q.; Li, W.J.; Chai, Y.; Yang, L.F.; Liu, K.Y.; Lu, H.Y.; et al. A Haplotype Map of Genomic Variations and Genome-Wide Association Studies of Agronomic Traits in Foxtail Millet (Setaria Italica). Nat. Genet. 2013, 45, 957–961. [Google Scholar] [CrossRef]
- Wang, Q.X.; Xie, W.B.; Xing, H.K.; Yan, J.; Meng, X.Z.; Li, X.L.; Fu, X.K.; Xu, J.Y.; Lian, X.M.; Yu, S.B.; et al. Genetic Architecture of Natural Variation in Rice Chlorophyll Content Revealed by a Genome-Wide Association Study. Mol. Plant 2015, 8, 946–957. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Yang, Y.M.; Zhang, S.Y.; Che, Z.J.; Yuan, W.J.; Yu, D.Y. GWAS Reveals Two Novel Loci for Photosynthesis-Related Traits in Soybean. Mol. Genet. Genomics 2020, 295, 705–716. [Google Scholar] [CrossRef]
- Zhang, M.Y.; Xue, C.; Hu, H.J.; Li, J.M.; Xue, Y.S.; Wang, R.Z.; Fan, J.; Zou, C.; Tao, S.T.; Qin, M.F.; et al. Genome-Wide Association Studies Provide Insights into the Genetic Determination of Fruit Traits of Pear. Nat. Commun. 2021, 12, 1144. [Google Scholar] [CrossRef]
- Zahid, G.; Aka Kaçar, Y.; Dönmez, D.; Kuden, A.; Giordani, T. Perspectives and Recent Progress of Genome-Wide Association Studies (GWAS) in Fruits. Mol. Biol. Rep. 2022, 49, 5341–5352. [Google Scholar] [CrossRef]
- Huang, X.M. Genetic Structure of Hexaploid Wild Camellia oleifera in Mount Jinggang and Lu based on Microsatellite Markers. Master’s Thesis, Nanchang University, Jiangxi, China, 2016. [Google Scholar]
- Zhu, Y.Y.; Yao, Y.; Xu, J.J.; Wang, G. Analysis on Phenotypic Diversity of Fruit and Seed Characters in Camellia meiocarpa. Guizhou Agric. Sci. 2016, 44, 125–129. [Google Scholar]
- Li, L.; Xu, F.; Zhang, G.F. Studies and Application of Gene Resources and Molecular Breeding of Aquatic Animals. J. Agric. Sci. Technol. 2011, 13, 102–110. [Google Scholar]
- Yan, M.M.; Wei, G.C.; Pan, X.H.; Ma, H.L.; Li, W.Z. A Method Suitable for Extracting Genomic DNA from Animal and Plant――Modified CTAB Method. Agric. Sci. Technol. 2008, 9, 39–41. [Google Scholar] [CrossRef]
- Zou, M.L.; Xia, Z.Q. Hyper-Seq: A Novel, Effective, and Flexible Marker-Assisted Selection and Genotyping Approach. The Innovation 2022, 3, 100254. [Google Scholar] [CrossRef]
- Chen, S.F.; Zhou, Y.Q.; Chen, Y.R.; Gu, J. Fastp: An Ultra-Fast All-in-One FASTQ Preprocessor. Bioinformatics 2018, 34, i884–i890. Available online: https://hub.uu2025.xyz/10.1093/bioinformatics/bty560. [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. 1000 Genome Project Data Processing Subgroup. The Sequence Alignment/Map Format and SAMtools. Bioinformatics 2009, 25, 2078–2079. Available online: https://hub.uu2025.xyz/10.1093/bioinformatics/btp352. [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and Accurate Short Read Alignment with Burrows-Wheeler Transform. Bioinformatics 2009, 25, 1754–1760. Available online: https://hub.uu2025.xyz/10.1093/bioinformatics/btp324. [CrossRef]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce Framework for Analyzing next-Generation DNA Sequencing Data. Genome Res. 2010, 20, 1297–1303. Available online: https://hub.uu2025.xyz/10.1101/gr.107524.110. [CrossRef]
- Wang, K.; Li, M.Y.; Hakonarson, H. ANNOVAR: Functional Annotation of Genetic Variants from High-Throughput Sequencing Data. Nucleic Acids Res. 2010, 38, e164–e164. Available online: https://hub.uu2025.xyz/10.1093/nar/gkq603. [CrossRef] [PubMed]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.R.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.W.; Daly, M.J.; et al. PLINK: A Tool Set for Whole-Genome Association and Population-Based Linkage Analyses. Am. J. Hum. Genet. 2007, 81, 559–575. Available online: https://hub.uu2025.xyz/10.1086/519795. [CrossRef] [PubMed]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. Fasttree: Computing Large Minimum Evolution Trees with Profiles Instead of a Distance Matrix. Mol. Biol. Evol. 2009, 26, 1641–1650. Available online: https://hub.uu2025.xyz/10.1093/molbev/msp077. [CrossRef]
- Broström, G. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013; Available online: http://www.R-project.org/ (accessed on 25 January 2026)ISBN 3-900051-07-0.
- Alexander, D.H.; Novembre, J.; Lange, K.L. Fast Model-Based Estimation of Ancestry in Unrelated Individuals. Genome Res. 2009, 19, 1655–1664. Available online: https://hub.uu2025.xyz/10.1101/gr.094052.109. [CrossRef]
- Francis, R.M. Pophelper: An R Package and Web App to Analyse and Visualize Population Structure. Mol. Ecol. Resour. 2017, 17, 27–32. Available online: https://hub.uu2025.xyz/10.1111/1755-0998.12509. [CrossRef]
- Browning, B.L.; Tian, X.; Zhou, Y.; Browning, S.R. Fast Two-Stage Phasing of Large-Scale Sequence Data. Am. J. Hum. Genet. 2021, 108, 1880–1890. [Google Scholar] [CrossRef]
- Guo, Z.F.; Yang, Q.N.; Huang, F.F.; Zheng, H.J.; Sang, Z.Q.; Xu, Y.F.; Zhang, C.; Wu, K.S.; Tao, J.J.; Prasanna, B.M.; et al. Development of High-Resolution Multiple-SNP Arrays for Genetic Analy-Ses and Molecular Breeding through Genotyping by Target Sequencing and Liquid Chip. Plant Commun. 2021, 2, 100230. Available online: https://hub.uu2025.xyz/10.1016/j.xplc.2021.100230. [CrossRef] [PubMed]
- Younessi-Hamzekhanlu, M.; Gailing, O. Genome-Wide SNP Markers Accelerate Perennial Forest Tree Breeding Rate for Disease Resistance through Marker-Assisted and Genome-Wide Selection. Int. J. Mol. Sci. 2022, 23, 12315. Available online: https://hub.uu2025.xyz/10.3390/ijms232012315. [CrossRef]
- Zhang, E.H.; Wang, X.Y.; Qin, Z.H.; Zhao, W.D.; Wei, C.J.; Wang, P.L. Genetic diversity analysis of Camellia oleifera in Guangxi using SSR markers. Guihaia 2016, 36, 806–811. [Google Scholar]
- Jiang, D.Z.; Fang, Y.Y.; Xiao, X.; Cheng, H.; Zhang, X.H.; Cheng, J.Y.; Li, L.L. Identifying Camellia oleifera Germplasm Accessions with Inter Simple Sequence Repeat Markers. Hubei Agric. Sci. 2018, 57, 119–125. [Google Scholar] [CrossRef]
- Chen, X.; Yun, Y.; Wu, Y.J.; Qi, H.S.; Yang, L.R.; Chen, J.L.; Zheng, D.J. Genetic Diversity Analysis of Camellia Oleifera Resources Based on SRAP Markers in Hainan Island. J. Trop. Subtrop. Bot. 2019, 27, 659–668. [Google Scholar] [CrossRef]
- Liu, K.; Wang, D.X.; Jiang, Z.P.; Mo, J.F.; Zeng, W.J.; Zhang, N.Y.; Liang, G.X. Development of SNP Markers of Camellia Oleifera Based on SLAF-Seq and Early Identification of Hybrids. Guangxi For. Sci. 2018, 47, 13–17. [Google Scholar] [CrossRef]
- Lin, P.; Wang, K.L.; Yao, X.H.; Ren, H.D. Development of DNA Molecular ID in Camellia Oleifera Germplasm Based on Transcriptome-Wide SNPs. Sci. Agric. Sin. 2023, 56, 217–235. [Google Scholar] [CrossRef]
- Zhang, J. Detection of Mutations of PAX9 and MSX1 Gene in Xinjiang Uyghur Patients with Non-syndrome Tooth Agenesis. Master’s Thesis, Xinjiang Medical University, Xinjiang, China, 2011. [Google Scholar]
- Huang, Y. Population Genetic Structure and Interspecific Introgressive Hybridization between Camellia meiocarpa and C. oleifera. Chin. J. Appl. Ecol. 2013, 24, 2345–2352. [Google Scholar] [CrossRef]
- Sun, Q. GWAS Analysis of Agronomic Traits of Cassava Germplasm in Brazil. Master’s Thesis, Guangxi University, Guangxi, China, 2020. [Google Scholar] [CrossRef]
- Zhu, Y.Z.; Liang, D.Y.; Song, Z.J.; Tan, Y.; Guo, X.L.; Wang, D.L. Genetic Diversity Analysis and Core Germplasm Collection Construction of Camellia Oleifera Based on Fruit Phenotype and SSR Data. Genes 2022, 13, 2351. [Google Scholar] [CrossRef] [PubMed]
- Li, G.L.; Zhang, H.L.; Li, J.J.; Zhang, Z.Y.; Li, Z.C. Genetic Control of Panicle Architecture in Rice. Crop J. 2021, 9, 590–597. [Google Scholar] [CrossRef]
- Kumar, S.; Molloy, C.; Hunt, M.; Deng, C.H.; Wiedow, C.; Andre, C.; Dare, A.; McGhie, T. GWAS Provides New Insights into the Genetic Mechanisms of Phytochemicals Production and Red Skin Colour in Apple. Hortic. Res. 2022, 9, uhac218. [Google Scholar] [CrossRef]
- Chen, P.P. Genome-wide Association Study of Common Walnut (Juglans regia L.) Fruit Traits. Ph.D. Dissertation, Northwest University, Shan’xi, China, 2022. [Google Scholar] [CrossRef]
- Xiao, Y. Preliminary Study on Population Genomics and Conservation Genomics of Oryza rufipogon. Master’s Thesis, South China Agricultural University, Guangzhou, China, 2020. [Google Scholar] [CrossRef]
- Li, Z.J.; Qin, Y. Chromatin Accessibility and the Gene Expression Regulation in Plants. Chin. Bull. Bot. 2021, 56, 664–675. [Google Scholar] [CrossRef]
- Dai, H. Comparative Analysis of the Association Between Major Agronomic Traits and SSR Molecular Markers in Camellia oleifera. Master’s Thesis, Guizhou Normal University, Guizhou, China, 2024. [Google Scholar] [CrossRef]
| Trait | SNP | Gene ID | KEGG pathway |
|---|---|---|---|
| Fruit shape index | Chr02:20212286 | maker-Chr2-snap-gene-202.6-mRNA-1 | Messenger RNA biogenesis |
| Number of seeds per fruit | Chr01:84240256 | augustus_masked-Chr1-processed-gene-841.29-mRNA-1 | Transcription factors |
| Chr03:105301961 | augustus_masked-Chr3-processed-gene-1053.14-mRNA-1 | Ribosome | |
| Chr03:167365123 | maker-Chr3-snap-gene-1673.21-mRNA-1 | Ribosome biogenesis | |
| Chr08:96873006 | maker-Chr8-snap-gene-969.6-mRNA-1 | Protein kinases | |
| Chr08:114566238 | maker-Chr8-snap-gene-1146.2-mRNA-1 | Carotenoid biosynthesis | |
| Chr08:118268465 | genemark-Chr8-processed-gene-1182.39-mRNA-1 | Cytochrome P450 | |
| Chr10:63527409 | maker-Chr10-snap-gene-635.3-mRNA-1 | Ubiquitin system | |
| Chr10:134255167 | maker-Chr10-snap-gene-1342.19-mRNA-1 | Enzymes with EC numbers | |
| Chr11:153463775 | snap_masked-Chr11-processed-gene-1534.13-mRNA-1 | Chromosome and associated proteins | |
| Chr14:35355883 | augustus_masked-Chr14-processed-gene-353.13-mRNA-1 | Cytochrome P450 | |
| Chr14:114275276 | snap_masked-Chr14-processed-gene-1142.45-mRNA-1 | Transcription factors | |
| Peel thickness | Chr01:6227267 | maker-Chr1-snap-gene-62.12-mRNA-1 | Cysteine and methionine metabolism |
| Chr04:159617039 | maker-Chr4-snap-gene-1596.0-mRNA-2 | Membrane trafficking | |
| Chr05:167572639 | maker-Chr5-snap-gene-1676.3-mRNA-1 | Exosome | |
| Chr07:41713895 | maker-Chr7-snap-gene-417.23-mRNA-1 | Transcription factors | |
| Chr09:72920012 | maker-Chr9-snap-gene-729.27-mRNA-1 | Protein kinases | |
| Chr13:135481819 | augustus_masked-Chr13-processed-gene-1354.42-mRNA-1 | Membrane trafficking | |
| Fruit height | Chr07:49799009 | maker-Chr7-snap-gene-497.36-mRNA-2 | Peptidases and inhibitors |
| Chr09:85815245 | maker-Chr9-snap-gene-858.43-mRNA-1 | Ubiquitin system | |
| Chr14:97790223 | maker-Chr14-snap-gene-977.10-mRNA-1 | Glycosyltransferases | |
| Fresh seed yield | Chr02:158899204 | maker-Chr2-snap-gene-1588.1-mRNA-1 | Chaperones and folding catalysts |
| Chr08:106777293 | augustus_masked-Chr8-processed-gene-1068.0-mRNA-1 | Transcription factors |
| Candidate gene | GO Category | Annotation |
|---|---|---|
| maker-Chr4-snap-gene-1582.40-mRNA-1 | molecular_function | function unknown |
| maker-Chr7-snap-gene-82.12-mRNA-1 | molecular_function | heterocyclic compound binding |
| maker-Chr12-snap-gene-379.34-mRNA-1 | molecular_function | nucleotide binding |
| maker-Chr14-snap-gene-1071.26-mRNA-1 | molecular_function | nucleotide binding |
| augustus_masked-Chr10-processed-gene-2066.42-mRNA-1 | molecular_function | heterocyclic compound binding |
| augustus_masked-Chr11-processed-gene-1405.6-mRNA-1 | molecular_function | function unknown |
| maker-Chr2-snap-gene-1558.2-mRNA-1 | cellular_component | bounding membrane of organelle |
| augustus_masked-Chr3-processed-gene-1053.14-mRNA-1 | cellular_component | protein-containing complex |
| maker-Chr3-snap-gene-1673.21-mRNA-1 | cellular_component | protein-containing complex |
| maker-Chr8-snap-gene-1008.6-mRNA-2 | cellular_component | bounding membrane of organelle |
| genemark-Chr8-processed-gene-1182.39-mRNA-1 | cellular_component | endoplasmic reticulum subcompartment |
| augustus_masked-Chr10-processed-gene-634.18-mRNA-1 | cellular_component | bounding membrane of organelle |
| maker-Chr10-snap-gene-635.3-mRNA-1 | cellular_component | ubiquitin ligase complex |
| maker-Chr10-snap-gene-1524.3-mRNA-1 | cellular_component | endoplasmic reticulum subcompartment |
| maker-Chr2-snap-gene-1588.1-mRNA-1 | cellular_component | bounding membrane of organelle |
| maker-Chr2-snap-gene-202.6-mRNA-1 | biological_process | reproduction |
| augustus_masked-Chr2-processed-gene-593.9-mRNA-1 | biological_process | RNA biosynthetic process |
| maker-Chr5-snap-gene-268.33-mRNA-1 | biological_process | RNA biosynthetic process |
| augustus_masked-Chr1-processed-gene-841.29-mRNA-1 | biological_process | reproduction |
| maker-Chr4-snap-gene-1150.47-mRNA-1 | biological_process | reproduction |
| maker-Chr5-snap-gene-1262.30-mRNA-1 | biological_process | reproduction |
| snap_masked-Chr6-processed-gene-475.2-mRNA-1 | biological_process | defense response to other organism |
| augustus_masked-Chr6-processed-gene-670.4-mRNA-1 | biological_process | reproduction |
| maker-Chr8-snap-gene-969.6-mRNA-1 | biological_process | cellular response to hormone stimulus |
| maker-Chr8-snap-gene-1146.2-mRNA-1 | biological_process | monocarboxylic acid metabolic process |
| maker-Chr10-snap-gene-1342.19-mRNA-1 | biological_process | transition metal ion transport |
| maker-Chr11-snap-gene-1345.57-mRNA-1 | biological_process | RNA biosynthetic process |
| augustus_masked-Chr12-processed-gene-1709.3-mRNA-1 | biological_process | reproduction |
| augustus_masked-Chr14-processed-gene-353.13-mRNA-1 | biological_process | monocarboxylic acid metabolic process |
| snap_masked-Chr14-processed-gene-1142.45-mRNA-1 | biological_process | reproduction |
| maker-Chr15-snap-gene-1364.34-mRNA-1 | biological_process | reproduction |
| maker-Chr1-snap-gene-62.12-mRNA-1 | biological_process | reproduction |
| maker-Chr4-snap-gene-1596.0-mRNA-2 | biological_process | autophagosome assembly |
| maker-Chr5-snap-gene-1275.2-mRNA-1 | biological_process | reproduction |
| maker-Chr5-snap-gene-1676.3-mRNA-1 | biological_process | transition metal ion transport |
| maker-Chr7-snap-gene-376.6-mRNA-1 | biological_process | regulation of shoot system morphogenesis |
| maker-Chr7-snap-gene-417.23-mRNA-1 | biological_process | reproduction |
| maker-Chr9-snap-gene-729.27-mRNA-1 | biological_process | reproduction |
| maker-Chr9-snap-gene-1211.11-mRNA-1 | biological_process | autophagosome assembly |
| augustus_masked-Chr13-processed-gene-1354.42-mRNA-1 | biological_process | regulation of localization |
| maker-Chr5-snap-gene-439.0-mRNA-1 | biological_process | reproduction |
| maker-Chr7-snap-gene-497.36-mRNA-2 | biological_process | monocarboxylic acid metabolic process |
| maker-Chr11-snap-gene-848.23-mRNA-1 | biological_process | reproduction |
| maker-Chr14-snap-gene-977.10-mRNA-1 | biological_process | reproduction |
| augustus_masked-Chr11-processed-gene-881.28-mRNA-1 | biological_process | plant organ development |
| maker-Chr5-snap-gene-439.0-mRNA-1 | biological_process | reproduction |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




