Submitted:
08 February 2026
Posted:
10 February 2026
You are already at the latest version
Abstract
Keywords:
1. Background: The Bioenergetic Evolution of Oncology
2. Aims and Rationale
- Deconstruct the molecular mechanisms by which the PI3K/Akt/mTOR and IGF-1 axes regulate mitochondrial dynamics and bioenergetics.
- Define the “Triple-Hit” strategy: a multi-tiered approach targeting glucose sensing, organelle morphology, and mitochondrial epigenetics.
- Evaluate the role of mitochondrial retrograde signaling (the “mito-to-nucleus” talk) in driving tumor progression and resistance.
- Propose the concept of “Nutri-phenotyping” as the solution to the translational gap, moving toward precision oncology.
3. The Growth Factor-Mitochondrial Nexus: A Molecular Deep-Dive
3.1. PI3K/Akt/mTOR: The Master Regulator of Glycolytic Commitment
3.2. IGF-1 Signaling: Driving Quantity over Quality
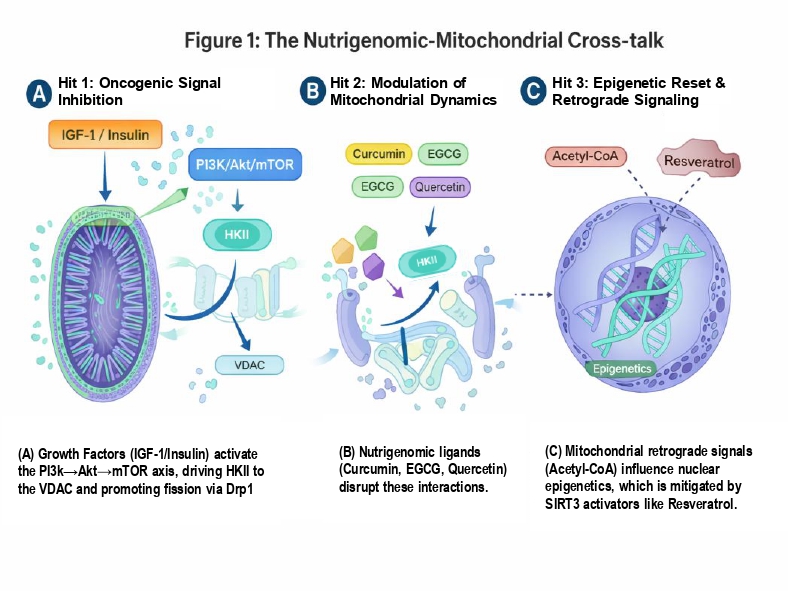
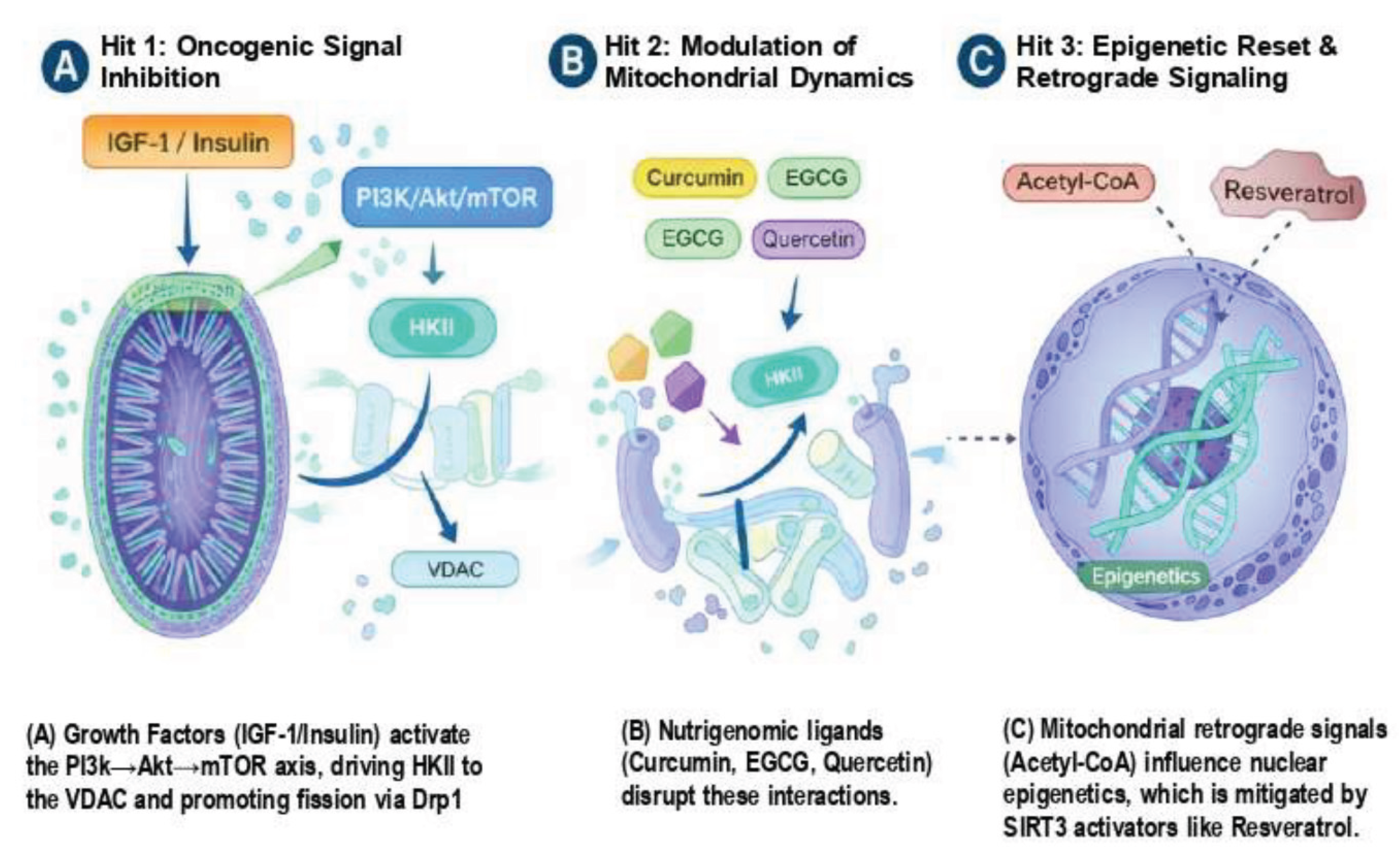
4. The “Triple-Hit” Strategy: A Framework for Therapeutic Intervention
4.1. Hit 1: Disruption of Glucose Sensing and “Metabolic Uncoupling”
4.2. Hit 2: Restoration of Mitochondrial Dynamics (Fusion vs. Fission)
4.3. Hit 3: Reversing “Mito-Epigenetic” Silencing of mtDNA
5. Retrograde Signaling: The “Mito-to-Nucleus” Dialogue
6. The Translational Gap: Challenges and Bioengineered Solutions
7. Conclusions and Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AMPK | AMP-activated protein kinase |
| DNMT | DNA methyltransferase |
| ETC | Electron transport chain |
| GFS | Growth factor signaling |
| HDAC | Histone deacetylase |
| HKII | Hexokinase II |
| IGF-1 | Insulin-like growth factor 1 |
| mtDNA | Mitochondrial DNA |
| OXPHOS | Oxidative phosphorylation |
| ROS | Reactive oxygen species |
| VDAC | Voltage-dependent anion channel |
References
- Warburg, O. On the origin of cancer cells. Science 1956, 123, 309–14. [Google Scholar] [CrossRef]
- Hanahan, D. Hallmarks of cancer: New dimensions. Cancer Discov. 2022, 12, 31–46. [Google Scholar] [CrossRef]
- Wallace, DC. Mitochondria and cancer. Nat Rev Cancer 2012, 12, 685–98. [Google Scholar] [CrossRef]
- Weinberg, SE; Chandel, NS. Targeting mitochondria metabolism for cancer therapy. Nat Chem Biol. 2015, 11, 9–15. [Google Scholar] [CrossRef]
- Pollak, M. Insulin and insulin-like growth factor signalling in neoplasia. Nat Rev Cancer 2008, 8, 915–28. [Google Scholar] [CrossRef]
- Pavlova, NN; Thompson, CB. The emerging hallmarks of cancer metabolism. Cell Metab. 2016, 23, 27–47. [Google Scholar] [CrossRef] [PubMed]
- Kaput, J. Nutrigenomics research: A review. Mol Nutr Food Res. 2014, 58, 1–12. [Google Scholar]
- Gholipour Maralan, F. Nutrigenomic mitochondrial landscapes. J Biomed Sci. 2024, 31, 112. [Google Scholar]
- Ristow, M. Mitohormesis: Promoting health and lifespan. Dose Response 2014, 12, 255–68. [Google Scholar] [CrossRef] [PubMed]
- Roberts, DJ; et al. Hexokinase II integrates glycolysis and autophagy. Cell Cycle 2015, 14, 2475–84. [Google Scholar]
- Bohn, T; et al. Mind the gap: From nutrition to health. Nutrients 2015, 7, 2745–58. [Google Scholar]
- Rein, MJ; et al. Bioavailability of polyphenols. Nutrients 2013, 5, 1–46. [Google Scholar]
- Sabatini, DM. Twenty-five years of mTOR. PNAS 2017, 114, 11818–25. [Google Scholar] [CrossRef]
- Vander Heiden, MG; et al. Understanding the Warburg Effect. Science 2009, 324, 1029–33. [Google Scholar] [CrossRef]
- Ravindran, J; et al. Curcumin and cancer cells. Cell Cycle 2009, 8, 1327–33. [Google Scholar]
- Galluzzi, L; et al. Metabolic control of cell death. Nat Rev Mol Cell Biol. 2014, 15, 755–72. [Google Scholar]
- Zhang, Y; et al. Berberine inhibits mTORC1. J Biol Chem. 2010, 285, 22251–62. [Google Scholar]
- Baur, JA; Sinclair, DA. Therapeutic potential of resveratrol. Nat Rev Drug Discov. 2006, 5, 493–506. [Google Scholar] [CrossRef] [PubMed]
- Scarpulla, RC. Transcriptional paradigms in mammalian mitochondrial biogenesis. Physiol Rev. 2008, 88, 611–38. [Google Scholar] [CrossRef] [PubMed]
- Yun, J; Finkel, T. Mitohormesis. Cell Metab. 2014, 19, 757–73. [Google Scholar] [CrossRef] [PubMed]
- Van Aller, GS; et al. EGCG inhibits PI3K. Lipids 2010, 45, 889–96. [Google Scholar]
- Semenza, GL. HIF-1 and cancer metabolism. Nat Rev Cancer 2013, 13, 672–82. [Google Scholar]
- Beevers, CS; et al. Curcumin inhibits mTOR signaling. Cancer Res. 2006, 66, 6139–46. [Google Scholar]
- Lu, C; Thompson, CB. Metabolic regulation of epigenetics. Cell Metab. 2012, 16, 9–17. [Google Scholar] [CrossRef]
- Zhao, J; et al. Drp1-dependent fission in cancer. Cell Death Dis. 2013, 4, e613. [Google Scholar]
- Youle, RJ; van der Bliek, AM. Mitochondrial fission, fusion, and stress. Science 2012, 337, 1062–5. [Google Scholar] [CrossRef] [PubMed]
- Singh, BN; et al. Green tea catechin EGCG. Life Sci. 2011, 88, 107–25. [Google Scholar]
- Vara-Messler, M; et al. Mfn2-mediated fusion by quercetin. Mol Nutr Food Res. 2021, 65, 2000851. [Google Scholar]
- Li, Y; et al. Quercetin, inflammation and immunity. Nutrients 2016, 8, 167. [Google Scholar] [CrossRef]
- Chen, H; Chan, DC. Mitochondrial dynamics in mammals. Curr Top Dev Biol. 2004, 59, 119–44. [Google Scholar]
- Shock, LS; et al. DNA methyltransferase 1 in mitochondria. PNAS 2011, 108, 3633–5. [Google Scholar] [CrossRef]
- Iacobazzi, V; et al. Mitochondrial DNA methylation targets. Int J Mol Sci. 2022, 23, 3098. [Google Scholar]
- Fang, MZ; et al. Dietary constituents as epigenetic modulators. J Nutr. 2007, 137, 223S. [Google Scholar] [CrossRef]
- Myzak, MC; et al. Sulforaphane inhibits HDAC. Exp Biol Med. 2006, 231, 1700–7. [Google Scholar]
- Dashwood, RH; Ho, E. Dietary HDAC inhibitors. Semin Cancer Biol. 2007, 17, 363–9. [Google Scholar] [CrossRef]
- Li, Y; et al. EGCG inhibits pancreatic cancer stem cells. Cancer Lett. 2011, 301, 249–58. [Google Scholar]
- Sanna, V; et al. Nanoencapsulation of EGCG. Nanomedicine 2013, 9, 392–405. [Google Scholar]
- Wiechert, W. 13C metabolic flux analysis. Metab Eng. 2001, 3, 195–206. [Google Scholar] [CrossRef] [PubMed]
- Caro, P; et al. Metabolic signatures in lymphoma. Cancer Cell. 2012, 22, 547–60. [Google Scholar] [CrossRef]
- Thimmulappa, RK; et al. Nrf2 is a critical regulator. J Clin Invest. 2006, 116, 984–95. [Google Scholar] [CrossRef] [PubMed]
- Clarke, JD; et al. Sulforaphane and cancer prevention. J Agric Food Chem. 2011, 59, 1099–105. [Google Scholar]
- Kikuno, N; et al. Genistein and DNMT inhibition. Cancer Res. 2008, 68, 7513–20. [Google Scholar]
- Butow, RA; Avadhani, NG. Mitochondrial to nuclear retrograde signaling. Mol Cell. 2004, 14, 1–15. [Google Scholar] [CrossRef]
- Shadel, GS; Horvath, TL. Mitochondrial ROS signaling in organismal homeostasis. Cell. 2015, 163, 560–9. [Google Scholar] [CrossRef] [PubMed]
- Wellen, KE; et al. ATP-citrate lyase links metabolism to histone acetylation. Science 2009, 324, 1076–80. [Google Scholar] [CrossRef]
- Hirschey, MD; et al. SIRT3 regulates mitochondrial fatty-acid oxidation. Nature 2010, 464, 121–5. [Google Scholar] [CrossRef] [PubMed]
- Finley, LW; et al. SIRT3 opposes the Warburg effect. Cancer Cell. 2011, 19, 416–28. [Google Scholar] [CrossRef]
- Manach, C; et al. Polyphenols: Food sources and bioavailability. Am J Clin Nutr. 2004, 79, 727–47. [Google Scholar] [CrossRef]
- Yallapu, MM; et al. Nano-curcumin for cancer therapy. Cancer Res. 2010, 70, 200. [Google Scholar]
- Sales, NM; et al. Nutrigenomics: Definitions and advances of this new science. Nutr Clin Metab. 2014, 28, 37–44. [Google Scholar] [CrossRef]
| Strategy | Bioactive Agent | Molecular Target | Mitochondrial/Oncogenic Outcome | Ref |
|---|---|---|---|---|
| Hit 1 | Curcumin | mTORC1 / HKII | Dissociates HKII from VDAC; initiates apoptosis | [15,24] |
| Hit 1 | Berberine | AMPK / Complex I | Activates energy-sensing; inhibits GFS growth | [26,28] |
| Hit 2 | Quercetin | SIRT1 / Mfn2 | Promotes mitochondrial fusion and quality | [30,34] |
| Hit 2 | EGCG | Drp1 (Ser616) | Inhibits excessive fission; reduces ROS leak | [31,33] |
| Hit 3 | Sulforaphane | HDAC / DNMT1 | De-methylates D-loop; restores OXPHOS | [40,41] |
| Hit 3 | Genistein | DNMTs | Reverses metabolic silencing of mtDNA | [38,42] |
| Challenge | Mechanistic Basis | Proposed Precision Solution | Ref |
|---|---|---|---|
| Bioavailability | Rapid glucuronidation | Nano-nutrigenomic encapsulation | [48,49] |
| Resistance | Metabolic Plasticity | Dual targeting of IGF-1R and OXPHOS | [19,23] |
| Retrograde Flux | Acetyl-CoA buildup | SIRT3 activators to clear nuclear substrates | [46,47] |
| Heterogeneity | “Mitotype” variation | Nutri-phenotyping via Liquid Biopsy | [50] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




