Submitted:
08 February 2026
Posted:
09 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
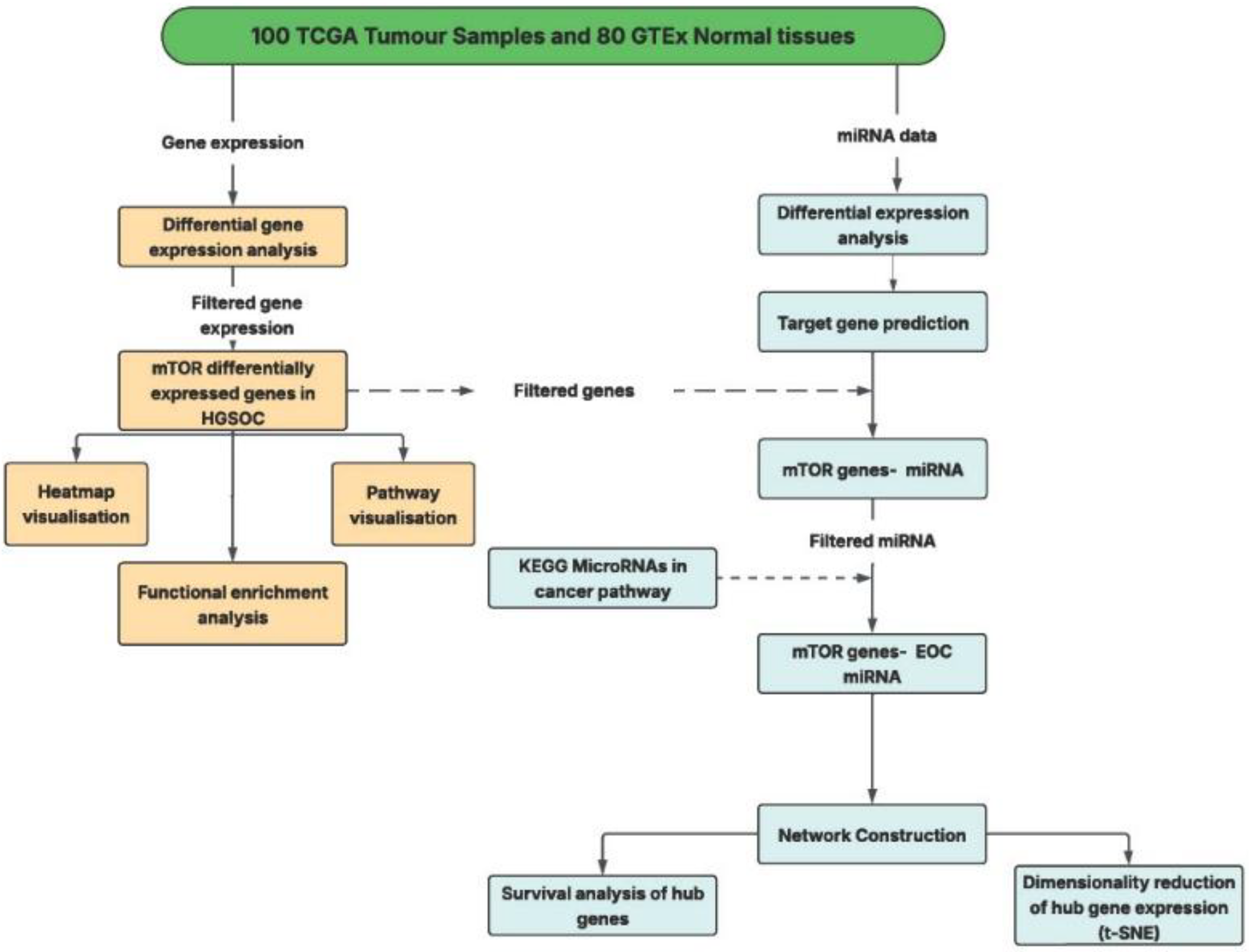
2. Materials and Methods
2.1. Dataset Selection
2.2. Data Processing
2.3. Differential Expression Analysis
2.4. Transcriptomic Analysis
2.5. Prediction and Integration of miRNA–mRNA Regulatory Interactions
2.6. Network Construction and Topological Analysis of mTOR Pathway miRNA–mRNA Interactions
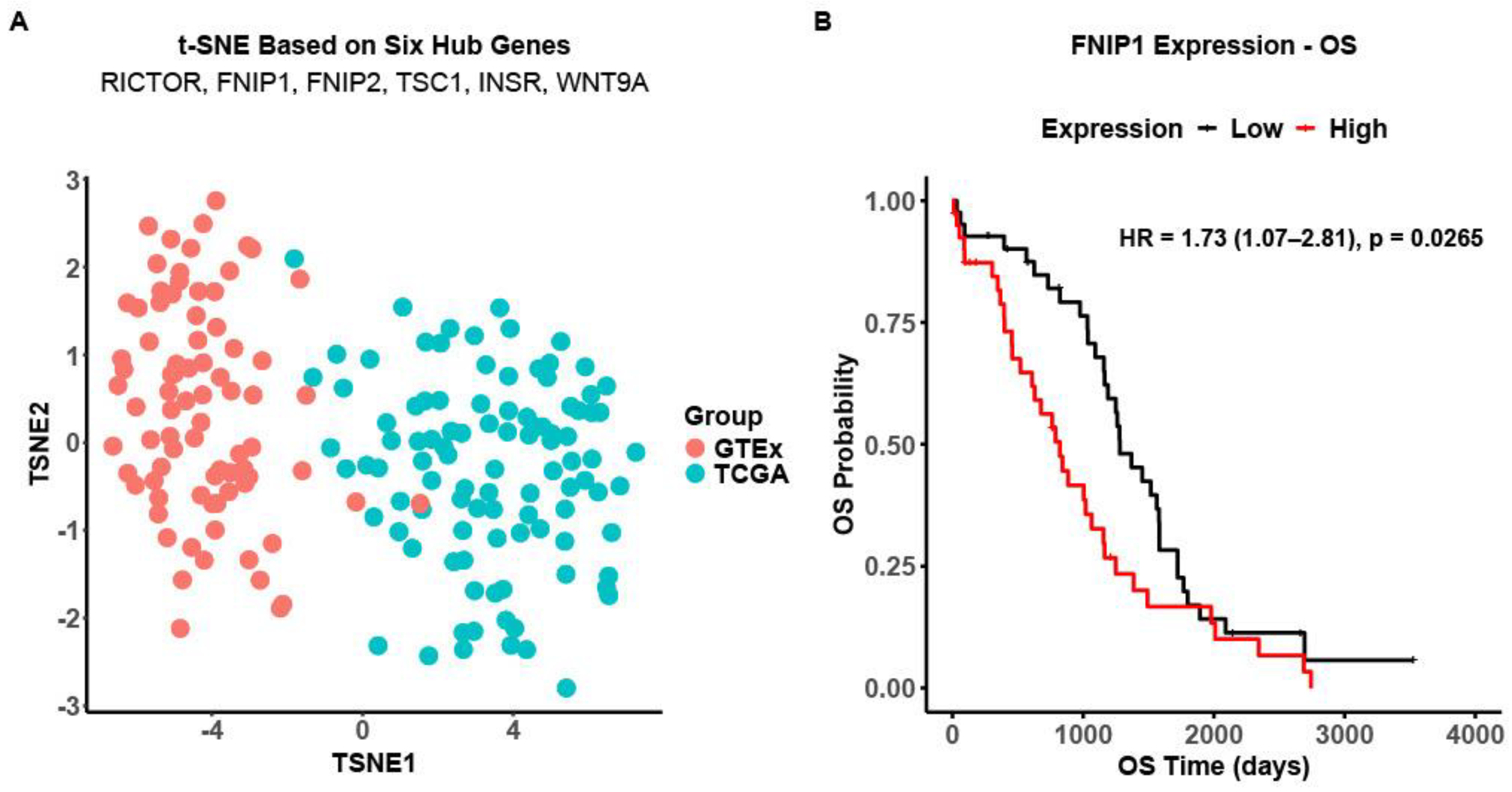
2.7. Survival Analysis
2.8. Dimensionality Reduction of Hub Gene Expression Analysis
3. Results
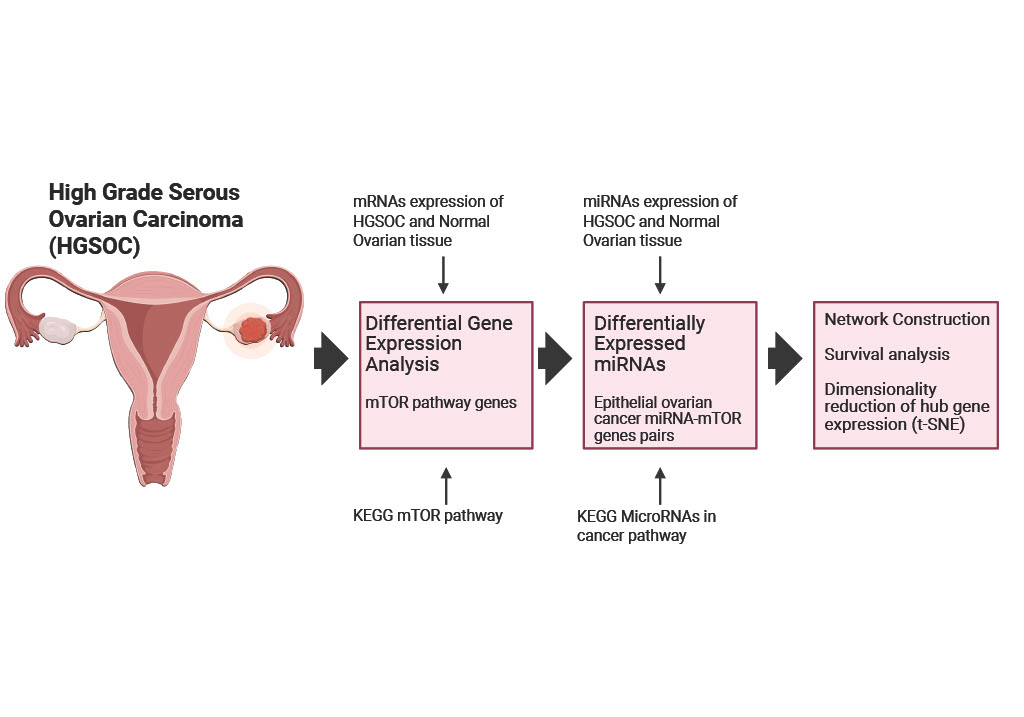
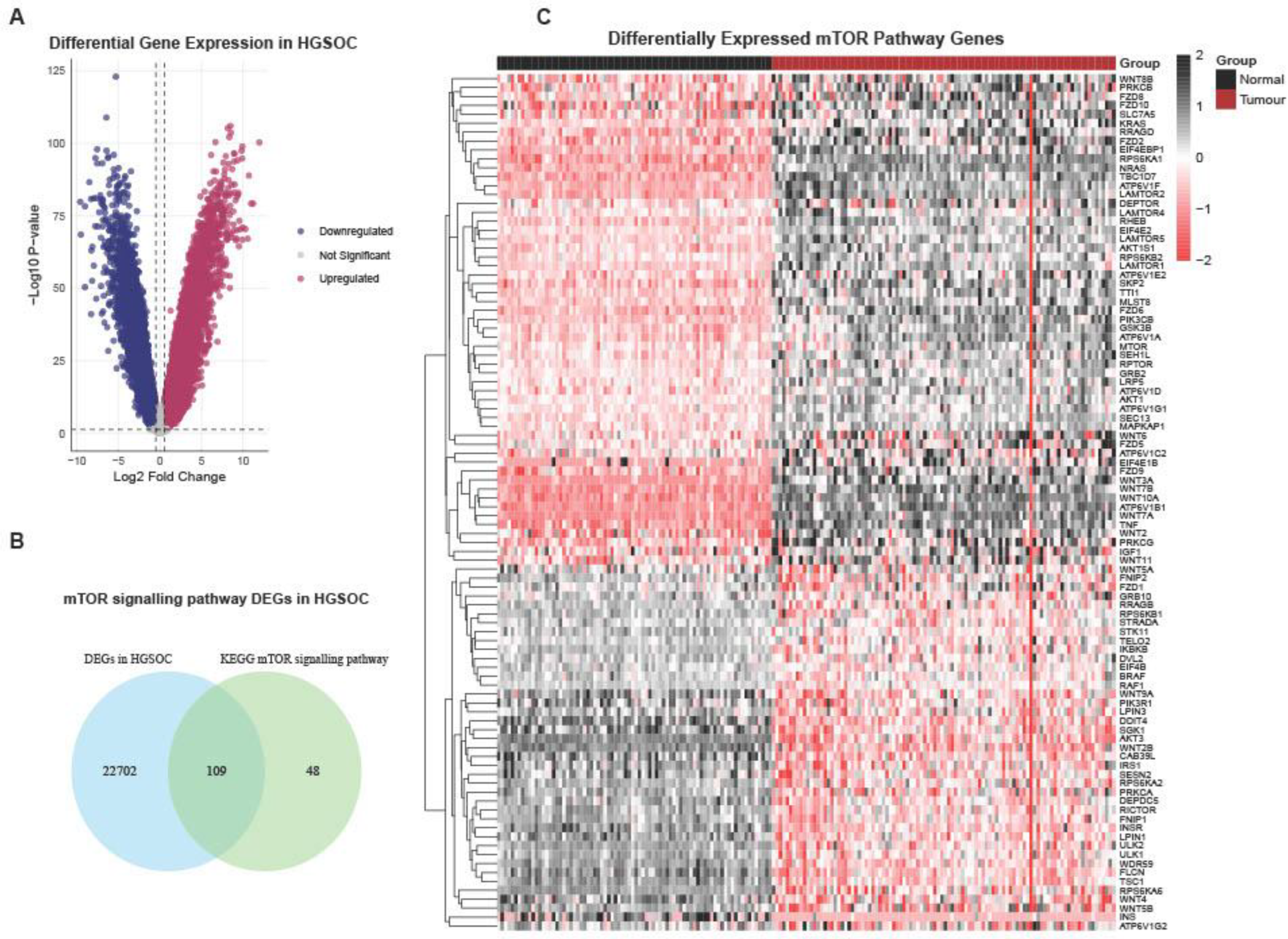
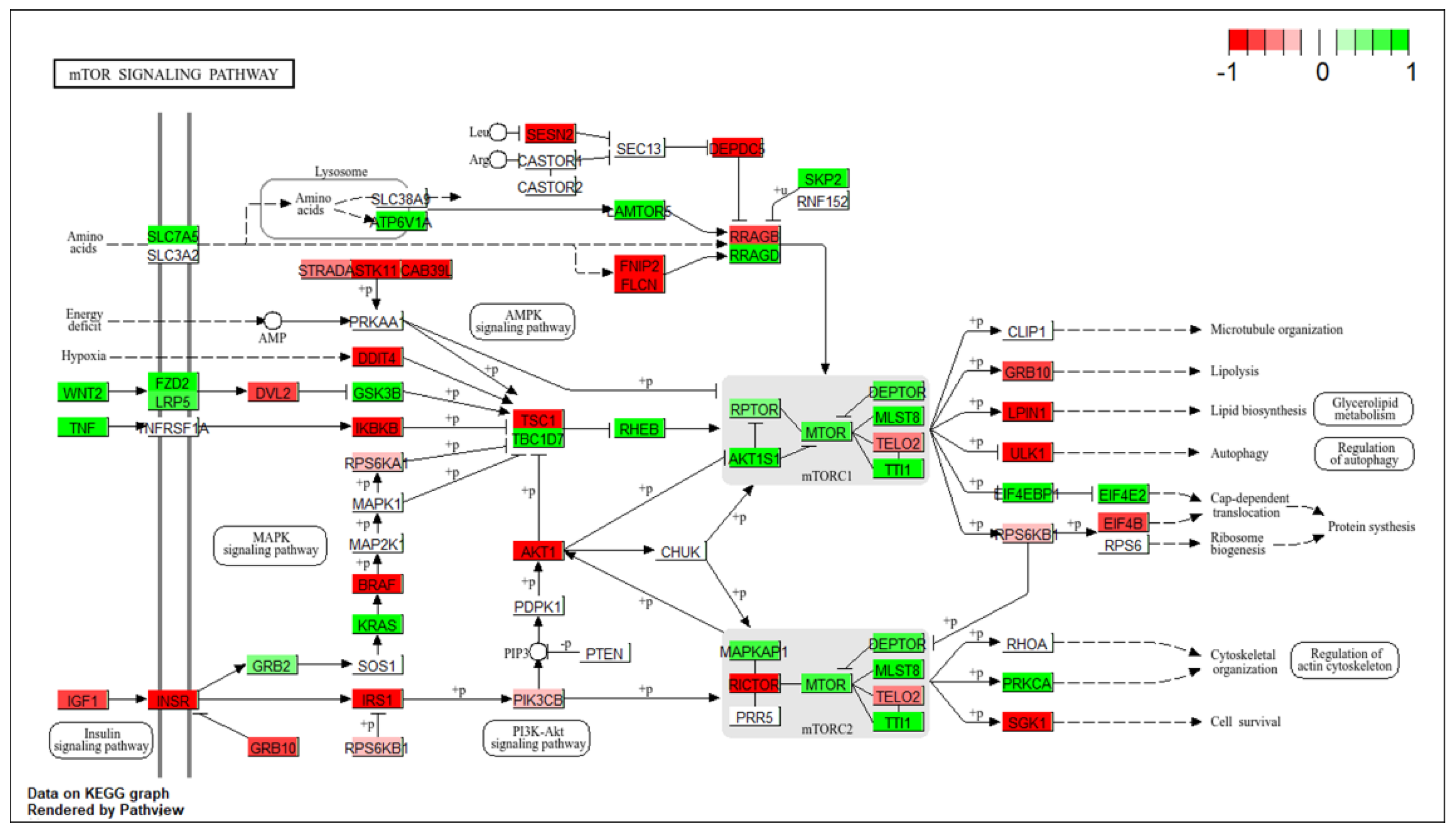
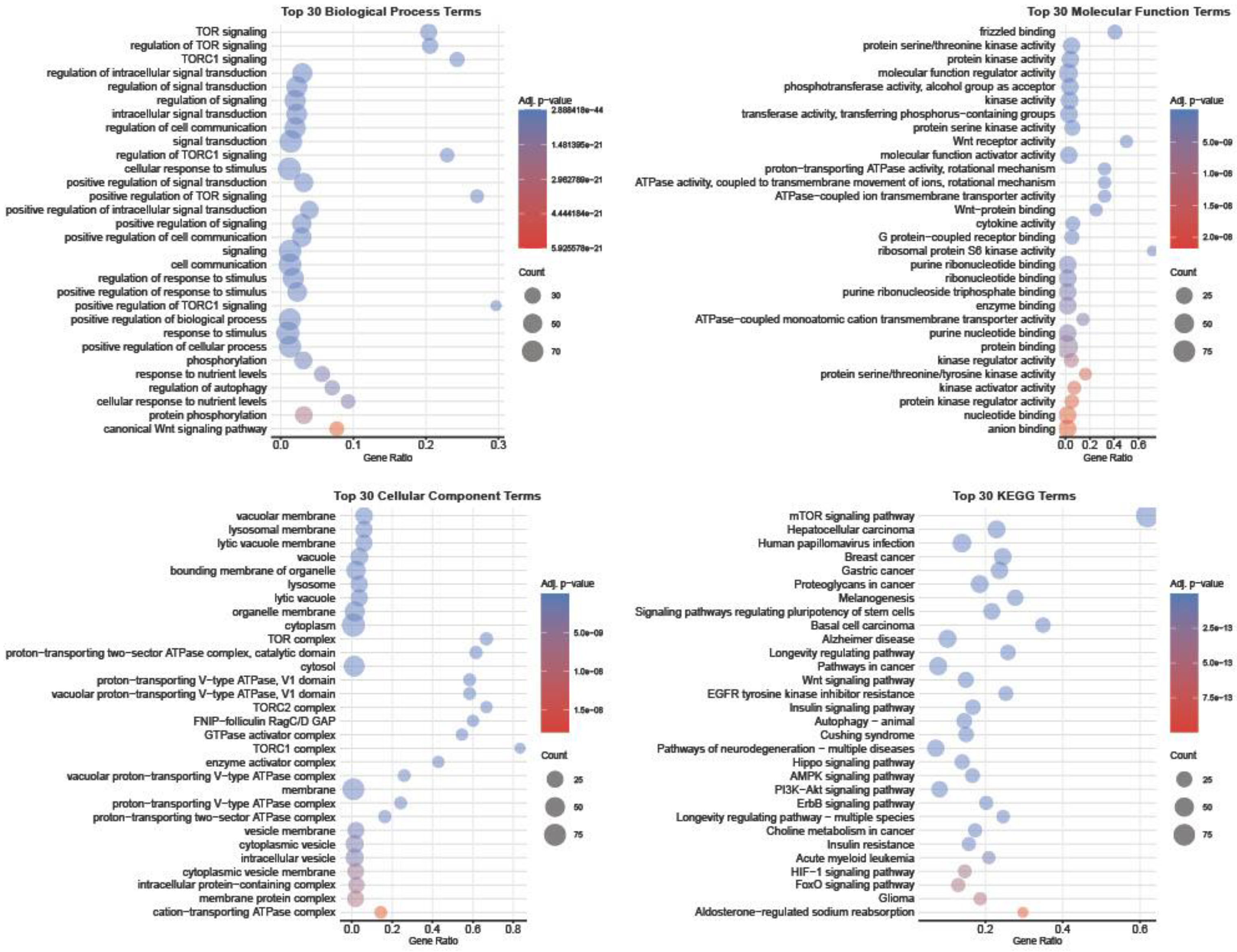
3.1. Transcriptomic Profiling of mTOR Pathway in High-Grade Serous Ovarian Cancer: A Pathway-Level Analysis
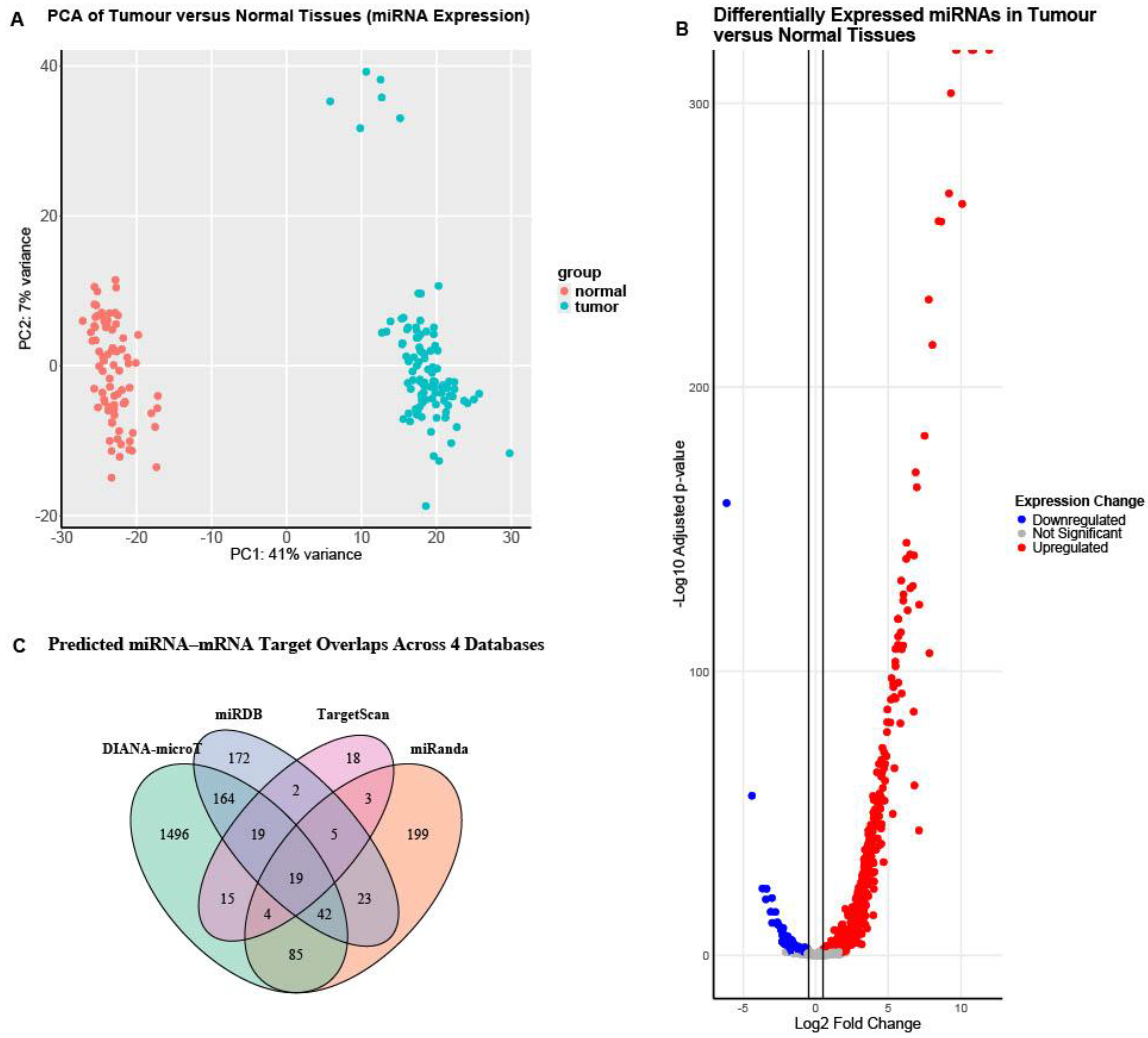
3.2. The Regulatory miRNA Network of mTOR Pathway Genes in High-Grade Serous Ovarian Cancer
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- International Agency for Research on Cancer GLOBAL CANCER OBSERVATORY. Ovary-Fact-Sheet. Available online: https://gco.iarc.who.int/media/globocan/factsheets/cancers/25-ovary-fact-sheet.pdf (accessed on 3 October 2025).
- De Leo, A.; Santini, D.; Ceccarelli, C.; Santandrea, G.; Palicelli, A.; Acquaviva, G.; Chiarucci, F.; Rosini, F.; Ravegnini, G.; Pession, A.; et al. What Is New on Ovarian Carcinoma: Integrated Morphologic and Molecular Analysis Following the New 2020 World Health Organization Classification of Female Genital Tumors. Diagnostics (Basel). 2021, 11. [Google Scholar] [CrossRef]
- Luvero, D.; Plotti, F.; Aloisia, A.; Montera, R.; Terranova; De Cicco Nardone, C.; Carlo; Scaletta, G.; Lopez, S.; Miranda, A.; Capriglione, S.; et al. Ovarian Cancer Relapse: From the Latest Scientific Evidence to the Best Practice. Crit. Rev. Oncol. Hematol. 2019, 140, 28–38. [Google Scholar] [CrossRef]
- Hablase, R.; Kyrou, I.; Randeva, H.; Karteris, E.; Chatterjee, J. The “Road” to Malignant Transformation from Endometriosis to Endometriosis-Associated Ovarian Cancers (EAOCs): An MTOR-Centred Review. Cancers (Basel). 2024, 16, 2160. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, R.; Wei, L.; Huang, J. MTOR Signaling, Function, Novel Inhibitors, and Therapeutic Targets. J. Nucl. Med. 2011, 52, 497–500. [Google Scholar] [CrossRef]
- Stuttfeld, E.; Aylett, C.H.S.; Imseng, S.; Boehringer, D.; Scaiola, A.; Sauer, E.; Hall, M.N.; Maier, T.; Ban, N. Architecture of the Human MTORC2 Core Complex. Elife 2018, 7. [Google Scholar] [CrossRef]
- Yip, C.K.; Murata, K.; Walz, T.; Sabatini, D.M.; Kang, S.A. Structure of the Human MTOR Complex I and Its Implications for Rapamycin Inhibition. Mol. Cell 2010, 38, 768–774. [Google Scholar] [CrossRef]
- Laplante, M.; Sabatini, D.M. MTOR Signaling in Growth Control and Disease. Cell 2012, 149, 274–293. [Google Scholar] [CrossRef] [PubMed]
- Rogers-Broadway, K.R.; Kumar, J.; Sisu, C.; Wander, G.; Mazey, E.; Jeyaneethi, J.; Pados, G.; Tsolakidis, D.; Klonos, E.; Grunt, T.; et al. Differential Expression of MTOR Components in Endometriosis and Ovarian Cancer: Effects of Rapalogues and Dual Kinase Inhibitors on MTORC1 and MTORC2 Stoichiometry. Int. J. Mol. Med. 2018, 43, 47. [Google Scholar] [CrossRef]
- Differential Expression of MTOR Signalling Components in Drug Resistance in Ovarian Cancer - PubMed. Available online: https://pubmed.ncbi.nlm.nih.gov/20944133/ (accessed on 15 December 2025).
- Altomare, D.A.; Hui, Q.W.; Skele, K.L.; De Rienzo, A.; Klein-Szanto, A.J.; Godwin, A.K.; Testa, J.R. AKT and MTOR Phosphorylation Is Frequently Detected in Ovarian Cancer and Can Be Targeted to Disrupt Ovarian Tumor Cell Growth. Oncogene KWRD. 2004, 23, 5853–5857. [Google Scholar] [CrossRef] [PubMed]
- Tang, L.; Bian, C. Research Progress in Endometriosis-Associated Ovarian Cancer. Front. Oncol. 2024, 14, 1381244. [Google Scholar] [CrossRef]
- Rinne, N.; Christie, E.L.; Ardasheva, A.; Kwok, C.H.; Demchenko, N.; Low, C.; Tralau-Stewart, C.; Fotopoulou, C.; Cunnea, P. Targeting the PI3K/AKT/MTOR Pathway in Epithelial Ovarian Cancer, Therapeutic Treatment Options for Platinum-Resistant Ovarian Cancer. Cancer Drug Resistance 2021, 4, 573. [Google Scholar] [CrossRef]
- Verhaak, R.G.W.; Tamayo, P.; Yang, J.Y.; Hubbard, D.; Zhang, H.; Creighton, C.J.; Fereday, S.; Lawrence, M.; Carter, S.L.; Mermel, C.H.; et al. Prognostically Relevant Gene Signatures of High-Grade Serous Ovarian Carcinoma. J. Clin. Invest. 2012, 123, 517. [Google Scholar] [CrossRef]
- GTEx Portal. Available online: https://www.gtexportal.org/home/ (accessed on 17 January 2026).
- GDC Data Portal Homepage. Available online: https://portal.gdc.cancer.gov/ (accessed on 17 January 2026).
- Beddows, I.; Fan, H.; Heinze, K.; Johnson, B.K.; Leonova, A.; Senz, J.; Djirackor, S.; Cho, K.R.; Pearce, C.L.; Huntsman, D.G.; et al. Cell State of Origin Impacts Development of Distinct Endometriosis-Related Ovarian Carcinoma Histotypes. Cancer Res. 2024, 184, 26–38. [Google Scholar] [CrossRef] [PubMed]
- Hashemi Sheikhshabani, S.; Amini-Farsani, Z.; Kazemifard, N.; Modarres, P.; Amini-Farsani, Z.; Omrani, M.D.; Ghafouri-Fard, S. Meta-Analysis of Microarray Data to Determine Gene Indicators Involved in the Cisplatin Resistance in Ovarian Cancer. In Cancer Rep.; PAGE:STRING:ARTICLE/CHAPTER, 2023; Volume 6, p. e1884. [Google Scholar] [CrossRef]
- Vivian, J.; Rao, A.A.; Nothaft, F.A.; Ketchum, C.; Armstrong, J.; Novak, A.; Pfeil, J.; Narkizian, J.; Deran, A.D.; Musselman-Brown, A.; et al. Toil Enables Reproducible, Open Source, Big Biomedical Data Analyses. Nat. Biotechnol. 2017, 35, 314. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated Estimation of Fold Change and Dispersion for RNA-Seq Data with DESeq2. Genome Biol. 2014, 15. [Google Scholar] [CrossRef]
- Kanehisa, M. Toward Understanding the Origin and Evolution of Cellular Organisms. Protein Sci. 2019, 28, 1947–1951. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef] [PubMed]
- Luo, W.; Brouwer, C. Pathview: An R/Bioconductor Package for Pathway-Based Data Integration and Visualization. Bioinformatics 2013, 29, 1830–1831. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Wang, X. MiRDB: An Online Database for Prediction of Functional MicroRNA Targets. Nucleic Acids Res. 2020, 48, D127–D131. [Google Scholar] [CrossRef]
- Ru, Y.; Kechris, K.J.; Tabakoff, B.; Hoffman, P.; Radcliffe, R.A.; Bowler, R.; Mahaffey, S.; Rossi, S.; Calin, G.A.; Bemis, L.; et al. The MultiMiR R Package and Database: Integration of MicroRNA–Target Interactions along with Their Disease and Drug Associations. Nucleic Acids Res. 2014, 42, e133–e133. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 2003, 13, 2498. [Google Scholar] [CrossRef]
- Agarwal, V.; Bell, G.W.; Nam, J.W.; Bartel, D.P. Predicting Effective MicroRNA Target Sites in Mammalian MRNAs. Elife 2015, 4. [Google Scholar] [CrossRef] [PubMed]
- John, B.; Enright, A.J.; Aravin, A.; Tuschl, T.; Sander, C.; Marks, D.S. Human MicroRNA Targets. PLoS Biol. 2004, 2. [Google Scholar] [CrossRef]
- Paraskevopoulou, M.D.; Georgakilas, G.; Kostoulas, N.; Vlachos, I.S.; Vergoulis, T.; Reczko, M.; Filippidis, C.; Dalamagas, T.; Hatzigeorgiou, A.G. DIANA-MicroT Web Server v5.0: Service Integration into MiRNA Functional Analysis Workflows. Nucleic Acids Res. 2013, 41. [Google Scholar] [CrossRef] [PubMed]
- Gasparri, M.L.; Bardhi, E.; Ruscito, I.; Papadia, A.; Farooqi, A.A.; Marchetti, C.; Bogani, G.; Ceccacci, I.; Mueller, M.D.; Benedetti Panici, P. PI3K/AKT/MTOR Pathway in Ovarian Cancer Treatment: Are We on the Right Track? Geburtshilfe Frauenheilkd. 2017, 77, 1095. [Google Scholar] [CrossRef]
- Rozengurt, E.; Soares, H.P.; Sinnet-Smith, J. Suppression of Feedback Loops Mediated by PI3K/MTOR Induces Multiple over-Activation of Compensatory Pathways: An Unintended Consequence Leading to Drug Resistance. Mol. Cancer Ther. 2014, 13, 2477. [Google Scholar] [CrossRef]
- King, E.R.; Zu, Z.; Tsang, Y.T.M.; Deavers, M.T.; Malpica, A.; Mok, S.C.; Gershenson, D.M.; Wong, K.K. The Insulin-like Growth Factor 1 Pathway Is a Potential Therapeutic Target for Low-Grade Serous Ovarian Carcinoma. Gynecol. Oncol. 2011, 123, 13. [Google Scholar] [CrossRef]
- Montero, J.C.; Chen, X.; Ocaña, A.; Pandiella, A. Predominance of MTORC1 over MTORC2 in the Regulation of Proliferation of Ovarian Cancer Cells: Therapeutic Implications. Mol. Cancer Ther. 2012, 11, 1342–1352. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhang, X.; Tang, W.; Lin, Z.; Xu, L.; Dong, R.; Li, Y.; Li, J.; Zhang, Z.; Li, X.; et al. MiR-130a Upregulates MTOR Pathway by Targeting TSC1 and Is Transactivated by NF-ΚB in High-Grade Serous Ovarian Carcinoma. Cell Death & Differentiation 2017 2017, 24 24, 2089–2100. [Google Scholar] [CrossRef]
- Glibo, M.; Serman, A.; Karin-Kujundzic, V.; Vlatkovic, I.B.; Miskovic, B.; Vranic, S.; Serman, L. The Role of Glycogen Synthase Kinase 3 (GSK3) in Cancer with Emphasis on Ovarian Cancer Development and Progression: A Comprehensive Review. Bosn. J. Basic Med. Sci. 2021, 21, 5. [Google Scholar] [CrossRef]
- Nicklin, P.; Bergman, P.; Zhang, B.; Triantafellow, E.; Wang, H.; Nyfeler, B.; Yang, H.; Hild, M.; Kung, C.; Wilson, C.; et al. Bidirectional Transport of Amino Acids Regulates MTOR and Autophagy. Cell 2009, 136, 521–534. [Google Scholar] [CrossRef]
- Zoncu, R.; Bar-Peled, L.; Efeyan, A.; Wang, S.; Sancak, Y.; Sabatini, D.M. MTORC1 Senses Lysosomal Amino Acids through an Inside-out Mechanism That Requires the Vacuolar H(+)-ATPase. Science 2011, 334, 678–683. [Google Scholar] [CrossRef]
- Lawrence, R.E.; Cho, K.F.; Rappold, R.; Thrun, A.; Tofaute, M.; Kim, D.J.; Moldavski, O.; Hurley, J.H.; Zoncu, R. A Nutrient-Induced Affinity Switch Controls MTORC1 Activation by Its Rag GTPase-Ragulator Lysosomal Scaffold. Nat. Cell Biol. 2018, 20, 1052–1063. [Google Scholar] [CrossRef]
- Bar-Peled, L.; Chantranupong, L.; Cherniack, A.D.; Chen, W.W.; Ottina, K.A.; Grabiner, B.C.; Spear, E.D.; Carter, S.L.; Meyerson, M.; Sabatini, D.M. A Tumor Suppressor Complex with GAP Activity for the Rag GTPases That Signal Amino Acid Sufficiency to MTORC1. Science (1979). 2013, 340, 1100–1106. [Google Scholar] [CrossRef]
- Hasumi, H.; Baba, M.; Hasumi, Y.; Lang, M.; Huang, Y.; Oh, H.B.F.; Matsuo, M.; Merino, M.J.; Yao, M.; Ito, Y.; et al. Folliculin-Interacting Proteins Fnip1 and Fnip2 Play Critical Roles in Kidney Tumor Suppression in Cooperation with Flcn. Proc. Natl. Acad. Sci. U. S. A. 2015, 112, E1624–E1631. [Google Scholar] [CrossRef]
- Custode, B.M.; Annunziata, F.; Dos Santos Matos, F.; Schiano, V.; Maffia, V.; Lillo, M.; Colonna, R.; De Cegli, R.; Ballabio, A.; Pastore, N. Folliculin Depletion Results in Liver Cell Damage and Cholangiocarcinoma through MiT/TFE Activation. Cell Death & Differentiation 2025 32:8 2025, 32, 1460–1472. [Google Scholar] [CrossRef]
- Napolitano, G.; Di Malta, C.; Esposito, A.; de Araujo, M.E.G.; Pece, S.; Bertalot, G.; Matarese, M.; Benedetti, V.; Zampelli, A.; Stasyk, T.; et al. A Substrate-Specific MTORC1 Pathway Underlies Birt-Hogg-Dubé Syndrome. Nature 2020, 585, 597–602. [Google Scholar] [CrossRef]
- Di Malta, C.; Siciliano, D.; Calcagni, A.; Monfregola, J.; Punzi, S.; Pastore, N.; Eastes, A.N.; Davis, O.; De Cegli, R.; Zampelli, A.; et al. TRANSCRIPTIONAL ACTIVATION OF RAGD GTPASE CONTROLS MTORC1 AND PROMOTES CANCER GROWTH. Science 2017, 356, 1188. [Google Scholar] [CrossRef] [PubMed]
- Ramirez Reyes, J.M.J.; Cuesta, R.; Pause, A. Folliculin: A Regulator of Transcription Through AMPK and MTOR Signaling Pathways. Front. Cell Dev. Biol. 2021, 9, 667311. [Google Scholar] [CrossRef] [PubMed]
- He, L.; Hannon, G.J. MicroRNAs: Small RNAs with a Big Role in Gene Regulation. Nature Reviews Genetics 2004, 5 5, 522–531. [Google Scholar] [CrossRef]
- Roush, S.; Slack, F.J. The Let-7 Family of MicroRNAs. Trends Cell Biol. 2008, 18, 505–516. [Google Scholar] [CrossRef] [PubMed]
- Chirshev, E.; Oberg, K.C.; Ioffe, Y.J.; Unternaehrer, J.J. Let-7 as Biomarker, Prognostic Indicator, and Therapy for Precision Medicine in Cancer. Clin. Transl. Med. 2019, 8, 24. [Google Scholar] [CrossRef]
- Fan, H.; Jiang, M.; Li, B.; He, Y.; Huang, C.; Luo, D.; Xu, H.; Yang, L.; Zhou, J. MicroRNA-Let-7a Regulates Cell Autophagy by Targeting Rictor in Gastric Cancer Cell Lines MGC-803 and SGC-7901. Oncol. Rep. 2018, 39, 1207–1214. [Google Scholar] [CrossRef]
- Jiang, S. A Regulator of Metabolic Reprogramming: MicroRNA Let-7. Transl. Oncol. 2019, 12, 1005–1013. [Google Scholar] [CrossRef]
- Li, C.H.; Liao, C.C. The Metabolism Reprogramming of MicroRNA Let-7-Mediated Glycolysis Contributes to Autophagy and Tumor Progression. Int. J. Mol. Sci. 2021, 23, 113. [Google Scholar] [CrossRef] [PubMed]
- Frederick, M.I.; Siddika, T.; Zhang, P.; Balasuriya, N.; Turk, M.A.; O’donoghue, P.; Heinemann, I.U. MiRNA-Dependent Regulation of AKT1 Phosphorylation. Cells 2022, 11, 821. [Google Scholar] [CrossRef] [PubMed]
- Expression of FNIP1 in Cancer - Summary - The Human Protein Atlas. Available online: https://www.proteinatlas.org/ENSG00000217128-FNIP1/cancer (accessed on 9 November 2025).
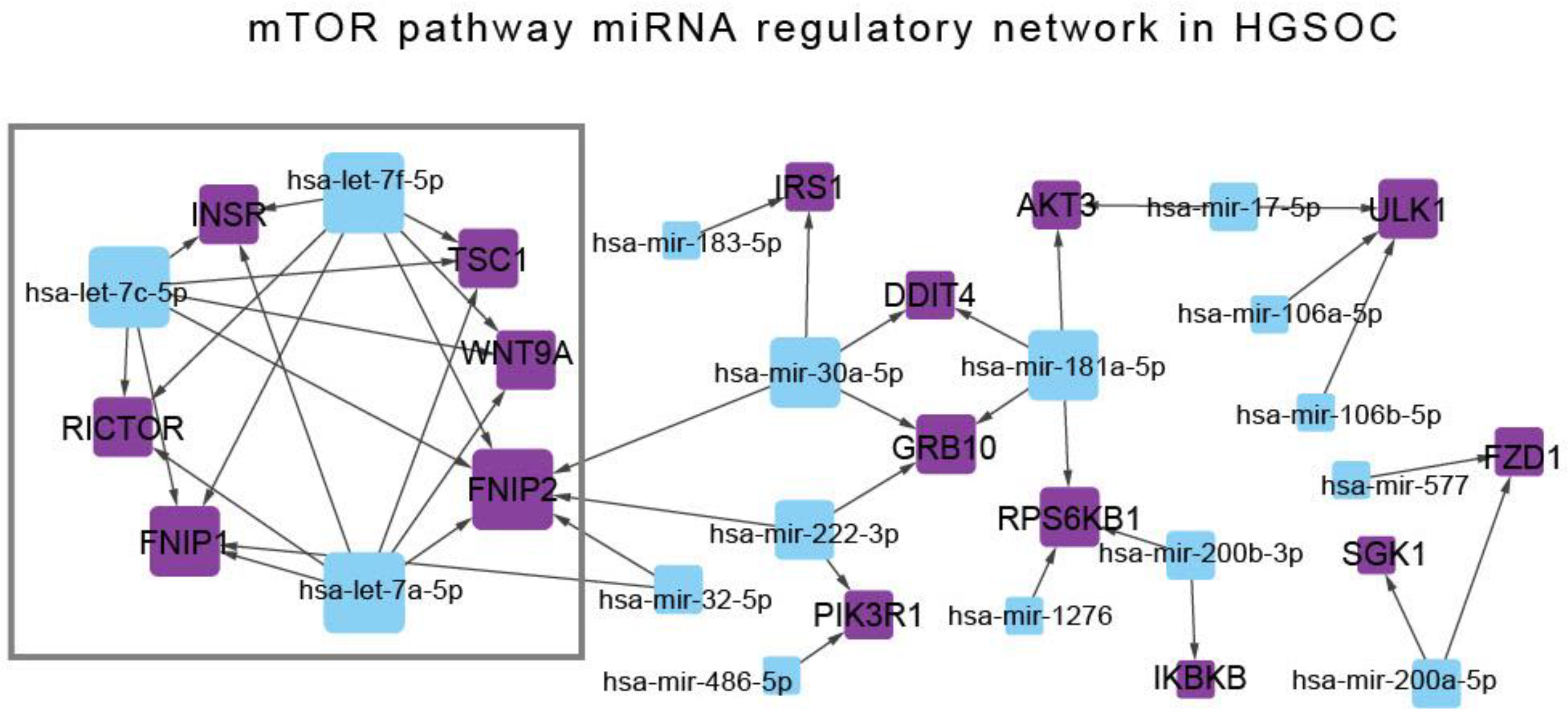
| Node | Node Type | Degree centrality |
|---|---|---|
| hsa-let-7a-5p | miRNA | 6 |
| hsa-let-7c-5p | miRNA | 6 |
| hsa-let-7f-5p | miRNA | 6 |
| hsa-mir-181a-5p | miRNA | 4 |
| hsa-mir-30a-5p | miRNA | 4 |
| hsa-mir-222-3p | miRNA | 3 |
| FNIP2 | mRNA | 6 |
| FNIP1 | mRNA | 4 |
| GRb10 | mRNA | 3 |
| INSR | mRNA | 3 |
| RICTOR | mRNA | 3 |
| RBS6KB1 | mRNA | 3 |
| TSC1 | mRNA | 3 |
| ULK1 | mRNA | 3 |
| WNT9A | mRNA | 3 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




