Submitted:
06 February 2026
Posted:
09 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
3. Results
4. Discussion
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Lее, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar] [CrossRef] [PubMed]
- Mehanna, E.T; Ghattas, M.H.; Mesbah, N.M.; Saleh, S.M.; Abo-Elmatty, D.M. Association of MicroRNA-146a rs2910164 Gene Polymorphism with Metabolic Syndrome. Folia Biol. 2015, 61(1), 43–48. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Yan, Y.; Xv, W.; Qian, G.; Li, C.; Zou, H.; Li, Y. A New Insight into the Roles of MiRNAs in Metabolic Syndrome. Biomed Res Int. 2018, 2018, 7372636. [Google Scholar] [CrossRef] [PubMed]
- Vorobeva, E.V.; Kyyaly, M.A.; Sones, C.L.; He, P.J.W.; Arshad, S.H.; Sanchez-Elsner, T.; Kurukulaaratchy, R.J. Circulating microRNAs as Potential Diagnostic Tools for Asthma and for Indicating Severe Asthma Risk. Int. J. Mol. Sci. 2025, 26(14), 6676. [Google Scholar] [CrossRef] [PubMed]
- Stevanovic, J.; Mitic, N.; Penezic, A.; Radojicic, O.; Ardalic, D.; Mandic, M.; Mandic-Markovic, V.; Mikovic, Ž.; Brkušanin, M.; Nedic, O.; et al. Upregulation of the Antioxidant Response-Related microRNAs miR-146a-5p and miR-21-5p in Gestational Diabetes: An Analysis of Matched Samples of Extracellular Vesicles and PBMCs. Int. J. Mol. Sci. 2025, 26(14), 6902. [Google Scholar] [CrossRef] [PubMed]
- Khalyfa, A.; Verma, M.; Alexander, M.M.; Qiao, Z.; Rood, T.; Kapoor, R.; Joshi, T.; Gozal, D.; Francisco, B.D. Childhood Asthma Biomarkers Derived from Plasma and Saliva Exosomal miRNAs. Int. J. Mol. Sci. 2025, 26(15), 7043. [Google Scholar] [CrossRef] [PubMed]
- Dobre, M.; Manuc, T.E.; Manuc, M.; Matei, I.-C.; Dobre, A.-M.; Dragne, A.-D.; Maffioletti, E.; Pelisenco, I.A.; Milanesi, E. Circulating miRNA Profile in Inflammatory Bowel Disease Patients with Stress, Anxiety, and Depression. Int. J. Mol. Sci. 2025, 26(15), 7321. [Google Scholar] [CrossRef] [PubMed]
- Ivashchenko, A.; Berillo, O.; Pyrkova, A.; Niyazova, R.; Atambayeva, S. MiR-3960 binding sites with mRNA of human genes. Bioinformation 2014, 10(7), 423–427. [Google Scholar] [CrossRef] [PubMed]
- Kool, E.T. Hydrogen bonding, base stacking, and steric effects in DNA replication. Annu. Rev. Biophys. Biomol. Struct. 2001, 30, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Leontis, N.B.; Stombaugh, J.; Westhof, E. The Non-watson-crick Base Pairs and their associated isostericity matrices. Nucleic Acids Res. 2002, 30(16), 3497–3531. [Google Scholar] [CrossRef] [PubMed]
- Davis, E.; Caiment, F.; Tordoir, X.; Cavaillé, J.; Ferguson-Smith, A.; Cockett, N.; Georges, M.; Charlier, C. RNAi-mediated allelic trans-interaction at the imprinted Rtl1/ Peg11 locus. Curr. Biol. 2005, 15(8), 743–749. [Google Scholar] [CrossRef] [PubMed]
- Garg, A.; Heinemann, U.A. Novel form of RNA double helix based on G·U and C·A+ wobble base pairing. RNA 2018, 24(2), 209–218. [Google Scholar] [CrossRef] [PubMed]
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S. A.; Driver, S. E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998, 391(6669), 806–811. [Google Scholar] [CrossRef] [PubMed]
- Atambayeva, S.; Niyazova, R.; Ivashchenko, A.; Pyrkova, A.; Pinsky, I.; Akimniyazova, A.; Labeit, S. The binding sites of miR-619-5p in the mRNAs of human and orthologous genes. BMC Genomics 2017, 18, 428. [Google Scholar] [CrossRef] [PubMed]
- Yurikova, O.Yu.; Aisina, D.E.; Niyazova, R.E.; Atambayeva, S.A.; Labeit, S.; Ivashchenko, A.T. The interaction of miR-5p and miR-3p with the mRNAs of orthologous genes. Mol. Biol. (Mosk) 2019, 53(4), 692–704. [Google Scholar] [CrossRef] [PubMed]
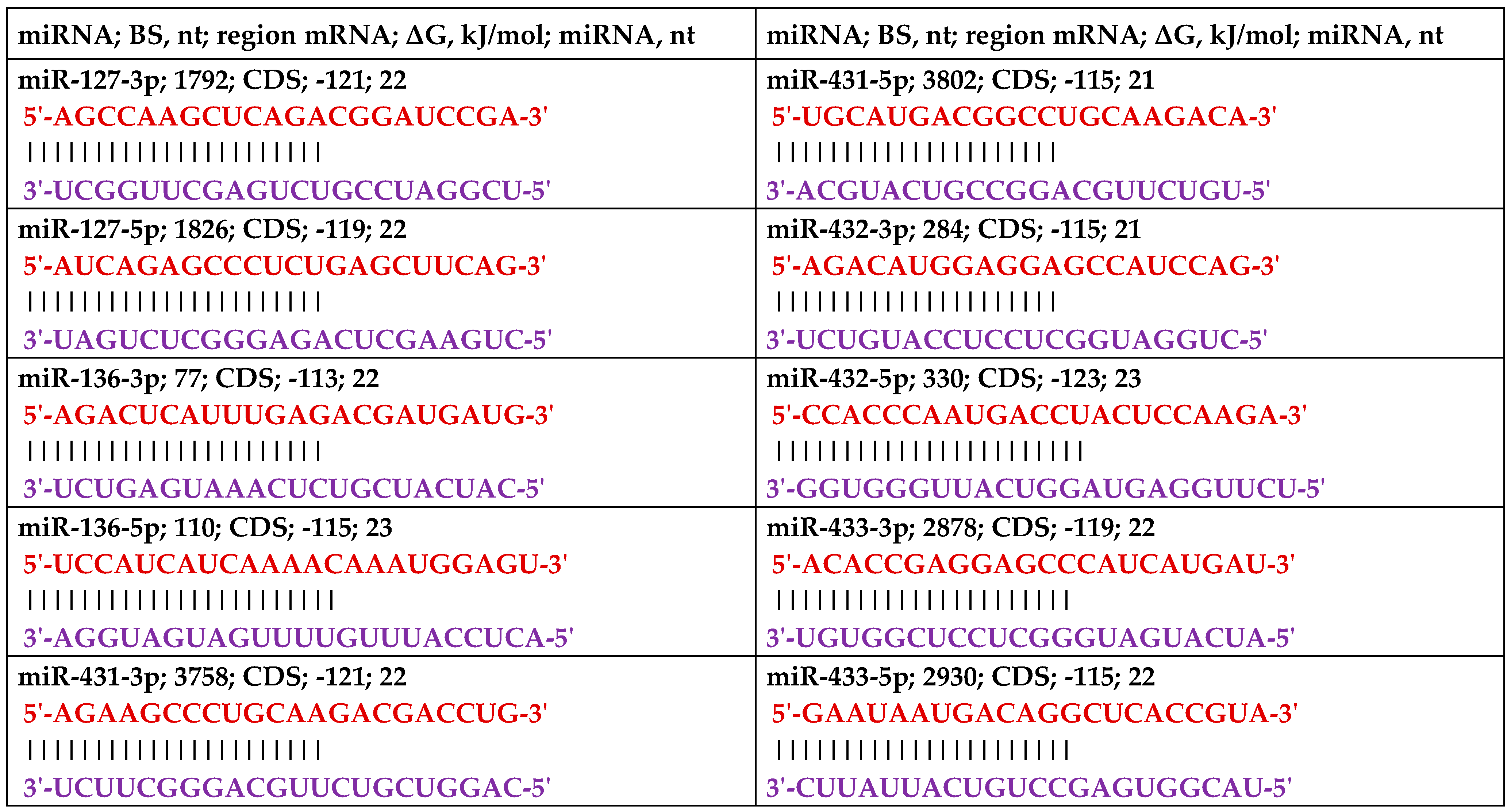
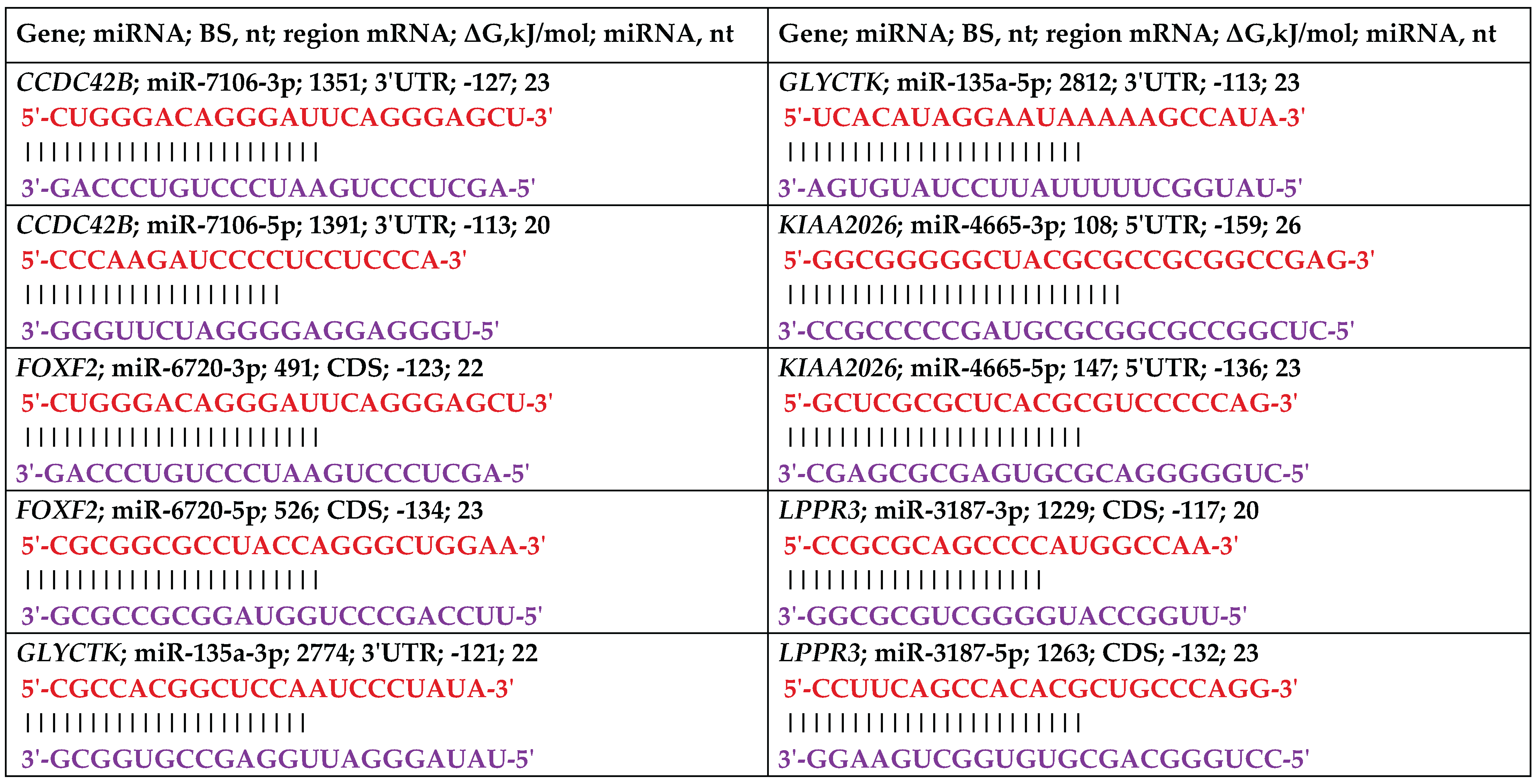
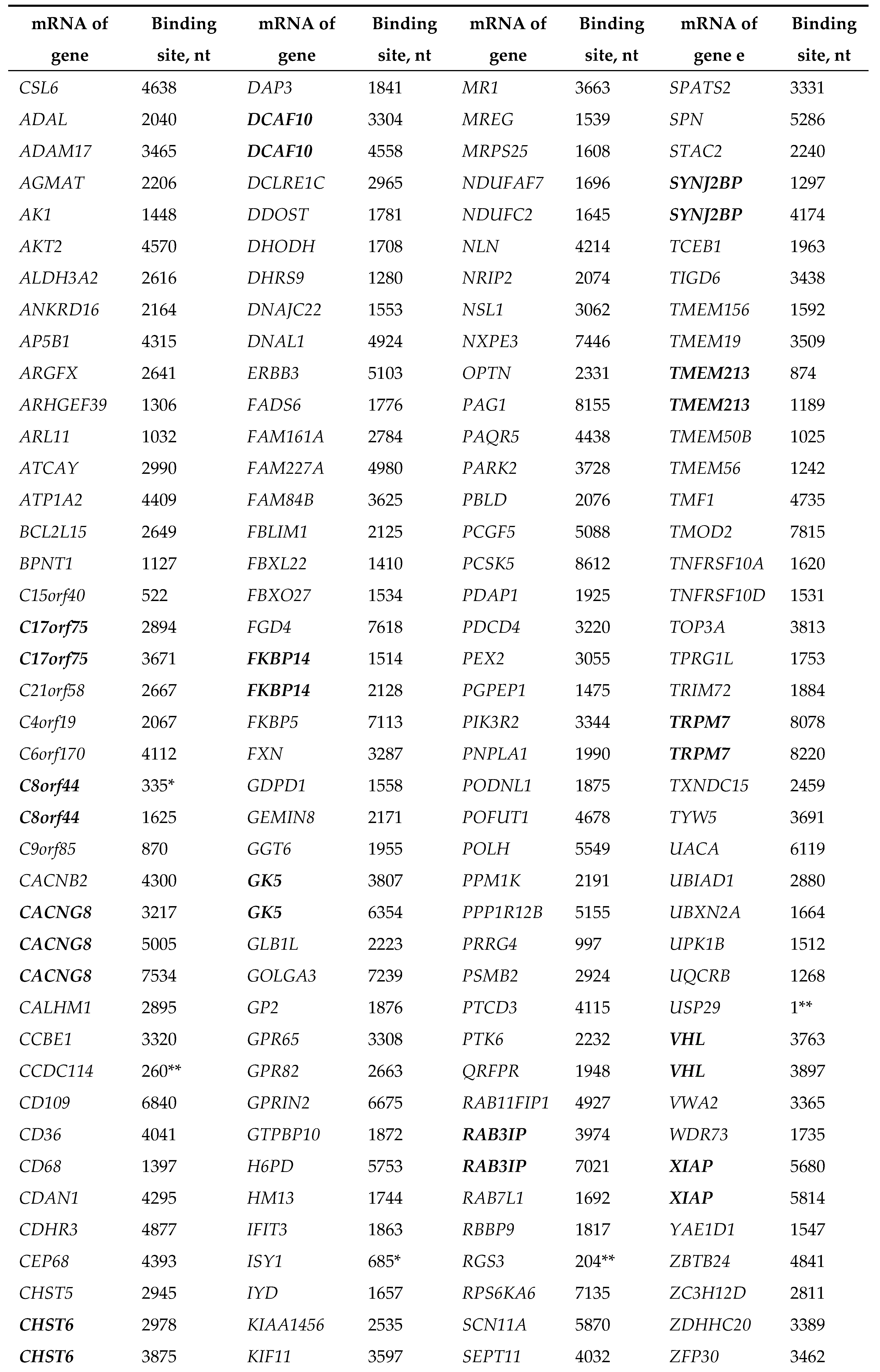 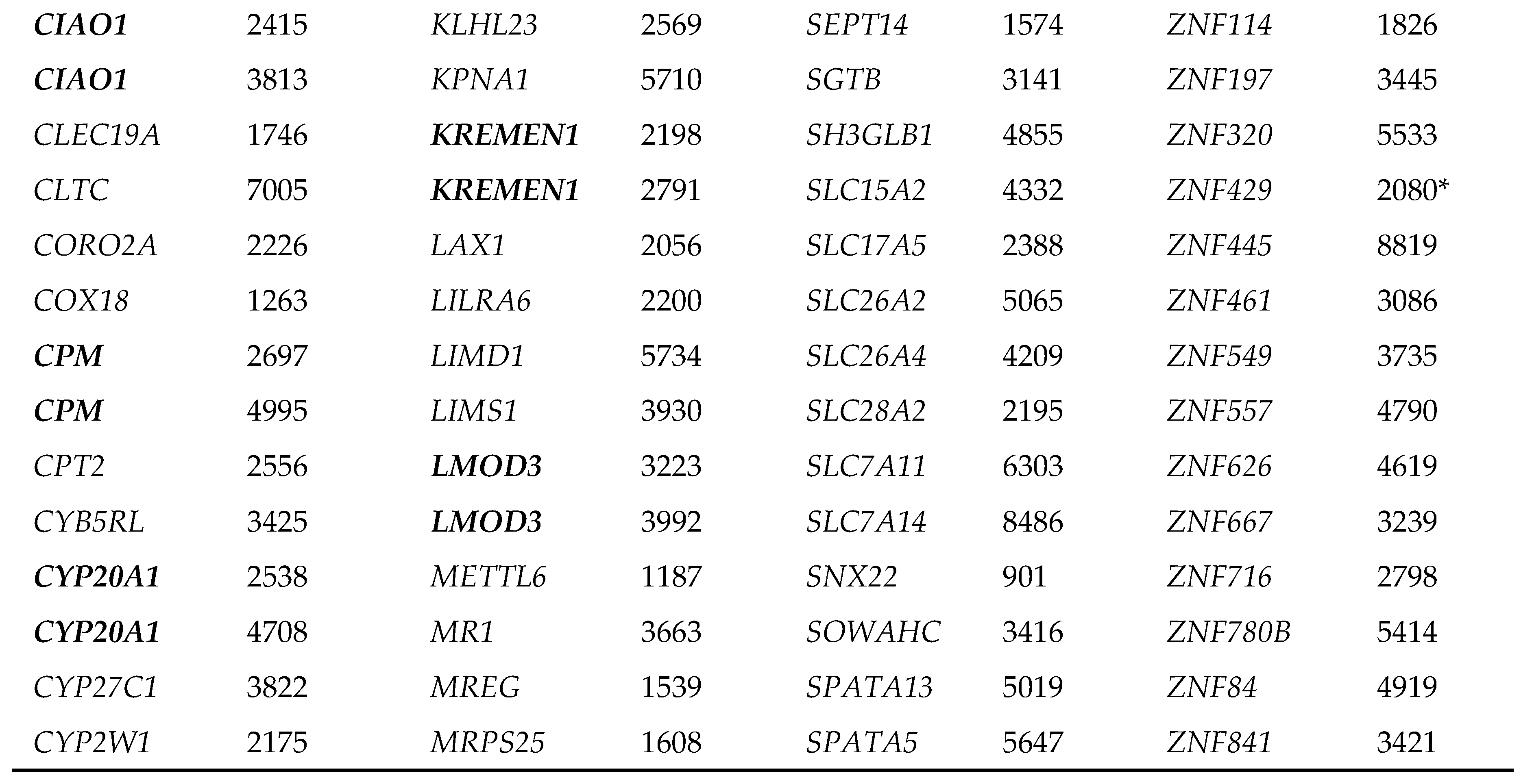
|
 |
| mRNA of gene | miRNA | Binding site, nt | ΔG,kJ/mole | Region of mRNA |
| ASXL1-v-1 | miR-1-155-3p | 340 | -138 | 5'UTR |
| CPT1A-v-2 | miR-1-155-3p | 97 | -138 | 5'UTR |
| CXXC4-v-1 | miR-1-155-3p | 742 | -138 | CDS |
| DISP2-v-1 | miR-1-155-3p | 20 | -138 | 5'UTR |
| DNAJC21- | miR-1-155-3p | 62 | -138 | 5'UTR |
| ESPN-v-1 | miR-1-155-3p | 1966 | -138 | CDS |
| FAM131C- | miR-1-155-3p | 1 | -138 | 5'UTR |
| 1HTT-v-1 | miR-1-155-3p | 263 | -138 | CDS |
| >IRX4-v-1 | miR-1-155-3p | 1243 | -138 | CDS |
| USP25-v-1 | miR-1-155-3p | 183 | -138 | 5'UTR |
| ACTN2 | miR-1-356-5p | 99 | -146 | 5'UTR |
| BMP8B | miR-1-356-5p | 250 | -146 | 5'UTR |
| BMP8B | miR-1-356-5p | 251 | -146 | 5'UTR |
| mRNA of gene | miRNA | Binding site, nt | ΔG,kJ/mole | Region of mRNA |
| KATNAL1 | miR-548aa | 3945 | -123 | 3'UTR |
| TCEANC2 | miR-548aa | 2658 | -123 | 3'UTR |
| KATNAL1 | miR-548am-5p | 3534 | -110 | 3'UTR |
| LG10 | miR-548ap-3p | 2116 | -89 | 3'UTR |
| ANKRD5 | miR-548ap-3p | 3406 | -89 | 3'UTR |
| KATNAL1 | miR-548ap-3p | 3501 | -89 | 3'UTR |
| TCEANC2 | miR-548ap-3p | 2664 | -89 | 3'UTR |
| ANKRD5 | miR-548au-5p | 3444 | -104 | 3'UTR |
| KATNAL1 | miR-548au-5p | 3535 | -104 | 3'UTR |
| KATNAL1 | miR-548c-5p | 3534 | -110 | 3'UTR |
| SMU1 | miR-548h-3p | 6756 | -115 | 3'UTR |
| ANKRD5 | miR-548t-3p | 3400 | -123 | 3'UTR |
| KATNAL1 | miR-548t-3p | 3495 | -123 | 3'UTR |
| TCEANC2 | miR-548t-3p | 2658 | -123 | 3'UTR |
| mRNA of gene | miRNA | Binding sate, nt | ΔG,kJ/mole | Region of mRNA |
| ACACB | miR-1273f | 7996 | -104 | 3'UTR |
| ATF7IP2 | miR-1273f | 2817 | -104 | 3'UTR |
| CA13 | miR-1273f | 2779 | -104 | 3'UTR |
| COX18 | miR-1273f | 1851 | -104 | 3'UTR |
| GTPBP3 | miR-1273f | 2143 | -104 | 3'UTR |
| KRAS | miR-1273f | 3208 | -104 | 3'UTR |
| LOC100996605 | miR-1273f | 238 | -104 | CDS |
| MAP3K13 | miR-1273f | 4636 | -104 | 3'UTR |
| RNF125 | miR-1273f | 2810 | -104 | 3'UTR |
| SLA2 | miR-1273f | 1910 | -104 | 3'UTR |
| SPTSSA | miR-1273f | 1564 | -104 | 3'UTR |
| XPNPEP3 | miR-1273f | 5392 | -104 | 3'UTR |
| AP3S2 | miR-1273g-3p | 4111 | -117 | 3'UTR |
| C15orf38 | miR-1273g-3p | 4433 | -117 | 3'UTR |
| C20orf203 | miR-1273g-3p | 1938 | -117 | 3'UTR |
| CRLF3 | miR-1273g-3p | 2030 | -117 | 3'UTR |
| EIF1AD | miR-1273g-3p | 2165 | -117 | 3'UTR |
| EVI5 | miR-1273g-3p | 4673 | -117 | 3'UTR |
| GJC1 | miR-1273g-3p | 4830 | -117 | 3'UTR |
| TRAF6 | miR-1273g-3p | 4719 | -117 | 3'UTR |
| ZNF621 | miR-1273g-3p | 4815 | -117 | 3'UTR |
| CASP10 | miR-1273h-5p | 2622 | -117 | 3'UTR |
| GINS3 | miR-1273h-5p | 442 | -117 | CDS |
| MXRA7 | miR-1273h-5p | 1379 | -117 | 3'UTR |
| ZNF445 | miR-1273h-5p | 5930 | -117 | 3'UTR |
| mRNA of gene | miRNA | Binding sate, nt | ΔG,kJ/mole | Region of mRNA |
| FBA1 | miR-1184 | 1016 | -127 | CDS |
| FBA2 | miR-1184 | 1001 | -127 | CDS |
| FBA3 | miR-1184 | 1001 | -127 | CDS |
| ERP29 | miR-1260a | 377 | -100 | CDS |
| NGB | miR-1260a | 1118 | -100 | 5'UTR |
| MAML2 | miR-1260a | 1715 | -104 | CDS |
| SC5DL | miR-1260a | 4212 | -108 | 3'UTR |
| MAML2 | miR-1260b | 1715 | -104 | CDS |
| CENPL | miR-1285-5p | 2316 | -113 | 3'UTR |
| FAM73A | miR-1285-5p | 2974 | -113 | 3'UTR |
| LRAT | miR-1285-5p | 3475 | -113 | 3'UTR |
| TSN | miR-1285-5p | 1964 | -113 | 3'UTR |
| GPR179 | miR-4478 | 2280 | -96 | CDS |
| KHSRP | miR-4478 | 66 | -96 | 5'UTR |
| PRR12 | miR-4478 | 4396 | -96 | CDS |
| RNF111 | miR-4478 | 2436 | -96 | CDS |
| KANK3 | miR-4508 | 510 | -106 | CDS |
| OXR1 | miR-4508 | 103 | -106 | 5'UTR |
| PXDNL | miR-4508 | 2686 | -106 | CDS |
| ALG1 | miR-4526 | 864 | -127 | CDS |
| ALG1L2 | miR-4526 | 400 | -127 | CDS |
| ITGAL | miR-5095 | 4708 | -117 | 3'UTR |
| KANK4 | miR-5095 | 4753 | -117 | 3'UTR |
| RBBP4 | miR-5095 | 4279 | -117 | 3'UTR |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





