Submitted:
06 February 2026
Posted:
06 February 2026
You are already at the latest version
Abstract
Keywords:
Introduction
Repeated Measures ANOVA
| Pairwise comparison | Description of method |
| Bonferroni | Single step. Strong control of Type I error/family-wise error rate (FWER). However, strong control may lead to Type II error. |
| Dunnett | Single step. Designed to compare treatment groups to a control group. |
| Benjamini-Hochberg/False Discovery Rate (BH/FDR) | Controls false discovery rate (FDR) rather than FWER; more powerful than Bonferroni, Holm. |
| Benjamini-Yekutieli (BY) | Same as above. |
| Hochberg | Step-up. Similar to Tukey, but better for unbalanced designs. More powerful than Bonferroni. |
| Holm | More powerful, step-down modification of Bonferroni procedure, while controlling FWER. |
| Hommel | Step up. Generally more powerful than Holm. |
| Sidak | Single step, though a step down version also exists. Stringent Type I error control, but more powerful than Bonferroni. |
| Scheffe | Single step. Flexible, with lower power compared to Tukey for pairwise comparisons. |
| Tukey HSD (Honestly Significant Difference) | Single step. Widely used method across several fields; use for balanced designs. |
| Games-Howell | Single step. Use when the assumption for equal variances is violated. |
| Dunn | Single step. Non-parametric test; use with non-parametric ANOVA. |
| Fisher’s Least Significant Differences | Single step. Does not control for inflation of Type I error; generally not recommended for post hoc testing. |
The R environment
Repeated Measures ANOVA in R
Objects
Data
Method
| Syntax | Purpose |
| library(tidyr) | Loads the package |
| pivot_wider(rma, names_from = phase, values_from = score) | Converts data from long format to wide format. |
| rma_wide<-pivot_wider(rma, names_from = phase, values_from = score) | Assigns a new object to the newly created data. |
| View(rma_wide) | Checks to make sure dataset was properly converted. |
| rma_long<-pivot_longer(rma_wide, cols = -ID, names_to = "phase", values_to = "score") | Converts data from wide to long format. |
Parametric Repeated Measures ANOVA
Repeated Measures ANOVA Using the Rstatix Package
| Package | Syntax | Purpose |
| library(rstatix) library(afex) library(emmeans) |
||
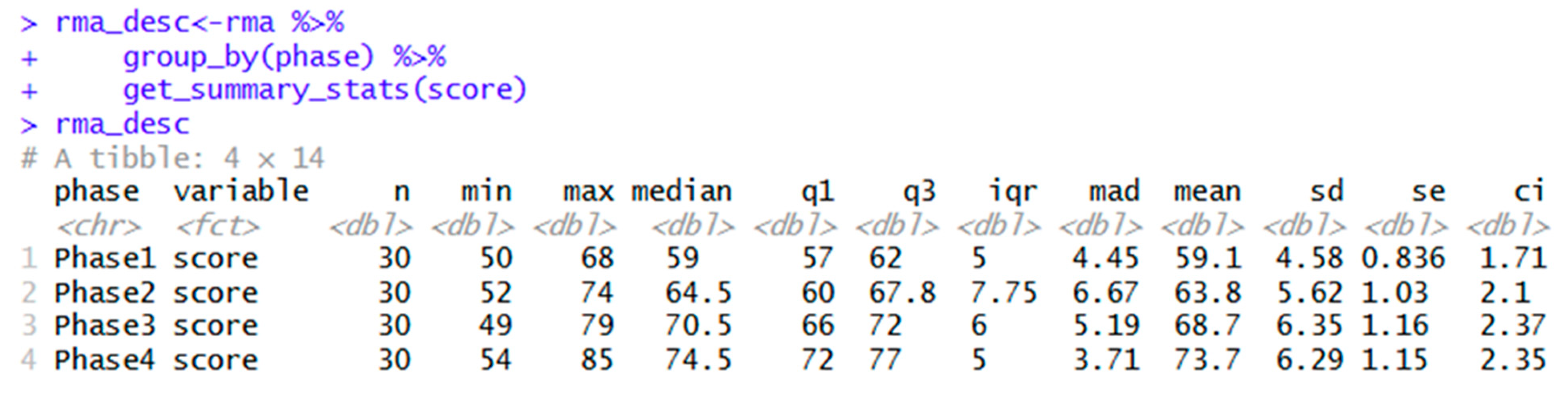
| rstatix | rma_desc<-rma %>% group_by(phase) %>% get_summary_stats(score) |
Creates object for descriptive statistics table. |
| rma_desc | Shows descriptive statistics. | |
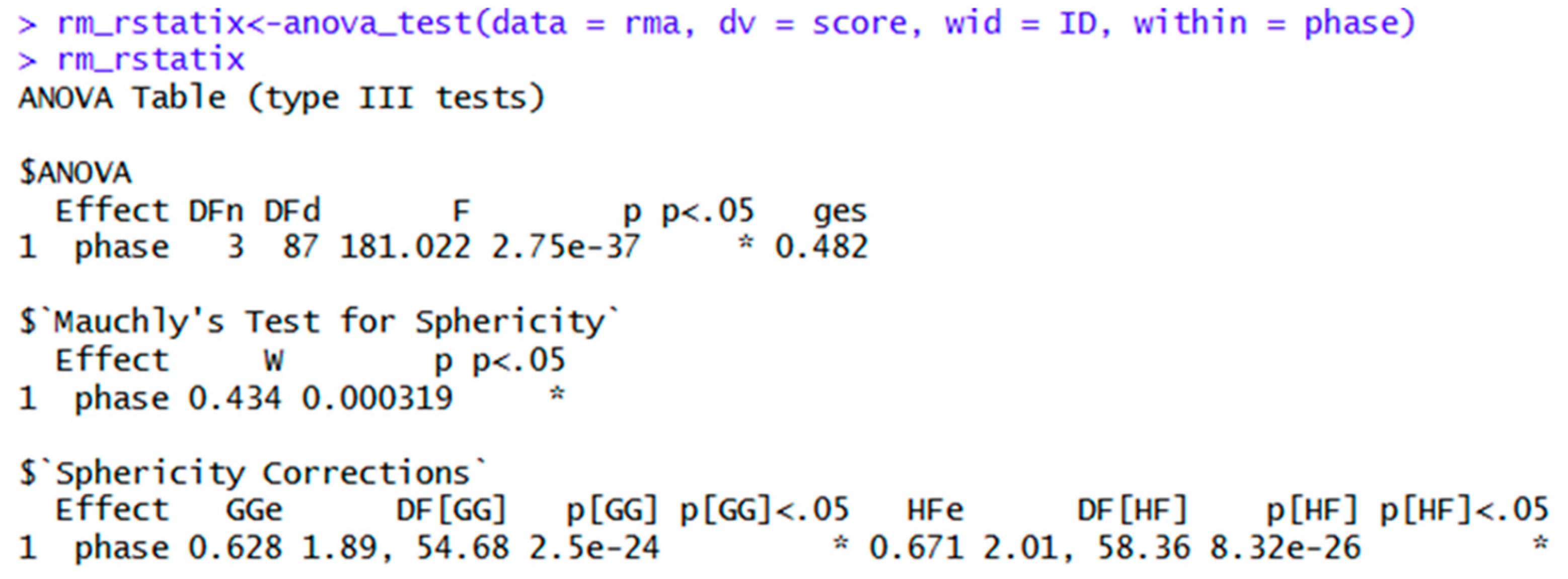
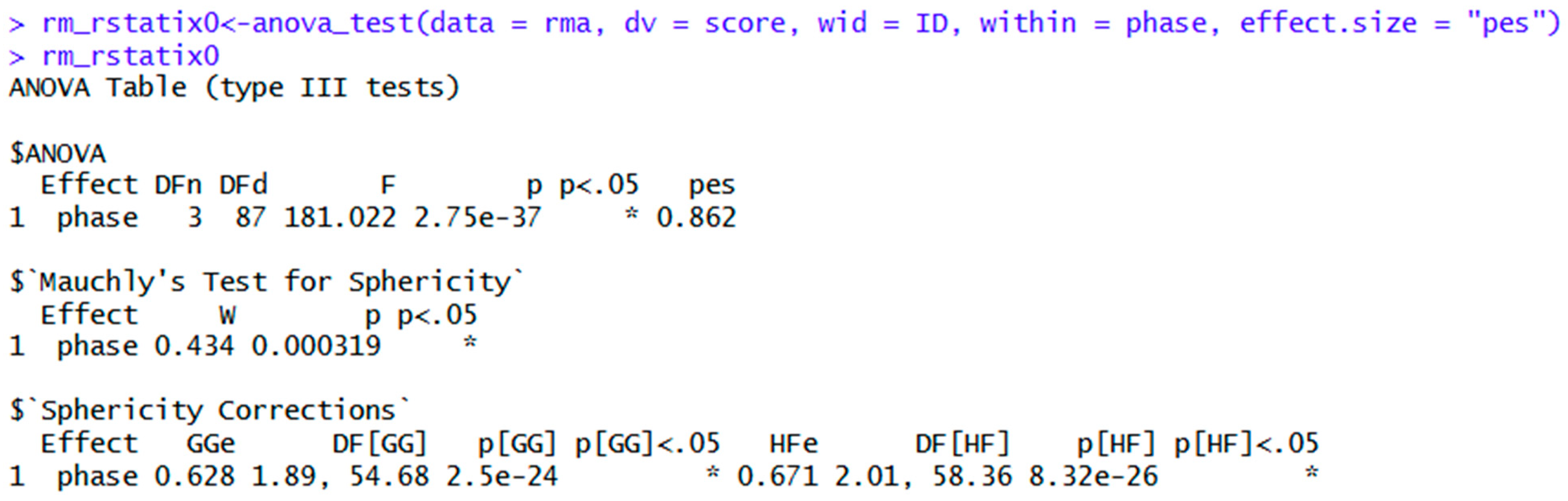
| rm_rstatix<-anova_test(data = rma, dv = score, wid = ID, within = phase) | Repeated measures ANOVA in rstatix. General eta squared is the default effect size. | |
| rm_rstatix0<-anova_test(data = rma, dv = score, wid = ID, within = phase, effect.size = "pes") | Similar to the above. Although the general interpretation is similar, note the difference between general and partial eta squared. | |
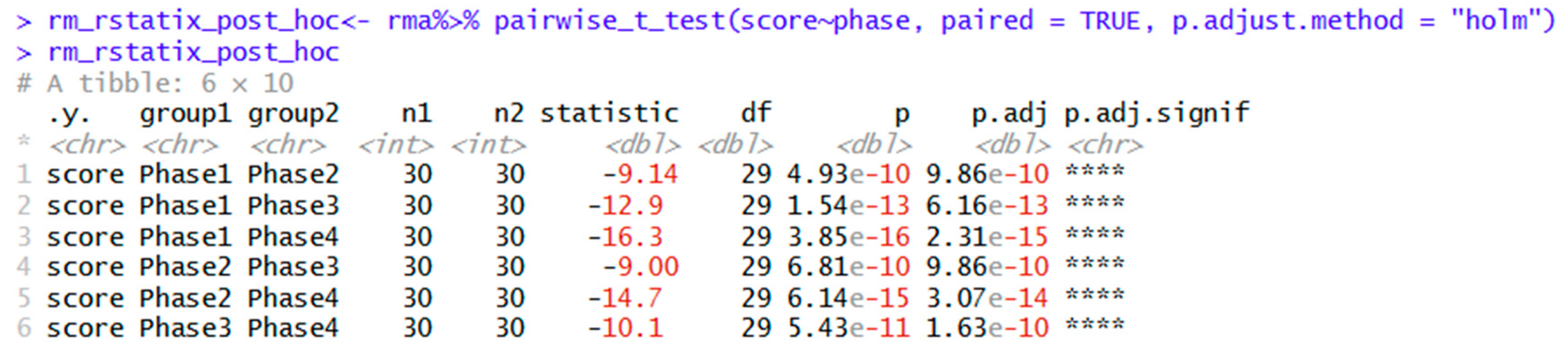
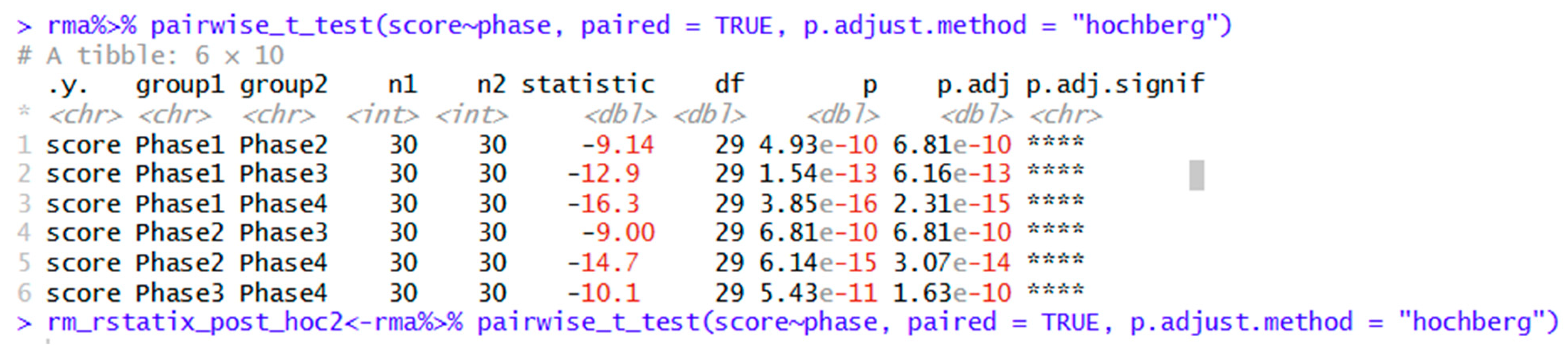
| rm_rstatix_post_hoc<- rma%>% pairwise_t_test(score~phase, paired = TRUE, p.adjust.method = "holm") | Post hoc tests using holm adjustment. Other options include Bonferroni, Hochberg, Hommel, and others. |


Repeated Measures ANOVA Using Afex and Emmeans
| Package | Syntax | Purpose |
| library(afex) library(datawizard) library(emmeans) library(ggstatsplot) |
||
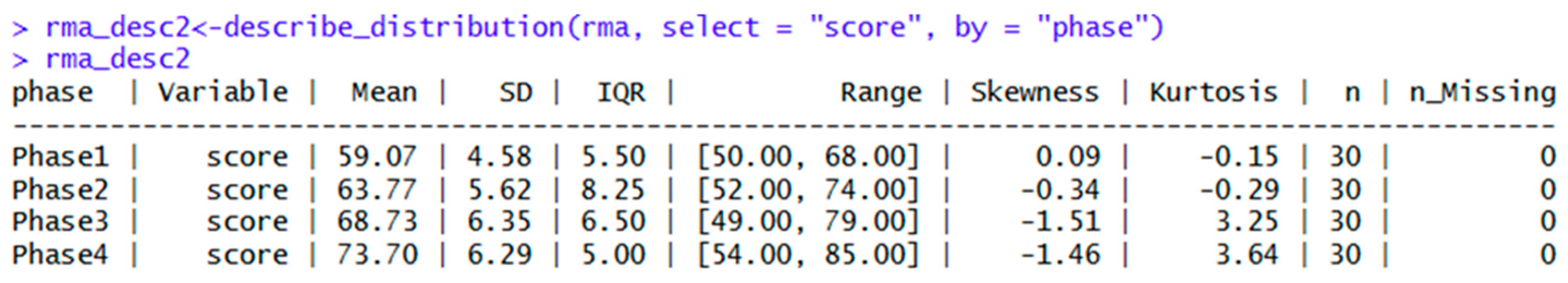
| datawizard | describe_distribution(rma, select = "score", by = "phase") | Descriptive statistics |
| afex | rm_afex<-aov_ez(data = rma, id = "ID", dv = "score", within = "phase", anova_table = list(es = "pes")) | Repeated measures ANOVA in afex. |
| rm_afex | Shows ANOVA table with any adjustments. Greenhouse-Geisser is the default method. | |
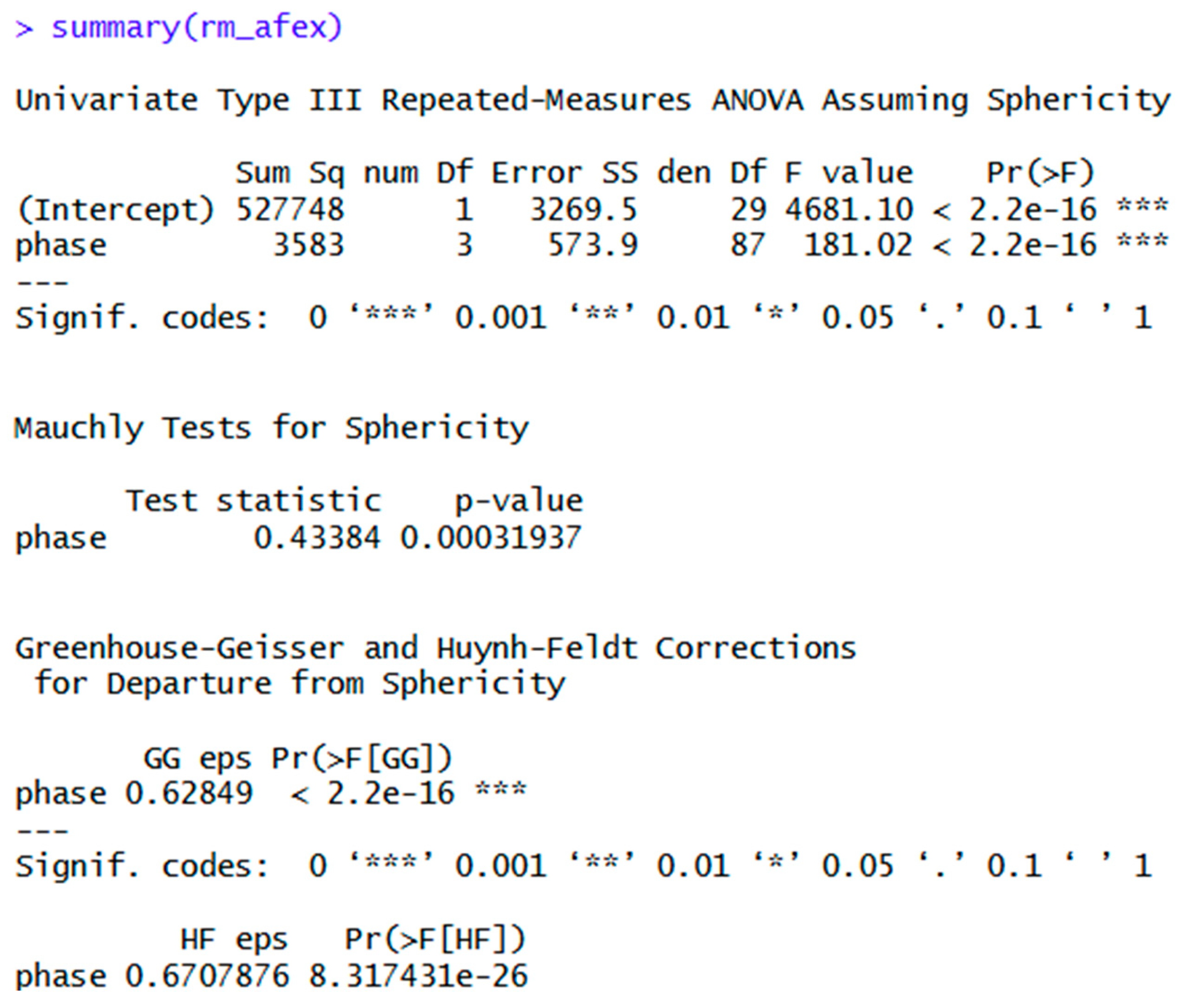
| summary(rm_afex) | Same as above, but with more detail. | |
| rm_afex_HF<- aov_ez(data = rma, id = "ID", dv = "score", within = "phase", anova_table = list(es = "pes", correction = "HF")) | Use this syntax to get Huynh-Feldt adjustment instead. | |
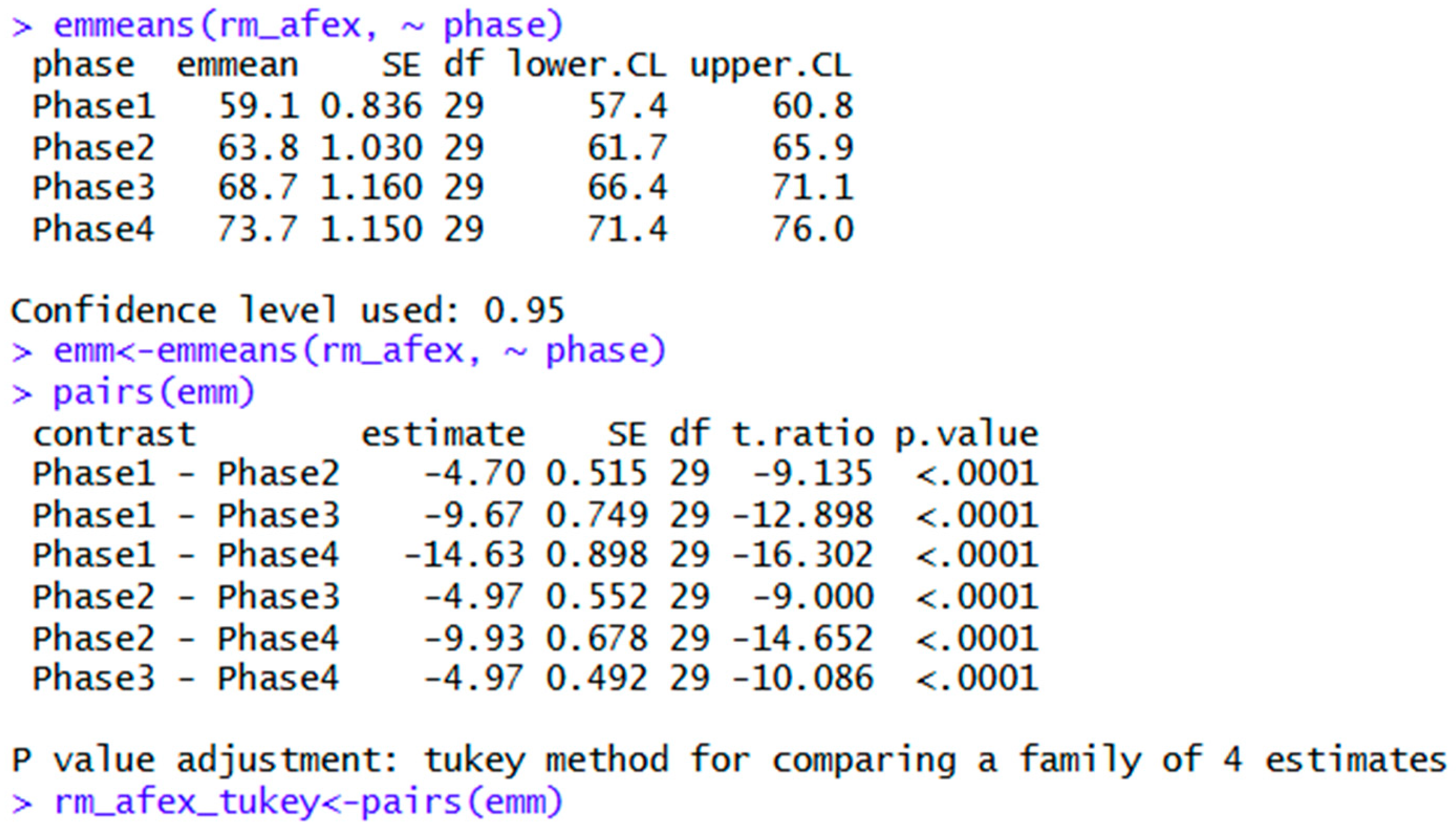
| emmeans | emm<-emmeans(rm_afex, ~ phase) | Shows means and other parameters for each level of the ANOVA. |
| pairs(emm) | Pairwise comparisons; Tukey post hoc is the default. | |
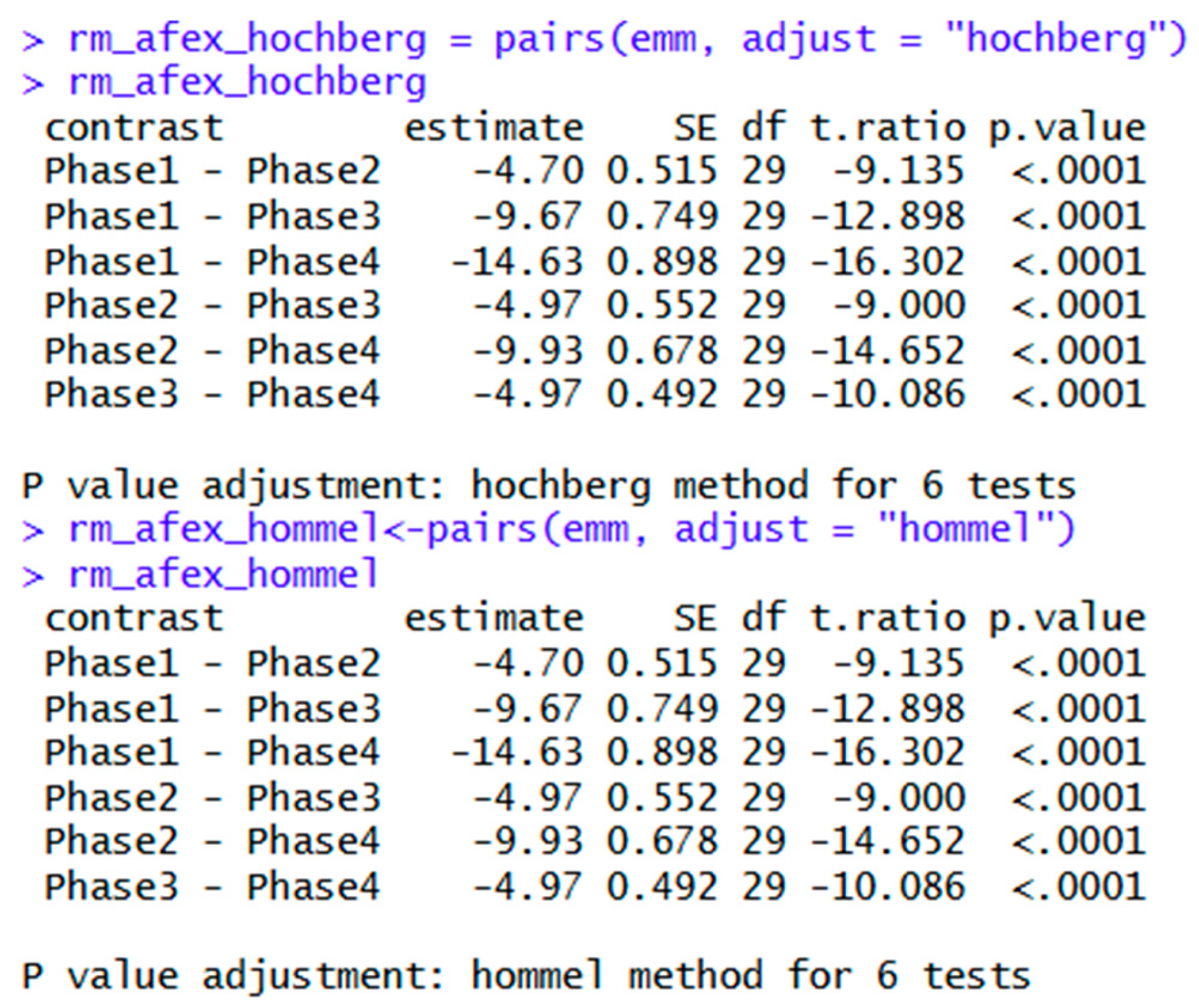
| rm_afex_hochberg = pairs(emm, adjust = "hochberg") | Creates an object for Hochberg post hoc tests. Bonferroni, Hommel, Holm, Scheffe, Sidak, and other options may be found here. | |
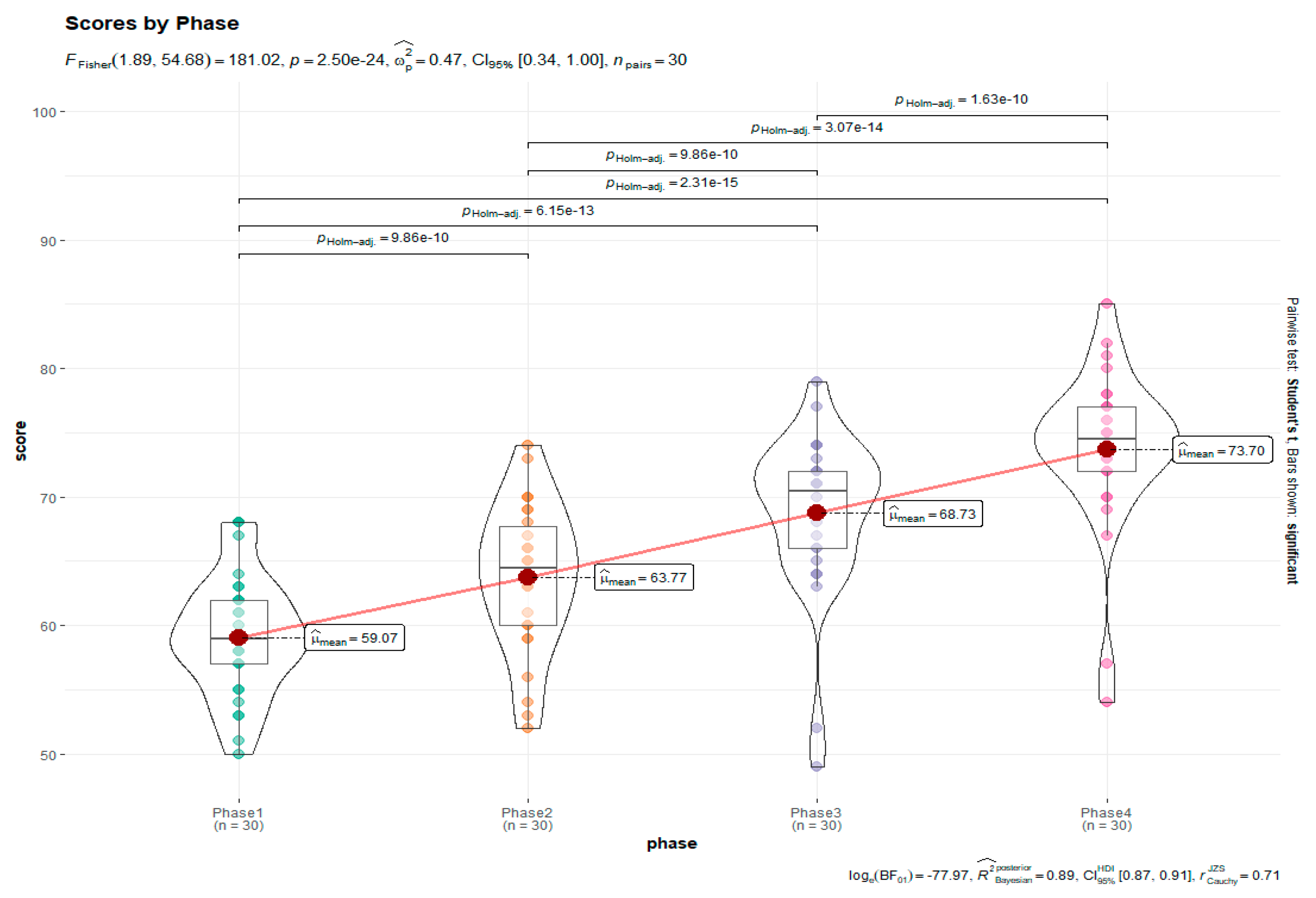
| ggstatsplot | rmplot<-ggwithinstats(data = rma, x = phase, y = score, type = "parametric", pairwise.comparisons = TRUE, p.adjust.method = "holm", title = "Scores by Phase") |
Visualization. |



Options for Violated Normality Assumptions
Friedman’s ANOVA Using Afex
| Package | Syntax | Purpose |
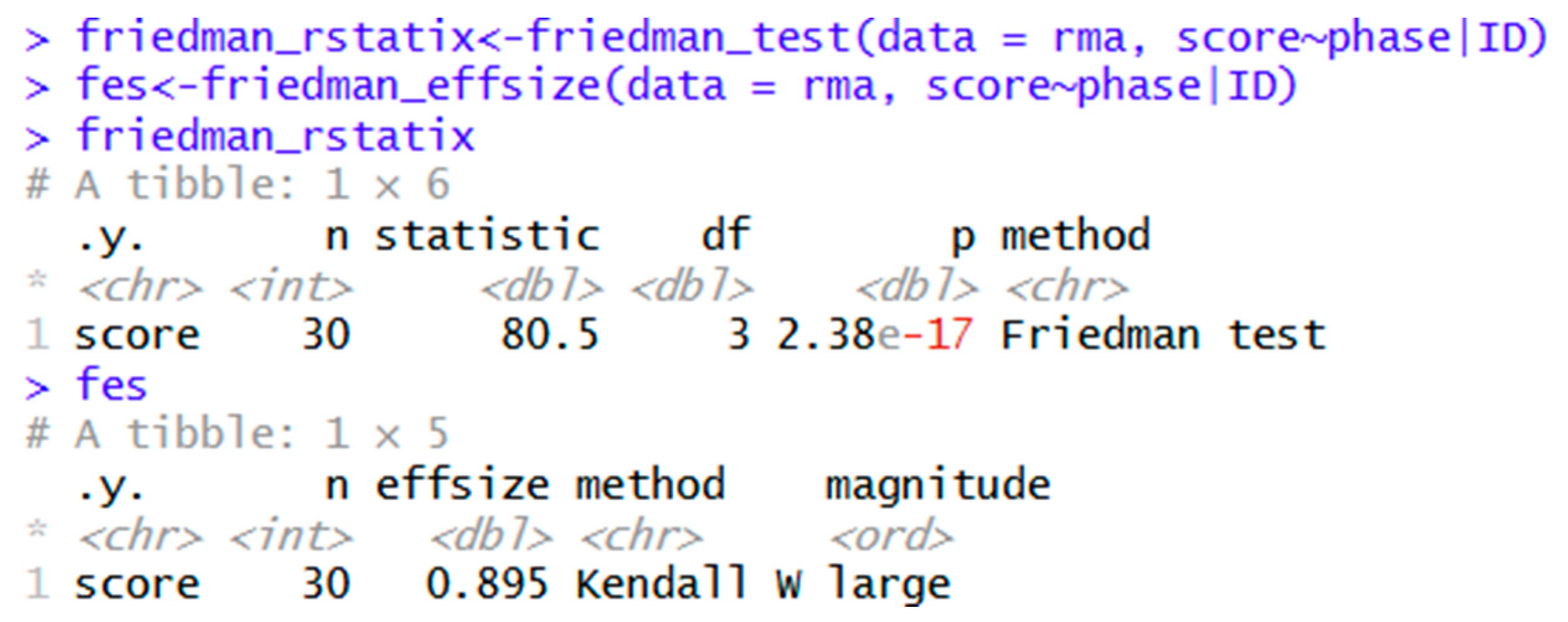
| rstatix | friedman_rstatix<-friedman_test(data = rma, score~phase|ID) | Friedman’s ANOVA in rstatix. |
| fes<-friedman_effsize(data = rma, score~phase|ID) | Generates Kendall’s W. | |
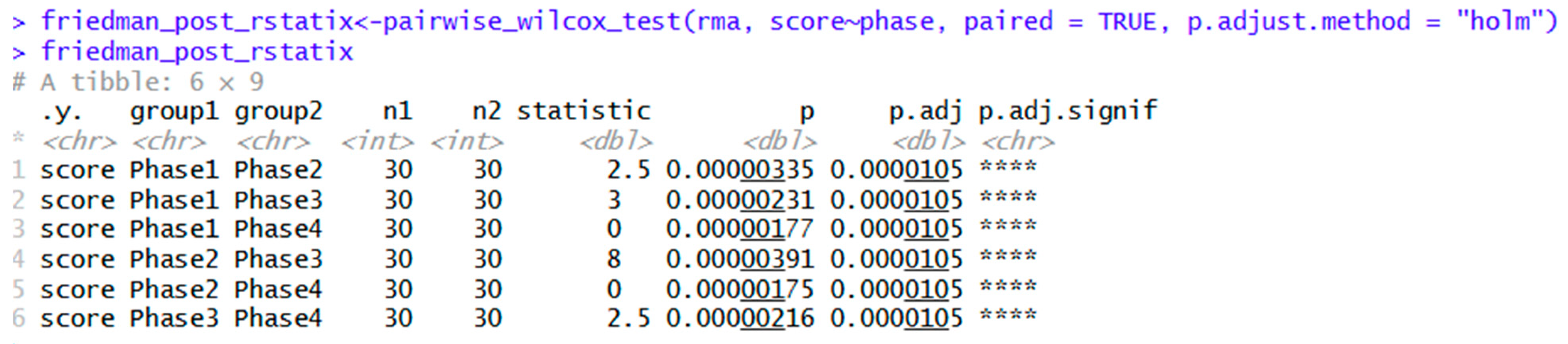
| friedman_post_rstatix<-pairwise_wilcox_test(rma, score~phase, paired = TRUE, p.adjust.method = "holm") | Pairwise Wilcoxon tests with Holm adjustments. | |
| pairwise_wilcox_test(rma, score~phase, paired = TRUE, p.adjust.method = "bonferroni") | Same as above, except with Bonferroni adjustment. | |
| ggstatsplot | friedmanplot<-ggwithinstats(data = rma, x = phase, y = score, type = "nonparametric", pairwise.comparisons = TRUE, p.adjust.method = "holm", title = "Scores by Phase") | Visualization. This package is helpful as it also gives an APA style report and post hoc comparisons. |
Robust Repeated Measures ANOVA Syntax Using WRS2
| Package | Syntax | Purpose |
| library(WRS2) | ||
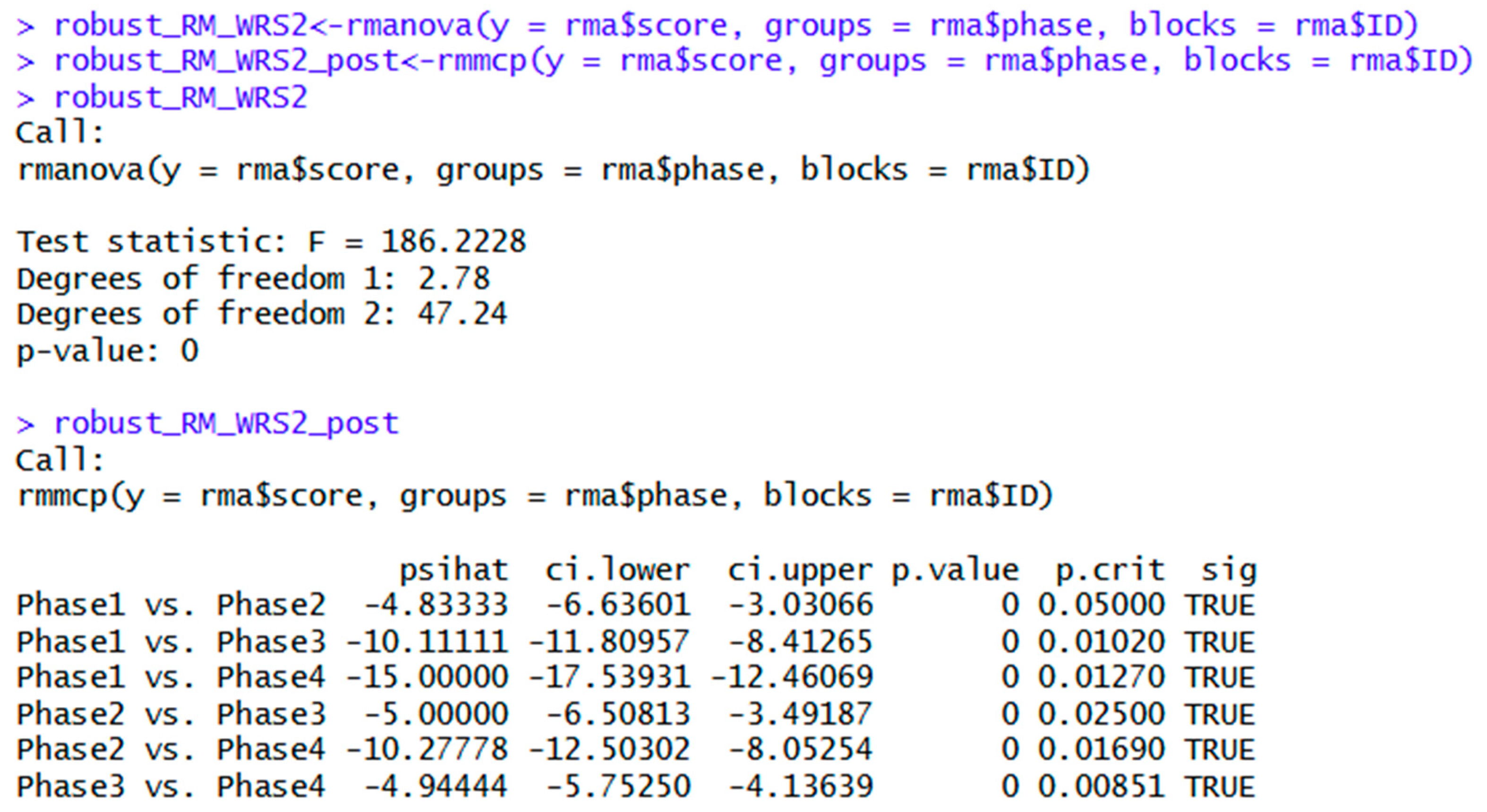
| WRS2 | robust_RM_WRS2<-rmanova(y = rma$score, groups = rma$phase, blocks = rma$ID) | Robust repeated measures ANOVA |
| with(rma, (rmanova(y = score, groups = phase, blocks = ID))) | Alternate syntax for above. | |
| rmmcp(y = rma$score, groups = rma$phase, blocks = rma$ID) | Post hoc tests for trimmed means. | |
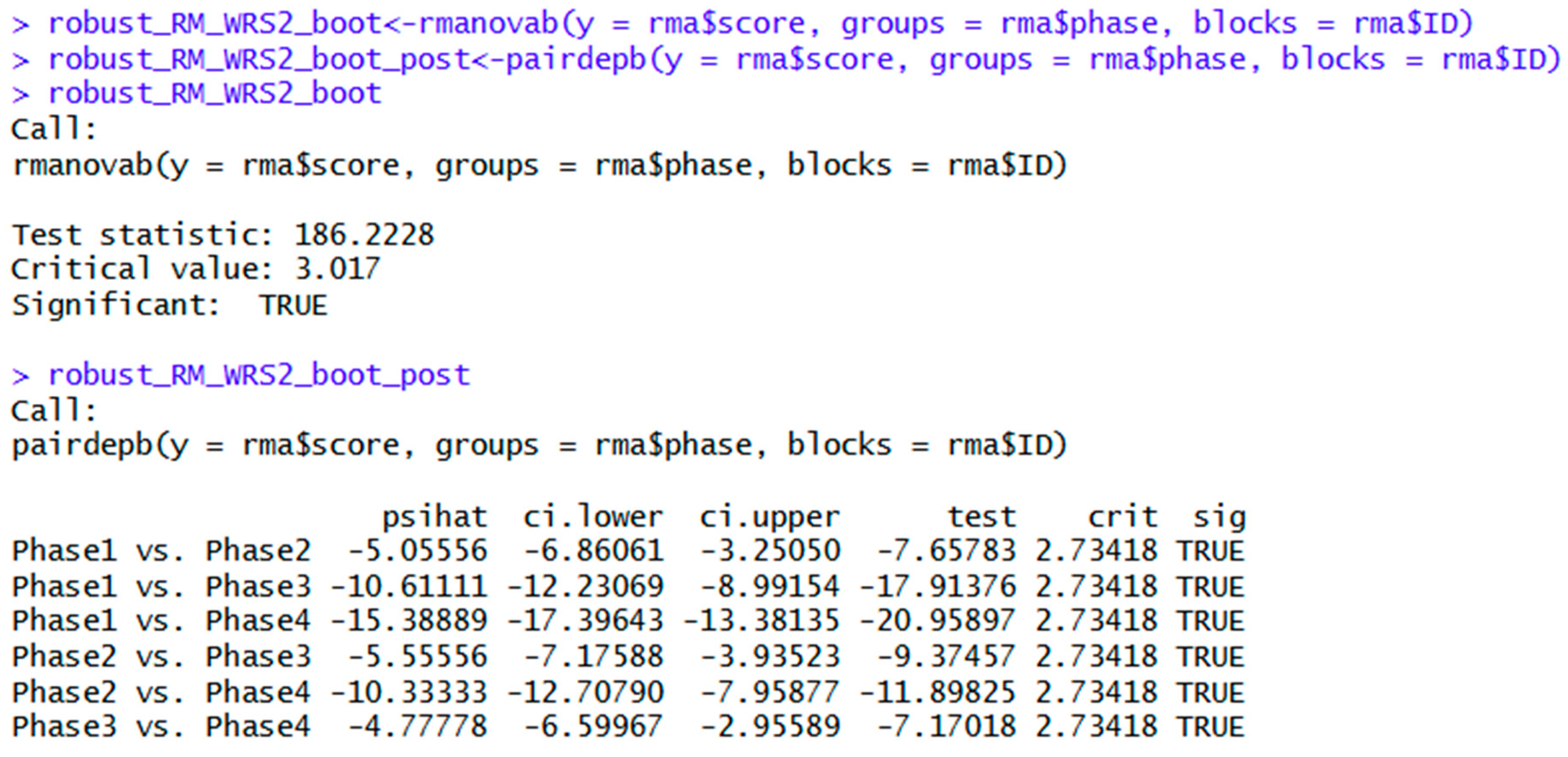
| rmanovab(y = rma$score, groups = rma$phase, blocks = rma$ID) | Bootstrapped robust ANOVA. | |
| rmanovab(y = rma$score, groups = rma$phase, blocks = rmaf$ID, nboot = 2000) | Same as above, specifying 2000 bootstrapped samples. | |
| pairdepb(y= rma$score, groups = rma$phase, blocks = rma$ID, nboot = 2000) | Bootstrapped post hoc tests for the above ANOVA. |
Aligned Rank Transformed ANOVA Using ARTool
| Package | Syntax | Purpose |
| library(ARTool) | ||
| rma$ID <-factor(rma$ID) | Convert the ID variable into a factor. | |
| rma$phase <-factor(rma$phase) | Same for independent variable. | |
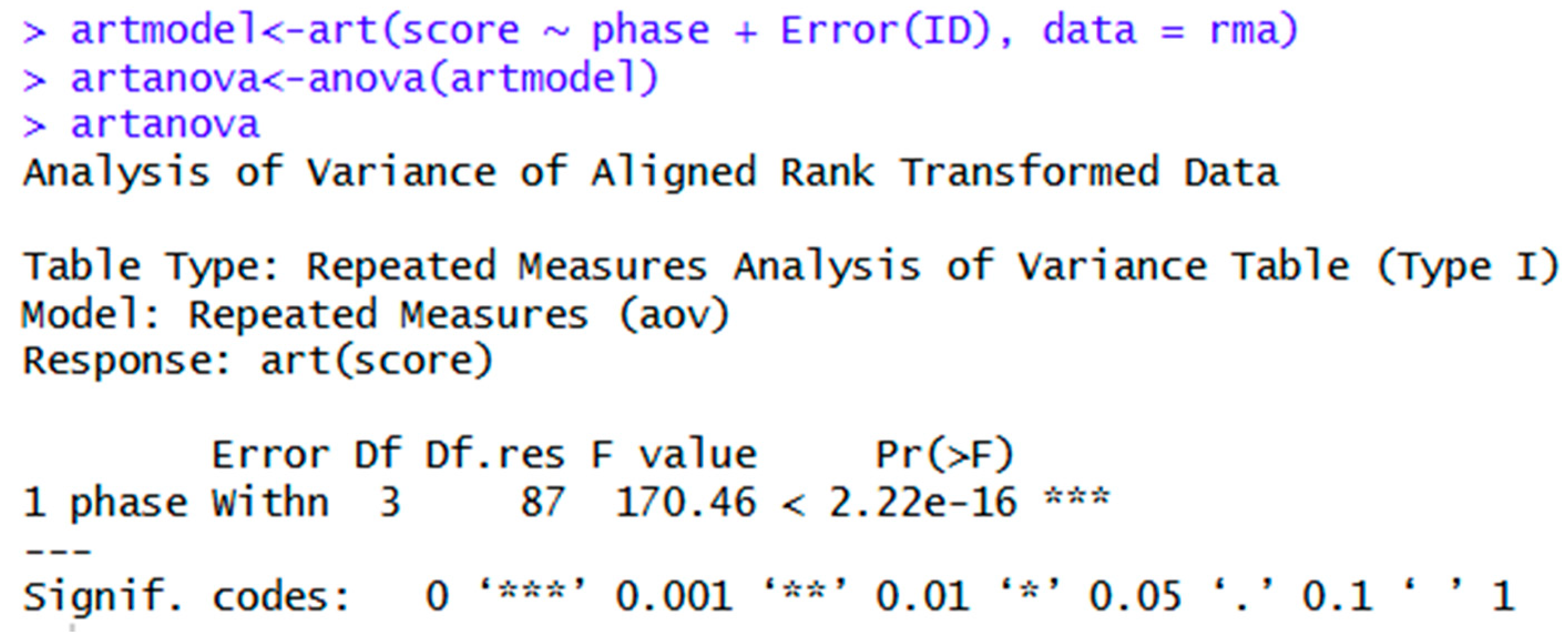
| ARTool | artmodel<-art(score ~ phase + Error(ID), data = rma) | Transformation of repeated measures data using ARTool. |
| artanova<- anova(artmodel) | Repeated measures ANOVA of aligned rank transformed data in ARTool. | |
| artanova | Shows ANOVA table. | |
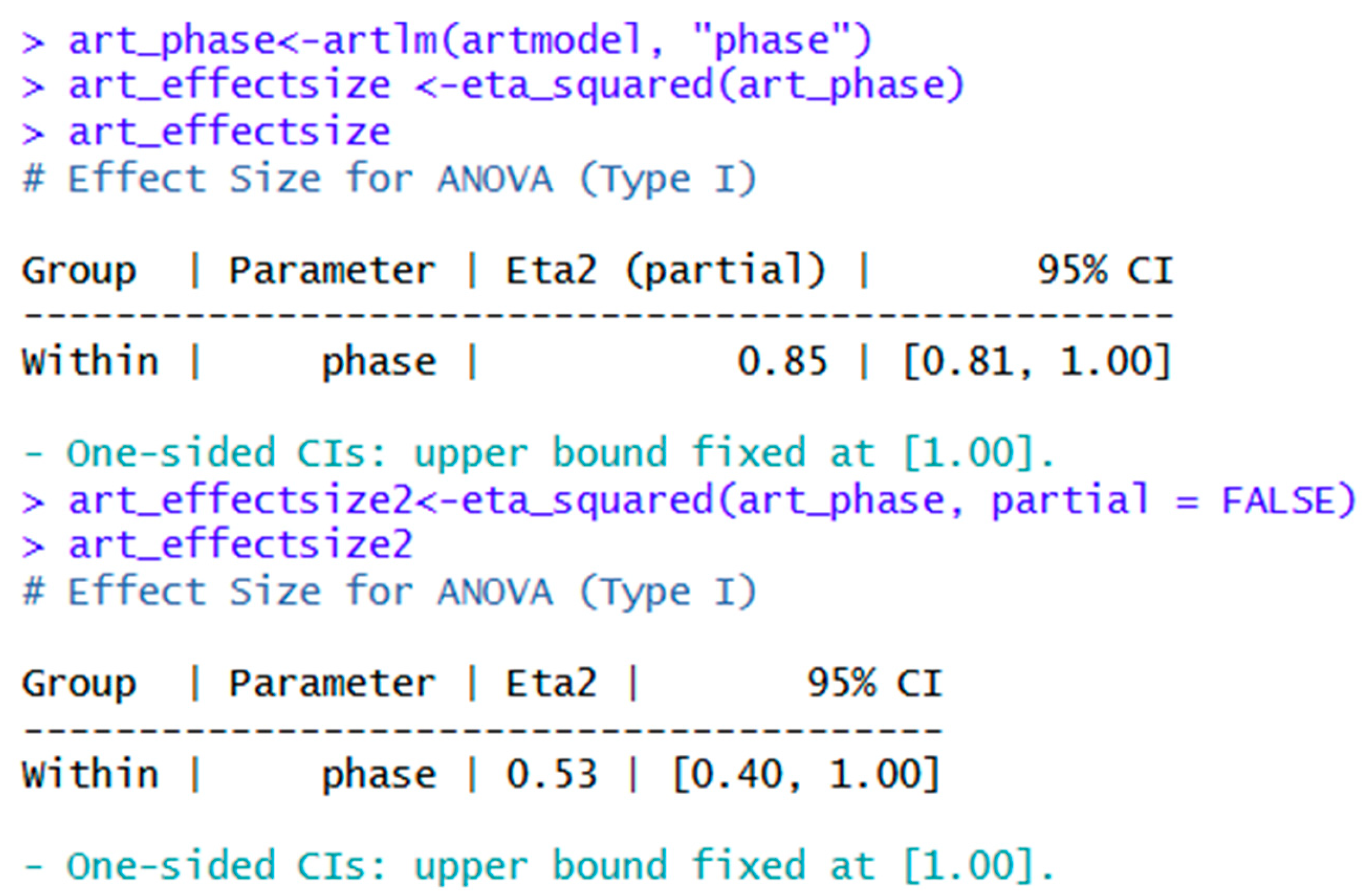
| art_phase<-artlm(artmodel, "phase") | ||
| effectsize | eta_squared(art_phase) | Partial eta squared. |
| art_effectsize2<-eta_squared(art_phase, partial = FALSE) | Generalized eta squared. | |
| eta_squared(art_phase, partial = FALSE, generalized = TRUE) | Same as above. | |
| emmeans | art_post <-emmeans(art_phase, pairwise ~ phase) | Pairwise comparisons with Tukey adjustment as the default method. Multiple post hoc tests are shown for demonstration only. Choose one post hoc test ahead of time. |
| emmeans(art_phase, pairwise ~ phase, adjust = "bonferroni") | Bonferroni adjustment. | |
| art_post_holm<-emmeans(art_phase, pairwise ~ phase, adjust = "holm") | Holm adjustment. Also offers Scheffe, Sidak, and others. |


Discussion
Limitations
Conclusions
References
- Ben-Shachar, M., Lüdecke, D., & Makowski, D. (2020). effectsize: Estimation of effect size indices and standardized parameters. Journal of Open Source Software, 5(56), 2815. [CrossRef]
- Cohen, J. (1988). Statistical Power Analysis for the Behavioral Sciences (2nd ed.). Hillsdale, NJ: Lawrence Erlbaum Associates, Publishers.
- Ellis, P. D. (2017). The essential guide to effect sizes : statistical power, meta-analysis and the interpretation of research results. Cambridge University Press.
- Field, A. (2013). Discovering statistics using IBM SPSS statistics (4th ed.). Sage Publications.
- Field, A., Miles, J., & Field, Z. (2012). Discovering Statistics Using R. SAGE.
- Kassambara, A. (2025). rstatix: Pipe-friendly framework for basic statistical tests. R package version 0.7.2.
- Kay, M., Elkin, L., Higgins, J., & Wobbrock, J. (2025). ARTool: Aligned rank transformation for nonparametric factorial ANOVAs. [CrossRef]
- Lenth, R., & Piaskowski, J. (2025). Emmeans: Estimated marginal means, aka least-squares means. [CrossRef]
- Mair, P., & Wilcox, R. (2019). Robust statistical methods in R using the WRS2 package. Behavior Research Methods, 52. [CrossRef]
- Patil, I. (2021). Visualizations with statistical details: The 'ggstatsplot' approach. Journal of Open Source Software, 6(61), 3167. [CrossRef]
- Patil, I., Makowski, D., Ben-Shachar, M. S., Wiernik, B. M., Bacher, E., & Ludecke, D. (2022). datawizard: An R Package for Easy Data Preparation and Statistical Transformations. Journal of Open Source Software, 7(78), 4684. [CrossRef]
- Posit (2025). RStudio Desktop. Posit. https://posit.co/download/rstudio-desktop/.
- R Core Team (2025). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/.
- Singmann H., Bolker B., Westfall J., Aust F., & Ben-Shachar, M. (2025). afex: Analysis of factorial experiments. R package version 1.5-0, https://github.com/singmann/afex.
- Wickham, H., Vaughan, D., & Girlich, M. (2024). tidyr: Tidy Messy Data [R package tidyr version 1.1.3] https://doi.org/10.32614/CRAN.package.tidyr. R package version 1.3.1. [CrossRef]
- Wobbrock, J. O., Findlater, L., Gergle, D., & Higgins, J. J. (2011). The aligned rank transform for nonparametric factorial analyses using only ANOVA procedures. Proceedings of the 2011 Annual Conference on Human Factors in Computing Systems - CHI ’11. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




