1. Introduction
Breast cancer affects approximately one in eight women worldwide and remains the second leading cause of cancer-related mortality among women in the United States [
1,
2]. Although breast cancer mortality has declined over recent decades due to advances in early detection and treatment, it remains the most frequently diagnosed cancer in women globally [
2]. Moreover, standard cancer therapies often impose substantial physical toxicity and do not uniformly improve patient prognosis [
3]. These challenges highlight the need for improved, biologically informed treatment strategies that account for tumor-specific and patient-specific genetic differences.
Radiotherapy is a cornerstone of non-surgical cancer treatment and is administered in more than half of all breast cancer cases [
3]. Ionizing radiation induces cytotoxicity by exploiting the genomic instability of tumor cells, which accumulate mutations that are inefficiently repaired due to defective DNA damage response mechanisms [
4]. As a result, malignant cells are generally more susceptible to radiation-induced damage than normal tissues. However, healthy tissues are frequently affected as well, limiting the therapeutic window of radiotherapy [
3].
Radiation-induced toxicities are commonly classified as acute or late effects. Acute toxicities typically develop during or shortly after treatment and may persist for several months, primarily affecting rapidly proliferating tissues such as the epidermis and hair follicles [
3,
5]. These effects include dermatitis, breast pain, and hyperpigmentation. Late toxicities may arise months to years after treatment, are often irreversible, and include fibrosis, collagen remodeling, vascular damage, and structural changes in irradiated tissues and adjacent organs [
3,
5]. Because of these adverse effects, radiation dose escalation is frequently constrained despite marked interpatient variability in treatment response [
3]. Despite the well-documented potential toxicities that come with breast irradiation, there is limited understanding of an individual patient’s sensitivity to that radiation. While most patients will experience some of the known toxicities associated with breast cancer radiation, a small number of patients will experience either very minimal or much more severe radiation toxicities. The clinical challenge, then, is sufficiently consenting a patient to a wide array of possible reactions to their treatment. It also means that Radiation Oncologists are forced to take a reactive approach to radiation toxicity as opposed to a proactive one. Having knowledge upfront of a particular patient’s radiation sensitivity would allow Physicians to not only plan their treatment in a way that may reduce toxicity but also take steps to prophylactically manage expected higher grade side effects. The ability to predict an individual’s toxicity response to breast radiation would be an incredible step forward in creating more tailored and less toxic breast cancer therapy.
Although several candidate genes have been implicated in radiosensitivity, reliable biomarkers capable of predicting individual responses to radiotherapy are currently lacking [
6,
7].
MicroRNAs (miRNAs) have emerged as key regulators of gene expression and represent a rapidly expanding area of cancer research. MiRNAs are short, non-coding RNA molecules (18–25 nucleotides) that regulate post-transcriptional gene expression by binding to complementary sequences in target messenger RNAs, leading to mRNA destabilization or translational repression [
8,
9]. Beyond their intracellular functions, miRNAs can mediate intercellular communication and influence cellular behavior in both local and distant tissues [
10]. They are detectable in a wide range of biological fluids, including plasma, serum, saliva, urine, and breast milk, either freely associated with RNA-binding proteins such as AGO2 or encapsulated within extracellular vesicles (EVs) [
8,
9,
11]. More than 2,000 mature miRNAs have been identified in humans, many of which are dysregulated in cancer and contribute to tumor growth, invasion, and metastasis [
10].
Among EVs, exosomes have garnered particular interest due to their role in transporting miRNAs and other bioactive cargo between cells. Exosomes are membrane-bound vesicles approximately 30–150 nm in diameter that are released by most eukaryotic cells and are present in diverse biological fluids [
11,
12]. They originate from the endosomal pathway, where intraluminal vesicles form within multivesicular bodies through mechanisms involving the ESCRT machinery [
12,
13]. Selective miRNA loading into exosomes occurs through both passive and active mechanisms, including recognition of specific sequence motifs by RNA-binding proteins [
10]. Following secretion, exosomes interact with recipient cells through multiple pathways, including receptor-ligand interactions, membrane fusion, and endocytosis, ultimately delivering their cargo and modulating cellular function [
13,
14,
15].
The clinical potential of exosomal miRNAs continues to expand as their biogenesis, targeting, and functional roles become better understood. Exosomal miRNA profiling has shown promise for cancer detection, monitoring disease progression, and characterizing tumor molecular profiles [
16]. Furthermore, therapeutic strategies aimed at inhibiting oncogenic miRNAs or restoring tumor-suppressive miRNAs are actively being explored [
9,
11].
Despite this progress, the role of miRNAs – particularly exosomal miRNAs – in determining radiosensitivity and radiation-induced toxicity remains relatively underexplored. Tumor heterogeneity contributes to variable radiotherapy responses, yet accurately predicting this variability remains challenging [
16]. Prior studies have identified miRNAs involved in tumor radiosensitivity and radiation response pathways, including miRNAs associated with DNA damage repair, oxidative stress, and apoptosis [
17,
18]. However, investigations into miRNAs associated with normal tissue toxicity are limited. A small number of studies have implicated specific miRNAs in radiation-induced cardiovascular and tissue toxicities, but these analyses have largely focused on total circulating miRNAs rather than those encapsulated within EVs [
6,
19]. This gap underscores the need for studies specifically examining exosomal miRNA expression in the context of radiotherapy-induced toxicity in breast cancer patients.
The present study aims to identify exosomal miRNAs whose baseline expression may predict individual susceptibility to radiation-induced toxicity in normal tissues. By analyzing plasma-derived exosomal miRNAs from breast cancer patients with varying degrees of radiotoxicity, this study seeks to identify reliable biomarkers that can inform personalized radiotherapy strategies. Using quantitative PCR as a high-throughput screening approach, this work evaluates the differential expression of hundreds of known miRNAs. The identified miRNAs may not only serve as predictive biomarkers but also provide mechanistic insight into the molecular pathways governing radiation response, ultimately contributing to improved risk stratification and individualized patient care.
2. Materials and Methods
Participants and blood samples:
The patients included in this study were enrolled on MCC-12181, an IRB-approved, prospective clinical trial that collected blood samples from patients undergoing definitive radiation therapy for cancer. Informed consent was obtained from all subjects. Per the trial protocol, blood samples were collected one day prior to the start of radiation and at a variety of time points post the start of treatment. For this current analysis, blood samples included were from those patients definitively treated with adjuvant radiation after surgery for breast cancer. Patients underwent either breast-conserving surgery or mastectomy with or without reconstruction. Radiation techniques included both external beam and brachytherapy approaches. All of the samples analyzed were collected prior to the patient starting radiation. Blood samples collected at additional time points were not included in the presented data. In total, samples from 89 patients were included in the analysis.
Clinical outcomes, including acute and long-term toxicities, were evaluated and data collected for those patients for whom we had blood samples. Patients were divided into four groups based on their observed radiation side effects. Group 1 contained patients who had very minimal to virtually no side effects from breast radiation. Group 2 contained patients who had an expected level of appropriate side effects from radiation (Grade 1- mild Grade 2 toxicities). Group 3 contained patients who had higher-grade acute toxicities with subsequent resolution (Grade 2-3 dermatitis, edema, pigmentation changes), while Group 4 had those with higher-grade long-term toxicity (Grade 2-3 fibrosis/fat necrosis/pigmentation changes). For the purposes of this study, possible toxicities meeting criteria for inclusion included acute radiation dermatitis, fibrosis, hyperpigmentation and fat necrosis. Toxicity grading was based on CTCAE version 5 criteria.
The blood samples were collected in the tubes coated on the inside with K2EDTA (Potassium Ethylene Diamine Tetra Acetic acid). On the same day of collection, blood samples were centrifuged at 3,000x g for 15 minutes at 4 °C. After centrifugation plasma part of all samples was aliquoted and saved in vapor phase of liquid nitrogen. For statistical analysis of the clinical data, Pearson’s chi-square (χ2) test was applied.
Exosome isolation
Exosomes were extracted from plasma samples by a two-step method. The size exclusion chromatography (SEC) method using the qEVoriginal columns (Izon Science) was applied as a first step. Every 0.5 ml of plasma yielded 1.5 ml of purified exosomes in the filtered (100 nm filter) 1xPBS buffer. This method has been described in detail previously [
20]. As the second step, exosomes were precipitated using the PureExo
® Exosome Isolation Kit for Serum and Plasma (101Bio, Palo Alto, CA, USA) according to the manufacturer’s instructions. Precipitated exosomes were reconstituted in 0.25 ml of the filtered (100 nm filter) 1xPBS buffer for subsequent analysis and RNA extraction.
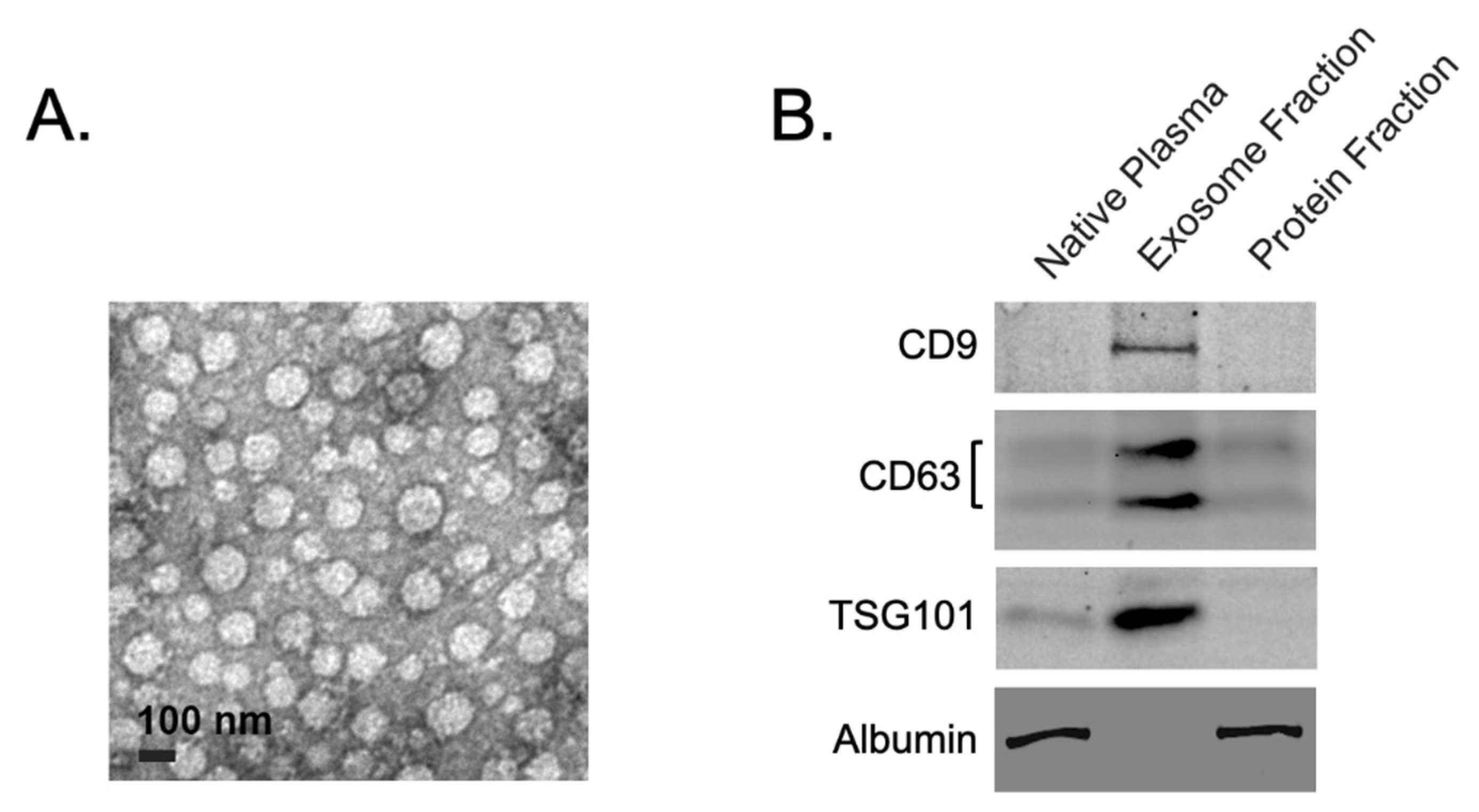
Western blotting (WB) and Antibodies:
Equal amount of the total protein from each sample was loaded and separated by SDS-PAGE gel and transferred to nitrocellulose membranes. The membranes were exposed to antibodies at specific dilutions. Primary antibodies used for WB: anti-CD9 (dilution 1:1000, Novus Biologicals), anti-CD63 (dilution 1:1000, ThermoFisher), anti-TSG101 (dilution 1:500, Cell Signaling), anti-Albumin (dilution 1:1000, Cell Signaling). Specific protein bands were detected using infrared-emitting conjugated secondary antibodies: anti-rabbit DyLight™ 800 4X PEG Conjugate (dilution 1:10,000, Cell Signaling). WB images were generated and analyzed using the ChemiDoc Infrared Imaging System (Bio-Rad).
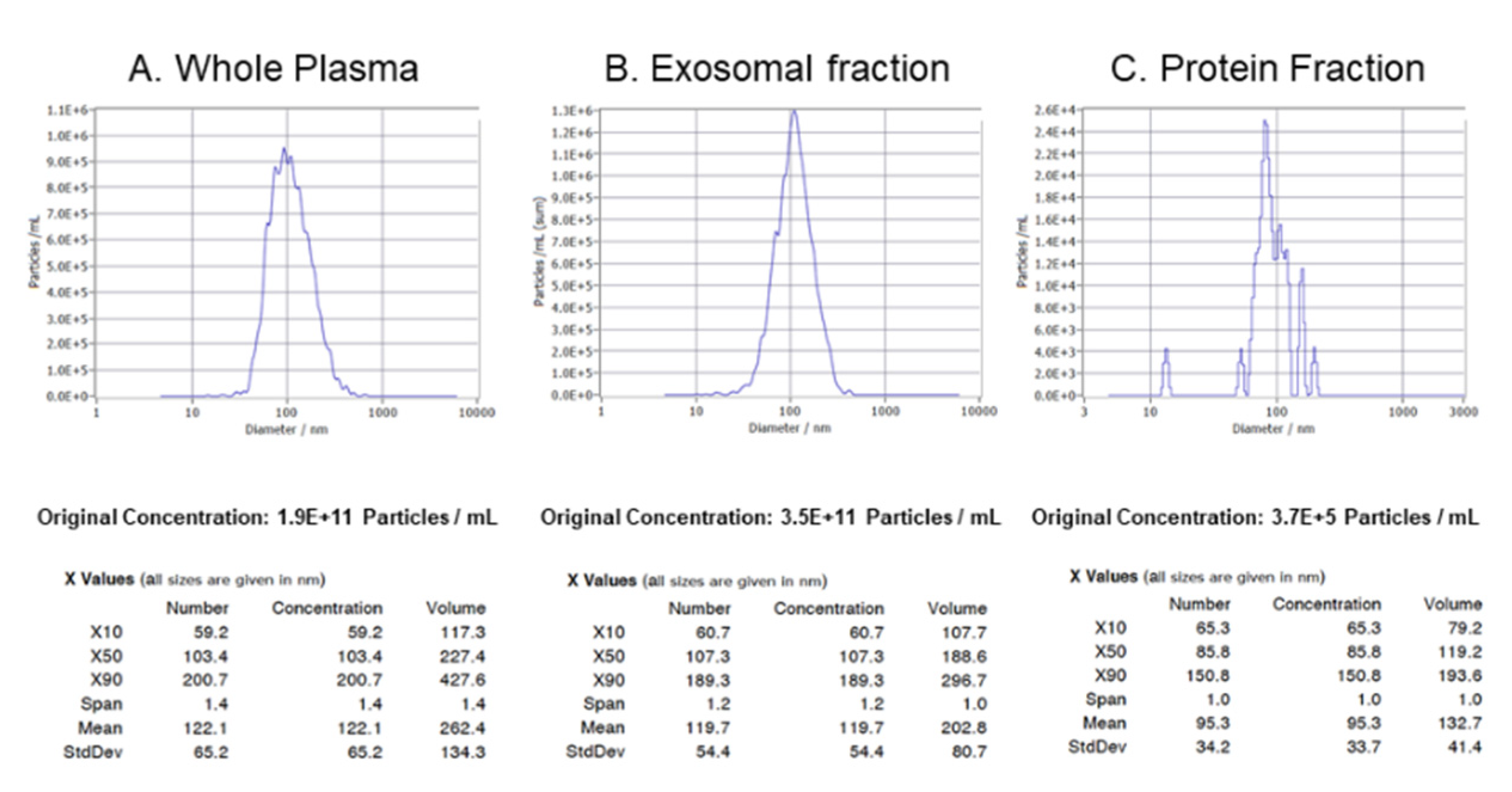
Nanoparticle tracking analysis (NTA)
The concentration and size of the extracted exosomes were measured using ZetaView® NTA. All samples were diluted in PBS to a final volume of 2 ml. Ideal measurement concentrations were found by pre-testing the ideal particle per frame value (140–200 particles/frame). The manufacturer’s default software settings for exosomes were selected accordingly. For each measurement, three cycles were performed by scanning 11 cell positions each and capturing 80 frames per position under following settings: Focus: autofocus; Camera sensitivity for all samples: 78; Shutter: 100; Scattering Intensity: detected automatically; Cell temperature: 25 °C. After capture, the videos were analyzed by the in-built ZetaView Software 8.04.02 SP2 with specific analysis parameters: Maximum area: 1000, Minimum area 5, Minimum brightness: 25. Hardware: embedded laser: 40 mW at 488 nm; camera: CMOS. The number of completed tracks in NTA measurements was always greater than the proposed minimum of 1000 to minimize data skewing based on single large particles.
Transmission electron microscopy (TEM)
Isolated exosomes were fixed and prepared for the TEM as was previously described [
21]. The TEM analysis was carried out by the Microscopy Core of VCU.
Exosomal RNA isolation and RT-PCR
Total RNA was isolated from the exosomal samples following the manufacturer’s instructions with the miRNeasy MicroRNA Extraction Kit (Qiagen) and then converted to cDNA using the miRCURY LNA RT Kit (Qiagen). The samples with total exosomal RNA and cDNA were stored in a freezer at −80 °C. The concentration and purity of the exosomal RNA samples were estimated using NanoDrop ND-1000 spectrophotometer (Thermo Scientific, Wilmington DE).
All cDNA samples were pooled into four groups (clinical Groups 1-4) and then assayed using the Serum/Plasma Focus PCR panel (Qiagen). This commercially available microarray platform contains 179 LNA miRNA primer sets of miRNAs that have been commonly found in human plasma. The Serum/Plasma Focus miRNA PCR Panels include potential reference genes and probes for evaluating potential sample contamination or damage (e.g., hemolysis). Each PCR panel also contains set of the negative control (H2O) and five sets of RNA Spike-In controls to evaluate RNA extraction (Spike-Ins 2-4-5; concentration ratio 1:100:10000) and cDNA synthesis (Spike-In 6) and to perform inter-plate calibration (Spike-In 3). The amplification was performed in a QuantStudio 5 Real-Time PCR System (Thermo Fisher Scientific, USA). Samples were amplified using RT2 SYBR® Green qPCR Mastermix with ROX (carboxy-X-rhodamine) passive reference dye from QIAGEN. The real-time PCR data were normalized by ROX passive reference. The amplification curves were analyzed using the QuantStudio™ Design & Analysis Software v1.4.2, both for determination of Ct values and for melting curve analysis. All assays were inspected for distinct melting curves and the Tm was checked to be within known specifications for each assay. The 2-ΔΔCt method was used for the calculation of relative miRNA expression levels.
RT-PCR data analysis
The amplification curves were analyzed using the QuantStudio™ Design & Analysis Software v1.4.2, both for determination of Ct values and for melting curve analysis. All assays were inspected for distinct melting curves (
Figure S1) and the Tm was checked to be within known specifications for each particular assay. Potential reference genes were identified by comparing the overall stability and relative expression of candidate miRNAs across all groups. Unless indicated otherwise, data are demonstrated as mean ± standard deviation (SD). The significance of the difference between groups was determined by the one-way analysis of variance (ANOVA). The Data Analysis Toolpak for Excel was used for the calculation of the ANOVA. Differences were considered significant for
p-values < 0.05.
3. Results
Table 1 provides a summary of the clinical characteristics of patients in the four study groups. Among the 89 participants included in this study, 9 (10.1%) exhibited minimal or no radiotherapy (RT) – associated toxicity, 45 (52.9%) experienced expected RT-associated toxicity, 7 (7.9%) developed acute Garde 2-3 RT-associated toxicity, and 28 (31.5%) demonstrated late-onset Grade 2-3 RT-associated toxicity.
Analysis of miRNAs from Plasma-Derived Exosomes
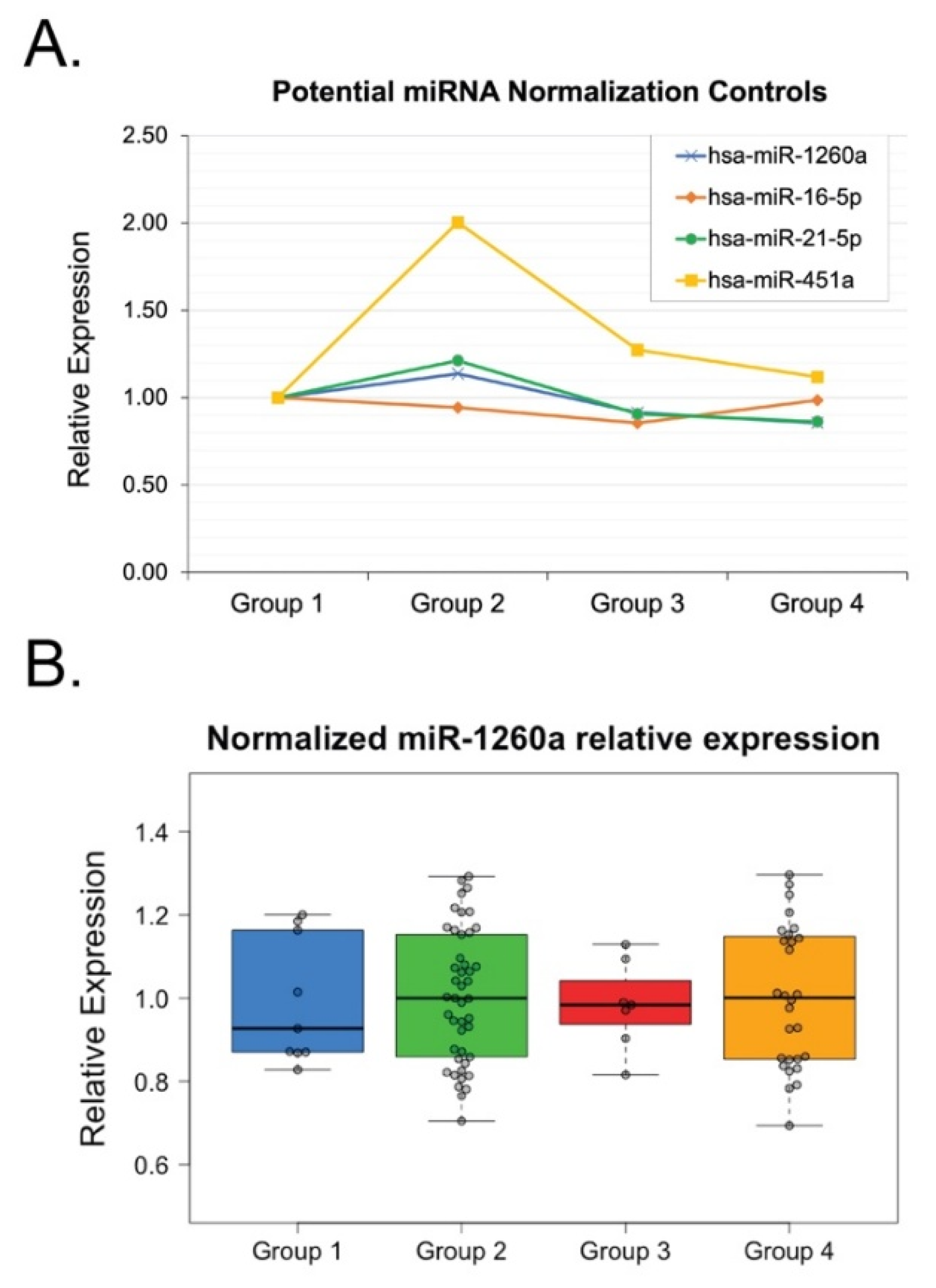
The group-based microarray analysis identified 164 miRNAs, 4 of which showed the most stable expression and were identified as a potential normalization controls: miR-16-5p, miR-21-5p, miR-451a, and miR-1260a (
Figure 3). Analysis of these miRNAs in all individual samples revealed that the most stable expression was demonstrated by miR-16-5p and miR-1260a (
Figure 3). For further qPCR data analysis, miR-16-5p was used as a normalization control. Out of 164 analyzed miRNAs, 22 showed at least a 50-fold difference in relative expression between at least two groups. These 22 miRNAs were followed up with individual-level measurement via qPCR. Analysis of individual samples showed no statistically significant difference between groups of patients for 18 out of the remaining 22 miRNAs (
Figure S2). As presented in
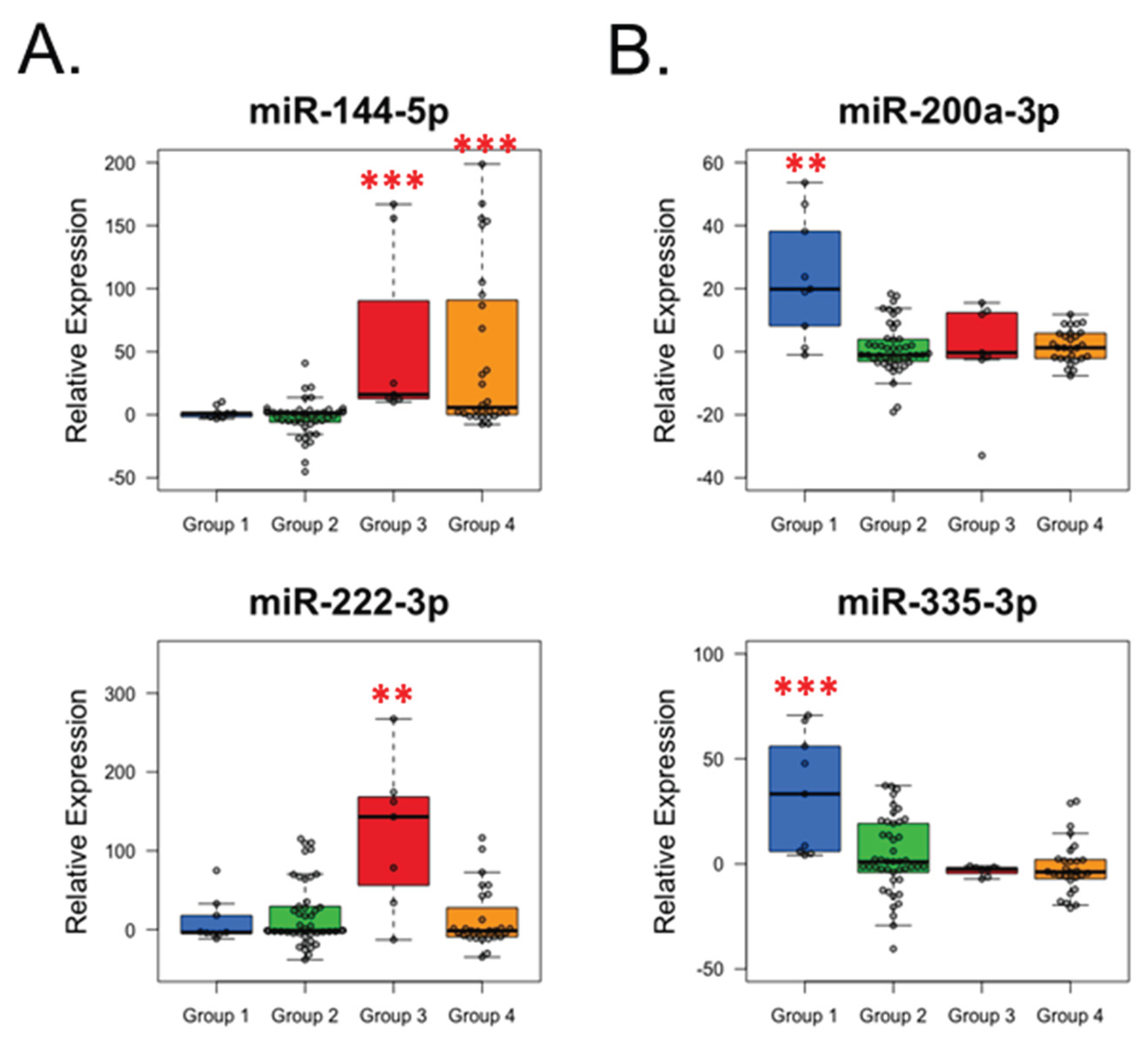
Figure 4 and
Figure 4 of 22 miRNAs (miR-144-5p, miR-200a-3p, miR-222-3p, and miR-335-3p) demonstrated statistically significant differential expression across the analyzed groups.
Two miRNAs exhibited significantly elevated expression in the acute and late high post-radiotherapy (post-RT) toxicity groups (Groups 3 and 4) (
Figure 4A). Specifically, miR-144-5p showed markedly higher relative expression in Group 3 (57.04 ± 71.49,
p < 0.0001) and Group 4 (46.82 ± 64.41,
p < 0.0001) compared with Group 1 (2.73 ± 3.69) and Group 2 (3.60 ± 7.58). MiR-222-3p was significantly overexpressed exclusively in the acute high post-RT toxicity group (Group 3; 122.78 ± 91.67,
p < 0.05) relative to Group 1 (14.12 ± 25.26), Group 2 (23.64 ± 34.84), and Group 4 (18.33 ± 33.45). In contrast, two miRNAs were significantly upregulated in the minimal toxicity group (Group 1) (
Figure 4B). MiR-200a-3p demonstrated higher relative expression in Group 1 (23.19 ± 19.16,
p < 0.05) compared with Group 2 (3.71 ± 5.37), Group 3 (6.32 ± 6.54), and Group 4 (3.31 ± 3.54). Similarly, miR-335-3p expression was significantly increased in Group 1 (32.09 ± 29.49,
p < 0.0001) relative to Group 2 (8.82 ± 12.14), Group 3 (0.46 ± 0.29), and Group 4 (4.18 ± 8.38).
4. Discussion
NTA demonstrated that the isolated particles exhibited a mean diameter consistent with EVs. Based on adherence to the SEC protocol and the observed particle size distribution, the isolated vesicles are presumed to be predominantly exosomes. However, definitive vesicle subtype identification cannot be achieved by NTA alone, as this technique does not discriminate EV subtypes based on defining characteristics such as surface marker expression [
22].
Although the particle size and concentration observed in plasma in the present study are comparable to those reported in some previous publications, substantial variability has been described in the literature. Reported exosome concentrations range from 0.88×10
8 to 13.38×10
8 exosomes/mL of plasma in certain studies, whereas others have reported concentrations between 8.35×10
9 and 10.9×10
9 exosomes/mL in breast cancer patients, with mean particle diameters of approximately 95 nm in localized disease and 125 nm in metastatic disease [
23,
24].
Differences in reported exosome concentrations are likely attributable, at least in part, to variability in isolation methodologies [
25,
26]. For example, a study employing the PureExo system with serum from healthy donors reported recovery of only 7.1×10
7 particles per 0.5 mL of input material [
27]. Additional factors contributing to variability in EV concentration and morphology include the use of serum versus plasma, differences in patient disease status, variability in particle characterization techniques, and deviations in protocol execution [
28,
29].
In the present study, a combination of SEC and the PureExo isolation kit was selected to achieve high-purity exosome preparations suitable for downstream miRNA isolation and quantitative PCR analysis [
27]. NTA performed on native plasma and isolated exosome fractions demonstrated that this combined isolation strategy enabled recovery of more than 90% of total plasma exosomes (
Figure 2).
Several previous studies have identified specific miRNAs as suitable endogenous controls for normalization of miRNA expression data. In particular, let-7a-5p, miR-21-5p, and miR-16-5p have been reported to exhibit stable expression across healthy, benign, and malignant tissues [
30]. In the present study, miR-16-5p demonstrated the highest expression stability in exosomes isolated from plasma samples and was therefore selected as the endogenous control, consistent with prior reports supporting its suitability for studies of metastatic breast cancer [
30]. These miRNAs are commonly referred to as “housekeeping” miRNAs, reflecting their involvement in fundamental cellular processes and their evolutionary conservation, which contributes to relatively uniform expression across cell types and conditions [
31]. The use of such stable reference miRNAs enables normalization of expression data to account for biological variability and technical variation introduced during sample processing and analysis [
31].
This study further identified two exosomal miRNAs that may represent candidate biomarkers associated with increased susceptibility to radiation-induced normal tissue toxicity in breast cancer patients. Exosomal miR-222-3p was upregulated exclusively in patients who developed acute high-grade toxicity, whereas miR-144-5p exhibited increased expression in both acute and late high-toxicity groups. Notably, both miRNAs are predicted to target manganese superoxide dismutase (SOD2), also known as manganese-dependent superoxide dismutase, a key mitochondrial antioxidant enzyme responsible for the detoxification of reactive oxygen species. SOD2 plays a critical role in mitigating oxidative stress induced by ionizing radiation and inflammatory cytokines [
32,
33]. Upregulation of miR-144-5p and miR-222-3p targeting this pathway may therefore contribute to impaired oxidative stress responses and heightened radiation sensitivity in normal tissues.
An additional finding of this study was the identification of two exosomal miRNAs, miR-222-3p and miR-335-3p, that were associated with patients who exhibited no or minimal normal tissue toxicity following radiotherapy. Although this subgroup included a relatively small number of patients, the observed differential expression of these miRNAs may reflect biological mechanisms contributing to tissue tolerance or adaptive responses to radiation-induced injury. Validation of these associations and clarification of their functional significance will require investigation in larger, independent cohorts.
The significance of the present findings lies in the identification of novel candidate biomarkers associated with radiotherapy-induced toxicity. Improved prediction of individual patient susceptibility to radiotoxicity could ultimately support treatment personalization and optimization of radiotherapy planning. Previous studies addressing tumor radioresistance have suggested that radiotherapy may be less effective in patients whose tumors express miRNAs associated with resistance [
3]. Our findings extend this concept by suggesting that patients exhibiting specific miRNA expression profiles associated with severe radiation-induced toxicity may benefit from individualized treatment modifications, such as optimization of radiotherapy techniques, adjustment of treatment planning parameters, or implementation of proactive supportive care strategies prior to the onset of clinical side effects. There is also a potential future in the use of miRNA as a therapeutic agent for sensitizing tumor tissues to radiation while protecting surrounding tissues. Administration of miRNAs associated with minimal adverse reaction or perhaps inhibiting those that promote toxicity are possible avenues for therapy. Beyond breast cancer, the data obtained in this study may have broader implications for other types of cancers as well as individuals exposed to radiation in other scenarios, such as industrial accidents [
34].
To date, few studies are directly comparable to the present investigation. Existing studies examining miRNA biomarkers associated with radiotherapy-induced toxicity in breast cancer patients frequently employ different isolation and analytical methodologies or focus on circulating miRNAs not encapsulated within EVs. In addition, many published studies include patient cohorts treated with heterogeneous radiotherapy doses and fractionation schedules and assess toxicity using non-uniform criteria, further limiting comparability across studies. For example, Esplugas et al. evaluated cardiotoxicity in relation to circulating miRNAs that are not EV-associated, rather than exosomal miRNAs [
19]. Other studies have compared patients with and without radiotherapy exposure but have not stratified toxicity outcomes according to temporal patterns, such as acute versus late effects [
7]. These methodological and clinical differences likely contribute to the limited availability of directly comparable data. Reproducibility remains a major challenge in exosomal miRNA research, owing in part to the wide range of available EV isolation techniques, reagents, and analytical platforms [
35]. Furthermore, there is currently no consensus regarding the optimal endogenous miRNAs for normalization, which further complicates cross-study validation [
35]. Addressing these technical and methodological limitations will be essential before exosomal miRNA biomarkers can be reliably translated into clinical practice.
Overall, further investigation of the miRNAs identified in this study is warranted to validate their association with radiotherapy-induced toxicity and to elucidate their mechanistic roles. Future studies employing standardized isolation, normalization, and analytical protocols, similar to those used here, will facilitate cross-study comparisons and strengthen the evidence base needed to advance exosomal miRNAs toward clinical application.
Supplementary Materials
The following supporting information can be downloaded at the website of this paper posted on Preprints.org, Figure S1: “Melting curves for miRNAs extracted from plasma exosomes of the breast cancer patients”, Figure S2: “qPCR analysis of miRNAs extracted from plasma exosomes”.
Author Contributions
Conceptualization, B.Q. and V.Y.; methodology, B.Q., D.A., and V.Y.; software, E.Z. and M.M.; validation, B.Q., M.M. and V.Y.; formal analysis, M.M. and V.Y.; investigation, M.M., C.R., and V.Y.; resources, D.A. and V.A.; data curation, M.M. and V.Y.; writing—original draft preparation, M.M. and V.Y; writing—review and editing, B.Q.; visualization, M.M. and V.Y; supervision, E.Z. and D.A.; project administration, D.A.; funding acquisition, V.Y. All authors have read and agreed to the published version of the manuscript.
Funding
EM analysis was performed by the Microscopy Core of VCU Massey Comprehensive Cancer Center funded from NIH-NCI Cancer Center Support Grant P30 CA016059. Dr. Yakovlev was supported by the internal Value Efficiency Training and Research (VETAR) fund of Virginia Commonwealth University.
Institutional Review Board Statement
The study was conducted in accordance with the Declaration of Helsinki, and approved by the Institutional Review Board of Virginia Commonwealth University (Protocol #MCC-12181).
Informed Consent Statement
Written informed consent has been obtained from the patient(s) to publish this paper.
Data Availability Statement
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Acknowledgments
The authors have reviewed and edited the output and take full responsibility for the content of this publication.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| EVs |
Extracellular Vesicles |
| SEC |
Size Exclusion Chromatography |
| NTA |
Nanoparticle tracking analysis |
| TEM |
Transmission Electron Microscopy |
| SOD2 |
manganese superoxide dismutase 2 |
References
- Siegel, R.L.; Giaquinto, A.N.; Jemal, A. Cancer statistics, 2024. CA Cancer J Clin 2024, 74, 12–49. [Google Scholar] [CrossRef]
- Sun, Y.S.; Zhao, Z.; Yang, Z.N.; Xu, F.; Lu, H.J.; Zhu, Z.Y.; Shi, W.; Jiang, J.; Yao, P.P.; Zhu, H.P. Risk Factors and Preventions of Breast Cancer. Int J Biol Sci 2017, 13, 1387–1397. [Google Scholar] [CrossRef]
- Barnett, G.C.; West, C.M.; Dunning, A.M.; Elliott, R.M.; Coles, C.E.; Pharoah, P.D.; Burnet, N.G. Normal tissue reactions to radiotherapy: towards tailoring treatment dose by genotype. Nat Rev Cancer 2009, 9, 134–142. [Google Scholar] [CrossRef] [PubMed]
- Kiwerska, K.; Szyfter, K. DNA repair in cancer initiation, progression, and therapy-a double-edged sword. J Appl Genet 2019, 60, 329–334. [Google Scholar] [CrossRef]
- Shaitelman, S.F.; Schlembach, P.J.; Arzu, I.; Ballo, M.; Bloom, E.S.; Buchholz, D.; Chronowski, G.M.; Dvorak, T.; Grade, E.; Hoffman, K.E.; et al. Acute and Short-term Toxic Effects of Conventionally Fractionated vs Hypofractionated Whole-Breast Irradiation: A Randomized Clinical Trial. JAMA Oncol 2015, 1, 931–941. [Google Scholar] [CrossRef]
- Speers, C.; Pierce, L.J. Molecular Signatures of Radiation Response in Breast Cancer: Towards Personalized Decision-Making in Radiation Treatment. Int J Breast Cancer 2017, 2017, 4279724. [Google Scholar] [CrossRef]
- Stepanovic, A.; Nikitovic, M.; Stanojkovic, T.P.; Grujicic, D.; Bukumiric, Z.; Srbljak, I.; Ilic, R.; Milosevic, S.; Arsenijevic, T.; Petrovic, N. Association between microRNAs 10b/21/34a and acute toxicity in glioblastoma patients treated with radiotherapy and temozolomide. Sci Rep 2022, 12, 7505. [Google Scholar] [CrossRef]
- Chong, Z.X.; Yeap, S.K.; Ho, W.Y. Roles of circulating microRNA(s) in human breast cancer. Arch Biochem Biophys 2020, 695, 108583. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Odenthal, M.; Fries, J.W. Exosomes as miRNA Carriers: Formation-Function-Future. Int J Mol Sci 2016, 17. [Google Scholar] [CrossRef] [PubMed]
- Ingenito, F.; Roscigno, G.; Affinito, A.; Nuzzo, S.; Scognamiglio, I.; Quintavalle, C.; Condorelli, G. The Role of Exo-miRNAs in Cancer: A Focus on Therapeutic and Diagnostic Applications. Int J Mol Sci 2019, 20. [Google Scholar] [CrossRef]
- Doyle, L.M.; Wang, M.Z. Overview of Extracellular Vesicles, Their Origin, Composition, Purpose, and Methods for Exosome Isolation and Analysis. Cells 2019, 8. [Google Scholar] [CrossRef]
- Stahl, A.L.; Johansson, K.; Mossberg, M.; Kahn, R.; Karpman, D. Exosomes and microvesicles in normal physiology, pathophysiology, and renal diseases. Pediatr Nephrol 2019, 34, 11–30. [Google Scholar] [CrossRef]
- Bunggulawa, E.J.; Wang, W.; Yin, T.; Wang, N.; Durkan, C.; Wang, Y.; Wang, G. Recent advancements in the use of exosomes as drug delivery systems. J Nanobiotechnology 2018, 16, 81. [Google Scholar] [CrossRef]
- Famta, P.; Shah, S.; Khatri, D.K.; Guru, S.K.; Singh, S.B.; Srivastava, S. Enigmatic role of exosomes in breast cancer progression and therapy. Life Sci 2022, 289, 120210. [Google Scholar] [CrossRef] [PubMed]
- Gurung, S.; Perocheau, D.; Touramanidou, L.; Baruteau, J. The exosome journey: from biogenesis to uptake and intracellular signalling. Cell Commun Signal 2021, 19, 47. [Google Scholar] [CrossRef] [PubMed]
- Alimirzaie, S.; Bagherzadeh, M.; Akbari, M.R. Liquid biopsy in breast cancer: A comprehensive review. Clin Genet 2019, 95, 643–660. [Google Scholar] [CrossRef]
- Grinan-Lison, C.; Olivares-Urbano, M.A.; Jimenez, G.; Lopez-Ruiz, E.; Del Val, C.; Morata-Tarifa, C.; Entrena, J.M.; Gonzalez-Ramirez, A.R.; Boulaiz, H.; Zurita Herrera, M.; et al. miRNAs as radio-response biomarkers for breast cancer stem cells. Mol Oncol 2020, 14, 556–570. [Google Scholar] [CrossRef]
- Pajic, M.; Froio, D.; Daly, S.; Doculara, L.; Millar, E.; Graham, P.H.; Drury, A.; Steinmann, A.; de Bock, C.E.; Boulghourjian, A.; et al. miR-139-5p Modulates Radiotherapy Resistance in Breast Cancer by Repressing Multiple Gene Networks of DNA Repair and ROS Defense. Cancer Res 2018, 78, 501–515. [Google Scholar] [CrossRef] [PubMed]
- Esplugas, R.; Arenas, M.; Serra, N.; Belles, M.; Bonet, M.; Gascon, M.; Vallve, J.C.; Linares, V. Effect of radiotherapy on the expression of cardiovascular disease-related miRNA-146a, -155, -221 and -222 in blood of women with breast cancer. PLoS One 2019, 14, e0217443. [Google Scholar] [CrossRef]
- McGinn, M.; Rabender, C.; Mikkelsen, R.; Yakovlev, V. Hepatocyte-derived extracellular vesicles regulate liver regeneration through a negative feedback mechanism. J Extracell Biol 2024, 3, e70023. [Google Scholar] [CrossRef]
- Jung, M.K.; Mun, J.Y. Sample Preparation and Imaging of Exosomes by Transmission Electron Microscopy. J Vis Exp 2018. [Google Scholar] [CrossRef]
- Di Bella, M.A. Overview and Update on Extracellular Vesicles: Considerations on Exosomes and Their Application in Modern Medicine. Biology (Basel) 2022, 11. [Google Scholar] [CrossRef]
- Konoshenko, M.; Sagaradze, G.; Orlova, E.; Shtam, T.; Proskura, K.; Kamyshinsky, R.; Yunusova, N.; Alexandrova, A.; Efimenko, A.; Tamkovich, S. Total Blood Exosomes in Breast Cancer: Potential Role in Crucial Steps of Tumorigenesis. Int J Mol Sci 2020, 21. [Google Scholar] [CrossRef]
- Vardaki, I.; Ceder, S.; Rutishauser, D.; Baltatzis, G.; Foukakis, T.; Panaretakis, T. Periostin is identified as a putative metastatic marker in breast cancer-derived exosomes. Oncotarget 2016, 7, 74966–74978. [Google Scholar] [CrossRef]
- El-Khoury, V.; Pierson, S.; Kaoma, T.; Bernardin, F.; Berchem, G. Assessing cellular and circulating miRNA recovery: the impact of the RNA isolation method and the quantity of input material. Sci Rep 2016, 6, 19529. [Google Scholar] [CrossRef] [PubMed]
- Sriram, H.; Khanka, T.; Kedia, S.; Tyagi, P.; Ghogale, S.; Deshpande, N.; Chatterjee, G.; Rajpal, S.; Patkar, N.V.; Subramanian, P.G.; et al. Improved protocol for plasma microRNA extraction and comparison of commercial kits. Biochem Med (Zagreb) 2021, 31, 030705. [Google Scholar] [CrossRef] [PubMed]
- Baranyai, T.; Herczeg, K.; Onodi, Z.; Voszka, I.; Modos, K.; Marton, N.; Nagy, G.; Mager, I.; Wood, M.J.; El Andaloussi, S.; et al. Isolation of Exosomes from Blood Plasma: Qualitative and Quantitative Comparison of Ultracentrifugation and Size Exclusion Chromatography Methods. PLoS One 2015, 10, e0145686. [Google Scholar] [CrossRef] [PubMed]
- Helwa, I.; Cai, J.; Drewry, M.D.; Zimmerman, A.; Dinkins, M.B.; Khaled, M.L.; Seremwe, M.; Dismuke, W.M.; Bieberich, E.; Stamer, W.D.; et al. A Comparative Study of Serum Exosome Isolation Using Differential Ultracentrifugation and Three Commercial Reagents. PLoS One 2017, 12, e0170628. [Google Scholar] [CrossRef]
- Wang, K.; Yuan, Y.; Cho, J.H.; McClarty, S.; Baxter, D.; Galas, D.J. Comparing the MicroRNA spectrum between serum and plasma. PLoS One 2012, 7, e41561. [Google Scholar] [CrossRef]
- Rinnerthaler, G.; Hackl, H.; Gampenrieder, S.P.; Hamacher, F.; Hufnagl, C.; Hauser-Kronberger, C.; Zehentmayr, F.; Fastner, G.; Sedlmayer, F.; Mlineritsch, B.; et al. miR-16-5p Is a Stably-Expressed Housekeeping MicroRNA in Breast Cancer Tissues from Primary Tumors and from Metastatic Sites. Int J Mol Sci 2016, 17. [Google Scholar] [CrossRef]
- Rice, J.; Roberts, H.; Rai, S.N.; Galandiuk, S. Housekeeping genes for studies of plasma microRNA: A need for more precise standardization. Surgery 2015, 158, 1345–1351. [Google Scholar] [CrossRef]
- Becuwe, P.; Ennen, M.; Klotz, R.; Barbieux, C.; Grandemange, S. Manganese superoxide dismutase in breast cancer: from molecular mechanisms of gene regulation to biological and clinical significance. Free Radic Biol Med 2014, 77, 139–151. [Google Scholar] [CrossRef]
- Pias, E.K.; Ekshyyan, O.Y.; Rhoads, C.A.; Fuseler, J.; Harrison, L.; Aw, T.Y. Differential effects of superoxide dismutase isoform expression on hydroperoxide-induced apoptosis in PC-12 cells. J Biol Chem 2003, 278, 13294–13301. [Google Scholar] [CrossRef] [PubMed]
- Halimi, M.; Shahabi, A.; Moslemi, D.; Parsian, H.; Asghari, S.M.; Sariri, R.; Yeganeh, F.; Zabihi, E. Human serum miR-34a as an indicator of exposure to ionizing radiation. Radiat Environ Biophys 2016, 55, 423–429. [Google Scholar] [CrossRef] [PubMed]
- Moldovan, L.; Batte, K.E.; Trgovcich, J.; Wisler, J.; Marsh, C.B.; Piper, M. Methodological challenges in utilizing miRNAs as circulating biomarkers. J Cell Mol Med 2014, 18, 371–390. [Google Scholar] [CrossRef] [PubMed]
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).