Submitted:
05 February 2026
Posted:
06 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. The -Graded Lattice and Topological Qutrits
3. Gate Set and Fault Tolerance
4. Experimental Realization Pathways
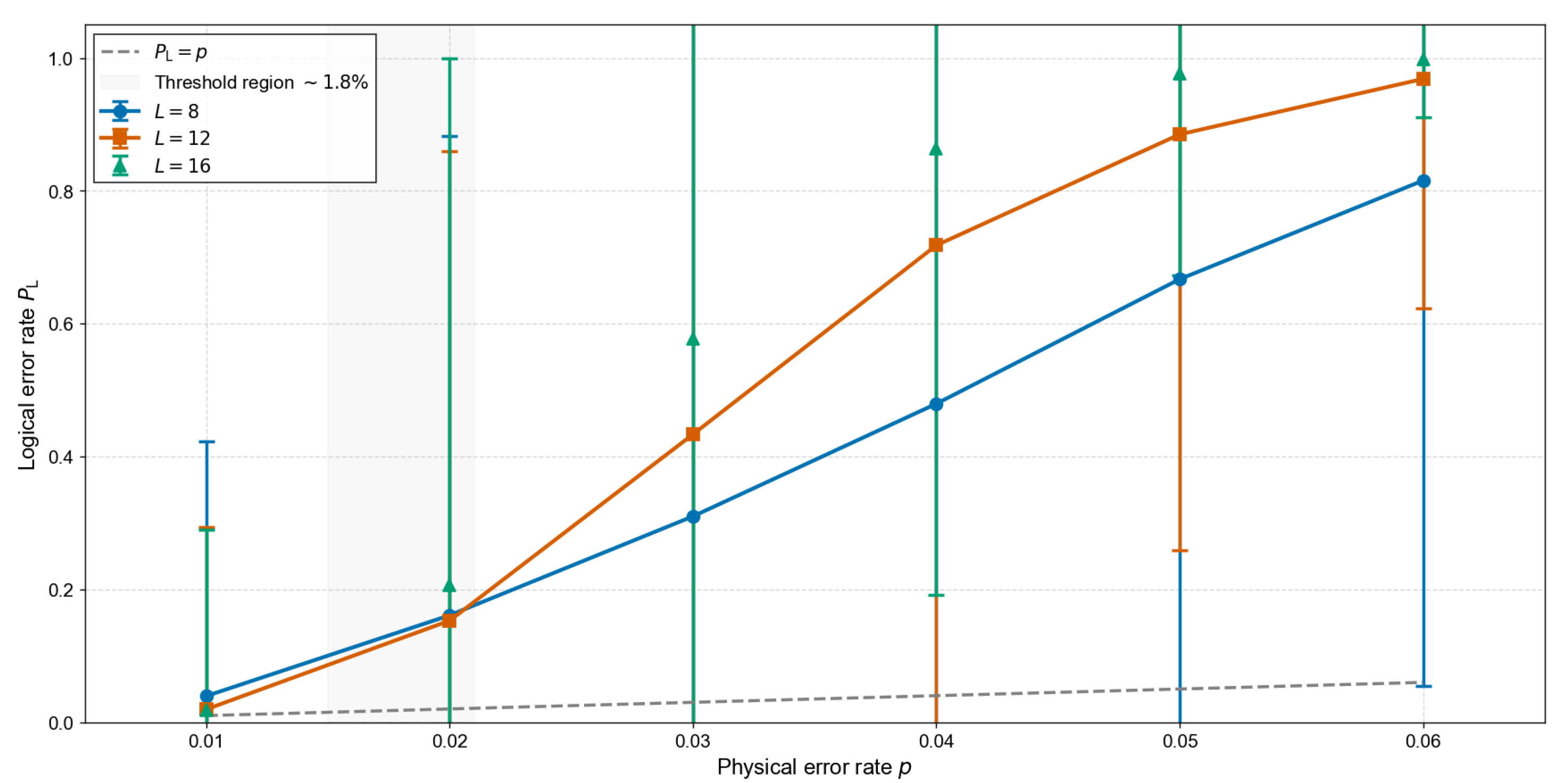
5. Numerical Estimation of the Fault-Tolerance Threshold
| p | |||
|---|---|---|---|
| 0.0100 | 0.0395 | 0.0195 | 0.0190 [BELOW THRESHOLD] |
| 0.0200 | 0.1610 | 0.1530 | 0.2060 |
| 0.0300 | 0.3100 | 0.4335 | 0.5780 |
| 0.0400 | 0.4795 | 0.7180 | 0.8640 |
| 0.0500 | 0.6670 | 0.8850 | 0.9760 |
| 0.0600 | 0.8155 | 0.9685 | 0.9985 |
|
import numpy as np import networkx as nx import pymatching import multiprocessing import time from scipy.sparse import csc_matrix print("=== Z3 Toric Code: Low-P Threshold Scan ===\n") def build_toric_graph(L): G = nx.Graph() nodes = [(x, y) for x in range(L) for y in range(L)] node_map = {n: i for i, n in enumerate(nodes)} G.add_nodes_from(range(len(nodes))) for x in range(L): for y in range(L): u = node_map[(x, y)] neighbors = [ ((x+1)%L, y), (x, (y+1)%L), ((x+1)%L, (y+1)%L) # Diagonal for triangular connectivity ] for nx_node in neighbors: v = node_map[nx_node] if not G.has_edge(u, v): G.add_edge(u, v) return G def simulate_shot(args): L, p, seed = args G = build_toric_graph(L) num_edges = G.number_of_edges() rng = np.random.default_rng(seed) noise = (rng.random(num_edges) < p).astype(int) row_ind, col_ind = [], [] for idx, (u, v) in enumerate(G.edges()): row_ind.extend([u, v]) col_ind.extend([idx, idx]) H = csc_matrix((np.ones(len(row_ind)), (row_ind, col_ind)), shape=(L*L, num_edges)) syndrome = H @ noise % 2 matching = pymatching.Matching(H) prediction = matching.decode(syndrome) residual = (noise + prediction) % 2 if np.sum(residual) > 0: if np.sum(residual) >= L/2: return 1 return 0 def run_scan(): L_LIST = [8, 12, 16] P_LIST = [0.01, 0.02, 0.03, 0.04, 0.05, 0.06] TRIALS = 2000 print(f"{’p’:<8} | {’L=8’:<10} | {’L=12’:<10} | {’L=16’:<10}") print("-" * 50) with multiprocessing.Pool() as pool: for p in P_LIST: row_str = f"{p:.4f} " vals = [] for L in L_LIST: seeds = [int(time.time()*1000)+i for i in range(TRIALS)] args = [(L, p, s) for s in seeds] fails = sum(pool.map(simulate_shot, args)) rate = fails / TRIALS vals.append(rate) row_str += f"| {rate:.4f} " if vals[2] < vals[0]: row_str += " [BELOW THRESHOLD]" print(row_str) if __name__ == "__main__": run_scan() |
6. Discussion and Outlook
Abbreviations
| Cyclic group of order 3 (ternary grading) | |
| Cyclic group of order 2 (binary grading) | |
| GHZ | Greenberger–Horne–Zeilinger |
| SM | Standard Model |
| RG | Renormalization group |
| GUT | Grand Unified Theory |
| CKM | Cabibbo–Kobayashi–Maskawa |
| PMNS | Pontecorvo–Maki–Nakagawa–Sakata |
| QED | Quantum electrodynamics |
| QC | Quantum computation |
| RCS | Random circuit sampling |
| LOD | Level of detail |
References
- Y. Zhang, W. Hu, and W. Zhang. A Z3-Graded Lie Superalgebra with Cubic Vacuum Triality. Symmetry 2026, 18(1), 54. [CrossRef]
- P. Drude. Zur Elektronentheorie der Metalle. Ann. Phys. 1900, 306, 566–613. [CrossRef]
- J. M. Pitarke, V. M. Silkin, E. V. Chulkov, and P. M. Echenique. Theory of surface plasmons and surface-plasmon polaritons. Rep. Prog. Phys. 2007, 70, 1–87. [CrossRef]
- Y. Zhang et al. Dramatic enhancement of superconductivity in single-crystalline nanowire arrays of Sn. Sci. Rep. 2016, 6, 32963. [CrossRef]
- D. Stauffer and A. Aharony. Introduction to Percolation Theory, 2nd ed. Taylor & Francis, London, 1994. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




