Submitted:
05 February 2026
Posted:
06 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Material and Methods
2.1. Study Design and Patient Cohort
2.2. Tissue and Urine Sample Collection
2.3. Multiparametric MRI Acquisition and Interpretation
2.4. Urinary Proteomic Sample Preparation
2.5. NanoLC–MS/MS And DIA Proteomic Analysis
2.6. Protein Identification and Quantification
2.7. Statistical and Differential Expression Analysis
2.8. Comparative Tissue-Urine Proteomic Analysis
2.9. Survival Analysis Using the TCGA-PRAD Cohort
3. Results
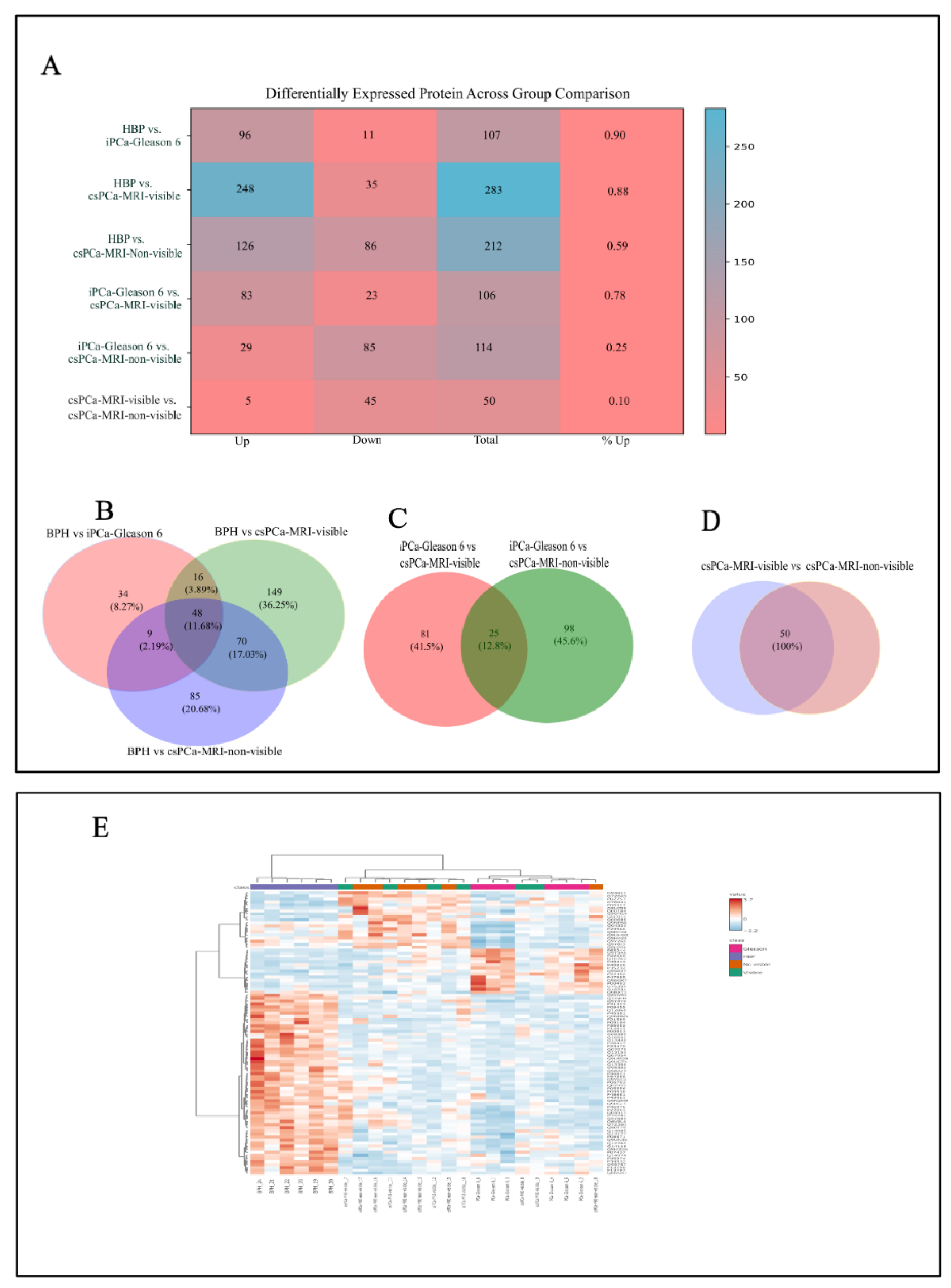
3.1. Tissue Proteome: Pairwise Comparisons
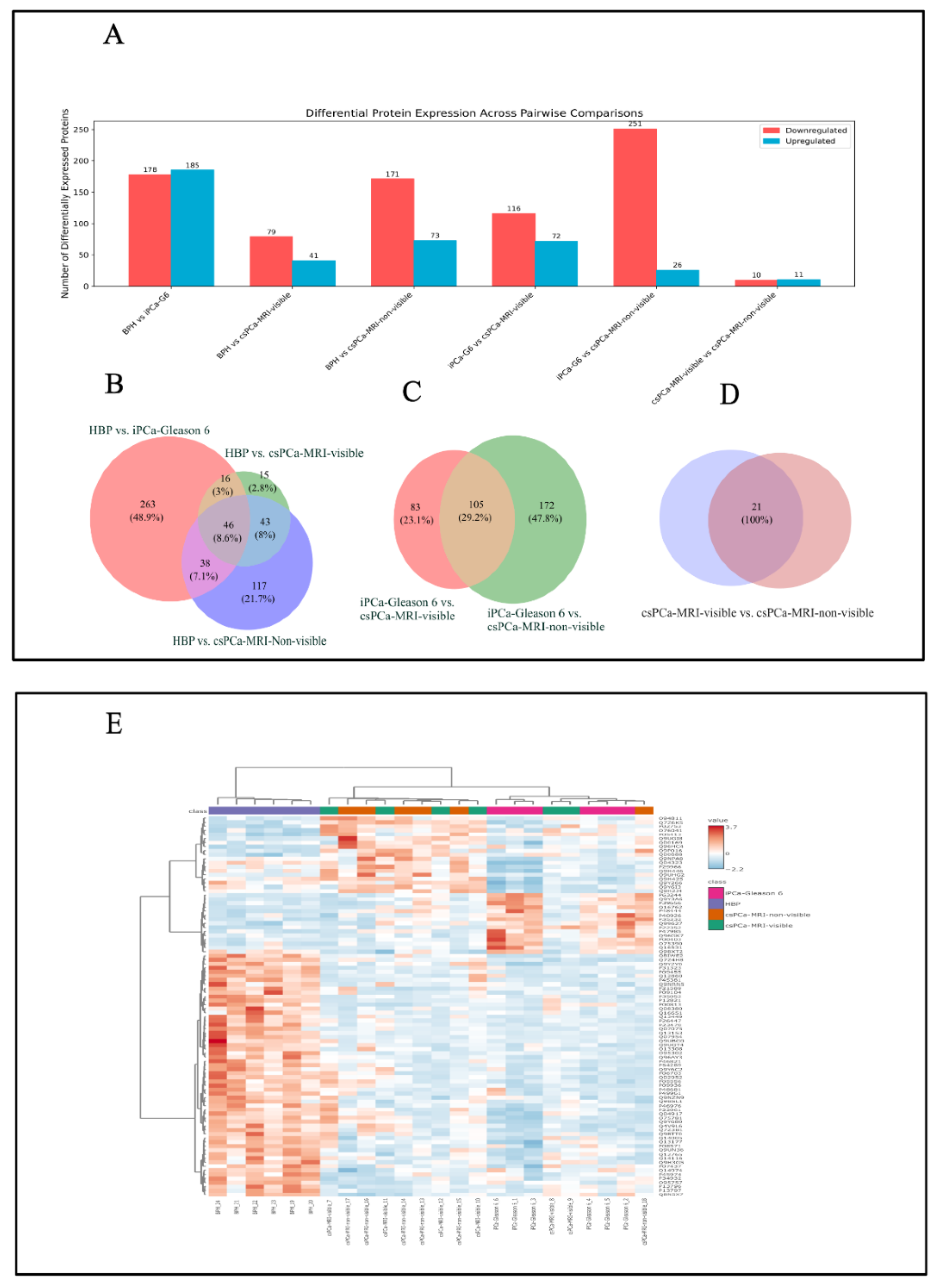
3.2. Tissue Proteome: Overlap and ANOVA
3.3. Urinary Proteome: Overlap and ANOVA
| Functional category | UniProt codes |
|---|---|
| Cytoskeleton & motility | O75503, P59998, P07360, Q07075, P13797, P26447, P55083, P13796, P05787, P08729, Q9BXS5 |
| Translation/RNA | P05387, P05386, P15586, O00264, P38571, Q12841, Q14108, P13639, P62330, Q9Y3B3, Q14240 |
| Metabolism/Redox | Q02083, P23526, P22392, Q00796, P00338, P00441, P09417, P14618 |
| Signaling & regulation | P22352, P06454, Q9BY67, Q14847, P08294, P08582, Q02952 |
| Cell adhesion/ECM | Q12860, Q16610, P51884, P05556, P16070, P43121 |
| Immunity/Inflammation | P80188, P05362, P04439, P08236, P06702, P19801 |
| Vesicle trafficking/Endocytosis | Q9UMX5, Q7Z3B1, Q9BRA2, P51149, P08962, P11234 |
| ER stress/Protein folding | O43598, Q14894, P34932, Q9UM22, P18827 |
| Protease regulation/Innate defense | P30740, P07384, P21291, P81605 |
| Lipid/Small-molecule transport | P02753, P05413, P02654, P12724 |
| Ubiquitin-proteasome system | Q9BRT3, Q9Y5K6, O14618 |
| Cell cycle/DNA replication & repair | P41222, Q9NR45 |
| Cell cycle/Nucleotide metabolism | P61916, P07996 |
| Mitochondria/Stress response | Q15185, Q92485 |
| Peptidase/Extracellular processing | P12821, P08473 |
| Lysosome/Autophagy | P13473 |
| Membrane organization | Q9UQB8 |
| Cell death/Differentiation | P31944 |
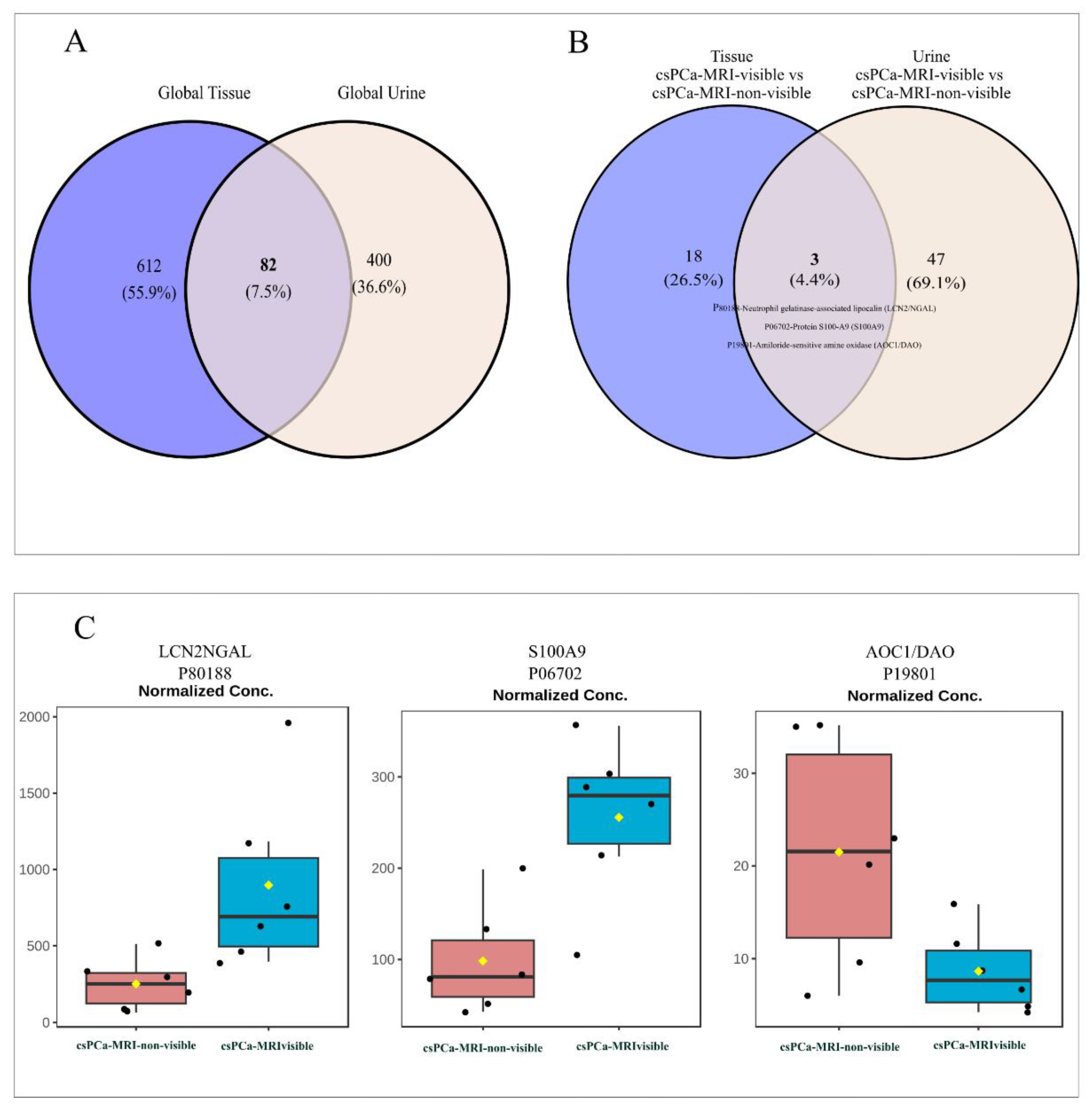
3.4. MRI-Visible vs. MRI-Non-Visible Overlap: Differential Urinary Protein Abundance
3.5. TCGA Validation
4. Discussion
Limitations
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Acknowledgments
Conflicts of Interest
References
- Siegel, R.L., A.N. Giaquinto, and A. Jemal, Cancer statistics, 2024. CA Cancer J Clin, 2024. 74(1): p. 12-49.
- Loeb, S., et al., Systematic review of complications of prostate biopsy. Eur Urol, 2013. 64(6): p. 876-92. [CrossRef]
- Barry, M.J., Screening for prostate cancer--the controversy that refuses to die. N Engl J Med, 2009. 360(13): p. 1351-4. [CrossRef]
- Sathianathen, N.J., et al., Negative Predictive Value of Multiparametric Magnetic Resonance Imaging in the Detection of Clinically Significant Prostate Cancer in the Prostate Imaging Reporting and Data System Era: A Systematic Review and Meta-analysis. Eur Urol, 2020. 78(3): p. 402-414. [CrossRef]
- Hamm, C.A., et al., Oncological Safety of MRI-Informed Biopsy Decision-Making in Men With Suspected Prostate Cancer. JAMA Oncol, 2025. 11(2): p. 145-153. [CrossRef]
- Haider, M.A., et al., Multiparametric Magnetic Resonance Imaging in the Diagnosis of Clinically Significant Prostate Cancer: an Updated Systematic Review. Clin Oncol (R Coll Radiol), 2021. 33(12): p. e599-e612. [CrossRef]
- Johnson, D.C., et al., Detection of Individual Prostate Cancer Foci via Multiparametric Magnetic Resonance Imaging. Eur Urol, 2019. 75(5): p. 712-720. [CrossRef]
- Pausch, A.M., et al., Risk factors for prostate cancer in men with false-negative mpMRI: A retrospective single center cohort study of image quality scores and clinical parameters. European Journal of Radiology, 2024. 170. [CrossRef]
- Schoots, I.G., et al., Analysis of Magnetic Resonance Imaging-directed Biopsy Strategies for Changing the Paradigm of Prostate Cancer Diagnosis. Eur Urol Oncol, 2020. 3(1): p. 32-41. [CrossRef]
- McNally, C.J., et al., Biomarkers That Differentiate Benign Prostatic Hyperplasia from Prostate Cancer: A Literature Review. Cancer Manag Res, 2020. 12: p. 5225-5241. [CrossRef]
- Iglesias-Gato, D., et al., The Proteome of Primary Prostate Cancer. Eur Urol, 2016. 69(5): p. 942-52.
- Crocetto, F., et al., Blood and urine-based biomarkers in prostate cancer: Current advances, clinical applications, and future directions. J Liq Biopsy, 2025. 9: p. 100305. [CrossRef]
- Bologna, E., et al., Tissue-Based Genomic Testing in Prostate Cancer: 10-Year Analysis of National Trends on the Use of Prolaris, Decipher, ProMark, and Oncotype DX. Clin Pract, 2024. 14(2): p. 508-520. [CrossRef]
- Zhou, B., et al., Quantitative proteomic analysis of prostate tissue specimens identifies deregulated protein complexes in primary prostate cancer. Clin Proteomics, 2019. 16: p. 15. [CrossRef]
- Ajami, T., et al., Assessing the Molecular Heterogeneity of Prostate Cancer Biopsy Sampling: Insights from the MAST Trial. Eur Urol Focus, 2025. 11(2): p. 287-293. [CrossRef]
- Fiorella, D., et al., Role of PCA3 and SelectMDx in the optimization of active surveillance in prostate cancer. Actas Urol Esp (Engl Ed), 2021. 45(6): p. 439-446.
- Frantzi, M., et al., Validation of a Urine-Based Proteomics Test to Predict Clinically Significant Prostate Cancer: Complementing mpMRI Pathway. Pathobiology, 2025. 92(2): p. 99-108. [CrossRef]
- Hatano, K. and K. Fujita, Extracellular vesicles in prostate cancer: a narrative review. Transl Androl Urol, 2021. 10(4): p. 1890-1907. [CrossRef]
- Frantzi, M., et al., Recent progress in urinary proteome analysis for prostate cancer diagnosis and management. Expert Rev Mol Diagn, 2015. 15(12): p. 1539-54. [CrossRef]
- Plas, S., et al., The impact of urine biomarkers for prostate cancer detection-A systematic state of the art review. Crit Rev Oncol Hematol, 2025. 210: p. 104699. [CrossRef]
- Latosinska, A., et al., Molecular Changes in Tissue Proteome during Prostate Cancer Development: Proof-of-Principle Investigation. Diagnostics (Basel), 2020. 10(9). [CrossRef]
- Khoo, A., et al., Prostate cancer multiparametric magnetic resonance imaging visibility is a tumor-intrinsic phenomena. J Hematol Oncol, 2022. 15(1): p. 48. [CrossRef]
- Bernardino, R., et al., Profiling of urinary extracellular vesicle protein signatures from patients with cribriform and intraductal prostate carcinoma in a cross-sectional study. Sci Rep, 2024. 14(1): p. 25065. [CrossRef]
- Heidegger, I., et al., Prediction of Clinically Significant Prostate Cancer by a Specific Collagen-related Transcriptome, Proteome, and Urinome Signature. Eur Urol Oncol, 2025. 8(3): p. 652-662. [CrossRef]
- Rossi, R., et al., Uncovering New Biomarkers for Prostate Cancer Through Proteomic and Network Analysis. Biology (Basel), 2025. 14(3). [CrossRef]
- Cancer Genome Atlas Research, N., The Molecular Taxonomy of Primary Prostate Cancer. Cell, 2015. 163(4): p. 1011-25.
- Pavlovic, B., et al., Urine biomarkers can predict prostate cancer and PI-RADS score prior to biopsy. Scientific Reports, 2024. 14(1): p. 18148. [CrossRef]
- Houlahan, K.E., et al., Molecular Hallmarks of Multiparametric Magnetic Resonance Imaging Visibility in Prostate Cancer. Eur Urol, 2019. 76(1): p. 18-23. [CrossRef]
- Carretero, F.J., et al., Frequent HLA class I alterations in human prostate cancer: molecular mechanisms and clinical relevance. Cancer Immunol Immunother, 2016. 65(1): p. 47-59. [CrossRef]
- Jasim, A., A. Mohammed, and A. Ibrahim, Functional Analysis of Beta 2 Microglobulin Protein in Patients with Prostate Cancer Using Bioinformatics Methods. Indian Journal of Public Health Research & Development, 2019. 10: p. 519. [CrossRef]
- Iczkowski, K.A., Cell adhesion molecule CD44: its functional roles in prostate cancer. Am J Transl Res, 2010. 3(1): p. 1-7.
- Su, C., et al., Integrinbeta-1 in disorders and cancers: molecular mechanisms and therapeutic targets. Cell Commun Signal, 2024. 22(1): p. 71. [CrossRef]
- Srivastava, M., et al., Diverse effects of ANXA7 and p53 on LNCaP prostate cancer cells are associated with regulation of SGK1 transcription and phosphorylation of the SGK1 target FOXO3A. Biomed Res Int, 2014. 2014: p. 193635. [CrossRef]
- Tan, S.H., et al., Prognostic features of Annexin A2 expression in prostate cancer. Pathology, 2021. 53(2): p. 205-213. [CrossRef]
- Yu, L., et al., Proteomics of Prostate Cancer Tissue Small Extracellular Vesicles Reveal Alteration of Metabolism. Proteomics, 2025: p. e70081. [CrossRef]
- Lima, T., et al., Application of Proteogenomics to Urine Analysis towards the Identification of Novel Biomarkers of Prostate Cancer: An Exploratory Study. Cancers (Basel), 2022. 14(8). [CrossRef]
- Chakraborty, S., et al., The multifaceted roles of neutrophil gelatinase associated lipocalin (NGAL) in inflammation and cancer. Biochim Biophys Acta, 2012. 1826(1): p. 129-69. [CrossRef]
- Muslu, N., et al., Neutrophil gelatinase-associated lipocalin as a screening test in prostate cancer. Turk J Urol, 2017. 43(1): p. 30-35. [CrossRef]
- Candido, S., et al., Roles of neutrophil gelatinase-associated lipocalin (NGAL) in human cancer. Oncotarget, 2014. 5(6): p. 1576-94. [CrossRef]
- Chen, Y., Z. Wu, and X. Yi, Elucidating the pan-oncologic landscape of S100A9: prognostic and therapeutic corollaries from an integrative bioinformatics and Mendelian randomization analysis. Scientific Reports, 2024. 14(1): p. 19071. [CrossRef]
- Chen, Y., Z. Wu, and X. Yi, Elucidating the pan-oncologic landscape of S100A9: prognostic and therapeutic corollaries from an integrative bioinformatics and Mendelian randomization analysis. Sci Rep, 2024. 14(1): p. 19071. [CrossRef]
- Tidehag, V., et al., High density of S100A9 positive inflammatory cells in prostate cancer stroma is associated with poor outcome. Eur J Cancer, 2014. 50(10): p. 1829-1835. [CrossRef]
- He, D., et al., Diamine oxidase acts as a novel risk factor in abnormal inflammation via mediating “cytosolic ROS-autophagy-IFN-gamma” axis in NK cells. Life Sci, 2025. 377: p. 123775. [CrossRef]
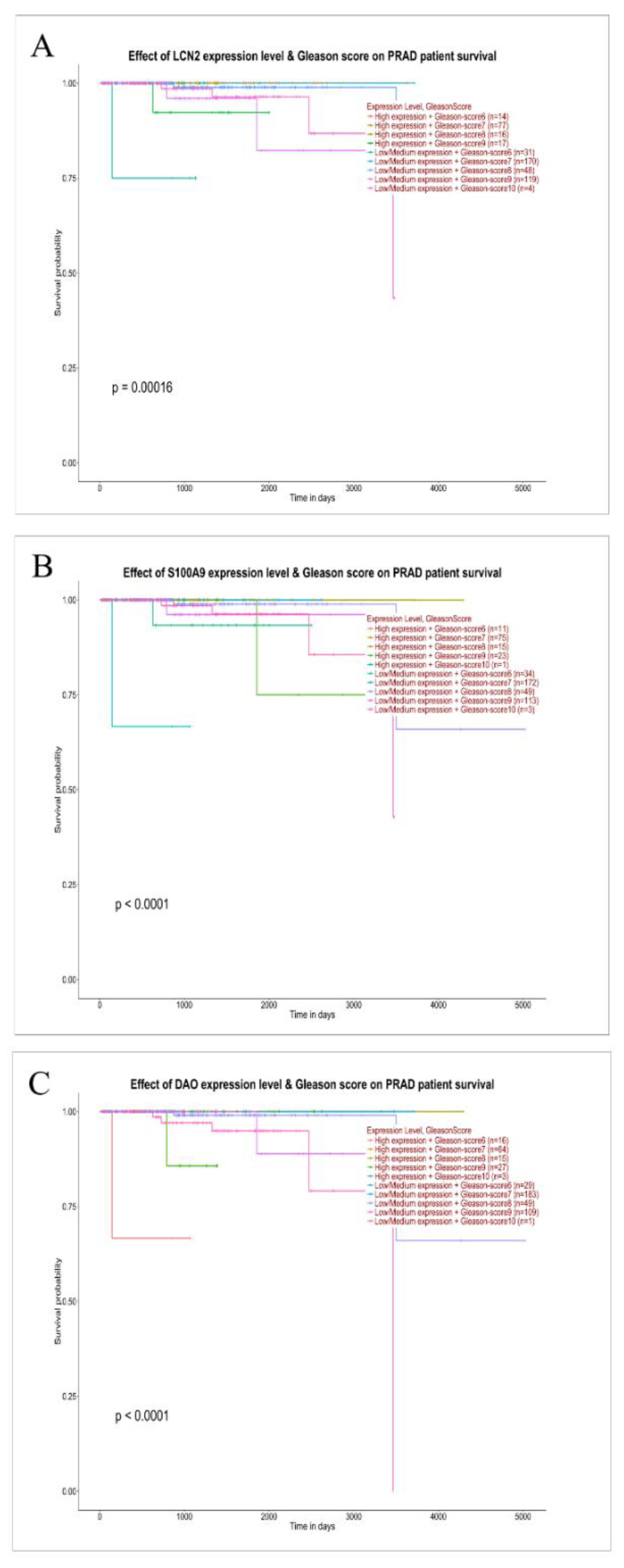
| Variable | Value |
|---|---|
| Age, years | 62.09 ± 6.07 (median: 63) |
| Total, PSA, ng/mL | 6.21 ± 3.52 (median: 4.9) |
| Free PSA, % | 13.88 ± 4.76 (median: 14) |
| Prostate volume, cc | 61.33 ± 40.32 (median: 50) |
| Benign prostatic hyperplasia (BPH), n (%) | 6 (25%) |
| iPCa-Gleason score 6, n (%) | 6 (25%) |
| cs-PCa-MRI-visible, n (%) | 6 (25%) |
| cs-PCa-MRI-non-visible, n (%) | 6 (25%) |
|
cs-PCa-MRI-visible, n (%) ISUP 2 3 |
6 (25%) 3 3 |
|
Cs-Pca-MRI-non-visible, n (%) ISUP 2 3 |
6 (25%) 4 2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




