Submitted:
05 February 2026
Posted:
06 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction and Background
1.1. Introduction
1.2. VERDICT MRI: Overview and Clinical Applications
Overview.
Clinical relevance.
Acquisition and practical considerations.
1.3. Computational Challenges in VERDICT Parameter Estimation
- 1.
- Computational complexity: expensive iterative solvers with costly forward models/Jacobians [16].
- 2.
- 3.
- Local minima and initialization sensitivity: non convexity yields inconsistent fits across starting points.
- 4.
- Noise sensitivity: high-b acquisitions are SNR-limited, degrading stability and biasing estimates [7].
- 5.
- Clinical feasibility: per-slice runtimes on the order of minutes hinder real-time mapping and workflow integration.
1.4. Deep Learning for Medical Parameter Prediction
1.5. Research Gap and Motivation
1.6. Thesis Contributions and Objectives
- Comprehensive architecture evaluation: feedforward, convolutional, recurrent, Transformer-based, and advanced (Variational Auto-encoder, Mixture of Experts) models under a unified pipeline.
- Standardized evaluation: rigorous protocols with bootstrap confidence intervals and significance testing for fair, meaningful comparisons.
- Reproducible platform: open-source implementations, configurations, and scripts for community use and extension.
2. Data
2.1. Overview and Rationale
Acquisition Scheme [14]
Model parameterization and constraints.
- Intracellular volume fraction (): with , enforcing ; a noninvasive surrogate for cellularity [1].
- Vascular volume fraction (): , indexing the fraction of signal arising from the vascular space.
- Intracellular diffusivity (): , reflecting apparent intracellular diffusion.
- Cell radius (R): , sensitive to cell size distribution/morphology [1].
- Extracellular axial diffusivity (): along the Zeppelin axis.
- Extracellular radial diffusivity (): in the deep learning implementation, this is obtained indirectly by predicting a multiplier , such that . This constrains by construction.
- Orientation angles: and define the Zeppelin axis [19].
- Vascular pseudo-diffusivity (): fixed to in this implementation (as in [14]), but still a parameter of the vascular compartment.
2.2. Acquisition Model and Units
2.3. VERDICT Signal Model
Restricted spheres (intra-cellular).
Hindered zeppelin (extra-cellular/extravascular).
Astrosticks (vascular).
2.4. Signal Synthesis for One Acquisition Scheme
- 1.
- Sample a parameter tuple .
- 2.
2.5. Context Within VERDICT Literature
3. Models and Methodology
3.1. Experimental Framework
3.2. Benchmark Neural Network Architectures
3.2.1. Multi-Layer Perceptron (MLP)
Implementation.
- Construct a list of layer widths .
- For each hidden transition , append Linear(,) and the chosen activation .
- Append the final Linear(,) without activation.
Capacity and complexity.
Instantiation in our benchmark.
3.2.2. ResNet
Motivation.
Architecture.
- 1.
- An input stem ,
- 2.
- Bresidual blocks with identity skip connections,
- 3.
- A linear head .
Implementation details.
- Stem: Linear(, d) followed by .
- Residual stack: B blocks, each with Linear(d, d)→→Linear(d, d), then skip-add and .
- Head: Linear(d, ) without activation.
Capacity and compute.
Training considerations.
Instantiation in our benchmark.
3.2.3. RNN-Based Regressor
Motivation.
Architecture.
Implementation details.
- 1.
- (Optional) linear projection if ,
- 2.
- reshape to ,
- 3.
- recurrent stack (RNN/LSTM/GRU, N layers, optional dropout),
- 4.
- last-timestep pooling, activation, and a final linear layer.
Capacity and complexity.
Training considerations.
Instantiation in our benchmark.
3.2.4. Transformer Regressor
Motivation.
Architecture.
- 1.
- Embedding (tokenization): a linear projection We treat the entire feature vector as a single token (sequence length ), so the encoder processes a tensor (batch dimension suppressed).
- 2.
- Encoder stack: N identical layers, each with (i) multi-head self-attention (MHSA) and (ii) a position-wise feed-forward network (FFN), both wrapped with residual connections and layer normalization:where with hidden width and activation (ReLU or GELU).
- 3.
- Pooling and head: mean pooling over the (single) token and a linear regressor:
Implementation details.
Complexity.
Instantiation in our benchmark.
Notes on sequence construction.
3.2.5. Advanced 1D CNN Regressor with Multi-Scale Attention
Motivation.
Architecture.
- 1.
- Embedding stem: , where is the chosen activation.
- 2.
- Feature stages (): each stage stacks a MultiScaleConvBlock and (optionally) a ResidualBlock, followed (except the last stage) by adaptive downsampling:where and (doubling across stages).
- 3.
- Global aggregation: global average and max pooling on the final feature map :
- 4.
- Head: a two-layer MLP with batch normalization, activation, and dropout, followed by a linear regressor to :
Attention modules.
Implementation notes.
Complexity.
Instantiation in our benchmark.
Training.
3.2.6. Variational Autoencoder (VAE) Regressor
Motivation.
Architecture.
- Encoder: MLP with hidden widths specified by hidden_dims, activation (e.g., ReLU/GELU), and optional dropout; linear heads produce and of size .
- Decoder: mirrored MLP mapping back to .
- Regressor: small MLP mapping to .
Objective.
Implementation details.
Training considerations.
Instantiation in our benchmark.
3.2.7. Mixture of Experts (MoE) Regressor
Motivation.
Architecture.
Implementation details.
- Experts: each expert is a feedforward MLP with hidden widths expert_hidden_dims and dropout between hidden layers; the last layer is linear to preserve regression fidelity.
- Gating network: a shallow MLP (input →hidden→hidden/2→ E) with dropout outputs unnormalized logits; softmax yields mixture weights.
- Pre/post processing: input LayerNorm and a residual linear projection improve conditioning; output LayerNorm stabilizes training across experts.
- Sparsity: if top_k < num_experts, only the selected experts are evaluated/combined; otherwise, all experts contribute.
Capacity and compute.
Training considerations.
Instantiation in our benchmark.
3.2.8. TabNet Regressor with Ghost Batch Normalization
Motivation.
Building blocks.
End-to-end step-wise computation.
Interpretability.
Implementation details.
Instantiation in our benchmark.
3.3. Training Methodology
3.3.1. Hyperparameter Configuration
3.3.2. Learning Rate Scheduling
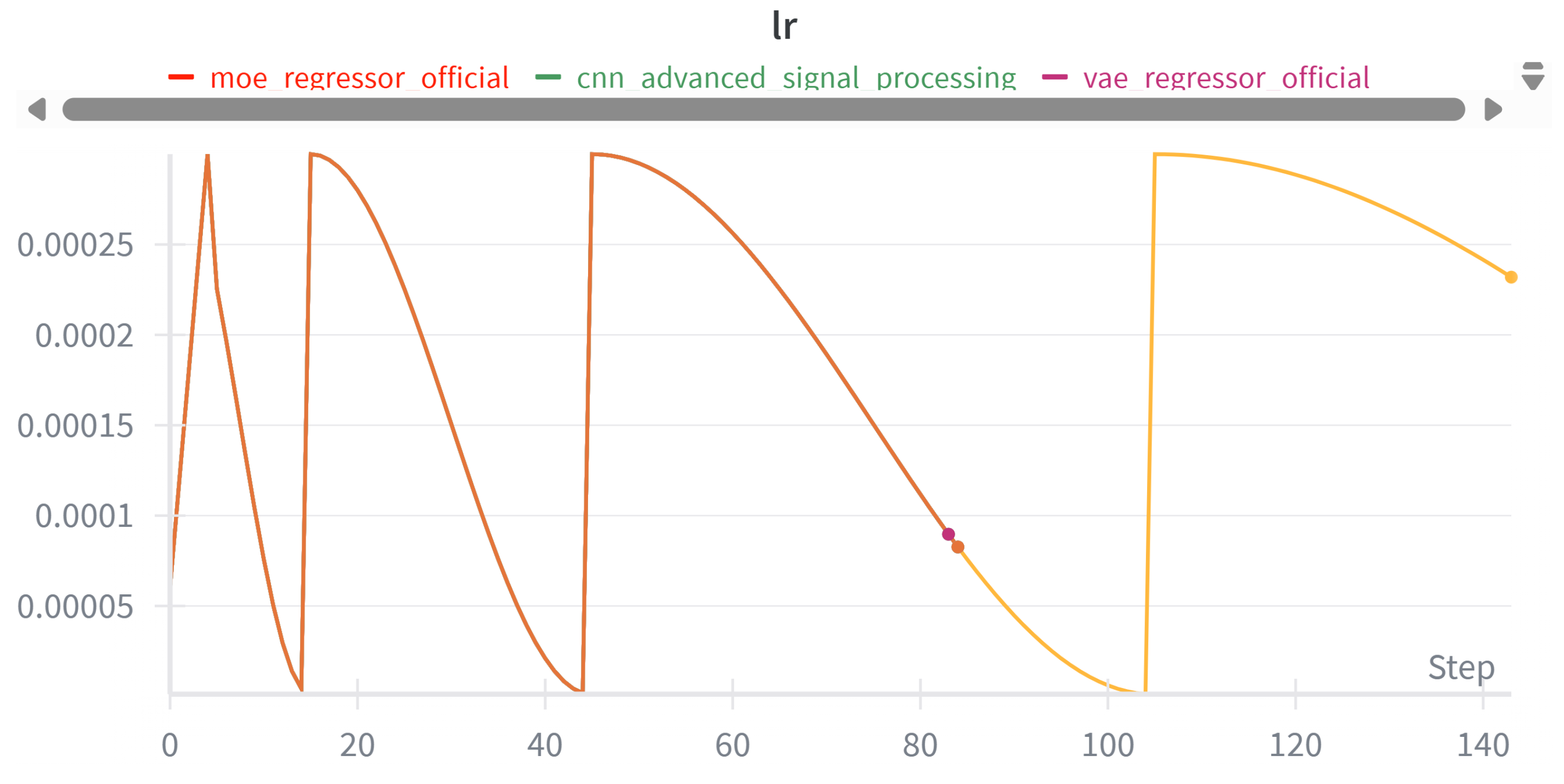
3.3.3. Data Preprocessing
3.4. Evaluation Metrics
3.5. Implementation Details
4. Results and Analysis
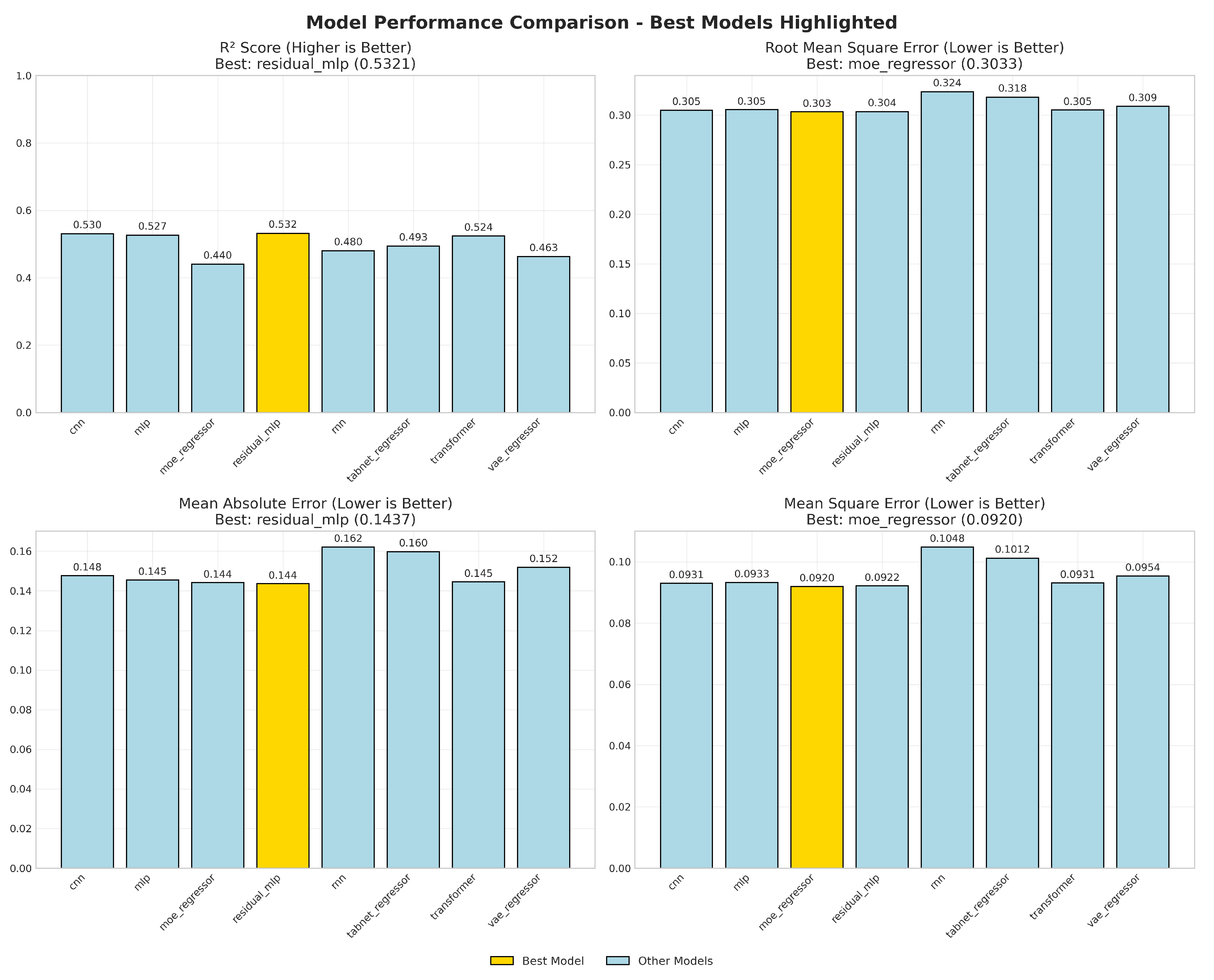
4.1. Model Performance
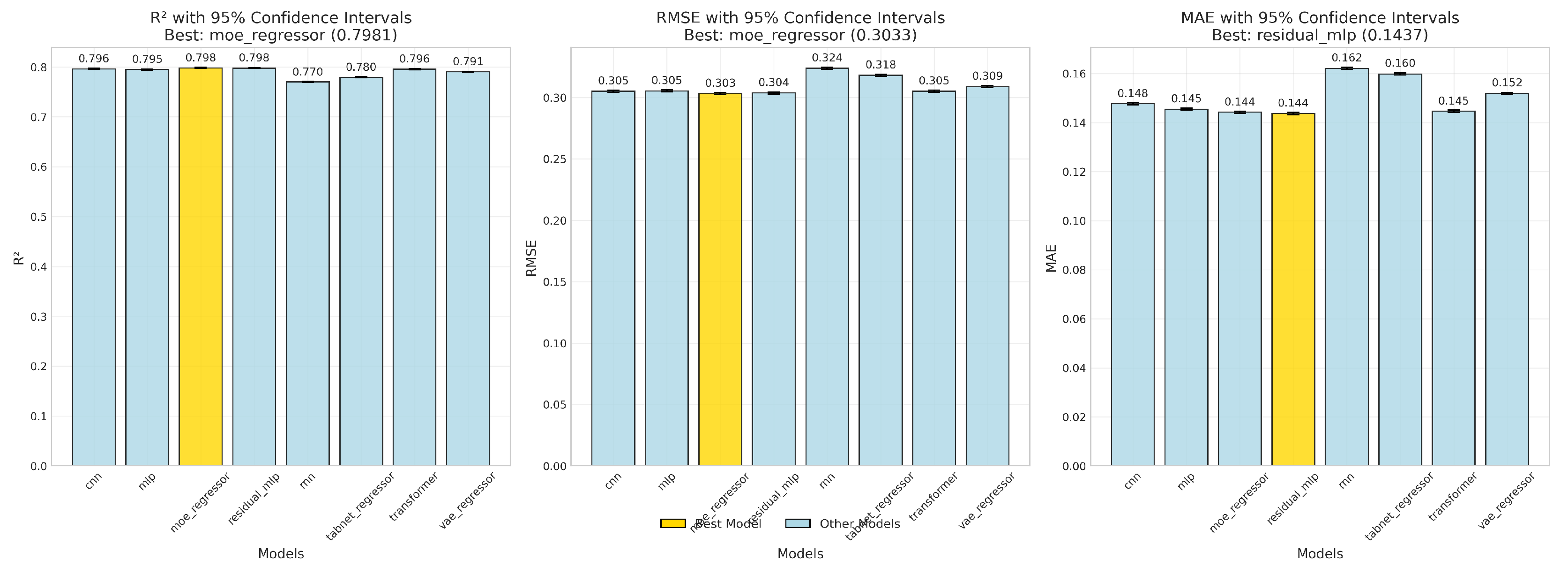
4.1.1. Uncertainty via Bootstrap Confidence Intervals
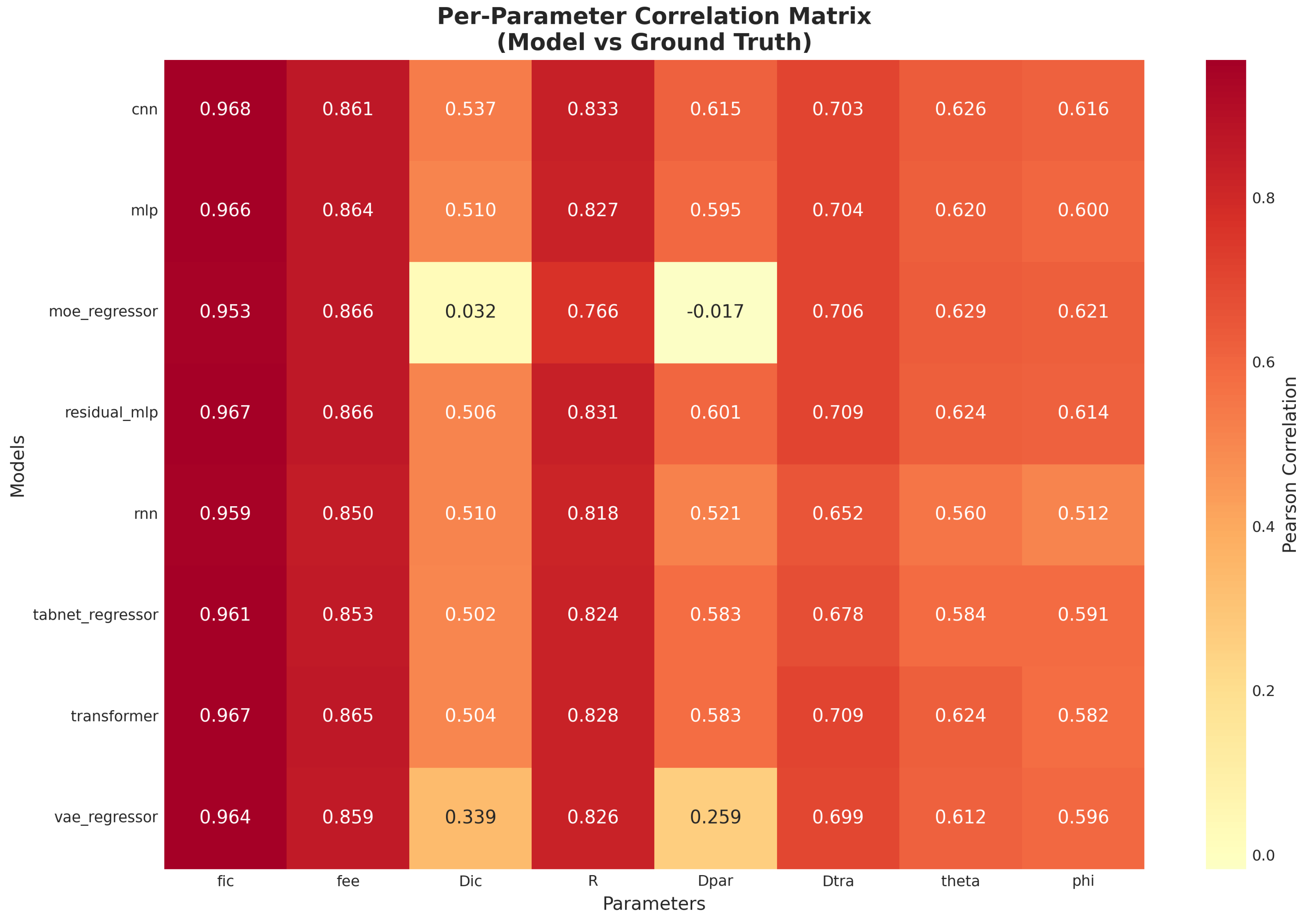
4.1.2. Correlation Structure Across Parameters
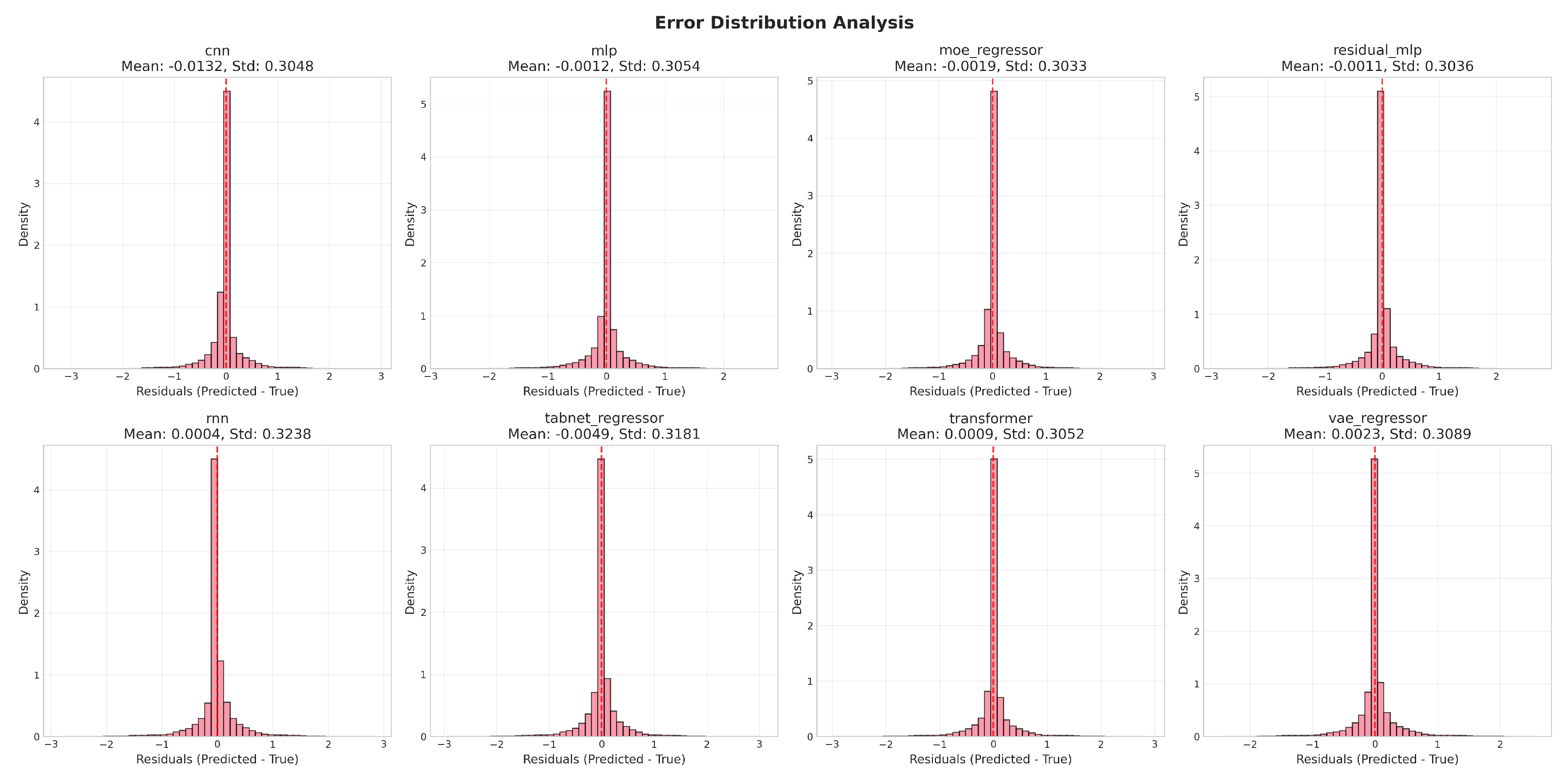
4.1.3. Error Behavior and Residual Diagnostics
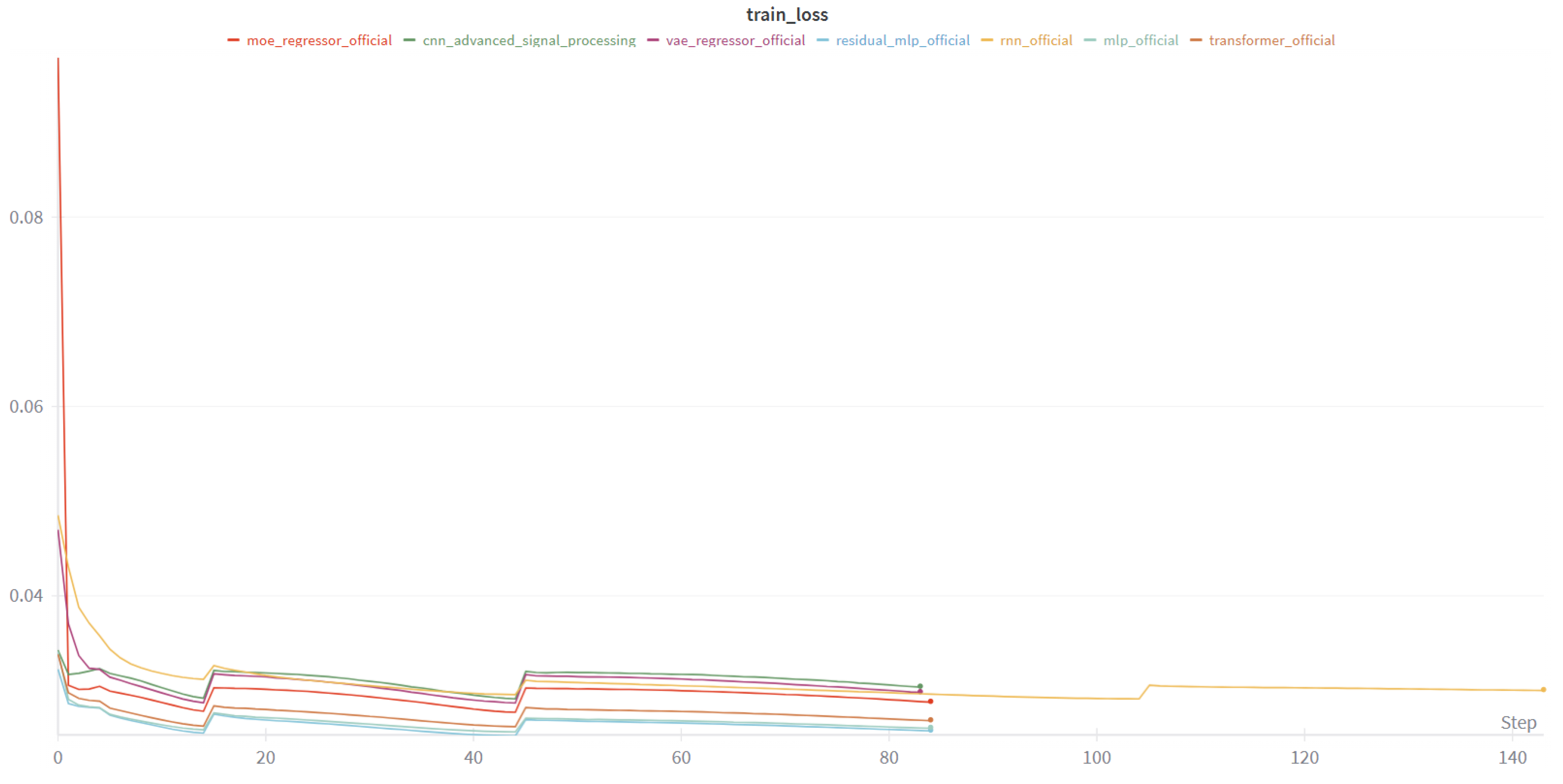
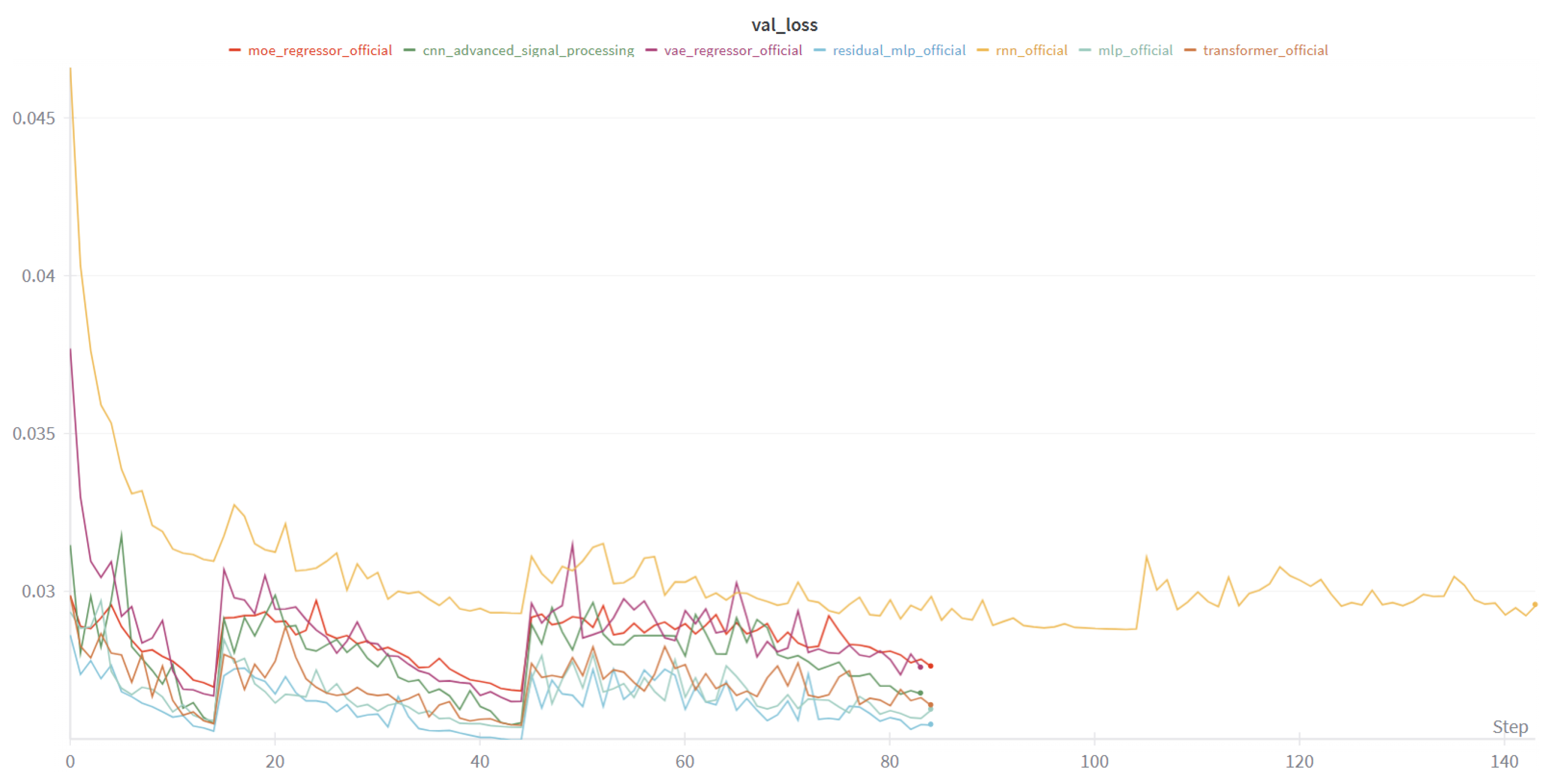
4.1.4. Learning Dynamics
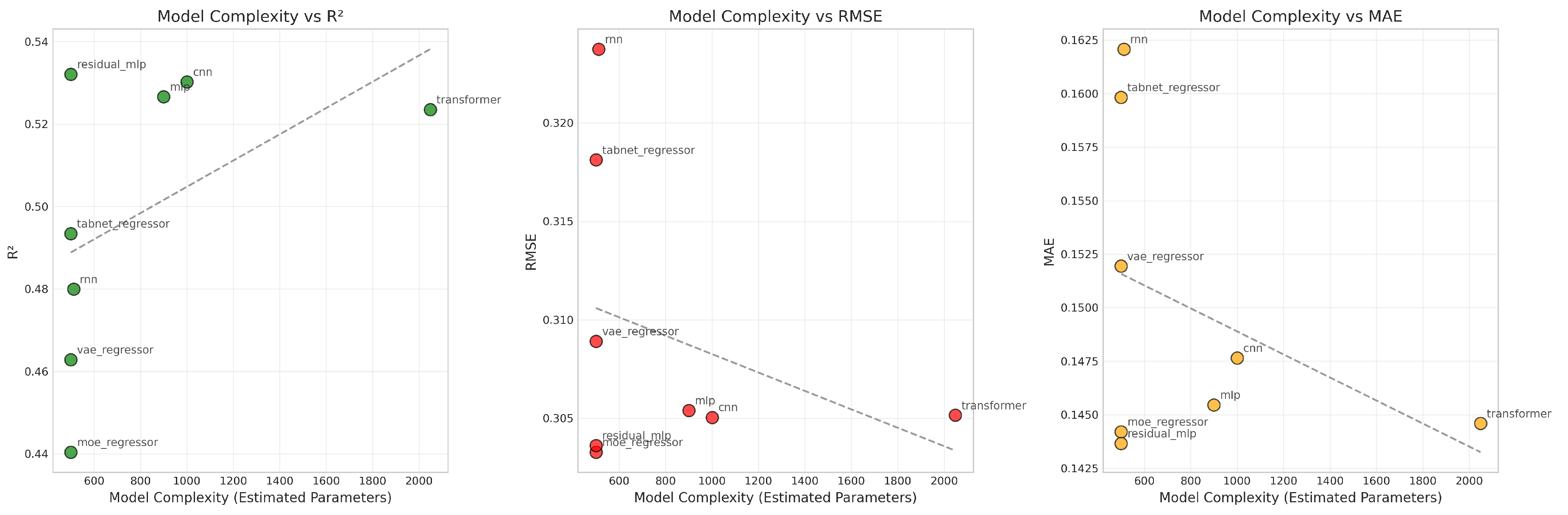
4.1.5. Complexity and Performance Trade-Off
R2.
RMSE and MAE.
Takeaway.
4.2. Rankings and Summary Table
4.2.1. Overall Ranking Methodology
Individual Metric Rankings
- Score Ranking: Models are ranked in descending order, where rank 1 corresponds to the highest score (best performance)
- RMSE Ranking: Models are ranked in ascending order, where rank 1 corresponds to the lowest RMSE (best performance)
- MAE Ranking: Models are ranked in ascending order, where rank 1 corresponds to the lowest MAE (best performance)
- MSE Ranking: Models are ranked in ascending order, where rank 1 corresponds to the lowest MSE (best performance)
- Mean Correlation Ranking: Models are ranked in descending order, where rank 1 corresponds to the highest mean correlation (best performance)
Overall Rank Calculation
Final Position Assignment
Advantages of This Approach
- 1.
- Balanced Assessment: Equal weight is given to each metric, preventing any single measure from dominating the evaluation
- 2.
- Robustness: Models that perform consistently well across all metrics are favored over those that excel in only one area
- 3.
- Interpretability: The ranking system is transparent and easy to understand
- 4.
- Flexibility: The approach can easily accommodate additional metrics if needed
4.3. Results in Real Patients Data
4.3.1. Multi-Patient Clinical Assessment
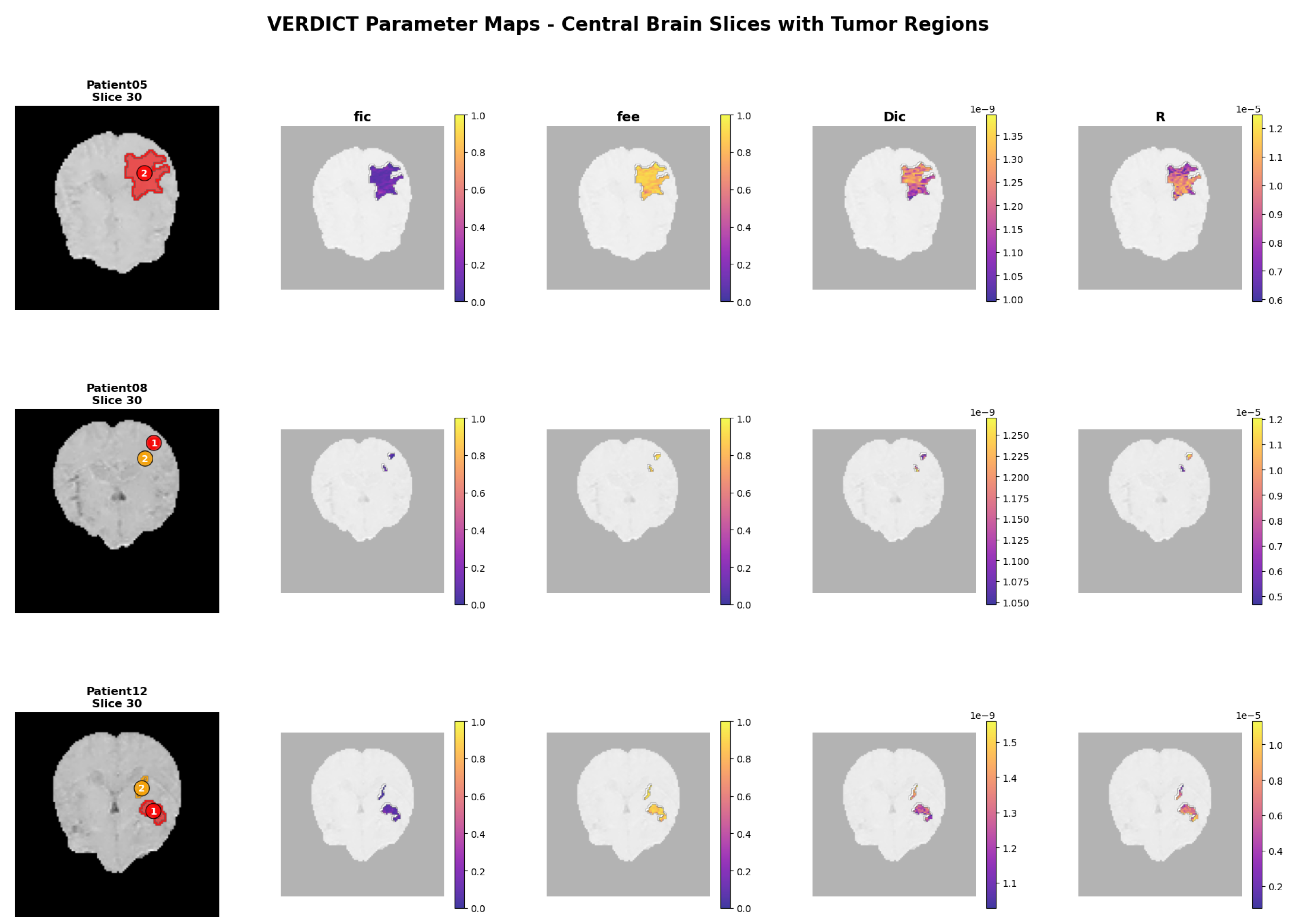
4.3.2. Patient Parameter Maps
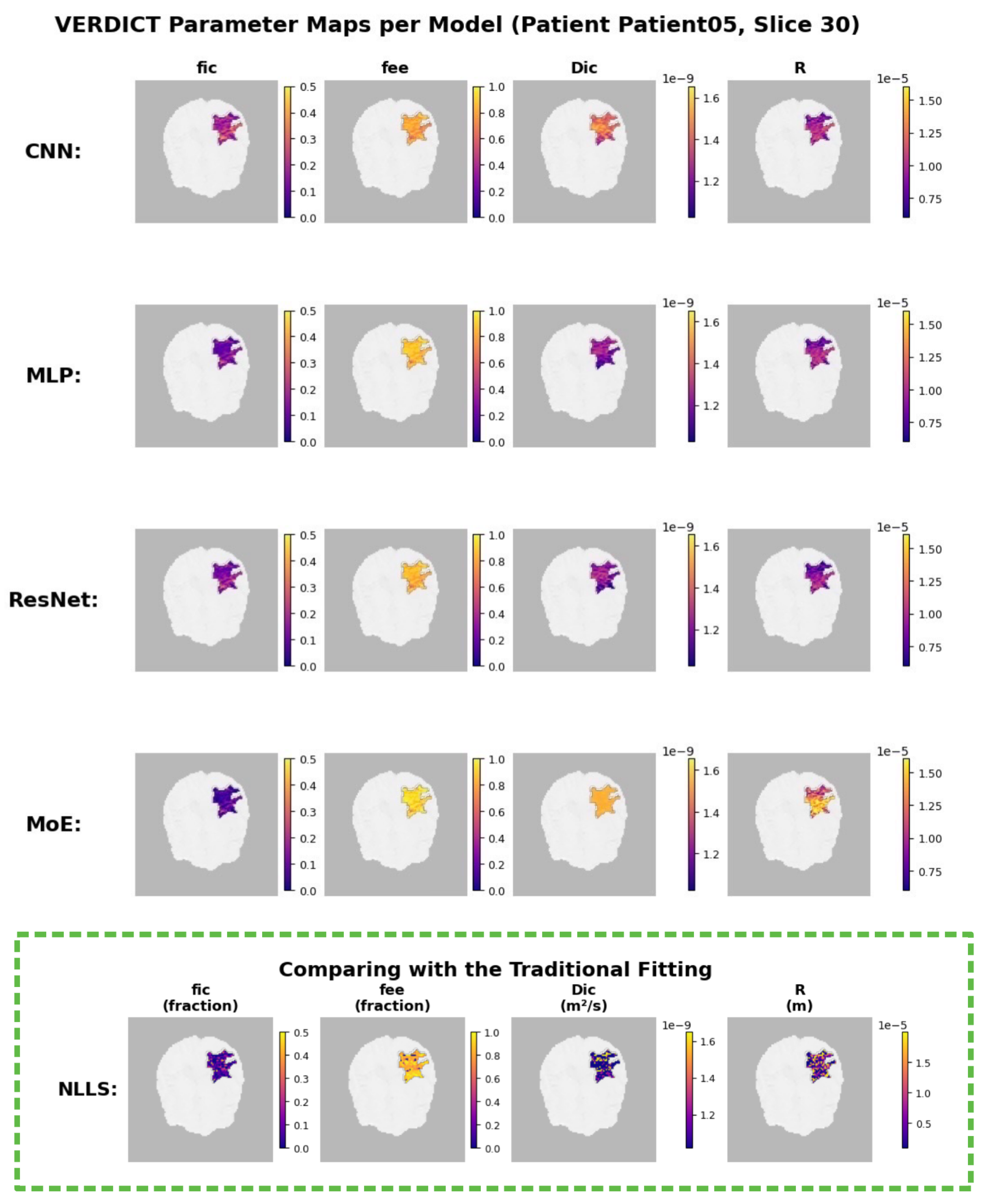
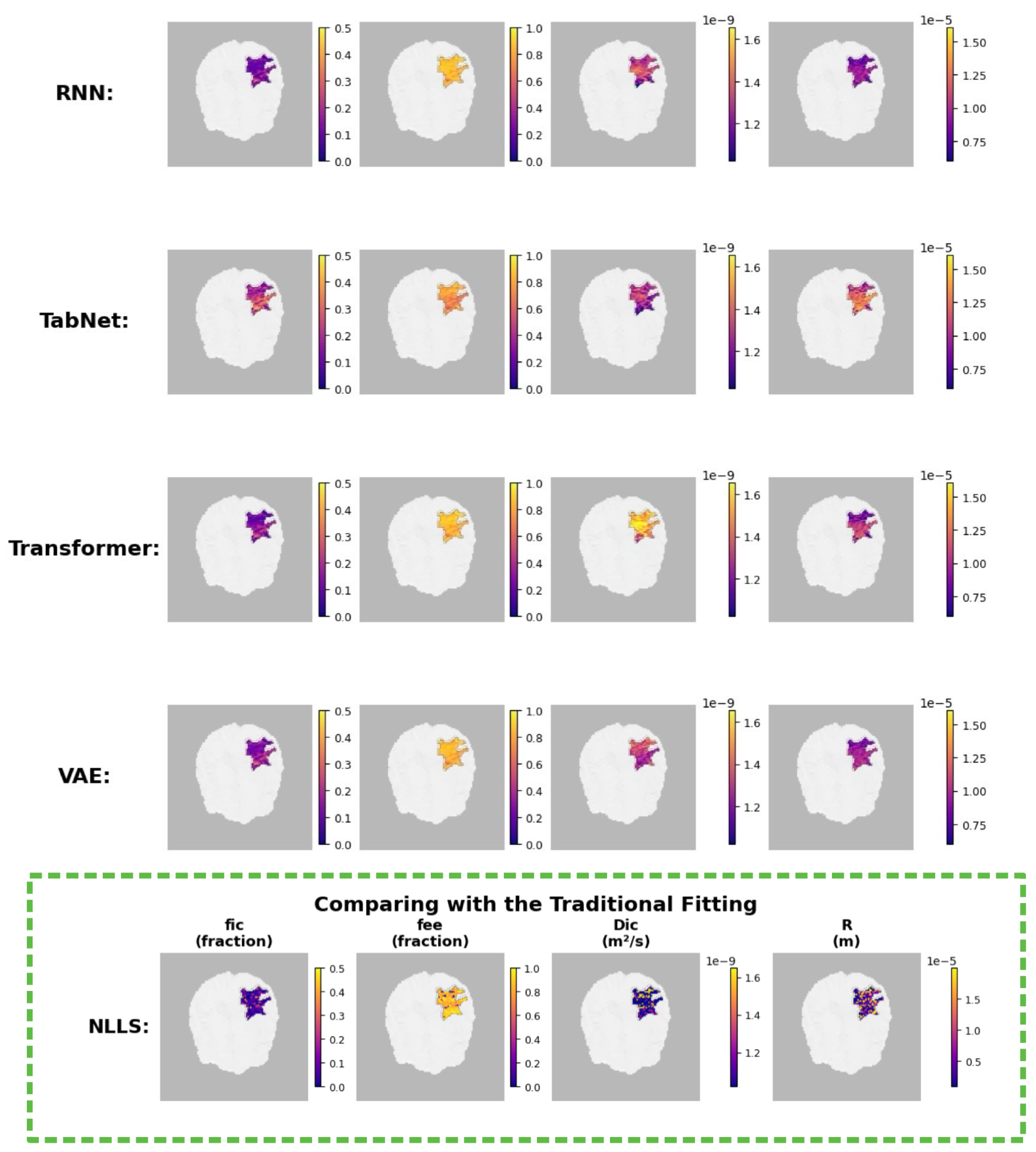
4.3.3. Model-Specific Parameter Reconstructions
Interpretation.
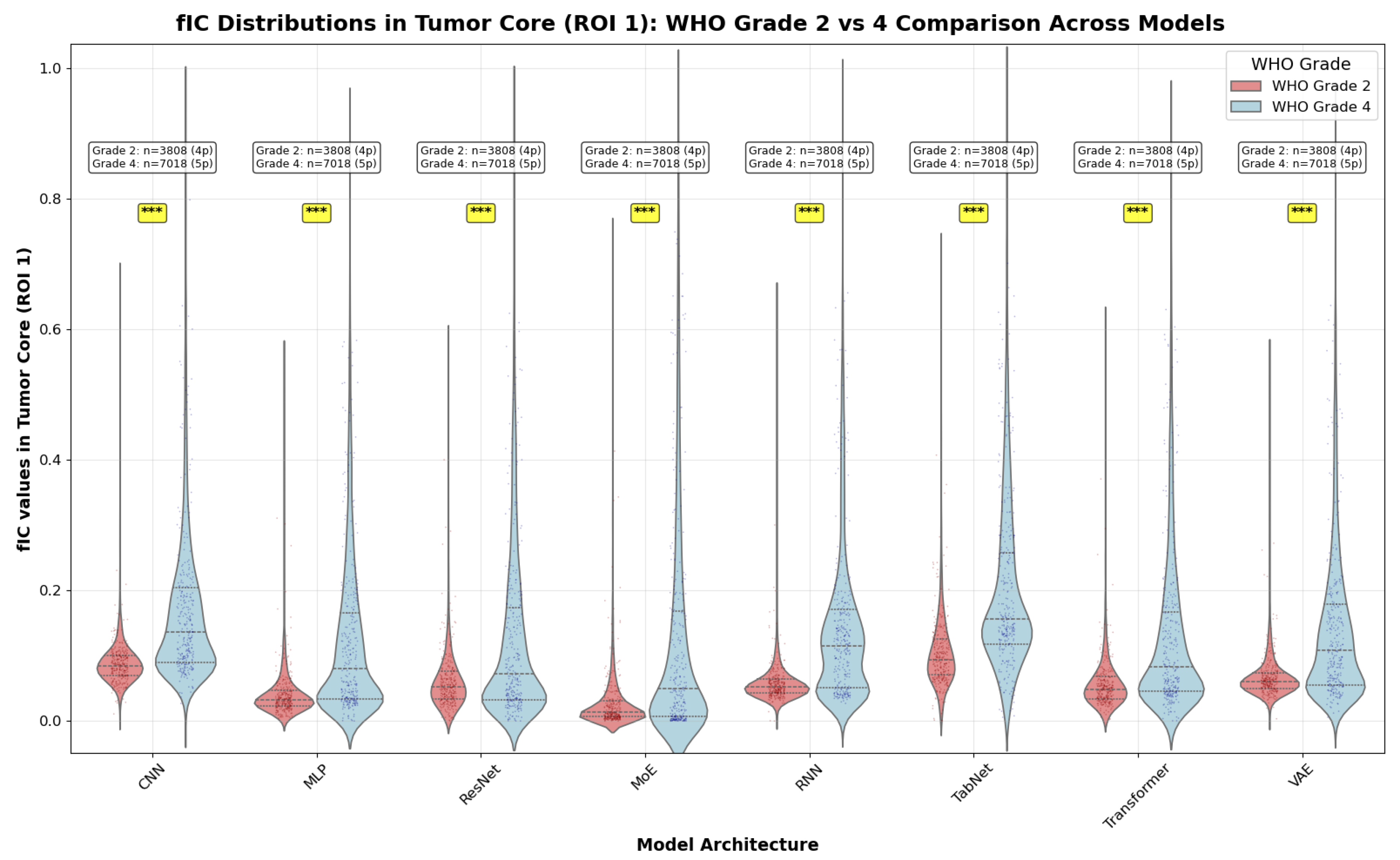
4.3.4. WHO Grade Comparison: Group Analysis
Methodology
Statistical Analysis
- Grade 2 vs Grade 4: Direct comparison between low-grade and high-grade gliomas
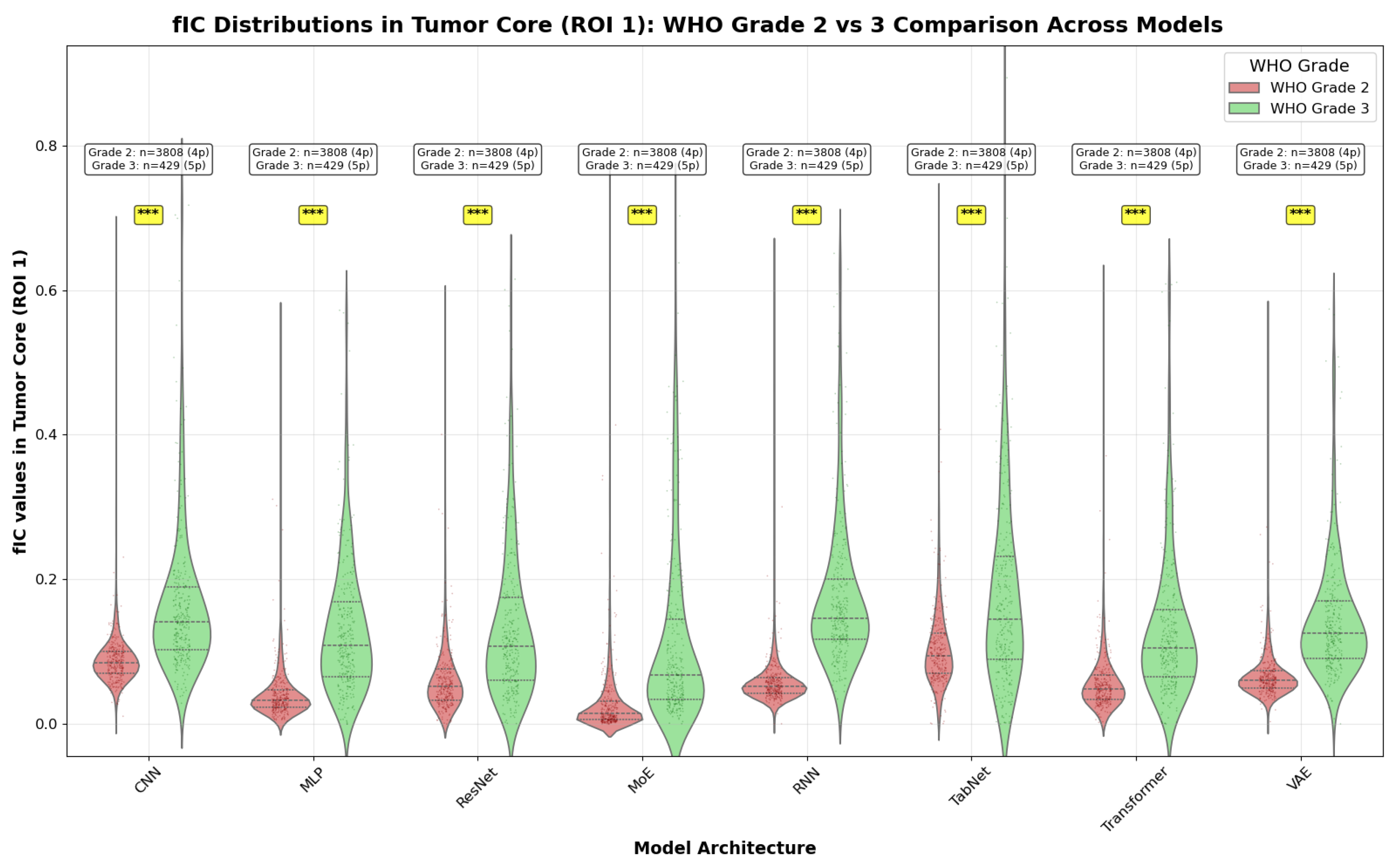
- Grade 2 vs Grade 3: Comparison within the broader low-grade category
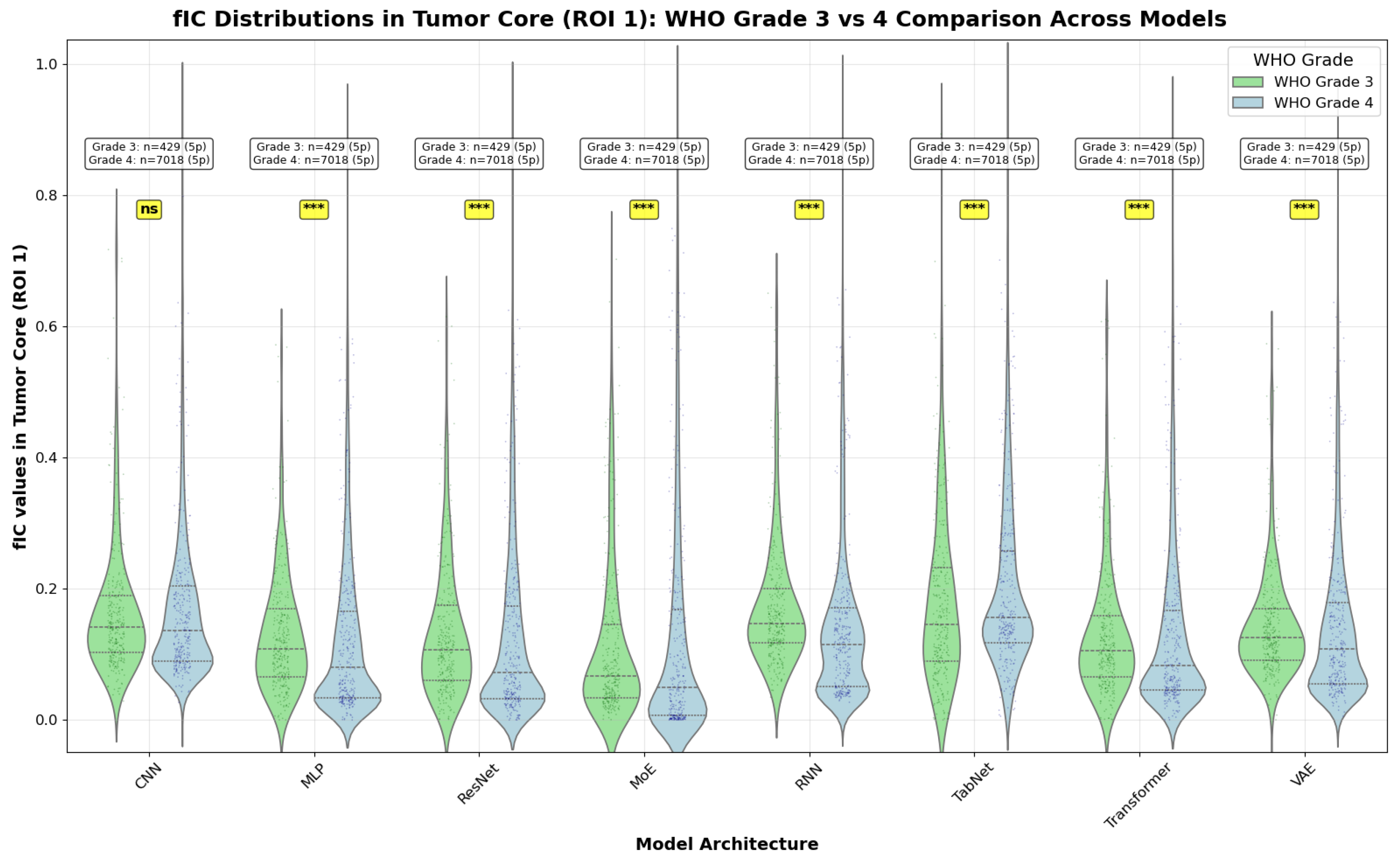
- Grade 3 vs Grade 4: Comparison within the malignant glioma spectrum
Interpretation
Clinical Relevance
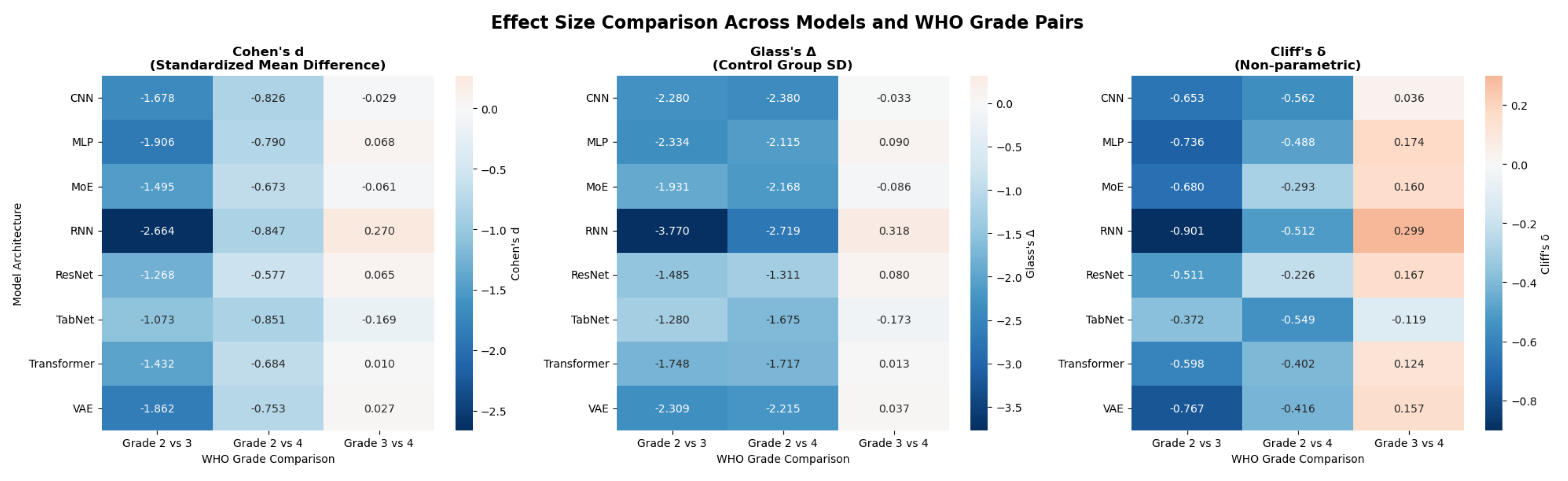
4.4. Effect Size Analysis
4.4.1. Effect Size Metrics
Cohen’s d
Glass’s Delta
Cliff’s Delta
4.4.2. Hierarchical Analysis Approach
- 1.
- Voxel level: Direct comparison of all C values. This reflects overall distributional differences but may inflate effect sizes due to within-patient correlation.
- 2.
- Patient level: Comparison using patient-averaged values, accounting for clustering and yielding more conservative, clinically interpretable estimates.
4.4.3. Clinical Significance Framework
- Statistical significance: (Mann-Whitney U).
- Practical significance: medium or large effect ( for Cohen’s d).
4.4.4. Results Summary
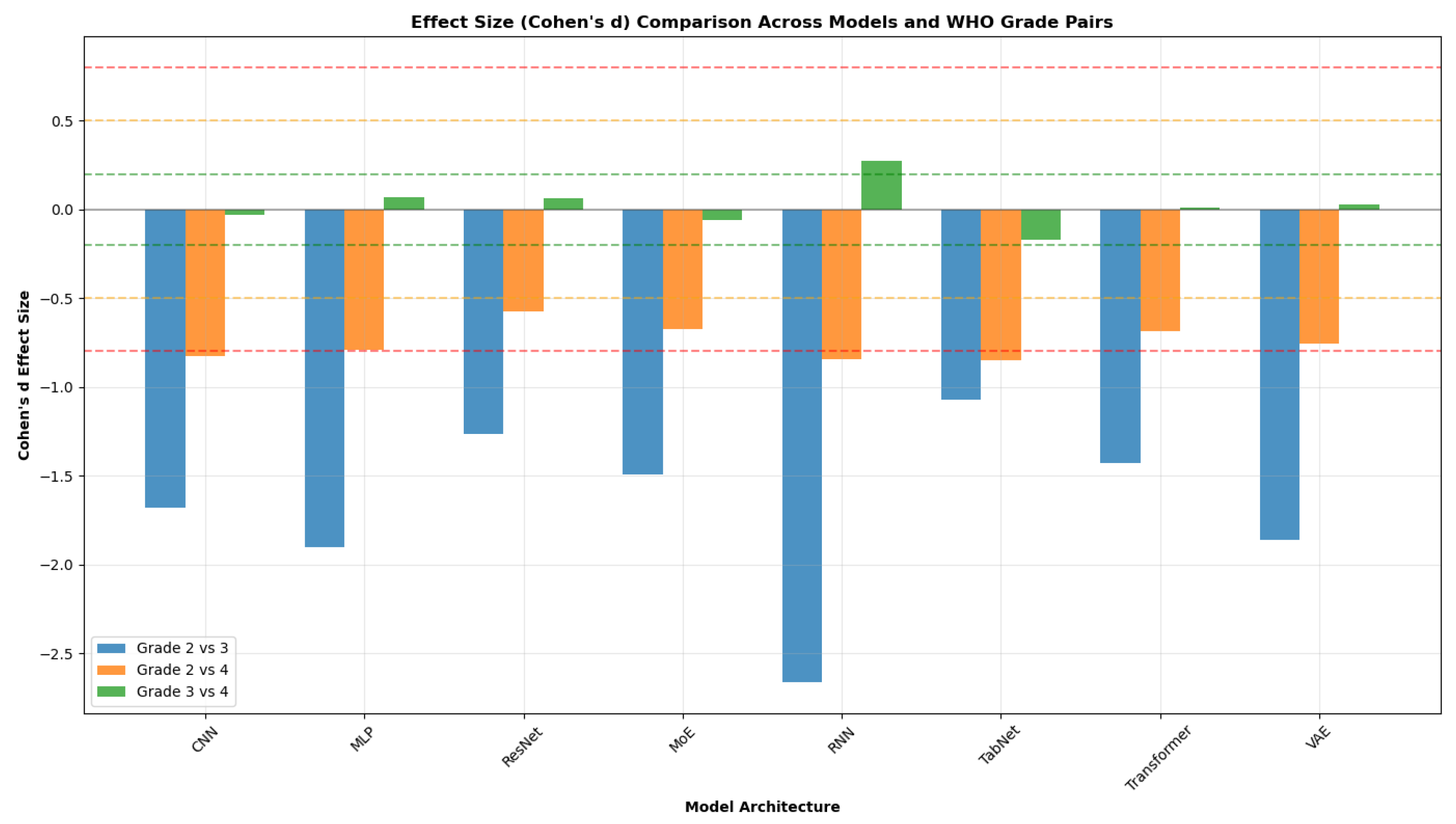
4.4.4.1. Cross-architecture patterns (see Figure 16 and Figure 17).
- Grade 2 vs 4 yields the largest across-the-board separations. Cohen’s d values span approximately (medium to large) across models, with Transformer, RNN, and MLP among the strongest.
- Grade 3 vs 4 generally shows small effects. Cohen’s d is near zero for most models, with RNN the only architecture showing a clear (small) separation (); Cliff’s mirrors this pattern with small magnitudes.
- Grade 2 vs 3 exhibits uniformly large effects across all models. Cohen’s d magnitudes range from about to , with the RNN showing the largest separation across all three metrics.
Model-specific observations.
- RNN consistently provides the largest Grade 2 vs 3 separation (all metrics) and the strongest, albeit still small, Grade 3 vs 4 effect among models.
- Transformer and CNN show reliable, consistent discrimination for the easier pairings (Grade 2 vs 3 and Grade 2 vs 4), indicating robustness across metrics.
- MLP and ResNet deliver intermediate-to-strong effects for Grade 2 vs 3 and Grade 2 vs 4, with limited separation for Grade 3 vs 4.
- Specialized architectures (TabNet, VAE, MoE) perform variably across pairs, but follow the same global ordering: Grade 2 vs 3 ≫ Grade 2 vs 4 > Grade 3 vs 4.
Clinical significance assessment.
4.4.5. Implications for Model Selection
- 1.
- Discrimination capability. Models with consistently large effects (e.g., RNN, Transformer, CNN) are preferable when clear grade differentiation is critical.
- 2.
- Sensitivity vs. robustness. Large effects should be balanced with generalizability; models that are consistently strong across metrics and grade pairs (e.g., Transformer, CNN) may offer greater reliability.
- 3.
- Grade-specific performance. Some models (e.g., RNN) excel at Grade 2 vs 3 and provide the best (though still small) separation for Grade 3 vs 4; selection may be tailored to the clinically relevant decision boundary.
5. Discussion
5.1. Benchmark Performance Across Architectures
5.1.1. Advanced Architectures Underperformed
5.1.2. Non-Monotonic Relationship Between Complexity and Performance
5.1.3. Performance Summary
5.2. Parameter-Wise Prediction Difficulty
5.2.1. Easy Parameters
5.2.2. Hard Parameters
5.3. Implications for Clinical Application
5.3.1. Computational Feasibility and Speed
5.3.2. Robustness and Reliability
5.4. Interpretability and Biophysical Plausibility
5.5. Limitations
- It was performed in a specific setting, using a particular implementation of the VERDICT model with a specific acquisition protocol. While justified, this may limit generalizability to other implementations or imaging conditions.
- The evaluation was conducted on a relatively small real dataset, without access to a reliable ground truth, which restricts the robustness of the conclusions.
5.6. Future Directions
5.6.1. Real-World Validation and Transfer Learning
5.6.2. Advanced Multi-Task Learning and Clinical Integration
5.6.3. Extension to Other Microstructure Models and Data
5.6.4. Clinical Deployment and User Studies
6. Conclusion
Key Findings
- Model complexity vs. performance: Simple feed-forward networks (e.g., multilayer perceptrons) performed on par with – and occasionally outperformed – more complex architectures (such as recurrent or attention-based models). Increased architectural complexity did not consistently translate to better predictive accuracy for VERDICT parameter estimation.
- Parameter-wise difficulty: Prediction accuracy varied across the individual microstructural parameters. The intracellular and extracellular volume fractions were estimated with the highest fidelity, whereas certain parameters (notably some diffusivity and geometrical metrics) proved more challenging. For example, the model correlations for intracellular diffusivity and extravascular diffusion coefficients were significantly lower than for the volume fractions, indicating that these targets may require specialized training strategies or additional features.
- Speed and robustness: All deep learning models delivered near-instant inference and stable performance, in contrast to the slow, iterative, and noise-sensitive conventional fitting. The learned predictors effectively eliminate the minutes-per-slice computational burden of standard methods, enabling rapid and robust VERDICT mapping that is far more compatible with clinical workflow.
Future Directions
- 1.
- Real-data validation: Rigorously evaluate the trained models on in-vivo clinical VERDICT MRI datasets to confirm that their performance generalizes beyond the simulated training data.
- 2.
- Improving difficult parameters: Develop model and training improvements to better estimate the parameters that remain challenging (for instance, incorporating tailored loss weighting or architecture changes to capture intracellular diffusivity and anisotropic diffusion components more effectively).
- 3.
- Clinical translation: Conduct prospective clinical studies to integrate the deep-learning pipeline into real imaging workflows and assess its impact on diagnostic accuracy and patient outcomes in practice.
Acknowledgments
Appendix A. Source Code
References
- Panagiotaki, E.; Walker-Samuel, S.; Siow, B.; Johnson, S.P.; Rajkumar, V.; Pedley, R.B.; Lythgoe, M.F.; Alexander, D.C. Noninvasive Quantification of Solid Tumor Microstructure Using VERDICT MRI. Cancer Research 2014, 74, 1902–1912. [CrossRef]
- Zaccagna, F.; Riemer, F.; Priest, A.N.; McLean, M.A.; Allinson, K.; Grist, J.T.; Dragos, C.; Matys, T.; Gillard, J.H.; Watts, C.; et al. Non-invasive assessment of glioma microstructure using VERDICT MRI: correlation with histology. European Radiology 2019, 29, 5559–5566. [CrossRef]
- Grussu, F.; Battiston, M.; Palombo, M.; Schneider, T.; Gandini Wheeler-Kingshott, Claudia A. M.; Alexander, D.C. Deep Learning Model Fitting for Diffusion-Relaxometry: A Comparative Study. In Computational Diffusion MRI; Mathematics and Visualization, Springer International Publishing: Cham, Switzerland, 2021; pp. 159–172. [CrossRef]
- O’Connor, J.P.B.; Rose, C.J.; Waterton, J.C.; Carano, R.A.D.; Parker, G.J.M.; Jackson, A. Imaging intratumor heterogeneity: role in therapy response, resistance, and clinical outcome. Clinical Cancer Research 2015, 21, 249–257. [CrossRef]
- Messina, C.; Bignone, E.; Bruno, E.; Bruno, F.; Nazarian, E.; Reginelli, A.; Calandri, S.; Tagliafico, A.; Ulivieri, A.; Guglielmi, G.; et al. Diffusion Weighted Imaging in Oncology: An Update. Eur. Radiol. Exp. 2020, 4, 55. [CrossRef]
- Drake-Pérez, M.; Boto, J.; Fitsiori, F.; Lovblad, R.; Vargas, R.T. Clinical applications of diffusion weighted imaging in neuroradiology. Insights Imaging 2018, 9, 535–547. [CrossRef]
- Jelescu, I.O.; Budde, M.D. Design and validation of diffusion MRI microstructure models. NMR in Biomedicine 2017, 30, e3729. [CrossRef]
- Litjens, G.; Kooi, T.; Ehteshami Bejnordi, B.; Setio, A.A.A.; Ciompi, F.; et al. A survey on deep learning in medical image analysis: the era of big data and deep learning. Medical Image Analysis 2017, 42, 60–88. [CrossRef]
- Golkov, V.; Dosovitskiy, A.; Sperl, J.I.; Menzel, M.I.; Czisch, M.; Sämann, P.; Brox, T.; Cremers, D. q-Space Deep Learning: Twelve-Fold Shorter and Model-Free Diffusion MRI Scans. IEEE Transactions on Medical Imaging 2016, 35, 1344–1351. [CrossRef]
- Figini, M.; Palombo, M.; Bailo, M.; Alexander, D.C.; Cercignani, M.; Castellano, A.; Panagiotaki, E. Towards a clinical acquisition protocol for VERDICT MRI in brain tumours. In Proceedings of the Proceedings of the International Society for Magnetic Resonance in Medicine (ISMRM), 2025, Vol. 33, p. 4068.
- Figini, M.; Palombo, M.; Bailo, M.; Callea, M.; Mortini, P.; Falini, A.; Alexander, D.C.; Cercignani, M.; Castellano, A.; Panagiotaki, E. Accelerated Glioma characterization with VERDICT MRI: a comparison between deep learning and non-linear least squares fitting. In Proceedings of the Proceedings of the International Society for Magnetic Resonance in Medicine (ISMRM), 2024, Vol. 32, p. 3502.
- Johnston, E.W.; Bonet-Carne, E.; Ferizi, U.; Yvernault, B.; Pye, H.; Patel, D.; Clemente, J.; Piga, W.; Heavey, S.; Sidhu, H.S.; et al. VERDICT MRI for Prostate Cancer: Intracellular Volume Fraction versus Apparent Diffusion Coefficient. Radiology 2019, 291, 391–397. [CrossRef]
- Panagiotaki, E.; Chan, R.W.; Dikaios, N.; Ahmed, H.U.; O’Callaghan, J.; Freeman, A.; Atkinson, D.; Punwani, S.; Hawkes, D.J.; Alexander, D.C. Microstructural characterization of normal and malignant human prostate tissue with vascular, extracellular, and restricted diffusion for cytometry in tumours magnetic resonance imaging. Investigative Radiology 2015, 50, 218–227. [CrossRef]
- Figini, M.; Castellano, A.; Bailo, M.; Callea, M.; Cadioli, M.; Bouyagoub, S.; Palombo, M.; Pieri, V.; Mortini, P.; Falini, A.; et al. Comprehensive Brain Tumour Characterisation with VERDICT-MRI: Evaluation of Cellular and Vascular Measures Validated by Histology. Cancers 2023, 15, 2490. [CrossRef]
- Bailey, C.; Collins, D.J.; Tunariu, N.; Orton, M.R.; Morgan, V.A.; Feiweier, T.; Hawkes, D.J.; Leach, M.O.; Alexander, D.C.; Panagiotaki, E. Microstructure Characterization of Bone Metastases from Prostate Cancer with Diffusion MRI: Preliminary Findings. Frontiers in Oncology 2018, 8, 26. [CrossRef]
- Novikov, D.S.; Kiselev, V.G.; Jespersen, S.N. Quantifying brain microstructure with diffusion MRI: Theory and parameter estimation. NMR in Biomedicine 2019, 32, e3998. [CrossRef]
- Goodfellow, I.; Bengio, Y.; Courville, A. Deep Learning; MIT Press: Cambridge, MA, 2016.
- Ye, C.; Cui, Y.; Li, X. q-Space Learning with Synthesized Training Data. In Computational Diffusion MRI; Mathematics and Visualization, Springer: Cham, 2019; pp. 123–132. [CrossRef]
- Behrens, T.E.J.; Woolrich, M.W.; Jenkinson, M.; Johansen-Berg, H.; Nunes, R.G.; Clare, S.; Matthews, P.M.; Brady, M.; Smith, S.M. Characterization and propagation of uncertainty in diffusion-weighted MR imaging. Magnetic Resonance in Medicine 2003, 50, 1077–1088. [CrossRef]
- Stejskal, E.O.; Tanner, J.E. Spin Diffusion Measurements: Spin Echoes in the Presence of a Time-Dependent Field Gradient. Journal of Chemical Physics 1965, 42, 288–292. [CrossRef]
- Basser, P.J.; Mattiello, J.; LeBihan, D. MR Diffusion Tensor Spectroscopy and Imaging. Biophysical Journal 1994, 66, 259–267. [CrossRef]
- Afzali, M.; Aja-Fernández, S.; Jones, D.K. Direction-averaged Diffusion-weighted MRI Signal Using Different Axisymmetric B-tensor Encoding Schemes. Magnetic Resonance in Medicine 2020, 84, 1579–1591. [CrossRef]
- Paszke, A.; Gross, S.; Massa, F.; others. PyTorch: An Imperative Style, High-Performance Deep Learning Library. In Proceedings of the Advances in Neural Information Processing Systems (NeurIPS), 2019, Vol. 32, pp. 8024–8035.
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2016, pp. 770–778. [CrossRef]
- Hochreiter, S.; Schmidhuber, J. Long Short-Term Memory. Neural Computation 1997, 9, 1735–1780. [CrossRef]
- Cho, K.; van Merriënboer, B.; Gülçehre, Ç.; Bahdanau, D.; Bougares, F.; Schwenk, H.; Bengio, Y. Learning Phrase Representations using RNN Encoder–Decoder for Statistical Machine Translation. In Proceedings of the Proceedings of the 2014 Conference on Empirical Methods in Natural Language Processing (EMNLP), Doha, Qatar, 2014; pp. 1724–1734. [CrossRef]
- Vaswani, A.; Shazeer, N.; others. Attention Is All You Need. In Proceedings of the Proceedings of the 31st International Conference on Neural Information Processing Systems (NeurIPS), 2017, pp. 5998–6008.
- LeCun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document recognition. Proceedings of the IEEE 1998, 86, 2278–2324. [CrossRef]
- Szegedy, C.; Liu, W.; Jia, Y.; Sermanet, P.; Reed, S.; Anguelov, D.; Erhan, D.; Vanhoucke, V.; Rabinovich, A. Going Deeper with Convolutions. In Proceedings of the Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2015, pp. 1–9. [CrossRef]
- Woo, S.; Park, J.; Lee, J.Y.; Kweon, I.S. CBAM: Convolutional Block Attention Module. In Proceedings of the Proceedings of the European Conference on Computer Vision (ECCV), 2018, pp. 3–19. [CrossRef]
- Hu, J.; Shen, L.; Sun, G. Squeeze-and-Excitation Networks. In Proceedings of the Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2018, pp. 7132–7141. [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Delving Deep into Rectifiers: Surpassing Human-Level Performance on ImageNet Classification. In Proceedings of the Proceedings of the IEEE International Conference on Computer Vision (ICCV), 2015, pp. 1026–1034. [CrossRef]
- Ioffe, S.; Szegedy, C. Batch Normalization: Accelerating Deep Network Training by Reducing Internal Covariate Shift. In Proceedings of the Proceedings of the 32nd International Conference on Machine Learning (ICML), 2015, pp. 448–456.
- Lin, M.; Chen, Q.; Yan, S. Network in Network. In Proceedings of the International Conference on Learning Representations (ICLR), 2014, [1312.4400].
- Kingma, D.P.; Welling, M. Auto-Encoding Variational Bayes. In Proceedings of the International Conference on Learning Representations (ICLR), 2014, [1312.6114].
- Bishop, C.M. Pattern Recognition and Machine Learning; Information Science and Statistics, Springer: New York, 2006.
- Jacobs, R.A.; Jordan, M.I.; Nowlan, S.J.; Hinton, G.E. Adaptive Mixtures of Local Experts. Neural Computation 1991, 3, 79–87. [CrossRef]
- Shazeer, N.; Mirhoseini, A.; Maziarz, K.; Davis, A.; Le, Q.V.; Hinton, G.E.; Dean, J.; et al. Outrageously Large Neural Networks: The Sparsely-Gated Mixture-of-Experts Layer. arXiv preprint arXiv:1701.06538 2017, [arXiv:cs.LG/1701.06538].
- Arik, S.Ö.; Pfister, T. TabNet: Attentive Interpretable Tabular Learning. In Proceedings of the Proceedings of the AAAI Conference on Artificial Intelligence, 2021, Vol. 35, pp. 6679–6687.
- Hoffer, E.; Hubara, I.; Soudry, D. Train longer, generalize better: Closing the generalization gap in large batch training of neural networks. arXiv preprint arXiv:1705.08741 2017.
- Prechelt, L. Early Stopping—But When? In Neural Networks: Tricks of the Trade; Orr, G.B.; Müller, K.R., Eds.; Springer: Berlin, Heidelberg, 1998; Vol. 1524, Lecture Notes in Computer Science, pp. 55–69. [CrossRef]
- Loshchilov, I.; Hutter, F. SGDR: Stochastic Gradient Descent with Warm Restarts. In Proceedings of the International Conference on Learning Representations (ICLR), 2017, [1608.03983].
- Efron, B.; Tibshirani, R.J. An Introduction to the Bootstrap; CRC Press: Boca Raton, FL, 1994.
- Biewald, L. Experiment Tracking with Weights and Biases. https://www.wandb.com, 2020. Software available from wandb.com.
- Kenny, M.; Schoen, I. Violin SuperPlots: visualizing replicate heterogeneity in large data sets. Molecular Biology of the Cell 2021, 32, 1333–1334. [CrossRef]
- Phuttharak, W.; Thammaroj, J.; Wara-Asawapati, S.; Panpeng, K. Grading Gliomas Capability: Comparison between Visual Assessment and Apparent Diffusion Coefficient (ADC) Value Measurement on Diffusion-Weighted Imaging (DWI). Asian Pacific Journal of Cancer Prevention 2020, 21, 385–390. [CrossRef]
- Kang, X.; et al. Grading of Glioma: combined diagnostic value of amide proton transfer (APT), diffusion-weighted imaging (DWI) and arterial spin labeling (ASL). BMC Medical Imaging 2020. [CrossRef]
- Cohen, J. Statistical Power Analysis for the Behavioral Sciences, 2nd ed.; Lawrence Erlbaum Associates: Hillsdale, NJ, 1988.
- Fritz, C.O.; Morris, P.E.; Richler, J.J. Effect size estimates: Current use, calculations, and interpretation. Journal of Experimental Psychology: General 2012, 141, 2–18. [CrossRef]
- Cliff, N. Dominance statistics: Ordinal analyses to answer ordinal questions; Vol. 114, Psychological Bulletin, 1993; pp. 494–509. [CrossRef]
- Hedges, L.V. Effect sizes in cluster-randomized designs. In Statistical Methods for Meta-Analysis; Cooper, H.; Hedges, L.V.; Valentine, J.C., Eds.; American Psychological Association, 2007; pp. 279–294. [CrossRef]
- Kirk, R.E. Practical Significance: A Concept Whose Time Has Come; Vol. 56, Educational and Psychological Measurement, 1996; pp. 746–759. [CrossRef]
- Sen, S.; Singh, S.; Pye, H.; Moore, C.M.; Whitaker, H.C.; Punwani, S.; Atkinson, D.; Panagiotaki, E.; Slator, P.J. ssVERDICT: Self-supervised VERDICT-MRI for enhanced prostate tumor characterization. Magnetic Resonance in Medicine 2024, 92, 2181–2192. [CrossRef]
- Li, H.; Liu, H.; von Busch, H.; Grimm, R.; Huisman, H.; Tong, A.; Winkel, D.; Penzkofer, T.; Shabunin, I.; Choi, M.H.; et al. Deep Learning-based Unsupervised Domain Adaptation via a Unified Model for Prostate Lesion Detection Using Multisite Bi-parametric MRI Datasets. Radiology: Artificial Intelligence 2024. Published online Aug. 8, 2024 (preprint DOI available).
- Wu, X.; Zhang, S.; Zhang, Z.; He, Z.; Xu, Z.; Wang, W.; Jin, Z.; You, J.; Guo, Y.; Zhang, L.; et al. Biologically interpretable multi-task deep learning pipeline predicts molecular alterations, grade, and prognosis in glioma patients. npj Precision Oncology 2024, 8, 181. [CrossRef]
- Tan, e.a. Artificial Intelligence for Diffusion MRI-Based Tissue Microstructure Estimation: Architectures, Pitfalls, and Future Perspectives. Frontiers in Neurology 2023, 14, 1168833. [CrossRef]
- Arslan, A.; Alis, D.; Erdemli, S.; Şeker, M.E.; Zeybel, G.; Sirolu, S.; Kurtçan, S.; Karaarslan, E. Does Deep Learning Software Improve the Consistency and Performance of Radiologists with Various Levels of Experience in Assessing Bi-Parametric Prostate MRI? Insights into Imaging 2023, 14. [CrossRef]
- Faiyaz, A.; Doyley, M.M.; Schifitto, G.; Uddin, M.N. Artificial Intelligence for Diffusion MRI-Based Tissue Microstructure Estimation in the Human Brain: An Overview. Frontiers in Neurology 2023, 14, 1168833. [CrossRef]
| 1 | The idea of leveraging synthetic diffusion signals to enable learning or model-driven estimation when dense training acquisitions are unavailable was introduced in the dMRI literature in [18]. |
| b (s/mm2) | 50 | 70 | 90 | 110 | 350 | 1000 | 1500 | 2500 | 3000 | 3500 | 711 | 3000 |
| TE (ms) | 45 | 53 | 43 | 43 | 54 | 78 | 118 | 88 | 103 | 123 | 78 | 78 |
| (ms) | 5 | 5 | 5 | 5 | 10 | 10 | 10 | 20 | 15 | 15 | 20 | 20 |
| (ms) | 22 | 30 | 20 | 20 | 26 | 50 | 90 | 50 | 70 | 90 | 42 | 42 |
| Ndir | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 38 | 63 |
| Parameter | min | max |
|---|---|---|
| 0 | ||
| 0 | ||
| R () | ||
| () | ||
| () | ||
| () | 0 | 1 |
| () | 0 | |
| () | 0 |
| Parameter | Tumour core (2 voxels) | Peritumoural area (6202 voxels) |
| Cell density | ||
| Extracellular | ||
| Cell diffusivity | ||
| Cell radius R | ||
| Parallel diffusivity | ||
| Transverse diffusivity | ||
| Polar angle | ||
| Azimuthal angle |
| Parameter | Tumour core (1040 voxels) | Peritumoural area (50 voxels) |
| Cell density | ||
| Extracellular | ||
| Cell diffusivity | ||
| Cell radius R | ||
| Parallel diffusivity | ||
| Transverse diffusivity | ||
| Polar angle | ||
| Azimuthal angle |
| Parameter | Tumour core (1230 voxels) | Peritumoural area (606 voxels) |
| Cell density | ||
| Extracellular | ||
| Cell diffusivity | ||
| Cell radius R | ||
| Parallel diffusivity | ||
| Transverse diffusivity | ||
| Polar angle | ||
| Azimuthal angle |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




