Submitted:
31 January 2026
Posted:
03 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
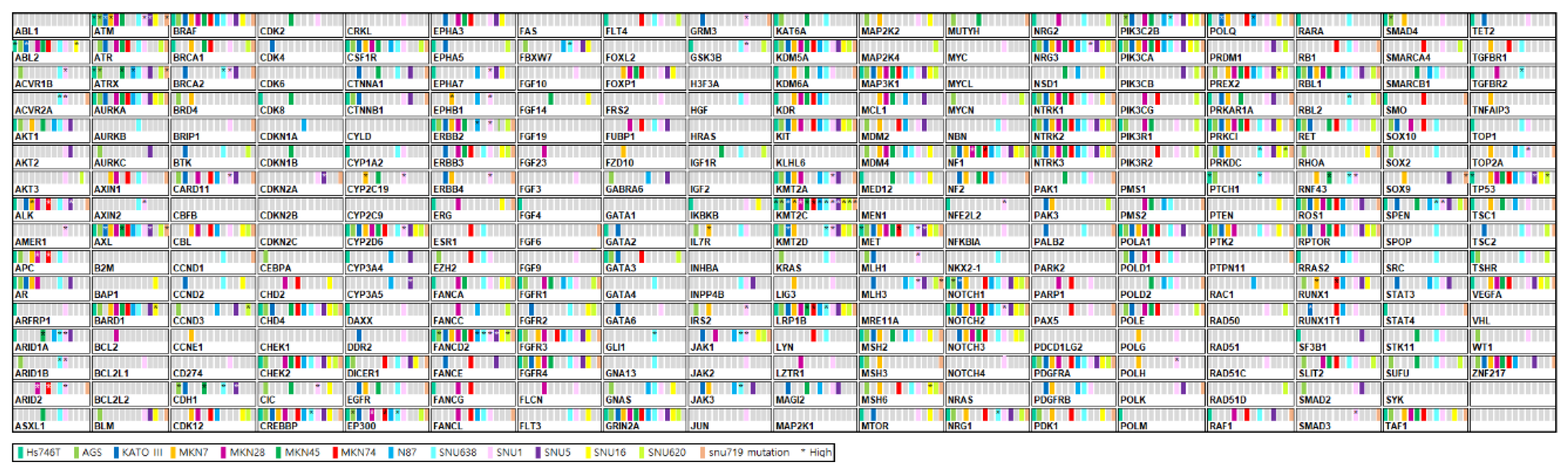
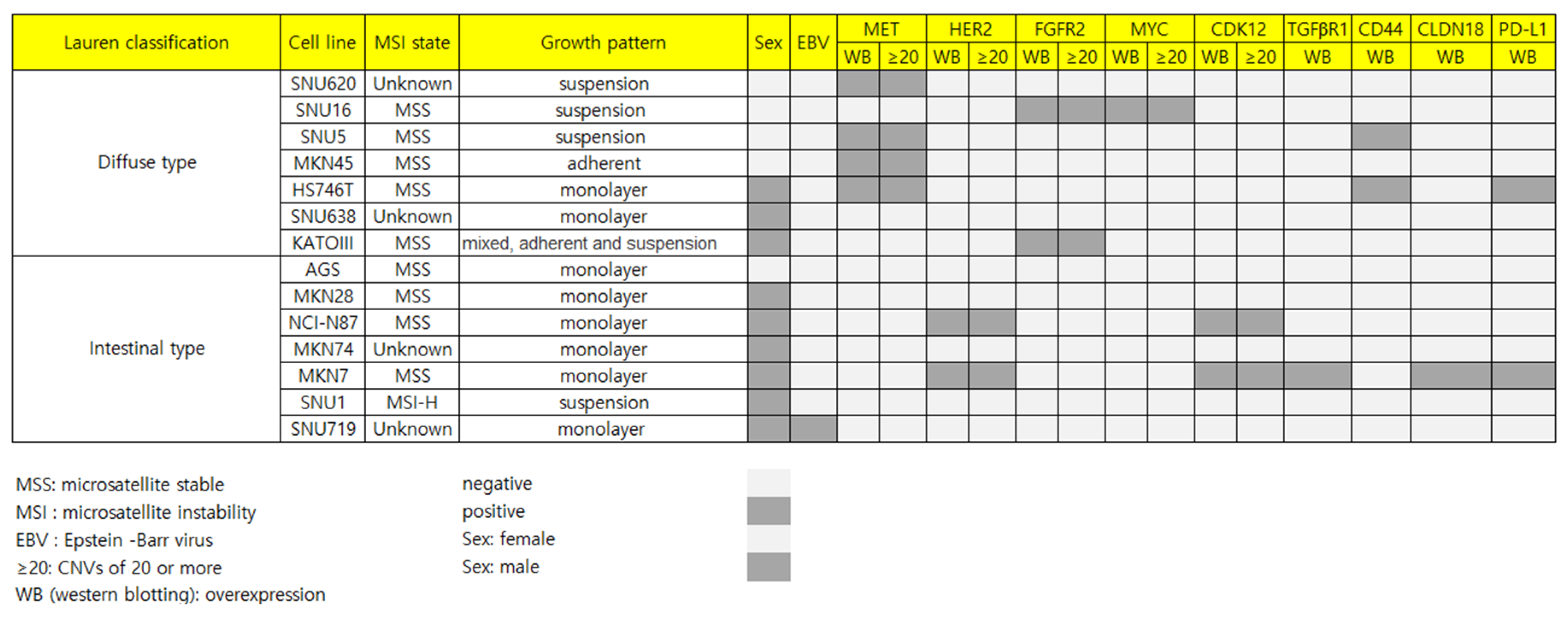
2.1. Characteristics of 14 GC Cell Lines
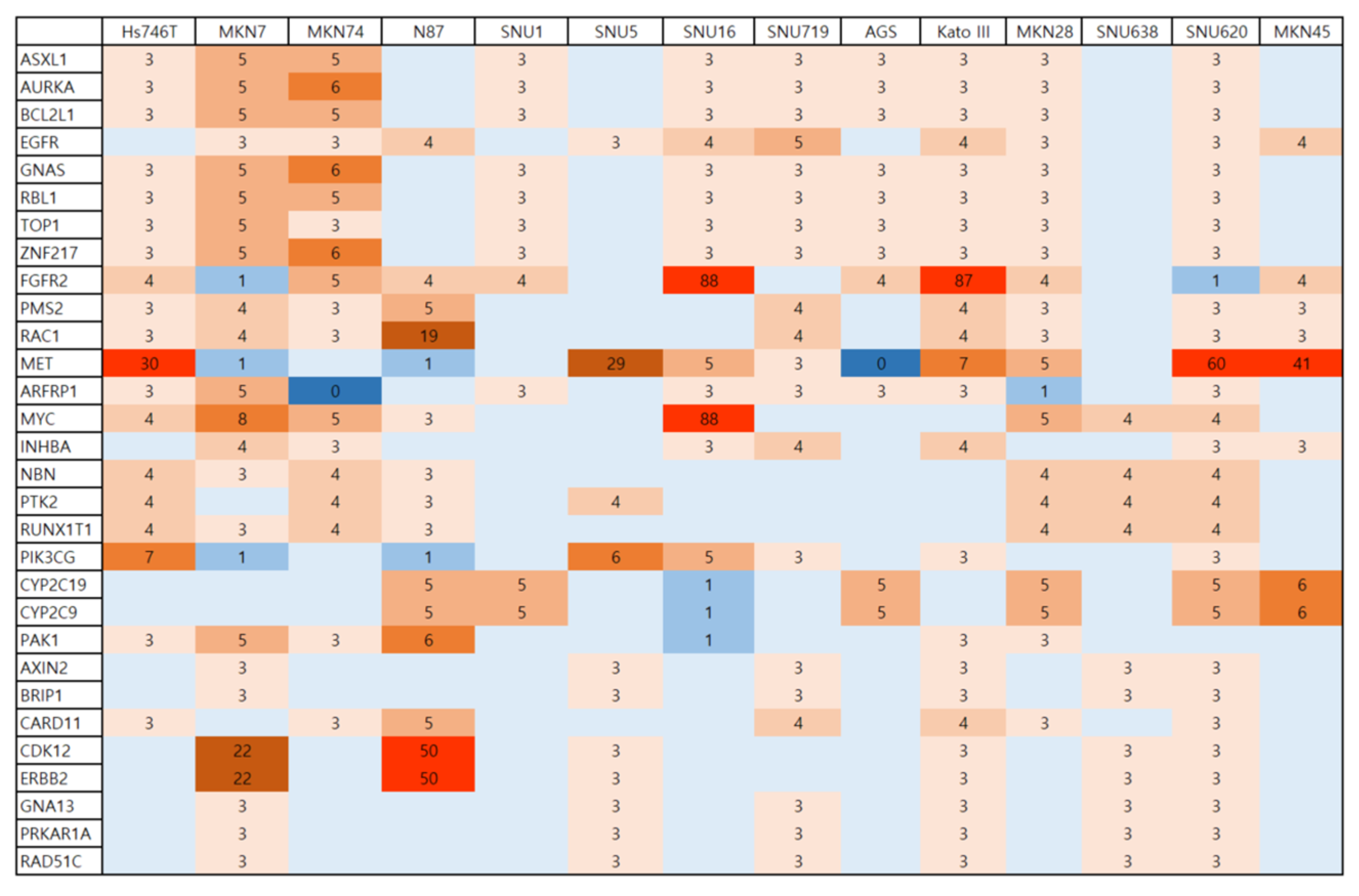
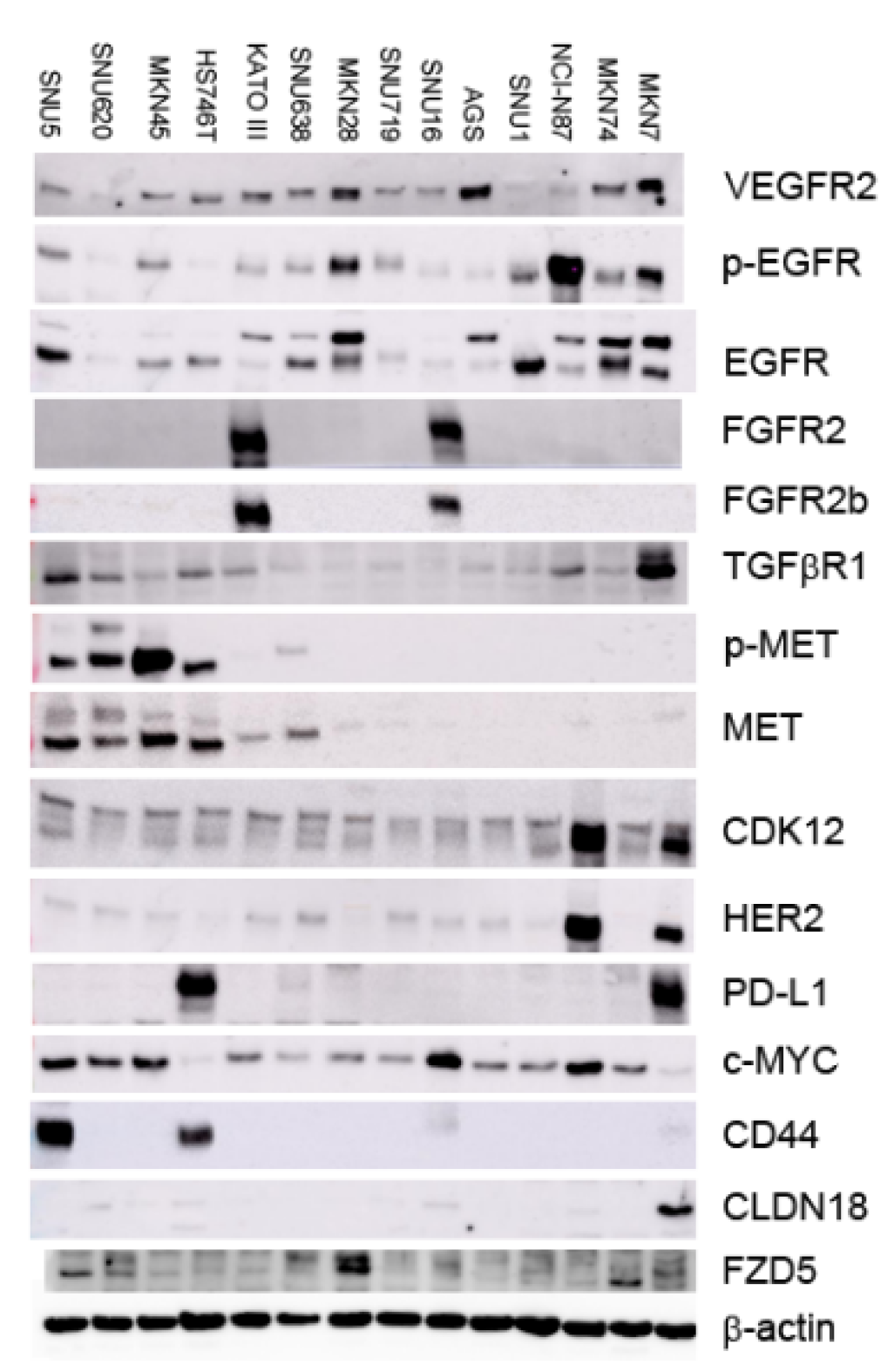
2.2. Patterns of RTKs and GC-Related Targets in the 14 GC Cell Lines
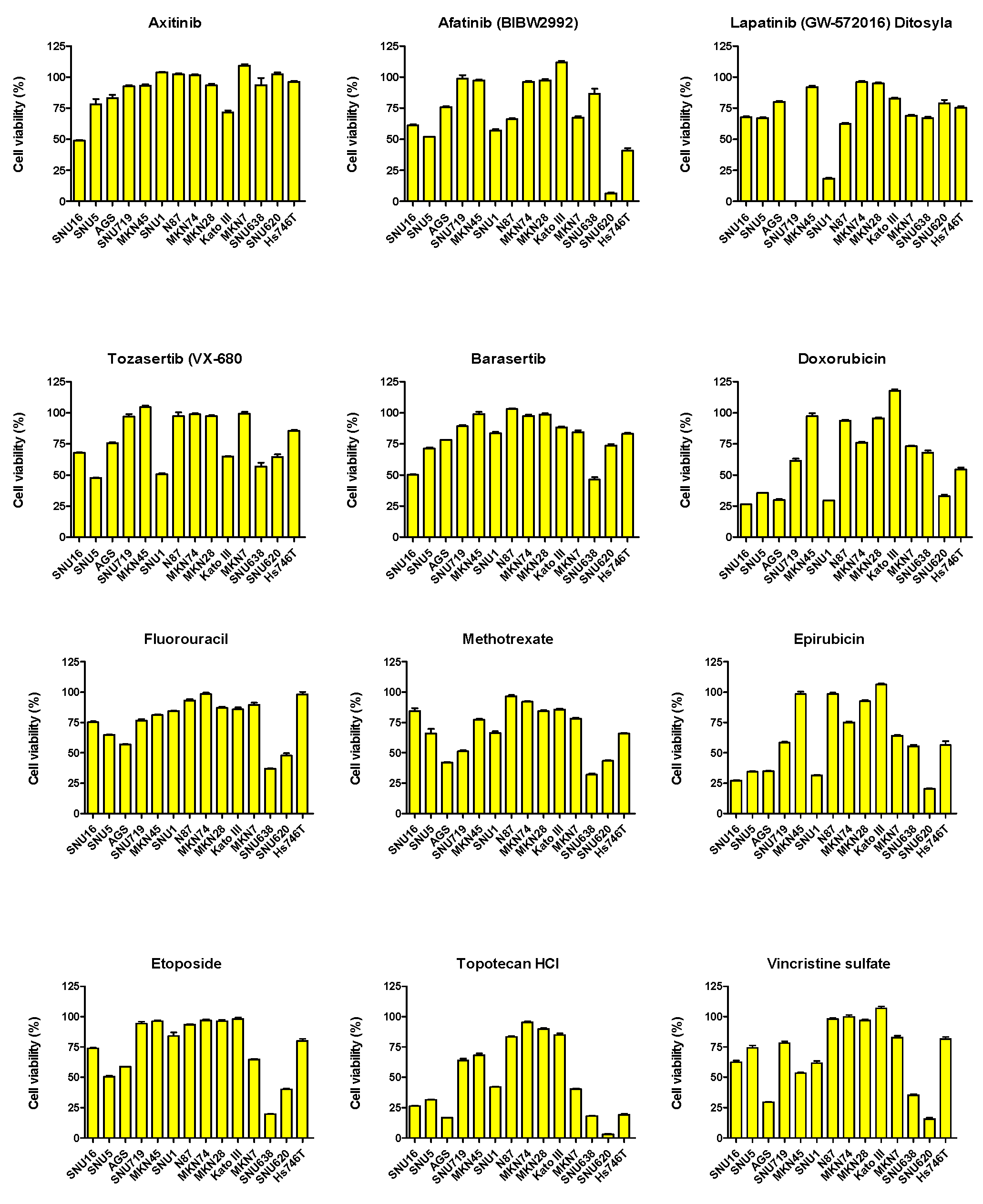
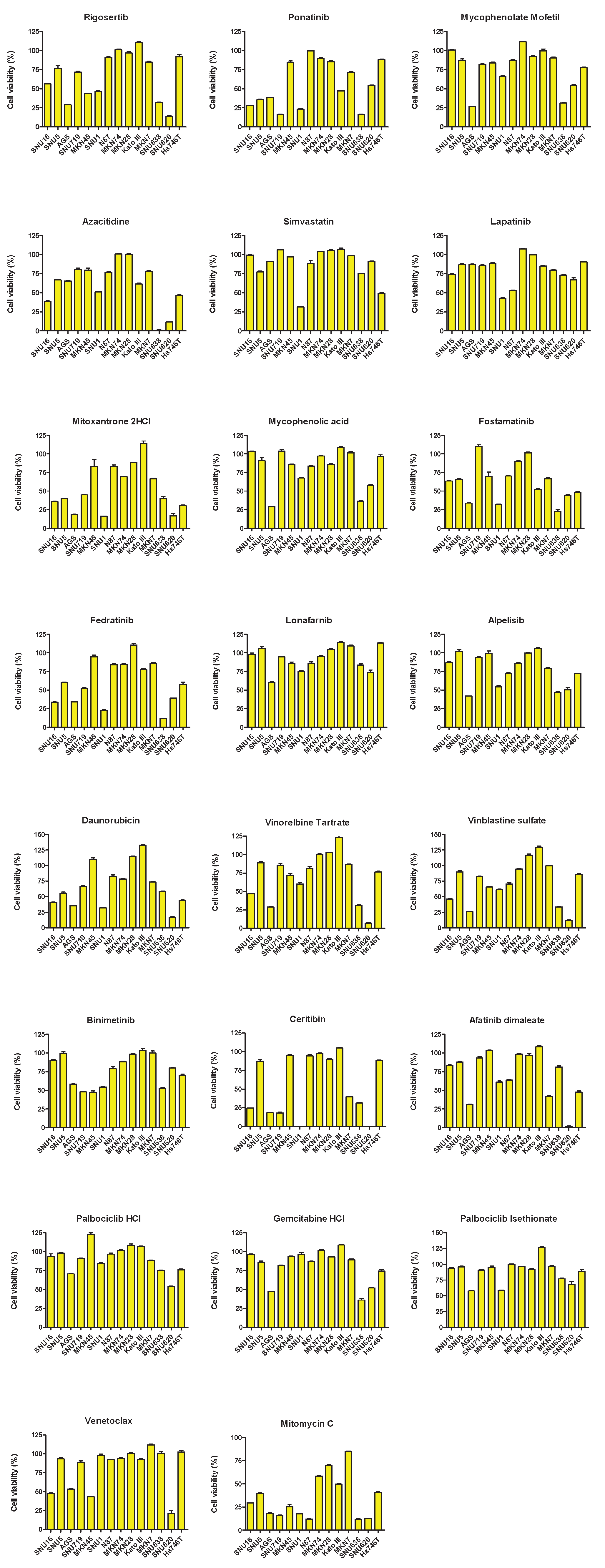
2.3. Sensitivity of 35 Drugs in the 14 GC Cell Lines


3. Discussion
4. Materials and Methods
4.1. Reagents
4.2. Cell Lines and Cell Culture
4.3. Target DNA-Sequencing Analysis
4.4. Cell Viability Assay
4.5. Western Blotting
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| GC | gastric cancer |
| GEJC | gastric ductal carcinoma in situ |
| RTKs | receptor tyrosine kinases |
| MET | mesenchymal–epithelial transition |
| FGFR2 | fibroblast growth factor receptor 2 |
| EGFR/ERBB1/HER1 | epidermal growth factor receptor |
| CNV | copy number variations |
| CLDN18.2 | isoform 2 of CLDN18 |
References
- Jung, K.W.; Won, Y.J.; Kong, H.J.; Lee, E.S.; Community of Population-Based Regional Cancer, R. Cancer Statistics in Korea: Incidence, Mortality, Survival, and Prevalence in 2015. Cancer Res Treat 2018, 50, 303–316. [Google Scholar] [CrossRef]
- Sitarz, R.; Skierucha, M.; Mielko, J.; Offerhaus, G.J.A.; Maciejewski, R.; Polkowski, W.P. Gastric cancer: epidemiology, prevention, classification, and treatment. Cancer Manag Res 2018, 10, 239–248. [Google Scholar] [CrossRef]
- Yeu, Y.; Yoon, Y.; Park, S. Protein localization vector propagation: a method for improving the accuracy of drug repositioning. Mol Biosyst 2015, 11, 2096–2102. [Google Scholar] [CrossRef]
- Xue, H.; Li, J.; Xie, H.; Wang, Y. Review of Drug Repositioning Approaches and Resources. Int J Biol Sci 2018, 14, 1232–1244. [Google Scholar] [CrossRef]
- Maroun, C.R.; Rowlands, T. The Met receptor tyrosine kinase: a key player in oncogenesis and drug resistance. Pharmacol Ther 2014, 142, 316–338. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, K.; Arao, T.; Hamaguchi, T.; Shimada, Y.; Kato, K.; Oda, I.; Taniguchi, H.; Koizumi, F.; Yanagihara, K.; Sasaki, H.; et al. FGFR2 gene amplification and clinicopathological features in gastric cancer. Br J Cancer 2012, 106, 727–732. [Google Scholar] [CrossRef] [PubMed]
- Arienti, C.; Pignatta, S.; Tesei, A. Epidermal Growth Factor Receptor Family and its Role in Gastric Cancer. Front Oncol 2019, 9, 1308. [Google Scholar] [CrossRef] [PubMed]
- Tanner, M.; Hollmen, M.; Junttila, T.T.; Kapanen, A.I.; Tommola, S.; Soini, Y.; Helin, H.; Salo, J.; Joensuu, H.; Sihvo, E.; et al. Amplification of HER-2 in gastric carcinoma: association with Topoisomerase IIalpha gene amplification, intestinal type, poor prognosis and sensitivity to trastuzumab. Ann Oncol 2005, 16, 273–278. [Google Scholar] [CrossRef]
- Lang, S.A.; Klein, D.; Moser, C.; Gaumann, A.; Glockzin, G.; Dahlke, M.H.; Dietmaier, W.; Bolder, U.; Schlitt, H.J.; Geissler, E.K.; et al. Inhibition of heat shock protein 90 impairs epidermal growth factor-mediated signaling in gastric cancer cells and reduces tumor growth and vascularization in vivo. Mol Cancer Ther 2007, 6, 1123–1132. [Google Scholar] [CrossRef]
- Sohn, S.H.; Sul, H.J.; Kim, B.J.; Zang, D.Y. Responses to the Tepotinib in Gastric Cancers with MET Amplification or MET Exon 14 Skipping Mutations and High Expression of Both PD-L1 and CD44. Cancers (Basel) 2022, 14. [Google Scholar] [CrossRef]
- Kawakami, H.; Okamoto, I.; Arao, T.; Okamoto, W.; Matsumoto, K.; Taniguchi, H.; Kuwata, K.; Yamaguchi, H.; Nishio, K.; Nakagawa, K.; et al. MET amplification as a potential therapeutic target in gastric cancer. Oncotarget 2013, 4, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Turner, N.; Grose, R. Fibroblast growth factor signalling: from development to cancer. Nat Rev Cancer 2010, 10, 116–129. [Google Scholar] [CrossRef] [PubMed]
- Moasser, M.M. The oncogene HER2: its signaling and transforming functions and its role in human cancer pathogenesis. Oncogene 2007, 26, 6469–6487. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Yao, X.; Li, S.N.; Suo, A.L.; Tian, T.; Ruan, Z.P.; Guo, H.; Yao, Y. Detection of prostate cancer related copy number variations with SNP genotyping array. Eur Rev Med Pharmacol Sci 2013, 17, 2916–2922. [Google Scholar]
- Shlien, A.; Malkin, D. Copy number variations and cancer. Genome Med 2009, 1, 62. [Google Scholar] [CrossRef]
- Xu, H.; Zhu, X.; Xu, Z.; Hu, Y.; Bo, S.; Xing, T.; Zhu, K. Non-invasive Analysis of Genomic Copy Number Variation in Patients with Hepatocellular Carcinoma by Next Generation DNA Sequencing. J Cancer 2015, 6, 247–253. [Google Scholar] [CrossRef]
- Sohn, S.H.; Sul, H.J.; Kim, B.J.; Zang, D.Y. Comparison of Tepotinib, Paclitaxel, or Ramucirumab Efficacy According to the Copy Number or Phosphorylation Status of the MET Gene: Doublet Treatment versus Single Agent Treatment. Int J Mol Sci 2024, 25. [Google Scholar] [CrossRef]
- Sertkaya, A.; Beleche, T.; Jessup, A.; Sommers, B.D. Costs of Drug Development and Research and Development Intensity in the US, 2000-2018. JAMA Netw Open 2024, 7, e2415445. [Google Scholar] [CrossRef]
- Jourdan, J.P.; Bureau, R.; Rochais, C.; Dallemagne, P. Drug repositioning: a brief overview. J Pharm Pharmacol 2020, 72, 1145–1151. [Google Scholar] [CrossRef]
- Panahizadeh, R.; Panahi, P.; Asghariazar, V.; Makaremi, S.; Noorkhajavi, G.; Safarzadeh, E. A literature review of recent advances in gastric cancer treatment: exploring the cross-talk between targeted therapies. Cancer Cell Int 2025, 25, 23. [Google Scholar] [CrossRef]
- Becker, J.C.; Muller-Tidow, C.; Serve, H.; Domschke, W.; Pohle, T. Role of receptor tyrosine kinases in gastric cancer: new targets for a selective therapy. World J Gastroenterol 2006, 12, 3297–3305. [Google Scholar] [CrossRef]
- Hashimoto, I.; Oshima, T. Claudins and Gastric Cancer: An Overview. Cancers (Basel) 2022, 14. [Google Scholar] [CrossRef]
- Agnarelli, A.; Vella, V.; Samuels, M.; Papanastasopoulos, P.; Giamas, G. Incorporating Immunotherapy in the Management of Gastric Cancer: Molecular and Clinical Implications. Cancers (Basel) 2022, 14. [Google Scholar] [CrossRef] [PubMed]
- Qin, S.; Li, A.; Yi, M.; Yu, S.; Zhang, M.; Wu, K. Recent advances on anti-angiogenesis receptor tyrosine kinase inhibitors in cancer therapy. J Hematol Oncol 2019, 12, 27. [Google Scholar] [CrossRef] [PubMed]
- Zhong, L.; Li, Y.; Xiong, L.; Wang, W.; Wu, M.; Yuan, T.; Yang, W.; Tian, C.; Miao, Z.; Wang, T.; et al. Small molecules in targeted cancer therapy: advances, challenges, and future perspectives. Signal Transduct Target Ther 2021, 6, 201. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.Z.; Wang, J.Q.; Chen, J.L.; Zhang, X.L.; Zhang, M.M.; Wang, S.L.; Zhao, C. CDK12 Promotes the Proliferation, Migration, and Angiogenesis of Gastric Carcinoma via Activating the PI3K/AKT/mTOR Signaling Pathway. Appl Biochem Biotechnol 2023, 195, 6913–6926. [Google Scholar] [CrossRef]
- Ikushima, H.; Miyazono, K. TGFbeta signalling: a complex web in cancer progression. Nat Rev Cancer 2010, 10, 415–424. [Google Scholar] [CrossRef]
- Jin, Z.; Huang, Z.; Wu, C.; Zhang, F.; Gao, Y.; Guo, S.; Tao, X.; Lu, S.; Zhang, J.; Huang, J.; et al. Molecular insights into gastric cancer: The impact of TGFBR2 and hsa-mir-107 revealed by microarray sequencing and bioinformatics. Comput Biol Med 2024, 172, 108221. [Google Scholar] [CrossRef]
- Sahin, U.; Koslowski, M.; Dhaene, K.; Usener, D.; Brandenburg, G.; Seitz, G.; Huber, C.; Tureci, O. Claudin-18 splice variant 2 is a pan-cancer target suitable for therapeutic antibody development. Clin Cancer Res 2008, 14, 7624–7634. [Google Scholar] [CrossRef]
- Boger, C.; Behrens, H.M.; Mathiak, M.; Kruger, S.; Kalthoff, H.; Rocken, C. PD-L1 is an independent prognostic predictor in gastric cancer of Western patients. Oncotarget 2016, 7, 24269–24283. [Google Scholar] [CrossRef]
- Jung, W.Y.; Kang, Y.; Lee, H.; Mok, Y.J.; Kim, H.K.; Kim, A.; Kim, B.H. Expression of moesin and CD44 is associated with poor prognosis in gastric adenocarcinoma. Histopathology 2013, 63, 474–481. [Google Scholar] [CrossRef]
- Hou, W.; Kong, L.; Hou, Z.; Ji, H. CD44 is a prognostic biomarker and correlated with immune infiltrates in gastric cancer. BMC Med Genomics 2022, 15, 225. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



