Submitted:
01 February 2026
Posted:
02 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods:
2.1. Chicken Samples Collection and Processing
2.2. The Total Viral DNA Isolation and Testing for the ILTV by Real-Time PCR
2.3. The Next Generation Sequencing (NGS)
2.4. Multiple Sequence Alignment (MSA) and Phylogenetic Analysis
2.5. Comparative Divergence Analysis of the Key ILTV Proteins
2.6. In Silico Structural Modeling of Some Key ILTV Proteins
2.7. Statistical Analysis
3. Results:
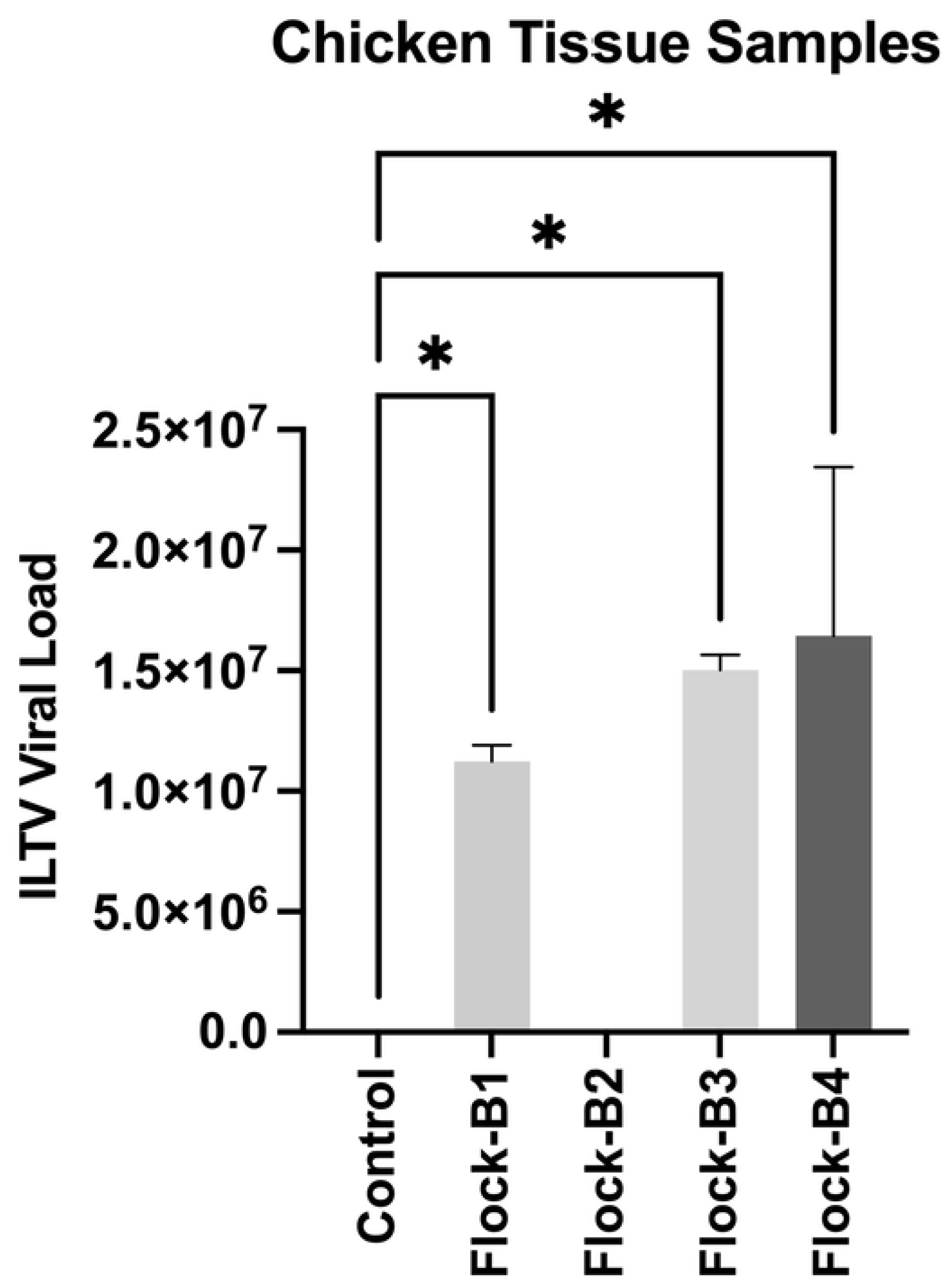
3.1. Molecular Detection and Selection of ILTV Isolates
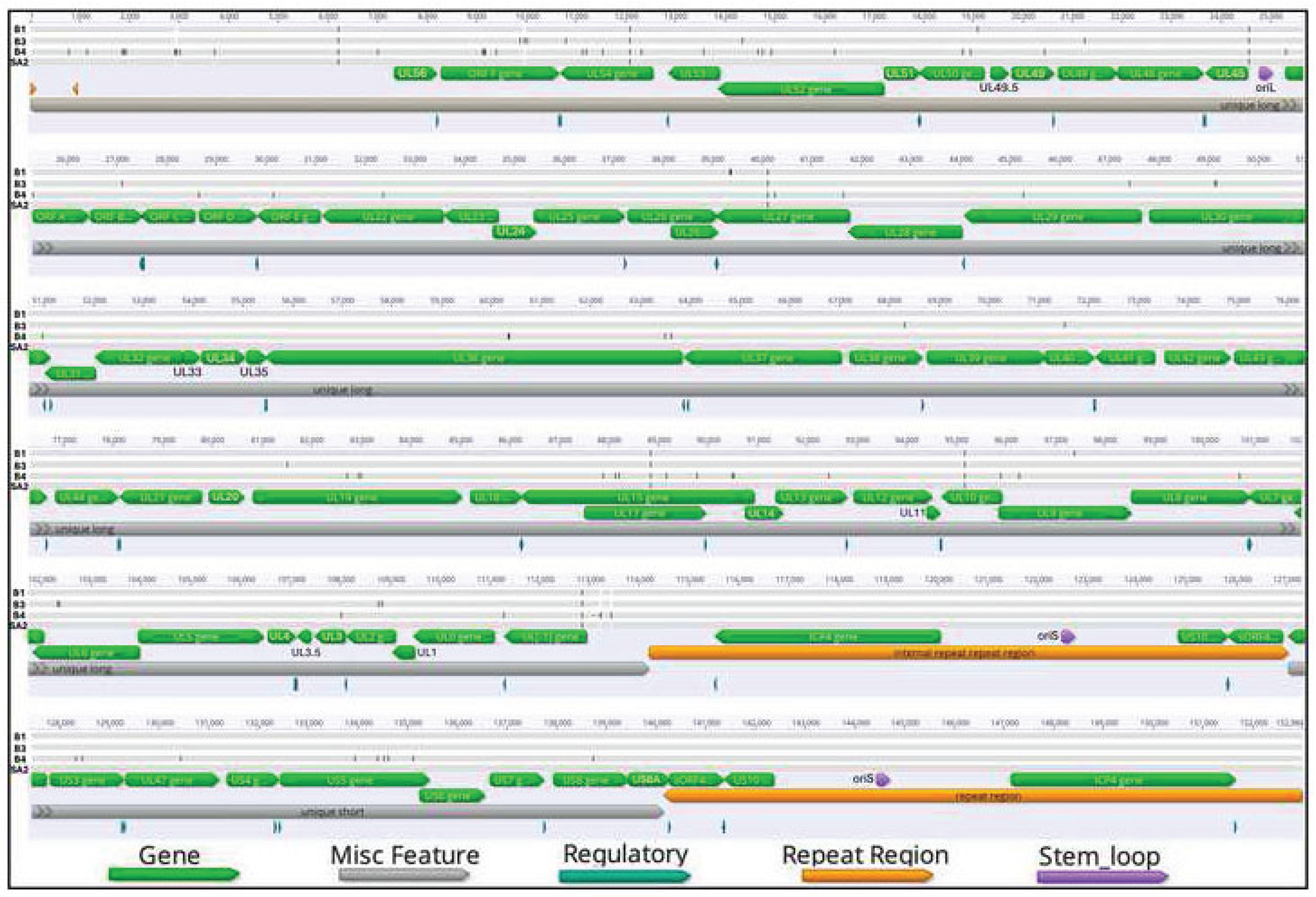
3.2. The Genome Structure and Organization of ILTV Isolates Reported in This Study
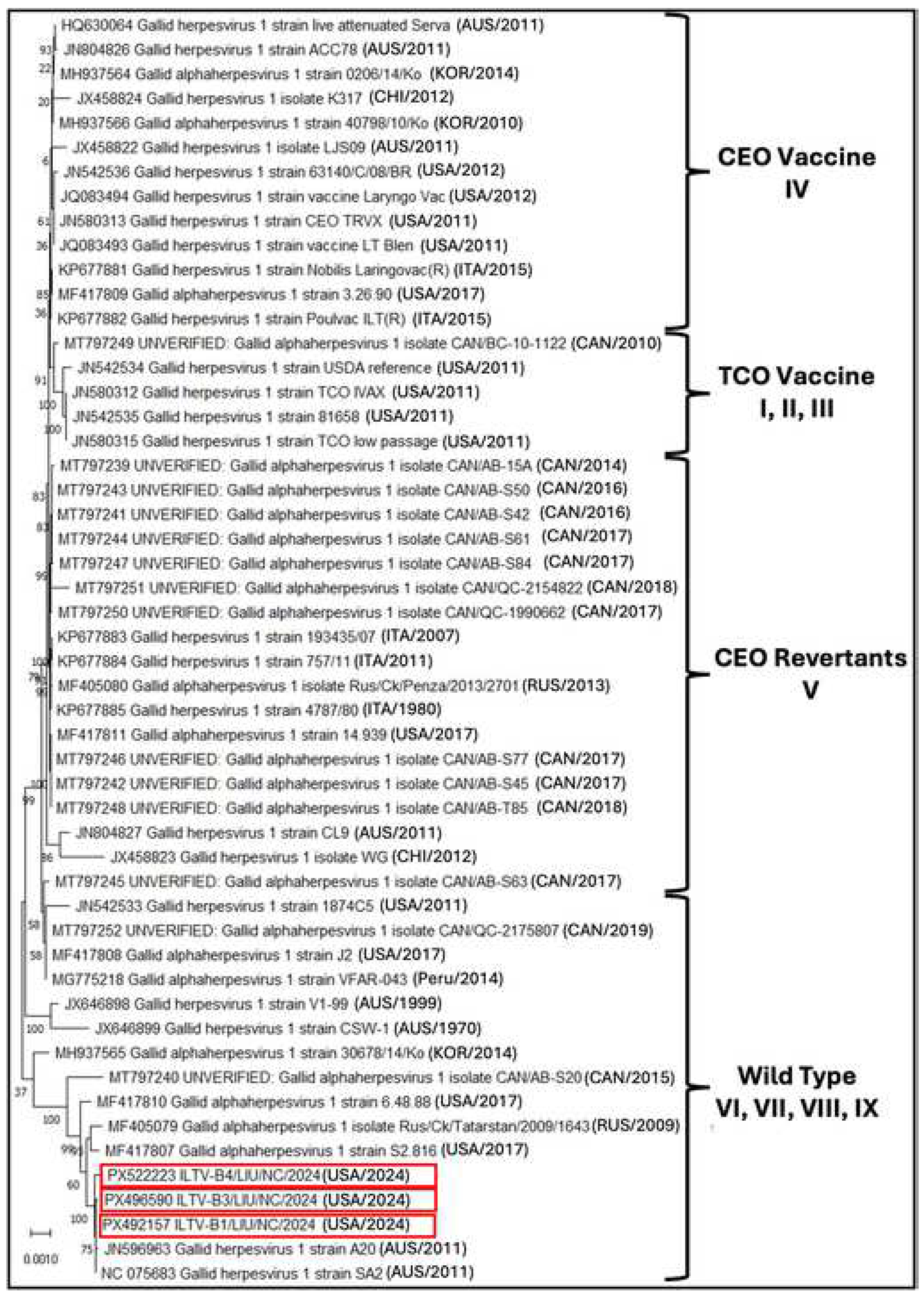
3.3. Phylogenetic Analysis Based on the Full-Length Genome Sequences of ILTV-B1, B3, and B4
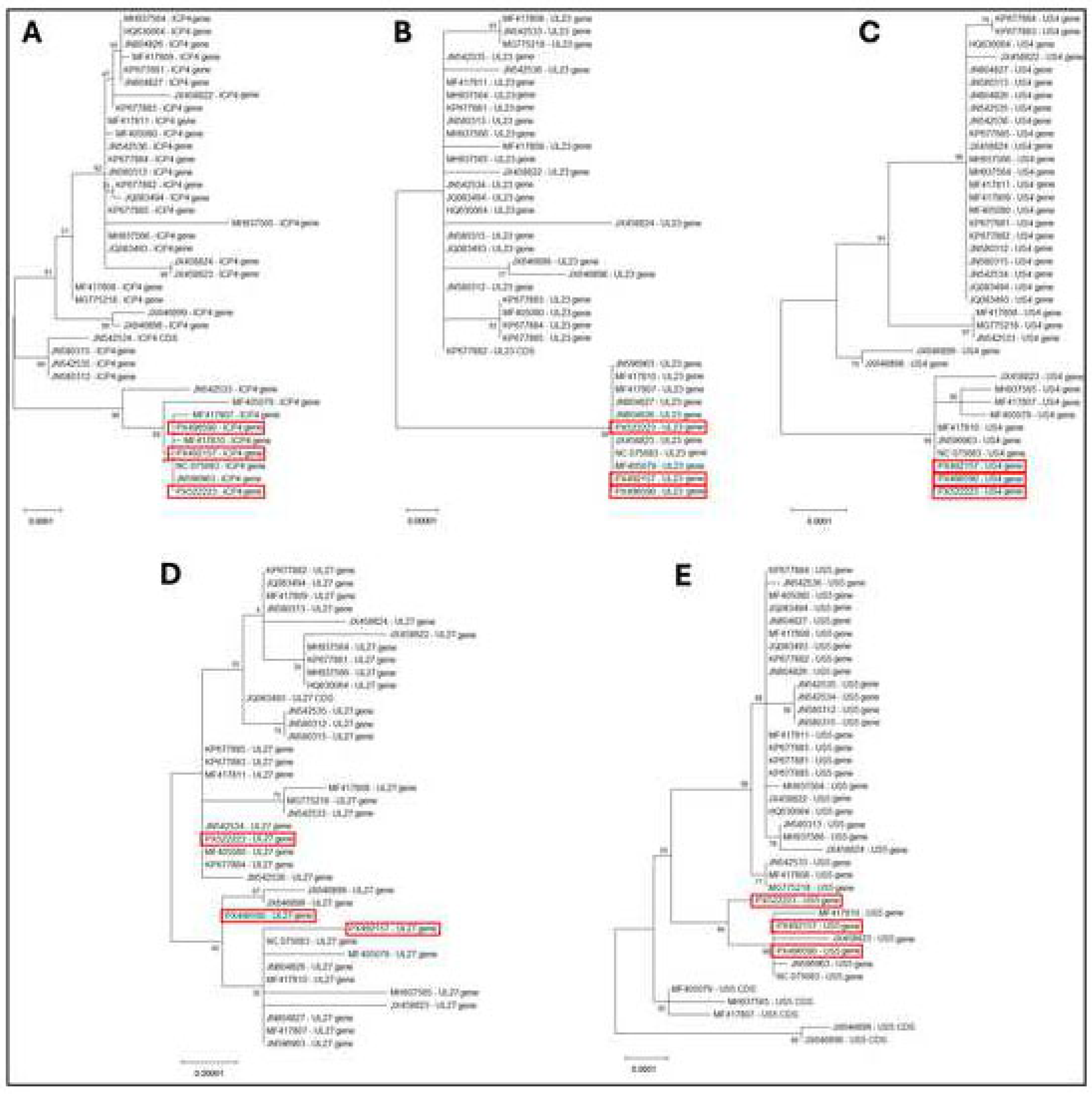
3.4. Phylogenetic Analysis of Some Key Genes of ILTV-B1, B3, and B4 Isolates
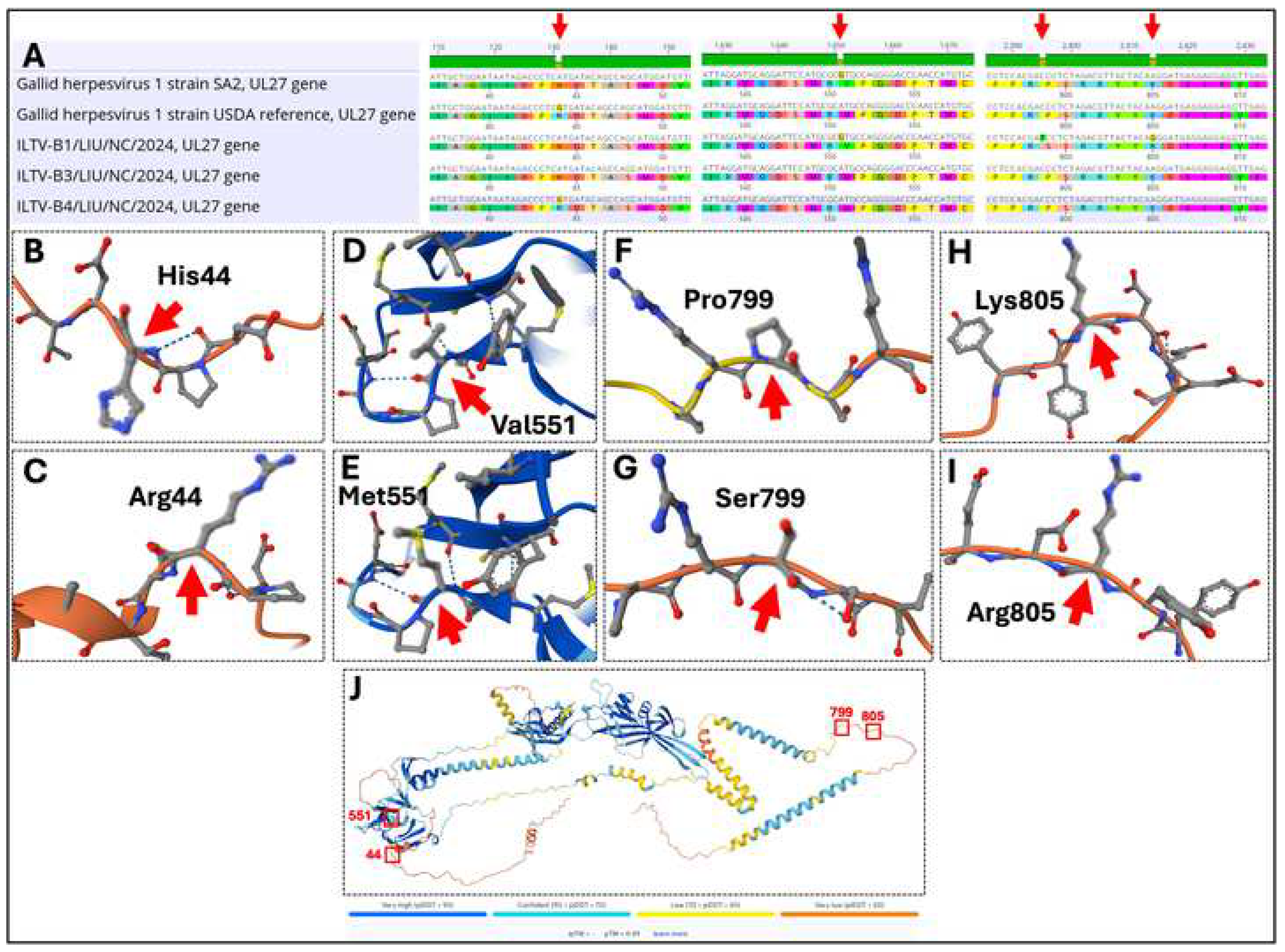
3.5. Mapping the Notable Mutations Within the UL27 (gB) Genes of ILTV-(B1, B3, and B4) Isolates Reported in This Study
3.6. Mutations and Substitution Analysis Within the US5 (gJ) Gene of ILTV- Isolates Reported in This Study
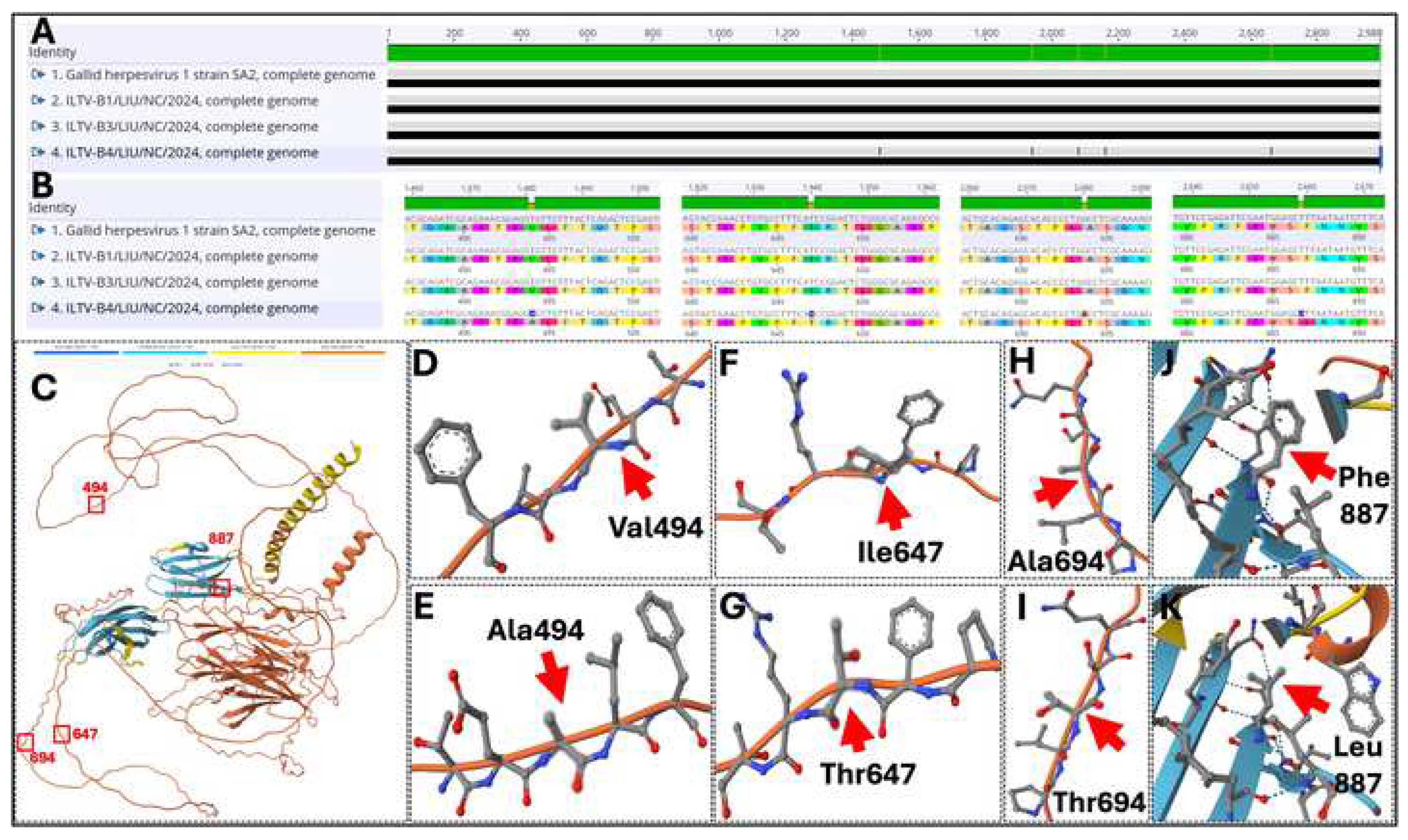
3.7. Mutations and Substitution Analysis Within the Structural Proteins of ILTV-B1, ILTV-B3, ILTV-B4 Compared to the ILTV Vaccine Strains
4. Discussion
Acknowledgments
References
- Cover, M.S., The early history of infectious laryngotracheitis. Avian Dis, 1996. 40(3): p. 494-500. [CrossRef]
- Davison, A.J., et al., The order Herpesvirales. Arch Virol, 2009. 154(1): p. 171-7. [CrossRef]
- Mettenleiter, T.C., B.G. Klupp, and H. Granzow, Herpesvirus assembly: an update. Virus Res, 2009. 143(2): p. 222-34. [CrossRef]
- Fuchs, W., et al., Molecular biology of avian infectious laryngotracheitis virus. Vet Res, 2007. 38(2): p. 261-79. [CrossRef]
- Garcia, M., Current and future vaccines and vaccination strategies against infectious laryngotracheitis (ILT) respiratory disease of poultry. Vet Microbiol, 2017. 206: p. 157-162. [CrossRef]
- Crawshaw, G.J. and B.R. Boycott, Infectious laryngotracheitis in peafowl and pheasants. Avian Dis, 1982. 26(2): p. 397-401. [CrossRef]
- Winterfield, R.W. and I.G. So, Susceptibility of turkeys to infectious laryngotracheitis. Avian Dis, 1968. 12(1): p. 191-202. [CrossRef]
- Ou, S.C. and J.J. Giambrone, Infectious laryngotracheitis virus in chickens. World J Virol, 2012. 1(5): p. 142-9. [CrossRef]
- Watrach, A.M., L.E. Hanson, and M.A. Watrach, The Structure of Infectious Laryngotracheitis Virus. Virology, 1963. 21: p. 601-8. [CrossRef]
- Mo, J. and J. Mo, Infectious Laryngotracheitis Virus and Avian Metapneumovirus: A Comprehensive Review. Pathogens, 2025. 14(1). [CrossRef]
- McGeoch, D.J., A. Dolan, and A.C. Ralph, Toward a comprehensive phylogeny for mammalian and avian herpesviruses. J Virol, 2000. 74(22): p. 10401-6. [CrossRef]
- Morales Ruiz, S., et al., Full-Genome Sequence of Infectious Laryngotracheitis Virus (Gallid Alphaherpesvirus 1) Strain VFAR-043, Isolated in Peru. Genome Announc, 2018. 6(10). [CrossRef]
- Lee, S.W., et al., First complete genome sequence of infectious laryngotracheitis virus. BMC Genomics, 2011. 12: p. 197. [CrossRef]
- Granzow, H., et al., Egress of alphaherpesviruses: comparative ultrastructural study. J Virol, 2001. 75(8): p. 3675-84. [CrossRef]
- Piccirillo, A., et al., Full Genome Sequence-Based Comparative Study of Wild-Type and Vaccine Strains of Infectious Laryngotracheitis Virus from Italy. PLoS One, 2016. 11(2): p. e0149529. [CrossRef]
- Fuchs, W. and T.C. Mettenleiter, DNA sequence and transcriptional analysis of the UL1 to UL5 gene cluster of infectious laryngotracheitis virus. J Gen Virol, 1996. 77 ( Pt 9): p. 2221-9. [CrossRef]
- Wild, M.A., S. Cook, and M. Cochran, A genomic map of infectious laryngotracheitis virus and the sequence and organization of genes present in the unique short and flanking regions. Virus Genes, 1996. 12(2): p. 107-16. [CrossRef]
- McGeoch, D.J., et al., The complete DNA sequence of the long unique region in the genome of herpes simplex virus type 1. J Gen Virol, 1988. 69 ( Pt 7): p. 1531-74. [CrossRef]
- Thureen, D.R. and C.L. Keeler, Jr., Psittacid herpesvirus 1 and infectious laryngotracheitis virus: Comparative genome sequence analysis of two avian alphaherpesviruses. J Virol, 2006. 80(16): p. 7863-72. [CrossRef]
- Nadimpalli, M., et al., Impairment of infectious laryngotracheitis virus replication by deletion of the UL[-1] gene. Arch Virol, 2017. 162(6): p. 1541-1548. [CrossRef]
- McGeoch, D.J., F.J. Rixon, and A.J. Davison, Topics in herpesvirus genomics and evolution. Virus Res, 2006. 117(1): p. 90-104. [CrossRef]
- Kelly, B.J., et al., Functional roles of the tegument proteins of herpes simplex virus type 1. Virus Res, 2009. 145(2): p. 173-86. [CrossRef]
- Menendez, K.R., et al., Molecular epidemiology of infectious laryngotracheitis: a review. Avian Pathol, 2014. 43(2): p. 108-17. [CrossRef]
- Blakey, J., et al., Retrospective analysis of infectious laryngotracheitis in backyard chicken flocks in California, 2007-2017, and determination of strain origin by partial ICP4 sequencing. J Vet Diagn Invest, 2019. 31(3): p. 350-358. [CrossRef]
- Devlin, J.M., et al., Glycoprotein G is a virulence factor in infectious laryngotracheitis virus. J Gen Virol, 2006. 87(Pt 10): p. 2839-2847. [CrossRef]
- Perez Contreras, A., et al., Analysis of Whole-Genome Sequences of Infectious laryngotracheitis Virus Isolates from Poultry Flocks in Canada: Evidence of Recombination. Viruses, 2020. 12(11). [CrossRef]
- Hernandez-Divers, S.M., et al., Backyard chicken flocks pose a disease risk for neotropic birds in Costa Rica. Avian Dis, 2008. 52(4): p. 558-66. [CrossRef]
- Dormitorio, T.V., J.J. Giambrone, and K.S. Macklin, Detection and isolation of infectious laryngotracheitis virus on a broiler farm after a disease outbreak. Avian Dis, 2013. 57(4): p. 803-7. [CrossRef]
- Parra, S.H., et al., Persistence of the tissue culture origin vaccine for infectious laryngotracheitis virus in commercial chicken flocks in Brazil. Poult Sci, 2015. 94(11): p. 2608-15. [CrossRef]
- Chacon, J.L., et al., Persistence and spreading of field and vaccine strains of infectious laryngotracheitis virus (ILTV) in vaccinated and unvaccinated geographic regions, in Brazil. Trop Anim Health Prod, 2015. 47(6): p. 1101-8. [CrossRef]
- Garcia, M. and G. Zavala, Commercial Vaccines and Vaccination Strategies Against Infectious Laryngotracheitis: What We Have Learned and Knowledge Gaps That Remain. Avian Dis, 2019. 63(2): p. 325-334. [CrossRef]
- Barboza-Solis, C., et al., Genotyping of Infectious Laryngotracheitis Virus (ILTV) Isolates from Western Canadian Provinces of Alberta and British Columbia Based on Partial Open Reading Frame (ORF) a and b. Animals (Basel), 2020. 10(9). [CrossRef]
- Ojkic, D., et al., Characterization of field isolates of infectious laryngotracheitis virus from Ontario. Avian Pathol, 2006. 35(4): p. 286-92. [CrossRef]
- Al-Mubarak, A.I.A., et al., Detection of Avian Orthoavulavirus-1 genotypes VI.2.1 and VII.1.1 with neuro-viscerotropic tropism in some backyard pigeons (Columbidae) in Eastern Saudi Arabia. Front Vet Sci, 2024. 11: p. 1352636. [CrossRef]
- Al-Mubarak, A.I.A., et al., Avian encephalomyelitis virus in backyard chickens. Vet World, 2023. 16(9): p. 1866-1870. [CrossRef]
- Alsultan, M.A., M.A. Alhammadi, and M.G. Hemida, Infectious bronchitis virus from chickens in Al-Hasa, Saudi Arabia 2015-2016. Vet World, 2019. 12(3): p. 424-433. [CrossRef]
- Ba Abduallah, M.M. and M.G. Hemida, Comparative analysis of the genome structure and organization of the Middle East respiratory syndrome coronavirus (MERS-CoV) 2012 to 2019 revealing evidence for virus strain barcoding, zoonotic transmission, and selection pressure. Rev Med Virol, 2021. 31(1): p. 1-12. [CrossRef]
- Hemida, M.G., The next-generation coronavirus diagnostic techniques with particular emphasis on the SARS-CoV-2. J Med Virol, 2021. 93(7): p. 4219-4241. [CrossRef]
- Hemida, M.G., et al., MERS coronavirus in dromedary camel herd, Saudi Arabia. Emerg Infect Dis, 2014. 20(7): p. 1231-4. [CrossRef]
- Hemida, M.G., et al., Phylogenetic Analysis of MERS-CoV in a Camel Abattoir, Saudi Arabia, 2016-2018. Emerg Infect Dis, 2020. 26(12): p. 3089-3091. [CrossRef]
- Hemida, M.G., et al., Foot-and-mouth disease virus O/ME-SA/Ind 2001 lineage outbreak in vaccinated Holstein Friesian cattle in Saudi Arabia in 2016. Vet Q, 2018. 38(1): p. 88-98. [CrossRef]
- Malik, Y.S., et al., SARS-CoV-2 Spike Protein Extrapolation for COVID Diagnosis and Vaccine Development. Front Mol Biosci, 2021. 8: p. 607886. [CrossRef]
- Shah, A.U., et al., Comparative Genome Sequencing Analysis of Some Novel Feline Infectious Peritonitis Viruses Isolated from Some Feral Cats in Long Island. Viruses, 2025. 17(2). [CrossRef]
- Shah, A.U., P. Gauger, and M.G. Hemida, Isolation and molecular characterization of an enteric isolate of the genotype-Ia bovine coronavirus with notable mutations in the receptor binding domain of the spike glycoprotein. Virology, 2025. 603: p. 110313. [CrossRef]
- Singh, K., Y.S. Malik, and M.G. Hemida, Comprehensive Evolutionary and Structural Analysis of the H5N1 Clade 2.4.3.4b Influenza a Virus Based on the Sequences and Data Mining of the Hemagglutinin, Nucleoprotein and Neuraminidase Genes Across Multiple Hosts. Pathogens, 2025. 14(9). [CrossRef]
- Madsen, J.M., et al., Evaluation of Maryland backyard flocks and biosecurity practices. Avian Dis, 2013. 57(2): p. 233-7. [CrossRef]
- Spatz, S.J., et al., MinION sequencing to genotype US strains of infectious laryngotracheitis virus. Avian Pathol, 2019. 48(3): p. 255-269. [CrossRef]
- Oldoni, I., et al., Pathogenicity and growth characteristics of selected infectious laryngotracheitis virus strains from the United States. Avian Pathol, 2009. 38(1): p. 47-53. [CrossRef]
- Chacon, J.L., et al., Molecular Characterization of the Infectious Laryngotracheitis Virus (ILTV) Involved in Poultry Outbreaks Reveals the Virus Origin and Estimated Spreading Route. Viruses, 2025. 17(2). [CrossRef]
- Shah, A.U., et al., Some novel field isolates belonging to lineage-1 of the genotype GI-avian infectious bronchitis virus (AIBV) show strong evidence of recombination with field/vaccinal strains. Infect Genet Evol, 2025. 129: p. 105723. [CrossRef]
- Rozen, S. and H. Skaletsky, Primer3 on the WWW for general users and for biologist programmers. Methods Mol Biol, 2000. 132: p. 365-86.
- Shah, A.U., et al., From nasal to basal: single-cell sequencing of the bursa of Fabricius highlights the IBDV infection mechanism in chickens. Cell Biosci, 2021. 11(1): p. 212. [CrossRef]
- Livak, K.J. and T.D. Schmittgen, Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods, 2001. 25(4): p. 402-8. [CrossRef]
- Kumar, S., et al., MEGA12: Molecular Evolutionary Genetic Analysis Version 12 for Adaptive and Green Computing. Mol Biol Evol, 2024. 41(12). [CrossRef]
- Abramson, J., et al., Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature, 2024. 630(8016): p. 493-500. [CrossRef]
- Thiry, E., et al., Recombination in alphaherpesviruses. Rev Med Virol, 2005. 15(2): p. 89-103. [CrossRef]
- Uchida, H., et al., A double mutation in glycoprotein gB compensates for ineffective gD-dependent initiation of herpes simplex virus type 1 infection. J Virol, 2010. 84(23): p. 12200-9. [CrossRef]
- Dufour-Zavala, L., Epizootiology of infectious laryngotracheitis and presentation of an industry control program. Avian Dis, 2008. 52(1): p. 1-7. [CrossRef]
- Kong, C., et al., Complete genome sequence of the first Chinese virulent infectious laryngotracheitis virus. PLoS One, 2013. 8(7): p. e70154. [CrossRef]
- Godowski, P.J. and D.M. Knipe, Transcriptional control of herpesvirus gene expression: gene functions required for positive and negative regulation. Proc Natl Acad Sci U S A, 1986. 83(2): p. 256-60. [CrossRef]
- Watson, R.J. and J.B. Clements, A herpes simplex virus type 1 function continuously required for early and late virus RNA synthesis. Nature, 1980. 285(5763): p. 329-30. [CrossRef]
- Wagner, L.M., et al., The N terminus and C terminus of herpes simplex virus 1 ICP4 cooperate to activate viral gene expression. J Virol, 2012. 86(12): p. 6862-74. [CrossRef]
- Huang, W., et al., The influence of stress factors on the reactivation of latent herpes simplex virus type 1 in infected mice. Cell Biochem Biophys, 2011. 61(1): p. 115-22. [CrossRef]
- Wang, J., et al., Complete genome sequence of virulent duck enteritis virus (DEV) strain 2085 and comparison with genome sequences of virulent and attenuated DEV strains. Virus Res, 2011. 160(1-2): p. 316-25. [CrossRef]
- Williams, R.K. and S.E. Straus, Specificity and affinity of binding of herpes simplex virus type 2 glycoprotein B to glycosaminoglycans. J Virol, 1997. 71(2): p. 1375-80. [CrossRef]
- Bender, F.C., et al., Herpes simplex virus glycoprotein B binds to cell surfaces independently of heparan sulfate and blocks virus entry. J Virol, 2005. 79(18): p. 11588-97. [CrossRef]
- Mardberg, K., et al., Basic amino acids as modulators of an O-linked glycosylation signal of the herpes simplex virus type 1 glycoprotein gC: functional roles in viral infectivity. Glycobiology, 2004. 14(7): p. 571-81. [CrossRef]
- Adamiak, B., et al., Human antibodies to herpes simplex virus type 1 glycoprotein C are neutralizing and target the heparan sulfate-binding domain. Virology, 2010. 400(2): p. 197-206. [CrossRef]
- Whitbeck, J.C., et al., Glycoprotein D of herpes simplex virus (HSV) binds directly to HVEM, a member of the tumor necrosis factor receptor superfamily and a mediator of HSV entry. J Virol, 1997. 71(8): p. 6083-93. [CrossRef]
- Whalley, J.M., et al., Host cell tropism of equine herpesviruses: glycoprotein D of EHV-1 enables EHV-4 to infect a non-permissive cell line. Arch Virol, 2007. 152(4): p. 717-25. [CrossRef]
- Westra, D.F., et al., Glycoprotein H of herpes simplex virus type 1 requires glycoprotein L for transport to the surfaces of insect cells. J Virol, 1997. 71(3): p. 2285-91. [CrossRef]
- Chen, J., T.S. Jardetzky, and R. Longnecker, The large groove found in the gH/gL structure is an important functional domain for Epstein-Barr virus fusion. J Virol, 2013. 87(7): p. 3620-7. [CrossRef]
- Heldwein, E.E. and C. Krummenacher, Entry of herpesviruses into mammalian cells. Cell Mol Life Sci, 2008. 65(11): p. 1653-68. [CrossRef]
- Uchida, H., et al., Novel mutations in gB and gH circumvent the requirement for known gD Receptors in herpes simplex virus 1 entry and cell-to-cell spread. J Virol, 2013. 87(3): p. 1430-42. [CrossRef]
- Veits, J., T.C. Mettenleiter, and W. Fuchs, Five unique open reading frames of infectious laryngotracheitis virus are expressed during infection but are dispensable for virus replication in cell culture. J Gen Virol, 2003. 84(Pt 6): p. 1415-1425. [CrossRef]
- Davison, A.J., Herpesvirus systematics. Vet Microbiol, 2010. 143(1): p. 52-69. [CrossRef]
- Li, C., et al., The Roles of Envelope Glycoprotein M in the Life Cycle of Some Alphaherpesviruses. Front Microbiol, 2021. 12: p. 631523. [CrossRef]
- García, M. and S. Spatz, Infectious Laryngotracheitis, in Diseases of Poultry. 2020. p. 189-209.
- Ziemann, K., T.C. Mettenleiter, and W. Fuchs, Gene arrangement within the unique long genome region of infectious laryngotracheitis virus is distinct from that of other alphaherpesviruses. J Virol, 1998. 72(1): p. 847-52. [CrossRef]
- Igarashi, K., et al., Construction and properties of a recombinant herpes simplex virus 1 lacking both S-component origins of DNA synthesis. J Virol, 1993. 67(4): p. 2123-32. [CrossRef]
- Polvino-Bodnar, M., P.K. Orberg, and P.A. Schaffer, Herpes simplex virus type 1 oriL is not required for virus replication or for the establishment and reactivation of latent infection in mice. J Virol, 1987. 61(11): p. 3528-35. [CrossRef]
- Przech, A.J., D. Yu, and S.K. Weller, Point mutations in exon I of the herpes simplex virus putative terminase subunit, UL15, indicate that the most conserved residues are essential for cleavage and packaging. J Virol, 2003. 77(17): p. 9613-21. [CrossRef]
- Higgs, M.R., V.G. Preston, and N.D. Stow, The UL15 protein of herpes simplex virus type 1 is necessary for the localization of the UL28 and UL33 proteins to viral DNA replication centres. J Gen Virol, 2008. 89(Pt 7): p. 1709-1715. [CrossRef]

| Isolate Name | Accession No. | Genome length (bp) | UL (bp) | IR (bp) | US (bp) | TR (bp) | G+C content |
|---|---|---|---|---|---|---|---|
| ILTV-B1/LIU/NC/2024 | PX492157 | 152,975 | 114,179 | 12,835 | 13,126 | 12,835 | 48.1% |
| ILTV-B3/LIU/NC/2024 | PX496590 | 152,978 | 114,182 | 12,835 | 13,126 | 12,835 | 48.1% |
| ILTV-B4/LIU/NC/2024 | PX522223 | 152,978 | 114,182 | 12,835 | 13,126 | 12,835 | 48.1% |
| Gene | Protein | Position | Amino Acid Substitution |
|---|---|---|---|
| UL22 | Envelope glycoprotein H | 1202 | H → R/H |
| 1696 | I → V | ||
| 1703 | L → S | ||
| 1813 | N → H | ||
| UL23 | Thymidine kinase | 755 | T → M |
| UL27 | Envelope glycoprotein B | 131 | R → H |
| 1651 | M → V/M | ||
| 1931 | I → T | ||
| 2395 | P → P/S | ||
| 2414 | K → R/K | ||
| UL1 | Envelope glycoprotein L | 377 | P → Q |
| UL53 | Envelope glycoprotein K | 646 | V → I |
| UL49.5 | Envelope glycoprotein N | 46 | V → M |
| 118 | Y → H | ||
| US4 | Envelope glycoprotein G | 133 | I → L |
| 173 | V → G | ||
| 292 | H → N | ||
| 344 | V → G | ||
| 353 | A → V | ||
| 387 | Q → H | ||
| 850 | F → L | ||
| 872 | Q → R | ||
| US5 | Envelope glycoprotein J | 23 | R → H |
| 903 | M → I | ||
| 1276 | T → A | ||
| 1481 | A → V | ||
| US6 | Envelope glycoprotein D | 61 | K → E |
| 580 | V → L | ||
| 592 | P → S | ||
| 644 | T → N | ||
| 752 | Q → R | ||
| US7 | Envelope glycoprotein I | 115 | I → V |
| 533 | A → D | ||
| 555 | L → F | ||
| 631 | A → T | ||
| 670 | V → I | ||
| US8 | Envelope glycoprotein E | 629 | K → R |
| ICP4 | Transcriptional regulator | 815 | A → T |
| 2240 | N → D | ||
| 2523 | L → P | ||
| 2531 | S → P | ||
| 3426 | V → G | ||
| 3909 | L → P | ||
| 4302 | P → L |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





