Submitted:
29 January 2026
Posted:
30 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Oligopaint Probe Synthesis
2.2. Oligopaint Probe Design for Spatial Resolution Calibration in Multi-Color FISH
2.3. Construction and Oligopaint Targeting of Synthetic Reporter Sequences
2.4. Transgene Design for Defined 3D Genomic Distance Measurements
2.5. Oligopaint Target Length Variants for Assessing Spatial Resolution
2.6. Modulation of Oligopaint Probe Density
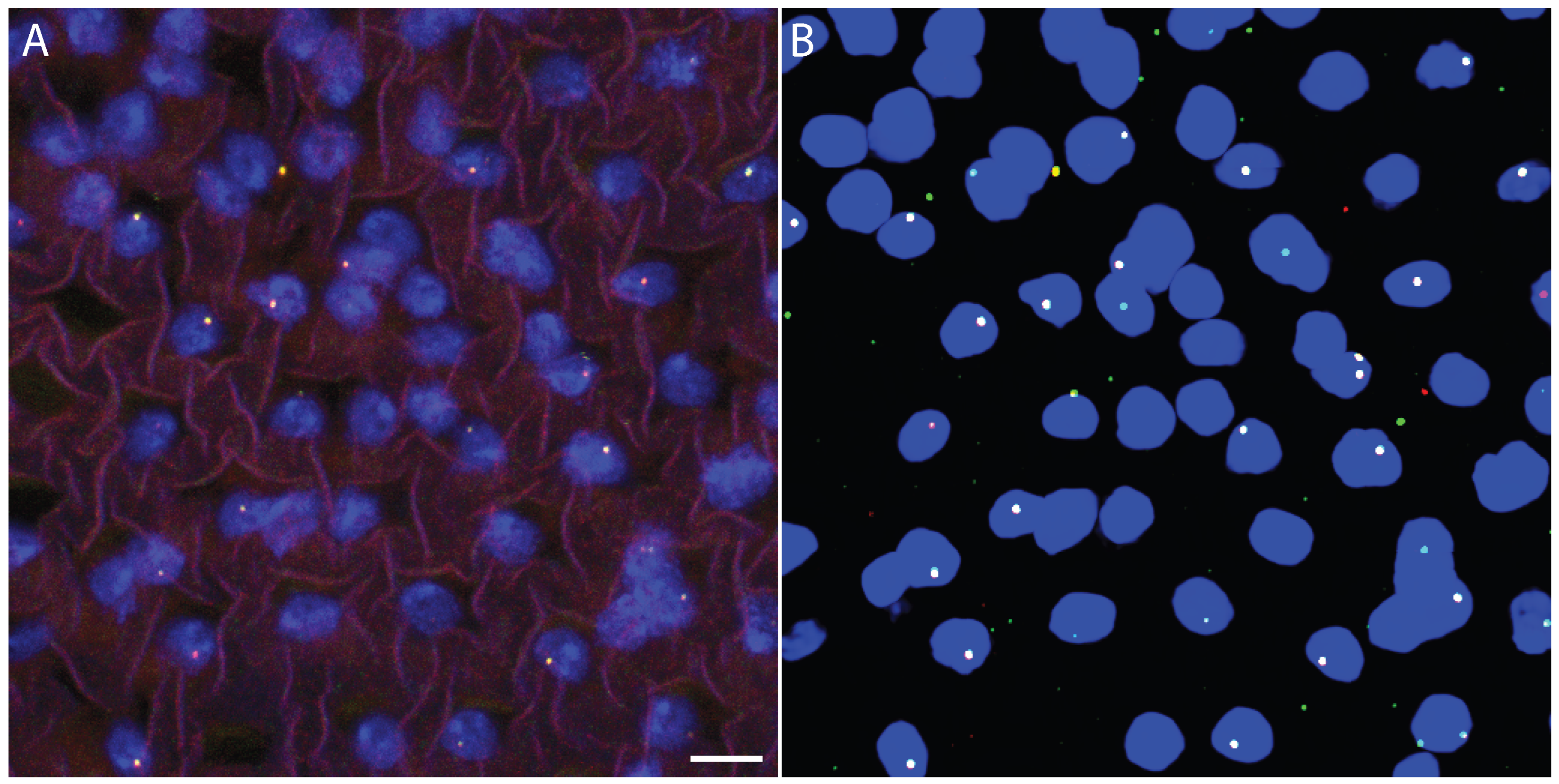
2.7. Fluorescence In Situ Hybridization on late Pupal Wings
2.8. Nanorulers Calibration Standards
2.9. Confocal Imaging
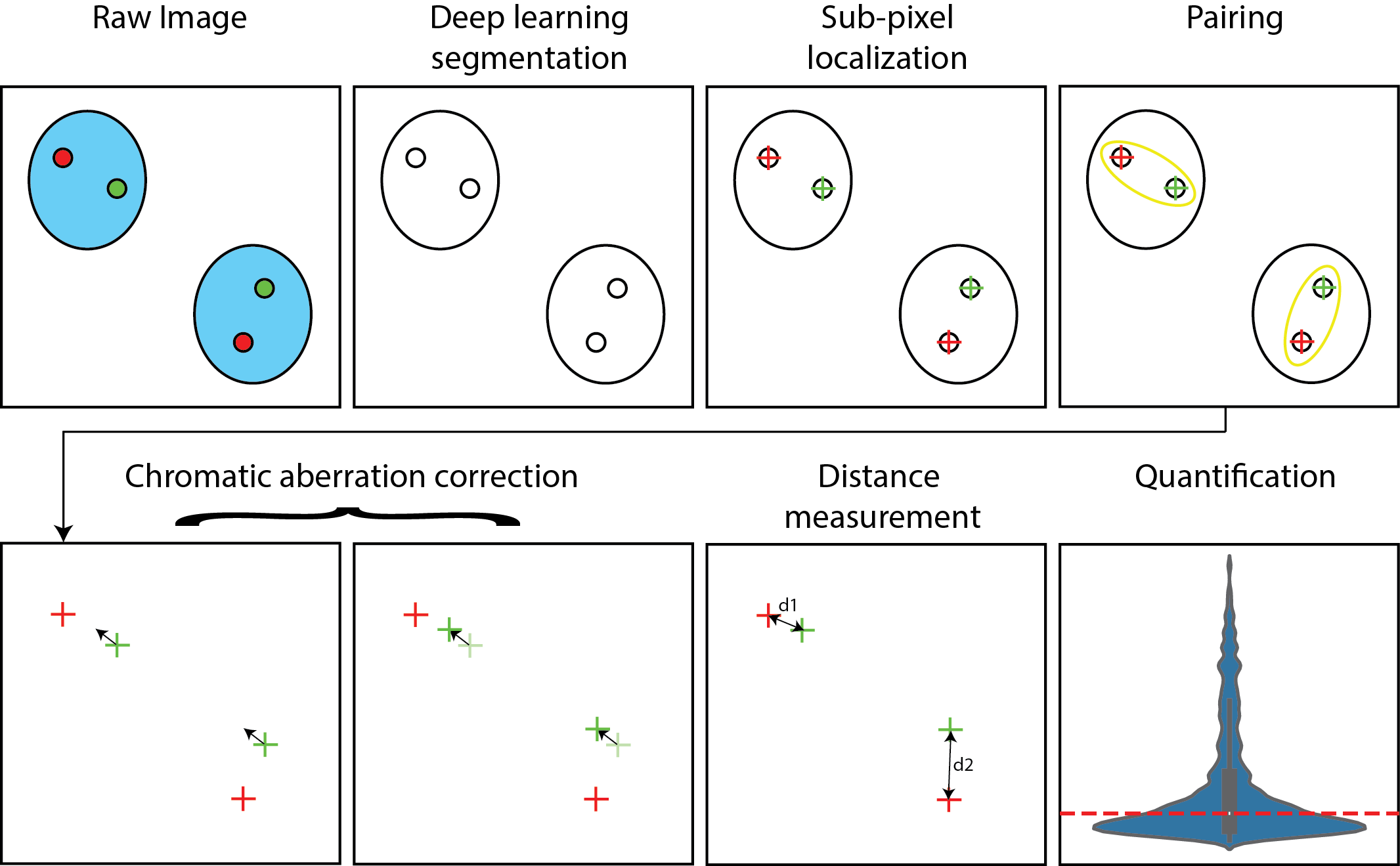
2.10. Deep Learning–Based Detection of Nuclei and Spots
2.11. Subpixel 3D Spot Centroid Extraction via Gaussian Fitting
2.12. Spot Pairing Across Channels
2.13. Linear Chromatic Correction (LCC)
2.14. Affine Chromatic Correction (ACC)
2.15. Distance Measurements and Resolution Assessment
3. Results
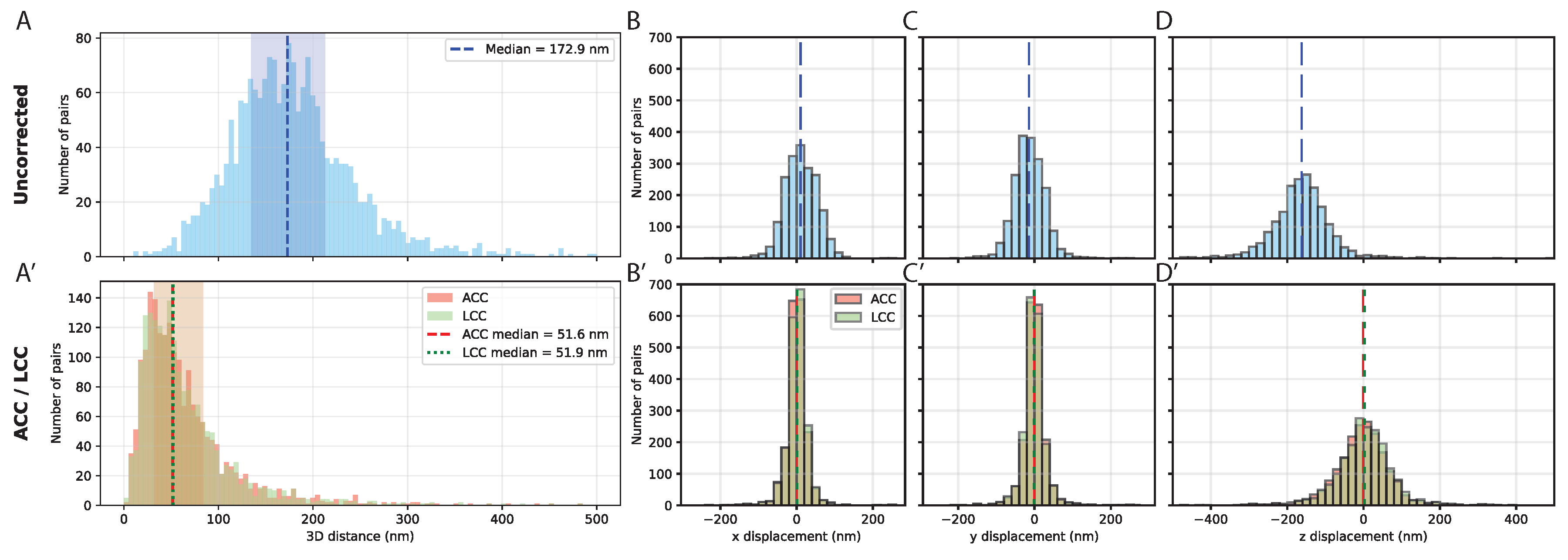
3.1. Quantifying and Correcting Chromatic Aberration to Establish the Baseline Resolution
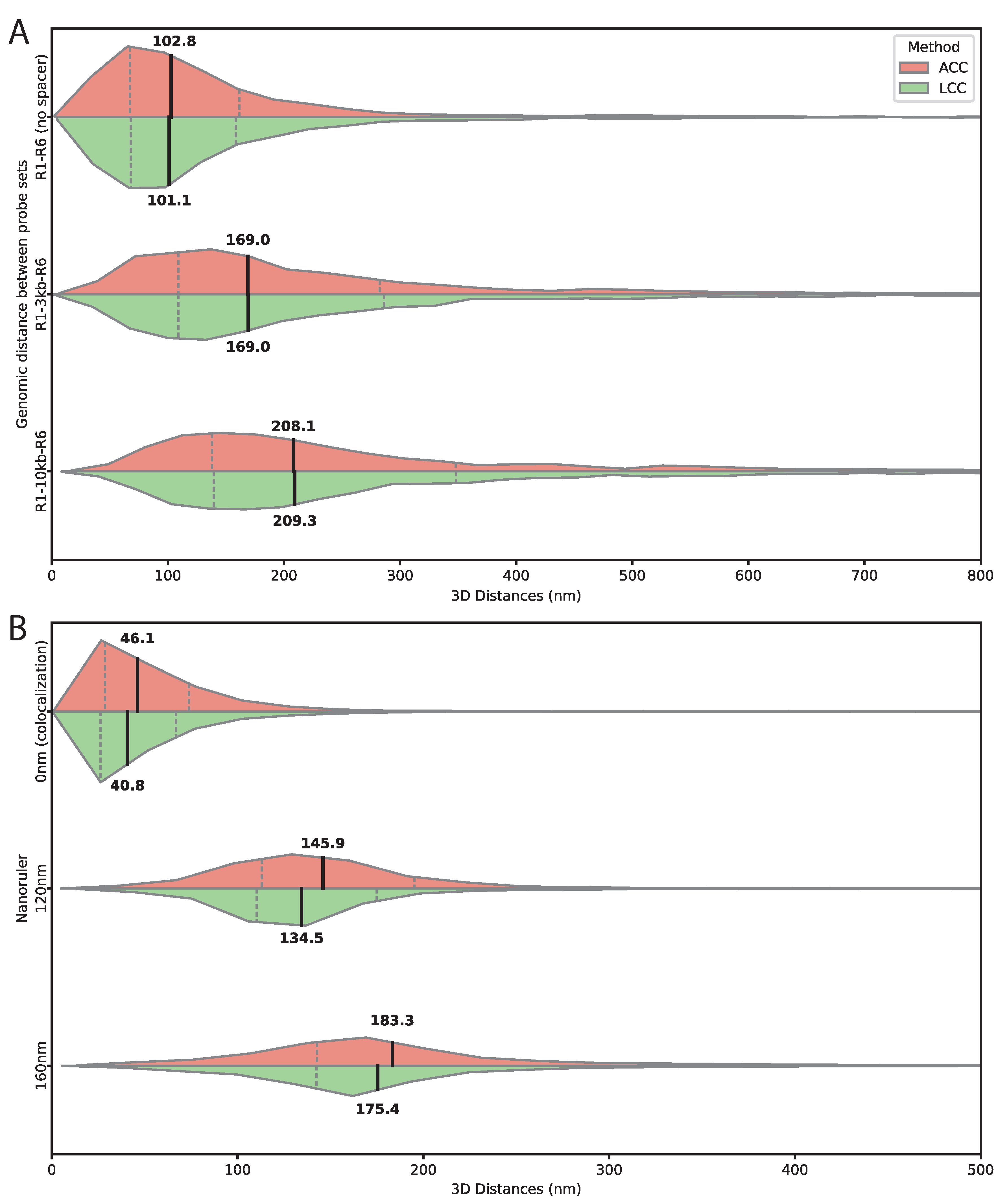
3.2. Validation of 3D Distance Measurements Using Genomic Standards and DNA Origami Nanorulers
3.3. Target Sequence Length And Composition Affect Spatial Resolution
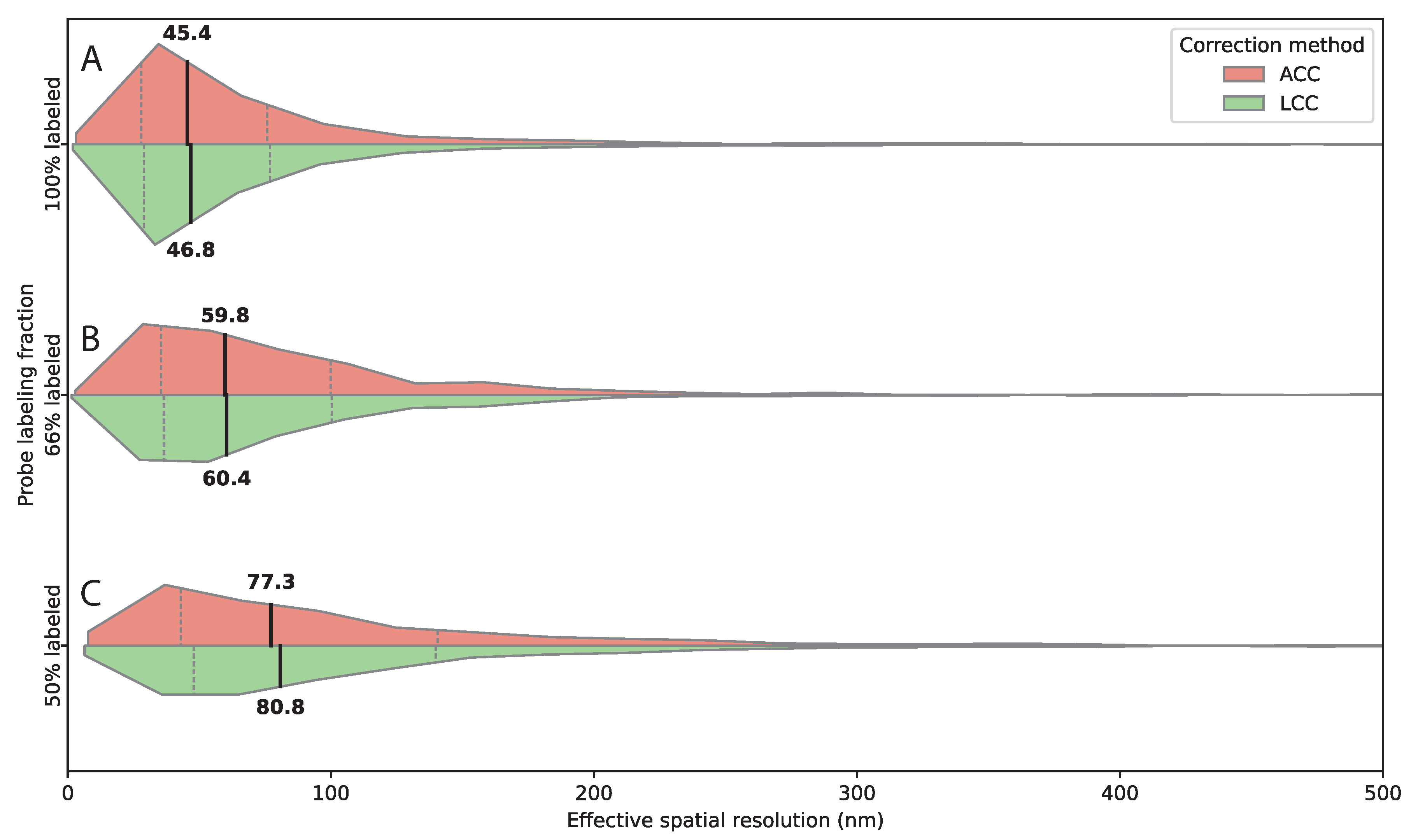
3.4. Probe Density Influences Detection Efficiency and Spatial Resolution
4. Discussion
4.1. FISH-Dist Enables Accurate Short-Range Genomic Distance Measurements
4.2. Probe Design Guidelines Emerge from Systematic Parameter Testing
4.3. Methodological Considerations and Limitations
4.4. Comparison with Existing Methods
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Popay, T.M.; Dixon, J.R. Coming full circle: On the origin and evolution of the looping model for enhancer–promoter communication. 298. Elsevier: Publisher. [CrossRef]
- Kim, S.; Shendure, J. Mechanisms of Interplay between Transcription Factors and the 3D Genome. 76, 306–319. [CrossRef]
- Beliveau, B.J.; Boettiger, A.N.; Avendaño, M.S.; Jungmann, R.; McCole, R.B.; Joyce, E.F.; Kim-Kiselak, C.; Bantignies, F.; Fonseka, C.Y.; Erceg, J.; et al. Single-molecule super-resolution imaging of chromosomes and in situ haplotype visualization using Oligopaint FISH probes. 6, 7147. [CrossRef] [PubMed]
- Bolzer, A.; Kreth, G.; Solovei, I.; Koehler, D.; Saracoglu, K.; Fauth, C.; Müller, S.; Eils, R.; Cremer, C.; Speicher, M.R.; et al. Three-Dimensional Maps of All Chromosomes in Human Male Fibroblast Nuclei and Prometaphase Rosettes. 3, e157. [CrossRef] [PubMed]
- Fields, B.D.; Nguyen, S.C.; Nir, G.; Kennedy, S. A multiplexed DNA FISH strategy for assessing genome architecture in Caenorhabditis elegans. 8, e42823. [CrossRef] [PubMed]
- Shirley, J.W.; Ty, S.; Takebayashi, S.i.; Liu, X.; Gilbert, D.M. FISH Finder: a high-throughput tool for analyzing FISH images. 27, 933–938. [CrossRef]
- Lieberman-Aiden, E.; van Berkum, N.L.; Williams, L.; Imakaev, M.; Ragoczy, T.; Telling, A.; Amit, I.; Lajoie, B.R.; Sabo, P.J.; Dorschner, M.O.; et al. Comprehensive Mapping of Long-Range Interactions Reveals Folding Principles of the Human Genome. 326, 289–293. [CrossRef]
- Mateo, L.J.; Murphy, S.E.; Hafner, A.; Cinquini, I.S.; Walker, C.A.; Boettiger, A.N. Visualizing DNA folding and RNA in embryos at single-cell resolution. 568, 49–54. [CrossRef]
- Harke, J.; Lee, J.R.; Nguyen, S.C.; Arab, A.; Rakowiecki, S.M.; Hugelier, S.; Paliou, C.; Rauseo, A.; Yunker, R.; Xu, K.; et al. Multiple allelic configurations govern long-range Shh enhancer-promoter communication in the embryonic forebrain. 84, 4698–4710.e6. [CrossRef]
- Gizzi, A.M.C.; Cattoni, D.I.; Fiche, J.B.; Espinola, S.M.; Gurgo, J.; Messina, O.; Houbron, C.; Ogiyama, Y.; Papadopoulos, G.L.; Cavalli, G.; et al. Microscopy-Based Chromosome Conformation Capture Enables Simultaneous Visualization of Genome Organization and Transcription in Intact Organisms. 74, 212–222.e5. [CrossRef]
- Barinov, L.; Ryabichko, S.; Bialek, W.; Gregor, T. Transcription-dependent spatial organization of a gene locus. 2012.15819.
- Urban, E.A.; Chernoff, C.; Layng, K.V.; Han, J.; Anderson, C.; Konzman, D.; Johnston, R.J. Activating and repressing gene expression between chromosomes during stochastic fate specification. 42, 111910. [CrossRef]
- Galouzis, C.C.; Prud’homme, B. Transvection regulates the sex-biased expression of a fly X-linked gene. 371, 396–400. [CrossRef]
- Delker, R.K.; Munce, R.H.; Hu, M.; Mann, R.S. Fluorescent labeling of genomic loci in Drosophila imaginal discs with heterologous DNA-binding proteins. [CrossRef]
- Viets, K.; Sauria, M.E.G.; Chernoff, C.; Viales, R.R.; Echterling, M.; Anderson, C.; Tran, S.; Dove, A.; Goyal, R.; Voortman, L.; et al. Characterization of Button Loci that Promote Homologous Chromosome Pairing and Cell-Type-Specific Interchromosomal Gene Regulation. 51, 341–356.e7. [CrossRef]
- Arbor, A. Labeling Protocol for myTags Immortal Libraries.
- Aigouy, B.; Cortes, C.; Liu, S.; Prud’Homme, B. EPySeg: a coding-free solution for automated segmentation of epithelia using deep learning. 147. [CrossRef]
- Imbert, A.; Ouyang, W.; Safieddine, A.; Coleno, E.; Zimmer, C.; Bertrand, E.; Walter, T.; Mueller, F. FISH-quant v2: a scalable and modular tool for smFISH image analysis. 28, 786–795. [CrossRef]
- Brückner, D.B.; Chen, H.; Barinov, L.; Zoller, B.; Gregor, T. Stochastic motion and transcriptional dynamics of pairs of distal DNA loci on a compacted chromosome. 380 1357–1362. [CrossRef] [PubMed]
- Leiwe, M.N.; Fujimoto, S.; Imai, T. Post hoc Correction of Chromatic Aberrations in Large-Scale Volumetric Images in Confocal Microscopy. 15. [CrossRef] [PubMed]
- Waskom, M.L. seaborn: statistical data visualization. Journal of Open Source Software 2021, 6, 3021. [Google Scholar] [CrossRef]
- Hunter, J.D. Matplotlib: A 2D graphics environment. Computing in Science & Engineering 2007, 9, 90–95. [Google Scholar] [CrossRef]
- Ershov, D.; Phan, M.S.; Pylvänäinen, J.W.; Rigaud, S.U.; Le Blanc, L.; Charles-Orszag, A.; Conway, J.R.W.; Laine, R.F.; Roy, N.H.; Bonazzi, D.; et al. Bringing TrackMate into the era of machine-learning and deep-learning. [CrossRef]
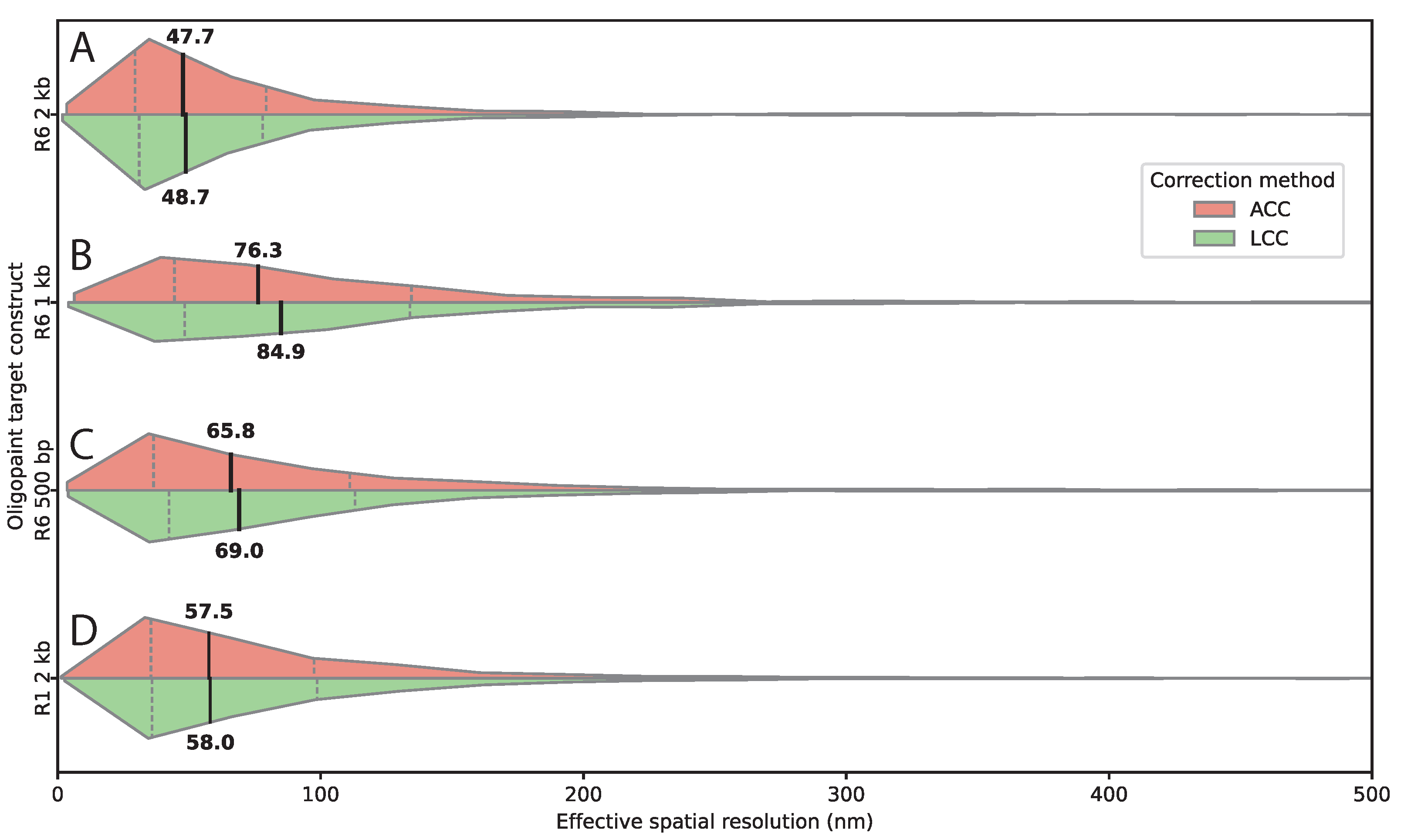
| Construct | Target size | Detected pairs per image |
|---|---|---|
| R6 | 2 kb | 190.5 |
| R6 | 1 kb | 57.4 |
| R6 | 500 bp | 34 |
| R1 | 2 kb | 128.8 |
| Labeled probe fraction |
Fluorescent / unlabeled oligonucleotides per locus |
Detected pairs per image |
|---|---|---|
| 100% | 87 / 0 | 190.5 |
| 66% | 57 / 30 | 133.2 |
| 50% | 43 / 43 | 51.1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




