Submitted:
28 January 2026
Posted:
29 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results and Discussion
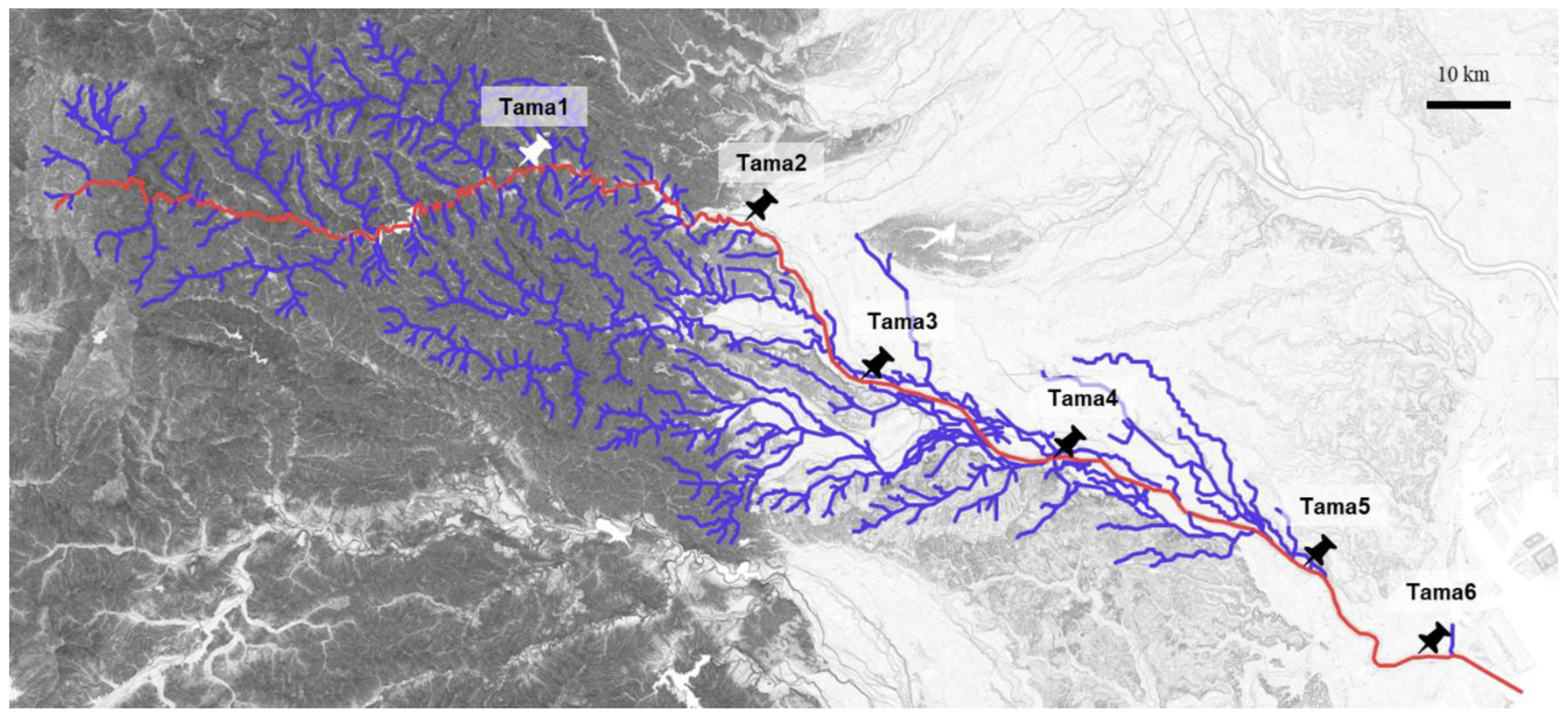
2.1. Various Plasmids Carrying ARGs Were Captured from the Tama River
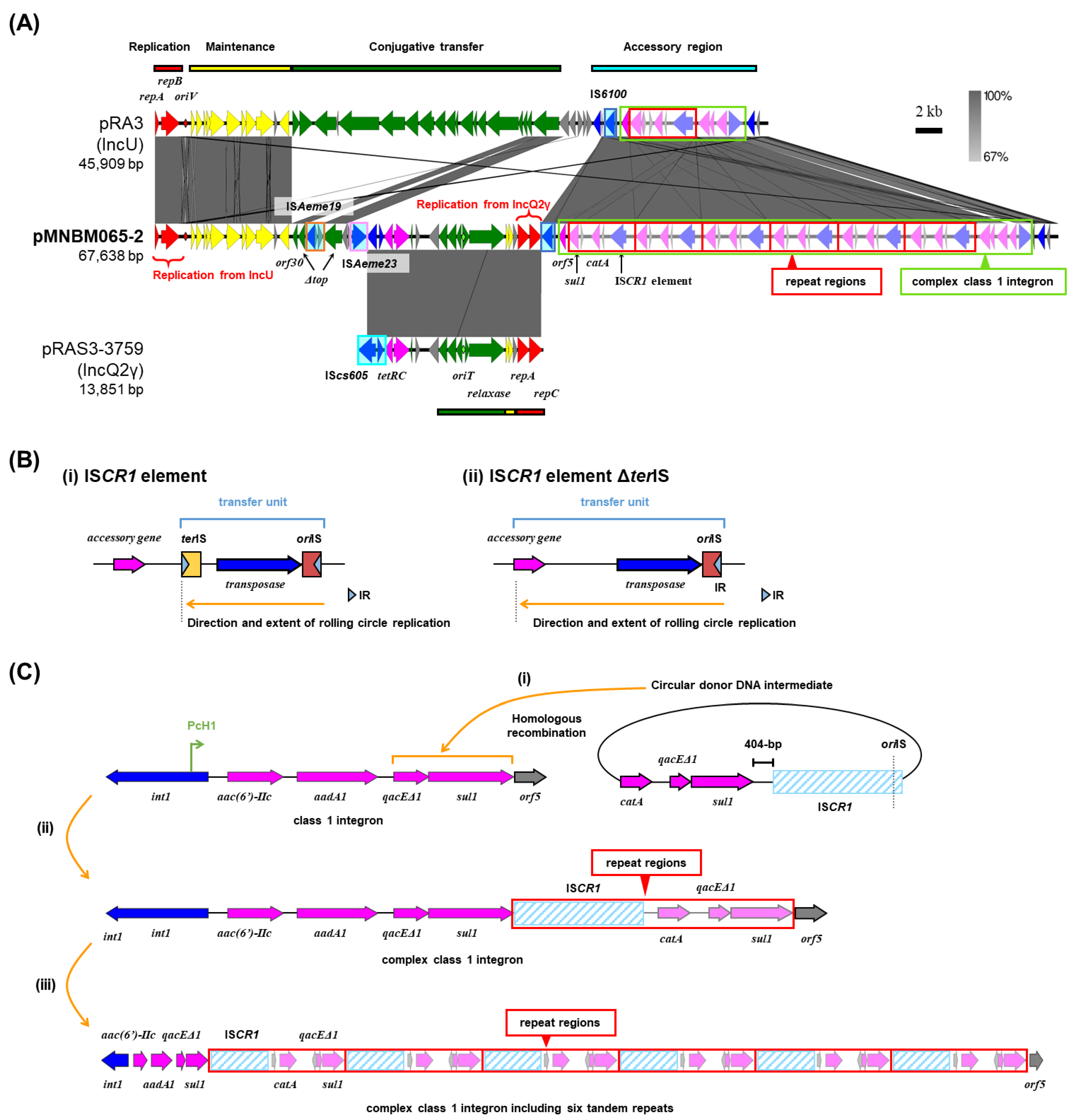
2.2. Multi-Replicon Plasmid pMNBM065-2
2.3. Mobilization of pMNBM065-2
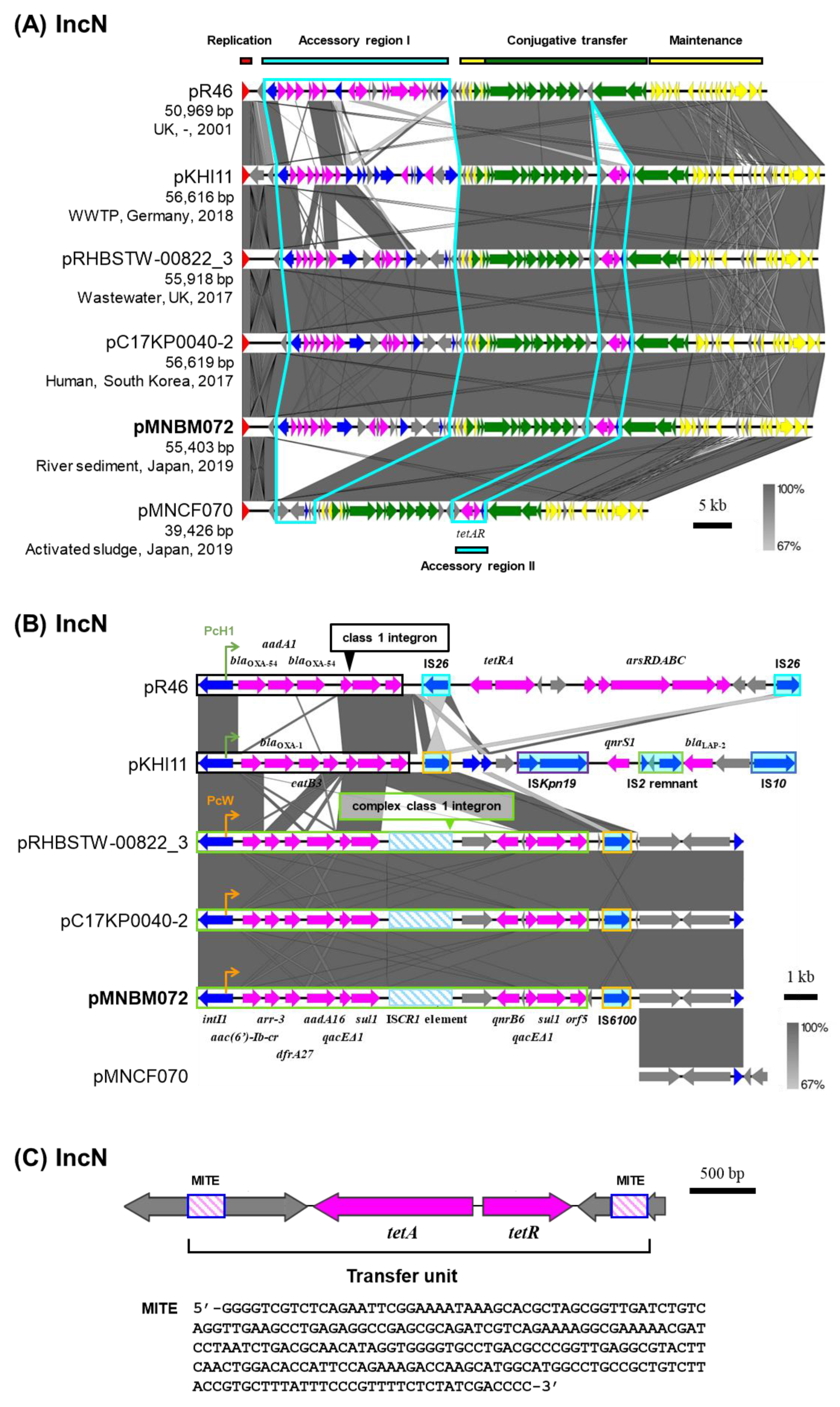
2.4. IncN Plasmid pMNBM072
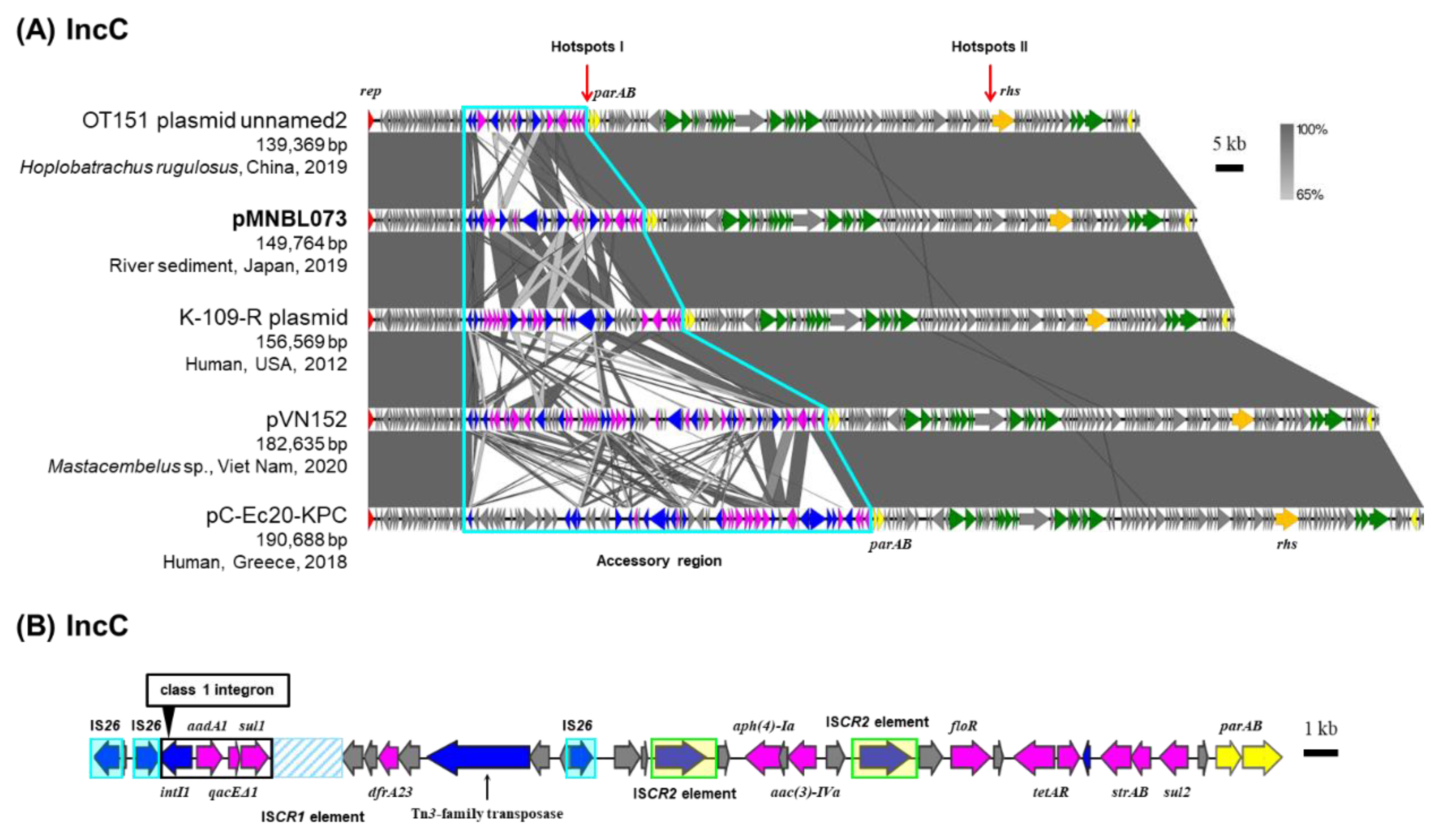
2.5. IncC Plasmid pMNBL073
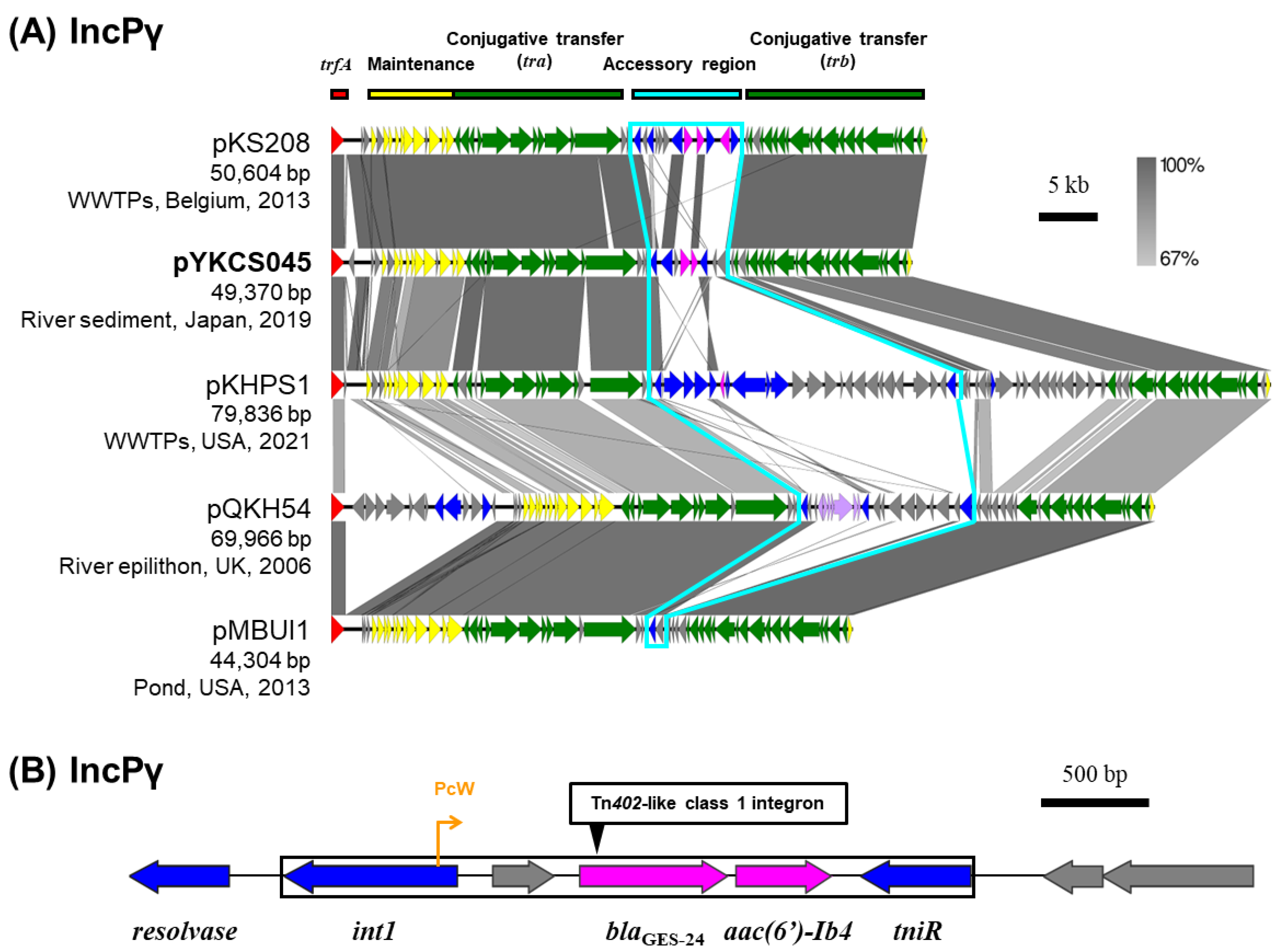
2.6. IncP Plasmids
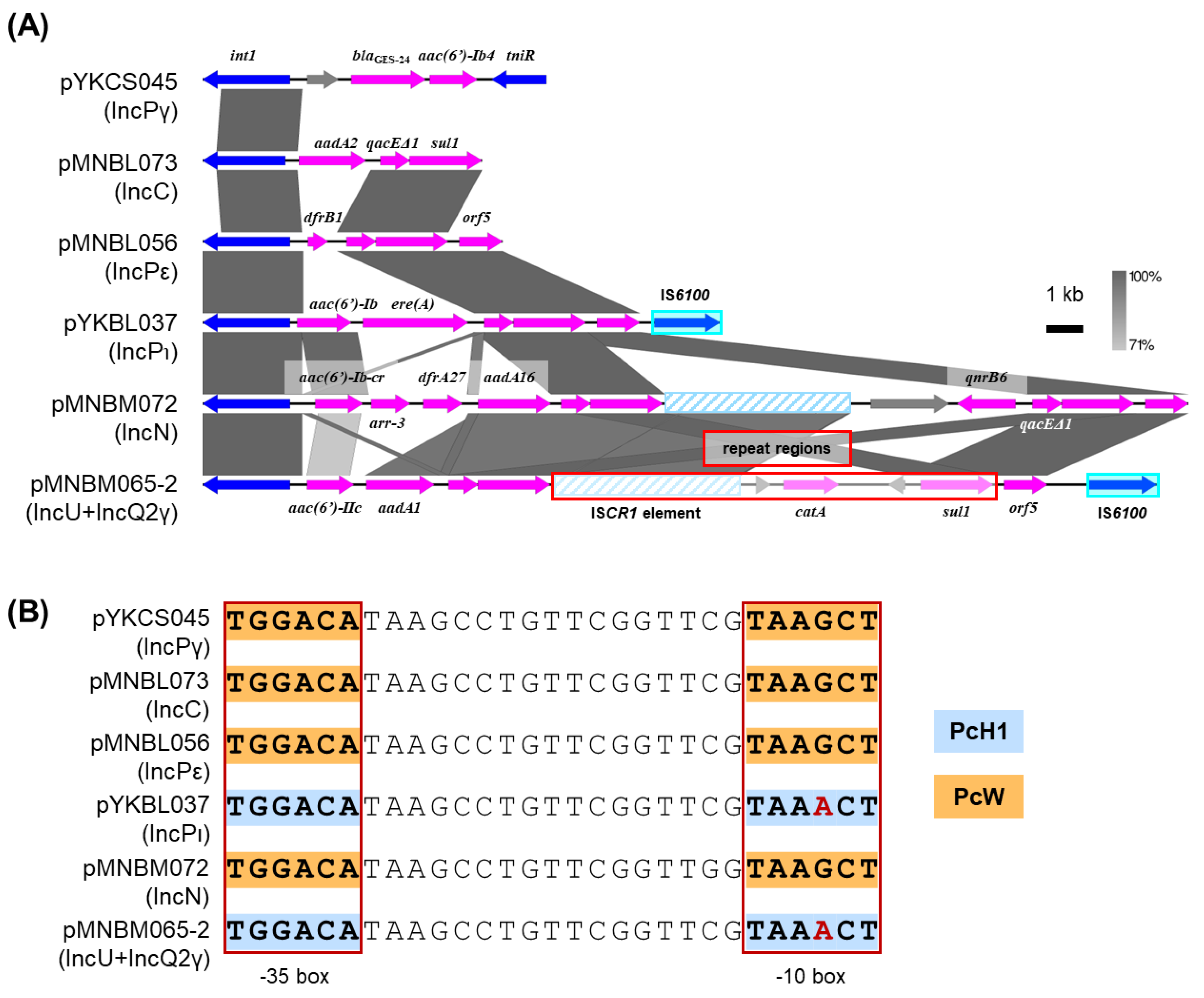
2.7. Class 1 Integrons
3. Materials and Methods
3.1. Bacterial Strains, Plasmids, and Culture Conditions
3.2. Collection of Environmental Samples
3.3. Resistance Testing
3.4. Plasmid Sequencing and Annotation
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ARG | Antimicrobial resistance gene |
| Cm | Chloramphenicol |
| CS | Conserved segment |
| DDBJ | DNA Data Bank of Japan |
| ENA | European Nucleotide Archive |
| GFP | Green fluorescent protein |
| IS | Insertion sequence |
| ISCR | Insertion sequence common region |
| kb | Kilobase(s) |
| MGE | Mobile genetic element |
| MIC | Minimum inhibitory concentration |
| oriIS | Origin of insertion sequence replication |
| PCR | Polymerase chain reaction |
| PLSDB | Plasmid database |
| Rep | Replication protein |
| RND | Resistance–nodulation–cell division |
| Tn | Transposon |
Appendix A
| Sample | Recipients | No. of isolates | IncP | PromA | pSN1216-29 | IncA or IncC | IncL or IncM | IncN | IncW | PCR-negative |
| Tama1 | M. resinovorans | 20 | 0 | 4 | - | - | - | - | - | 16 |
| E. coli | 0 | - | - | - | - | - | - | - | - | |
| Tama2 | M. resinovorans | 20 | 0 | 1 | - | - | - | - | - | 19 |
| E. coli | 0 | - | - | - | - | - | - | - | - | |
| Tama3 | M. resinovorans | 9 | 0 | 0 | - | - | - | - | - | 9 |
| E. coli | 0 | - | - | - | - | - | - | - | - | |
| Tama4 | M. resinovorans | 20 | 0 | 0 | 12 | - | - | - | - | 8 |
| E. coli | 4 | 0 | 0 | 2 | 0 | 0 | 0 | 1 | ||
| Tama5 | M. resinovorans | 40 | 1 | 6 | - | - | - | - | - | 33 |
| E. coli | 17 | 7 | 1 | 0 | 0 | 0 | 2 | 0 | 8 | |
| Tama6 | M. resinovorans | 36 | 3 | 0 | - | - | - | - | - | 34 |
| E. coli | 1 | 1 | 0 | 0 | 0 | 0 | 0 | - | 0 | |
| total | 167 | 16 | 24 | 12 | 2 | 0 | 2 | 0 | 127 | |
References
- Castañeda-Barba, S.; Top, E.M.; Stalder, T. Plasmids, a Molecular Cornerstone of Antimicrobial Resistance in the One Health Era. Nat. Rev. Microbiol. 2024, 22, 18–32. [Google Scholar] [CrossRef]
- Flach, C.-F.; Johnning, A.; Nilsson, I.; Smalla, K.; Kristiansson, E.; Larsson, D.G.J. Isolation of Novel IncA/C and IncN Fluoroquinolone Resistance Plasmids from an Antibiotic-Polluted Lake. J. Antimicrob. Chemother. 2015, 70, 2709–2717. [Google Scholar] [CrossRef]
- Yu, Z.; Wang, Q.; Pinilla-Redondo, R.; Madsen, J.S.; Clasen, K.A.D.; Ananbeh, H.; Olesen, A.K.; Gong, Z.; Yang, N.; Dechesne, A.; et al. Horizontal Transmission of a Multidrug-Resistant IncN Plasmid Isolated from Urban Wastewater. Ecotoxicol. Environ. Saf. 2024, 271, 115971. [Google Scholar] [CrossRef]
- Nguyen, T.N.; Kasuga, I.; Liu, M.; Katayama, H. Occurrence of Antibiotic Resistance Genes as Emerging Contaminants in Watersheds of Tama River and Lake Kasumigaura in Japan. IOP Conf. Ser. Earth Environ. Sci. 2019, 266, 012003. [Google Scholar] [CrossRef]
- Shintani, M.; Vestergaard, G.; Milaković, M.; Kublik, S.; Smalla, K.; Schloter, M.; Udiković-Kolić, N. Integrons, Transposons and IS Elements Promote Diversification of Multidrug Resistance Plasmids and Adaptation of Their Hosts to Antibiotic Pollutants from Pharmaceutical Companies. Environ. Microbiol. 2023, 25, 3035–3051. [Google Scholar] [CrossRef]
- Hauschild, K.; Suzuki, M.; Wolters, B.; Tokuda, M.; Yamazaki, R.; Masumoto, M.; Moriuchi, R.; Dohra, H.; Bunk, B.; Spröer, C.; et al. The Transferable Resistome of Biosolids-Plasmid Sequencing Reveals Carriage of Clinically Relevant Antibiotic Resistance Genes. mBio 2025, 16, e0206825. [Google Scholar] [CrossRef]
- Wolters, B.; Hauschild, K.; Blau, K.; Mulder, I.; Heyde, B.J.; Sørensen, S.J.; Siemens, J.; Jechalke, S.; Smalla, K.; Nesme, J. Biosolids for Safe Land Application: Does Wastewater Treatment Plant Size Matters When Considering Antibiotics, Pollutants, Microbiome, Mobile Genetic Elements and Associated Resistance Genes? Environ. Microbiol. 2022, 24, 1573–1589. [Google Scholar] [CrossRef] [PubMed]
- Blau, K.; Bettermann, A.; Jechalke, S.; Fornefeld, E.; Vanrobaeys, Y.; Stalder, T.; Top, E.M.; Smalla, K. The Transferable Resistome of Produce. mBio 2018, 9, e01300-18. [Google Scholar] [CrossRef] [PubMed]
- Hölzel, C.S.; Tetens, J.L.; Schwaiger, K. Unraveling the Role of Vegetables in Spreading Antimicrobial-Resistant Bacteria: A Need for Quantitative Risk Assessment. Foodborne Pathog. Dis. 2018, 15, 671–688. [Google Scholar] [CrossRef] [PubMed]
- Hayakawa, M.; Tokuda, M.; Kaneko, K.; Nakamichi, K.; Yamamoto, Y.; Kamijo, T.; Umeki, H.; Chiba, R.; Yamada, R.; Mori, M.; et al. Hitherto-Unnoticed Self-Transmissible Plasmids Widely Distributed among Different Environments in Japan. Appl. Environ. Microbiol. 2022, 88, e0111422. [Google Scholar] [CrossRef] [PubMed]
- Szuplewska, M.; Ludwiczak, M.; Lyzwa, K.; Czarnecki, J.; Bartosik, D. Mobility and Generation of Mosaic Non-Autonomous Transposons by Tn3-Derived Inverted-Repeat Miniature Elements (TIMEs). PLoS One 2014, 9, e105010. [Google Scholar] [CrossRef]
- Kulinska, A.; Czeredys, M.; Hayes, F.; Jagura-Burdzy, G. Genomic and Functional Characterization of the Modular Broad-Host-Range RA3 Plasmid, the Archetype of the IncU Group. Appl. Environ. Microbiol. 2008, 74, 4119–4132. [Google Scholar] [CrossRef]
- Lewicka, E.; Dolowy, P.; Godziszewska, J.; Litwin, E.; Ludwiczak, M.; Jagura-Burdzy, G. Transcriptional Organization of the Stability Module of Broad-Host-Range Plasmid RA3, from the IncU Group. Appl. Environ. Microbiol. 2020, 86, e00847-20. [Google Scholar] [CrossRef]
- L’Abée-Lund, T.M.; Sørum, H. A Global Non-Conjugative Tet C Plasmid, pRAS3, from Aeromonas salmonicida. Plasmid 2002, 47, 172–181. [Google Scholar] [CrossRef]
- Loftie-Eaton, W.; Rawlings, D.E. Comparative Biology of Two Natural Variants of the IncQ-2 Family Plasmids, pRAS3.1 and pRAS3.2. J. Bacteriol. 2009, 191, 6436–6446. [Google Scholar] [CrossRef] [PubMed]
- Fournier, K.C.; Paquet, V.E.; Attéré, S.A.; Farley, J.; Marquis, H.; Gantelet, H.; Ravaille, C.; Vincent, A.T.; Charette, S.J. Expansion of the pRAS3 Plasmid Family in Aeromonas salmonicida Subsp. salmonicida and Growing Evidence of Interspecies Connections for These Plasmids. Antibiotics (Basel) 2022, 11, 1047. [Google Scholar] [CrossRef] [PubMed]
- Mendiola, M.V.; de la Cruz, F. IS91 Transposase Is Related to the Rolling-Circle-Type Replication Proteins of the pUB110 Family of Plasmids. Nucleic Acids Res. 1992, 20, 3521. [Google Scholar] [CrossRef]
- Mendiola, M.V.; Bernales, I.; de la Cruz, F. Differential Roles of the Transposon Termini in IS91 Transposition. Proc. Natl. Acad. Sci. U. S. A. 1994, 91, 1922–1926. [Google Scholar] [CrossRef]
- Tavakoli, N.; Comanducci, A.; Dodd, H.M.; Lett, M.C.; Albiger, B.; Bennett, P. IS1294, a DNA Element That Transposes by RC Transposition. Plasmid 2000, 44, 66–84. [Google Scholar] [CrossRef]
- Garcillán-Barcia, M.P.; De La Cruz, F.; Bernales, I.; Mendiola, M.V. IS91 Rolling-Circle Transposition. In Mobile DNA II; American Society of Microbiology, 2002; pp. 891–904. ISBN 9781555812096. [Google Scholar]
- Toleman, M.A.; Walsh, T.R. Combinatorial Events of Insertion Sequences and ICE in Gram-Negative Bacteria. FEMS Microbiol. Rev. 2011, 35, 912–935. [Google Scholar] [CrossRef] [PubMed]
- TnPedia Team TnPedia: IS91 and ISCR Families of Prokaryotic Insertion Sequences . Zenodo 2025. [CrossRef]
- Andersen, J.B.; Sternberg, C.; Poulsen, L.K.; Bjorn, S.P.; Givskov, M.; Molin, S. New Unstable Variants of Green Fluorescent Protein for Studies of Transient Gene Expression in Bacteria. Appl. Environ. Microbiol. 1998, 64, 2240–2246. [Google Scholar] [CrossRef] [PubMed]
- Tokuda, M.; Yuki, M.; Ohkuma, M.; Kimbara, K.; Suzuki, H.; Shintani, M. Transconjugant Range of PromA Plasmids in Microbial Communities Is Predicted by Sequence Similarity with the Bacterial Host Chromosome. Microb. Genom. 2023, 9, mgen.0.001043. [Google Scholar] [CrossRef]
- Eikmeyer, F.; Hadiati, A.; Szczepanowski, R.; Wibberg, D.; Schneiker-Bekel, S.; Rogers, L.M.; Brown, C.J.; Top, E.M.; Pühler, A.; Schlüter, A. The Complete Genome Sequences of Four New IncN Plasmids from Wastewater Treatment Plant Effluent Provide New Insights into IncN Plasmid Diversity and Evolution. Plasmid 2012, 68, 13–24. [Google Scholar] [CrossRef]
- Gomi, R.; Yano, H. Miniature Inverted-Repeat Transposable Elements Mobilize Diverse Antibiotic Resistance Genes in Enterobacteriaceae. bioRxiv 2025. [Google Scholar] [CrossRef]
- Zhang, F.; Ye, X.; Yin, Z.; Hu, M.; Wang, B.; Liu, W.; Li, B.; Ren, H.; Jin, Y.; Yue, J. Comparative Genomics Reveals New Insights into the Evolution of the IncA and IncC Family of Plasmids. Front. Microbiol. 2022, 13, 1045314. [Google Scholar] [CrossRef]
- Galata, V.; Fehlmann, T.; Backes, C.; Keller, A. PLSDB: A Resource of Complete Bacterial Plasmids. Nucleic Acids Res. 2018, 47, D195–D202. [Google Scholar] [CrossRef]
- Schmartz, G.P.; Hartung, A.; Hirsch, P.; Kern, F.; Fehlmann, T.; Müller, R.; Keller, A. PLSDB: Advancing a Comprehensive Database of Bacterial Plasmids. Nucleic Acids Res. 2022, 50, D273–D278. [Google Scholar] [CrossRef]
- Molano, L.-A.G.; Hirsch, P.; Hannig, M.; Müller, R.; Keller, A. The PLSDB 2025 Update: Enhanced Annotations and Improved Functionality for Comprehensive Plasmid Research. Nucleic Acids Res. 2025, 53, D189–D196. [Google Scholar] [CrossRef] [PubMed]
- Haines, A.S.; Akhtar, P.; Stephens, E.R.; Jones, K.; Thomas, C.M.; Perkins, C.D.; Williams, J.R.; Day, M.J.; Fry, J.C. Plasmids from Freshwater Environments Capable of IncQ Retrotransfer Are Diverse and Include pQKH54, a New IncP-1 Subgroup Archetype. Microbiology 2006, 152, 2689–2701. [Google Scholar] [CrossRef]
- Yano, H.; Rogers, L.M.; Knox, M.G.; Heuer, H.; Smalla, K.; Brown, C.J.; Top, E.M. Host Range Diversification within the IncP-1 Plasmid Group. Microbiology 2013, 159, 2303–2315. [Google Scholar] [CrossRef] [PubMed]
- Fonseca, É.L.; Vicente, A.C. Integron Functionality and Genome Innovation: An Update on the Subtle and Smart Strategy of Integrase and Gene Cassette Expression Regulation. Microorganisms 2022, 10, 224. [Google Scholar] [CrossRef]
- Jové, T.; Da Re, S.; Denis, F.; Mazel, D.; Ploy, M.-C. Inverse Correlation between Promoter Strength and Excision Activity in Class 1 Integrons. PLoS Genet. 2010, 6, e1000793. [Google Scholar] [CrossRef]
- Ghaly, T.M.; Chow, L.; Asher, A.J.; Waldron, L.S.; Gillings, M.R. Evolution of Class 1 Integrons: Mobilization and Dispersal via Food-Borne Bacteria. PLoS One 2017, 12, e0179169. [Google Scholar] [CrossRef]
- Sambrook, J.; Russell, D. Molecular Cloning: A Laboratory Manual, 3rd ed.; Cold Spring Harbor Laboratory: Cold Spring Harbor, NY, 2001; ISBN 9780879695774. [Google Scholar]
- Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria That Grow Aerobically; Approved Standard-12th Edition. CLSI Document M07; Wayne, PA, 2024.
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A New Genome Assembly Algorithm and Its Applications to Single-Cell Sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [PubMed]
- Tanizawa, Y.; Fujisawa, T.; Nakamura, Y. DFAST: A Flexible Prokaryotic Genome Annotation Pipeline for Faster Genome Publication. Bioinformatics 2018, 34, 1037–1039. [Google Scholar] [CrossRef] [PubMed]
- Sullivan, M.J.; Petty, N.K.; Beatson, S.A. Easyfig: A Genome Comparison Visualizer. Bioinformatics 2011, 27, 1009–1010. [Google Scholar] [CrossRef]
| Plasmid | Inc_group | Size (bp) | Source | Methods* | Accessory genes | Accession number |
| pMNBM065-2 | IncU, IncQ2γ | 67,638 | Tama5 | B |
tetRC, IS6100, class 1 integron [aac(6’)-IIc, aadA1, qacEΔ1- sul1]; ISCR1 element; catA, qacEΔ1- sul1 (repeat unit x6), orf5 |
LC895900.1 |
| pMNBM072 | IncN | 55,403 | Tama5 | B | Class 1 integron [aac(6’)-Ib-cr, arr-3, dfrA27, aadA16, qacEΔ1-sul1, ISCR1 element qnrB6, qacEΔ1-sul1-orf5], tetAR |
LC663726.1 |
| pMNBL073 | IncC | 149,764 | Tama4 | B | Class 1 integron (aadA2, qacEΔ1, sul1), dfrA23, aph(4)-Ia, aac(3)-IVa, floR, tetAR, strAB, sul2 |
LC663722.1 |
| pYKCS045 | IncPγ | 49,370 | Tama5 | T | Tn402-like class 1 integron [blaGES-24, aac(6’)-Ib4] | LC623929.1 |
| pMNBL056 | IncPε | 52,432 | Tama4 | B | Class 1 integron (dfrB1, qacEΔ1-sul1-orf5, tetRA) | LC623890.1 |
| pYKBL037 | IncPι | 64,506 | Tama4 | T | Transposon (strAB, class 1 integron [aac(6’)-Ib, ere(A), qacEΔ1-sul1-orf5], transposon (blaA, qac) |
LC623919.1 |
| pMNDW109 | IncPβ | 98,131 | Tama5 | B | Tn3-family transposon (mer operon, IS1071, dmfR, IS1071), IS6100, sul1, qacEΔ1, aadA2, IS6 composite transposon (mphE, msrE, IS26, tetRC, IS26, tetRC), blaD |
LC895901.1 |
| pMNDX110 | IncPβ | 77,896 | Tama6 | B |
CusA/CzcA, HlyD, heavy metal resistance, blaC, IS26, msrE, mphE, IS26, tetCR, IS26 |
LC895902.1 |
| pMNBM077 | IncPκ | 53,339 | Tama5 | B | Transposon (tetAR, transposon) | LC623892.1 |
| pYKTC011-1 | PromAβ-1 | 57,620 | Tama6 | T | Tn501(remnant)(mer operon), IS21, Tn3-family (blaNPS), relE | LC623931.1 |
| pMNBL076-1 | Not classified | 16,094 | Tama4 | B | tetA | LC663723.1 |
| Plasmid | Inc_group | Tc(12.5) | Gm(30) | Ap(50) | Km(50) | Cm(30) | Sm(25) | Sm(50) | Em(25) |
| pMNBM065-2 | IncU, IncQ2γ | R | - | - | - | R | - | - | - |
| pMNBM072 | IncN | R | S | R | R | - | - | R | - |
| pMNBL073 | IncC | R | R | S | S | R | - | - | - |
| pYKCS045 | IncPγ | - | S | R | S | - | - | - | - |
| pMNBL056 | IncPε | R | S | S | - | - | - | - | - |
| pYKBL037 | IncPι | - | S | R | R | - | R | S | R |
| pMNDW109 | IncPβ | R | S | R | - | - | - | - | - |
| pMNDX110 | IncPβ | R | S | S | - | - | - | - | - |
| pMNBM077 | IncPκ | R | - | S | - | - | - | - | - |
| pYKTC011-1 | PromAβ-1 | - | - | R | - | - | - | - | - |
| pMNBL076-1 | Not classified | - | - | - | - | - | - | - | - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




