Submitted:
22 March 2026
Posted:
23 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Clinical and Pathological Motivation
1.2. Purpose and Scope of the Review
2. Overview of the Five Neurodegeneration-Associated Proteins and Their Intrinsic Disorder Status
2.1. Amyloid-β (Aβ)
2.2. τ-Protein (Tau)
2.3. α-Synuclein
2.4. TAR DNA-Binding Protein 43 (TDP-43)
2.5. Fused in Sarcoma (FUS)
2.6. Commonality of Individualities
2.7. Mightly Alliance: Beyond the Individual Armies
3. And Intrinsic Disorder to Rule Them All
3.1. Intrinsic Disorder and LLPS
3.2. Intrinsic Disorder and Aggregation
3.3. Shared Disorder Features Across Neurodegeneration-Associated Proteins
3.4. Prion-like Behaviors in Disordered Protein Systems
3.5. Convergence of Aggregation Pathways in Mixed Pathology
3.6. Intrinsic Disorder, Aging, and Proteostasis Failure
3.7. Aberrant Stabilization of Condensates and Irreversible Aggregation
3.8. Systems-Level Failure to Regulate Disorder in Long-Lived Neurons
3.9. Implications for Disease Classification and Mechanistic Understanding: Rethinking Neurodegenerative Diseases as Disorder-Driven Network Failures
4. Potential Therapeutic Implementations of Intrinsic Disorder
4.1. Challenges of Targeting IDPs
4.2. Disorder-Aware Drug Discovery and Intervention Approaches
4.3. Integrating Intrinsic Disorder into Models of Protein-Specific Diseases
4.4. Biomarkers in Neurodegeneration Considering Intrinsic Disorder
5. Open Questions and Future Directions
5.1. Quantitative Thresholds Between Functional Disorder and Pathology
6. Conclusions
6.1. Intrinsic Disorder as a Unifying Biophysical Principle
6.2. From Isolated Proteinopathies to Interacting Disorder-Driven Networks
6.3. Outlook for Disorder-Centric Neurodegeneration Research
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Aβ | Amyloid-β |
| AD | Alzheimer's disease |
| AGD | Argyrophilic Grain Disease |
| ALS | Amyotrophic lateral sclerosis |
| APP | Amyloid precursor protein |
| CAA | Cerebral amyloid angiopathy |
| CBD | Corticobasal degeneration |
| CSF | Cerebrospinal fluid |
| CTE | Chronic traumatic encephalopathy |
| FTLD | Frontotemporal lobar degeneration |
| FTLD-FUS | Frontotemporal lobar degeneration with FUS pathology |
| FTLD-TDP | Frontotemporal lobar degeneration with TDP-43 pathology |
| FUS | Fused in sarcoma |
| IDP | Intrinsically disordered protein |
| IDR | Intrinsically disordered region |
| LATE | Limbic-predominant age-related TDP-43 encephalopathy |
| LLPS | Liquid–liquid phase separation |
| MSA | Multiple system atrophy |
| NFT | Neurofibrillary tangle |
| NAC | Non-amyloid component |
| NACP | Non-amyloid component precursor |
| PAF PD |
Pure Autonomic Failure Parkinson's Disease |
| PID | Protein intrinsic disorder |
| PiD | Pick’s disease |
| PSP | Progressive supranuclear palsy |
| PTM RBD |
Post-translational modification Rapid Eye Movement (REM) Sleep Behavior Disorder |
| RNA | Ribonucleic acid |
| SNCA | Synuclein alpha gene |
| TDP-43 | TAR DNA-binding protein 43 |
References
- Goedert, M.; Jakes, R.; Spillantini, M.G. The Synucleinopathies: Twenty Years On. J Parkinsons Dis 2017, 7, S51–S69. [Google Scholar] [CrossRef] [PubMed]
- Hattiholi, A.; Hegde, H.; Shetty, S.K. Tauopathies: Emerging discoveries on tau protein, with a special focus on Alzheimer's disease. Neuropeptides 2025, 112, 102536. [Google Scholar] [CrossRef] [PubMed]
- Padilla-Godínez, F.J.; Vázquez-García, E.R.; Trujillo-Villagrán, M.I.; Soto-Rojas, L.O.; Palomero-Rivero, M.; Hernández-González, O.; Pérez-Eugenio, F.; Collazo-Navarrete, O.; Arias-Carrión, O.; Guerra-Crespo, M. A-Synuclein and Tau: Interactions, Cross-Seeding, and the Redefinition of Synucleinopathies as Complex Proteinopathies. Frontiers in Neuroscience 2025, 19. [Google Scholar] [CrossRef] [PubMed]
- Forrest, S.L.; Kovacs, G.G. Current Concepts of Mixed Pathologies in Neurodegenerative Diseases. Can J Neurol Sci 2023, 50, 329–345. [Google Scholar] [CrossRef]
- Brenowitz, W.D.; Hubbard, R.A.; Keene, C.D.; Hawes, S.E.; Longstreth, W.T., Jr.; Woltjer, R.L.; Kukull, W.A. Mixed neuropathologies and estimated rates of clinical progression in a large autopsy sample. Alzheimers Dement 2017, 13, 654–662. [Google Scholar] [CrossRef]
- Attems, J.; Jellinger, K.A. The overlapping spectrum of Alzheimer disease, Lewy body disease and vascular pathology. Lancet Neurology 2014. [Google Scholar]
- Ling, S.C.; Polymenidou, M.; Cleveland, D.W. Converging Mechanisms in ALS and FTD: Disrupted RNA and Protein Homeostasis. Neuron 2013, 79, 416–438. [Google Scholar] [CrossRef]
- Spires-Jones, T.L.; Attems, J.; Thal, D.R. Interactions of pathological proteins in neurodegenerative diseases. Acta Neuropathologica 2017, 134, 187–205. [Google Scholar] [CrossRef]
- Wright, P.E.; Dyson, H.J. Intrinsically unstructured proteins: re-assessing the protein structure-function paradigm. J Mol Biol 1999, 293, 321–331. [Google Scholar] [CrossRef]
- Uversky, V.N.; Gillespie, J.R.; Fink, A.L. Why are "natively unfolded" proteins unstructured under physiologic conditions? Proteins 2000, 41, 415–427. [Google Scholar] [CrossRef]
- Dunker, A.K.; Lawson, J.D.; Brown, C.J.; Williams, R.M.; Romero, P.; Oh, J.S.; Oldfield, C.J.; Campen, A.M.; Ratliff, C.M.; Hipps, K.W.; et al. Intrinsically disordered protein. J Mol Graph Model 2001, 19, 26–59. [Google Scholar] [CrossRef] [PubMed]
- Dunker, A.K.; Obradovic, Z. The protein trinity--linking function and disorder. Nat Biotechnol 2001, 19, 805–806. [Google Scholar] [CrossRef] [PubMed]
- Tompa, P. Intrinsically unstructured proteins. Trends Biochem Sci 2002, 27, 527–533. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N. Natively unfolded proteins: a point where biology waits for physics. Protein Sci 2002, 11, 739–756. [Google Scholar] [CrossRef]
- Uversky, V.N. What does it mean to be natively unfolded? Eur J Biochem 2002, 269, 2–12. [Google Scholar] [CrossRef]
- Dyson, H.J.; Wright, P.E. Intrinsically unstructured proteins and their functions. Nat Rev Mol Cell Biol 2005, 6, 197–208. [Google Scholar] [CrossRef]
- Uversky, V.N.; Dunker, A.K. Understanding Protein Non-Folding. Biochimica Et Biophysica Acta (Bba) - Proteins and Proteomics 2010, 1804, 1231–1264. [Google Scholar] [CrossRef]
- Uversky, V.N. The mysterious unfoldome: structureless, underappreciated, yet vital part of any given proteome. J Biomed Biotechnol 2010, 2010, 568068. [Google Scholar] [CrossRef]
- Dunker, A.K.; Oldfield, C.J.; Meng, J.; Romero, P.; Yang, J.Y.; Chen, J.W.; Vacic, V.; Obradovic, Z.; Uversky, V.N. The unfoldomics decade: an update on intrinsically disordered proteins. BMC Genomics 2008, 9 Suppl 2, S1. [Google Scholar] [CrossRef]
- Dunker, A.K.; Silman, I.; Uversky, V.N.; Sussman, J.L. Function and structure of inherently disordered proteins. Curr Opin Struct Biol 2008, 18, 756–764. [Google Scholar] [CrossRef]
- Dunker, A.K.; Obradovic, Z.; Romero, P.; Garner, E.C.; Brown, C.J. Intrinsic protein disorder in complete genomes. Genome Inform Ser Workshop Genome Inform 2000, 11, 161–171. [Google Scholar] [PubMed]
- Ward, J.J.; Sodhi, J.S.; McGuffin, L.J.; Buxton, B.F.; Jones, D.T. Prediction and functional analysis of native disorder in proteins from the three kingdoms of life. J Mol Biol 2004, 337, 635–645. [Google Scholar] [CrossRef] [PubMed]
- Oldfield, C.J.; Cheng, Y.; Cortese, M.S.; Brown, C.J.; Uversky, V.N.; Dunker, A.K. Comparing and combining predictors of mostly disordered proteins. Biochemistry 2005, 44, 1989–2000. [Google Scholar] [CrossRef] [PubMed]
- Xue, B.; Dunker, A.K.; Uversky, V.N. Orderly order in protein intrinsic disorder distribution: disorder in 3500 proteomes from viruses and the three domains of life. Journal of biomolecular structure & dynamics 2012, 30, 137–149. [Google Scholar] [CrossRef]
- Peng, Z.; Oldfield, C.J.; Xue, B.; Mizianty, M.J.; Dunker, A.K.; Kurgan, L.; Uversky, V.N. A creature with a hundred waggly tails: intrinsically disordered proteins in the ribosome. Cellular and molecular life sciences: CMLS 2014, 71, 1477–1504. [Google Scholar] [CrossRef]
- Peng, Z.; Yan, J.; Fan, X.; Mizianty, M.J.; Xue, B.; Wang, K.; Hu, G.; Uversky, V.N.; Kurgan, L. Exceptionally abundant exceptions: comprehensive characterization of intrinsic disorder in all domains of life. Cellular and molecular life sciences: CMLS 2015, 72, 137–151. [Google Scholar] [CrossRef]
- Uversky, V.N. Protein folding revisited. A polypeptide chain at the folding-misfolding-nonfolding cross-roads: which way to go? Cellular and molecular life sciences: CMLS 2003, 60, 1852–1871. [Google Scholar] [CrossRef]
- Uversky, V.N. Intrinsic disorder-based protein interactions and their modulators. Curr Pharm Des 2013, 19, 4191–4213. [Google Scholar] [CrossRef]
- Williams, R.M.; Obradovi, Z.; Mathura, V.; Braun, W.; Garner, E.C.; Young, J.; Takayama, S.; Brown, C.J.; Dunker, A.K. The protein non-folding problem: amino acid determinants of intrinsic order and disorder. Pac Symp Biocomput 2001, 89–100. [Google Scholar]
- Romero, P.; Obradovic, Z.; Li, X.; Garner, E.C.; Brown, C.J.; Dunker, A.K. Sequence complexity of disordered protein. Proteins 2001, 42, 38–48. [Google Scholar] [CrossRef]
- Radivojac, P.; Iakoucheva, L.M.; Oldfield, C.J.; Obradovic, Z.; Uversky, V.N.; Dunker, A.K. Intrinsic disorder and functional proteomics. Biophys J 2007, 92, 1439–1456. [Google Scholar] [CrossRef]
- Vacic, V.; Uversky, V.N.; Dunker, A.K.; Lonardi, S. Composition Profiler: a tool for discovery and visualization of amino acid composition differences. BMC Bioinformatics 2007, 8, 211. [Google Scholar] [CrossRef] [PubMed]
- Hemmings, H.C., Jr.; Nairn, A.C.; Aswad, D.W.; Greengard, P. DARPP-32, a dopamine- and adenosine 3':5'-monophosphate-regulated phosphoprotein enriched in dopamine-innervated brain regions. II. Purification and characterization of the phosphoprotein from bovine caudate nucleus. J Neurosci 1984, 4, 99–110. [Google Scholar] [CrossRef] [PubMed]
- Gast, K.; Damaschun, H.; Eckert, K.; Schulze-Forster, K.; Maurer, H.R.; Muller-Frohne, M.; Zirwer, D.; Czarnecki, J.; Damaschun, G. Prothymosin alpha: a biologically active protein with random coil conformation. Biochemistry 1995, 34, 13211–13218. [Google Scholar] [CrossRef] [PubMed]
- Weinreb, P.H.; Zhen, W.; Poon, A.W.; Conway, K.A.; Lansbury, P.T., Jr. NACP, a protein implicated in Alzheimer's disease and learning, is natively unfolded. Biochemistry 1996, 35, 13709–13715. [Google Scholar] [CrossRef]
- Freiberger, M.I.; Wolynes, P.G.; Ferreiro, D.U.; Fuxreiter, M. Frustration in Fuzzy Protein Complexes Leads to Interaction Versatility. J Phys Chem B 2021, 125, 2513–2520. [Google Scholar] [CrossRef]
- Fuxreiter, M. Fuzziness: linking regulation to protein dynamics. Mol Biosyst 2012, 8, 168–177. [Google Scholar] [CrossRef]
- Fuxreiter, M. Towards a Stochastic Paradigm: From Fuzzy Ensembles to Cellular Functions. Molecules 2018, 23. [Google Scholar] [CrossRef]
- Fuxreiter, M. Fuzzy protein theory for disordered proteins. Biochem Soc Trans 2020, 48, 2557–2564. [Google Scholar] [CrossRef]
- Fuxreiter, M. Context-dependent, fuzzy protein interactions: Towards sequence-based insights. Curr Opin Struct Biol 2024, 87, 102834. [Google Scholar] [CrossRef]
- Fuxreiter, M.; Tompa, P. Fuzzy complexes: a more stochastic view of protein function. Adv Exp Med Biol 2012, 725, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Miskei, M.; Gregus, A.; Sharma, R.; Duro, N.; Zsolyomi, F.; Fuxreiter, M. Fuzziness enables context dependence of protein interactions. FEBS Lett 2017, 591, 2682–2695. [Google Scholar] [CrossRef] [PubMed]
- Sharma, R.; Raduly, Z.; Miskei, M.; Fuxreiter, M. Fuzzy complexes: Specific binding without complete folding. FEBS Lett 2015, 589, 2533–2542. [Google Scholar] [CrossRef] [PubMed]
- Tompa, P.; Fuxreiter, M. Fuzzy complexes: polymorphism and structural disorder in protein-protein interactions. Trends Biochem Sci 2008, 33, 2–8. [Google Scholar] [CrossRef]
- Welch, G.R. Fuzziness" in the celular interactome: a historical perspective. Adv Exp Med Biol 2012, 725, 184–190. [Google Scholar] [CrossRef]
- Dosztanyi, Z.; Chen, J.; Dunker, A.K.; Simon, I.; Tompa, P. Disorder and sequence repeats in hub proteins and their implications for network evolution. J Proteome Res 2006, 5, 2985–2995. [Google Scholar] [CrossRef]
- Dunker, A.K.; Cortese, M.S.; Romero, P.; Iakoucheva, L.M.; Uversky, V.N. Flexible nets. The roles of intrinsic disorder in protein interaction networks. FEBS J 2005, 272, 5129–5148. [Google Scholar] [CrossRef]
- Haynes, C.; Oldfield, C.J.; Ji, F.; Klitgord, N.; Cusick, M.E.; Radivojac, P.; Uversky, V.N.; Vidal, M.; Iakoucheva, L.M. Intrinsic disorder is a common feature of hub proteins from four eukaryotic interactomes. PLoS Comput Biol 2006, 2, e100. [Google Scholar] [CrossRef]
- Hu, G.; Wu, Z.; Uversky, V.N.; Kurgan, L. Functional Analysis of Human Hub Proteins and Their Interactors Involved in the Intrinsic Disorder-Enriched Interactions. Int J Mol Sci 2017, 18. [Google Scholar] [CrossRef]
- Oldfield, C.J.; Meng, J.; Yang, J.Y.; Yang, M.Q.; Uversky, V.N.; Dunker, A.K. Flexible nets: disorder and induced fit in the associations of p53 and 14-3-3 with their partners. BMC Genomics 2008, 9 Suppl 1, S1. [Google Scholar] [CrossRef]
- Oldfield, C.J.; Dunker, A.K. Intrinsically disordered proteins and intrinsically disordered protein regions. Annu Rev Biochem 2014, 83, 553–584. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N. Intrinsically disordered proteins from A to Z. Int J Biochem Cell Biol 2011, 43, 1090–1103. [Google Scholar] [CrossRef] [PubMed]
- Wright, P.E.; Dyson, H.J. Intrinsically disordered proteins in cellular signalling and regulation. Nat Rev Mol Cell Biol 2015, 16, 18–29. [Google Scholar] [CrossRef] [PubMed]
- Dunker, A.K.; Brown, C.J.; Obradovic, Z. Identification and functions of usefully disordered proteins. Adv Protein Chem 2002, 62, 25–49. [Google Scholar]
- Tompa, P.; Csermely, P. The role of structural disorder in the function of RNA and protein chaperones. Faseb J 2004, 18, 1169–1175. [Google Scholar] [CrossRef]
- Bondos, S.E.; Dunker, A.K.; Uversky, V.N. On the roles of intrinsically disordered proteins and regions in cell communication and signaling. Cell Commun Signal 2021, 19, 88. [Google Scholar] [CrossRef]
- Bondos, S.E.; Dunker, A.K.; Uversky, V.N. Intrinsically disordered proteins play diverse roles in cell signaling. Cell Commun Signal 2022, 20, 20. [Google Scholar] [CrossRef]
- Kulkarni, P.; Bhattacharya, S.; Achuthan, S.; Behal, A.; Jolly, M.K.; Kotnala, S.; Mohanty, A.; Rangarajan, G.; Salgia, R.; Uversky, V. Intrinsically Disordered Proteins: Critical Components of the Wetware. Chem Rev 2022, 122, 6614–6633. [Google Scholar] [CrossRef]
- Kulkarni, P.; Leite, V.B.P.; Roy, S.; Bhattacharyya, S.; Mohanty, A.; Achuthan, S.; Singh, D.; Appadurai, R.; Rangarajan, G.; Weninger, K.; et al. Intrinsically disordered proteins: Ensembles at the limits of Anfinsen's dogma. Biophys Rev (Melville) 2022, 3, 011306. [Google Scholar] [CrossRef]
- Fuxreiter, M.; Tompa, P.; Simon, I.; Uversky, V.N.; Hansen, J.C.; Asturias, F.J. Malleable machines take shape in eukaryotic transcriptional regulation. Nat Chem Biol 2008, 4, 728–737. [Google Scholar] [CrossRef]
- Fuxreiter, M.; Toth-Petroczy, A.; Kraut, D.A.; Matouschek, A.; Lim, R.Y.; Xue, B.; Kurgan, L.; Uversky, V.N. Disordered proteinaceous machines. Chem Rev 2014, 114, 6806–6843. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N. The multifaceted roles of intrinsic disorder in protein complexes. FEBS Lett 2015, 589, 2498–2506. [Google Scholar] [CrossRef] [PubMed]
- El Hadidy, N.; Uversky, V.N. Intrinsic Disorder of the BAF Complex: Roles in Chromatin Remodeling and Disease Development. Int J Mol Sci 2019, 20. [Google Scholar] [CrossRef] [PubMed]
- Toth-Petroczy, A.; Oldfield, C.J.; Simon, I.; Takagi, Y.; Dunker, A.K.; Uversky, V.N.; Fuxreiter, M. Malleable machines in transcription regulation: the mediator complex. PLoS Comput Biol 2008, 4, e1000243. [Google Scholar] [CrossRef]
- Nesterov, S.V.; Ilyinsky, N.S.; Plokhikh, K.S.; Manuylov, V.D.; Chesnokov, Y.M.; Vasilov, R.G.; Kuznetsova, I.M.; Turoverov, K.K.; Gordeliy, V.I.; Fonin, A.V.; et al. Order wrapped in chaos: On the roles of intrinsically disordered proteins and RNAs in the arrangement of the mitochondrial enzymatic machines. Int J Biol Macromol 2024, 267, 131455. [Google Scholar] [CrossRef]
- Coelho Ribeiro Mde, L.; Espinosa, J.; Islam, S.; Martinez, O.; Thanki, J.J.; Mazariegos, S.; Nguyen, T.; Larina, M.; Xue, B.; Uversky, V.N. Malleable ribonucleoprotein machine: protein intrinsic disorder in the Saccharomyces cerevisiae spliceosome. PeerJ 2013, 1, e2. [Google Scholar] [CrossRef]
- Peng, Z.; Mizianty, M.J.; Xue, B.; Kurgan, L.; Uversky, V.N. More than just tails: intrinsic disorder in histone proteins. Mol Biosyst 2012, 8, 1886–1901. [Google Scholar] [CrossRef]
- Iakoucheva, L.M.; Radivojac, P.; Brown, C.J.; O'Connor, T.R.; Sikes, J.G.; Obradovic, Z.; Dunker, A.K. The importance of intrinsic disorder for protein phosphorylation. Nucleic Acids Res 2004, 32, 1037–1049. [Google Scholar] [CrossRef]
- Pejaver, V.; Hsu, W.L.; Xin, F.; Dunker, A.K.; Uversky, V.N.; Radivojac, P. The structural and functional signatures of proteins that undergo multiple events of post-translational modification. Protein Sci 2014, 23, 1077–1093. [Google Scholar] [CrossRef]
- Darling, A.L.; Uversky, V.N. Intrinsic Disorder and Posttranslational Modifications: The Darker Side of the Biological Dark Matter. Front Genet 2018, 9, 158. [Google Scholar] [CrossRef]
- Meng, F.; Na, I.; Kurgan, L.; Uversky, V.N. Compartmentalization and Functionality of Nuclear Disorder: Intrinsic Disorder and Protein-Protein Interactions in Intra-Nuclear Compartments. Int J Mol Sci 2015, 17. [Google Scholar] [CrossRef] [PubMed]
- Darling, A.L.; Liu, Y.; Oldfield, C.J.; Uversky, V.N. Intrinsically Disordered Proteome of Human Membrane-Less Organelles. Proteomics 2018, 18, e1700193 (1700112 pages. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N. The roles of intrinsic disorder-based liquid-liquid phase transitions in the "Dr. Jekyll-Mr. Hyde" behavior of proteins involved in amyotrophic lateral sclerosis and frontotemporal lobar degeneration. Autophagy 2017, 13, 2115–2162. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N. Intrinsically disordered proteins in overcrowded milieu: Membrane-less organelles, phase separation, and intrinsic disorder. Curr Opin Struct Biol 2017, 44, 18–30. [Google Scholar] [CrossRef]
- Uversky, V.N. Protein intrinsic disorder-based liquid-liquid phase transitions in biological systems: Complex coacervates and membrane-less organelles. Adv Colloid Interface Sci 2017, 239, 97–114. [Google Scholar] [CrossRef]
- Antifeeva, I.A.; Fonin, A.V.; Fefilova, A.S.; Stepanenko, O.V.; Povarova, O.I.; Silonov, S.A.; Kuznetsova, I.M.; Uversky, V.N.; Turoverov, K.K. Liquid-liquid phase separation as an organizing principle of intracellular space: overview of the evolution of the cell compartmentalization concept. Cellular and molecular life sciences: CMLS 2022, 79, 251. [Google Scholar] [CrossRef]
- Fonin, A.V.; Antifeeva, I.A.; Kuznetsova, I.M.; Turoverov, K.K.; Zaslavsky, B.Y.; Kulkarni, P.; Uversky, V.N. Biological soft matter: intrinsically disordered proteins in liquid-liquid phase separation and biomolecular condensates. Essays Biochem 2022, 66, 831–847. [Google Scholar] [CrossRef]
- Fonin, A.V.; Darling, A.L.; Kuznetsova, I.M.; Turoverov, K.K.; Uversky, V.N. Intrinsically disordered proteins in crowded milieu: when chaos prevails within the cellular gumbo. Cellular and molecular life sciences: CMLS 2018, 75, 3907–3929. [Google Scholar] [CrossRef]
- Turoverov, K.K.; Kuznetsova, I.M.; Fonin, A.V.; Darling, A.L.; Zaslavsky, B.Y.; Uversky, V.N. Stochasticity of Biological Soft Matter: Emerging Concepts in Intrinsically Disordered Proteins and Biological Phase Separation. Trends Biochem Sci 2019, 44, 716–728. [Google Scholar] [CrossRef]
- Uversky, V.N.; Kuznetsova, I.M.; Turoverov, K.K.; Zaslavsky, B. Intrinsically disordered proteins as crucial constituents of cellular aqueous two phase systems and coacervates. FEBS Lett 2015, 589, 15–22. [Google Scholar] [CrossRef]
- Darling, A.L.; Zaslavsky, B.Y.; Uversky, V.N. Intrinsic Disorder-Based Emergence in Cellular Biology: Physiological and Pathological Liquid-Liquid Phase Transitions in Cells. Polymers 2019, 11, 990. [Google Scholar] [CrossRef] [PubMed]
- Darling, A.L.; Shorter, J. Combating deleterious phase transitions in neurodegenerative disease. Biochim Biophys Acta Mol Cell Res 2021, 1868, 118984. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N.; Dave, V.; Iakoucheva, L.M.; Malaney, P.; Metallo, S.J.; Pathak, R.R.; Joerger, A.C. Pathological unfoldomics of uncontrolled chaos: intrinsically disordered proteins and human diseases. Chem Rev 2014, 114, 6844–6879. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N.; Oldfield, C.J.; Dunker, A.K. Intrinsically Disordered Proteins in Human Diseases: Introducing the D2Concept. Annual Review of Biophysics 2008, 37, 215–246. [Google Scholar] [CrossRef]
- Gadhave, K.; Gehi, B.R.; Kumar, P.; Xue, B.; Uversky, V.N.; Giri, R. The dark side of Alzheimer's disease: unstructured biology of proteins from the amyloid cascade signaling pathway. Cellular and molecular life sciences: CMLS 2020, 77, 4163–4208. [Google Scholar] [CrossRef]
- Uversky, V.N. Wrecked regulation of intrinsically disordered proteins in diseases: pathogenicity of deregulated regulators. Front Mol Biosci 2014, 1, 6. [Google Scholar] [CrossRef]
- Uversky, V.N. Rebellion of the deregulated regulators: What is the clinical relevance of studying intrinsically disordered proteins? Expert Rev Proteomics 2022, 19, 279–282. [Google Scholar] [CrossRef]
- Rahimi, J.; Kovacs, G.G. Prevalence of mixed pathologies in the aging brain. Alzheimers Res Ther 2014, 6, 82. [Google Scholar] [CrossRef]
- Kovacs, G.G.; Milenkovic, I.; Wohrer, A.; Hoftberger, R.; Gelpi, E.; Haberler, C.; Honigschnabl, S.; Reiner-Concin, A.; Heinzl, H.; Jungwirth, S.; et al. Non-Alzheimer neurodegenerative pathologies and their combinations are more frequent than commonly believed in the elderly brain: a community-based autopsy series. Acta Neuropathol 2013, 126, 365–384. [Google Scholar] [CrossRef]
- Sagare, A.P.; Deane, R.; Zloković, B.V. Low-Density Lipoprotein Receptor-Related Protein 1: A Physiological Aβ Homeostatic Mechanism With Multiple Therapeutic Opportunities. Pharmacology & Therapeutics 2012, 136, 94–105. [Google Scholar] [CrossRef]
- Xin, S.-H.; Tan, L.; Cao, X.P.; Yu, J.T.; Tan, L. Clearance of Amyloid Beta and Tau in Alzheimer’s Disease: From Mechanisms to Therapy. Neurotoxicity Research 2018, 34, 733–748. [Google Scholar] [CrossRef]
- Alvira, X.; Carro, E. Clearance of Amyloid-Β Peptide Across the Choroid Plexus in Alzheimers Disease. Current Aging Science 2010, 3, 219–229. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.-f.; Xu, T.-H.; Yan, Y.; Zhou, Y.-r.; Jiang, Y.; Melcher, K.; Xu, H.E. Amyloid Beta: Structure, Biology and Structure-Based Therapeutic Development. Acta Pharmacologica Sinica 2017, 38, 1205–1235. [Google Scholar] [CrossRef] [PubMed]
- Eisenberg, D.; Jucker, M. The Amyloid State of Proteins in Human Diseases. Cell 2012, 148, 1188–1203. [Google Scholar] [CrossRef] [PubMed]
- Petkova, A.T.; Yau, W.M.; Tycko, R. Experimental Constraints on Quaternary Structure in Alzheimer's Β-Amyloid Fibrils. Biochemistry 2005, 45, 498–512. [Google Scholar] [CrossRef]
- Selkoe, D.J. Translating Cell Biology Into Therapeutic Advances in Alzheimer's Disease. Nature 1999, 399, A23–A31. [Google Scholar] [CrossRef]
- Bitan, G.; Kirkitadze, M.; Lomakin, A.; Vollers, S.S.; Benedek, G.B.; Teplow, D.B. Amyloid Β-Protein (Aβ) Assembly: Aβ40 and Aβ42 Oligomerize Through Distinct Pathways. Proceedings of the National Academy of Sciences 2002, 100, 330–335. [Google Scholar] [CrossRef]
- Hamley, I.W. The Amyloid Beta Peptide: A Chemist’s Perspective. Role in Alzheimer’s and Fibrillization. Chemical Reviews 2012, 112, 5147–5192. [Google Scholar] [CrossRef]
- Vivekanandan, S.; Brender, J.R.; Lee, S.Y.; Ramamoorthy, A. A partially folded structure of amyloid-beta(1-40) in an aqueous environment. Biochem Biophys Res Commun 2011, 411, 312–316. [Google Scholar] [CrossRef]
- Hou, L.; Shao, H.; Zhang, Y.; Li, H.; Menon, N.K.; Neuhaus, E.B.; Brewer, J.M.; Byeon, I.J.; Ray, D.G.; Vitek, M.P.; et al. Solution NMR studies of the A beta(1-40) and A beta(1-42) peptides establish that the Met35 oxidation state affects the mechanism of amyloid formation. J Am Chem Soc 2004, 126, 1992–2005. [Google Scholar] [CrossRef]
- Lazo, N.D.; Grant, M.A.; Condron, M.C.; Rigby, A.C.; Teplow, D.B. On the nucleation of amyloid beta-protein monomer folding. Protein Sci 2005, 14, 1581–1596. [Google Scholar] [CrossRef] [PubMed]
- Saieva, S.; Morales, R. Misfolded amyloid-beta conformational variants (strains) as drivers of Alzheimer's disease neuropathology. Neural Regen Res 2025, 20, 3219–3220. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.L.; Lee, V.M. Cell-to-cell transmission of pathogenic proteins in neurodegenerative diseases. Nat Med 2014, 20, 130–138. [Google Scholar] [CrossRef] [PubMed]
- Jucker, M.; Walker, L.C. Self-propagation of pathogenic protein aggregates in neurodegenerative diseases. Nature 2013, 501, 45–51. [Google Scholar] [CrossRef] [PubMed]
- Morales, R.; Green, K.M.; Soto, C. Cross currents in protein misfolding disorders: interactions and therapy. CNS Neurol Disord Drug Targets 2009, 8, 363–371. [Google Scholar] [CrossRef]
- Zhang, X.; Fu, Z.; Meng, L.; He, M.; Zhang, Z. The Early Events That Initiate beta-Amyloid Aggregation in Alzheimer's Disease. Front Aging Neurosci 2018, 10, 359. [Google Scholar] [CrossRef]
- Dey, A.; Verma, A.; Bhaskar, U.; Sarkar, B.; Kallianpur, M.; Vishvakarma, V.; Das, A.K.; Garai, K.; Mukherjee, O.; Ishii, K.; et al. A Toxicogenic Interaction Between Intracellular Amyloid-Β and Apolipoprotein-E. Acs Chemical Neuroscience 2024, 15, 1265–1275. [Google Scholar] [CrossRef]
- Busche, M.A.; Hyman, B.T. Synergy between amyloid-β and tau in Alzheimer's disease. Nat Neurosci 2020, 23, 1183–1193. [Google Scholar] [CrossRef]
- Porta, S.; Xu, Y.; Restrepo, C.R.; Kwong, L.K.; Zhang, B.; Brown, H.J.; Lee, E.B.; Trojanowski, J.Q.; Lee, V.M. Patient-Derived Frontotemporal Lobar Degeneration Brain Extracts Induce Formation and Spreading of TDP-43 Pathology in Vivo. Nature Communications 2018, 9. [Google Scholar] [CrossRef]
- Baumkotter, F.; Schmidt, N.; Vargas, C.; Schilling, S.; Weber, R.; Wagner, K.; Fiedler, S.; Klug, W.; Radzimanowski, J.; Nickolaus, S.; et al. Amyloid precursor protein dimerization and synaptogenic function depend on copper binding to the growth factor-like domain. J Neurosci 2014, 34, 11159–11172. [Google Scholar] [CrossRef]
- Oughtred, R.; Rust, J.; Chang, C.; Breitkreutz, B.J.; Stark, C.; Willems, A.; Boucher, L.; Leung, G.; Kolas, N.; Zhang, F.; et al. The BioGRID database: A comprehensive biomedical resource of curated protein, genetic, and chemical interactions. Protein Sci 2021, 30, 187–200. [Google Scholar] [CrossRef] [PubMed]
- Breitkreutz, B.J.; Stark, C.; Reguly, T.; Boucher, L.; Breitkreutz, A.; Livstone, M.; Oughtred, R.; Lackner, D.H.; Bahler, J.; Wood, V.; et al. The BioGRID Interaction Database: 2008 update. Nucleic Acids Res 2008, 36, D637–640. [Google Scholar] [CrossRef] [PubMed]
- Curtain, C.C.; Ali, F.; Volitakis, I.; Cherny, R.A.; Norton, R.S.; Beyreuther, K.; Barrow, C.J.; Masters, C.L.; Bush, A.I.; Barnham, K.J. Alzheimer's disease amyloid-beta binds copper and zinc to generate an allosterically ordered membrane-penetrating structure containing superoxide dismutase-like subunits. J Biol Chem 2001, 276, 20466–20473. [Google Scholar] [CrossRef] [PubMed]
- Hardenberg, M.; Horvath, A.; Ambrus, V.; Fuxreiter, M.; Vendruscolo, M. Widespread occurrence of the droplet state of proteins in the human proteome. Proc Natl Acad Sci U S A 2020, 117, 33254–33262. [Google Scholar] [CrossRef]
- Chen, J.; Li, S.; Zhang, F.; Chen, J.; Cai, C.; Guo, Y.; Lei, Z.; Zeng, L.H.; Zi, D.; Shen, Y.; et al. The pathogenic APP N-terminal Val225Ala mutation alters tau protein liquid-liquid phase separation and exacerbates synaptic damage. Mol Psychiatry 2025, 30, 2316–2334. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Zidek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Oates, M.E.; Romero, P.; Ishida, T.; Ghalwash, M.; Mizianty, M.J.; Xue, B.; Dosztanyi, Z.; Uversky, V.N.; Obradovic, Z.; Kurgan, L.; et al. D(2)P(2): database of disordered protein predictions. Nucleic Acids Res 2013, 41, D508–516. [Google Scholar] [CrossRef]
- Baas, P.W.; Qiang, L. Tau: It's Not What You Think. Trends Cell Biol 2019, 29, 452–461. [Google Scholar] [CrossRef]
- van der Lee, R.; Buljan, M.; Lang, B.; Weatheritt, R.J.; Daughdrill, G.W.; Dunker, A.K.; Fuxreiter, M.; Gough, J.; Gsponer, J.; Jones, D.T.; et al. Classification of intrinsically disordered regions and proteins. Chem Rev 2014, 114, 6589–6631. [Google Scholar] [CrossRef]
- Uversky, V.N. A Decade and a Half of Protein Intrinsic Disorder: Biology Still Waits for Physics. Protein Science 2013, 22, 693–724. [Google Scholar] [CrossRef]
- Uversky, V.N. Intrinsically Disordered Proteins and Their “Mysterious” (Meta)Physics. Frontiers in Physics 2019, 7. [Google Scholar] [CrossRef]
- Uversky, V.N. Intrinsically Disordered Proteins and Their Environment: Effects of Strong Denaturants, Temperature, pH, Counter Ions, Membranes, Binding Partners, Osmolytes, and Macromolecular Crowding. The Protein Journal 2009, 28, 305–325. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Lin, Y.; Eschmann, N.A.; Zhou, H.; Rauch, J.N.; Hernandez, I.; Guzman, E.; Kosik, K.S.; Han, S. RNA stores tau reversibly in complex coacervates. PLoS Biol 2017, 15, e2002183. [Google Scholar] [CrossRef]
- Wegmann, S.; Eftekharzadeh, B.; Tepper, K.; Zoltowska, K.M.; Bennett, R.E.; Dujardin, S.; Laskowski, P.R.; MacKenzie, D.; Kamath, T.; Commins, C.; et al. Tau protein liquid-liquid phase separation can initiate tau aggregation. EMBO J 2018, 37. [Google Scholar] [CrossRef]
- Boyko, S.; Qi, X.; Chen, T.H.; Surewicz, K.; Surewicz, W.K. Liquid-liquid phase separation of tau protein: The crucial role of electrostatic interactions. J Biol Chem 2019, 294, 11054–11059. [Google Scholar] [CrossRef] [PubMed]
- Margittai, M. Driving tau into phase-separated liquid droplets. J Biol Chem 2019, 294, 11060–11061. [Google Scholar] [CrossRef] [PubMed]
- Wegmann, S. Liquid-Liquid Phase Separation of Tau Protein in Neurobiology and Pathology. Adv Exp Med Biol 2019, 1184, 341–357. [Google Scholar] [CrossRef]
- Boyko, S.; Surewicz, K.; Surewicz, W.K. Regulatory mechanisms of tau protein fibrillation under the conditions of liquid-liquid phase separation. Proc Natl Acad Sci U S A 2020, 117, 31882–31890. [Google Scholar] [CrossRef]
- Kanaan, N.M.; Hamel, C.; Grabinski, T.; Combs, B. Liquid-liquid phase separation induces pathogenic tau conformations in vitro. Nat Commun 2020, 11, 2809. [Google Scholar] [CrossRef]
- Ukmar-Godec, T.; Wegmann, S.; Zweckstetter, M. Biomolecular condensation of the microtubule-associated protein tau. Semin Cell Dev Biol 2020, 99, 202–214. [Google Scholar] [CrossRef]
- Zhang, X.; Vigers, M.; McCarty, J.; Rauch, J.N.; Fredrickson, G.H.; Wilson, M.Z.; Shea, J.E.; Han, S.; Kosik, K.S. The proline-rich domain promotes Tau liquid-liquid phase separation in cells. J Cell Biol 2020, 219. [Google Scholar] [CrossRef]
- Dong, X.; Bera, S.; Qiao, Q.; Tang, Y.; Lao, Z.; Luo, Y.; Gazit, E.; Wei, G. Liquid-Liquid Phase Separation of Tau Protein Is Encoded at the Monomeric Level. J Phys Chem Lett 2021, 12, 2576–2586. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Fichou, Y.; Longhini, A.P.; Llanes, L.C.; Yin, P.; Bazan, G.C.; Kosik, K.S.; Han, S. Liquid-Liquid Phase Separation of Tau Driven by Hydrophobic Interaction Facilitates Fibrillization of Tau. J Mol Biol 2021, 433, 166731. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, P.; Zweckstetter, M. Phase separation of the microtubule-associated protein tau. Essays Biochem 2022, 66, 1013–1021. [Google Scholar] [CrossRef] [PubMed]
- Prince, P.R.; Hochmair, J.; Brognaro, H.; Gevorgyan, S.; Franck, M.; Schubert, R.; Lorenzen, K.; Yazici, S.; Mandelkow, E.; Wegmann, S.; et al. Initiation and modulation of Tau protein phase separation by the drug suramin. Sci Rep 2023, 13, 3963. [Google Scholar] [CrossRef]
- Cuevas-Velazquez, C.L.; Vellosillo, T.; Guadalupe, K.; Schmidt, H.B.; Yu, F.; Moses, D.; Brophy, J.A.N.; Cosio-Acosta, D.; Das, A.; Wang, L.; et al. Intrinsically Disordered Protein Biosensor Tracks the Physical-Chemical Effects of Osmotic Stress on Cells. Nature Communications 2021, 12. [Google Scholar] [CrossRef]
- Bianchi, G.; Mangiagalli, M.; Ami, D.; Ahmed, J.; Lombardi, S.; Longhi, S.; Natalello, A.; Tompa, P.; Brocca, S. Condensation of the N-Terminal Domain of Human Topoisomerase 1 Is Driven by Electrostatic Interactions and Tuned by Its Charge Distribution. International Journal of Biological Macromolecules 2024, 254, 127754. [Google Scholar] [CrossRef]
- Bravo, C.; Krukowski, K.; Barker, S.; Wang, C.; Li, Y.; Fan, L.; Vázquez-Rosa, E.; Shin, M.K.; Wong, M.Y.; McCullough, L.D.; et al. Anti-Acetylated-Tau Immunotherapy Is Neuroprotective in Tauopathy and Brain Injury. Molecular Neurodegeneration 2024, 19. [Google Scholar] [CrossRef]
- Ayers, J.I.; Giasson, B.I.; Borchelt, D. Prion-Like Spreading in Tauopathies. Biological Psychiatry 2018, 83, 337–346. [Google Scholar] [CrossRef]
- Arriagada, P.V.; Growdon, J.H.; Hedley-Whyte, E.T.; Hyman, B.T. Neurofibrillary Tangles but Not Senile Plaques Parallel Duration and Severity of Alzheimer's Disease. Neurology 1992, 42, 631–631. [Google Scholar] [CrossRef]
- Eisele, Y.S. From Soluble Aβ to Progressive Aβ Aggregation: Could Prion-Like Templated Misfolding Play a Role? Brain Pathology 2013, 23, 333–341. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.H.; Chan, H.S. Phase Separation and Single-Chain Compactness of Charged Disordered Proteins Are Strongly Correlated. Biophysical Journal 2017, 112, 2043–2046. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N. Looking at the recent advances in understanding alpha-synuclein and its aggregation through the proteoform prism. F1000Res 2017, 6, 525. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N. A protein-chameleon: conformational plasticity of alpha-synuclein, a disordered protein involved in neurodegenerative disorders. Journal of biomolecular structure & dynamics 2003, 21, 211–234. [Google Scholar] [CrossRef]
- Theillet, F.X.; Binolfi, A.; Bekei, B.; Martorana, A.; Rose, H.; Stuiver, M.; Verzini, S.; Lorenz, D.; Rossum, M.v.; Goldfarb, D.; et al. Structural Disorder of Monomeric A-Synuclein Persists in Mammalian Cells. Nature 2016, 530, 45–50. [Google Scholar] [CrossRef]
- Burmann, B.M.; Gerez, J.; Matečko-Burmann, I.; Campioni, S.; Kumari, P.; Ghosh, D.; Mazur, A.; Aspholm, E.E.; Šulskis, D.; Wawrzyniuk, M.; et al. Regulation of A-Synuclein by Chaperones in Mammalian Cells. Nature 2019, 577, 127–132. [Google Scholar] [CrossRef]
- Uversky, V.N.; Li, J.; Fink, A.L. Evidence for a partially folded intermediate in alpha-synuclein fibril formation. J Biol Chem 2001, 276, 10737–10744. [Google Scholar] [CrossRef]
- Moons, R.; Konijnenberg, A.; Mensch, C.; Elzen, R.V.; Johannessen, C.; Maudsley, S.; Lambeir, A.M.; Sobott, F. Metal Ions Shape A-Synuclein. Scientific Reports 2020, 10. [Google Scholar] [CrossRef]
- Lashuel, H.A.; Overk, C.; Oueslati, A.; Masliah, E. The Many Faces of A-Synuclein: From Structure and Toxicity to Therapeutic Target. Nature Reviews Neuroscience 2012, 14, 38–48. [Google Scholar] [CrossRef]
- Uversky, V.N.; Eliezer, D. Biophysics of Parkinsons Disease: Structure and Aggregation of &Amp;#945;- Synuclein. Current Protein and Peptide Science 2009, 10, 483–499. [Google Scholar] [CrossRef]
- Stephens, A.D.; Zacharopoulou, M.; Moons, R.; Fusco, G.; Seetaloo, N.; Chiki, A.; Woodhams, P.J.; Mela, I.; Lashuel, H.A.; Phillips, J.J.; et al. Extent of N-Terminus Exposure of Monomeric Alpha-Synuclein Determines Its Aggregation Propensity. Nature Communications 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N.; Li, J.; Fink, A.L. Metal-triggered structural transformations, aggregation, and fibrillation of human alpha-synuclein. A possible molecular NK between Parkinson's disease and heavy metal exposure. J Biol Chem 2001, 276, 44284–44296. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N.; Li, J.; Fink, A.L. Pesticides directly accelerate the rate of alpha-synuclein fibril formation: a possible factor in Parkinson's disease. FEBS Lett 2001, 500, 105–108. [Google Scholar] [CrossRef] [PubMed]
- Manning-Bog, A.B.; McCormack, A.L.; Li, J.; Uversky, V.N.; Fink, A.L.; Di Monte, D.A. The herbicide paraquat causes up-regulation and aggregation of alpha-synuclein in mice: paraquat and alpha-synuclein. J Biol Chem 2002, 277, 1641–1644. [Google Scholar] [CrossRef]
- Trexler, A.; Rhoades, E. A-Synuclein Binds Large Unilamellar Vesicles as an Extended Helix. Biochemistry 2009, 48, 2304–2306. [Google Scholar] [CrossRef]
- Jo, E.; McLaurin, J.; Yip, C.M.; St. George-Hyslop, P.; Fraser, P.E. α-Synuclein Membrane Interactions and Lipid Specificity *. Journal of Biological Chemistry 2000, 275, 34328–34334. [Google Scholar] [CrossRef]
- Davidson, W.S.; Jonas, A.; Clayton, D.F.; George, J.M. Stabilization of α-Synuclein Secondary Structure upon Binding to Synthetic Membranes *. Journal of Biological Chemistry 1998, 273, 9443–9449. [Google Scholar] [CrossRef]
- Ray, S.; Singh, N.; Kumar, R.; Patel, K.; Pandey, S.; Datta, D.; Mahato, J.; Panigrahi, R.; Navalkar, A.; Mehra, S.; et al. alpha-Synuclein aggregation nucleates through liquid-liquid phase separation. Nat Chem 2020, 12, 705–716. [Google Scholar] [CrossRef]
- Piroska, L.; Fenyi, A.; Thomas, S.; Plamont, M.A.; Redeker, V.; Melki, R.; Gueroui, Z. alpha-Synuclein liquid condensates fuel fibrillar alpha-synuclein growth. Sci Adv 2023, 9, eadg5663. [Google Scholar] [CrossRef]
- Rodriguez, L.C.; Foressi, N.N.; Celej, M.S. Liquid-liquid phase separation of tau and alpha-synuclein: A new pathway of overlapping neuropathologies. Biochem Biophys Res Commun 2024, 741, 151053. [Google Scholar] [CrossRef]
- Ruiz-Ortega, E.D.; Wilkaniec, A.; Adamczyk, A. Liquid-liquid phase separation and conformational strains of alpha-Synuclein: implications for Parkinson's disease pathogenesis. Front Mol Neurosci 2024, 17, 1494218. [Google Scholar] [CrossRef] [PubMed]
- Sternke-Hoffmann, R.; Sun, X.; Menzel, A.; Pinto, M.D.S.; Venclovaite, U.; Wordehoff, M.; Hoyer, W.; Zheng, W.; Luo, J. Phase Separation and Aggregation of alpha-Synuclein Diverge at Different Salt Conditions. Adv Sci (Weinh) 2024, 11, e2308279. [Google Scholar] [CrossRef] [PubMed]
- Winner, B.; Jappelli, R.; Maji, S.K.; Desplats, P.; Boyer, L.; Aigner, S.; Hetzer, C.; Loher, T.; Vilar, M.; Campioni, S.; et al. In Vivo Demonstration That A-Synuclein Oligomers Are Toxic. Proceedings of the National Academy of Sciences 2011, 108, 4194–4199. [Google Scholar] [CrossRef] [PubMed]
- Breydo, L.; Wu, J.; Uversky, V.N. A-Synuclein Misfolding and Parkinson's Disease. Biochimica Et Biophysica Acta (Bba) - Molecular Basis of Disease 2012, 1822, 261–285. [Google Scholar] [CrossRef]
- Maltsev, A.S.; Ying, J.; Bax, A. Impact of N-Terminal Acetylation of A-Synuclein on Its Random Coil and Lipid Binding Properties. Biochemistry 2012, 51, 5004–5013. [Google Scholar] [CrossRef]
- Picca, A.; Guerra, F.; Calvani, R.; Romano, R.; Coelho-Júnior, H.J.; Bucci, C.; Marzetti, E. Mitochondrial Dysfunction, Protein Misfolding and Neuroinflammation in Parkinson's Disease: Roads to Biomarker Discovery. Biomolecules 2021, 11. [Google Scholar] [CrossRef]
- Guo, J.; Covell, D.J.; Daniels, J.P.; Iba, M.; Stieber, A.; Zhang, B.; Riddle, D.M.; Kwong, L.K.; Xu, Y.; Trojanowski, J.Q.; et al. Distinct A-Synuclein Strains Differentially Promote Tau Inclusions in Neurons. Cell 2013, 154, 103–117. [Google Scholar] [CrossRef]
- Foressi, N.N.; Rodriguez, L.C.; Celej, M.S. Heterotypic liquid-liquid phase separation of tau and alpha-synuclein: Implications for overlapping neuropathologies. Biochim Biophys Acta Proteins Proteom 2023, 1871, 140950. [Google Scholar] [CrossRef]
- Parnetti, L.; Chiasserini, D.; Bellomo, G.; Giannandrea, D.; Carlo, C.D.; Qureshi, M.M.; Ardah, M.T.; Varghese, S.; Bonanni, L.; Borroni, B.; et al. Cerebrospinal Fluid Tau/A-synuclein Ratio in Parkinson's Disease and Degenerative Dementias. Movement Disorders 2011, 26, 1428–1435. [Google Scholar] [CrossRef]
- Pajkos, M.; Clerc, I.; Zanon, C.; Bernado, P.; Cortes, J. AFflecto: A web server to generate conformational ensembles of flexible proteins from AlphaFold models. J Mol Biol 2025, 437, 169003. [Google Scholar] [CrossRef]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: visual molecular dynamics. J Mol Graph 1996, 14, 33-38, 27-38. [Google Scholar] [CrossRef] [PubMed]
- Latimer, C.S.; Liachko, N.F. Tau and TDP-43 synergy: a novel therapeutic target for sporadic late-onset Alzheimer's disease. Geroscience 2021, 43, 1627–1634. [Google Scholar] [CrossRef] [PubMed]
- Ratti, A.; Buratti, E. Physiological functions and pathobiology of TDP-43 and FUS/TLS proteins. J Neurochem 2016, 138 Suppl 1, 95–111. [Google Scholar] [CrossRef] [PubMed]
- Ling, S.C.; Polymenidou, M.; Cleveland, D.W. Converging mechanisms in ALS and FTD: disrupted RNA and protein homeostasis. Neuron 2013, 79, 416–438. [Google Scholar] [CrossRef]
- Nasir, A.R.; Delpirou Nouh, C. TDP-43-proteinopathy at the crossroads of tauopathy: on copathology and current and prospective biomarkers. Front Cell Neurosci 2025, 19, 1671419. [Google Scholar] [CrossRef]
- Wright, P.E.; Dyson, H.J. Intrinsically Disordered Proteins in Cellular Signalling and Regulation. Nature Reviews Molecular Cell Biology 2014, 16, 18–29. [Google Scholar] [CrossRef]
- McAleese, K.E.; Walker, L.; Erskine, D. TDP-43 pathology in Alzheimer’s disease, dementia with Lewy bodies and aging. Brain Pathology 2017, 27, 472–479. [Google Scholar] [CrossRef]
- Santamaria, N.; Alhothali, M.; Alfonso, M.H.; Breydo, L.; Uversky, V.N. Intrinsic disorder in proteins involved in amyotrophic lateral sclerosis. Cellular and molecular life sciences: CMLS 2017, 74, 1297–1318. [Google Scholar] [CrossRef]
- Loughlin, F.E.; Wilce, J.A. TDP-43 and FUS-structural insights into RNA recognition and self-association. Curr Opin Struct Biol 2019, 59, 134–142. [Google Scholar] [CrossRef]
- Zhang, Y.P.; Kedia, S.; Klenerman, D. Rethinking neurodegeneration through a co-proteinopathy lens. Trends Neurosci 2025, 48, 952–963. [Google Scholar] [CrossRef]
- Igaz, L.M.; Kwong, L.K.; Lee, E.B.; Chen-Plotkin, A.; Swanson, E.A.; Unger, T.L.; Malunda, J.; Xu, Y.; Winton, M.J.; Trojanowski, J.Q.; et al. Dysregulation of the ALS-associated Gene TDP-43 Leads to Neuronal Death and Degeneration in Mice. Journal of Clinical Investigation 2011, 121, 726–738. [Google Scholar] [CrossRef] [PubMed]
- Lang, R.; Hodgson, R.E.; Shelkovnikova, T.A. TDP-43 in nuclear condensates: where, how, and why. Biochem Soc Trans 2024, 52, 1809–1825. [Google Scholar] [CrossRef] [PubMed]
- Conicella, A.E.; Zerze, G.H.; Mittal, J.; Fawzi, N.L. ALS Mutations Disrupt Phase Separation Mediated by alpha-Helical Structure in the TDP-43 Low-Complexity C-Terminal Domain. Structure 2016, 24, 1537–1549. [Google Scholar] [CrossRef]
- Gopal, P.P.; Nirschl, J.J.; Klinman, E.; Holzbaur, E.L. Amyotrophic lateral sclerosis-linked mutations increase the viscosity of liquid-like TDP-43 RNP granules in neurons. Proc Natl Acad Sci U S A 2017, 114, E2466–E2475. [Google Scholar] [CrossRef] [PubMed]
- Li, H.R.; Chiang, W.C.; Chou, P.C.; Wang, W.J.; Huang, J.R. TAR DNA-binding protein 43 (TDP-43) liquid-liquid phase separation is mediated by just a few aromatic residues. J Biol Chem 2018, 293, 6090–6098. [Google Scholar] [CrossRef]
- Babinchak, W.M.; Haider, R.; Dumm, B.K.; Sarkar, P.; Surewicz, K.; Choi, J.K.; Surewicz, W.K. The role of liquid-liquid phase separation in aggregation of the TDP-43 low-complexity domain. J Biol Chem 2019, 294, 6306–6317. [Google Scholar] [CrossRef]
- Mann, J.R.; Gleixner, A.M.; Mauna, J.C.; Gomes, E.; DeChellis-Marks, M.R.; Needham, P.G.; Copley, K.E.; Hurtle, B.; Portz, B.; Pyles, N.J.; et al. RNA Binding Antagonizes Neurotoxic Phase Transitions of TDP-43. Neuron 2019, 102, 321–338 e328. [Google Scholar] [CrossRef]
- Wang, C.; Duan, Y.; Duan, G.; Wang, Q.; Zhang, K.; Deng, X.; Qian, B.; Gu, J.; Ma, Z.; Zhang, S.; et al. Stress Induces Dynamic, Cytotoxicity-Antagonizing TDP-43 Nuclear Bodies via Paraspeckle LncRNA NEAT1-Mediated Liquid-Liquid Phase Separation. Mol Cell 2020, 79, 443–458 e447. [Google Scholar] [CrossRef]
- Webber, C.J.; Lei, S.E.; Wolozin, B. The pathophysiology of neurodegenerative disease: Disturbing the balance between phase separation and irreversible aggregation. Prog Mol Biol Transl Sci 2020, 174, 187–223. [Google Scholar] [CrossRef]
- Pakravan, D.; Michiels, E.; Bratek-Skicki, A.; De Decker, M.; Van Lindt, J.; Alsteens, D.; Derclaye, S.; Van Damme, P.; Schymkowitz, J.; Rousseau, F.; et al. Liquid-Liquid Phase Separation Enhances TDP-43 LCD Aggregation but Delays Seeded Aggregation. Biomolecules 2021, 11. [Google Scholar] [CrossRef]
- Bhopatkar, A.A.; Dhakal, S.; Abernathy, H.G.; Morgan, S.E.; Rangachari, V. Charge and redox states modulate granulin-TDP-43 coacervation toward phase separation or aggregation. Biophys J 2022, 121, 2107–2126. [Google Scholar] [CrossRef] [PubMed]
- Staderini, T.; Bigi, A.; Mongiello, D.; Cecchi, C.; Chiti, F. Biophysical characterization of full-length TAR DNA-binding protein (TDP-43) phase separation. Protein Sci 2022, 31, e4509. [Google Scholar] [CrossRef] [PubMed]
- Song, J. Molecular Mechanisms of Phase Separation and Amyloidosis of ALS/FTD-linked FUS and TDP-43. Aging Dis 2024, 15, 2084–2112. [Google Scholar] [CrossRef] [PubMed]
- Sergeeva, O.S.; Neklesova, M.V.; Reushev, V.A.; Artemov, A.V.; Kuznetsova, I.M.; Turoverov, K.K.; Uversky, V.N.; Fonin, A.V. On the potential roles of TDP-43 in the formation of membraneless organelles and their transformation into toxic aggregates. Biochem Biophys Res Commun 2025, 788, 152808. [Google Scholar] [CrossRef]
- Mosna, S.; Dormann, D. TDP-43 Phosphorylation: Pathological Modification or Protective Factor Antagonizing TDP-43 Aggregation in Neurodegenerative Diseases? Bioessays 2026, 48, e70084. [Google Scholar] [CrossRef]
- Cairns, N.J.; Neumann, M.; Bigio, E.H.; Holm, I.E.; Troost, D.; Hatanpaa, K.J.; Foong, C.; White, C.L.; Schneider, J.A.; Kretzschmar, H.A.; et al. TDP-43 in Familial and Sporadic Frontotemporal Lobar Degeneration With Ubiquitin Inclusions. American Journal of Pathology 2007, 171, 227–240. [Google Scholar] [CrossRef]
- Robinson, J.; Lee, E.B.; Xie, S.X.; Rennert, L.; Suh, E.; Bredenberg, C.; Caswell, C.; Deerlin, V.M.V.; Yan, N.; Yousef, A.H.; et al. Neurodegenerative Disease Concomitant Proteinopathies Are Prevalent, Age-Related and APOE4-associated. Brain 2018, 141, 2181–2193. [Google Scholar] [CrossRef]
- Benajiba, L.; Ber, I.L.; Camuzat, A.; Lacoste, M.; Thomas-Antérion, C.; Couratier, P.; Legallic, S.; Salachas, F.; Hannequin, D.; Décousus, M.; et al. Annals of Neurology 2009, 65, 470–473. [CrossRef]
- Josephs, K.A.; Murray, M.E.; Tosakulwong, N. TDP-43 pathology in aging and Alzheimer's disease. Lancet Neurology 2016, 15, 129–138. [Google Scholar] [CrossRef]
- Nelson, P.T.; Trojanowski, J.Q.; Jack, C.R.; Boyle, P.A.; Arfanakis, K.; Rademakers, R.; Alafuzoff, I.; Attems, J.; Brayne, C.; Coyle-Gilchrist, I.T.S.; et al. Limbic-predominant age-related TDP-43 encephalopathy (LATE). Brain 2019, 142, 1503–1527. [Google Scholar] [CrossRef]
- Neumann, M.; Mackenzie, I.R.A. Review: Neuropathology of TDP-43 proteinopathies. Neuropathology and Applied Neurobiology 2019, 45, 521–537. [Google Scholar] [CrossRef]
- Carey, J.L.; Guo, L. Liquid-Liquid Phase Separation of TDP-43 and FUS in Physiology and Pathology of Neurodegenerative Diseases. Frontiers in Molecular Biosciences 2022, 9. [Google Scholar] [CrossRef] [PubMed]
- Fujii, R.; Takumi, T. TLS facilitates transport of mRNA encoding an actin-stabilizing protein to dendritic spines. J Cell Sci 2005, 118, 5755–5765. [Google Scholar] [CrossRef] [PubMed]
- Bonucci, A.; Murrali, M.G.; Banci, L.; Pierattelli, R. A combined NMR and EPR investigation on the effect of the disordered RGG regions in the structure and the activity of the RRM domain of FUS. Sci Rep 2020, 10, 20956. [Google Scholar] [CrossRef]
- Lagier-Tourenne, C.; Cleveland, D.W. Rethinking ALS: The FUS about TDP-43. Cell 2009, 136, 1001–1004. [Google Scholar] [CrossRef]
- Kwiatkowski, T.J.; Bosco, D.A.; Leclerc, A.; Tamrazian, E.; Vanderburg, C.; Russ, C.; Davis, A.; Gilchrist, J.M.; Kasarskis, E.J.; Munsat, T.L.; et al. Mutations in The. Science 2009, 323, 1205–1208. [Google Scholar] [CrossRef]
- Kang, J.; Lim, L.; Lu, Y.; Song, J. A Unified Mechanism for LLPS of ALS/FTLD-causing FUS as Well as Its Modulation by ATP and Oligonucleic Acids. Plos Biology 2019, 17, e3000327. [Google Scholar] [CrossRef]
- Mehdiabadi, M. Modeling Intrinsically Disordered Regions From AlphaFold2 to AlphaFold3. Protein Science 2025, 35. [Google Scholar] [CrossRef]
- Lagier-Tourenne, C.; Cleveland, D.W. Rethinking ALS: The FUS/TDP-43 paradigm. Cold Spring Harbor Perspectives in Biology 2009, 1, a003689. [Google Scholar] [CrossRef]
- Neumann, M.; Rademakers, R.; Roeber, S.; Baker, M.; Kretzschmar, H.A.; Mackenzie, I.R.A. A New Subtype of Frontotemporal Lobar Degeneration With FUS Pathology. Brain 2009, 132, 2922–2931. [Google Scholar] [CrossRef]
- Molliex, A.; Temirov, J.; Lee, J.; Coughlin, M.; Kanagaraj, A.; Kim, H.J.; Mittag, T.; Taylor, J.P. Phase Separation by Low Complexity Domains Promotes Stress Granule Assembly and Drives Pathological Fibrillization. Cell 2015, 163, 123–133. [Google Scholar] [CrossRef] [PubMed]
- Hallegger, M.; Chakrabarti, A.M.; Lee, F.C.Y.; Lee, B.L.; Amalietti, A.G.; Odeh, H.M.; Copley, K.E.; Rubien, J.D.; Portz, B.; Kuret, K.; et al. TDP-43 condensation properties specify its RNA-binding and regulatory repertoire. Cell 2021, 184, 4680–4696.e4622. [Google Scholar] [CrossRef]
- Soranno, A.; Koenig, I.; Borgia, M.B.; Hofmann, H.; Zosel, F.; Nettels, D.; Schuler, B. Single-Molecule Spectroscopy Reveals Polymer Effects of Disordered Proteins in Crowded Environments. Proceedings of the National Academy of Sciences 2014, 111, 4874–4879. [Google Scholar] [CrossRef] [PubMed]
- Burke, K.A.; Janke, A.M.; Rhine, C.L.; Fawzi, N.L. Residue-by-Residue View of In Vitro FUS Granules that Bind the C-Terminal Domain of RNA Polymerase II. Mol Cell 2015, 60, 231–241. [Google Scholar] [CrossRef] [PubMed]
- Monahan, Z.; Ryan, V.H.; Janke, A.M.; Burke, K.A.; Rhoads, S.N.; Zerze, G.H.; O'Meally, R.; Dignon, G.L.; Conicella, A.E.; Zheng, W.; et al. Phosphorylation of the FUS low-complexity domain disrupts phase separation, aggregation, and toxicity. Embo j 2017, 36, 2951–2967. [Google Scholar] [CrossRef]
- Qamar, S.; Wang, G.; Randle, S.J.; Ruggeri, F.S.; Varela, J.A.; Lin, J.Q.; Phillips, E.C.; Miyashita, A.; Williams, D.; Strohl, F.; et al. FUS Phase Separation Is Modulated by a Molecular Chaperone and Methylation of Arginine Cation-pi Interactions. Cell 2018, 173, 720–734 e715. [Google Scholar] [CrossRef]
- Murthy, A.C.; Dignon, G.L.; Kan, Y.; Zerze, G.H.; Parekh, S.H.; Mittal, J.; Fawzi, N.L. Molecular interactions underlying liquid-liquid phase separation of the FUS low-complexity domain. Nat Struct Mol Biol 2019, 26, 637–648. [Google Scholar] [CrossRef]
- Boczek, E.E.; Fursch, J.; Niedermeier, M.L.; Jawerth, L.; Jahnel, M.; Ruer-Gruss, M.; Kammer, K.M.; Heid, P.; Mediani, L.; Wang, J.; et al. HspB8 prevents aberrant phase transitions of FUS by chaperoning its folded RNA-binding domain. Elife 2021, 10. [Google Scholar] [CrossRef]
- Davis, R.B.; Kaur, T.; Moosa, M.M.; Banerjee, P.R. FUS oncofusion protein condensates recruit mSWI/SNF chromatin remodeler via heterotypic interactions between prion-like domains. Protein Sci 2021, 30, 1454–1466. [Google Scholar] [CrossRef]
- Joshi, A.; Walimbe, A.; Avni, A.; Rai, S.K.; Arora, L.; Sarkar, S.; Mukhopadhyay, S. Single-molecule FRET unmasks structural subpopulations and crucial molecular events during FUS low-complexity domain phase separation. Nat Commun 2023, 14, 7331. [Google Scholar] [CrossRef]
- Esteban-Hofer, L.; Emmanouilidis, L.; Yulikov, M.; Allain, F.H.; Jeschke, G. Ensemble structure of the N-terminal domain (1-267) of FUS in a biomolecular condensate. Biophys J 2024, 123, 538–554. [Google Scholar] [CrossRef]
- Thirumalai, D.; Kumar, A.; Chakraborty, D.; Straub, J.E.; Mugnai, M.L. Conformational fluctuations and phases in fused in sarcoma (FUS) low-complexity domain. Biopolymers 2024, 115, e23558. [Google Scholar] [CrossRef]
- Miller, A.; Toprakcioglu, Z.; Qamar, S.; St George-Hyslop, P.; Ruggeri, F.S.; Knowles, T.P.J.; Vendruscolo, M. Nanoscale profiling of evolving intermolecular interactions in ageing FUS condensates. Commun Chem 2025, 8, 284. [Google Scholar] [CrossRef]
- Weng, S.L.; Mohanty, P.; Mittal, J. Elucidation of the Molecular Interaction Network Underlying Full-Length FUS Conformational Transitions and Its Phase Separation Using Atomistic Simulations. J Phys Chem B 2025, 129, 8843–8857. [Google Scholar] [CrossRef] [PubMed]
- Zheng, T.; Sojitra, K.A.; Cummings, S.; Chen, Q.; Mohanty, P.; Mittal, J.; Fawzi, N.L. RNA modulates FUS condensate assembly, dynamics, and aggregation through diverse molecular contacts. bioRxiv 2025. [Google Scholar] [CrossRef]
- Almalki, S.; Salama, M.; Taylor, M.J.; Ahmed, Z.; Tuxworth, R.I. FUS-related amyotrophic lateral sclerosis-frontotemporal dementia and links to the DNA damage response: a systematic review. Front Mol Neurosci 2025, 18, 1671910. [Google Scholar] [CrossRef] [PubMed]
- Selkoe, D.J.; Hardy, J. The Amyloid Hypothesis of Alzheimer’s Disease at 25 Years. EMBO Journal 2016, 35, 595–608. [Google Scholar] [CrossRef]
- Lashuel, H.A. Structural biology of alpha-synuclein and its aggregation. Nature Reviews Molecular Cell Biology 2020. [Google Scholar]
- Neumann, M.; Sampathu, D.M.; Kwong, L.K.; Truax, A.C.; Micsenyi, M.C.; Chou, T.T.; Bruce, J.; Schuck, T.; Grossman, M.; Clark, C.M.; et al. Ubiquitinated TDP-43 in Frontotemporal Lobar Degeneration and Amyotrophic Lateral Sclerosis. Science 2006, 314, 130–133. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Kirsch, R.; Koutrouli, M.; Nastou, K.; Mehryary, F.; Hachilif, R.; Gable, A.L.; Fang, T.; Doncheva, N.T.; Pyysalo, S.; et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res 2023, 51, D638–D646. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Nastou, K.; Koutrouli, M.; Kirsch, R.; Mehryary, F.; Hachilif, R.; Hu, D.; Peluso, M.E.; Huang, Q.; Fang, T.; et al. The STRING database in 2025: protein networks with directionality of regulation. Nucleic Acids Res 2025, 53, D730–D737. [Google Scholar] [CrossRef] [PubMed]
- von Mering, C.; Huynen, M.; Jaeggi, D.; Schmidt, S.; Bork, P.; Snel, B. STRING: a database of predicted functional associations between proteins. Nucleic Acids Res 2003, 31, 258–261. [Google Scholar] [CrossRef] [PubMed]
- Rajagopalan, K.; Mooney, S.M.; Parekh, N.; Getzenberg, R.H.; Kulkarni, P. A majority of the cancer/testis antigens are intrinsically disordered proteins. J Cell Biochem 2011, 112, 3256–3267. [Google Scholar] [CrossRef]
- Uversky, V.N. Analyzing IDPs in interactomes. In Intrinsically Disordered Proteins;Volume Methods in Molecular Biology; Kragelund, B.B., Skriver, K., Eds.; Humana New York, NY, 2020; pp. 895–945. [Google Scholar]
- Mohammed, A.S.; Uversky, V.N. Intrinsic Disorder as a Natural Preservative: High Levels of Intrinsic Disorder in Proteins Found in the 2600-Year-Old Human Brain. Biology (Basel) 2022, 11. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Xue, B.; Jones, W.T.; Rikkerink, E.; Dunker, A.K.; Uversky, V.N. A functionally required unfoldome from the plant kingdom: intrinsically disordered N-terminal domains of GRAS proteins are involved in molecular recognition during plant development. Plant Mol Biol 2011, 77, 205–223. [Google Scholar] [CrossRef]
- Xue, B.; Oldfield, C.J.; Van, Y.Y.; Dunker, A.K.; Uversky, V.N. Protein intrinsic disorder and induced pluripotent stem cells. Mol Biosyst 2012, 8, 134–150. [Google Scholar] [CrossRef]
- Mohan, A.; Sullivan, W.J., Jr.; Radivojac, P.; Dunker, A.K.; Uversky, V.N. Intrinsic disorder in pathogenic and non-pathogenic microbes: discovering and analyzing the unfoldomes of early-branching eukaryotes. Mol Biosyst 2008, 4, 328–340. [Google Scholar] [CrossRef]
- Huang, F.; Oldfield, C.; Meng, J.; Hsu, W.L.; Xue, B.; Uversky, V.N.; Romero, P.; Dunker, A.K. Subclassifying disordered proteins by the CH-CDF plot method. Pac Symp Biocomput 2012, 128–139. [Google Scholar]
- Sade, D.; Shaham-Niv, S.; Arnon, Z.A.; Tavassoly, O.; Gazit, E. Seeding of Proteins Into Amyloid Structures by Metabolite Assemblies May Clarify Certain Unexplained Epidemiological Associations. Open Biology 2018, 8, 170229. [Google Scholar] [CrossRef]
- Tsoi, P.S.; Quan, M.D.; Ferreon, J.C.; Ferreon, A.C.M. Aggregation of Disordered Proteins Associated with Neurodegeneration. International Journal of Molecular Sciences 2023, 24, 3380. [Google Scholar] [CrossRef]
- Tsoi, P.S.; Quan, M.D.; Ferreon, J.C.; Ferreon, A.C.M. Aggregation of Disordered Proteins Associated With Neurodegeneration. International Journal of Molecular Sciences 2023, 24, 3380. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Zhang, L.; Dai, T.; Qin, Z.; Lu, H.; Zhang, L.; Zhou, F. Liquid–liquid Phase Separation in Human Health and Diseases. Signal Transduction and Targeted Therapy 2021, 6. [Google Scholar] [CrossRef] [PubMed]
- Alberti, S.; Dormann, D. Liquid–Liquid Phase Separation in Disease. Annual Review of Genetics 2019, 53, 171–194. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Zhou, W. Phase separation as a new form of regulation in innate immunity. Mol Cell 2024, 84, 2410–2422. [Google Scholar] [CrossRef]
- Xie, C.C.; Wang, T.; Liu, X.R.; Wang, Y.; Dang, Q.; Ding, T.; Xu, J.Q.; Yu, X.J.; Lin, H.; Xu, X.W.; et al. Liquid-Liquid Phase Separation in Major Hallmarks of Cancer. Cell Prolif 2026, 59, e70122. [Google Scholar] [CrossRef]
- Thakur, D.K.; Padole, S.; Sarkar, T.; Arumugam, S.; Chattopadhyay, S. Liquid-Liquid Phase Separation: Mechanisms, Roles, and Implications in Cellular Function and Disease. FASEB Bioadv 2025, 7, e70054. [Google Scholar] [CrossRef]
- Hyman, A.A.; Weber, C.A.; Jülicher, F. Liquid-Liquid Phase Separation in Biology. Annual Review of Cell and Developmental Biology 2014, 30, 39–58. [Google Scholar] [CrossRef]
- Wang, J.; Choi, J.M.; Holehouse, A.S.; Lee, H.O.; Zhang, X.; Jahnel, M.; Maharana, S.; Lemaitre, R.; Pozniakovsky, A.; Drechsel, D.; et al. A Molecular Grammar Governing the Driving Forces for Phase Separation of Prion-Like RNA Binding Proteins. Cell 2018, 174, 688–699.e616. [Google Scholar] [CrossRef]
- Korsak, M.; Kozyreva, T. Beta Amyloid Hallmarks: From Intrinsically Disordered Proteins to Alzheimer's Disease. Adv Exp Med Biol 2015, 870, 401–421. [Google Scholar] [CrossRef]
- Coskuner, O.; Uversky, V.N. Intrinsically disordered proteins in various hypotheses on the pathogenesis of Alzheimer's and Parkinson's diseases. Prog Mol Biol Transl Sci 2019, 166, 145–223. [Google Scholar] [CrossRef]
- Nguyen, P.H.; Derreumaux, P. Structures of the intrinsically disordered Abeta, tau and alpha-synuclein proteins in aqueous solution from computer simulations. Biophys Chem 2020, 264, 106421. [Google Scholar] [CrossRef] [PubMed]
- Tsoi, P.S.; Quan, M.D.; Ferreon, J.C.; Ferreon, A.C.M. Aggregation of Disordered Proteins Associated with Neurodegeneration. Int J Mol Sci 2023, 24. [Google Scholar] [CrossRef] [PubMed]
- Skrabana, R.; Skrabanova, M.; Csokova, N.; Sevcik, J.; Novak, M. Intrinsically disordered tau protein in Alzheimer's tangles: a coincidence or a rule? Bratisl Lek Listy 2006, 107, 354–358. [Google Scholar] [PubMed]
- Sabbagh, J.J.; Dickey, C.A. The Metamorphic Nature of the Tau Protein: Dynamic Flexibility Comes at a Cost. Front Neurosci 2016, 10, 3. [Google Scholar] [CrossRef]
- Cehlar, O.; Bagarova, O.; Hornakova, L.; Skrabana, R. The structure of the unstructured: mosaic of tau protein linear motifs obtained by high-resolution techniques and molecular simulation. Gen Physiol Biophys 2021, 40, 479–493. [Google Scholar] [CrossRef]
- Chinnathambi, S.; Velmurugan, G.; Chandrashekar, M. Tau protein structure and dynamics. Adv Protein Chem Struct Biol 2025, 147, 241–258. [Google Scholar] [CrossRef]
- Sandeep, K.; Savastano, A.; Singh, P.; Mukhopadhyay, S.; Zweckstetter, M. Liquid–liquid Phase Separation of Tau: From Molecular Biophysics to Physiology and Disease. Protein Science 2021, 30, 1294–1314. [Google Scholar] [CrossRef]
- Ono, K.; Takahashi, R.; Ikeda, T.; Yamada, M. Cross-seeding Effects of Amyloid Β-protein and A-synuclein. Journal of Neurochemistry 2012, 122, 883–890. [Google Scholar] [CrossRef]
- Sengupta, U.; Kayed, R. Amyloid β, Tau, and α-Synuclein aggregates in the pathogenesis, prognosis, and therapeutics for neurodegenerative diseases. Prog Neurobiol 2022, 214, 102270. [Google Scholar] [CrossRef]
- Vasconcelos, B.; Stancu, I.C.; Buist, A.; Bird, M.; Wang, P.; Vanoosthuyse, A.; Kolen, K.V.; Verheyen, A.; Kienlen-Campard, P.; Octave, J.N.; et al. Heterotypic Seeding of Tau Fibrillization by Pre-Aggregated Abeta Provides Potent Seeds for Prion-Like Seeding and Propagation of Tau-Pathology in Vivo. Acta Neuropathologica 2016, 131, 549–569. [Google Scholar] [CrossRef]
- Moreno-González, I.; Edwards, G.A.; Salvadores, N.; Shahnawaz, M.; Díaz-Espinoza, R.; Soto, C. Molecular Interaction Between Type 2 Diabetes and Alzheimer’s Disease Through Cross-Seeding of Protein Misfolding. Molecular Psychiatry 2017, 22, 1327–1334. [Google Scholar] [CrossRef] [PubMed]
- Rahimi, J.; Kovács, G.G. Prevalence of Mixed Pathologies in the Aging Brain. Alzheimer S Research & Therapy 2014, 6. [Google Scholar] [CrossRef]
- Katorcha, E.; Makarava, N.; Lee, Y.J.; Lindberg, I.; Monteiro, M.J.; Kovács, G.G.; Baskakov, I.V. Cross-Seeding of Prions by Aggregated A-Synuclein Leads to Transmissible Spongiform Encephalopathy. Plos Pathogens 2017, 13, e1006563. [Google Scholar] [CrossRef] [PubMed]
- Cukalevski, R.; Yang, X.; Meisl, G.; Weininger, U.; Bernfur, K.; Frohm, B.; Knowles, T.P.J.; Linse, S. The Aβ40 and Aβ42 Peptides Self-Assemble Into Separate Homomolecular Fibrils in Binary Mixtures but Cross-React During Primary Nucleation. Chemical Science 2015, 6, 4215–4233. [Google Scholar] [CrossRef]
- Hipp, M.S.; Kasturi, P.; Hartl, F.U. The Proteostasis Network and Its Decline in Ageing. Nature Reviews Molecular Cell Biology 2019, 20, 421–435. [Google Scholar] [CrossRef]
- Hipp, M.S.; Park, S.H.; Hartl, F.U. Proteostasis impairment in protein-misfolding and aggregation diseases. Nature Reviews Molecular Cell Biology 2014. [Google Scholar] [CrossRef]
- Mukherjee, A.; Morales-Scheihing, D.; Butler, P.C.; Soto, C. Type 2 Diabetes as a Protein Misfolding Disease. Trends in Molecular Medicine 2015, 21, 439–449. [Google Scholar] [CrossRef]
- Tyedmers, J.; Mogk, A.; Bukau, B. Cellular Strategies for Controlling Protein Aggregation. Nature Reviews Molecular Cell Biology 2010, 11, 777–788. [Google Scholar] [CrossRef]
- Patel, A.; Lee, H.O.; Jawerth, L.; Maharana, S.; Jahnel, M.; Hein, M.Y.; Stoynov, S.; Mahamid, J.; Saha, S.; Franzmann, T.M.; et al. A Liquid-to-Solid Phase Transition of the ALS Protein FUS Accelerated by Disease Mutation. Cell 2015, 162, 1066–1077. [Google Scholar] [CrossRef]
- Babinchak, W.M.; Surewicz, W.K. Liquid-Liquid Phase Separation and Its Mechanistic Role in Pathological Protein Aggregation. J Mol Biol 2020, 432, 1910–1925. [Google Scholar] [CrossRef]
- Balchin, D.; Hayer-Hartl, M.; Hartl, F.U. In Vivo Aspects of Protein Folding and Quality Control. Science 2016, 353. [Google Scholar] [CrossRef]
- Powers, E.T.; Morimoto, R.I.; Dillin, A.; Kelly, J.W.; Balch, W.E. Biological and Chemical Approaches to Diseases of Proteostasis Deficiency. Annual Review of Biochemistry 2009, 78, 959–991. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Selvaratnam, T.; Lee, J.C.T.; Chao, Y.; Tan, E.K. Molecular Targets for Modulating the Protein Translation Vital to Proteostasis and Neuron Degeneration in Parkinson’s Disease. Translational Neurodegeneration 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Shorter, J. The Mammalian Disaggregase Machinery: Hsp110 Synergizes With Hsp70 and Hsp40 to Catalyze Protein Disaggregation and Reactivation in a Cell-Free System. Plos One 2011, 6, e26319. [Google Scholar] [CrossRef] [PubMed]
- Hipp, M.S.; Park, S.-H.; Hartl, F.U. Proteostasis Impairment in Protein-Misfolding and -Aggregation Diseases. Trends in Cell Biology 2014, 24, 506–514. [Google Scholar] [CrossRef]
- Uversky, V.N. Unusual biophysics of intrinsically disordered proteins. Biochim Biophys Acta 2013, 1834, 932–951. [Google Scholar] [CrossRef]
- Cheng, Y.; LeGall, T.; Oldfield, C.J.; Mueller, J.P.; Van, Y.Y.; Romero, P.; Cortese, M.S.; Uversky, V.N.; Dunker, A.K. Rational drug design via intrinsically disordered protein. Trends Biotechnol 2006, 24, 435–442. [Google Scholar] [CrossRef]
- Dunker, A.K.; Uversky, V.N. Drugs for 'protein clouds': targeting intrinsically disordered transcription factors. Curr Opin Pharmacol 2010, 10, 782–788. [Google Scholar] [CrossRef]
- Metallo, S.J. Intrinsically disordered proteins are potential drug targets. Curr Opin Chem Biol 2010, 14, 481–488. [Google Scholar] [CrossRef]
- Uversky, V.N. Intrinsically disordered proteins and novel strategies for drug discovery. Expert Opin Drug Discov 2012, 7, 475–488. [Google Scholar] [CrossRef]
- Hu, G.; Wu, Z.; Wang, K.; Uversky, V.N.; Kurgan, L. Untapped Potential of Disordered Proteins in Current Druggable Human Proteome. Curr Drug Targets 2016, 17, 1198–1205. [Google Scholar] [CrossRef]
- Uversky, V.N. How to drug a cloud? Targeting intrinsically disordered proteins. Pharmacol Rev 2024. [Google Scholar] [CrossRef] [PubMed]
- Cortese, M.S.; Uversky, V.N.; Dunker, A.K. Intrinsic disorder in scaffold proteins: getting more from less. Prog Biophys Mol Biol 2008, 98, 85–106. [Google Scholar] [CrossRef] [PubMed]
- Buday, L.; Tompa, P. Functional classification of scaffold proteins and related molecules. FEBS J 2010, 277, 4348–4355. [Google Scholar] [CrossRef] [PubMed]
- Barbar, E.; Nyarko, A. Polybivalency and disordered proteins in ordering macromolecular assemblies. Semin Cell Dev Biol 2015, 37, 20–25. [Google Scholar] [CrossRef]
- Clark, S.A.; Jespersen, N.; Woodward, C.; Barbar, E. Multivalent IDP assemblies: Unique properties of LC8-associated, IDP duplex scaffolds. FEBS Lett 2015, 589, 2543–2551. [Google Scholar] [CrossRef]
- Patil, A.; Kinoshita, K.; Nakamura, H. Hub promiscuity in protein-protein interaction networks. Int J Mol Sci 2010, 11, 1930–1943. [Google Scholar] [CrossRef]
- Carels, N.; Sgariglia, D.; Junior, M.G.V.; Lima, C.R.; Carneiro, F.R.G.; Silva, G.F.D.; Silva, F.; Scardini, R.; Tuszynski, J.A.; Andrade, C.V.; et al. A Strategy Utilizing Protein-Protein Interaction Hubs for the Treatment of Cancer Diseases. Int J Mol Sci 2023, 24. [Google Scholar] [CrossRef]
- Iakoucheva, L.M.; Brown, C.J.; Lawson, J.D.; Obradovic, Z.; Dunker, A.K. Intrinsic disorder in cell-signaling and cancer-associated proteins. J. Mol. Biol. 2002, 323, 573–584. [Google Scholar] [CrossRef]
- Dunker, A.K.; Brown, C.J.; Lawson, J.D.; Iakoucheva, L.M.; Obradovic, Z. Intrinsic disorder and protein function. Biochemistry 2002, 41, 6573–6582. [Google Scholar] [CrossRef]
- Dyson, H.J.; Wright, P.E. Insights into the structure and dynamics of unfolded proteins from nuclear magnetic resonance. Adv Protein Chem 2002, 62, 311–340. [Google Scholar]
- Fink, A.L. Natively unfolded proteins. Curr Opin Struct Biol 2005, 15, 35–41. [Google Scholar] [CrossRef]
- Uversky, V.N.; Oldfield, C.J.; Dunker, A.K. Showing your ID: intrinsic disorder as an ID for recognition, regulation and cell signaling. J Mol Recognit 2005, 18, 343–384. [Google Scholar] [CrossRef] [PubMed]
- Oldfield, C.J.; Cheng, Y.; Cortese, M.S.; Romero, P.; Uversky, V.N.; Dunker, A.K. Coupled folding and binding with alpha-helix-forming molecular recognition elements. Biochemistry 2005, 44, 12454–12470. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N. Multitude of binding modes attainable by intrinsically disordered proteins: a portrait gallery of disorder-based complexes. Chem Soc Rev 2011, 40, 1623–1634. [Google Scholar] [CrossRef] [PubMed]
- Hsu, W.L.; Oldfield, C.; Meng, J.; Huang, F.; Xue, B.; Uversky, V.N.; Romero, P.; Dunker, A.K. Intrinsic protein disorder and protein-protein interactions. Pac Symp Biocomput 2012, 116–127. [Google Scholar]
- Babu, M.M.; van der Lee, R.; de Groot, N.S.; Gsponer, J. Intrinsically disordered proteins: regulation and disease. Curr Opin Struct Biol 2011, 21, 432–440. [Google Scholar] [CrossRef]
- Kim, J.Y.; Chung, H.S. Disordered proteins follow diverse transition paths as they fold and bind to a partner. Science 2020, 368, 1253–1257. [Google Scholar] [CrossRef]
- Uversky, V.N. Dancing Protein Clouds: The Strange Biology and Chaotic Physics of Intrinsically Disordered Proteins. J Biol Chem 2016, 291, 6681–6688. [Google Scholar] [CrossRef]
- Uversky, V.N. Protein intrinsic disorder and structure-function continuum. Prog Mol Biol Transl Sci 2019, 166, 1–17. [Google Scholar] [CrossRef]
- Gupta, M.N.; Uversky, V.N. Protein structure-function continuum model: Emerging nexuses between specificity, evolution, and structure. Protein Sci 2024, 33, e4968. [Google Scholar] [CrossRef] [PubMed]
- Kulkarni, P.; Porter, L.; Chou, T.F.; Chong, S.; Chiti, F.; Schafer, J.W.; Mohanty, A.; Ramisetty, S.; Onuchic, J.N.; Tuite, M.; et al. Evolving concepts of the protein universe. iScience 2025, 28, 112012. [Google Scholar] [CrossRef] [PubMed]
- Uversky, V.N.; Dunker, A.K. Understanding protein non-folding. Biochim. Biophys. Acta 2010, 1804, 1231–1264. [Google Scholar] [CrossRef] [PubMed]
- Collins, M.O.; Yu, L.; Campuzano, I.; Grant, S.G.; Choudhary, J.S. Phosphoproteomic analysis of the mouse brain cytosol reveals a predominance of protein phosphorylation in regions of intrinsic sequence disorder. Molecular & cellular proteomics: MCP 2008, 7, 1331–1348. [Google Scholar] [CrossRef]
- Pejaver, V.; Hsu, W.L.; Xin, F.; Dunker, A.K.; Uversky, V.N.; Radivojac, P. The structural and functional signatures of proteins that undergo multiple events of post-translational modification. Protein Sci 2014. [Google Scholar] [CrossRef]
- Kurotani, A.; Tokmakov, A.A.; Kuroda, Y.; Fukami, Y.; Shinozaki, K.; Sakurai, T. Correlations between predicted protein disorder and post-translational modifications in plants. Bioinformatics 2014. [Google Scholar] [CrossRef]
- Romero, P.R.; Zaidi, S.; Fang, Y.Y.; Uversky, V.N.; Radivojac, P.; Oldfield, C.J.; Cortese, M.S.; Sickmeier, M.; LeGall, T.; Obradovic, Z.; et al. Alternative splicing in concert with protein intrinsic disorder enables increased functional diversity in multicellular organisms. Proc Natl Acad Sci U S A 2006, 103, 8390–8395. [Google Scholar] [CrossRef]
- Buljan, M.; Chalancon, G.; Dunker, A.K.; Bateman, A.; Balaji, S.; Fuxreiter, M.; Babu, M.M. Alternative splicing of intrinsically disordered regions and rewiring of protein interactions. Current opinion in structural biology 2013, 23, 443–450. [Google Scholar] [CrossRef]
- Buljan, M.; Chalancon, G.; Eustermann, S.; Wagner, G.P.; Fuxreiter, M.; Bateman, A.; Babu, M.M. Tissue-specific splicing of disordered segments that embed binding motifs rewires protein interaction networks. Molecular cell 2012, 46, 871–883. [Google Scholar] [CrossRef]
- Smith, L.M.; Kelleher, N.L.; Consortium for Top Down, P. Proteoform: a single term describing protein complexity. Nat Methods 2013, 10, 186–187. [Google Scholar] [CrossRef]
- Uversky, V.N. p53 Proteoforms and Intrinsic Disorder: An Illustration of the Protein Structure-Function Continuum Concept. Int J Mol Sci 2016, 17, 1874. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Zhao, S.; Dunker, A.K. Intrinsically Disordered Proteins Link Alternative Splicing and Post-translational Modifications to Complex Cell Signaling and Regulation. J Mol Biol 2018, 430, 2342–2359. [Google Scholar] [CrossRef] [PubMed]
- Ruan, H.; Sun, Q.; Zhang, W.; Liu, Y.; Lai, L. Targeting intrinsically disordered proteins at the edge of chaos. Drug Discov Today 2019, 24, 217–227. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.; Choi, S.; Kim, R.; Koh, J. Mechanisms of Macromolecular Interactions Mediated by Protein Intrinsic Disorder. Mol Cells 2020, 43, 899–908. [Google Scholar] [CrossRef]
- Morris, O.M.; Torpey, J.H.; Isaacson, R.L. Intrinsically disordered proteins: modes of binding with emphasis on disordered domains. Open Biol 2021, 11, 210222. [Google Scholar] [CrossRef]
- Bugge, K.; Sottini, A.; Ivanovic, M.T.; Buus, F.S.; Saar, D.; Fernandes, C.B.; Kocher, F.; Martinsen, J.H.; Schuler, B.; Best, R.B.; et al. Role of charges in a dynamic disordered complex between an IDP and a folded domain. Nat Commun 2025, 16, 3242. [Google Scholar] [CrossRef]
- Das, R.K.; Pappu, R.V. Conformations of Intrinsically Disordered Proteins Are Influenced by Linear Sequence Distributions of Oppositely Charged Residues. Proceedings of the National Academy of Sciences 2013, 110, 13392–13397. [Google Scholar] [CrossRef]
- Suskiewicz, M.J.; Sussman, J.L.; Silman, I.; Shaul, Y. Context-dependent resistance to proteolysis of intrinsically disordered proteins. Protein Sci 2011, 20, 1285–1297. [Google Scholar] [CrossRef]
- Manav, N.; Jit, B.P.; Kataria, B.; Sharma, A. Cellular and epigenetic perspective of protein stability and its implications in the biological system. Epigenomics 2024, 16, 879–900. [Google Scholar] [CrossRef]
- Conti, B.A.; Oppikofer, M. Biomolecular condensates: new opportunities for drug discovery and RNA therapeutics. Trends Pharmacol Sci 2022, 43, 820–837. [Google Scholar] [CrossRef]
- Igelmann, S.; Lessard, F.; Ferbeyre, G. Liquid-Liquid Phase Separation in Cancer Signaling, Metabolism and Anticancer Therapy. Cancers (Basel) 2022, 14. [Google Scholar] [CrossRef] [PubMed]
- Mitrea, D.M.; Mittasch, M.; Gomes, B.F.; Klein, I.A.; Murcko, M.A. Modulating biomolecular condensates: a novel approach to drug discovery. Nat Rev Drug Discov 2022, 21, 841–862. [Google Scholar] [CrossRef] [PubMed]
- Patel, A.; Mitrea, D.; Namasivayam, V.; Murcko, M.A.; Wagner, M.; Klein, I.A. Principles and functions of condensate modifying drugs. Front Mol Biosci 2022, 9, 1007744. [Google Scholar] [CrossRef] [PubMed]
- Ambadi Thody, S.; Clements, H.D.; Baniasadi, H.; Lyon, A.S.; Sigman, M.S.; Rosen, M.K. Small-molecule properties define partitioning into biomolecular condensates. Nat Chem 2024, 16, 1794–1802. [Google Scholar] [CrossRef] [PubMed]
- Qin, C.; Wang, Y.L.; Zheng, J.; Wan, X.B.; Fan, X.J. Current perspectives in drug targeting intrinsically disordered proteins and biomolecular condensates. BMC Biol 2025, 23, 118. [Google Scholar] [CrossRef]
- Deka, N.; Nag, N.; Tripathi, T. Intrinsically disordered proteins and liquid-liquid phase separation in drug discovery. Drug Discov Today 2026, 31, 104603. [Google Scholar] [CrossRef]
- Kim, D.H.; Han, K.H. Transient Secondary Structures as General Target-Binding Motifs in Intrinsically Disordered Proteins. Int J Mol Sci 2018, 19. [Google Scholar] [CrossRef]
- Wichapong, K.; Silvestre-Roig, C.; Braster, Q.; Schumski, A.; Soehnlein, O.; Nicolaes, G.A.F. Structure-based peptide design targeting intrinsically disordered proteins: Novel histone H4 and H2A peptidic inhibitors. Comput Struct Biotechnol J 2021, 19, 934–948. [Google Scholar] [CrossRef]
- Hadži, S.; Purič, S.; Zavrtanik, U.; Vranken, W. What stabilizes pre-folded structures in the intrinsically disordered α-helical binding motifs? bioRxiv 2022, 2022.2001. 2028.478151. [CrossRef]
- Fry, D.C. Targeting protein-protein interactions for drug discovery. Methods Mol Biol 2015, 1278, 93–106. [Google Scholar] [CrossRef]
- Modell, A.E.; Blosser, S.L.; Arora, P.S. Systematic Targeting of Protein-Protein Interactions. Trends Pharmacol Sci 2016, 37, 702–713. [Google Scholar] [CrossRef] [PubMed]
- Olah, J.; Szenasi, T.; Lehotzky, A.; Norris, V.; Ovadi, J. Challenges in Discovering Drugs That Target the Protein-Protein Interactions of Disordered Proteins. Int J Mol Sci 2022, 23. [Google Scholar] [CrossRef] [PubMed]
- Petri, L.; Abranyi-Balogh, P.; Vagrys, D.; Imre, T.; Varro, N.; Mandity, I.; Racz, A.; Wittner, L.; Toth, K.; Toth, E.Z.; et al. A covalent strategy to target intrinsically disordered proteins: Discovery of novel tau aggregation inhibitors. Eur J Med Chem 2022, 231, 114163. [Google Scholar] [CrossRef] [PubMed]
- Huang, F.; Han, X.; Xiao, X.; Zhou, J. Covalent Warheads Targeting Cysteine Residue: The Promising Approach in Drug Development. Molecules 2022, 27. [Google Scholar] [CrossRef]
- Sakamoto, K.M.; Kim, K.B.; Kumagai, A.; Mercurio, F.; Crews, C.M.; Deshaies, R.J. Protacs: Chimeric molecules that target proteins to the Skp1–Cullin–F box complex for ubiquitination and degradation. Proceedings of the National Academy of Sciences 2001, 98, 8554–8559. [Google Scholar] [CrossRef]
- Schapira, M.; Calabrese, M.F.; Bullock, A.N.; Crews, C.M. Targeted protein degradation: expanding the toolbox. Nat Rev Drug Discov 2019, 18, 949–963. [Google Scholar] [CrossRef]
- Banik, S.M.; Pedram, K.; Wisnovsky, S.; Ahn, G.; Riley, N.M.; Bertozzi, C.R. Lysosome-targeting chimaeras for degradation of extracellular proteins. Nature 2020, 584, 291–297. [Google Scholar] [CrossRef]
- Clift, D.; McEwan, W.A.; Labzin, L.I.; Konieczny, V.; Mogessie, B.; James, L.C.; Schuh, M. A Method for the Acute and Rapid Degradation of Endogenous Proteins. Cell 2017, 171, 1692–1706 e1618. [Google Scholar] [CrossRef]
- Li, Z.; Wang, C.; Wang, Z.; Zhu, C.; Li, J.; Sha, T.; Ma, L.; Gao, C.; Yang, Y.; Sun, Y.; et al. Allele-selective lowering of mutant HTT protein by HTT-LC3 linker compounds. Nature 2019, 575, 203–209. [Google Scholar] [CrossRef]
- He, H.; Zhou, C.; Chen, X. ATNC: versatile nanobody chimeras for autophagic degradation of intracellular unligandable and undruggable proteins. Journal of the American Chemical Society 2023, 145, 24785–24795. [Google Scholar] [CrossRef]
- Sun, X.; Wu, D.; Zhou, C.; Xia, S.; Chen, X. Degradation of intrinsically disordered proteins using proteolysis targeting nanobody conjugate. bioRxiv 2024, 2024.2011. 2027.622182.
- Pan, Y.; Wang, Y.; Gou, S. Proteolysis targeting chimera, molecular glue degrader and hydrophobic tag tethering degrader for targeted protein degradation: Mechanisms, strategies and application. Bioorg Chem 2025, 161, 108491. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Zhao, J.; Zhong, K.; Tong, A.; Jia, D. Targeted protein degradation: mechanisms, strategies and application. Signal Transduct Target Ther 2022, 7, 113. [Google Scholar] [CrossRef] [PubMed]
- Ganguly, D.; Otieno, S.; Waddell, B.; Iconaru, L.; Kriwacki, R.W.; Chen, J. Electrostatically accelerated coupled binding and folding of intrinsically disordered proteins. J Mol Biol 2012, 422, 674–684. [Google Scholar] [CrossRef]
- Saurabh, S.; Nadendla, K.; Purohit, S.S.; Sivakumar, P.M.; Cetinel, S. Fuzzy Drug Targets: Disordered Proteins in the Drug-Discovery Realm. ACS Omega 2023, 8, 9729–9747. [Google Scholar] [CrossRef]
- Lazar, T.; Connor, A.; DeLisle, C.F.; Burger, V.; Tompa, P. Targeting protein disorder: the next hurdle in drug discovery. Nat Rev Drug Discov 2025, 24, 743–763. [Google Scholar] [CrossRef]
- Coskuner, O.; Uversky, V.N. Current Stage and Future Perspectives for Homology Modeling, Molecular Dynamics Simulations, Machine Learning With Molecular Dynamics, and Quantum Computing for Intrinsically Disordered Proteins And Proteins With Intrinsically Disordered Regions. Current Protein and Peptide Science 2024, 25, 163–171. [Google Scholar] [CrossRef]
- Zhang, Z.; Nie, S.; Chen, L. Targeting prion-like protein spreading in neurodegenerative diseases. Neural Regen Res 2018, 13, 1875–1878. [Google Scholar] [CrossRef]
- Zafar, S.; Fatima, S.I.; Schmitz, M.; Zerr, I. Current Technologies Unraveling the Significance of Post-Translational Modifications (PTMs) as Crucial Players in Neurodegeneration. Biomolecules 2024, 14. [Google Scholar] [CrossRef]
- Hofweber, M.; Dormann, D. Friend or foe-Post-translational modifications as regulators of phase separation and RNP granule dynamics. J Biol Chem 2019, 294, 7137–7150. [Google Scholar] [CrossRef]
- Mathieu, C.; Pappu, R.V.; Taylor, J.P. Beyond aggregation: Pathological phase transitions in neurodegenerative disease. Science 2020, 370, 56–60. [Google Scholar] [CrossRef] [PubMed]
- Wheeler, R.J. Therapeutics-how to treat phase separation-associated diseases. Emerg Top Life Sci 2020, 4, 307–318. [Google Scholar] [CrossRef] [PubMed]
- Amzallag, E.; Hornstein, E. Crosstalk between Biomolecular Condensates and Proteostasis. Cells 2022, 11. [Google Scholar] [CrossRef] [PubMed]
- Vendruscolo, M.; Fuxreiter, M. Protein condensation diseases: therapeutic opportunities. Nat Commun 2022, 13, 5550. [Google Scholar] [CrossRef]
- Li, S.; Wang, Y.; Lai, L. Small molecules in regulating protein phase separation. Acta Biochim Biophys Sin (Shanghai) 2023, 55, 1075–1083. [Google Scholar] [CrossRef]
- Ding, M.; Xu, W.; Pei, G.; Li, P. Long way up: rethink diseases in light of phase separation and phase transition. Protein Cell 2024, 15, 475–492. [Google Scholar] [CrossRef]
- Liang, P.; Zhang, J.; Wang, B. Emerging Roles of Ubiquitination in Biomolecular Condensates. Cells 2023, 12. [Google Scholar] [CrossRef]
- Yerbury, J.J.; Ooi, L.; Dillin, A.; Saunders, D.N.; Hatters, D.M.; Beart, P.M.; Cashman, N.R.; Wilson, M.R.; Ecroyd, H. Walking the Tightrope: Proteostasis and Neurodegenerative Disease. Journal of Neurochemistry 2016, 137, 489–505. [Google Scholar] [CrossRef]
- Wood, L.B.; Winslow, A.R.; Strasser, S.D. Systems biology of neurodegenerative diseases. Integr Biol (Camb) 2015, 7, 758–775. [Google Scholar] [CrossRef]
- Uversky, V.N. Targeting intrinsically disordered proteins in neurodegenerative and protein dysfunction diseases: another illustration of the D(2) concept. Expert Rev Proteomics 2010, 7, 543–564. [Google Scholar] [CrossRef]
- Utami, K.H.; Morimoto, S.; Mitsukura, Y.; Okano, H. The roles of intrinsically disordered proteins in neurodegeneration. Biochim Biophys Acta Gen Subj 2025, 1869, 130772. [Google Scholar] [CrossRef] [PubMed]
- Martinelli, A.H.S.; Lopes, F.C.; John, E.B.O.; Carlini, C.R.; Ligabue-Braun, R. Modulation of Disordered Proteins with a Focus on Neurodegenerative Diseases and Other Pathologies. Int J Mol Sci 2019, 20. [Google Scholar] [CrossRef] [PubMed]
- Aftab, A.; Sil, S.; Nath, S.; Basu, A.; Basu, S. Intrinsic Disorder and Other Malleable Arsenals of Evolved Protein Multifunctionality. J Mol Evol 2024, 92, 669–684. [Google Scholar] [CrossRef] [PubMed]
- Chakrabarti, P.; Chakravarty, D. Intrinsically disordered proteins/regions and insight into their biomolecular interactions. Biophysical Chemistry 2022, 283, 106769. [Google Scholar] [CrossRef]
- Wesseling, H.; Mair, W.; Kumar, M.; Schlaffner, C.N.; Tang, S.; Beerepoot, P.; Fatou, B.; Guise, A.J.; Cheng, L.; Takeda, S.; et al. Tau PTM Profiles Identify Patient Heterogeneity and Stages of Alzheimer's Disease. Cell 2020, 183, 1699–1713 e1613. [Google Scholar] [CrossRef]
- Lin, W.; Phanse, S.; van der Spek, S.J.F.; Lampl, N.; Stephens, M.C.; Ortiz, A.R.; Taylor, A.; Hekman, R.; Roberts, R.; Jiang, L.; et al. Progressive Remodeling of Global Protein Interaction Networks in a Mouse Model of Tauopathy. bioRxiv 2025. [Google Scholar] [CrossRef]
- Gruet, A.; Dosnon, M.; Blocquel, D.; Brunel, J.; Gerlier, D.; Das, R.K.; Bonetti, D.; Gianni, S.; Fuxreiter, M.; Longhi, S.; et al. Fuzzy regions in an intrinsically disordered protein impair protein-protein interactions. FEBS J 2016, 283, 576–594. [Google Scholar] [CrossRef]
- Bhattacharya, S.; Lin, X. Recent Advances in Computational Protocols Addressing Intrinsically Disordered Proteins. Biomolecules 2019, 9. [Google Scholar] [CrossRef]
- Sharma, R.; Demeny, M.; Ambrus, V.; Kiraly, S.B.; Kurtan, T.; Gatti-Lafranconi, P.; Fuxreiter, M. Specific and Fuzzy Interactions Cooperate in Modulating Protein Half-Life. J Mol Biol 2019, 431, 1700–1707. [Google Scholar] [CrossRef]
- Fuxreiter, M.; Tompa, P. Fuzzy interactome: the limitations of models in molecular biology. Trends in Biochemical Sciences 2009, 34, 3. [Google Scholar] [CrossRef]
- Tsvetkov, P.; Reuven, N.; Shaul, Y. The nanny model for IDPs. Nat Chem Biol 2009, 5, 778–781. [Google Scholar] [CrossRef] [PubMed]
- Gajadhar, A.S.; White, F.M. System level dynamics of post-translational modifications. Curr Opin Biotechnol 2014, 28, 83–87. [Google Scholar] [CrossRef] [PubMed]
- Bah, A.; Forman-Kay, J.D. Modulation of Intrinsically Disordered Protein Function by Post-translational Modifications. J Biol Chem 2016, 291, 6696–6705. [Google Scholar] [CrossRef] [PubMed]
- Gluth, A.; Li, X.; Gritsenko, M.A.; Gaffrey, M.J.; Kim, D.N.; Lalli, P.M.; Chu, R.K.; Day, N.J.; Sagendorf, T.J.; Monroe, M.E. Integrative multi-PTM proteomics reveals dynamic global, redox, phosphorylation, and acetylation regulation in cytokine-treated pancreatic beta cells. Molecular & Cellular Proteomics 2024, 23, 100881. [Google Scholar]
- Sprunger, M.L.; Jackrel, M.E. Prion-Like Proteins in Phase Separation and Their Link to Disease. Biomolecules 2021, 11. [Google Scholar] [CrossRef]
- Cai, N.; Zhan, X.; Zhang, Q.; Di, H.; Chen, C.; Hu, Y.; Yan, X. Red Blood Cell-Derived Small Extracellular Vesicles Inhibit Influenza Virus Through Surface-Displayed Sialic Acids. Angewandte Chemie 2024, 137. [Google Scholar] [CrossRef]
- Zhang, Y.; Jin, C.; Xu, X.; Guo, J.; Wang, L. The role of liquid-liquid phase separation in the disease pathogenesis and drug development. Biomed Pharmacother 2024, 180, 117448. [Google Scholar] [CrossRef]
- Sun, J.; Chen, Y.; Bi, R.; Yuan, Y.; Yu, H. Bioinformatic approaches of liquid–liquid phase separation in human disease. Chinese medical journal 2024, 137, 1912–1925. [Google Scholar] [CrossRef]
- Ayyadevara, S.; Ganne, A.; Balasubramaniam, M.; Shmookler Reis, R.J. Intrinsically disordered proteins identified in the aggregate proteome serve as biomarkers of neurodegeneration. Metab Brain Dis 2022, 37, 147–152. [Google Scholar] [CrossRef]
- Manzanza, N.O.; Sedlackova, L.; Kalaria, R.N. Alpha-Synuclein Post-translational Modifications: Implications for Pathogenesis of Lewy Body Disorders. Front Aging Neurosci 2021, 13, 690293. [Google Scholar] [CrossRef]
- Oh, I.M.; Li, S.H.; Pereira, F.L.; Kaur, P.; Raju, A.; Jerfy, R.; Suemoto, C.K.; Leite, R.E.P.; Ribeiro Paes, V.; Rodriguez, R.D.; et al. Truncated tau accumulates before hyperphosphorylated tau and in relatively distinct neuronal subpopulations in entorhinal cortex and inferior temporal gyrus in Alzheimer’s disease patients. Alzheimer's & Dementia 2025, 20, e093137. [Google Scholar] [CrossRef]
- Snider, N.T.; Omary, M.B. Post-translational modifications of intermediate filament proteins: mechanisms and functions. Nat Rev Mol Cell Biol 2014, 15, 163–177. [Google Scholar] [CrossRef]
- Zhang, H.; Ge, Y. Comprehensive analysis of protein modifications by top-down mass spectrometry. Circ Cardiovasc Genet 2011, 4, 711. [Google Scholar] [CrossRef]
- Zetterberg, H.; Blennow, K. Chronic traumatic encephalopathy: fluid biomarkers. Handb Clin Neurol 2018, 158, 323–333. [Google Scholar] [CrossRef]
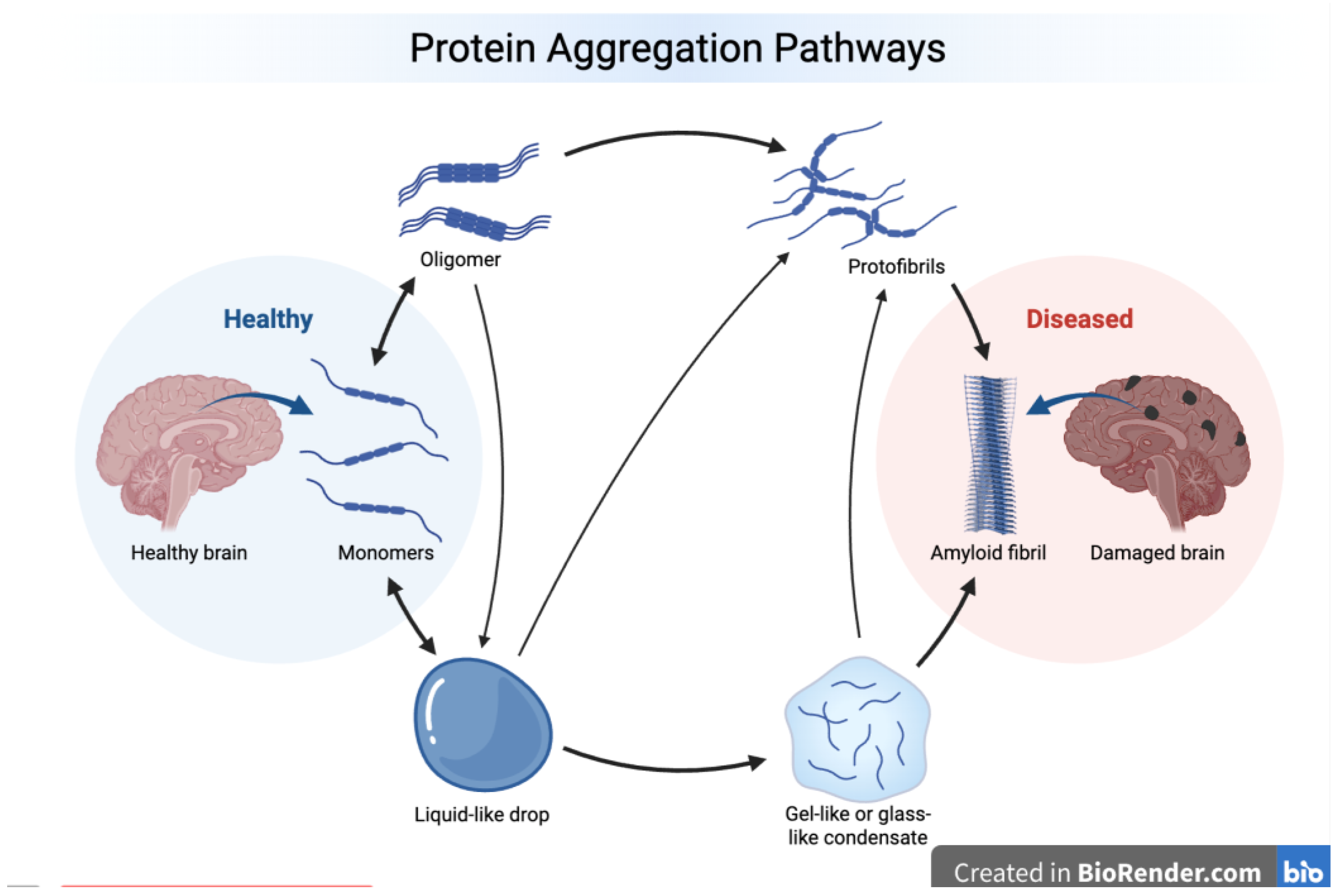
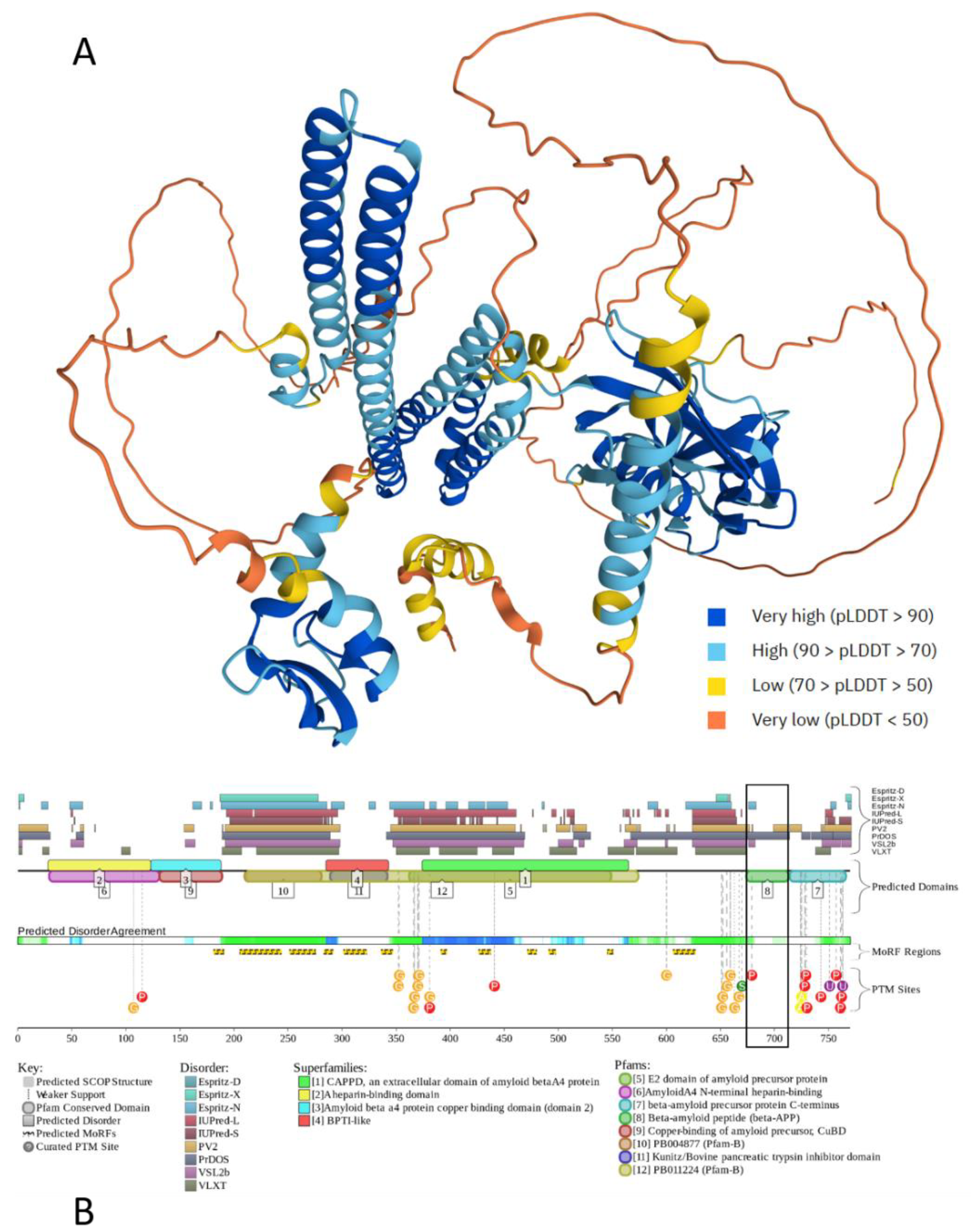
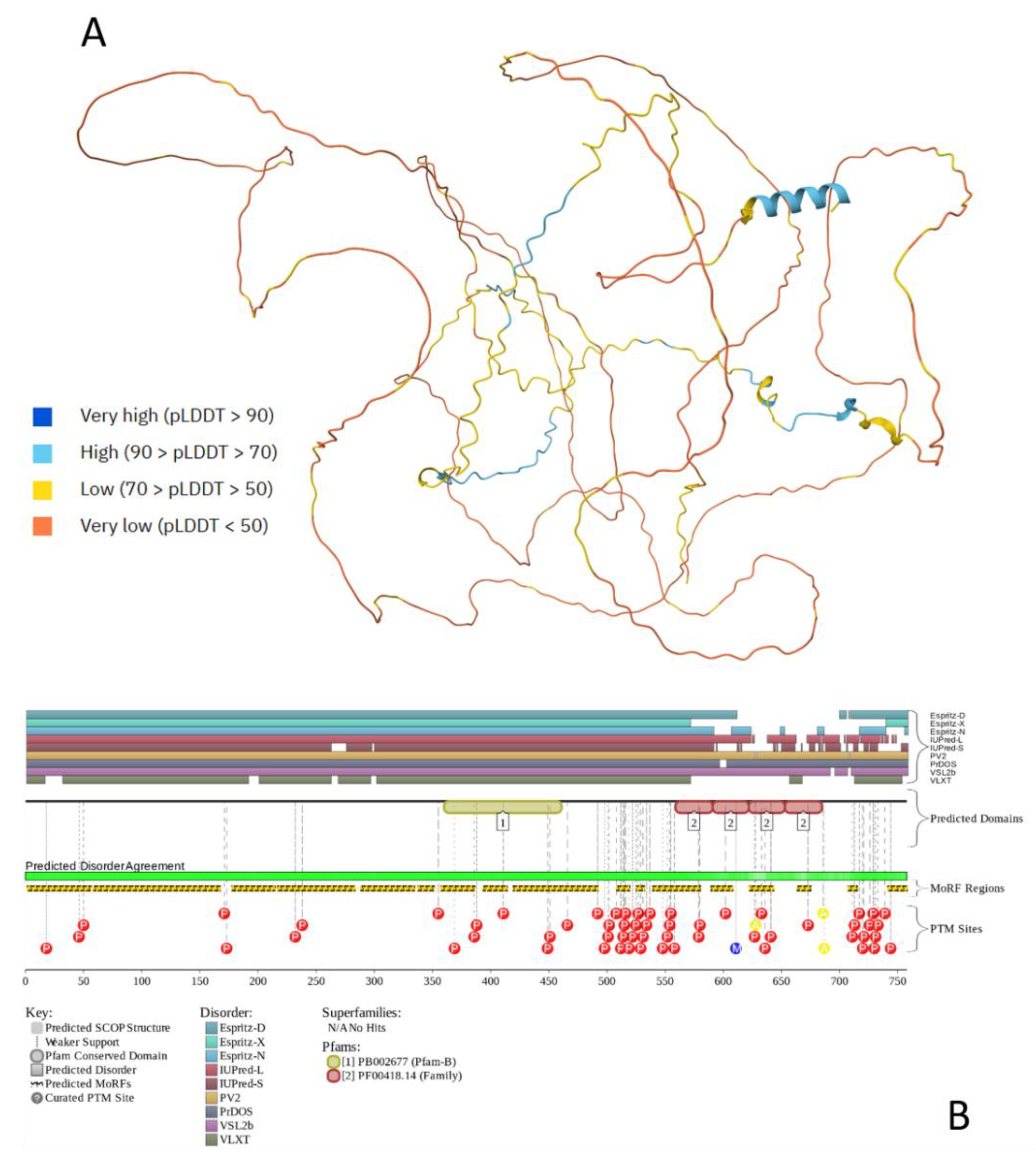
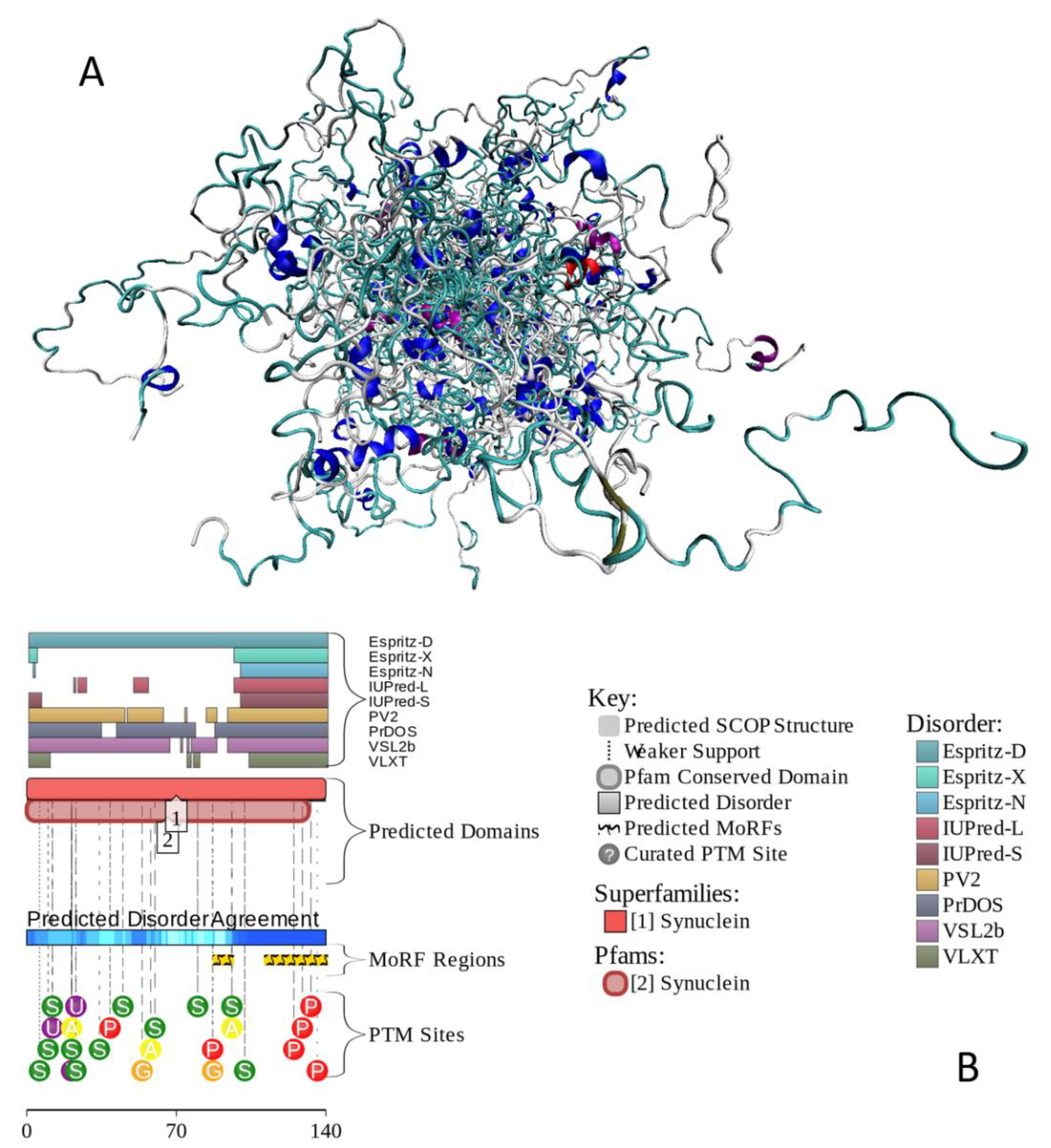
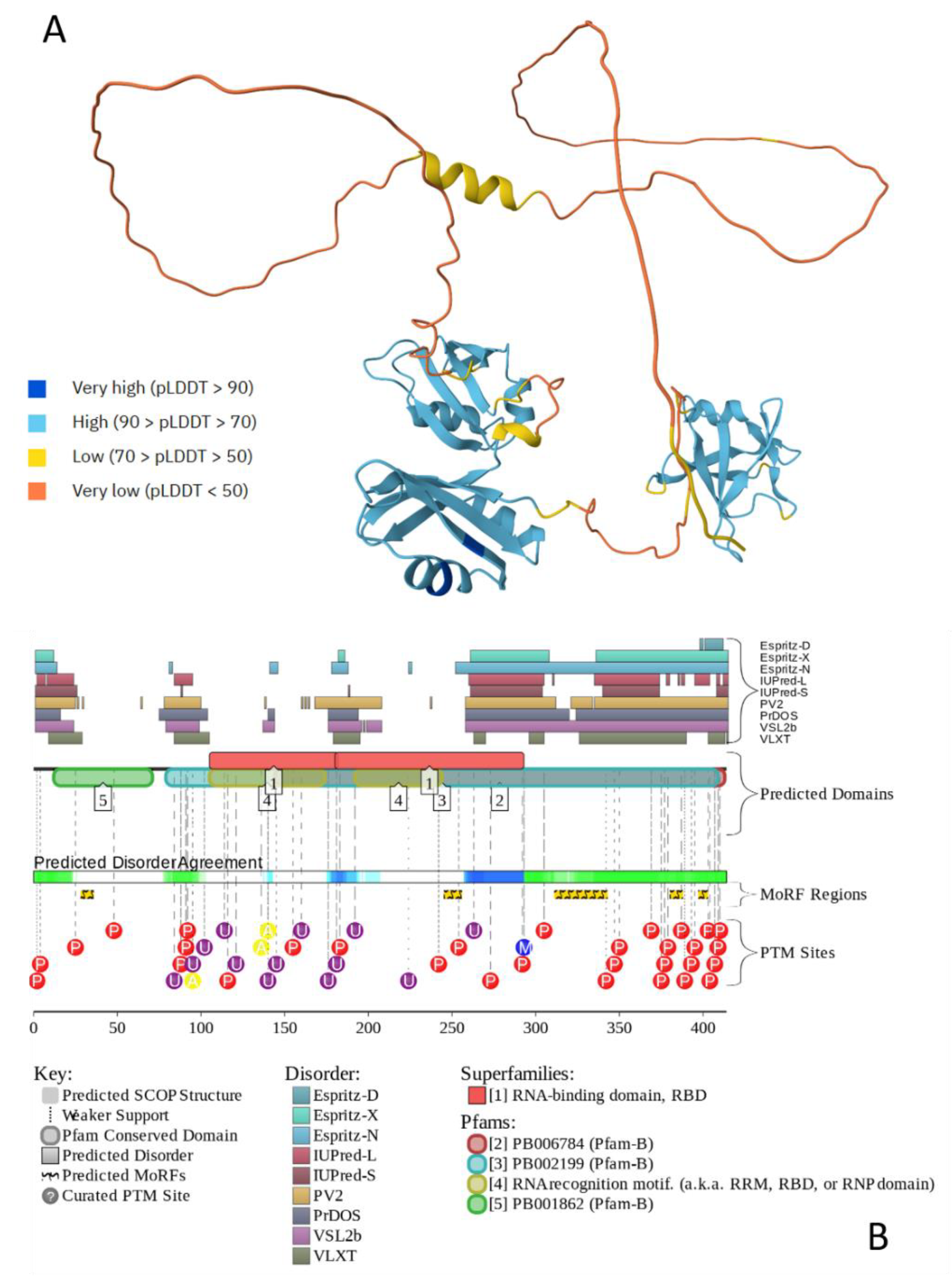
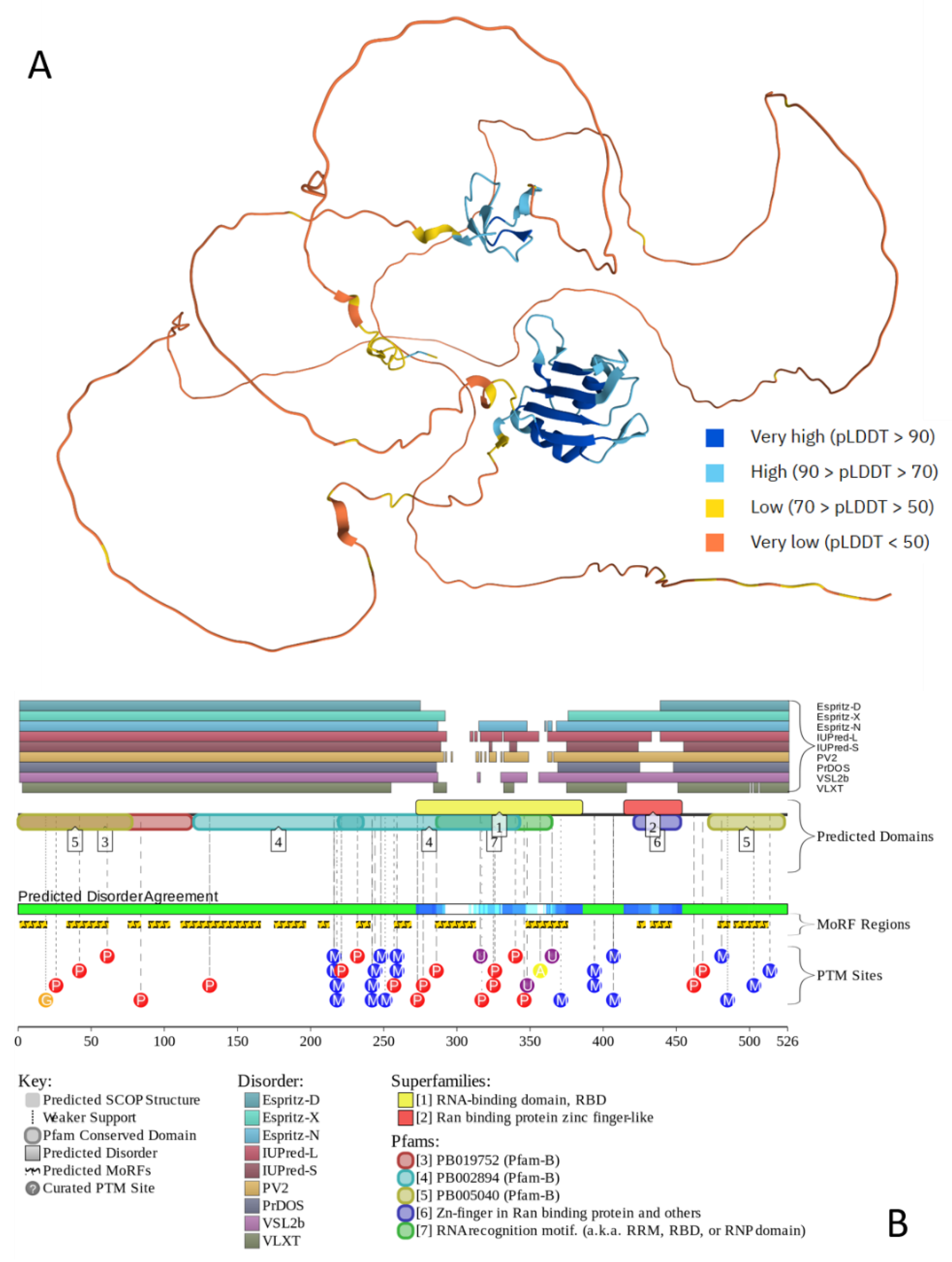
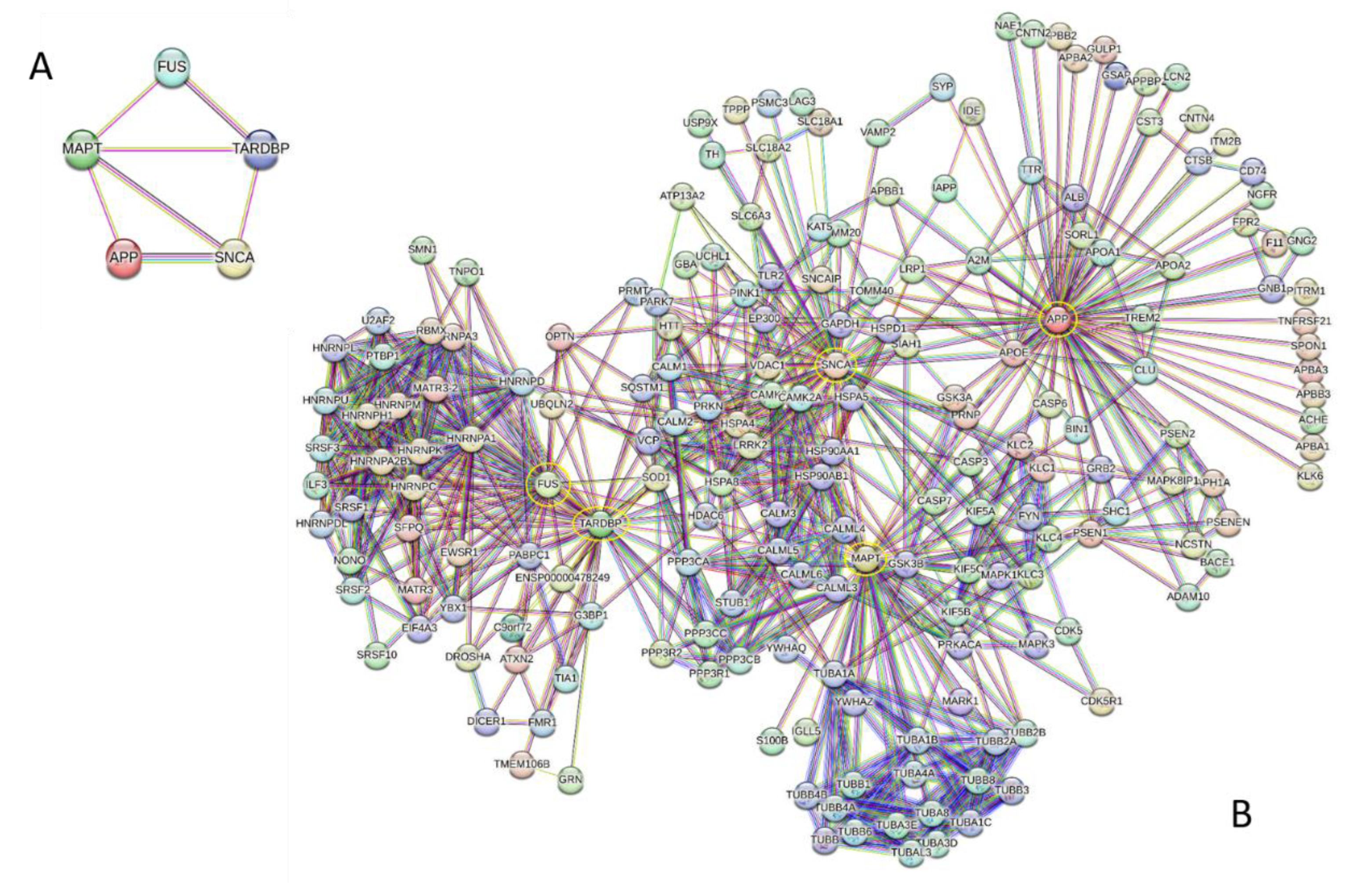
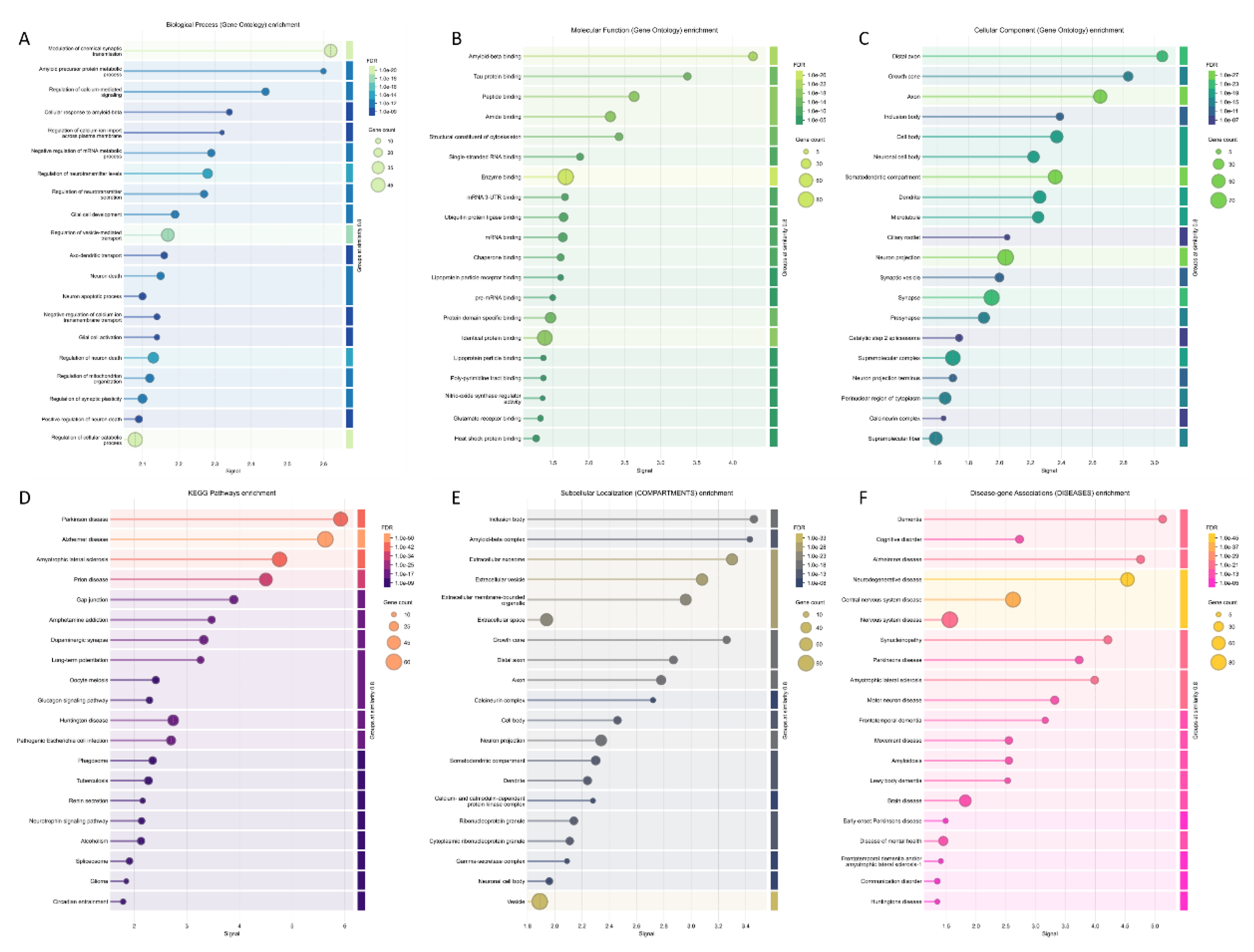
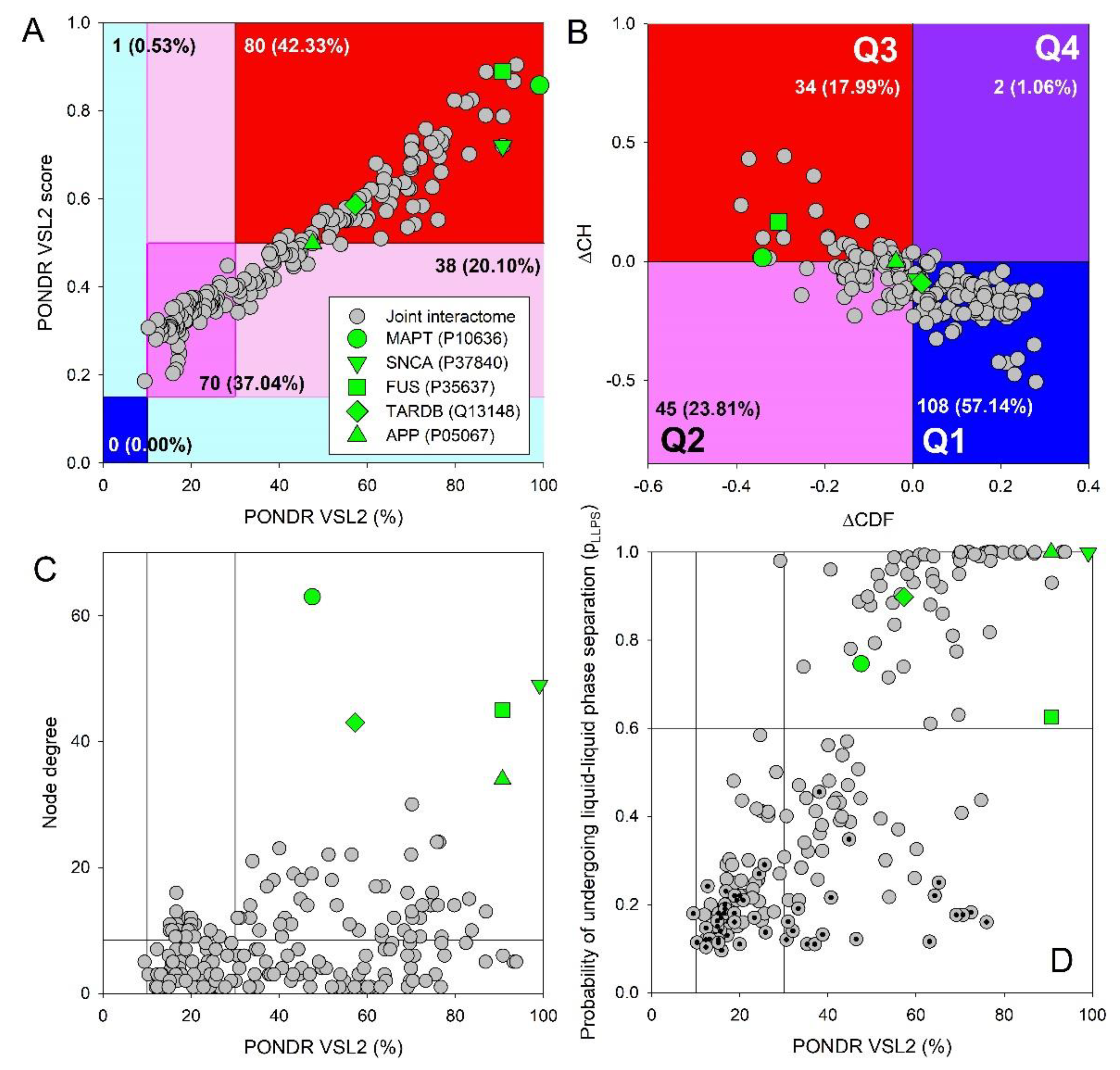
| Feature | Amyloid-β (Aβ) [94,96,98,227] | τ-protein (tau) [120,140] | α-synuclein [150,167,228] | TDP-43 [197,200,229] | FUS [122,201,209] |
| Physiological role | APP-derived peptide; modulates synaptic activity at low concentrations | Microtubule-associated protein stabilizing neuronal function | Presynaptic protein regulating vesicle trafficking | RNA-binding protein regulating splicing and RNA metabolism | RNA/DNA-binding protein involved in RNA metabolism and stress responses |
| Pathological aggregate | Extracellular plaques; soluble toxic oligomers | Intracellular neurofibrillary tangles (NFTs) | Lewy bodies and Lewy neurites; oligomers | Cytoplasmic inclusions with nuclear depletion | Cytoplasmic inclusions due to nuclear clearance defects |
| Intrinsic disorder features | Intrinsically disordered as a monomer | Highly intrinsically disordered; conformationally flexible | Almost entirely intrinsically disordered | Large intrinsically disordered regions | Highly intrinsically disordered; prion-like domains |
| Sequence determinants of disorder | Charged and flexible N-terminus; hydrophobic C-terminal motifs (Aβ42) | Low sequence complexity; polar and charged amino acid residues | NAC region; polar and charged amino acid residues, particularly in its N-terminal region | Low-complexity C-terminal domain with polar and charged amino acid residues; charge asymmetry | Low-complexity, Arg-Gly-Gly repeats in the RGG motif |
| Post-translational modifications (PTMs) | Phosphorylation and acetylation | Hyperphosphorylation and acetylation | N-terminal acetylation and phosphorylation | Hyperphosphorylation and ubiquitination | PTMs modulate phase behavior and aggregation |
| Phase separation (LLPS) | No classical LLPS; APP can undergo LLPS, aggregation emerges from disordered peptide plasticity | Undergoes LLPS; droplets can mature into fibrillar aggregates | Forms stress-induced condensates | Stress-granule–associated LLPS precedes aggregation | Prominent LLPS; condensate hardening linked to disease |
| Prevalence in aged brain | Very high | Very high | Moderate to high | Common, especially in elderly | Rare in community autopsy series |
| Common co-pathologies | Tau, α-synuclein, TDP-43, FUS, vascular pathology | Aβ, α-synuclein, TDP-43 | Aβ, tau | Aβ, tau, α-synuclein | TDP-43; ALS/FTLD spectrum |
| Cross-seeding/ synergy | Seeds tau misfolding; interacts with α-synuclein | Synergizes with Aβ; interacts with α-synuclein | Cross-seeds tau and Aβ; strain-dependent | Often superimposed on AD pathology | Stress-granule and RNA-metabolism interactions |
| Impact on mixed-pathology dementia | Early driver and amplifier of downstream proteinopathies | Strong determinant of cognitive decline and neuronal loss | Exacerbates cognitive and neuropsychiatric symptoms | Accelerates dementia severity and memory impairment | Associated with earlier onset and aggressive disease |
| Associated diseases | AD, cerebral amyloid angiopathy (CAA), Down syndrome, Aβ–related angiitis (ABRA), CAA-related inflammation, cerebral amyloidoma, dementia with Lewy bodies (DLB), retinal disorders, traumatic brain injury (TBI) | AD, progressive supranuclear palsy (PSP), corticobasal degeneration (CBD), Pick’s disease (PiD), frontotemporal dementia with parkinsonism-17 (FTDP-17), argyrophilic grain disease (AGD), chronic traumatic encephalopathy (CTE), down syndrome, Guam parkinsonism-dementia complex, postencephalitic parkinsonism, DLB, PD | PD, DLB, multiple system atrophy (MSA), pure autonomic failure (PAF), REM sleep behavior disorder (RBD), AD, Gaucher’s disease, neuroaxonal dystrophy, neurodegeneration with brain iron accumulation (NBIA) | ALS, FTLD-TDP, LATE, AD, CTE, LBD, Huntington’s disease (HD), multisystem proteinopathy (MSP), Perry syndrome, Alexander disease | ALS-FUS, FTLD-FUS, neuronal intermediate filament inclusion disease (NIFID), basophilic inclusion body disease (BIBD), essential tremor (ET), polyglutamine (PolyQ) diseases (HD and spinocerebellar ataxias SCA1 and SCA3), cancers (myxoid liposarcoma, ewing sarcoma, acute myeloid leukemia (AML) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




