Submitted:
05 February 2026
Posted:
06 February 2026
You are already at the latest version
Abstract
Keywords:
Part I. Data Sources and Analytical Framework
Pairwise FST Analysis
Global25 and Affinity Visualization
DNAgenics Admixture Modeling
Original Analyses and Derived Data Generated in This Study
Interpretive Framework
Rationale for Reanalysis Without New Sequencing
Part II. Reference Population Design and the Appearance of Genetic Intermediacy
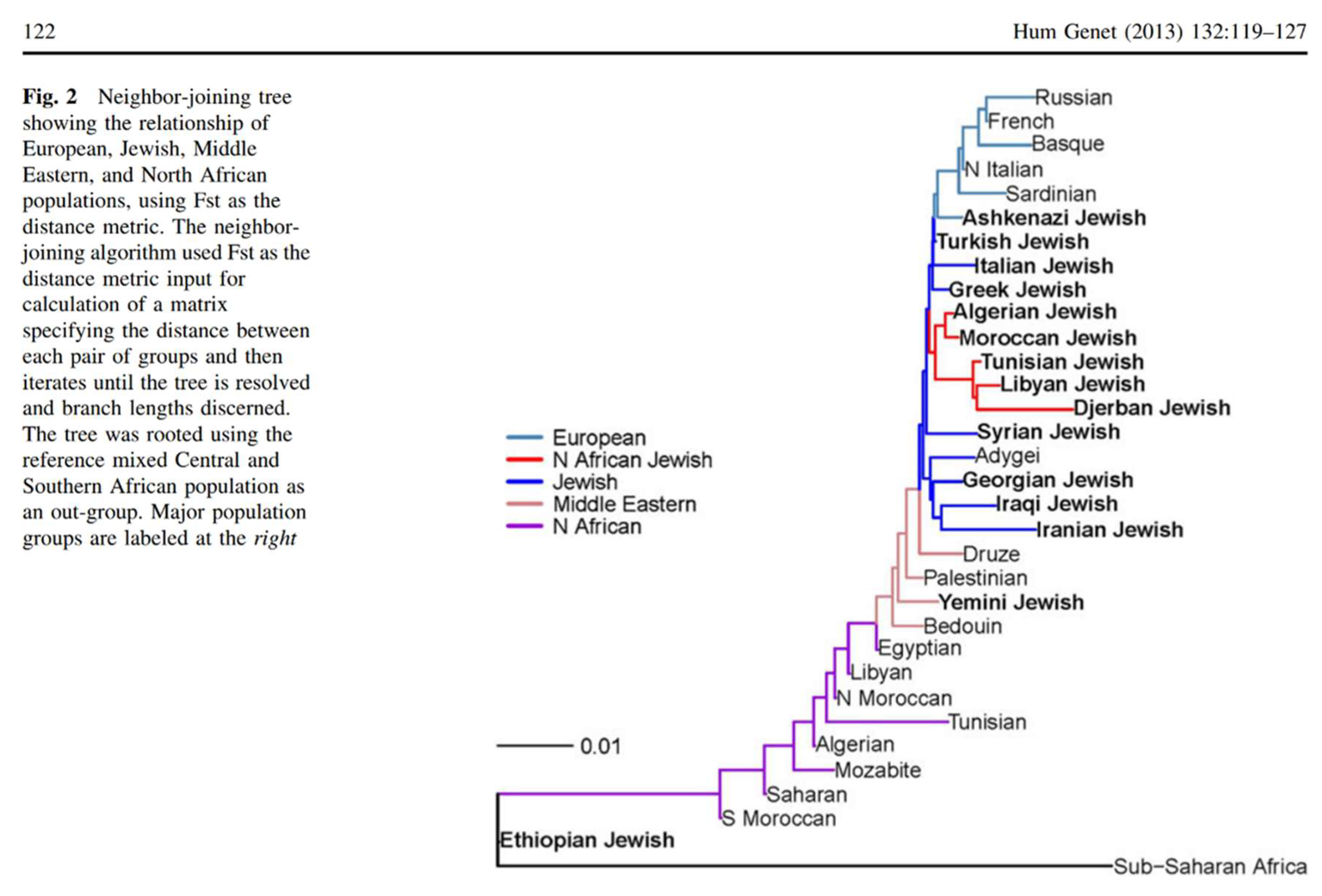
Part III. Y-Chromosomal Structure of Ashkenazi Jews in Mediterranean Context
Uniparental Misinterpretation and the Illusion of a Levantine Paternal Origin
Reassessment of Ashkenazi Y-Chromosomal Structure Using Southern Italian Comparators
Demonstration of Ashkenazi Divergence from Levantine Paternal Structure
| Population | E-M78 | E-M123 | J1-M267 | J2-M172 |
|---|---|---|---|---|
| Ashkenazi Jews | 5.2 | 11.7 | 14.6 | 23.2 |
| Lebanese | ~2 to 5 | ~4 to 5 | ~25 to 35 | ~10 to 15 |
| Palestinians | ~1 to 3 | ~2 to 4 | ~35 to 45 | ~10 to 15 |
| Iraqis | ~1 to 3 | ~2 to 3 | ~40 to 50 | ~10 to 15 |
Fine-Scale Southern European J1 Variation and Upper-Range Overlap
| Region | Population | J1-M267 |
|---|---|---|
| Italy | Sicily, Agrigento | 11.1 |
| Italy | Western Sicily | 10.13 |
| Greece | Crete, Nea Nikomedeia | 10.5 |
| Italy | Trapani, Sicily | 8.82 |
| Italy | Campania, Benevento | 8.3 |
| Greece | Crete | 8.3 |
| Italy | Sicily, Southwest | 7.0 |
| Italy | Calabria, Tyrrhenian Calabria | 6.8 |
Cohen Modal Haplotype Arguments and Comparator Omission
Part IV. Interpretive Framework and Constraints on Inference
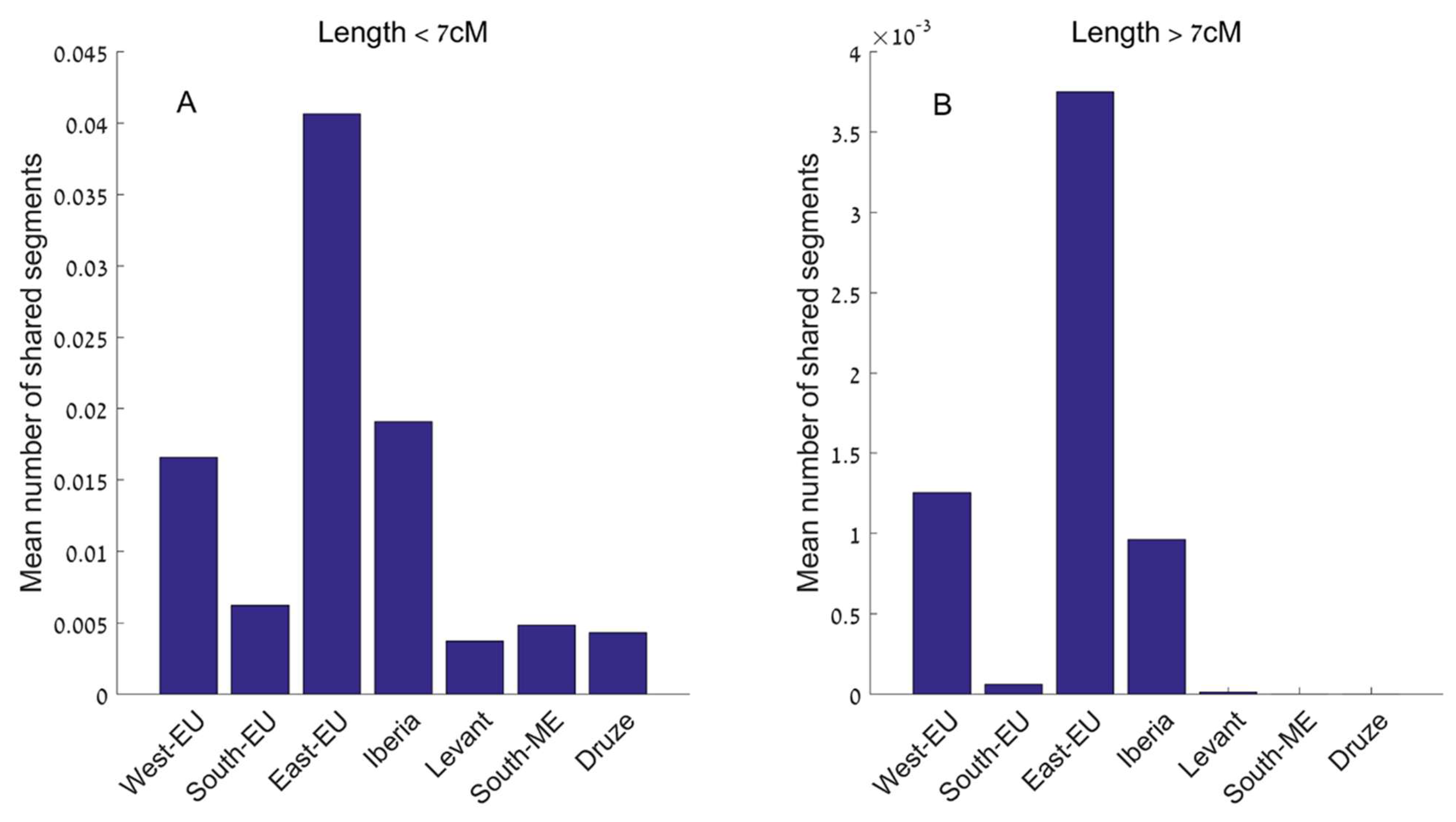
Part V. Convergent Autosomal Modeling Across Independent Analytical Frameworks
| Study | Methodology | Data type | Southern Italian / Mediterranean component | Additional components |
|---|---|---|---|---|
| Lerga-Jaso et al. (2025) | Local ancestry inference (Orchestra; IBD-based plus deep-learning smoothing) | Modern genomes | ~68% Italian | ~16.6% Levantine; ~7.2% Iraqi-Iranian-Caucasian-Turkish; ~2.4% Greek-Balkan; ~1.7% Eastern European |
| Waldman et al. (2022) | qpAdm (historically constrained model) | Modern genomes | ~65% Southern Italian | ~19% Lebanese; ~16% Eastern European |
| Brace et al. (2022) | qpAdm anchored to medieval ancient DNA (Chapelfield) | Medieval plus modern | ~67% Sicilian (Southern Italian proxy) | ~33% Turkish Jewish; ~0% French; ~0% Polish |
Convergent Autosomal FST Evidence for Southern European affinity of Ashkenazi Jews
| A | ||
|---|---|---|
| Comparison population | FST | |
| Italians | 0.0040 | |
| Greeks | 0.0042 | |
| Spanish | 0.0056 | |
| Tuscans | 0.0066 | |
| Germans | 0.0072 | |
| Druze | 0.0088 | |
| Palestinians | 0.0093 | |
| Irish | 0.0109 | |
| Swedes | 0.0120 | |
| Russians | 0.0137 | |
| Basque | 0.0144 | |
| B | ||
| Comparison population | Distance (x1000) | |
| Italians | 44 | |
| Greeks | 105 | |
| Turks | 170 | |
| Germans | 131 | |
| French | 144 | |
| British | 238 | |
| Poles | 195 | |
| Russians | 230 | |
| Palestinians | 277 | |
Part VI. Integrated Interpretation and Implications for Models of Ashkenazi Origins
| Population | Sample size | CCR5-Δ32 allele frequency |
|---|---|---|
| Swedish | 131 | 0.137 |
| Estonian | 158 | 0.133 |
| Polish | 30 | 0.133 |
| French | 230 | 0.089 |
| Italian | 172 | 0.055 |
| Greek | 160 | 0.044 |
| Ashkenazi Jews | 503 | 0.097 |
| Lebanese | 51 | 0.000 |
| Saudi Arabian | 100 | 0.000 |
| Chinese | 40 | 0.000 |
| African populations | various | 0.000 |
Part VII. Historical and Geographic Context of Jewish Settlement in Southern Italy and the Central Mediterranean
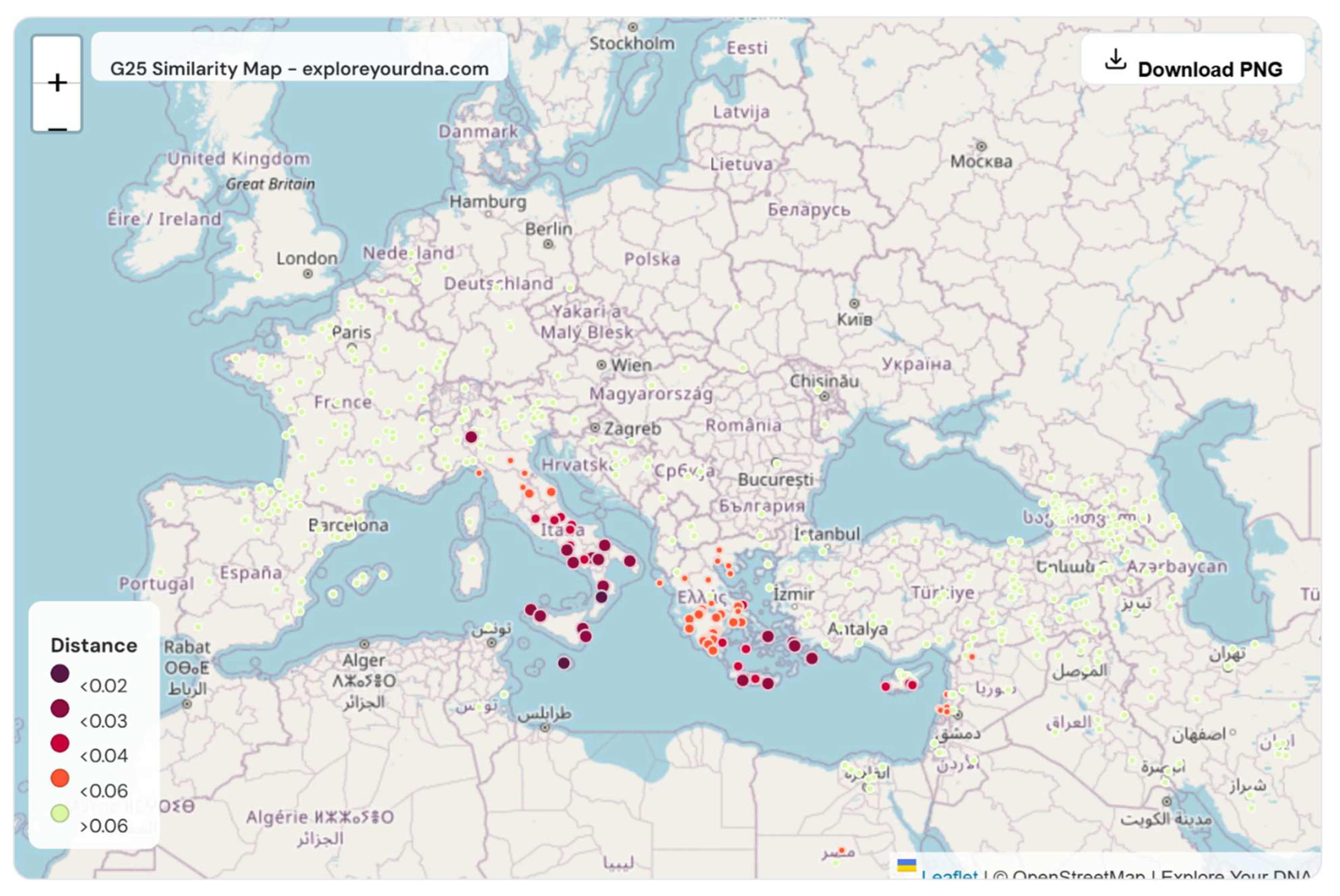
| # | Population | Distance |
|---|---|---|
| 1 | Maltese | 0.0180 |
| 2 | Italian Calabria | 0.0182 |
| 3 | Italian Campania | 0.0207 |
| 4 | Italian Campania Naples (Campanian) | 0.0224 |
| 5 | Sicilian Syracuse | 0.0236 |
| 6 | Sicilian Trapani | 0.0237 |
| 7 | Greek Crete | 0.0245 |
| 8 | Sicilian Central | 0.0245 |
| 9 | Sicilian East | 0.0245 |
| 10 | Italian Calabria (Cosentian) | 0.0248 |
| 11 | Greek Crete Lasithi | 0.0251 |
| 12 | Italian Basilicata (Lucanian) | 0.0253 |
| 13 | Italian Basilicata | 0.0253 |
| 14 | Greek Crete Rethymno | 0.0261 |
| 15 | Sicilian | 0.0270 |
| 16 | Greek Dodecanese Kos | 0.0273 |
| 17 | Greek Kos | 0.0273 |
| 18 | Italian Apulia | 0.0278 |
| 19 | Greek Dodecanese | 0.0280 |
| 20 | Sicilian West | 0.0288 |
| 21 | Italian Apulia (Apulian) | 0.0290 |
| 22 | Greek Cyclades Amorgos | 0.0298 |
| 23 | Greek Crete Chania | 0.0307 |
| 24 | Italian Abruzzo | 0.0313 |
| 25 | Italian Abruzzo (Abruzzese) | 0.0313 |
| 26 | Greek Euboea Central | 0.0318 |
| 27 | Greek Crete Heraklion | 0.0321 |
| 28 | Greek Cyclades Milos | 0.0334 |
| 29 | Italian Campania Salerno (Campanian) | 0.0353 |
| 30 | Italian Molise | 0.0353 |
Part VIII. Independent qpAdm and FST Analyses with External Validation
qpAdm Modeling Performed by the Author
- Mbuti.DG
- Yoruba.DG
- Ju_hoan_North.DG
- Han.DG
- Chukchi.DG
- Karitiana.DG
- Papuan.DG
- Iranian.DG
- Switzerland_Bichon_Epipaleolithic.SG
- Basque.DG
Autosomal FST Distances Calculated by the Author
FST and PCA Calibration Using the Canary Islands
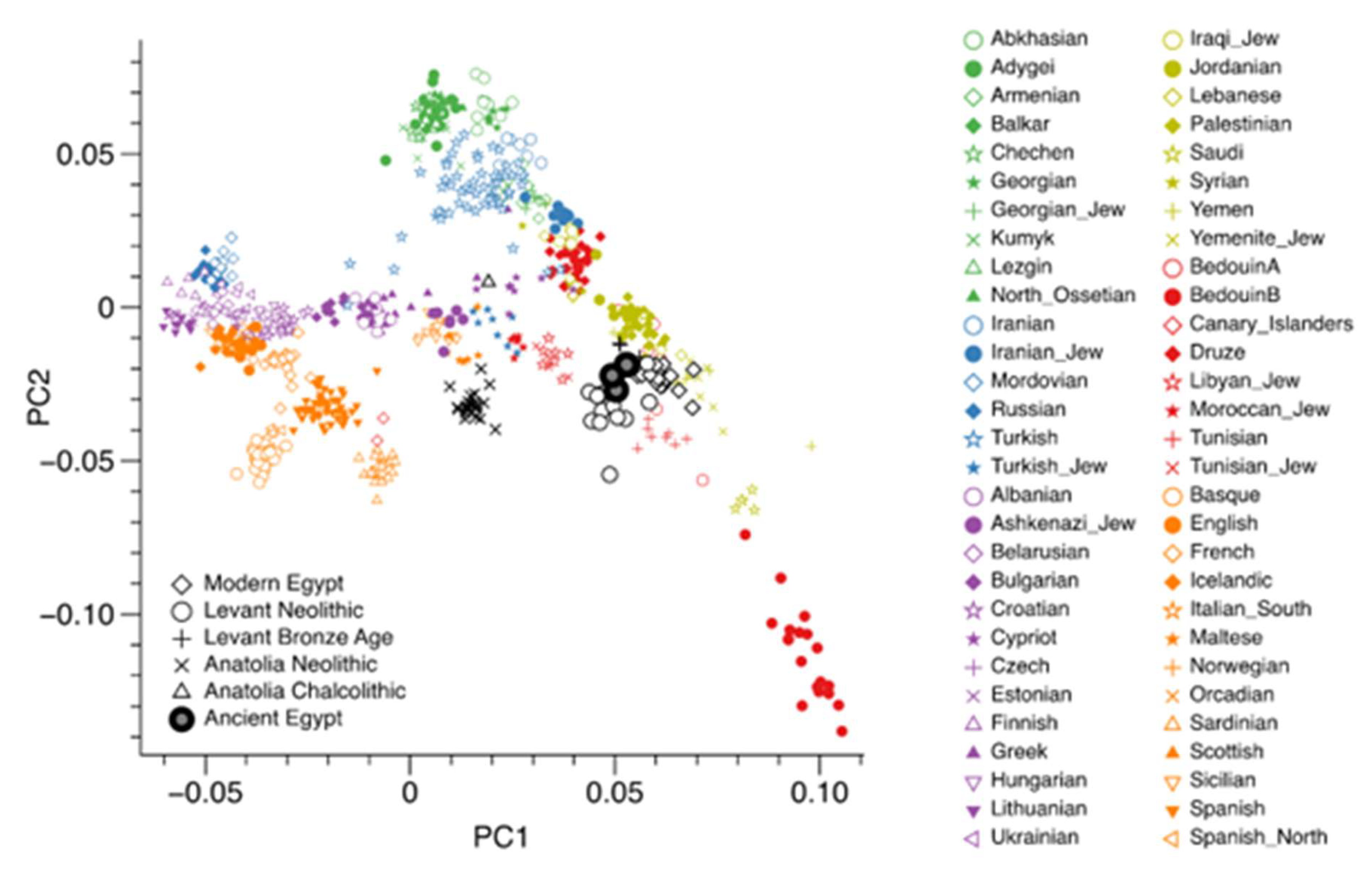
Ancient-Only G25 Similarity Analysis (Author-Generated)
Summary of Author-Generated Validation Analyses
Part IX. Independent Modern Population PCA and Full-Dimensional Global25 Distance Analysis
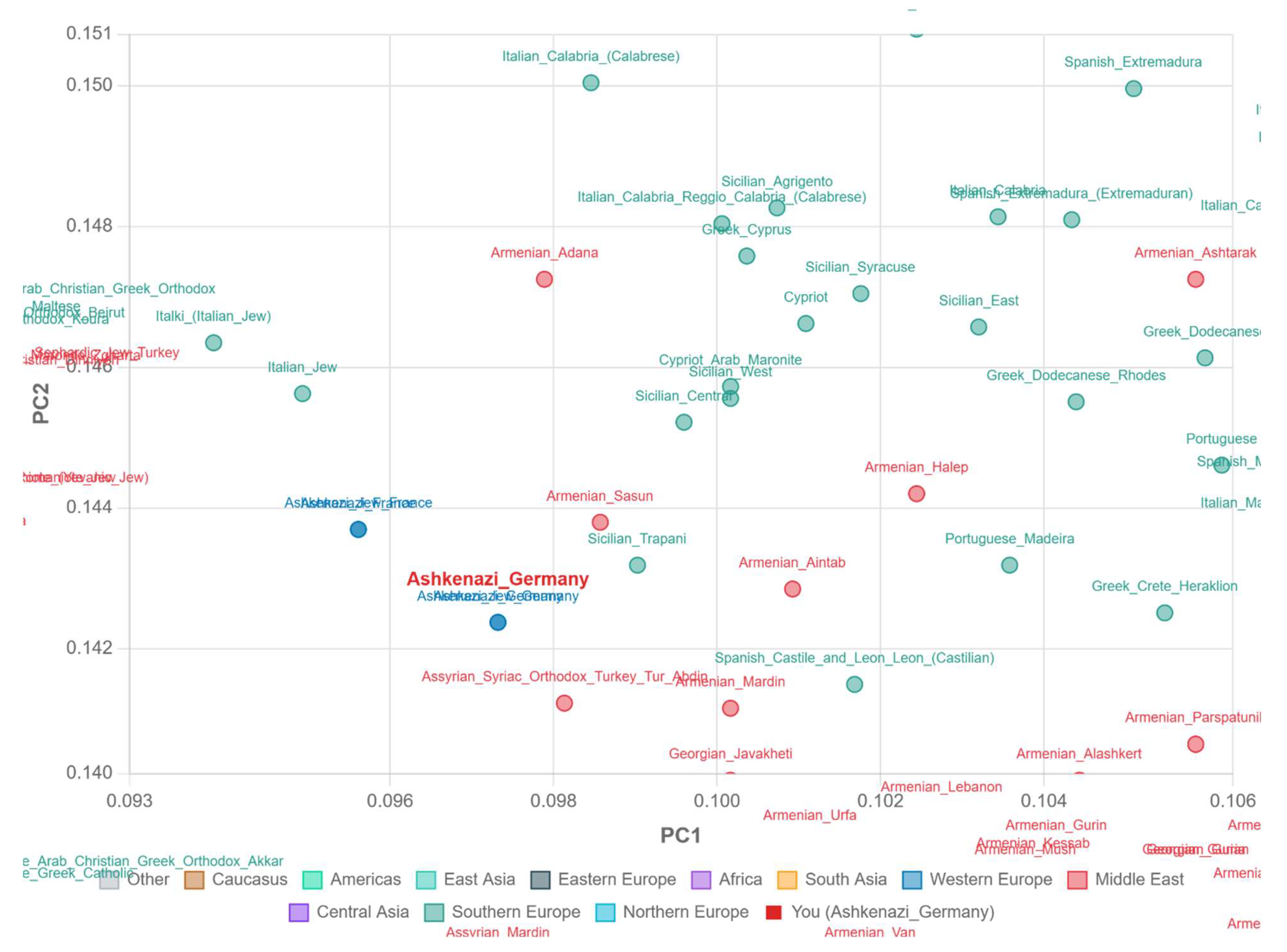
PCA Visualization (PC1 vs PC2)
Full 25-Dimensional Global25 Distance Rankings
Synthesis of Global25 Results
Part X. Italian Jews as a Falsification Test of Northern Italian and Convergent Origin Models
Part XI. Conclusions: Southern Italian Affinity and Reference Population Effects in Ashkenazi Origin Models
Supplementary Materials
References
- Atzmon, G.; Hao, L.; Pe’er, I.; Velez, C.; Pearlman, A.; Palamara, P.F.; Morrow, B.; Friedman, E.; Oddoux, C.; Burns, E.; Ostrer, H. Abraham’s children in the genome era: major Jewish diaspora populations comprise distinct genetic clusters with shared Middle Eastern ancestry. American Journal of Human Genetics 2010, 86, 850–859. [Google Scholar] [CrossRef] [PubMed]
- Behar, D.M.; Garrigan, D.; Kaplan, M.E.; Mobasher, Z.; Rosengarten, D.; Karafet, T.M.; Quintana-Murci, L.; Ostrer, H.; Skorecki, K.; Hammer, M.F. Contrasting patterns of Y chromosome variation in Ashkenazi Jewish and host non-Jewish European populations. Human Genetics 2004, 114, 354–365. [Google Scholar] [CrossRef]
- Behar, D.M.; Yunusbayev, B.; Metspalu, M.; Metspalu, E.; Rosset, S.; Parik, J.; Rootsi, S.; Chaubey, G.; Kutuev, I.; Yudkovsky, G.; Khusnutdinova, E.K.; Balanovsky, O.; Semino, O.; Pereira, L.; Comas, D.; Gurwitz, D.; Bonne-Tamir, B.; Parfitt, T.; Hammer, M.F.; Skorecki, K.; Villems, R. The genome-wide structure of the Jewish people. Nature 2010, 466, 238–242. [Google Scholar] [CrossRef]
- Boattini, A.; Martinez-Cruz, B.; Sarno, S.; Harmant, C.; Useli, A.; et al. Uniparental markers in Italy reveal a sex-biased genetic structure and different historical strata. PLOS ONE 2013, 8, e65441. [Google Scholar] [CrossRef]
- Bonfil, R. Jewish Life in Renaissance Italy; University of California Press: Berkeley, 1994. [Google Scholar]
- Botigué, L.R.; Henn, B.M.; Gravel, S.; Maples, B.K.; Gignoux, C.R.; Corona, E.; Atzmon, G.; Burns, E.; Ostrer, H.; Flores, C.; Bertranpetit, J.; Comas, D.; Bustamante, C.D. Gene flow from North Africa contributes to differential human genetic diversity in southern Europe. Proceedings of the National Academy of Sciences USA 2013, 110, 11791–11796. [Google Scholar] [CrossRef]
- Brace, S.; Diekmann, Y.; Booth, T.; et al. Genomes from a medieval mass burial show Ashkenazi-associated hereditary diseases pre-date the 12th century. Cell 2022, 185, 4350–4359.e6. [Google Scholar] [CrossRef]
- Bray, S.M.; Mulle, J.G.; Dodd, A.F.; Pulver, A.E.; Wooding, S.; Warren, S.T. Signatures of founder effects, admixture, and selection in the Ashkenazi Jewish population. Proceedings of the National Academy of Sciences USA 2010, 107, 16222–16227. [Google Scholar] [CrossRef] [PubMed]
- Campbell, C.L.; Palamara, P.F.; Dubrovsky, M.; Botigué, L.R.; Fellous, M.; Atzmon, G.; Oddoux, C.; Pearlman, A.; Hao, L.; Henn, B.M.; Burns, E.; Bustamante, C.D.; Comas, D.; Friedman, E.; Pe’er, I.; Ostrer, H. North African Jewish and non-Jewish populations form distinctive, orthogonal clusters. Proceedings of the National Academy of Sciences USA 2012, 109, 13865–13870. [Google Scholar] [CrossRef]
- Chazan, R. European Jewry and the First Crusade; University of California Press: Berkeley, 1987. [Google Scholar]
- Chiaroni, J.; King, R.J.; Myres, N.M.; Henn, B.M.; Ducourneau, A.; Mitchell, M.J.; Boetsch, G.; Sheikha, I.; et al. The emergence of Y-chromosome haplogroup J1e among Arabic-speaking populations. European Journal of Human Genetics 2010, 18, 348–353. [Google Scholar] [CrossRef]
- Di Gaetano, C.; Voglino, F.; Guarrera, S.; Fiorito, G.; Rosa, F.; Di Blasio, A.M.; Manzini, P.; Dianzani, I.; Betti, M.; Cusi, D.; Frau, F.; Barlassina, C.; Mirabelli, D.; Matullo, G. An overview of the genetic structure within the Italian population from genome-wide data. PLOS ONE 2012, 7, e43759. [Google Scholar] [CrossRef] [PubMed]
- Di Giacomo, F.; Luca, F.; Anagnou, N.; et al. Clinal patterns of human Y chromosomal diversity in continental Italy and Greece are dominated by drift and founder effects. Molecular Phylogenetics and Evolution 2003, 28, 387–395. [Google Scholar] [CrossRef]
- Drineas, P.; Tsetsos, F.; Plantinga, A.; et al. Genetic history of the population of Crete. Annals of Human Genetics 2019, 83, 373–388. [Google Scholar] [CrossRef] [PubMed]
- Fadhlaoui-Zid, K.; Martinez-Cruz, B.; Khodjet-El-Khil, H.; Mendizabal, I.; Benammar-Elgaaïed, A.; Comas, D. Genetic structure of Tunisian ethnic groups revealed by paternal lineages. American Journal of Physical Anthropology 2011, 146, 271–280. [Google Scholar] [CrossRef] [PubMed]
- Feldman, L.H. Jew and Gentile in the Ancient World; Princeton University Press: Princeton, 1993. [Google Scholar]
- Grossman, A. The Early Sages of Ashkenaz; Magnes Press: Jerusalem, 1995. [Google Scholar]
- Gruen, E.S. Diaspora: Jews Amidst Greeks and Romans; Harvard University Press: Cambridge, MA, 2002. [Google Scholar]
- Hammer, M.F.; Redd, A.J.; Wood, E.T.; et al. Jewish and Middle Eastern non-Jewish populations share a common pool of Y-chromosome biallelic haplotypes. Proceedings of the National Academy of Sciences USA 2000, 97, 6769–6774. [Google Scholar] [CrossRef]
- Hammer, M.F.; Behar, D.M.; Karafet, T.M.; Mendez, F.L.; Hallmark, B.; Erez, T.; Zhivotovsky, L.A.; Rosset, S.; Skorecki, K. Extended Y chromosome haplotypes resolve multiple and unique lineages of the Jewish priesthood. Human Genetics 2009, 126, 707–717. [Google Scholar] [CrossRef]
- King, R.J.; Ozcan, S.S.; Carter, T.; Kalfoğlu, E.; Atasoy, S.; Triantaphyllidis, C.; Kouvatsi, A.; Lin, A.A.; Chow, C.E.T.; Zhivotovsky, L.A.; Michalodimitrakis, M.; Underhill, P.A. Differential Y-chromosome Anatolian influences on the Greek and Cretan Neolithic. Annals of Human Genetics 2008, 72, 205–214. [Google Scholar] [CrossRef] [PubMed]
- Kopelman, N.M.; Stone, L.; Wang, C.; Gefel, D.; Feldman, M.W.; Hillel, J.; Rosenberg, N.A. Genomic microsatellites identify shared Jewish ancestry intermediate between Middle Eastern and European populations. BMC Genetics 2009, 10, 80. [Google Scholar] [CrossRef] [PubMed]
- Lerga-Jaso, J.; Novković, B.; Unnikrishnan, D.; et al. Tracing human genetic histories with precise local ancestry inference. Nature Communications 2025, 16, 4576. [Google Scholar] [CrossRef]
- Listman, J.B.; Hasin, D.; Kranzler, H.R.; Malison, R.T.; Mutirangura, A.; Sughondhabirom, A.; Aharonovich, E.; Spivak, B.; Gelernter, J. Identification of population substructure among Jews using STR markers and dependence on reference populations included. BMC Genetics 2010, 11, 48. [Google Scholar] [CrossRef]
- Nebel, A.; Filon, D.; Brinkmann, B.; Majumder, P.P.; Faerman, M.; Oppenheim, A. The Y chromosome pool of Jews as part of the genetic landscape of the Middle East. American Journal of Human Genetics 2001, 69, 1095–1112. [Google Scholar] [CrossRef]
- Ostrer, H.; Skorecki, K. The population genetics of the Jewish people. Human Genetics 2013, 132, 119–127. [Google Scholar] [CrossRef]
- Rodríguez-Varela, R.; Günther, T.; Krzewińska, M.; et al. Genomic analyses of pre-European conquest human remains from the Canary Islands. Current Biology 2017, 27, 3396–3402.e5. [Google Scholar] [CrossRef]
- Roth, C. History of the Jews in Italy; Jewish Publication Society: Philadelphia, 1946. [Google Scholar]
- Rutgers, L.V. The Jews in Late Ancient Rome; Brill: Leiden, 1995. [Google Scholar]
- Sarno, S.; Boattini, A.; Pagani, L.; Sazzini, M.; De Fanti, S.; Quagliariello, A.; Gnecchi-Ruscone, G.A.; Guichard, E.; Ciani, G.; Bortolini, E.; Barbieri, C.; Cilli, E.; Petrilli, R.; Mikerezi, I.; Sineo, L.; Vilar, M.; Wells, S.; Luiselli, D.; Pettener, D. Ancient and recent admixture layers in Sicily and Southern Italy trace multiple migration routes along the Mediterranean. Scientific Reports 2017, 7, 39844. [Google Scholar] [CrossRef] [PubMed]
- Schuenemann, V.J.; Krause, J.; et al. Ancient Egyptian mummy genomes suggest an increase of Sub-Saharan African ancestry in post-Roman periods. Nature Communications 2017, 8, 15694. [Google Scholar] [CrossRef]
- Semino, O.; Magri, C.; Benuzzi, G.; et al. Origin, diffusion, and differentiation of Y-chromosome haplogroups E and. J. American Journal of Human Genetics 2004, 74, 1023–1034. [Google Scholar] [CrossRef]
- Shen, P.; Lavi, T.; Kivisild, T.; et al. Reconstruction of patrilineages and matrilineages of Samaritans. Human Mutation 2004, 24, 248–260. [Google Scholar] [CrossRef]
- Tian, C.; Kosoy, R.; Nassir, R.; Lee, A.; Villoslada, P.; Klareskog, L.; Hammarström, L.; Garchon, H.J.; Pulver, A.E.; Ransom, M.; Gregersen, P.K.; Seldin, M.F. European population genetic substructure. Molecular Medicine 2009, 15, 371–383. [Google Scholar] [CrossRef] [PubMed]
- Waldman, S.; Backenroth, D.; Harney, É.; et al. Genome-wide data from medieval German Jews. Cell 2022, 185, 4703–4716.e16. [Google Scholar] [CrossRef] [PubMed]
- Xue, J.; Lencz, T.; Darvasi, A.; Pe’er, I.; Carmi, S. The time and place of European admixture in Ashkenazi Jewish history. PLOS Genetics 13(4), e1006644. [CrossRef]
- Zoossmann-Diskin, A. The origin of Eastern European Jews revealed by autosomal, sex chromosomal and mtDNA polymorphisms. Biology Direct 2010, 5, 57. [Google Scholar] [CrossRef]
| Study | Ashkenazi sample size | E total | E-M78 | E-M123 | J1-M267 | J2-M172 | Notes |
|---|---|---|---|---|---|---|---|
| Hammer et al. 2009 | 1,575 (pooled) | NR | ~3 | ~17 | ~17 | ~20 | Ashkenazi and non-Ashkenazi Jews pooled; subgroup frequencies not reported |
| Behar et al. 2004 | ~442 | ~16.1 | NR | NR | ~19 | ~19 | No Southern European comparators; coarse resolution |
| Semino et al. 2004 | 77 | NR | 5.2 | 11.7 | 14.6 | 23.2 | Only study including Southern Italians; higher subclade resolution |
| Nebel et al. 2001 | ~79 | 23 | NR | NR | ~19 | ~24 | No Southern European comparators |
| Shen et al. 2004 | ~20 | NR | ~10 | ~10 | ~20 | ~15 | Very small sample; limited resolution |
| Population | Sample size | E-M78 | E-M123 | J1-M267 | J2-M172 |
|---|---|---|---|---|---|
| Ashkenazi Jews | 77 | 5.2 | 11.7 | 14.6 | 23.2 |
| Italian (Apulia) | 86 | 0.0 | 11.6 | 2.3 | 29.1 |
| Italian (Calabria 1) | 80 | 16.3 | 2.5 | 1.8 | 22.8 |
| Italian (Calabria 2, Albanian community, Cosenza) | 68 | 5.9 | 13.2 | 0.0 | 20.0 |
| Italian (Sicily) | 55 | 12.7 | 3.6 | 7.1 | 16.7 |
| Model components | p value | Feasible | Italian component (%) | Lebanese (%) | Russian (%) |
|---|---|---|---|---|---|
| Italian_South.HO + Lebanese.HO + Russian.DG | 0.1445 | Yes | 69.97 | 22.06 | 7.97 |
| Italian_North.HO + Lebanese.HO + Russian.DG | 0.0256 | No | 54.53 | 42.57 | 2.90 |
| Italian_Central.HO + Lebanese.HO + Russian.DG | 0.0514 | Yes | 61.33 | 33.83 | 4.84 |
| Population | FST distance | Standard error |
|---|---|---|
| Cretan.DG | 0.0049 | 0.00124 |
| Italian_South.HO | 0.0049 | 0.00033 |
| Sicilian.HO | 0.0052 | 0.00041 |
| Greek.HO | 0.0055 | 0.00035 |
| Maltese.HO | 0.0066 | 0.00045 |
| Jew_Moroccan.HO | 0.0091 | 0.00058 |
| Jew_Iraqi.DG | 0.0122 | 0.00138 |
| Jew_Iranian.HO | 0.0125 | 0.00050 |
| Jew_Tunisian.HO | 0.0127 | 0.00056 |
| Jew_Libyan.HO | 0.0132 | 0.00053 |
| Jew_Georgian.HO | 0.0133 | 0.00056 |
| Jew_Yemenite.HO | 0.0174 | 0.00054 |
| Population pair | FST distance | Interpretation |
|---|---|---|
| Italian_South.HO vs Maltese.HO | 0.0036 | Closely related central Mediterranean populations |
| Jew_Ashkenazi.HO vs Italian_South.HO | 0.0049 | Strong regional affinity |
| IBS_CanaryIslands.DG vs Spanish.HO | 0.0038 | Iberian-derived population with additional North African ancestry |
| Rank | Population | Distance |
|---|---|---|
| 1 | Maltese | 0.0239 |
| 2 | Italian Calabria | 0.0242 |
| 3 | Sicilian Central | 0.0264 |
| 4 | Italian Campania | 0.0273 |
| 5 | Italian Campania Naples (Campanian) | 0.0277 |
| 6 | Greek Dodecanese | 0.0289 |
| 7 | Sicilian East | 0.0290 |
| 8 | Sicilian Syracuse | 0.0291 |
| 9 | Sicilian | 0.0295 |
| 10 | Greek Crete Lasithi | 0.0303 |
| 11 | Greek Crete | 0.0305 |
| 12 | Sicilian Trapani | 0.0309 |
| 13 | Greek Dodecanese Kos | 0.0310 |
| 14 | Greek Kos | 0.0310 |
| 15 | Italian Calabria (Cosentian) | 0.0310 |
| 16 | Greek Cyprus | 0.0322 |
| 17 | Cypriot | 0.0326 |
| 18 | Italian Basilicata | 0.0326 |
| 19 | Italian Basilicata (Lucanian) | 0.0328 |
| 20 | Greek Crete Rethymno | 0.0331 |
| 21 | Italian Apulia | 0.0341 |
| 22 | Turkish Cyprus | 0.0342 |
| 23 | Greek Cyclades Amorgos | 0.0350 |
| 24 | Sicilian West | 0.0357 |
| 25 | Greek Euboea Central | 0.0365 |
| 26 | Italian Apulia (Apulian) | 0.0368 |
| 27 | Greek Crete Heraklion | 0.0381 |
| 28 | Greek Crete Chania | 0.0385 |
| 29 | Italian Campania Benevento (Campanian) | 0.0389 |
| 30 | Italian Campania Salerno (Campanian) | 0.0391 |
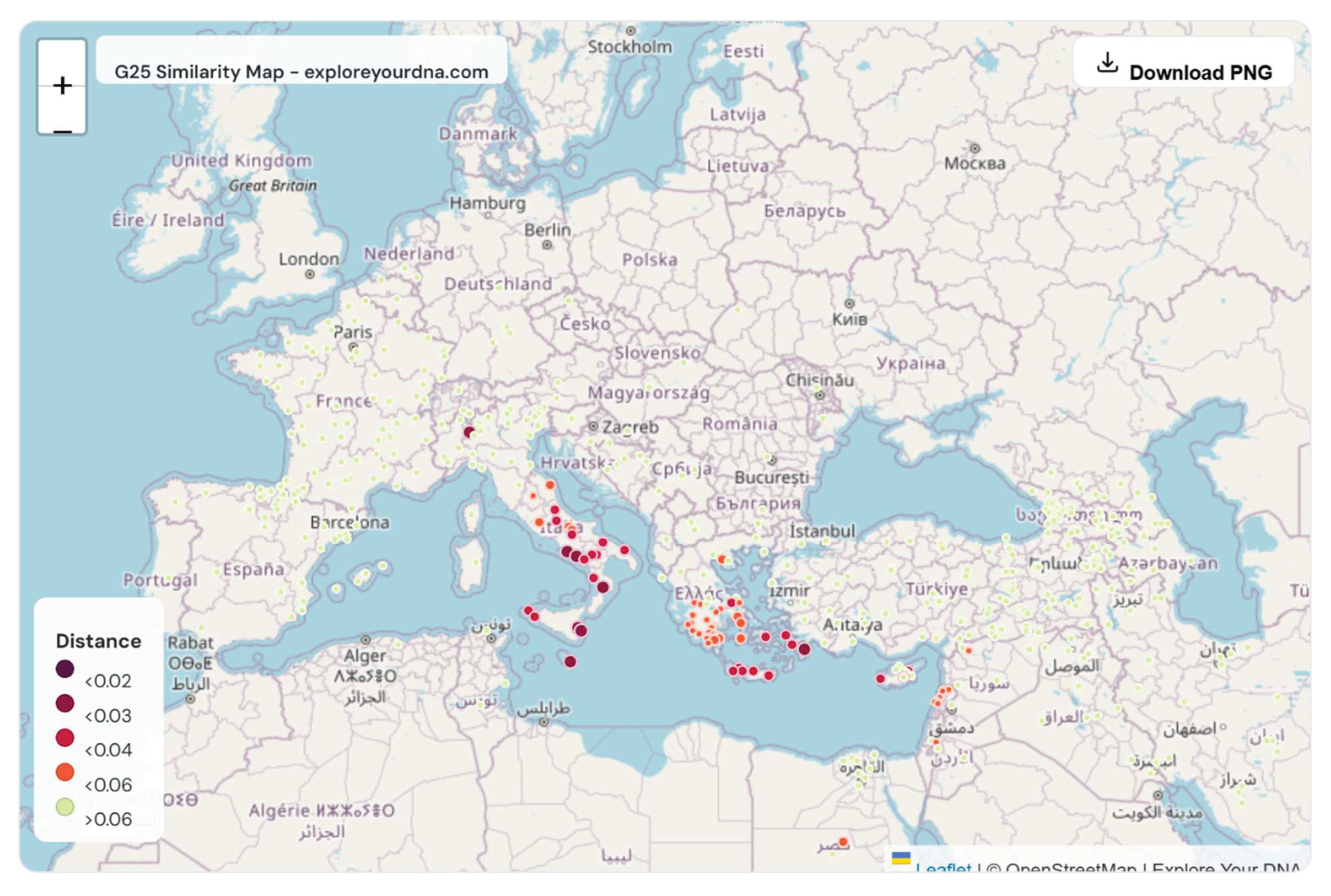
| The closest affinities localize overwhelmingly to southern Italy, Sicily, Malta, and the adjacent Aegean. Northern Italian populations are absent from the closest similarity ranks. This result reproduces the same Southern Italian anchoring observed under PCA, FST, qpAdm, and clustering analyses and does not depend on admixture proportion assumptions. Taken together, these findings impose a hard constraint on viable origin models. If Ashkenazi Jews derive approximately fifty percent of their ancestry from European sources and that European component is Italian, while Italian Jews remained resident in southern Italy and Sicily, then both populations must derive that Italian ancestry from the same Southern Italian genetic substrate. There is no defensible mechanism by which Ashkenazi Jews could acquire substantial Northern Italian, Western European, or Slavic ancestry while Italian Jews, who remained geographically closer to any Italian source population, did not. Models invoking convergent Levantine–European admixture do not resolve this contradiction, as they require independent mixtures to converge on nearly identical Southern Italian autosomal placement while entirely bypassing Northern Italian populations. Italian Jews therefore function as a population-level falsification test. Their autosomal position demonstrates that Italian ancestry acquired and retained within Italy manifests as Southern Italian and central Mediterranean structure rather than as Northern Italian or hybrid Northern European structure. Consequently, the Italian component consistently detected in Ashkenazi Jews must derive from Southern Italy and the adjacent Aegean genetic continuum, with Ashkenazi-specific founder effects and later Eastern European admixture shaping variation within that space rather than redefining its origin. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




