Submitted:
26 January 2026
Posted:
28 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Design and Data Collection
2.2. Inclusion and Exclusion Criteria
2.3. Sample Size and Sampling Procedure
2.4. Dependent and Independent Variables
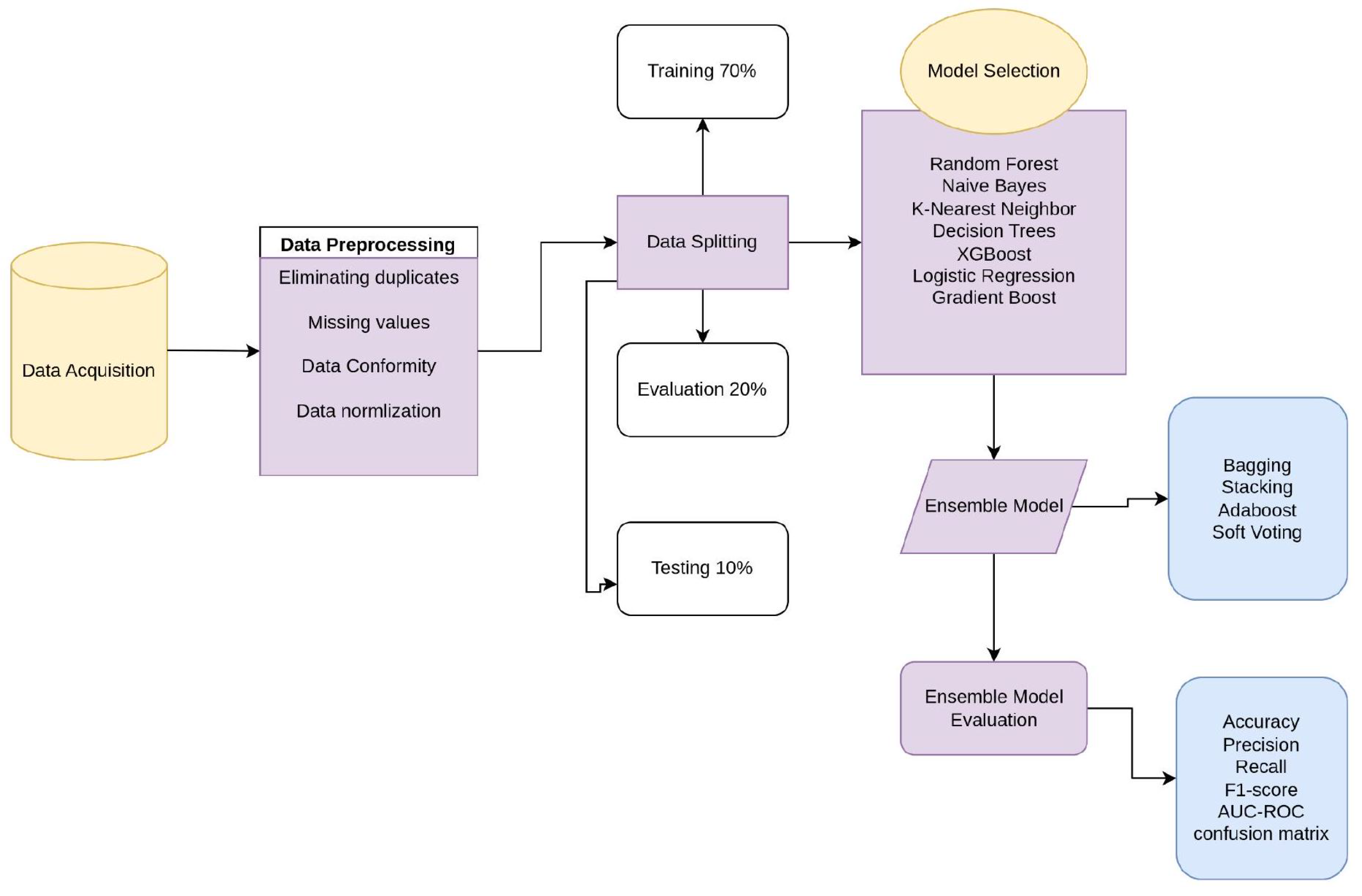
2.5. Data Preprocessing
2.6. Data Cleaning
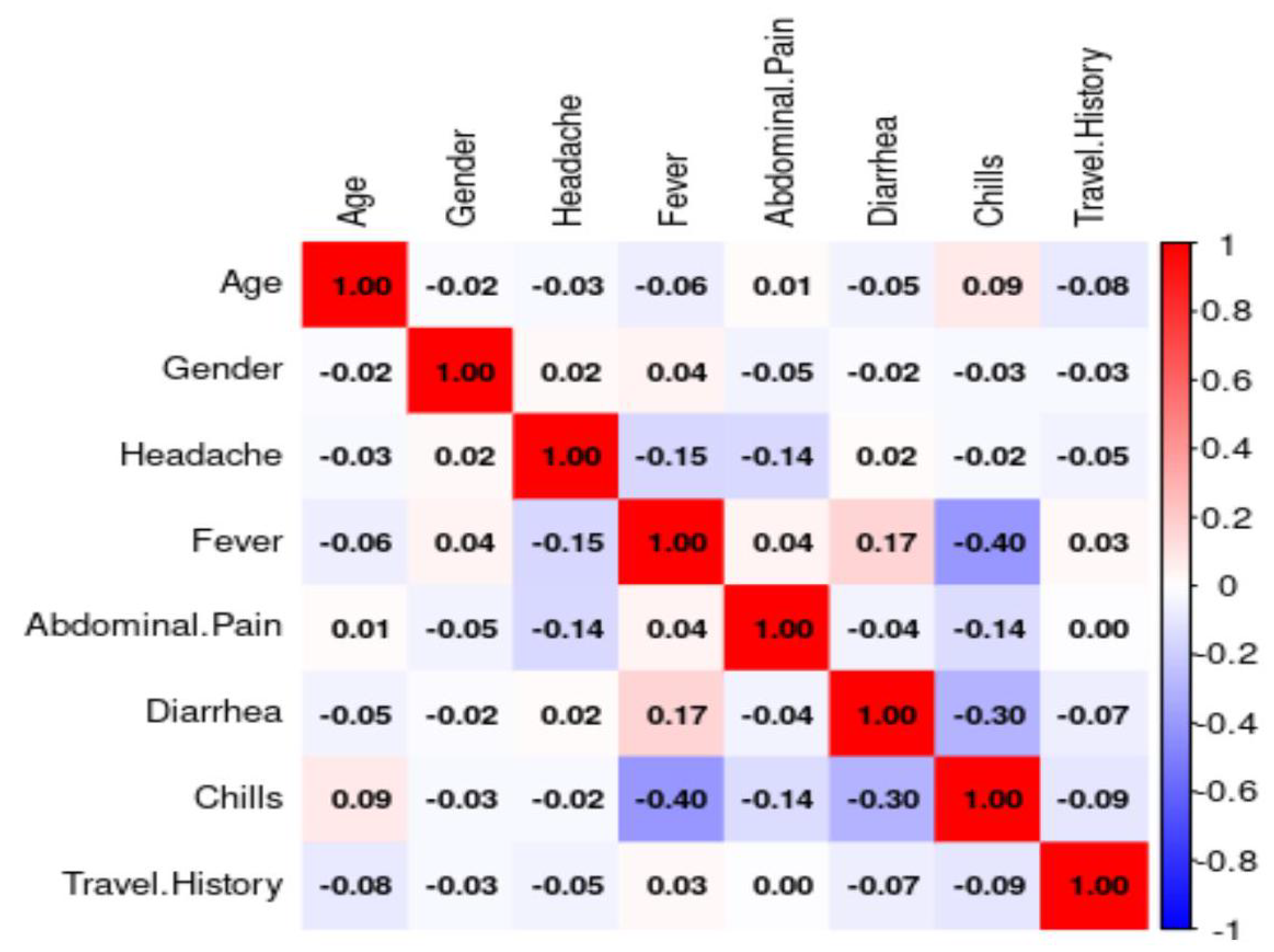
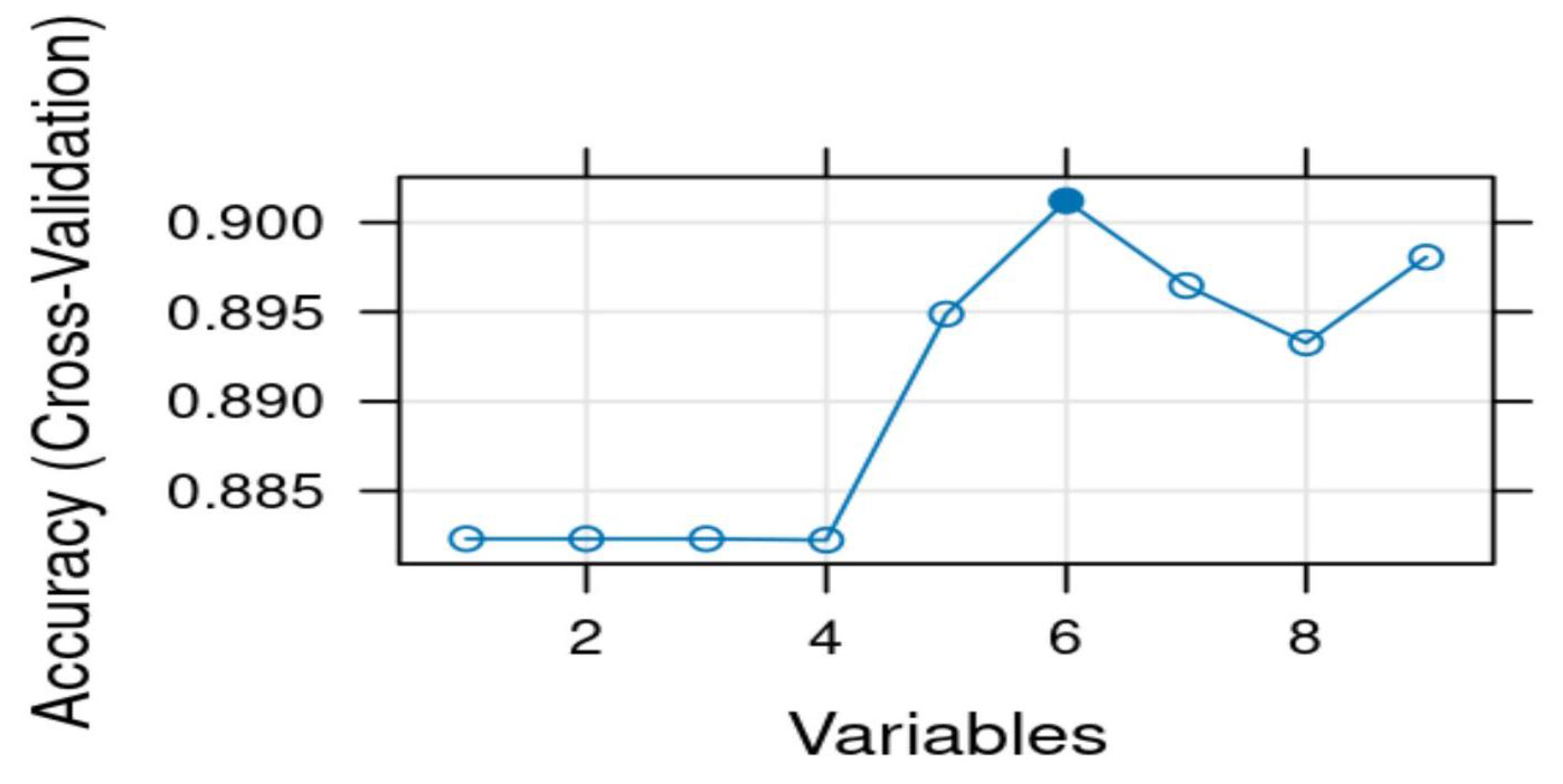
2.7. Feature Selection
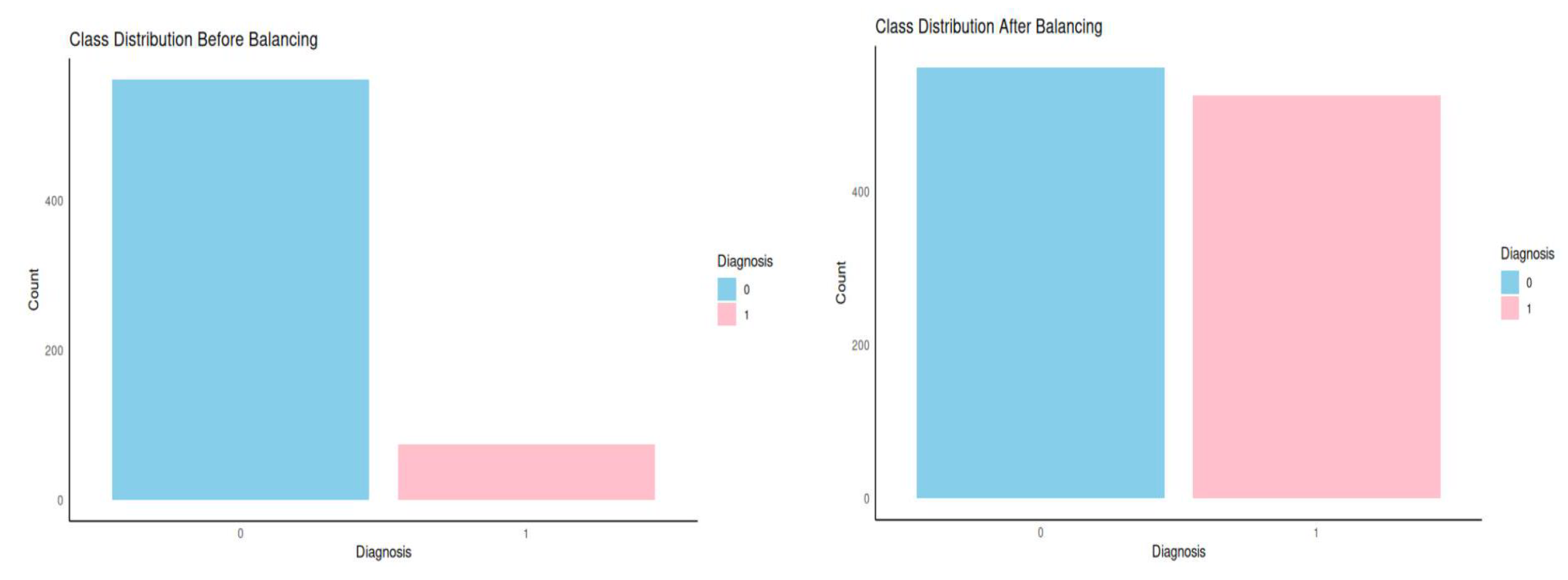
2.8. Handling Class Imbalance
2.9. Data Encoding
2.10. Data Splitting
2.11. Individual Model Selection and Training
2.12. Hyperparameter Tuning
2.13. Ensemble Model Selection and Building
2.14. Individual and Ensemble Model Performance Evaluation
2.15. Ethical Review
3. Results
3.1. Analysis of Demographic and Clinical Variables Associated with Malaria Diagnosis
3.2. Feature Selection
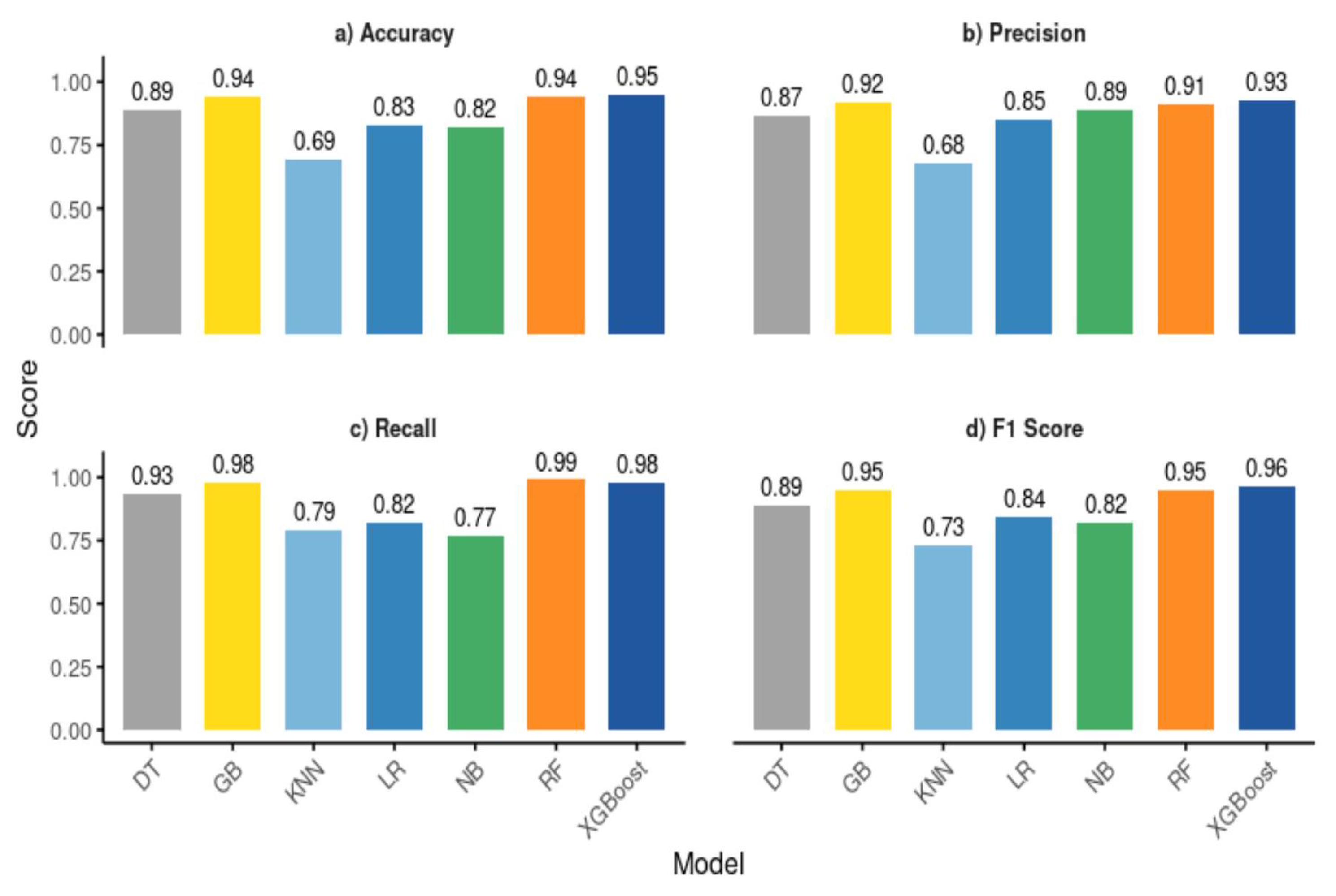
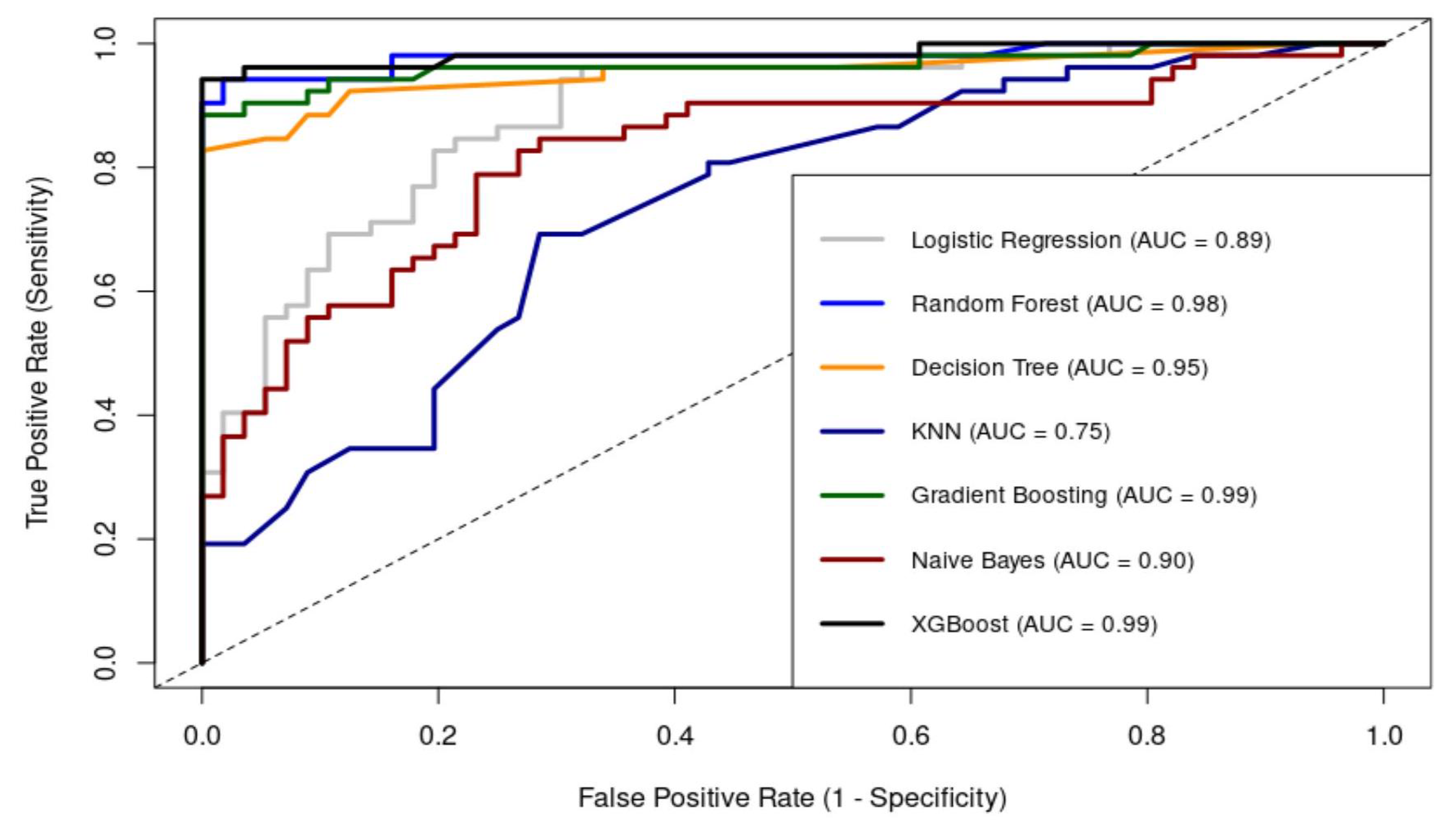
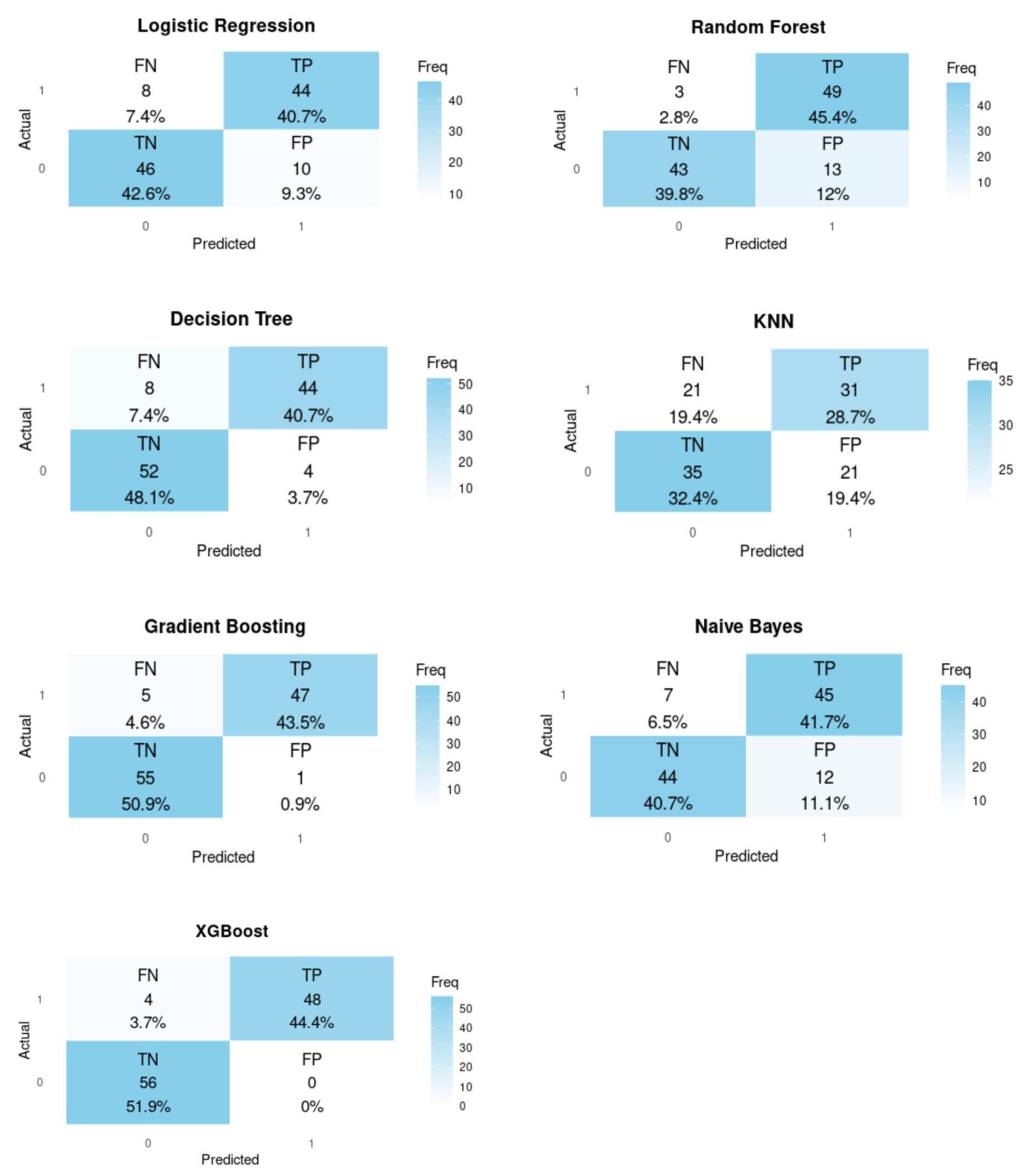
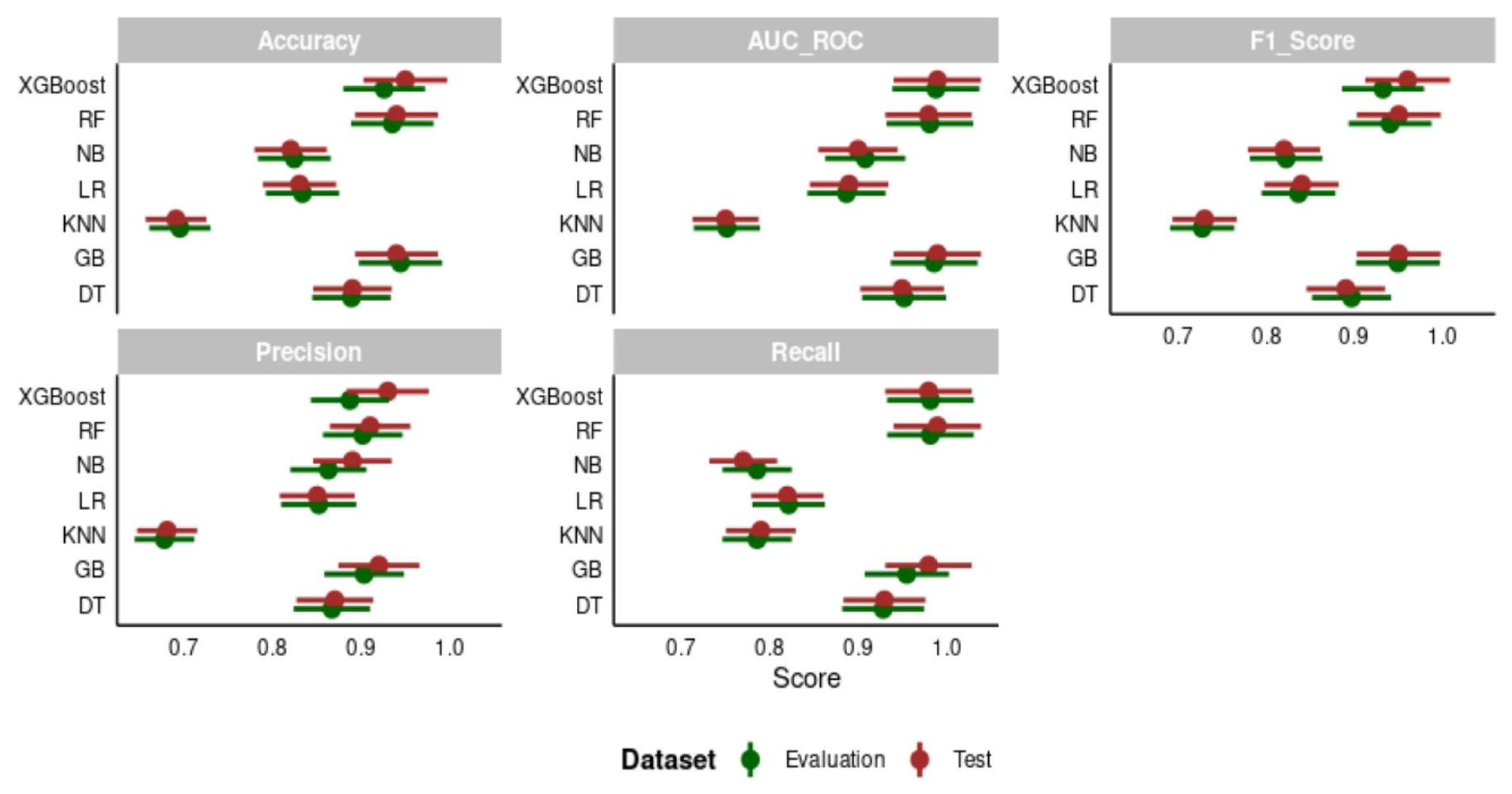
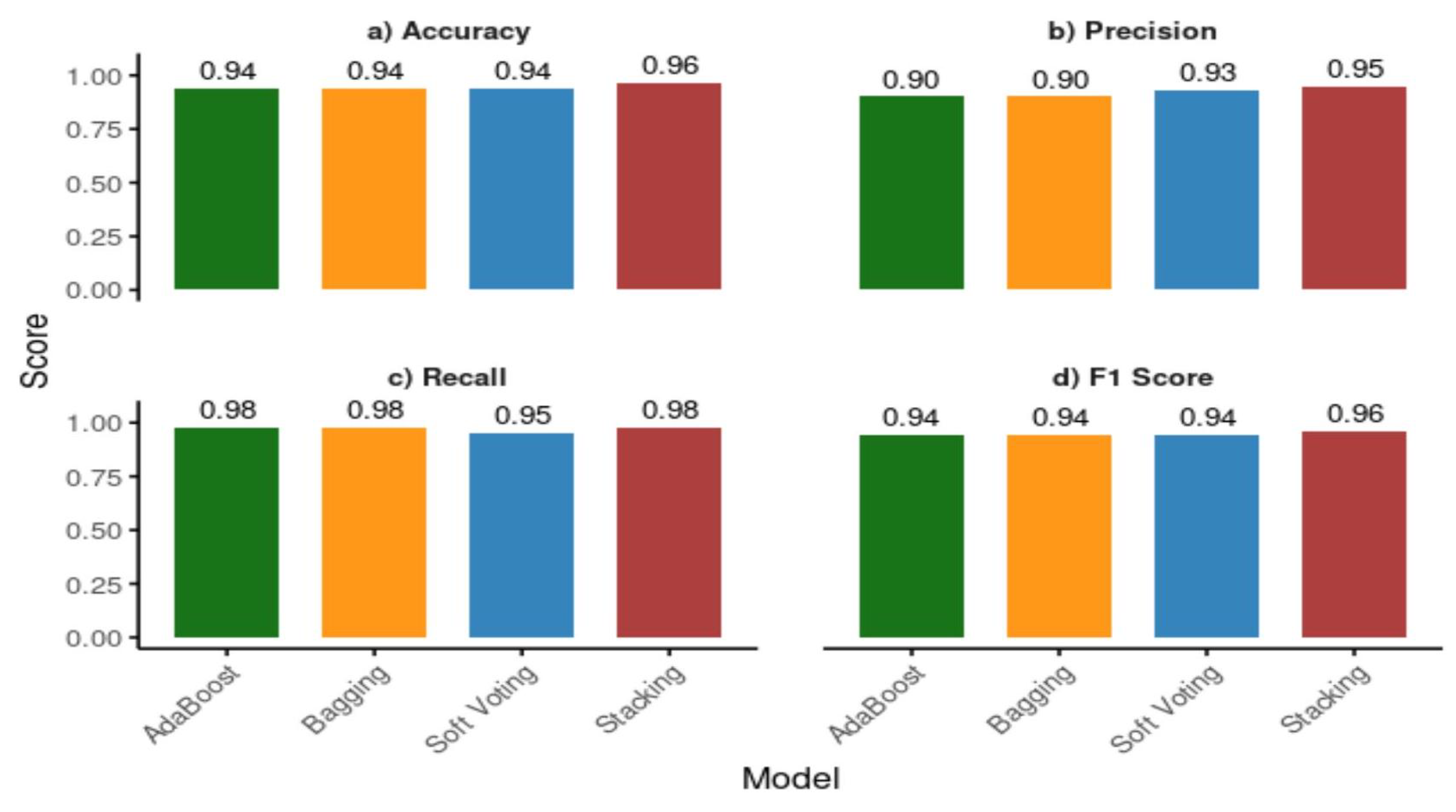
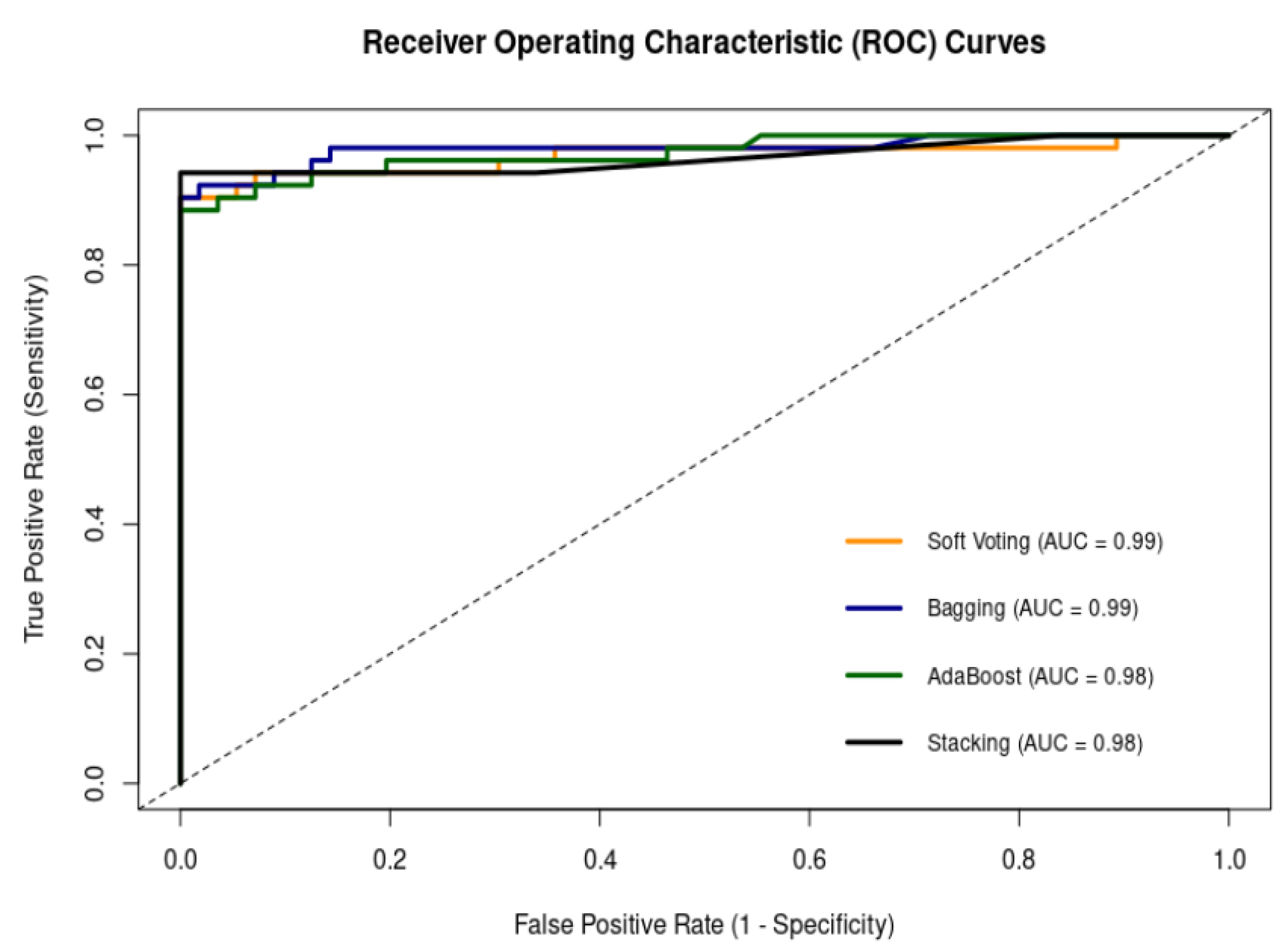
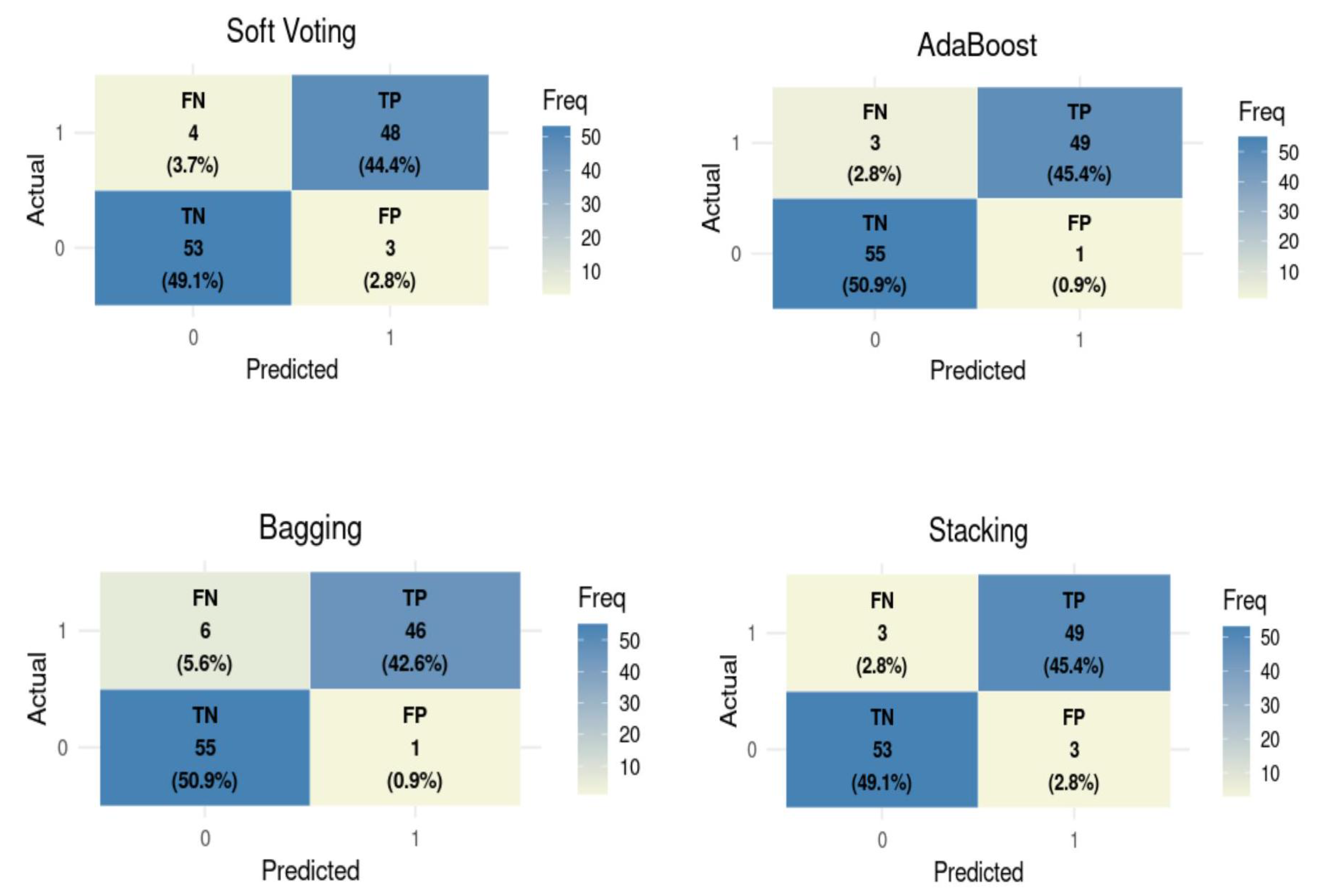
3.3. Machine Learning
4. Discussion
5. Conclusion
Funding
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ML | Machine Learning |
| AI | Artificial Intelligence |
| LR | Logistic Regression |
| RF | Random Forest |
| DT | Decision Trees |
| GB | Gradient Boosting |
| KNN | K-Nearest Neighbors |
| NB | Naive Bayes |
| AUC-ROC | Area Under the Receiver Operating Characteristic Curve |
| SMOTE | Synthetic Minority Oversampling Technique |
| mRDTs | malaria Rapid Diagnostic Tests |
| ARIMA | Autoregressive Integrated Moving Average |
| STL+ARIMA | Seasonal and Trend Decomposition using Loess |
| BP-ANN | Back Propagation Artificial Neural Network |
| LSTM | Long Short-Term Memory |
References
- Hamid, M.M.A.; Mohamed, A.O.; Mohammed, F.O.; Elaagip, A.; Mustafa, S.A.; Elfaki, T.; Jebreel, W.M.; Albsheer, M.M.; Dittrich, S.; Owusu, E.D.; et al. Diagnostic accuracy of an automated microscope solution (miLab™) in detecting malaria parasites in symptomatic patients at point-of-care in Sudan: a case–control study. Malaria Journal 2024, 23, 200. [CrossRef]
- Organization, W.H.; et al. WHO guidelines for malaria, 13 august 2025 2025.
- Rajab, S.; Nakibuule, R.; Marvin, G. Interpretable Ensemble Model-Agonistc Approaches for Malaria Prediction. In Proceedings of the Proceedings of the 2024 Sixteenth International Conference on Contemporary Computing, 2024, pp. 451–459.
- Kagaba Amina, G. Socio-Economic Determinants and Malaria Risk: Assessing the Impact of Poverty, Housing Conditions, and Healthcare Accessibility in High-Incidence Regions. SCIENCES (NIJRMS) 2024, 5, 120–4.
- Moyo-Ndlovu, T. Malaria claims three more lives in Zimbabwe, total deaths rise to 32. https://www.heraldonline.co.zw/malaria-claims-three-more-lives-in-zimbabwe-total-deaths-rise-to-32/, 2024. Accessed: 12 January 2025.
- Kabalu Tshiongo, J.; Zola Matuvanga, T.; Mitashi, P.; Maketa, V.; Schallig, H.D.; Mens, P.F.; Muhindo Mavoko, H.; Matangila Rika, J. Prevention of malaria in pregnant women and its effects on maternal and child health, the case of centre hospitalier de Kingasani II in the Democratic Republic of the congo. Tropical Medicine and Infectious Disease 2024, 9, 92. [CrossRef]
- Sato, S. Plasmodium—a brief introduction to the parasites causing human malaria and their basic biology. Journal of physiological anthropology 2021, 40, 1. [CrossRef]
- Wang, M.; Wang, H.; Wang, J.; Liu, H.; Lu, R.; Duan, T.; Gong, X.; Feng, S.; Liu, Y.; Cui, Z.; et al. A novel model for malaria prediction based on ensemble algorithms. PloS one 2019, 14, e0226910. [CrossRef]
- Mfuh, K.O.; Achonduh-Atijegbe, O.A.; Bekindaka, O.N.; Esemu, L.F.; Mbakop, C.D.; Gandhi, K.; Leke, R.G.; Taylor, D.W.; Nerurkar, V.R. A comparison of thick-film microscopy, rapid diagnostic test, and polymerase chain reaction for accurate diagnosis of Plasmodium falciparum malaria. Malaria journal 2019, 18, 73. [CrossRef]
- Varo, R.; Balanza, N.; Mayor, A.; Bassat, Q. Diagnosis of clinical malaria in endemic settings. Expert Review of Anti-infective Therapy 2021, 19, 79–92. [CrossRef]
- Wongsrichanalai, C.; Barcus, M.J.; Muth, S.; Sutamihardja, A.; Wernsdorfer, W.H. A review of malaria diagnostic tools: microscopy and rapid diagnostic test (RDT). Defining and Defeating the Intolerable Burden of Malaria III: Progress and Perspectives: Supplement to Volume 77 (6) of American Journal of Tropical Medicine and Hygiene 2007. [CrossRef]
- Ojeniyi, F.D.; Ayoola, A.O.; Ibitoye, O.; Opaleye, O.O.; Olowe, O.A.; Ehigie, L.O.; Thomas, B.N.; Ojurongbe, O. Performance and challenges of malaria rapid diagnostic tests in endemic regions of Africa. Scientific Reports 2025. [CrossRef]
- Maitland, K. Management of severe paediatric malaria in resource-limited settings. BMC medicine 2015, 13, 42. [CrossRef]
- Ayoka, T.O.; Nnadi, C.O. Application of machine learning models in predicting malaria prevalence in nigeria: an analysis of the 2015–2020 demographic and health surveys. Journal of Parasitic Diseases 2025, pp. 1–13. [CrossRef]
- Stephen, A.; Akomolafe, P.O.; Ogundoyin, K.I. A model for predicting malaria outbreak using machine learning technique. Annals Computer Science Series 2021, 19.
- Muriithi, D.; Lumumba, V.; Okongo, M. A Machine Learning-Based Prediction of Malaria Occurrence in Kenya. American Journal of Theoretical and Applied Statistics 2024, 13, 65–72. [CrossRef]
- Mariki, M. Machine learning model for prediction of malaria in low and high endemic areas of Tanzania. PhD thesis, NM-AIST, 2023.
- SAHU, P.; Priyadarshini, P.; Tripathy, S.; Das, Y.; Pradhan, S. Machine learning strategies for malaria risk prediction based on text-based clinical information 2023. [CrossRef]
- for Africa Network, D.D. The potential of digital molecular diagnostics for infectious diseases in sub-Saharan Africa. PLOS Digital Health 2022, 1, e0000064.
- Cunnington, A. The Digital Diagnostics for Africa Network (2022) The potential of digital molecular diagnostics for infectious diseases in sub-Saharan Africa. PLOS Digit Health 1 (6): e0000064, 2022.
- Devi, S.S.; Solanki, V.K.; Laskar, R.H. Recent advances on big data analysis for malaria prediction and various diagnosis methodologies. Handbook of data science approaches for biomedical engineering 2020, pp. 153–184.
- Cho, S.; Kim, S.; Kim, Y.; Park, Y. Optical imaging techniques for the study of malaria. Trends in biotechnology 2012, 30, 71–79. [CrossRef]
- Awe, O.O.; Mwangi, P.N.; Goudoungou, S.K.; Esho, R.V.; Oyejide, O.S. Explainable AI for enhanced accuracy in malaria diagnosis using ensemble machine learning models. BMC Medical Informatics and Decision Making 2025, 25, 162. [CrossRef]
- Muhammad, B.; Varol, A. A symptom-based machine learning model for malaria diagnosis in Nigeria. In Proceedings of the 2021 9th International Symposium on Digital Forensics and Security (ISDFS). IEEE, 2021, pp. 1–6.
- Mbunge, E.; Milham, R.C.; Sibiya, M.N.; Takavarasha Jr, S. Machine learning techniques for predicting malaria: Unpacking emerging challenges and opportunities for tackling malaria in sub-saharan Africa. In Proceedings of the Computer Science On-line Conference. Springer, 2023, pp. 327–344.
- Khan, O.; Ajadi, J.O.; Hossain, M.P. Predicting malaria outbreak in The Gambia using machine learning techniques. PLoS One 2024, 19, e0299386. [CrossRef]
- Ly, A.Y.; Allaya, M.M.; Loum, M.A.; Gning, L.D.; Sy, O.; Bakhoum, L.N.; Ndiaye, K.; Ndiop, M.; Sall, F.B.; Ndiaye, J.L.A.; et al. Forecasting malaria incidence in the Southeast districts of Senegal using a machine learning approach. BMC Artificial Intelligence 2025, 1, 9. [CrossRef]
- Brown, B.J.; Manescu, P.; Przybylski, A.A.; Caccioli, F.; Oyinloye, G.; Elmi, M.; Shaw, M.J.; Pawar, V.; Claveau, R.; Shawe-Taylor, J.; et al. Data-driven malaria prevalence prediction in large densely populated urban holoendemic sub-Saharan West Africa. Scientific Reports 2020, 10, 15918. [CrossRef]
- Rubuga, F.K.; Ahmed, A.; Siddig, E.; Sera, F.; Moirano, G.; Aimable, M.; Albert, T.; Gallican, N.R.; Nebié, E.I.; Kitema, G.F.; et al. Potential impact of climatic factors on malaria in Rwanda between 2012 and 2021: a time-series analysis. Malaria Journal 2024, 23, 274. [CrossRef]
- Gavi, S.; Tapera, O.; Mberikunashe, J.; Kanyangarara, M. Malaria incidence and mortality in Zimbabwe during the COVID-19 pandemic: analysis of routine surveillance data. Malaria journal 2021, 20, 233. [CrossRef]
- Chivasa, T.; Nunu, W.N.; Dhlamini, M.; Maviza, A.; Ndagurwa, G.N. Impact of Climate Change on Malaria Transmission and Management in Zimbabwe: A Scoping Review of the Literature. Health Services Insights 2025, 18, 11786329251374245. [CrossRef]
- Lakshmi, T.J.; Prasad, C.S.R. A study on classifying imbalanced datasets. In Proceedings of the 2014 First international conference on networks & soft computing (ICNSC2014). IEEE, 2014, pp. 141–145.
- Sameh, B.; Khodadadi, N.; Khodadadi, E.; Eid, M.; El-kenawy, E.S. Advancements and Future Directions in Machine Learning for Medical Diagnostics: A Comprehensive Review 2024. 7, 18–31. [CrossRef]
- Rahman, M.M.; Davis, D.N. Addressing the class imbalance problem in medical datasets. International Journal of Machine Learning and Computing 2013, 3, 224. [CrossRef]
- Chawla, N.V.; Bowyer, K.W.; Hall, L.O.; Kegelmeyer, W.P. SMOTE: synthetic minority over-sampling technique. Journal of artificial intelligence research 2002, 16, 321–357. [CrossRef]
- Ranjha, R.; Singh, K.; Mohan, M.; Anvikar, A. Age-specific Malaria Vulnerability and Transmission Reservoir among Children. Global Pediatrics 2023, 6, 100085. [CrossRef]
- Khagayi, S.; Desai, M.; Amek, N.; Were, V.; Onyango, E.; Otieno, K.; Bigogo, G.; Munga, S.; Odhiambo, F.; Hamel, M.; et al. Modelling the relationship between malaria prevalence as a measure of transmission and mortality across age groups. Malaria Journal 2019, 18. [CrossRef]
- Carneiro, I.; Roca-Feltrer, A.; Griffin, J.; Smith, L.; Tanner, M.; Schellenberg, J.; Greenwood, B.; Schellenberg, D. Age-Patterns of Malaria Vary with Severity, Transmission Intensity and Seasonality in Sub-Saharan Africa: A Systematic Review and Pooled Analysis. PloS one 2010, 5, e8988. [CrossRef]
- Yadav, S.S.; Kadam, V.J.; Jadhav, S.M.; Jagtap, S.; Pathak, P.R. Machine learning based malaria prediction using clinical findings. In Proceedings of the 2021 International Conference on Emerging Smart Computing and Informatics (ESCI). IEEE, 2021, pp. 216–222.
- Mienye, I.D.; Sun, Y. A survey of ensemble learning: Concepts, algorithms, applications, and prospects. Ieee Access 2022, 10, 99129–99149. [CrossRef]
- Shaikh, T.A.; Rasool, T.; Verma, P.; Mir, W.A. A fundamental overview of ensemble deep learning models and applications: systematic literature and state of the art. Annals of Operations Research 2024, pp. 1–77. [CrossRef]
- Grandvalet, Y. Bagging equalizes influence. Machine Learning 2004, 55, 251–270. [CrossRef]
- Sarmah, U.; Borah, P.; Bhattacharyya, D.K. Ensemble learning methods: An empirical study. SN Computer Science 2024, 5, 924. [CrossRef]
- Altman, N.; Krzywinski, M. Ensemble methods: bagging and random forests. Nature Methods 2017, 14, 933–935. [CrossRef]
- Kumar, M.; Singhal, S.; Shekhar, S.; Sharma, B.; Srivastava, G. Optimized stacking ensemble learning model for breast cancer detection and classification using machine learning. Sustainability 2022, 14, 13998. [CrossRef]
- Javed, D. MULTI-ENSEMBLE ARCHITECTURE FOR NETWORK INTRUSION DETECTION: A STACKING, VOTING, AND HYBRID ADABOOST-RANDOM FOREST APPROACH ON CIC-IDS2024. Spectrum of Engineering Sciences 2025, pp. 967–981.
- Li, X.; Wang, L.; Sung, E. A study of AdaBoost with SVM based weak learners. In Proceedings of the Proceedings. 2005 IEEE International Joint Conference on Neural Networks, 2005. IEEE, 2005, Vol. 1, pp. 196–201.
- Atukunda, A.; Deogratius, M.A.; Arinaitwe, E.; Orishaba, P.; Kamya, M.R.; Nankabirwa, J.I. Do clinicians in areas of declining malaria transmission adhere to malaria diagnosis guidelines? A cross-sectional study from Kampala, Uganda. Malaria Journal 2021, 20, 187. [CrossRef]
- Doreswamy, S.; Al Sudani, H. Malaria Falciparum: Relapse After a Decade. Cureus 2022, 14. [CrossRef]
- Lynch, C.A.; Bruce, J.; Bhasin, A.; Roper, C.; Cox, J.; Abeku, T.A. Association between recent internal travel and malaria in Ugandan highland and highland fringe areas. Tropical medicine & international health 2015, 20, 773–780.
- Brungard, C.W.; Boettinger, J.L.; Duniway, M.C.; Wills, S.A.; Edwards Jr, T.C. Machine learning for predicting soil classes in three semi-arid landscapes. Geoderma 2015, 239, 68–83. [CrossRef]
- Ekusai-Sebatta, D.; Arinaitwe, E.; Mpimbaza, A.; Nankabirwa, J.I.; Drakeley, C.; Rosenthal, P.J.; Staedke, S.G.; Muyinda, H. Challenges and opportunities for use of long-lasting insecticidal nets to prevent malaria during overnight travel in Uganda: a qualitative study. Malaria journal 2021, 20, 283. [CrossRef]
- Arinaitwe, E.; Dorsey, G.; Nankabirwa, J.I.; Kigozi, S.P.; Katureebe, A.; Kakande, E.; Rek, J.; Rosenthal, P.J.; Drakeley, C.; Kamya, M.R.; et al. Association between recent overnight travel and risk of malaria: a prospective cohort study at 3 sites in Uganda. Clinical Infectious Diseases 2019, 68, 313–320. [CrossRef]
- Mahajan, M.; Kalra, A.S.; Bansal, A.; Bhattacherjee, A.; Jain, E.; et al. Automated Malaria Detection Using Deep Learning: A Convolutional Neural Network Approach for Microscopic Cell Image Analysis. In Proceedings of the 2024 International Conference on Decision Aid Sciences and Applications (DASA). IEEE, 2024, pp. 1–6.
- Rajab, S.; Nakatumba-Nabende, J.; Marvin, G. Interpretable Machine Learning Models for Predicting Malaria. 04 2023, pp. 1–6. [CrossRef]
| Feature Name | Feature Description | DataType | Levels | Encoding |
|---|---|---|---|---|
| Residence | Living Environment | Categorical | Gutu = 1, Gweru = 2 | 1,2 |
| Age | Patient’s age in years | Integer | 0 to 95 | Continuous (Binning) |
| Gender | Biological sex of a patient | Categorical | Male , Female | 1,0 |
| Headache | Presence of headache symptom | Binary Integer | Yes , No | 1,0 |
| Fever | Presence of fever symptom | Binary Integer | Yes , No | 1,0 |
| Abdominal Pain | Presence of abdominal Pain symptom | Binary Integer | Yes , No | 1,0 |
| Diarrhea | Presence of diarrhea symptom | Binary Integer | Yes , No | 1,0 |
| Chills | Sudden cold sensations | Binary Integer | Yes , No | 1,0 |
| Travel History | Recent travel to malaria-endemic areas | Binary Integer | Yes , No | 1,0 |
| Diagnosis | Malaria diagnosis outcome | Categorical | Positive , Negative | 1,0 |
| Variable | Category | Negative () | Positive () | Total () | p-value |
|---|---|---|---|---|---|
| Gender | Male | 283 (50.4%) | 41 (54.7%) | 324 (50.9%) | 0.483 |
| Female | 279 (49.6%) | 34 (45.3%) | 313 (49.1%) | ||
| Fever | Yes | 371 (66.0%) | 63 (84.0%) | 434 (68.1%) | 0.002 ** |
| No | 191 (34.0%) | 12 (16.0%) | 203 (31.9%) | ||
| Chills | Yes | 350 (62.3%) | 61 (81.3%) | 411 (64.5%) | 0.001 ** |
| No | 212 (37.7%) | 14 (18.7%) | 226 (35.5%) | ||
| Headache | Yes | 521 (92.7%) | 69 (92.0%) | 590 (92.6%) | 0.826 |
| No | 41 (7.3%) | 6 (8.0%) | 47 (7.4%) | ||
| Diarrhea | Yes | 149 (26.5%) | 31 (41.3%) | 180 (28.3%) | 0.007 ** |
| No | 413 (73.5%) | 44 (58.7%) | 457 (71.7%) | ||
| Abdominal Pain | Yes | 166 (29.5%) | 38 (50.7%) | 204 (32.0%) | <0.001 ** |
| No | 396 (70.5%) | 37 (49.3%) | 433 (68.0%) | ||
| Travel History | Yes | 198 (35.2%) | 37 (49.3%) | 235 (36.9%) | 0.017 * |
| No | 364 (64.8%) | 38 (50.7%) | 402 (63.1%) | ||
| Location | Rural | 287 (51.1%) | 49 (65.3%) | 336 (52.7%) | 0.020 * |
| Urban | 275 (48.9%) | 26 (34.7%) | 301 (47.3%) | ||
| Age Group | 0-5 | 92 (16.4%) | 8 (10.7%) | 100 (15.7%) | 0.298 |
| 6-15 | 62 (11.0%) | 6 (8.0%) | 68 (10.7%) | ||
| 16-30 | 160 (28.5%) | 19 (25.3%) | 179 (28.1%) | ||
| 31-45 | 104 (18.5%) | 21 (28.0%) | 125 (19.6%) | ||
| 46-60 | 89 (15.8%) | 11 (14.7%) | 100 (15.7%) | ||
| >60 | 55 (9.8%) | 10 (13.3%) | 65 (10.2%) |
| Variable | VIF Value |
|---|---|
| Chills | 1.35 |
| Fever | 1.24 |
| Diarrhoea | 1.12 |
| Headache | 1.06 |
| Abdominal Pain | 1.06 |
| Residence | 1.04 |
| Travel History | 1.04 |
| Age | 1.03 |
| Gender | 1.01 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




