Submitted:
27 January 2026
Posted:
28 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Sampling
2.2. Micromorphological Characterization of Trichomes
2.2.1. Optical and Stereomicroscopy
2.2.2. Scanning Electron Microscopy (SEM)
2.3. Histological Analysis of Pericarp Thickness
2.4. Assessment of Ploidy Levels
2.5. Transcriptomic Profiling and Bioinformatic Analysis
2.6. Statistical Analysis
3. Results
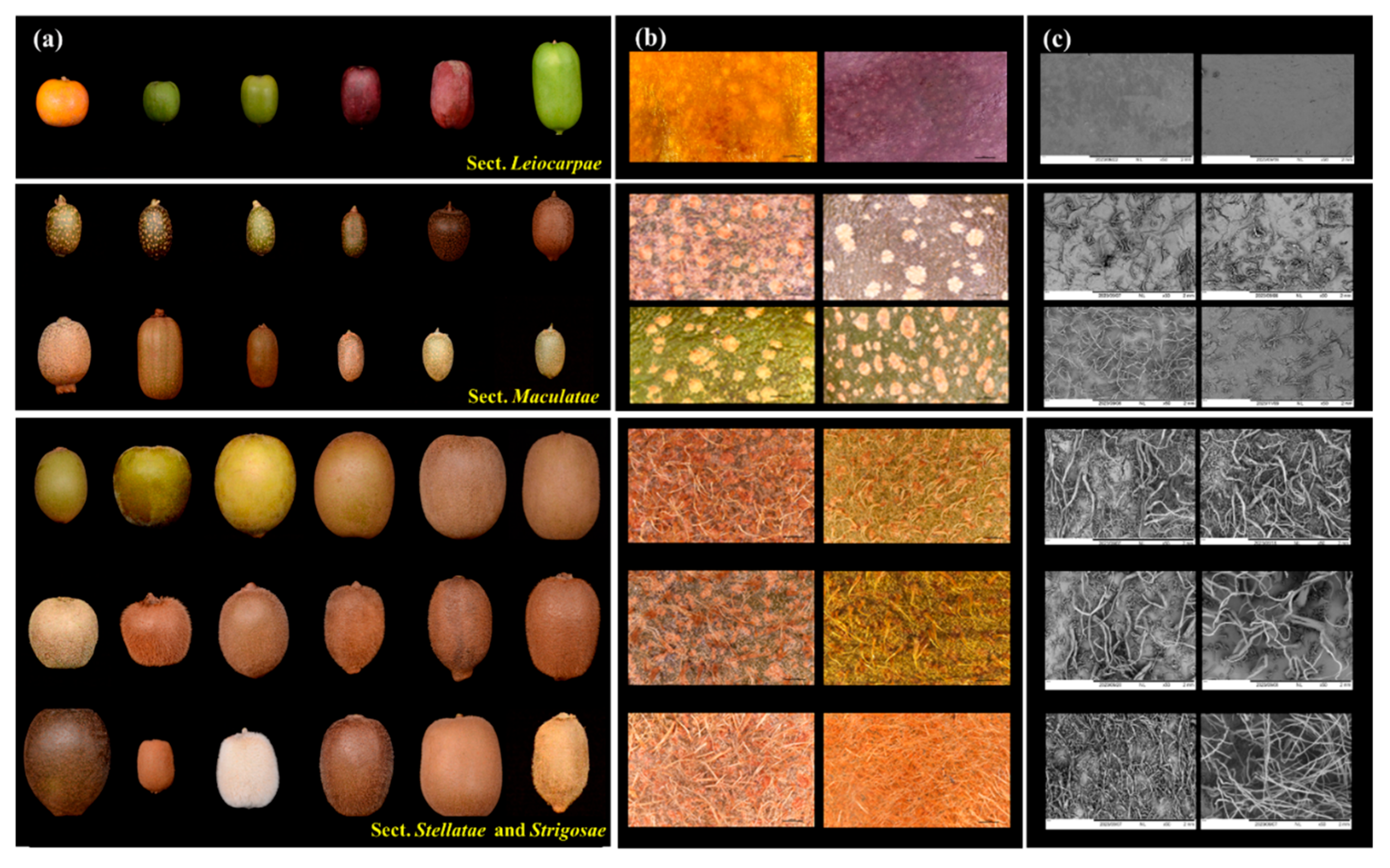
3.1. Morphological Diversity of Kiwifruit Epidermis and Trichomes
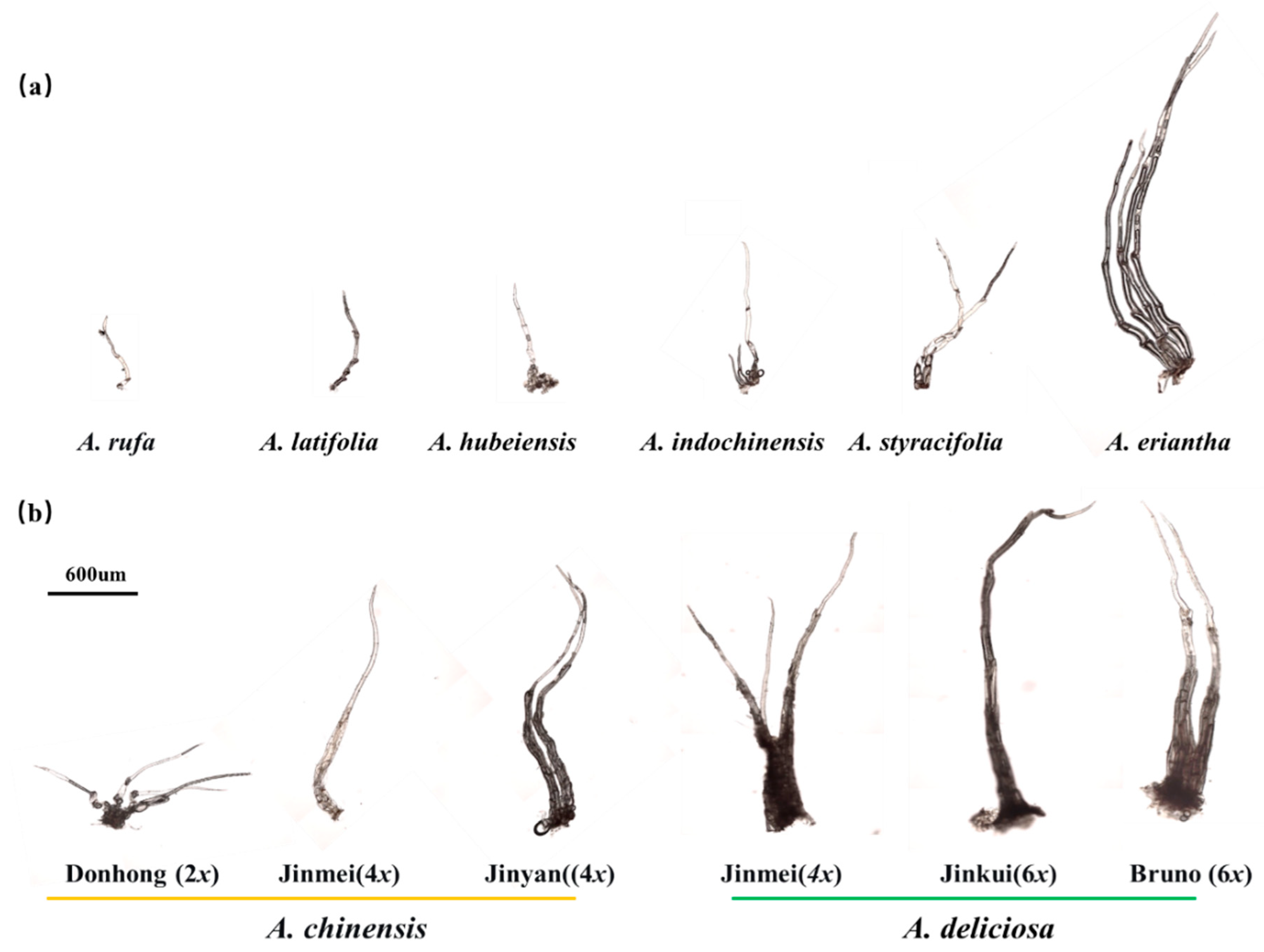
3.2. Cytological Architecture of Kiwifruit Multicellular Trichomes
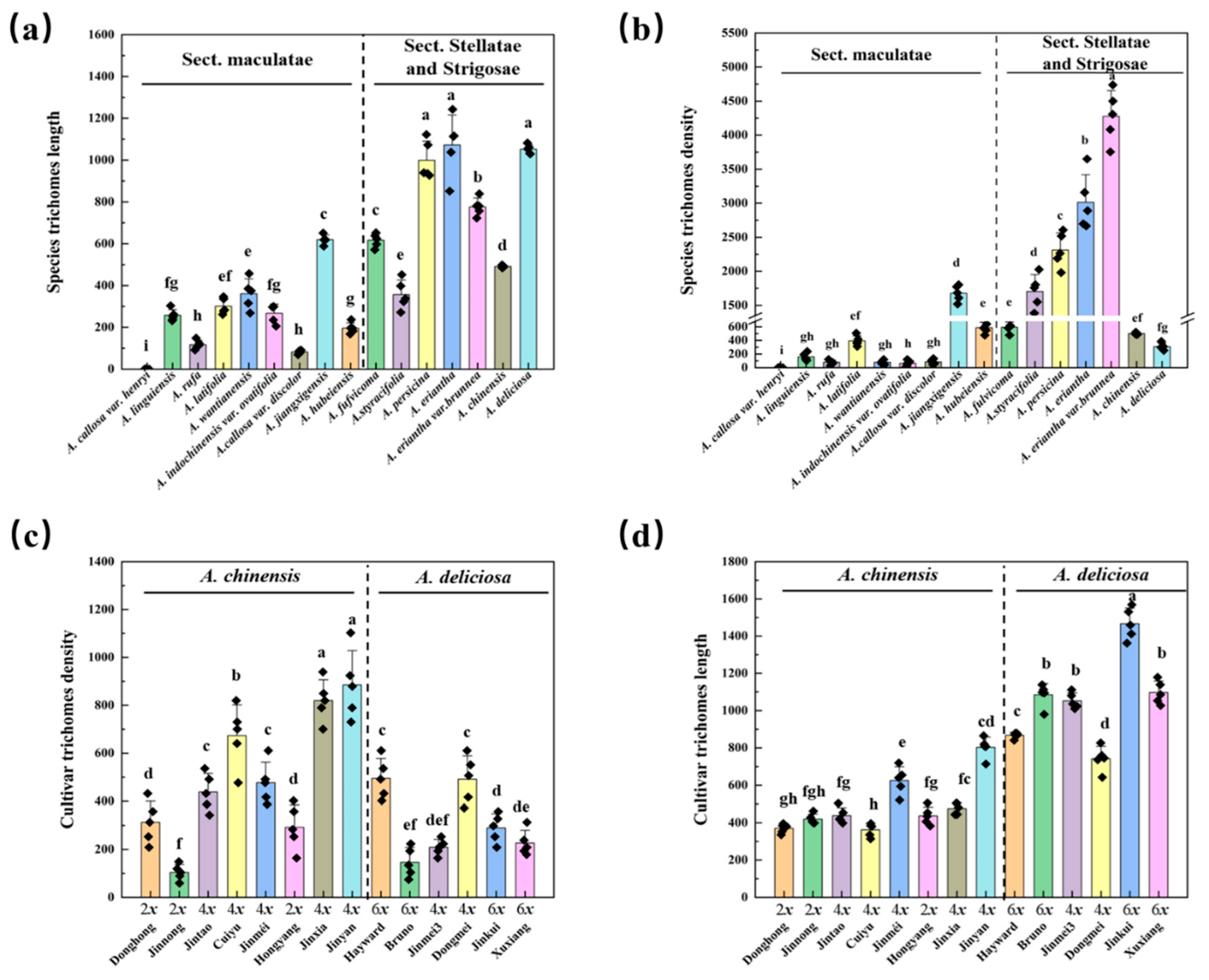
3.3. Micromorphological and Density Variation Across Actinidia Taxa
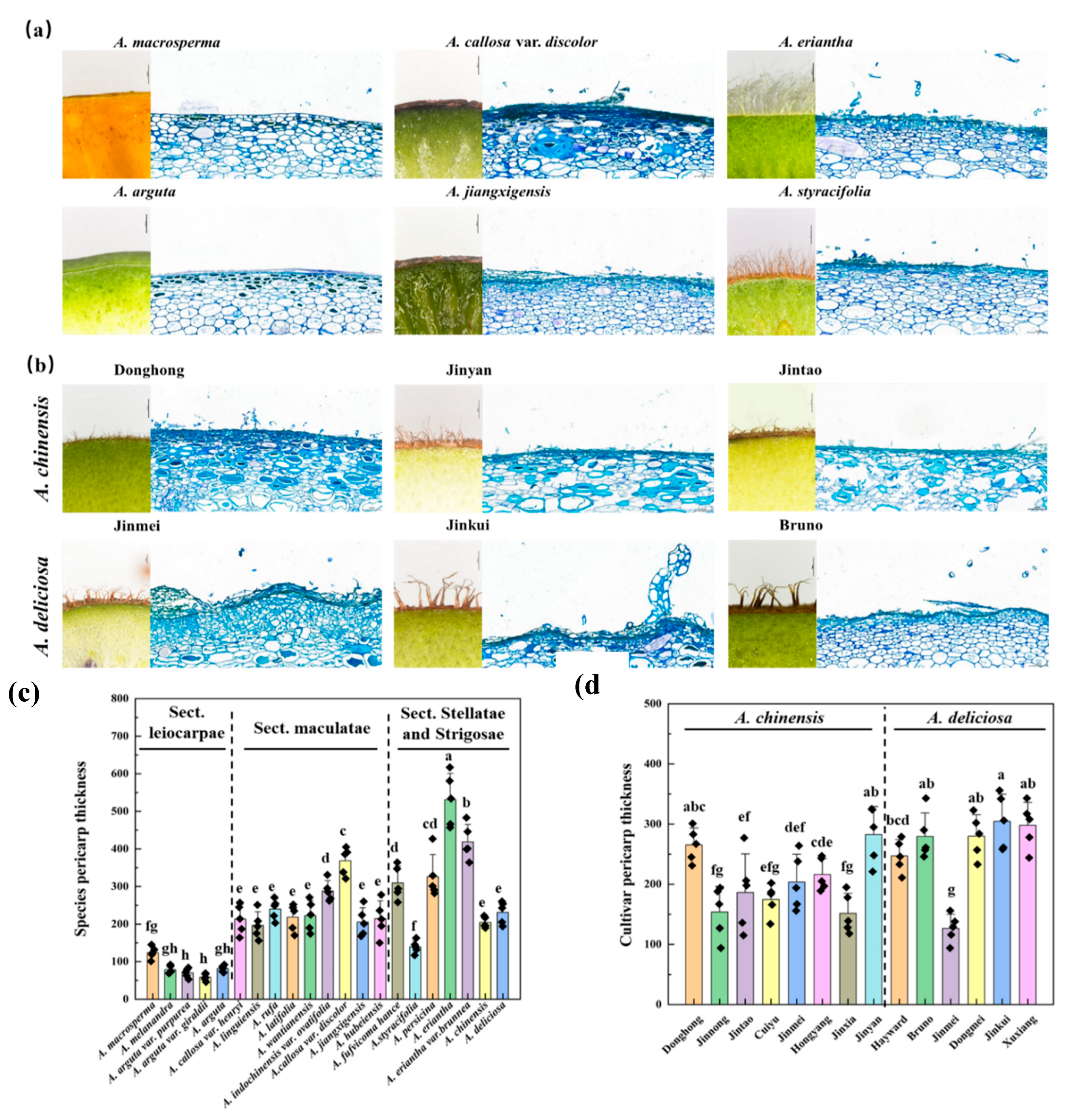
3.4. Comparative Analysis of Pericarp Thickness
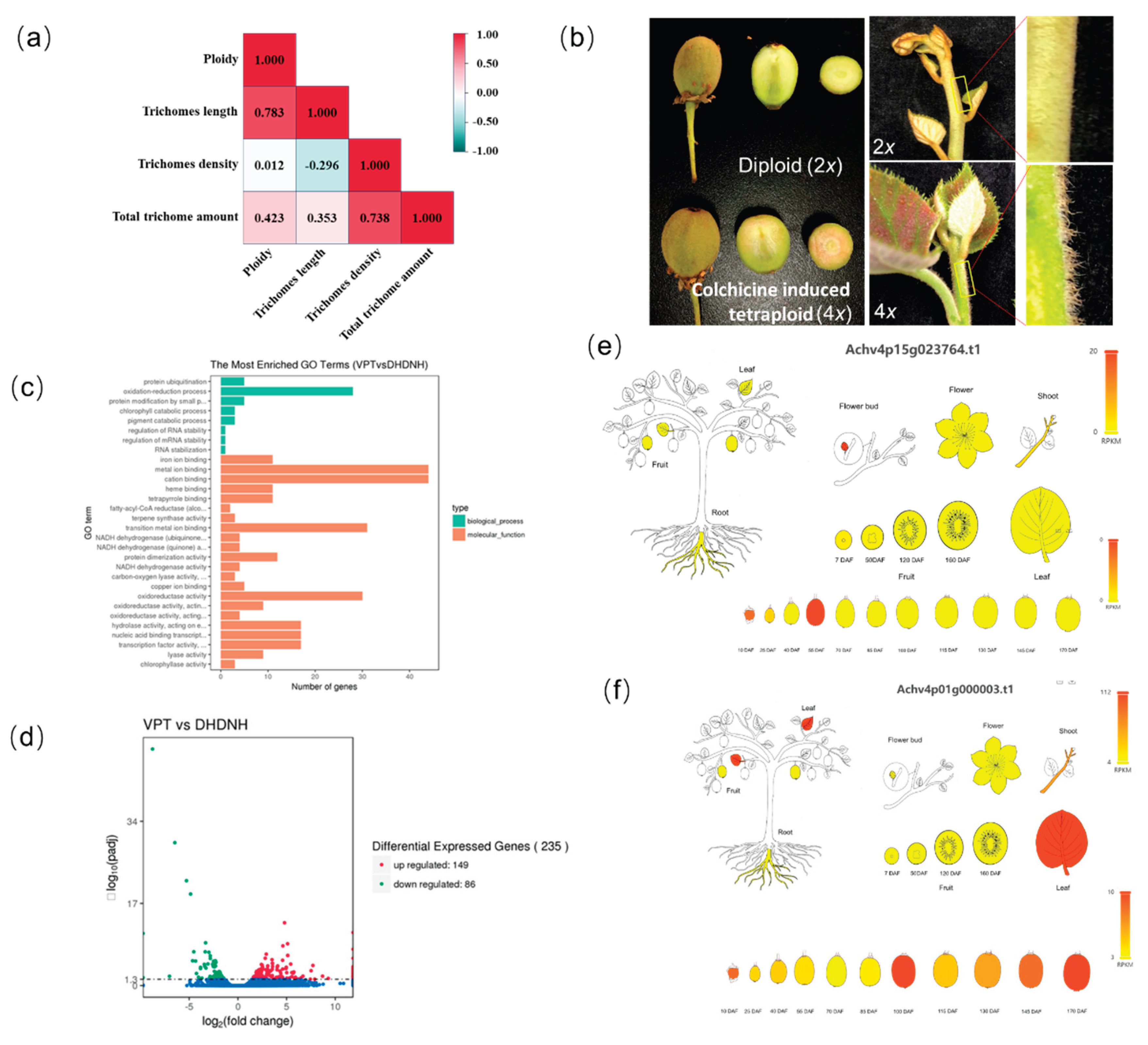
3.5. Transcriptomic Analysis of Ploidy-Induced Trichome Morphogenesis
4. Discussion
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Werker, E. Trichome diversity and development. Advances in Botanical Research 2000, 31, 1–35. [Google Scholar] [CrossRef]
- Chalvin, C.; Drevensek, S.; Dron, M.; Bendahmane, A.; Boualem, A. Genetic control of glandular trichome development. Trends in Plant Science 2020, 25, 477–487. [Google Scholar] [CrossRef] [PubMed]
- Mauricio, R. Costs of resistance to natural enemies in field populations of the annual plant Arabidopsis thaliana. The American Naturalist 1998, 152, 265–272. [Google Scholar] [CrossRef]
- Ran, R.; Li, X.; Zhang, J.; Zhao, J.; Zhao, X.; Cui, X.; Chen, G.; Zhao, P. Monocot-like leaf structure and trichome-water relations in early growth stages of the C3 plant sand rice (Agriophyllum squarrosum). Plant Science 2025, 112480. [Google Scholar] [CrossRef]
- Oksman-Caldentey, K.; Inzé, D. Plant cell factories in the post-genomic era: new ways to produce designer secondary metabolites. Trends in Plant Science 2004, 9, 433–440. [Google Scholar] [CrossRef]
- Cao, J.; Zhao, Y.; Tang, K. MYC: orchestrating secondary metabolism and glandular trichome formation. Trends in Plant Science 2025, 30, 821–825. [Google Scholar] [CrossRef]
- Wang, H.; Ren, J.; Zhou, S.; Duan, Y.; Zhu, C.; Chen, C.; Liu, Z.; Zheng, Q.; Xiang, S.; Xie, Z.; Wang, X.; Chai, J.; Ye, J.; Xu, Q.; Guo, W.; Deng, X.; Zhang, F. Molecular regulation of oil gland development and biosynthesis of essential oils in Citrus spp. Science 2024, 383, 659–666. [Google Scholar] [CrossRef] [PubMed]
- Feng, Z.; Bartholomew, E.S.; Liu, Z.; Cui, Y.; Dong, Y.; Li, S.; Wu, H.; Ren, Z.; Liu, X. Glandular trichomes: new focus on horticultural crops. Horticulture Research 2021, 8. [Google Scholar] [CrossRef]
- Fu, Y.; Li, M.; Zhang, W.; Liu, X.; Huang, L.; Zhang, S.; Liang, X.; Zhang, L.; Tang, K.; Jocelyn, K.; Shen, Q. The role, regulation and application of plant fruit trichomes. Molecular Horticulture 2025, 5, 41. [Google Scholar] [CrossRef]
- Li, X.; Li, J. Lectotypification of Actinidia. Nordic Journal of Botany 2007, 25(5-6), 294–295. [Google Scholar] [CrossRef]
- Liang, C. The genus Actinidia in China; Science Press: Beijing, 1984. [Google Scholar]
- Wu, M.; Bian, X.; Hu, S.; Huang, B.; Shen, J.; Du, Y.; Wang, Y.; Xu, M.; Xu, H.; Yang, M.; Wu, S. A gradient of the Hd-Zip regulator woolly regulates multicellular trichome morphogenesis in tomato. The Plant Cell 2024, 36, 2375–2392. [Google Scholar] [CrossRef]
- Wu, M.; Bian, X.; Huang, B.; Du, Y.; Hu, S.; Wang, Y.; Shen, J.; Wu, S. HD-Zip proteins modify floral structures for self-pollination in tomato. Science 2024, 384, 124–130. [Google Scholar] [CrossRef]
- Yu, N.; Cai, W.; Wang, S.; Shan, C.; Wang, L.; Chen, X. Temporal control of trichome distribution by microRNA156-targeted SPL genes in Arabidopsis thaliana. The Plant Cell 2010, 22, 2322–2335. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Yang, W.; Dong, G.; Hu, Q.; Li, D.; Liu, J. Construction of the super pan-genome for the genus Actinidia reveals structural variations linked to phenotypic diversity. Horticulture Research 2025, 12, uhaf067. [Google Scholar] [CrossRef]
- Wu, M.; Bian, X.; Hu, S.; Huang, B.; Shen, J.; Du, Y.; Wang, Y.; Xu, M.; Xu, H.; Yang, M.; Wu, S. A gradient of the HD-Zip regulator woolly regulates multicellular trichome morphogenesis in tomato. The Plant Cell 2024, 36, 2375–2392. [Google Scholar] [CrossRef]
- Wu, M.; Bian, X.; Huang, B.; Du, Y.; Hu, S.; Wang, Y.; Shen, J.; Wu, S. HD-Zip proteins modify floral structures for self-pollination in tomato. Science 2024, 384, 124–130. [Google Scholar] [CrossRef]
- Yu, N.; Cai, W.; Wang, S.; Shan, C.; Wang, L.; Chen, X. Temporal control of trichome distribution by microRNA156-targeted SPL genes in Arabidopsis thaliana. The Plant Cell 2010, 22, 2322–2335. [Google Scholar] [CrossRef]
- White, J. Ontogeny and morphology of ovarian and fruit hairs in kiwifruit. New Zealand Journal of Botany 1986, 24, 403–414. [Google Scholar] [CrossRef]
- Ferguson, A. The need for characterization and evaluation of germplasm: kiwifruit as an example. Euphytica 2007, 154, 371–382. [Google Scholar] [CrossRef]
- Huang, H.; Li, J.; Ferguson, A. The genus Actinidia: a review of morphological and taxonomic characteristics. Plant Diversity and Resources 2013, 35, 697–706. [Google Scholar]
- Qi, B.; Li, P.; Li, J.; Zha, M.; Wang, F. Kiwifruit Peelability (Actinidia spp.): A Review. Horticulturae 2025, 11, 927. [Google Scholar] [CrossRef]
- Celano, G.; Minnocci, A.; Sebastiani, L.; D’Auria, M.; Xiloyannis, C. Changes in the structure of the skin of kiwifruit in relation to water loss. The Journal of Horticultural Science and Biotechnology 2009, 84, 41–46. [Google Scholar] [CrossRef]
- Wei, N.; Cronn, R.; Liston, A.; Ashman, T. Functional trait divergence and trait plasticity confer polyploid advantage in heterogeneous environments. New Phytologist 2018, 221, 2286–2297. [Google Scholar] [CrossRef] [PubMed]
- Bharati, R.; Gupta, A.; Novy, P.; Severová, L.; Šrédl, K.; Žiarovská, J.; Fernández-Cusimamani, E. Synthetic polyploid induction influences morphological, physiological, and photosynthetic characteristics in Melissa officinalis L. Frontiers in Plant Science 2023, 14. [Google Scholar] [CrossRef]
- Wu, J.; Ferguson, A.; Murray, B.; Jia, Y.; Datson, P.; Zhang, J. Induced polyploidy dramatically increases the size and alters the shape of fruit in Actinidia chinensis. Annals of Botany 2012, 109, 169–179. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



