Submitted:
25 January 2026
Posted:
27 January 2026
You are already at the latest version
Abstract
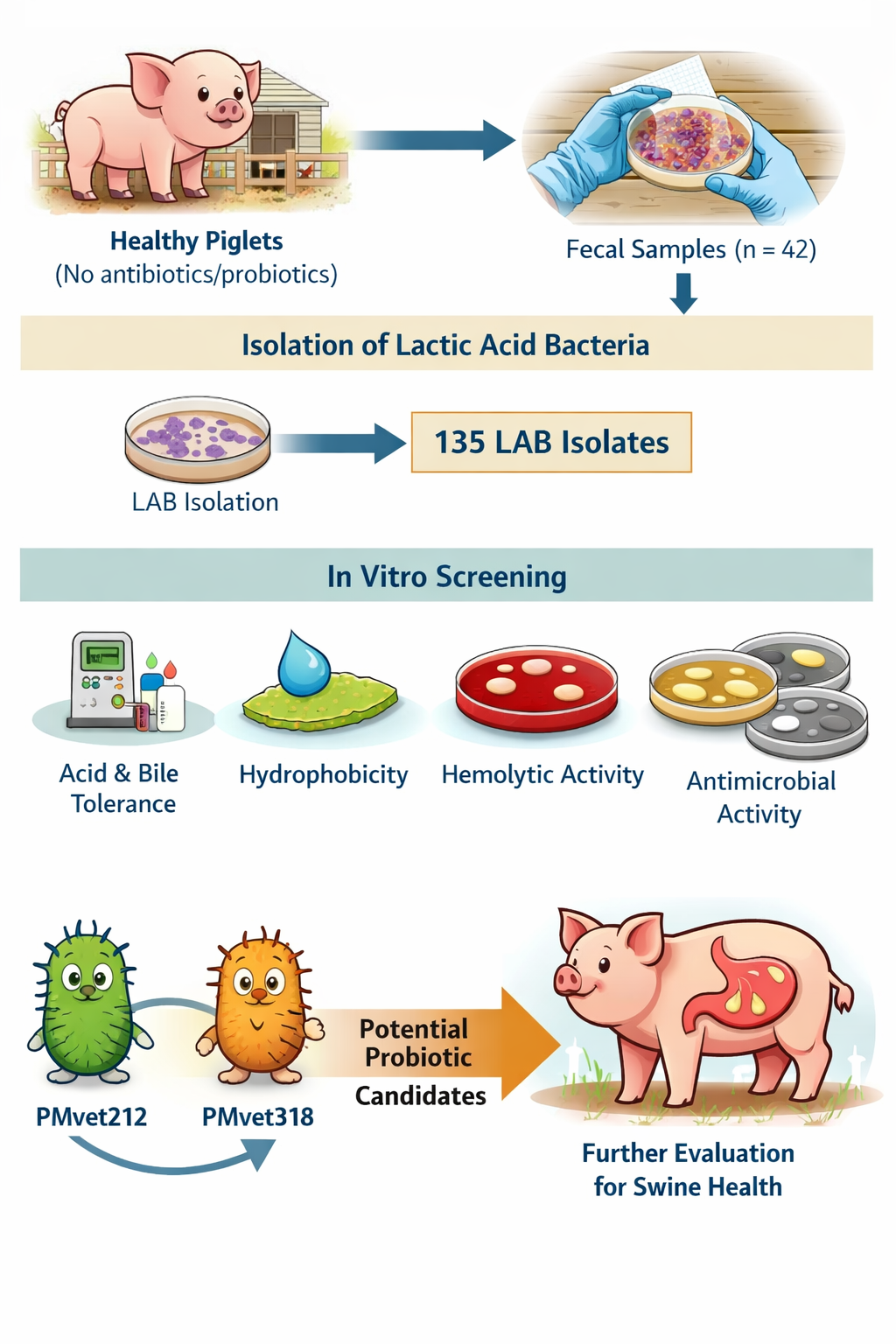
Keywords:
1. Introduction
2. Materials and Methods
2.1. Ethical Approval
2.2. Sample Collection
2.3. Isolation of Lactic Acid Bacteria (LAB)
2.4. Evaluation of Probiotic Properties
2.4.1. Acid Tolerance
2.4.2. Bile Salt Tolerance
2.4.3. Adhesion Ability to Intestinal Epithelium (Surface Hydrophobicity)
2.4.4. Hemolytic Activity
2.4.5. Antimicrobial Activity Against E. coli and S. aureus ATCC 6538
2.5. Data Interpretation Probiotic Properties
2.5.1. Acid Tolerance
2.5.2. Bile Salt Tolerance
2.5.3. Surface Adhesion Ability
2.5.4. Hemolytic Activity
2.5.5. Antimicrobial Activity Against E. coli and S. aureus ATCC 6538
2.6. Statistical Analysis
3. Results
3.1. Isolation of Lactic Acid Bacteria (LAB) from Piglet Feces
3.2. Probiotic Properties of Selected LAB Isolated
3.2.1. Acid Tolerance
3.2.2. Bile Salt Tolerance
3.2.3. Cell Surface Hydrophobicity (MATH Assay)
3.2.4. Hemolytic Activity (Safety Evaluation)
3.2.5. Antimicrobial Activity Against Escherichia coli and Staphylococcus aureus
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Fairbrother, J.M.; Nadeau, E.; Gyles, C.L. Escherichia coli in postweaning diarrhea in pigs: An update on bacterial types, pathogenesis, and prevention strategies. Anim. Health Res. Rev. 2005, 6(1), 17–39. [Google Scholar] [CrossRef]
- Luppi, A. Swine enteric colibacillosis: Diagnosis, therapy, and antimicrobial resistance. Porcine Health Manag. 2017, 3, 16. [Google Scholar] [CrossRef] [PubMed]
- Davis, M.E.; Maxwell, C.V.; Brown, D.C.; de Rodas, B.Z.; Johnson, Z.B.; Kegley, E.B.; Hellwig, D.H.; Dvorak, R.A. Effect of dietary mannan oligosaccharides and/or copper sulfate on growth and immunocompetence of pigs. J. Anim. Sci. 2002, 80, 2887–2894. [Google Scholar] [CrossRef]
- Moredo, F.A.; Pineyro, P.E.; Marquez, G.C.; Sanz, M.; Colello, R.; Etcheverria, A.; Padola, N.L.; Quiroga, M.A.; Perfumo, C.J.; Galli, L.; Leotta, G.A. Enterotoxigenic Escherichia coli subclinical infection in pigs: Bacteriological and genotypic characterization and antimicrobial resistance profiles. Foodborne Pathog. Dis. 2015, 12, 704–711. [Google Scholar] [CrossRef] [PubMed]
- Namkung, H.; Gong, M.; Yu, H.; Cottrill, M.; de Lange, C.F.M. Impact of feeding blends of organic acids and herbal extracts on growth performance, gut microbiota, and digestive function in weaned pigs. Can. J. Anim. Sci. 2004, 84, 697–704. [Google Scholar] [CrossRef]
- Bogere, P.; Choi, Y.J.; Heo, J. Probiotics as alternatives to antibiotics in treating post-weaning diarrhoea in pigs. S. Afr. J. Anim. Sci. 2019, 49, 403–416. [Google Scholar] [CrossRef]
- Laird, T.J.; Abraham, S.; Jordan, D.; Pluske, J.R.; Hampson, D.J.; Trott, D.J.; Dea, M.O. Porcine enterotoxigenic Escherichia coli: Antimicrobial resistance and development of microbial-based alternative control strategies. Vet. Microbiol. 2021, 258, 109117. [Google Scholar] [CrossRef]
- Boaventura, C.; Azevedo, R.; Uetanabaro, A.; Nicoli, J.; Braga, L. The benefits of probiotics in human and animal nutrition. In New Advances in Basic and Clinical Gastroenterology; Brzozowski, T., Ed.; InTech: Rijeka, Croatia, 2012; pp. 75–100. [Google Scholar] [CrossRef]
- Binns, N. Probiotics, Prebiotics and the Gut Microbiota; ILSI Europe Concise Monograph Series: Brussels, Belgium, 2013. [Google Scholar]
- Fouhse, J.M.; Zijlstra, R.T.; Willing, B.P. The role of gut microbiota in the health and disease of pigs. Anim. Front. 2016, 6(3), 30–36. [Google Scholar] [CrossRef]
- FAO/WHO. Probiotics in Food: Health and Nutritional Properties and Guidelines for Evaluation; World Health Organization: Rome, Italy, 2006. [Google Scholar]
- De Angelis, M.; Siragusa, S.; Berloco, M.; Caputo, L.; Settanni, L.; Alfonsi, G.; Amerio, M.; Grandi, A.; Ragni, A.; Gobbetti, M. Selection of potential probiotic lactobacilli from pig feces to be used as additives in pelleted feed. Res. Microbiol. 2006, 157, 792–801. [Google Scholar] [CrossRef]
- Mojgani, N.; Hussaini, F.; Vaseji, N. Characterization of indigenous Lactobacillus strains for probiotic properties. Jundishapur J. Microbiol. 2015, 8, e17523. [Google Scholar] [CrossRef]
- Ozugul, F.; Hamed, I. The importance of lactic acid bacteria for prevention of bacterial growth and biogenic amine formation: A review. Crit. Rev. Food Sci. Nutr. 2018, 58, 1660–1671. [Google Scholar] [CrossRef]
- Senok, A.C.; Ismaeel, A.Y.; Botta, G.A. Probiotics: Facts and myths. Clin. Microbiol. Infect. 2005, 11, 958–966. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Zuo, B.; Li, Q.; Zhao, F.; Wang, J.; Huang, W.; Sun, Z.; Chen, Y. Dietary supplementation with Lactiplantibacillus plantarum P-8 improves the growth performance and gut microbiota of weaned piglets. Microbiol. Spectr. 2024, 12(2), e02345-22. [Google Scholar] [CrossRef]
- Guo, X.H.; Kim, J.M.; Nam, H.M.; Park, S.Y.; Kim, J.M. Screening lactic acid bacteria from swine origins for multistrain probiotics based on in vitro functional properties. Anaerobe 2010, 16, 321–326. [Google Scholar] [CrossRef] [PubMed]
- Ren, D.; Li, C.; Qin, Y.; Yin, R.; Du, S.; Ye, F.; Liu, C.; Liu, H.; Wang, M.; Li, Y.; Sun, Y.; Li, X.; Tian, M.; Jin, N. In vitro evaluation of the probiotic and functional potential of Lactobacillus strains isolated from fermented food and human intestine. Anaerobe 2014, 30, 1–10. [Google Scholar] [CrossRef]
- Ekmekci, H. Characterization of vaginal lactobacilli coaggregation ability with Escherichia coli. Microbiol. Immunol. 2009, 53, 59–65. [Google Scholar] [CrossRef]
- Adimpong, D.B.; Nielsen, D.S.; Sørensen, K.I.; Derkx, P.M.F.; Jespersen, L. Genotypic characterization and safety assessment of lactic acid bacteria from indigenous African fermented food products. BMC Microbiol. 2012, 12, 75. [Google Scholar] [CrossRef]
- Sirichokchatchawan, W.; Pupa, P.; Praechansri, P.; Amin, N.; Tanasupawat, S.; Sonthayanon, P.; Prapasarakul, N. Autochthonous lactic acid bacteria isolated from pig feces in Thailand show probiotic properties and antibacterial activity against enteric pathogenic bacteria. Microb. Pathog. 2018, 119, 208–215. [Google Scholar] [CrossRef]
- Petsuriyawong, B.; Khunajakr, S. Screening of probiotic lactic acid bacteria from piglet feces. Kasetsart J. Nat. Sci. 2011, 45, 245–253. [Google Scholar]
- Buxton, R. Blood agar plates and hemolysis protocols. American Society for Microbiology , 2005. Available online: https://asm.org https://asm.org/getattachment/7ec0de2b-bb16-4f6e-ba07-2aea25a43e76/protocol-2885.pdf.
- Dowarah, R.; Verma, A.K.; Agarwal, N.; Singh, P.; Singh, B.R. Selection and characterization of probiotic lactic acid bacteria and its impact on growth, nutrient digestibility, and health in weaned piglets. PLoS ONE 2018, 13, e0192978. [Google Scholar] [CrossRef]
- Petri, D.; Hill, J.E.; Van Kessel, A.G. Microbial succession in the gastrointestinal tract of pre-weaned pigs. Livest. Sci. 2010, 133, 107–109. [Google Scholar] [CrossRef]
- Mustakim, M.; Sinawat, S.; Salleh, S.N.; Purwati, E.; Alias, R.; Mohamad, S.A.; Mat Issa, Z. Human milk as a potential source for isolation of probiotic lactic acid bacteria: A mini review. Food Res. 2019, 4, 274–285. [Google Scholar] [CrossRef]
- Czerucka, D.; Piche, T.; Rampal, P. Yeast as probiotics-Saccharomyces boulardii. Aliment. Pharmacol. Ther. 2007, 26, 767–778. [Google Scholar] [CrossRef]
- Elghandour, M.M.Y.; Tan, Z.L.; Abu Hafsa, S.H.; Adegbeye, M.J.; Greiner, R.; Ugbogu, E.A.; Cedillo Monroy, J.; Salem, A.Z.M. Saccharomyces cerevisiae as a probiotic feed additive to non- and pseudo-ruminants: A review. J. Appl. Microbiol. 2020, 128, 658–674. [Google Scholar] [CrossRef]
- Boontiam, W.; Bunchasak, C.; Kim, Y.Y.; Kitipongpysan, S.; Hong, J. Hydrolyzed yeast supplementation to newly weaned piglets: Growth performance, gut health, and microbial fermentation. Animals 2022, 12, 350. [Google Scholar] [CrossRef] [PubMed]
- Canibe, N.; Højberg, O.; Kongsted, H.; Vodolazska, D.; Lauridsen, C.; Nielsen, T.H.; Schönher, A.A. Preventive measures to reduce post-weaning diarrhoea in piglets: A review. Animals 2022, 12, 2585. [Google Scholar] [CrossRef]
- Yeo, S.; Lee, S.; Park, H.; Shin, H.; Holzapfel, W.; Huh, C.S. Development of putative probiotics as feed additives: Validation in a porcine-specific gastrointestinal tract model. Appl. Microbiol. Biotechnol. 2016, 100, 10043–10054. [Google Scholar] [CrossRef] [PubMed]
- Gotcheva, V.; Hristozova, E.; Hristozova, T.; Guo, M.; Roshkova, Z.; Angelov, A. Assessment of potential probiotic properties of lactic acid bacteria and yeast strains. Food Biotechnol. 2002, 16, 211–225. [Google Scholar] [CrossRef]
- Wang, H.; Xu, R.; Zhang, H.; Su, Y.; Zhu, W. Swine gut microbiota and its interaction with host nutrient metabolism. Anim. Nutr. 2020, 6, 410–420. [Google Scholar] [CrossRef]
- Wang, M.; Zhou, X.; Birch Hansen, L.H.; Sheng, Y.; Yu, B.; He, J.; Yu, J.; Zheng, P. Complex probiotics reduce diarrhea by enhancing immunity and balancing gut microbiota in weaned piglets. Front. Immunol. 2025, 16, 1629044. [Google Scholar] [CrossRef]
- Garcia-Cayuela, T.; Korany, A.; Bustos, I.; Cadinanos, L.; Requena, T.; Pelaez, C.; Martinez-Cuesta, M.C. Adhesion abilities of dairy Lactobacillus plantarum strains showing an aggregation phenotype. Food Res. Int. 2014, 57, 44–50. [Google Scholar] [CrossRef]
- Chokesajjawatee, N.; Santiyanont, P.; Chantarasakha, K.; Kocharin, K.; Thammarongtham, C.; Lertampaiporn, S.; Vorapreeda, T.; Srisuk, T.; Wongsurawat, T.; Jenjaroenpun, P.; Nookaew, I.; Visessanguan, W. Safety assessment of a Nham starter culture Lactobacillus plantarum BCC9546 via whole-genome analysis. Sci. Rep. 2020, 10, 10241. [Google Scholar] [CrossRef]
- Anadón, A.; Martínez-Larrañaga, M.R.; Martínez, M.A. Probiotics for animal nutrition in the European Union: Regulation and safety assessment. Regul. Toxicol. Pharmacol. 2006, 45, 91–95. [Google Scholar] [CrossRef]
- Zendeboodi, F.; Khorshidian, N.; Mortazavian, A.M. Probiotic: Conceptualization from a new approach. Curr. Opin. Food Sci. 2020, 32, 103–123. [Google Scholar] [CrossRef]
- Qiao, J.; Li, H.; Wang, Z.; Wang, W. Effects of Lactobacillus acidophilus dietary supplementation on performance, intestinal barrier function, rectal microflora, and serum immune function in weaned piglets challenged with Escherichia coli lipopolysaccharide. Antonie Van Leeuwenhoek 2015, 107, 883–891. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Zhang, M.; Zhao, L.; Ge, K.; Wang, Z.; Jun, L.; Ren, F. Growth performance and post-weaning diarrhea in piglets fed diets supplemented with probiotic complexes. J. Microbiol. Biotechnol. 2018, 28, 1791–1799. [Google Scholar] [CrossRef] [PubMed]
| Symbol | Interpretation | Inhibition Zone Diameter (mm) |
| - | Non-inhibition | <5 |
| + | Weak inhibition | >5 |
| ++ | Intermediate inhibition | >10 |
| +++ | Strong inhibition | >15 |
| ++++ | Very strong inhibition | >20 |
| Isolate ID | log CFU/mL | |
| Initial Count (average) | pH 3.1 | |
| 120 | 6.37±0.5 | 5.35±0.4 |
| 151 | 6.20±0.3 | |
| 183 | 6.15±0.7 | |
| 212 | 5.60±0.5 | |
| 318 | 5.56±0.6 | |
| Isolate ID | 0% (0 h) | 0% (24 h, Control) | 0.3% (24 h) | 0.5% (24 h) |
| 120 | 6.45 ± 0.4 | 9.34 ± 0.6 | 7.35 ± 0.5 | 3.54 ± 0.3 |
| 151 | 6.58 ± 0.3 | 9.02 ± 0.5 | 7.46 ± 0.4 | <3.00 |
| 183 | 6.47 ± 0.5 | 9.12 ± 0.6 | 7.54 ± 0.4 | <3.00 |
| 212 | 6.81 ± 0.4 | 9.66 ± 0.5 | 7.81 ± 0.3 | 6.56 ± 0.4 |
| 318 | 6.00 ± 0.6 | 9.08 ± 0.6 | 7.38 ± 0.5 | <3.00 |
| Isolate ID | OD<sub>600</sub> Before |
OD After (0 min) |
OD After (30 min) |
%Hydrophobicity (mean) |
| 120 | 0.937 ± 0.01 | 1.092 ± 0.02 | 1.014 ± 0.02 | 5.93 ± 0.03 |
| 151 | 0.952 ± 0.02 | 1.146 ± 0.03 | 1.042 ± 0.02 | 9.30 ± 0.04 |
| 183 | 0.937 ± 0.01 | 1.145 ± 0.03 | 1.062 ± 0.03 | 6.46 ± 0.05 |
| 212 | 0.871 ± 0.02 | 1.014 ± 0.02 | 0.924 ± 0.02 | 7.85 ± 0.02 |
| 318 | 0.935 ± 0.02 | 1.171 ± 0.03 | 1.010 ± 0.02 | 12.38 ± 0.03 |
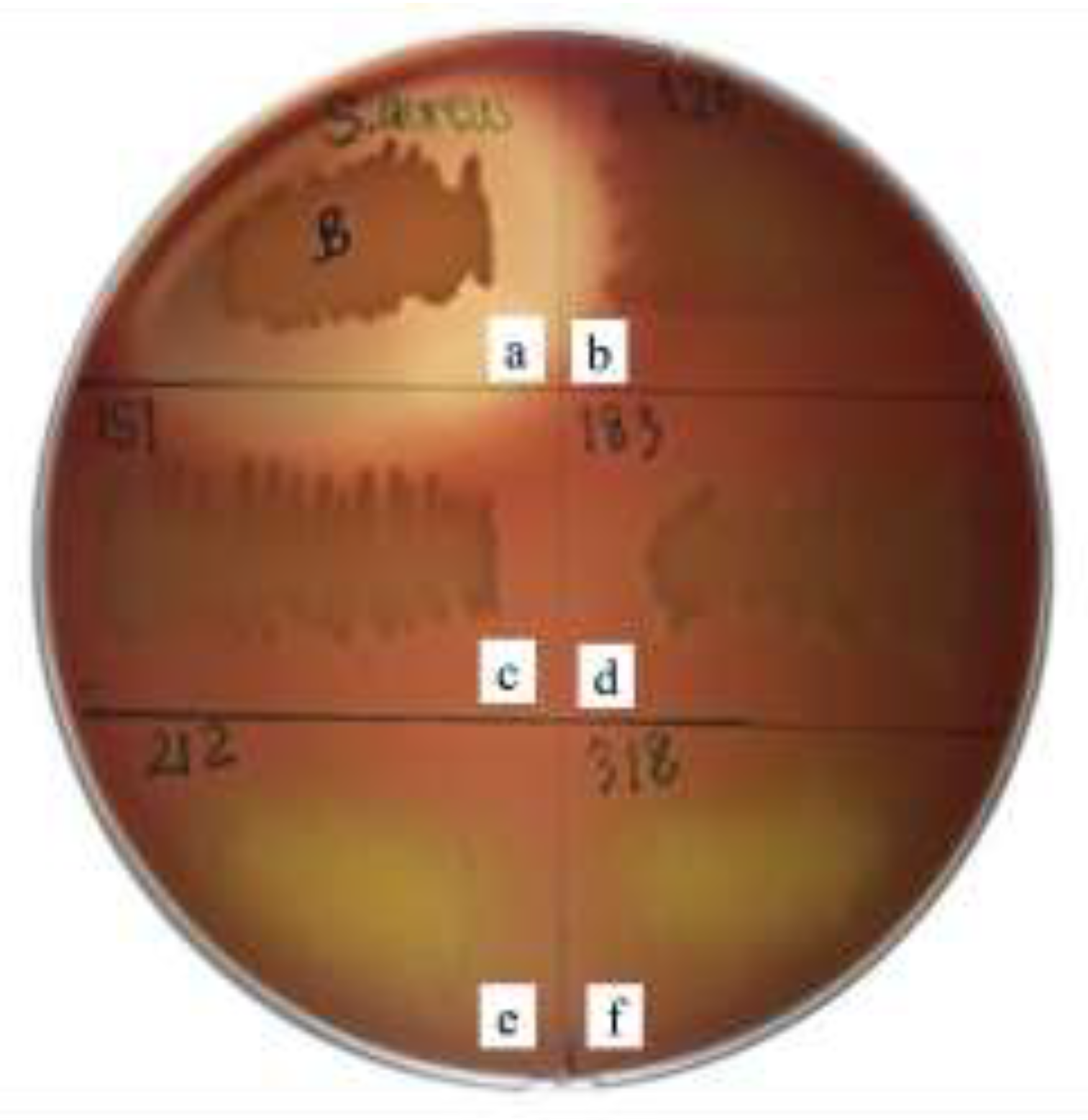
| Isolate ID | Clear zone (mm) - E. coli | Clear zone (mm) - S. aureus |
| PMvet120 | Incomplete inhibition (no clear zone) | 0.0 ± 0.0 |
| PMvet151 | Incomplete inhibition (no clear zone) | 0.0 ± 0.0 |
| PMvet183 | Incomplete inhibition (no clear zone) | 0.0 ± 0.0 |
| PMvet212 | 12.50 ± 0.4 | 11.75 ± 0.5 |
| PMvet318 | 11.50 ± 0.3 | 10.50 ± 0.4 |
| Control (2% lactic acid) | 18.50 ± 0.5 | 15.25 ± 0.4 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




