Submitted:
16 January 2026
Posted:
19 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. SULBA Framework
| Algorithm 1. Stepwise Upper and Lower Boundaries Augmentation (SULBA) |
| Input: |
2.2. Scaling of Generated Samples
2.3. SULBA Perfect Reversibility
2.4. Datasets and Preprocessing
2.5. Network Architectures
2.6. Training and Implementation Details
2.7. Evaluation Protocol and Statistical Analysis
3. Results
3.1. Benchmark Performance on 2D Medical Image Classification
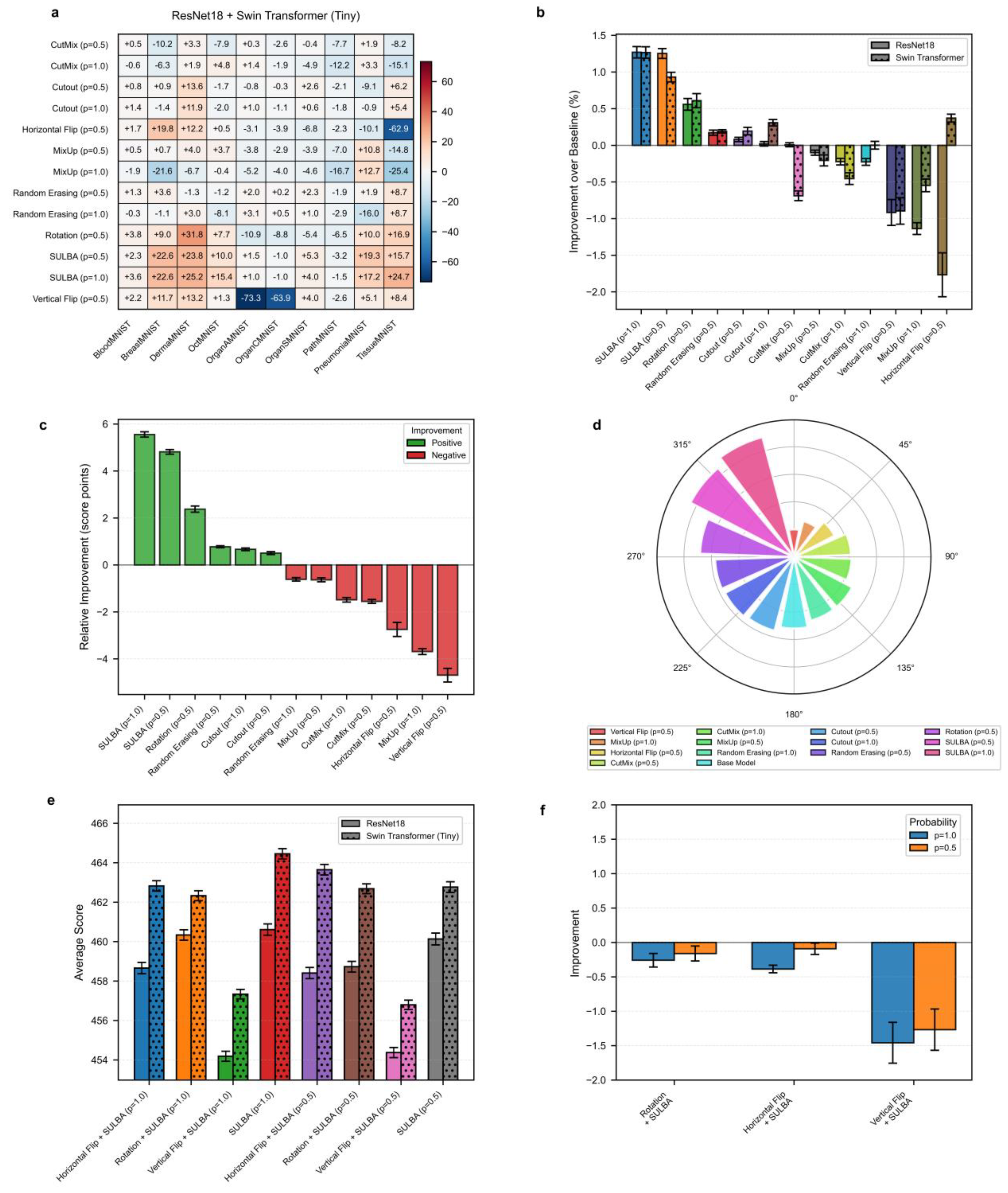
3.1.1. SULBA Provides Robust Performance Gains Across Diverse Datasets and Model Architectures
3.1.2. SULBA Demonstrates Superior and Consistent Performance Improvements
3.1.3. The Integration of SULBA with Traditional Augmentations Does Not Confer Synergistic Benefits
3.2. Benchmark Performance on 3D Medical Image Classification
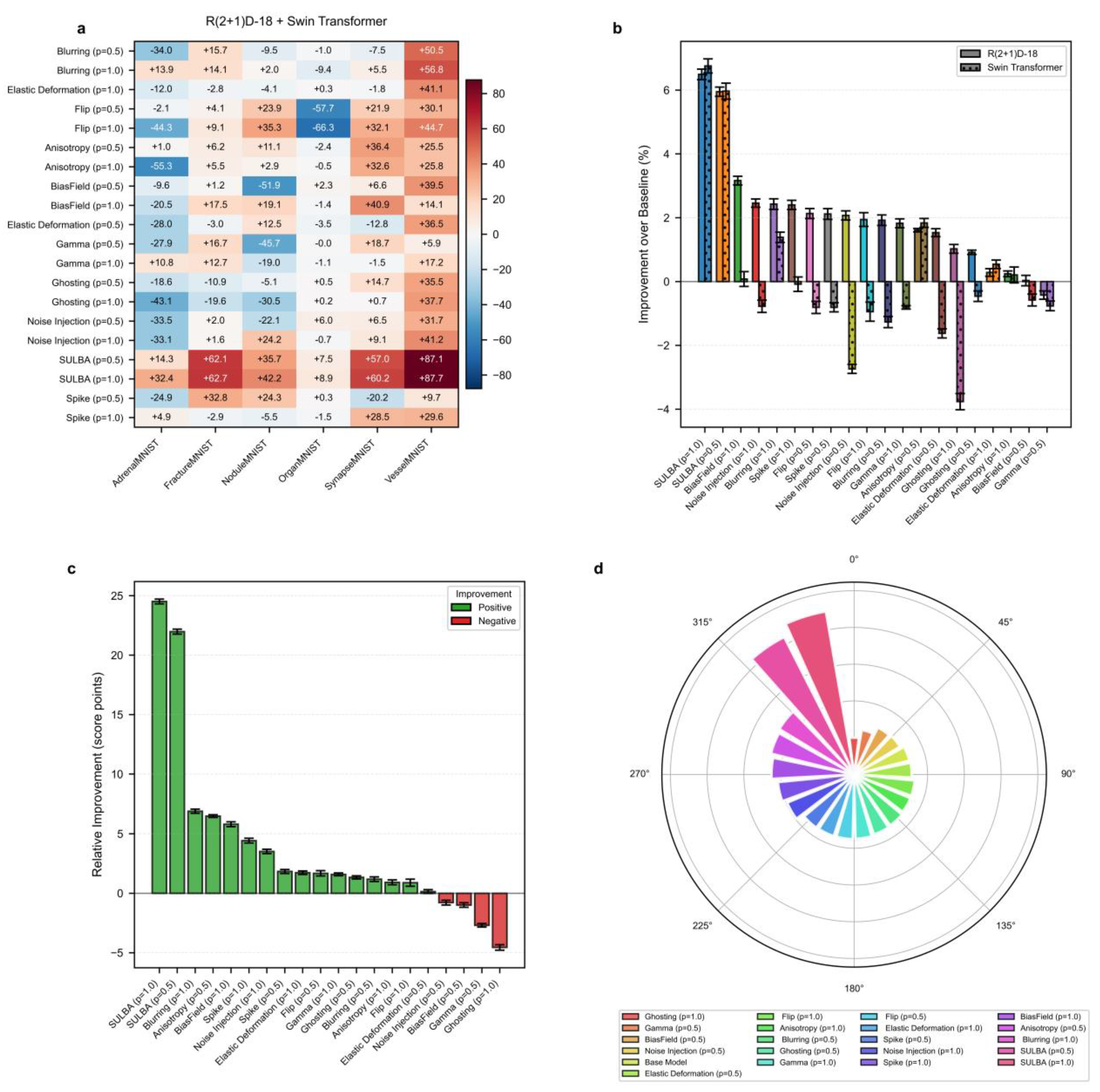
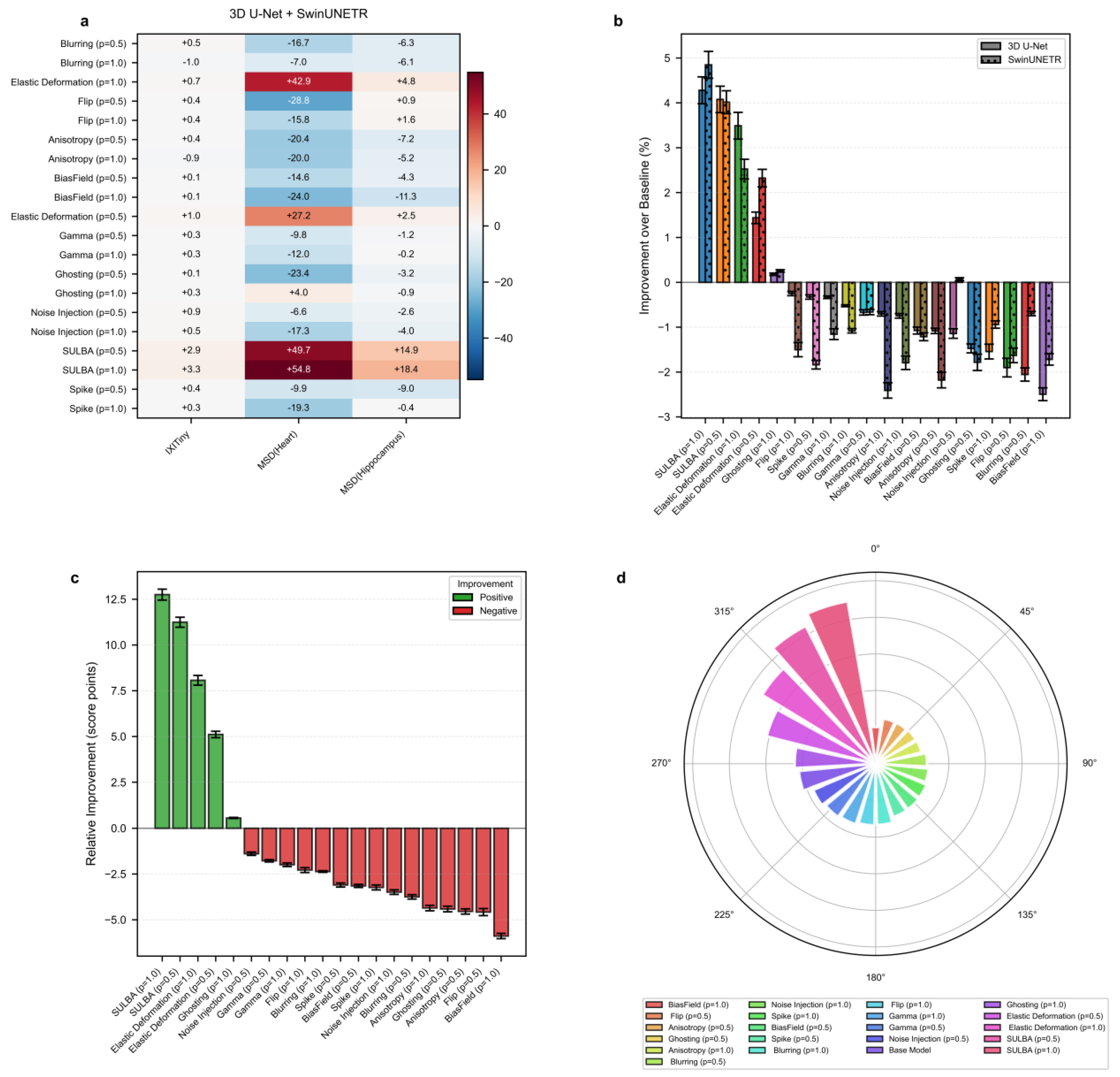
3.2.1. SULBA Delivers Consistent and Exceptionally Large Improvements Across All 3D Datasets
3.2.2. Traditional 3D Augmentations Show High Dataset-Specific Variance and Inconsistent Effects
3.2.3. SULBA Substantially Outperforms Standard Volumetric Augmentation Techniques in 3D Classification
3.3. Benchmark Performance on 2D Medical Image Segmentation
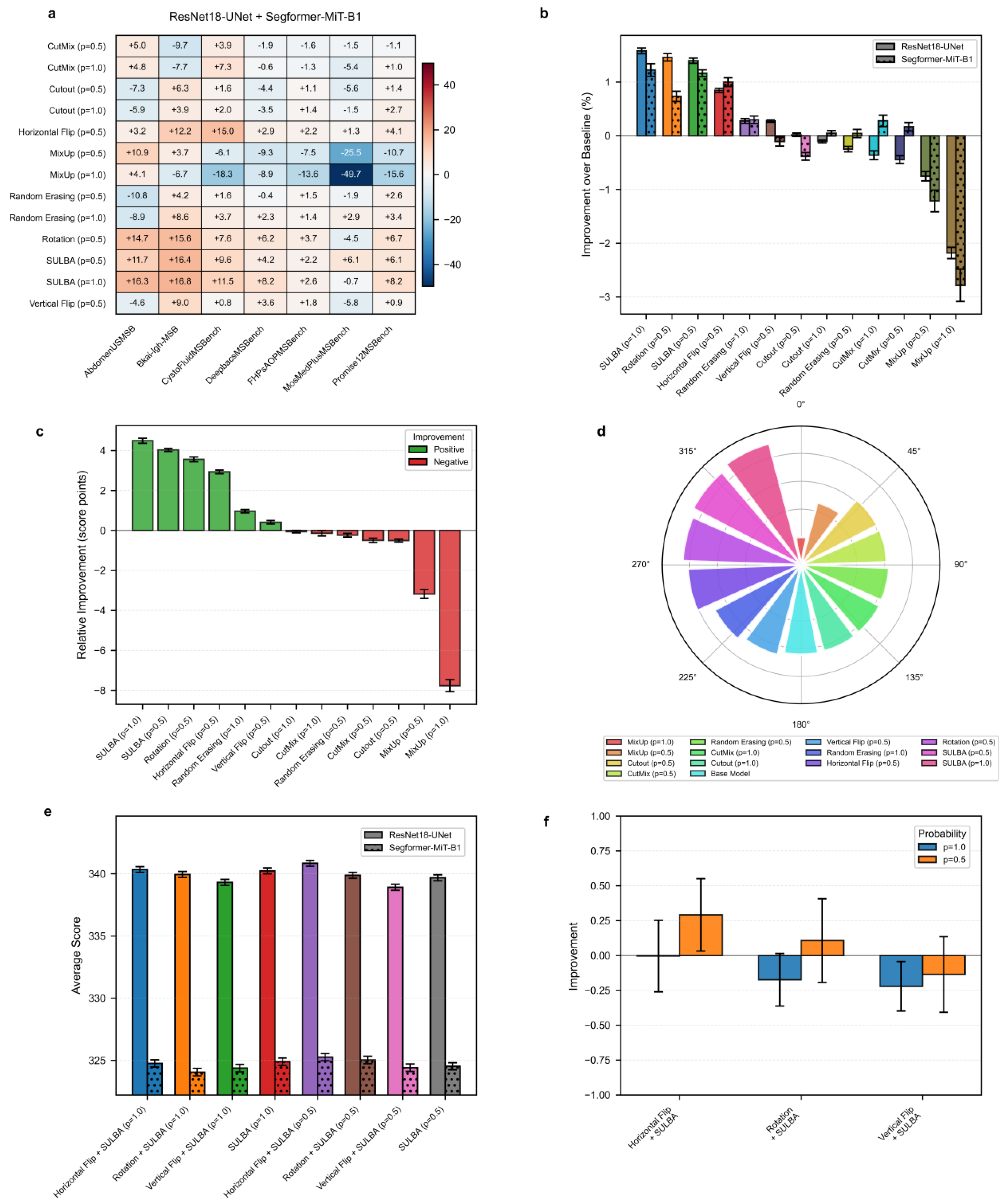
3.3.1. SULBA Provides Robust, Positive Improvements Across Diverse Segmentation Datasets
3.3.2. SULBA Ranks as the Top-Performing Augmentation Strategy for 2D Segmentation
3.3.3. Combining SULBA with Spatial Augmentations Provides Marginal and Inconsistent Benefits
3.4. Benchmark Performance on 3D Medical Image Segmentation
3.4.1. SULBA Delivers Consistent Improvements Across 3D Segmentation Datasets
3.4.2. Conventional 3D Augmentation Methods Exhibit Pronounced Dataset-Dependent Variability
3.4.3. SULBA Achieves the Highest Overall Ranking Among 3D Augmentation Strategies
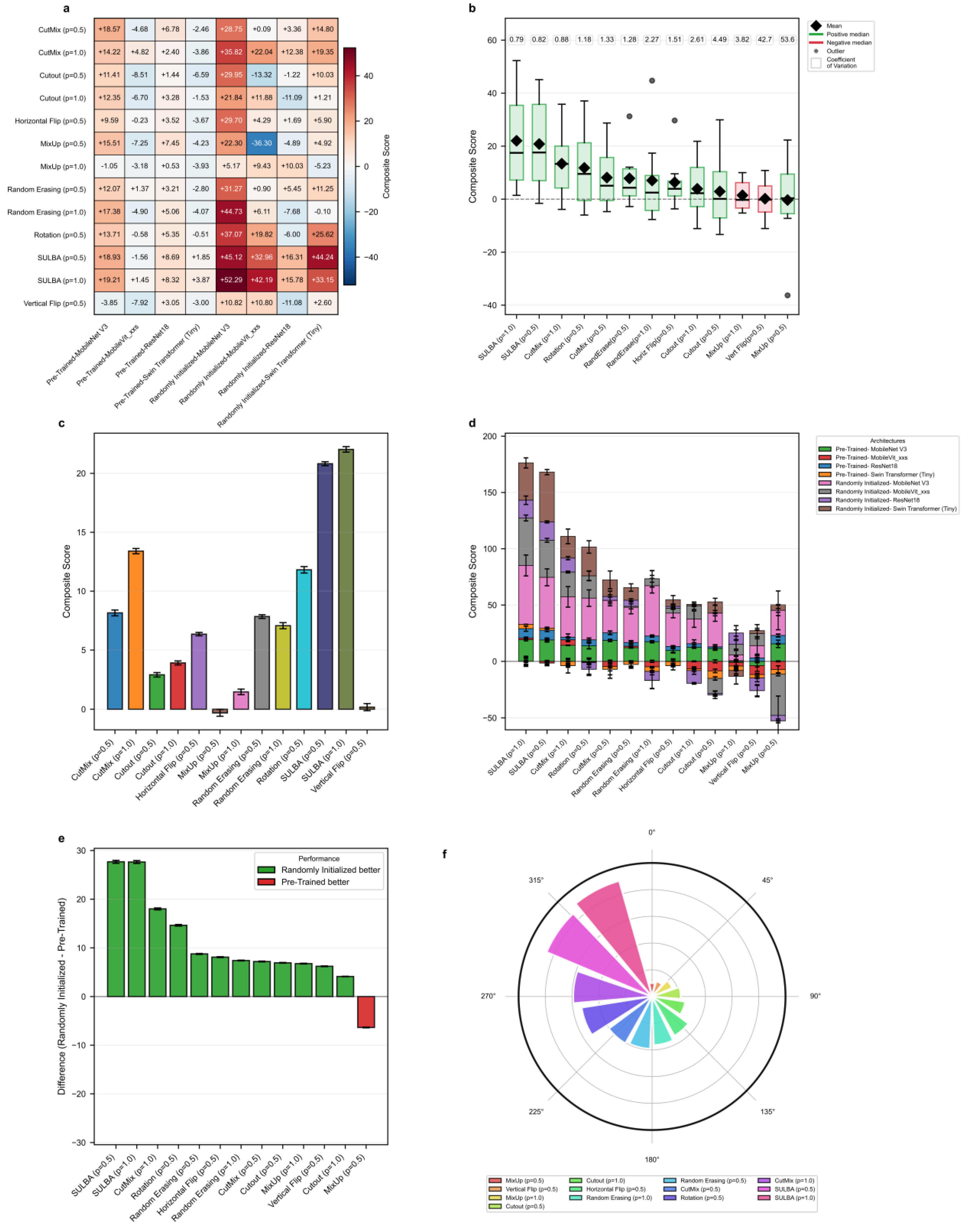
3.5. Generalization Performance Across Diverse Architectures
3.5.1. SULBA Delivers Superior Cross-Dataset Generalization
3.5.2. SULBA Provides Consistent Improvements Across Architectures
3.5.3. Training with Randomly Initialized Weights Amplifies SULBA’s Benefits
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Code Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| n | Number |
| C | Cumulative |
| M | Metric |
| 2D | Two Dimension |
| 3D | Three Dimension |
| DA | Data Augmentation |
| AI | Artificial Intelligent |
| MSD | Medical Segmentation Decalthlon |
| SULBA | Stepwise Upper and Lower Boundaries Augmentation |
| AUROC | Area Under the Receiver Operating Characteristic Curve |
References
- Tian, F.; Liu, D.; Wei, N.; Fu, Q.; Sun, L.; Liu, W.; Sui, X.; Tian, K.; Nemeth, G.; Feng, J.; Xu, J. Prediction of tumor origin in cancers of unknown primary origin with cytology-based deep learning. Nature Medicine 2024, 30, 1309–19. [Google Scholar] [CrossRef]
- Kumar, R.; Kumbharkar, P.; Vanam, S.; Sharma, S. Medical images classification using deep learning: a survey. Multimedia Tools and Applications 2024, 83, 19683–728. [Google Scholar] [CrossRef]
- Ma, J.; He, Y.; Li, F.; Han, L.; You, C.; Wang, B. Segment anything in medical images. Nature Communications 2024, 15, 654. [Google Scholar] [CrossRef] [PubMed]
- Kshatri, S.S.; Singh, D. Convolutional neural network in medical image analysis: a review. Archives of Computational Methods in Engineering 2023, 30, 2793–810. [Google Scholar] [CrossRef]
- Takahashi, S.; Sakaguchi, Y.; Kouno, N.; Takasawa, K.; Ishizu, K.; Akagi, Y.; Aoyama, R.; Teraya, N.; Bolatkan, A.; Shinkai, N.; Machino, H. Comparison of vision transformers and convolutional neural networks in medical image analysis: A systematic review. Journal of Medical Systems 2024, 48, 84. [Google Scholar] [CrossRef]
- Khan, R.F.; Lee, B.D.; Lee, M.S. Transformers in medical image segmentation: a narrative review. Quantitative Imaging in Medicine and Surgery 2023, 13, 8747. [Google Scholar] [CrossRef]
- Tudosiu, P.D.; Pinaya, W.H.; Ferreira Da Costa, P.; Dafflon, J.; Patel, A.; Borges, P.; Fernandez, V.; Graham, M.S.; Gray, R.J.; Nachev, P.; Ourselin, S. Realistic morphology-preserving generative modelling of the brain. Nature Machine Intelligence 2024, 6, 811–9. [Google Scholar] [CrossRef]
- Dhar, T.; Dey, N.; Borra, S.; Sherratt, R.S. Challenges of deep learning in medical image analysis—improving explainability and trust. IEEE Transactions on Technology and Society 2023, 4, 68–75. [Google Scholar] [CrossRef]
- Price, W.N.; Cohen, I.G. Privacy in the age of medical big data. Nature medicine 2019, 25, 37–43. [Google Scholar] [CrossRef]
- Xu, C.; Coen-Pirani, P.; Jiang, X. Empirical study of overfitting in deep learning for predicting breast cancer metastasis. Cancers 2023, 15, 1969. [Google Scholar] [CrossRef]
- Azizi, S.; Culp, L.; Freyberg, J.; Mustafa, B.; Baur, S.; Kornblith, S.; Chen, T.; Tomasev, N.; Mitrović, J.; Strachan, P.; Mahdavi, S.S. Robust and data-efficient generalization of self-supervised machine learning for diagnostic imaging. Nature Biomedical Engineering 2023, 7, 756–79. [Google Scholar] [CrossRef]
- Goceri, E. Medical image data augmentation: techniques, comparisons and interpretations. Artificial intelligence review 2023, 56, 12561–605. [Google Scholar] [CrossRef] [PubMed]
- Kebaili, A.; Lapuyade-Lahorgue, J.; Ruan, S. Deep learning approaches for data augmentation in medical imaging: a review. Journal of imaging 2023, 9, 81. [Google Scholar] [CrossRef] [PubMed]
- Makhlouf, A.; Maayah, M.; Abughanam, N.; Catal, C. The use of generative adversarial networks in medical image augmentation. Neural Computing and Applications 2023, 35, 24055–68. [Google Scholar] [CrossRef]
- Wang, J.; Wang, K.; Yu, Y.; Lu, Y.; Xiao, W.; Sun, Z.; Liu, F.; Zou, Z.; Gao, Y.; Yang, L.; Zhou, H.Y. Self-improving generative foundation model for synthetic medical image generation and clinical applications. Nature Medicine 2025, 31, 609–17. [Google Scholar] [CrossRef]
- Fujii, Y.; Uchida, D.; Sato, R.; Obata, T.; Akihiro, M.; Miyamoto, K.; Morimoto, K.; Terasawa, H.; Yamazaki, T.; Matsumoto, K.; Horiguchi, S. Effectiveness of data-augmentation on deep learning in evaluating rapid on-site cytopathology at endoscopic ultrasound-guided fine needle aspiration. Scientific reports 2024, 14, 22441. [Google Scholar] [CrossRef]
- Abe, A.A.; Nyathi, M. Lung Cancer Diagnosis From Computed Tomography Images Using Deep Learning Algorithms With Random Pixel Swap Data Augmentation: Algorithm Development and Validation Study. JMIR Bioinformatics and Biotechnology 2025, 6, e68848. [Google Scholar] [CrossRef]
- Saad, M.M.; O’Reilly, R.; Rehmani, M.H. A survey on training challenges in generative adversarial networks for biomedical image analysis. Artificial Intelligence Review 2024, 57, 19. [Google Scholar] [CrossRef]
- Islam, S.; Aziz, M.T.; Nabil, H.R.; Jim, J.R.; Mridha, M.F.; Kabir, M.M.; Asai, N.; Shin, J. Generative adversarial networks (GANs) in medical imaging: Advancements, applications, and challenges. IEEE Access. 2024, 12, 35728–53. [Google Scholar] [CrossRef]
- Rao, A.; Lee, J.Y.; Aalami, O. Studying the impact of augmentations on medical confidence calibration. In InProceedings of the IEEE/CVF International Conference on Computer Vision, 2023; pp. 2462–2472. [Google Scholar]
- Sun, D.; Dornaika, F. Data augmentation for deep visual recognition using superpixel based pairwise image fusion. Information Fusion. 2024, 107, 102308. [Google Scholar] [CrossRef]
- Pineau, J.; Vincent-Lamarre, P.; Sinha, K.; Larivière, V.; Beygelzimer, A.; d'Alché-Buc, F.; Fox, E.; Larochelle, H. Improving reproducibility in machine learning research (a report from the neurips 2019 reproducibility program). Journal of machine learning research 2021, 22, 1–20. [Google Scholar]
- Paszke, A.; Gross, S.; Massa, F.; Lerer, A.; Bradbury, J.; Chanan, G.; Killeen, T.; Lin, Z.; Gimelshein, N.; Antiga, L.; Desmaison, A. Pytorch: An imperative style, high-performance deep learning library. Advances in neural information processing systems 2019, 32. [Google Scholar]
- Pérez-García, F.; Sparks, R.; Ourselin, S. TorchIO: a Python library for efficient loading, preprocessing, augmentation and patch-based sampling of medical images in deep learning. Computer methods and programs in biomedicine 2021, 208, 106236. [Google Scholar] [CrossRef] [PubMed]
- Abadi, M.; Barham, P.; Chen, J.; Chen, Z.; Davis, A.; Dean, J.; Devin, M.; Ghemawat, S.; Irving, G.; Isard, M.; Kudlur, M. {TensorFlow}: a system for {Large-Scale} machine learning. In12th USENIX symposium on operating systems design and implementation (OSDI 16), 2016; pp. 265–283. [Google Scholar]
- Shorten, C.; Khoshgoftaar, T.M. A survey on image data augmentation for deep learning. Journal of big data 2019, 6, 1–48. [Google Scholar] [CrossRef]
- Zoph, B.; Cubuk, E.D.; Ghiasi, G.; Lin, T.Y.; Shlens, J.; Le, Q.V. Learning data augmentation strategies for object detection. In InEuropean conference on computer vision; Springer International Publishing: Cham, 23 Aug 2020; pp. 566–583. [Google Scholar]
- Cohen, T.; Welling, M. Group equivariant convolutional networks. InInternational conference on machine learning, 2016 Jun 11; PMLR; pp. 2990–2999. [Google Scholar]
- Zhang, Y.; Hare, J.; Prugel-Bennett, A. Deep set prediction networks. In Advances in Neural Information Processing Systems; 2019; p. 32. [Google Scholar]
- Gerken, J.E.; Aronsson, J.; Carlsson, O.; Linander, H.; Ohlsson, F.; Petersson, C.; Persson, D. Geometric deep learning and equivariant neural networks. Artificial Intelligence Review 2023, 56, 14605–62. [Google Scholar] [CrossRef]
- Diaz-Peregrino, R.; Robles, F.T.; Gonzalez, G.; Palma, R.; Escalante-Ramirez, B.; Olveres, J.; Reyes-Gonzalez, J.P.; Gomez-Coeto, J.A.; Rodriguez-Herrera, C.A. Enhancing generalization in whole-body MRI-based deep learning models: A novel data augmentation pipeline for cross-platform adaptation. Intelligence-Based Medicine 2025, 100277. [Google Scholar] [CrossRef]
- Yang, J.; Shi, R.; Wei, D.; Liu, Z.; Zhao, L.; Ke, B.; Pfister, H.; Ni, B. Medmnist v2-a large-scale lightweight benchmark for 2d and 3d biomedical image classification. Scientific Data 2023, 10, 41. [Google Scholar] [CrossRef]
- Kuş, Z.; Aydin, M. MedSegBench: A comprehensive benchmark for medical image segmentation in diverse data modalities. Scientific Data 2024, 11, 1283. [Google Scholar] [CrossRef]
- Antonelli, M.; Reinke, A.; Bakas, S.; Farahani, K.; Kopp-Schneider, A.; Landman, B.A.; Litjens, G.; Menze, B.; Ronneberger, O.; Summers, R.M.; Van Ginneken, B. The medical segmentation decathlon. Nature communications 2022, 13, 4128. [Google Scholar] [CrossRef]
- Deng, J.; Dong, W.; Socher, R.; Li, L.J.; Li, K.; Fei-Fei, L. Imagenet: A large-scale hierarchical image database. In2009 IEEE conference on computer vision and pattern recognition, 2009 Jun 20; Ieee; pp. 248–255. [Google Scholar]
- Kermany, D.S.; Goldbaum, M.; Cai, W.; Valentim, C.C.; Liang, H.; Baxter, S.L.; McKeown, A.; Yang, G.; Wu, X.; Yan, F.; Dong, J. Identifying medical diagnoses and treatable diseases by image-based deep learning. cell. 2018, 172, 1122–31. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In InProceedings of the IEEE conference on computer vision and pattern recognition, 2016; pp. 770–778. [Google Scholar]
- Liu, Z.; Lin, Y.; Cao, Y.; Hu, H.; Wei, Y.; Zhang, Z.; Lin, S.; Guo, B. Swin transformer: Hierarchical vision transformer using shifted windows. In InProceedings of the IEEE/CVF international conference on computer vision, 2021; pp. 10012–10022. [Google Scholar]
- Tran, D.; Wang, H.; Torresani, L.; Ray, J.; LeCun, Y.; Paluri, M. A closer look at spatiotemporal convolutions for action recognition. In InProceedings of the IEEE conference on Computer Vision and Pattern Recognition, 2018; pp. 6450–6459. [Google Scholar]
- Liu, Z.; Ning, J.; Cao, Y.; Wei, Y.; Zhang, Z.; Lin, S.; Hu, H. Video swin transformer. In InProceedings of the IEEE/CVF conference on computer vision and pattern recognition, 2022; pp. 3202–3211. [Google Scholar]
- Kay, W.; Carreira, J.; Simonyan, K.; Zhang, B.; Hillier, C.; Vijayanarasimhan, S.; Viola, F.; Green, T.; Back, T.; Natsev, P.; Suleyman, M. The kinetics human action video dataset. arXiv 2017, arXiv:1705.06950. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In InInternational Conference on Medical image computing and computer-assisted intervention; Springer international publishing: Cham, 2015; pp. 234–241. [Google Scholar]
- Xie, E.; Wang, W.; Yu, Z.; Anandkumar, A.; Alvarez, J.M.; Luo, P. SegFormer: Simple and efficient design for semantic segmentation with transformers. Advances in neural information processing systems 2021, 34, 12077–90. [Google Scholar]
- Çiçek, Ö.; Abdulkadir, A.; Lienkamp, S.S.; Brox, T.; Ronneberger, O. 3D U-Net: learning dense volumetric segmentation from sparse annotation. In InInternational conference on medical image computing and computer-assisted intervention; Springer International Publishing: Cham, 2 Oct 2016; pp. 424–432. [Google Scholar]
- Hatamizadeh, A.; Nath, V.; Tang, Y.; Yang, D.; Roth, H.R.; Xu, D. Swin unetr: Swin transformers for semantic segmentation of brain tumors in mri images. In InInternational MICCAI brainlesion workshop; Springer International Publishing: Cham, 27 Sep 2021; pp. 272–284. [Google Scholar]
- Cardoso, M.J.; Li, W.; Brown, R.; Ma, N.; Kerfoot, E.; Wang, Y.; Murrey, B.; Myronenko, A.; Zhao, C.; Yang, D.; Nath, V. Monai: An open-source framework for deep learning in healthcare. arXiv 2022, arXiv:2211.02701. [Google Scholar] [CrossRef]
- Howard, A.; Sandler, M.; Chu, G.; Chen, L.C.; Chen, B.; Tan, M.; Wang, W.; Zhu, Y.; Pang, R.; Vasudevan, V.; Le, Q.V. Searching for mobilenetv3. In InProceedings of the IEEE/CVF international conference on computer vision, 2019; pp. 1314–1324. [Google Scholar]
- Mehta, S.; Rastegari, M. Mobilevit: light-weight, general-purpose, and mobile-friendly vision transformer. arXiv 2021, arXiv:2110.02178. [Google Scholar]
- Zhong, Z.; Zheng, L.; Kang, G.; Li, S.; Yang, Y. Random erasing data augmentation. InProceedings of the AAAI conference on artificial intelligence 2020, Vol. 34(No. 07), 13001–13008. [Google Scholar] [CrossRef]
- DeVries, T.; Taylor, G.W. Improved regularization of convolutional neural networks with cutout. arXiv 2017, arXiv:1708.04552. [Google Scholar] [CrossRef]
- Yun, S.; Han, D.; Oh, S.J.; Chun, S.; Choe, J.; Yoo, Y. Cutmix: Regularization strategy to train strong classifiers with localizable features. In InProceedings of the IEEE/CVF international conference on computer vision, 2019; pp. 6023–6032. [Google Scholar]
- Zhang, H.; Cisse, M.; Dauphin, Y.N.; Lopez-Paz, D. mixup: Beyond empirical risk minimization. arXiv 2017, arXiv:1710.09412. [Google Scholar]
- Billot, B.; Robinson, E.; Dalca, A.V.; Iglesias, J.E. Partial volume segmentation of brain MRI scans of any resolution and contrast. In InInternational Conference on Medical image computing and computer-assisted intervention; Springer International Publishing: Cham, 29 Sep 2020; pp. 177–187. [Google Scholar]
- Sudre, CH; Cardoso, MJ; Ourselin, S; Alzheimer’s Disease Neuroimaging Initiative. Longitudinal segmentation of age-related white matter hyperintensities. Medical Image Analysis 2017, 38, 50–64. [Google Scholar] [CrossRef]
- Shackleford, J.; Kandasamy, N.; Sharp, G. High performance deformable image registration algorithms for manycore processors; Newnes, 2013; pp. pp1–12. [Google Scholar]
- Goceri, E. Medical image data augmentation: techniques, comparisons and interpretations. Artificial intelligence review 2023, 56, 12561–605. [Google Scholar] [CrossRef]
- Zhao, M.; Wei, Y.; Lu, Y.; Wong, K.K. A novel U-Net approach to segment the cardiac chamber in magnetic resonance images with ghost artifacts. Computer Methods and Programs in Biomedicine 2020, 196, 105623. [Google Scholar] [CrossRef]
- Kumar, T.; Brennan, R.; Mileo, A.; Bendechache, M. Image data augmentation approaches: A comprehensive survey and future directions. IEEE Access, 2024 Sep 30. [Google Scholar]
- Cubuk, E.D.; Zoph, B.; Mane, D.; Vasudevan, V.; Le, Q.V. Autoaugment: Learning augmentation strategies from data. In InProceedings of the IEEE/CVF conference on computer vision and pattern recognition, 2019; pp. 113–123. [Google Scholar]
- Theodoris, C.V.; Xiao, L.; Chopra, A.; Chaffin, M.D.; Al Sayed, Z.R.; Hill, M.C.; Mantineo, H.; Brydon, E.M.; Zeng, Z.; Liu, X.S.; Ellinor, P.T. Transfer learning enables predictions in network biology. Nature 2023, 618, 616–24. [Google Scholar] [CrossRef]
- Sabha, S.U.; Assad, A.; Din, N.M.; Bhat, M.R. From scratch or pretrained? An in-depth analysis of deep learning approaches with limited data. International Journal of System Assurance Engineering and Management 2024, 1–0. [Google Scholar] [CrossRef]
- Jiménez-Sánchez, A.; Avlona, N.R.; de Boer, S.; Campello, V.M.; Feragen, A.; Ferrante, E.; Ganz, M.; Gichoya, J.W.; Gonzalez, C.; Groefsema, S.; Hering, A. In the picture: Medical imaging datasets, artifacts, and their living review. In InProceedings of the 2025 ACM Conference on Fairness, Accountability, and Transparency, 2025; pp. 511–531. [Google Scholar]
- Dulaney, A.; Virostko, J. Disparities in the demographic composition of The Cancer Imaging Archive. Radiology: Imaging Cancer 2024, 6, e230100. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



